Summary
Much of the infection cycle of Mycobacterium tuberculosis (Mtb) is spent within its host cell, the macrophage. As a consequence of the chronic, enduring nature of the infection, this cell/cell interaction has become highly intimate, and the bacterium has evolved to detect, react to, and manipulate the evolving, immune-modulated phenotype of its host. In this review, we discuss the nature of the endosomal/lysosomal continuum, the characterization of the bacterium’s transcriptional responses during the infection cycle, and the dominant environmental cues that shape this response. We also discuss how the metabolism of both cells is modulated by the infection and the impact that this has on the progression of the granuloma. Finally, we detail how these transcriptional responses can be exploited to construct reporter bacterial strains to probe the temporal and spatial environmental shifts experienced by Mtb during the course of experimental infections. These reporter strains provide new insights into the fitness of Mtb under immune- and drug-mediated pressure.
Keywords: Mycobacterium tuberculosis, macrophage, phagosome, tuberculosis
Introduction
Mycobacterium tuberculosis (Mtb) represents one of the most successful infectious agents specific to humans. While it was thought previously to have evolved from the animal pathogen Mycobacterium bovis (1), it is now known to have accompanied mankind as we emerged from Africa approximately 50,000 years ago (2, 3). The success of Mtb as a pathogen is likely due in large part to its ability to infect many yet cause active disease in only a few, which, in the limited pockets of hunter-gatherer human populations present for much of human evolution, would ensure its continued maintenance in its chosen host.
Within the body, Mtb resides inside macrophages for much of its infection cycle (4), although the bacterium is also found in neutrophils, and extracellularly, in the caseous center of necrotic granulomas. The macrophage is regarded as a key player in both the innate and acquired arms of the immune response (5). A resting macrophage performs tasks critical to tissue homeostasis (6). The cell is highly degradative and clears apoptotic debris in a non-inflammatory manner. Activation of this cell through either pathogen-derived Toll-like receptor (TLR) ligands or cytokines such as interferon-γ (IFN-γ) leads to enhanced killing capacity through reactive oxygen and nitrogen intermediates and a reduction in proteolytic activity coupled with increased antigen presentation capabilities (6, 7). The innate defense role of phagocytes such as macrophages and neutrophils in the body means that they are among the most frequent cells that will encounter microbial invaders. While neutrophils are highly aggressive, short-lived cells, macrophages generate a more regulated antimicrobial response and are usually not driven to programmed cell death by a microbial encounter.
To appreciate how Mtb survives within these potentially hostile cells, it is important to have an understanding of the physiology of phagosome maturation. Any particle, whether it be inert or live, that enters a macrophage through ligation of phagocytic receptors triggers a cascade of events that can activate specific anti-microbial responses ranging from the assembly of the NADPH oxidase complex in the phagosomal membrane, to the regulated fusion with intracellular vesicles and remodeling of the phagosome with subsequent fusion to lysosomes. Most cellular microbiological studies have utilized immunofluorescent localization of known lysosomal markers such as rab7 or cathepsin D as a means of assessing the ‘lysosomal nature’ of an intracellular compartment, but these analyses are for the most part subjective and do not directly inform the investigator of the functionality of the marker or the compartment. To overcome this and develop a more in depth understanding of the kinetics of physiological changes occurring in the phagosome post-internalization, we developed a broad range of real-time, fluorescence-based readouts of changes that occur in the lumen of the phagosome (5) (Fig. 1).
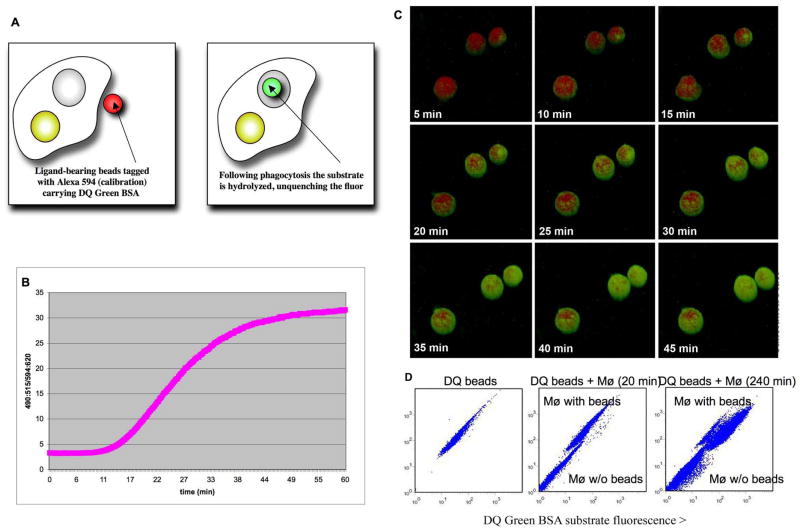
Fig. 1. Real-time measurement of changing phagosomal hydrolase activity in the phagosome during the maturation process.
Assays have been developed that measure a range of lysosomal hydrolase activities including bulk proteinase activity, cysteine proteinase activity, lipolysis and β-galactosidase activity. The assays involve fluorogenic substrates that are linked to silica beads (A). These beads are also coupled with an opsonizing molecule, such as IgG or mannosylated BSA, to facilitate their uptake by macrophages, and a calibration fluorophore. Results are usually expressed as a ratio of substrate fluorescence:calibration fluorescence. The examples illustrated show experiments with beads coupled with the proteinase substrate DQ Green BSA, which consists of albumin derivatized with a self-quenching fluorophore. The ensuing change in fluorescence generated through the release of fluorescent peptides can be measured by several different platforms, each of which provides its own unique insight. A spectrofluorometer (B) provides a kinetic readout that represents an average value across a population of cells mounted on glass coverslips in cuvettes. A confocal microscope (C) allows visualization and quantification of the increase in green fluorescence from the hydrolyzed substrate compared with the red fluorescence of the calibration fluorophore at the level of the individual phagosome. These frames are from a 45-minute movie. Flow cytometry (C) enables examination of the heterogeneity of the activity across the cell population at the level of each individual cell. Analysis by flow cytometry can be combined with immunofluorescence with antibodies against pathogens, or cell surface markers. This figure was modified from Russell et al. (5).
Phagosome maturation
The rate of acidification of a phagosome has long been utilized as an indicator of phagosomal maturation. pH-sensitive dyes such as carboxyfluorescein allow one to measure this parameter in real-time, and demonstrates that it takes between 12–15 minutes for a phagosome containing inert, IgG-opsonized particles to reach a pH equilibrium at around pH 4.8 in murine bone marrow-derived macrophages (8). The same experiment conducted on phagosomes containing Mtb reveals that the bacterium blocks the acidification of its vacuole, which shows markedly reduced acquisition of the proton-ATPase required for the acidification process (9, 10). The Mtb vacuole equilibrates to a pH of 6.4, still acidic but considerably less hostile than the pH of the lysosome.
Maturing phagosomes acquire lysosomal hydrolases through both fusion with pre-existing lysosomes and with trafficking vesicles that shuttle the enzymes, usually in an inactive pro-enzyme form, from the trans Golgi network (TGN). The activity of these enzymes can be assayed directly through the use of a range of fluorogenic substrates that provide a quantitative readout of enzymatic activity, which is a combination of enzyme concentration, the activation state of the enzyme, and the pH of the phago/lysosome (8, 11–13) (Fig. 1). The majority of lysosomal hydrolases, not surprisingly, have an acidic pH optimum. Most of these intraphagosomal enzyme assays are processive and only stop when the substrate is exhausted. In contrast, another assay developed to quantify phagosome maturation measures phagosome/lysosome fusion through the use of fluorescence resonance energy transfer (FRET) (8). The assay utilizes a donor fluorophore on the phagocytosed particle that, when excited, transfers its energy to an acceptor fluorophore that was pulse-chased into the macrophage lysosomes prior to initiation of the experiment. The assay measures the concentration of fluid-phase lysosomal cargo delivered to the phagosome as it matures. Interestingly, this assay reaches an equilibrium or steady state, which takes around 90 min in murine macrophages. The differential kinetics of the two steady state readouts, pH and phagosome/lysosome fusion, raise an interesting point regarding the mature phagosome. Terminal pH, pH 4.5–5.0, is reached within 12–15 min, yet maximal concentration of lysosomal constituents requires 90 min to equilibrate. Thus, the degree of maturation of a phagosome can be a somewhat subjective evaluation unless the relevant parameters are clearly defined. Immunofluorescent co-localization with markers such as LAMP1 is a singularly inadequate means of experimentally determining the lysosomal nature of a phagosomal compartment.
As mentioned previously, the vacuole in which Mtb resides exhibits partial acidification to pH 6.4. Further analysis of this vacuole demonstrated that it did acquire detectable amounts of cathepsin D but that the enzyme was predominantly in its pro-enzyme form and therefore had not come from fusion with lysosomes but presumably from vesicles derived from the TGN (14, 15). In addition, analysis of transferrin trafficking, as an indicator of the early recycling endosomal network, revealed that the Mtb-containing vacuole was readily accessible to transferrin, which passed through this vacuole as a pulse (14). Finally, the glycosphingolipids on the surface of the macrophage, such as the ganglioside GM1, also cycled through the Mtb-containing vacuole with considerable efficiency (16). All these data imply that the Mtb-containing vacuole is not an inert, sequestered compartment, but functions like a fully integrated compartment of the rapid, recycling endosomal network. This indicates that Mtb has arrested the maturation of its vacuole to prevent full acidification or acquisition of active lysosome hydrolases, yet it retains access to cargo delivered to the cell through the endosomal system. It is possible that this latter characteristic has significance for nutrient acquisition.
This blockage in phagosome maturation is dependent on the activation status of the host macrophage. Activation of the cell with cytokines such as IFN-γ leads to partial reversal of this blockage and the delivery of the bacterium into a compartment with a pH of approximately pH 5.2 (17, 18). Although Mtb is capable of surviving low pH environments transiently, it does undergo growth arrest at lower pH. So an activated macrophage represents an inhospitable environment that is capable of restricting bacterial survival. In addition to modulation of the Mtb-containing vacuole, in mice, the predominant characteristic responsible for restricting the survival of Mtb is the inducible nitric oxide synthase (NOS2) (19). Mice deficient in this gene are almost as susceptible to Mtb infection as mice deficient in either IFN-γ or the IFN-γ receptor. Data from human macrophages activated in vivo indicate that they are also capable of generating reactive nitrogen intermediates (20, 21); however, NOS expression cannot be readily induced in human macrophages by activation in vitro. The intracellular environment inhabited by Mtb is clearly highly plastic and is actively modulated by changes in the immune status of the host.
The transcriptional response of intracellular Mtb
While Mtb is undoubtedly able to prevent full maturation of the phagosome in which it resides, it remains subject to various intraphagosomal stresses during its life cycle. These environmental stresses can act as important cues for Mtb, informing the bacterium of its location. Proper sensing and response to such environmental cues is thus critical for Mtb’s ability to successfully colonize its host. Genome-wide transcriptional studies have been very important in providing insight into the signals that Mtb perceives during infection of its host. The first global transcriptional study of Mtb during macrophage infection of up to 48 h was conducted by Schnappinger and colleagues (22) and revealed signatures indicative of a nitrosative and oxidative environment. Since then, transcriptional profiles of intracellular Mtb have been studied in various contexts, including in murine bone marrow-derived macrophage and human macrophage-like cell line infections, in Mtb murine infection models, and in clinical lung samples (23–26).
Rohde and colleagues (23) found that host cell contact alone failed to elicit changes in Mtb’s transcriptome, a distinct difference as compared to other intracellular pathogens that respond to cell contact during infection, such as Yersinia, Salmonella, and Shigella (27–29). In line with the data from Schnappinger et al. (22) however, internalization of Mtb into the macrophage led to distinct transcriptional changes: 68 genes were significantly upregulated at 2 h post-infection, with a further 75 genes identified as demonstrating sustained upregulation by EDGE analysis, which takes into account the temporal aspect of the changes (23, 30). The authors delineated several regulons among the upregulated genes, including the PhoPR and DosR regulons, as well as members of the WhiB family of transcriptional regulators. These findings are of particular interest due to their relation to bacterial sensing and response to external stimuli. Acidic pH in the phagosome is sufficient to trigger PhoP expression in Salmonella enterica Typhimirium (31), and indeed Rohde et al. (23) found that treatment of Mtb-infected macrophages with concanamycin A, an inhibitor of the vacuolar H+-ATPase, abolished upregulation of the PhoPR regulon genes previously observed. The DosR regulon has been shown to be upregulated under conditions of hypoxia and high levels of nitric oxide (NO), and during dormancy induction in experimental models (32–38). Observation of upregulation of the dormancy-related genes is also in agreement with the early transcriptional studies of Schnappinger et al. (22). Finally, members of the WhiB family all contain an iron-sulfur (Fe-S) cluster component, a recognized basis for redox-responsive sensor proteins in bacteria (39). In the case of WhiB3, Singh and colleagues demonstrated that its response to hypoxia and NO occurred through its Fe-S cluster, in a manner similar to that of Escherichia coli’s fumarate nitrate regulator, previously shown to regulate gene expression in response to these same environmental cues (40–42).
Analysis of Mtb’s transcriptional response to its environment through an extended macrophage infection time course (14 days) revealed that the largest transcriptional changes occurred at day 2 post-infection, with the expression profiles of different regulons changing in distinct patterns (43) (Fig. 2). For example, genes that were induced by acidic pH mostly had sustained high expression through the 14-day infection. In contrast, many genes of the dosR regulon that showed upregulation as early as day 2 of infection then demonstrated marked downregulation at day 8 post-infection, even as expression of the dosRS/T genes remained constant (43) (Fig. 2). This pattern is particularly intriguing in light of the growth kinetics of Mtb during this extended macrophage infection. Specifically, by tracking not just colony -orming units (CFUs) but also Mtb population growth and death rates with the use of a ‘clock’ plasmid that is lost at a known rate as the bacteria replicate (44), Rohde and colleagues (43) defined an initial phase during which both Mtb replication and death rates were high (0–2 days). This was followed by a period of adaptation, where both Mtb replication and death rates slowed, and finally a period of growth (after day 6), where the slowed replication rate exceeded the death rate, leading to an increase in CFUs (43) (Fig. 2). The timing of the initial upregulation followed by marked downregulation of the dosR regulon genes at day 8 thus coincides with stress and adaptation periods first experienced by Mtb, before a transition from adaptation to net growth. These results also indicate interestingly that the initial high replication rate of Mtb was incompatible with intracellular survival.
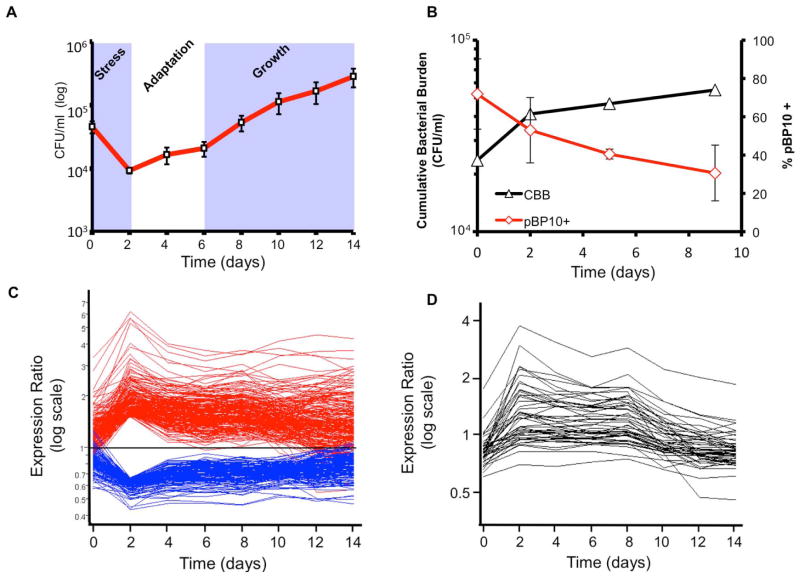
Fig. 2. Life and death dynamics during long-term intracellular survival of Mtb.
(A) Bacterial survival assays. Resting murine bone-marrow-derived macrophages were infected at low multiplicity of infection (~1:1) with CDC1551. Viable CFUs were quantified at day 0 and at 2-day intervals post-infection over a 14-day time course by lysis of monolayers, serial dilution, and plating on 7H10 medium. Error bars indicate standard error of the mean from two independent biological replicates, each consisting of three technical replicates per strain (total of six wells/strain). (B) Replication clock plasmid. The percentage of bacteria containing the pBP10 plasmid during growth in resting macrophages was determined by comparing CFUs (mean ± SD) recovered on kanamycin versus non-selective media (red). The cumulative bacterial burden (CBB) (black) was determined by mathematical modeling based on total viable CFUs and plasmid frequency data. Data shown represent two independent experiments, with each sample performed in quadruplicate (eight total wells/time point). (C) The ‘bottleneck’ response. Temporal expression profiles of genes differentially regulated at day 2 post-infection, shown as ratio of signal intensity relative to control. Note the maximal change in transcript levels at day 2 post-infection followed by the majority trending back toward control levels. (D) ‘Guilt by association’ analysis. Genes regulated in synch with known virulence regulons (i.e. the DosR regulon) were identified by using a highly regulated member of this regulon, hspX. This figure is reproduced from Rohde et al. (43)
There has been growing appreciation that genetic diversity within clinical isolates of Mtb has functional consequences that can impact on infection outcome (3, 45). In a study of two references strains (CDC1551 and H37Rv) and 15 clinical Mtb isolates representing 5 distinct genotypes, Homolka and colleagues (46) were able to identify a ‘core transcriptome’ but also observed differential abilities in macrophage colonization of the various strains. Intriguingly, they were able to tease apart different transcriptional responses between the strains in the presence of host cell stresses, and to correlate these differences with the macrophage colonization phenotype. For example, the West African strains tested infected macrophages very poorly, and were also subsequently found to fail to induce genes in the locus required for synthesis of the virulence-associated cell wall lipid phthiocerol dimycocerosate (PDIM) during infection (46). Given the role of PDIMs in aiding Mtb’s infection of its host, discussed in greater depth later in the chapter, this result suggests how genetic diversity among strains can underlie distinct transcriptional responses that in turn play a role in differential infection outcomes.
Environmental cues utilized by Mtb inside its host
It is evident from the studies described above that Mtb displays distinct transcriptional responses to host cell infection and that these responses can be matched to particular environmental signals ‘seen’ inside the host. The integration of the transcriptional-based studies and mutant studies with analysis of the physiology of the intraphagosomal environment have highlighted several candidate environmental cues that may be sensed and responded to by Mtb inside its host. One such major cue is pH. As mentioned earlier, the Mtb phagosome acidifies to a pH ~6.4 (9, 10), which represents a significant decrease from neutrality, even as it is much higher than the pH of a lysosome (pH 4.5–5.0). In essence, the ability to sense environmental pH, and other cues that likewise flux during phagosomal maturation, would act as a ‘global positioning system’ for Mtb in determining its location in the cell. Several groups have documented the profound transcriptional response of Mtb to pH (23, 47, 48). Notably, treatment of macrophages with concanamycin A prevented phagosomal acidification and resulted in elimination of the induction of almost half the genes (30/68) found to be upregulated by Mtb during initial macrophage infection (23) (Fig. 3). This result is an indication both of the importance of pH as an environmental cue for Mtb during colonization of its host, and also of the existence of yet other signals that play critical roles in allowing Mtb to sense its location in the host.
Fig. 3. Mtb utilizes pH and [Cl−] as environmental cues.
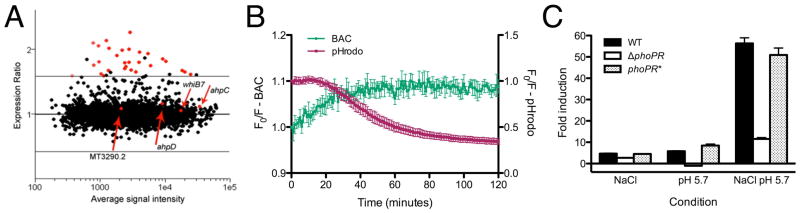
(A) Phagosome acidification is a key inducer of Mtb transcriptional changes. Murine bone marrow-derived macrophages were infected with CDC1551 for 2 h. For the test set, the macrophages were treated with 100 nM concanamycin A (CcA) to inhibit phagosomal acidification, before Mtb infection. Y-axis shows gene expression ratios from Mtb present in untreated phagosomes (pH 6.4) relative to CcA-treated phagosomes (pH 7.0). Mtb genes whose induction at 2 h was sensitive to CcA (>1.5-fold, p < 0.05) are shown in red. Red arrows indicate examples of genes induced at 2 h in untreated macrophages whose expression was insensitive to CcA. (B) [Cl−] and pH are inversely correlated during phagosome maturation. 10,10′-Bis[3-carboxylpropyl]-9,9′-biacridinium (BAC)/pHrodo beads were added to murine bone marrow-derived macrophages and BAC (green) and pHrodo (red) fluorescence tracked with a microplate reader over time. F0 is fluorescence at time = 0 min, and F is fluorescence at each given time point. BAC fluorescence decreases as [Cl−] increases. pHrodo signal increases as pH decreases. Data are shown as means ± SD from 4 wells. (C) Links between Mtb’s response to pH and Cl−. The indicated Mtb strains were grown in broth buffered at pH 7.0, pH 7.0 + 250 mM NaCl, pH 5.7, or pH 5.7 + 250 mM NaCl for 4 h. qRT-PCR of rv2390c expression in WT, ΔphoPR, and the complemented mutant (phoPR*) is shown. Fold induction is as compared to the corresponding strain grown in media at pH 7.0. Data are shown as means ± SD from 3 technical replicates. This figure is reproduced from Rohde et al. (23) and Tan et al. (54).
What are the consequences for Mtb if it fails to sense and respond to external pH in its environment? The two-component system PhoPR has been implicated as a central player in regulating Mtb’s transcriptional response to low pH (23, 49), and a phoP-deficient mutant is significantly attenuated both in macrophage and murine model infections (48). A locus unique to the tuberculosis complex, aprABC (‘acid and phagosome regulated’), was among the Mtb genes upregulated within 2 hours of macrophage infection that was sensitive to concanamycin A treatment (23, 49). An aprA-deficient mutant exhibited extensive transcriptional changes, suggesting a role as a transcriptional regulator (49). aprABC expression was regulated by phoP, as acidic pH induction of aprA expression was lost in a phoP mutant background (49). Like the phoP-deficient Mtb, the aprABC-deficient mutant was attenuated for growth in macrophages (48, 49). At acidic pH, Mtb increased synthesis of PDIMs, and both phoP and aprABC were found to be involved in regulation of PDIM synthesis: aprA-deficient mutants failed to induce phthiocerol A and phthiodiolone mycocerates at low pH, while phoP-deficient mutants overproduced these two lipids (49). These data are in agreement with results from screens for acidic pH-sensitive Mtb mutants, in that an ability to maintain cell wall integrity appears to be an important component of Mtb’s pH response. For example, in a screen for Mtb mutants unable to resist acidic pH, Vandal and colleagues identified rv3671c as a gene required for maintenance of intrabacterial pH homeostasis in the face of low external pH, with a mutant deficient in this gene severely attenuated in a murine model of Mtb infection (50). rv3671c encodes a membrane-bound protease, and the mutant possessed cell wall defects, as illustrated by an increased sensitivity to detergents, and to lipophilic antibiotics such as rifampicin and erythromycin. All of the above data demonstrate the importance of pH sensing and response to Mtb’s survival and growth in its host.
Phagosomal acidification is the influx of protons into the phagosome, and recent studies have begun to shed further insight into other environmental cues linked to this influx of positive charge that Mtb also responds to. In accord with data from endosomal maturation (51–53), chloride was found to act as a counter-anion that balances proton movement during phagosomal acidification: as pH decreased in the maturing phagosome, chloride concentration increased (54) (Fig. 3). Tan and colleagues (54) found that Mtb responds transcriptionally to high chloride concentration ([Cl−]), with many of the genes induced in high [Cl−] also induced in low pH. Strikingly, Mtb’s response to pH and [Cl−] was synergistic, allowing it exploit these two interlinked host signals to inform it of its location in the host. As a further link between pH and Cl−, the PhoPR two-component system known to be critical in Mtb’s response to pH was also found to play a role in the bacterium’s response to Cl−, with a phoPR-deficient mutant showing much reduced induction in expression of Cl−-responsive genes, in the presence of high [Cl−] (54), (Fig. 3).
In addition to the flux of ions such as H+ and Cl−, metal ions constitute a potentially important environmental aspect of the phagosome. Using X-ray fluorescence microscopy, Wagner and colleagues (55) determined the levels of 10 single elements in Mtb-infected macrophage phagosomes. Of note, they observed high Cl− levels in the Mtb-containing phagosomes, supporting the findings of Tan et al. regarding Cl− as an environmental cue for Mtb (54, 55). In resting macrophages, the Mtb-containing phagosome was found to have high iron concentrations; however, additional experiments with Mycobacterium avium showed that activation of macrophages led to a significant decrease in phagosomal iron concentration (55). These results match the transcriptional data of Homolka et al. (46), who reported upregulation of bfrB, encoding an iron storage protein, during Mtb infection of resting macrophages; conversely, upregulation of genes involved in iron acquisition (mycobactin mbtA-J) were observed during Mtb infection of activated macrophages. Interestingly, Wagner and colleagues (55) found high levels of copper and zinc in Mtb-containing phagosomes, which fits with the transcriptional data from Botella et al. (56, 57) identifying a heavy metal intoxication ‘signature’ in Mtb during infection of host macrophages. This was reflected in the sustained upregulation of several putative P1B-type ATPase-encoding genes (ctpC, ctpG, and ctpV), as well as that of the P2A-type ATPase-encoding gene ctpF, and the metal-responsive transcriptional regulators cmtR and csoR (56, 57). Using confocal and electron microscopy methods, they were able to visualize a ‘burst’ of free zinc during Mtb infection of macrophages, with zinc accumulation within the phagosome and in Mtb (56). Studies with a ctpC-deficient mutant showed that it accumulated more zinc intrabacterially than wildtype Mtb (56). Further, the mutant was exquisitely sensitive to zinc in broth, and was deficient in its ability to infect macrophages.
As with zinc, Mtb has a significant transcriptional response to copper and encodes a copper-responsive transcriptional regulator, csoR (58). CtpV and MctB are two proteins that have been implicated in copper efflux from Mtb (59, 60). Both ctpV-deficient and mctB-deficient mutants accumulated excess copper and were more sensitive to copper (59, 60), and the mctB-deficient mutants were severely attenuated for infection of guinea pigs (60). While the bacterial load for ctpV-deficient mutants in both murine and guinea pig models of Mtb infection was similar to wildtype bacteria, decreased histopathology was observed in the case of the mutant-infected lungs, with an associated increased survival time documented as well (59).
Mtb, like many microorganisms, must walk a knife edge with metal ions such as those described above, as these ions are both critical for Mtb growth, for example in their roles as cofactors in essential enzymes (61, 62), and yet simultaneously toxic at high levels. As evidenced from the many genes that Mtb possesses for combating metal intoxication, the bacterium has evolved to resist the host’s attempts at infection control via the concentration of metals such as zinc and copper within the phagosome.
In addition to the different ion concentrations within the phagosome, Mtb must also contend with reactive oxygen and nitrogen intermediates generated by their host macrophages. As described earlier, macrophages produce reactive oxygen intermediates (ROI) via the NADPH oxidase complex, with levels increased upon IFN-γ or LPS activation of the macrophages (7). Nonetheless, this superoxide burst on its own does not seem to exert much control on Mtb infection, as p47phox−/− mice demonstrate only an early and transient defect in restricting Mtb growth (63). Mtb does respond transcriptionally to oxidative stress (64), and several resistance mechanisms have been described. This includes the production of reactive oxygen scavenging enzymes such as superoxide dismutases (SodA, SodC) that convert superoxide to H2O2 (65–67), and catalase-peroxidase-peroxynitritase enzymes (KatG, and a AphC/AphD/DlaT/LpD complex) that can process H2O2 (68, 69). Mtb also synthesizes mycothiol (instead of glutathione), a low molecular weight thiol that acts as an antioxidant, maintaining a reduced intrabacterial environment (70, 71). In addition, the methionine sulfoxide reductases MsrA and MsrB work to reverse oxidative damage to methionine residues, although this mechanism is not critical for Mtb’s survival in its host, as a msrA/msrB double mutant displayed only a minimal phenotype in a murine infection (72).
While Mtb has clearly evolved multiple mechanisms for successfully dealing with oxidative stress such that the activities of the NADPH oxidase complex do not appear to play a sustained role in controlling Mtb infection, nitric oxide (NO) and its products are known to be extremely important for host control of Mtb. Most strikingly, NOS2−/− mice are almost as susceptible to Mtb infection as IFN-γ−/− mice, which develop a fatal disseminated infection (19, 73, 74). Work done in a macaque model of Mtb infection showed the upregulation of NOS activity in granulomatous tissue as compared to uninvolved tissue from the same host (75), and increased levels of NOS expression and NO production have also been demonstrated during Mtb infection in human lungs (76, 77), indicating the likely relevance of NO in human TB disease. Mtb demonstrates a significant transcriptional response to NO, including upregulation of genes in the aforementioned dosR regulon (33, 35, 64, 78). The peroxidase-peroxynitritase complex of AphC/AphD/DlaT/LpD that functions in ROI detoxification also plays a role in detoxifying reactive nitrogen intermediates (RNI) (68). In a screen for Mtb mutants unable to withstand RNI stress, Darwin and colleagues (79) identified the bacterial proteasome as an important player. An Mtb mutant deficient in the proteasome ATPase mpA was more susceptible to RNI-mediated damage and was attenuated in its ability to colonize a murine host, a phenotype that was partially rescued in NOS2−/− mice (79, 80).
Linked metabolism of host and pathogen
One area in which the intracellular environment has clearly molded the Mtb genome is in central carbon metabolism. The Mtb/macrophage interaction is a long-lived one and during this interplay the bacterium must sustain itself by utilizing carbon sources available within the host cell. In 2000, McKinney and colleagues (81) demonstrated that Mtb deficient in expression of isocitrate lyase (icl1) survived poorly in immune-competent mice. As isocitrate lyase was known to be the gating enzyme to the glyoxylate shunt and is required for retention of carbon when growing on fatty acids as a limiting carbon source, these data were thought to indicate that Mtb relied on host fatty acids for nutrition inside its host cell. Shortly afterwards, Sassetti and Rubin (82) performed a high density transposon insertion site mapping screen (TraSH) to identify genes essential for infection in the mouse, and in subsequent work, they went on to show that the mce4 gene encoded part of a cholesterol importation apparatus (83). Several laboratories have gone on to show that the ability of Mtb to acquire and process cholesterol is necessary for the optimum growth of Mtb, both in macrophages in culture and in experimental murine infections (84–87).
The degradation of cholesterol is a complex process, and the Mtb genome contains a region of 83 genes involved in this process (88). This locus encodes the majority, but not all, of the enzymes involved in the breakdown of the sterol rings and the β-oxidation of the acyl side chain. Degradation of cholesterol yields pyruvate, acetyl-CoA, and propionyl-CoA, the latter being potentially toxic to Mtb. Mtb has three possible routes of detoxification of propionyl-CoA, the methyl citrate cycle (MCC), the methyl malonyl pathway (MMP), and the utilization of methyl malonyl-CoA to build complex cell wall lipids such a pthiocerol dimycocerosate (PDIM) (89). McKinney and colleagues demonstrated that ICL1 is actually a bifunctional enzyme that is capable of processing both isocitrate and methylisocitrate, and as such the phenotype of a Δicl1 mutant in macrophages and mice is due to the inability of the MCC to adequately detoxify the products from propionyl-CoA (90–92). The interplay between the different routes of propionyl-CoA from cholesterol in intracellular Mtb was further probed in a recent study on pathways of rescue of the Δicl1 mutant in macrophages (93). The Δicl1 mutant exhibits minimal growth in murine macrophages in culture. On the basis of transcriptional profiling that shows marked upregulation of the cholesterol degradation locus and the genes of the MCC (43, 46), it was hypothesized that the bacterium was experiencing intoxication from the accumulation of propionyl-CoA, indicating that MCC is the primary route of processing of this toxic intermediate. Growth of the mutant could be restored via two different routes. First, the addition of exogenous vitamin B12 (VitB12) opens the MMP to facilitate conversion of propionyl-CoA into succinyl-CoA. The MutAB methylmalonyl mutase enzyme complex requires VitB12 as a co-factor and, although Mtb appears to possess the genes encoding the synthetic pathway for VitB12, conditions have not been found under which the bacterium synthesizes the vitamin. Second, the addition of oleic acid to the infected macrophage culture enhances the progression of the infected macrophage to a foamy phenotype, laden with lipid droplets. This fatty acid also rescues the impaired growth phenotype of the Δicl1 mutant. Metabolic labeling studies show that the addition of oleic acid expands the acetyl-CoA pool in the bacterium, providing the additional building blocks to expand the synthesis of PDIM and shuttle toxic propionyl-CoA, in the form of malonyl-CoA, into the peripheral lipids of the bacterial cell wall. The ability of intracellular Mtb to access and utilize host derived cholesterol and fatty acids has been demonstrated genetically, biochemically, and through the use of fluorescently tagged fatty acids that accumulate as droplets inside intracellular Mtb.
Modulation of host tissue metabolism at the site of infection
The infected macrophage does not exist in isolation, because the infection triggers a robust pro-inflammatory response at the infection site, and this leads to robust cell recruitment of neutrophils, other macrophages, and lymphocytes. The tissue undergoes extensive, pathogen-driven remodeling to generate a macrophage-rich granuloma (94–96). Within an infected individual these granulomas are extremely heterogeneous, and even in an individual with active disease, only some of the granulomas harbor a productive infection (75, 97). Nevertheless, it is generally accepted that active granulomas, that have live, replicating organisms and have the potential to progress, exhibit extensive central caseation. These granulomas are rich in foamy macrophages that have extensive lipid stores in the cytosol (Fig. 4). Interestingly, similar cellular responses can be induced by inoculation of mice with polystyrene beads coated with the Mtb cell wall lipid trehalose dimycolate (TDM) (94, 98). Moreover, transformation of the non-pathogenic Mycobacterium smegmatis with the Mtb gene encoding methyl mycolic acid synthase enables M. smegmatis to synthesize Mtb-like TDM that induces lipid droplets in macrophages (95). TDM is recognized by the macrophage receptors Mincle and MARCO (99, 100). It is thought that the interaction with MARCO stabilizes a weak interaction with TLR2 to facilitate activation of the macrophages that leads to foam cell formation (99). In culture, the induction of a foamy macrophage phenotype is not restricted to the infected macrophages but is exhibited by all the cells in the culture. The same is true in the granuloma where this lipid-loaded phenotype is common among those macrophages subtending the caseous center (Fig. 4). It had been noted previously that Mtb actively sheds its peripheral cell wall lipids and that these are released from infected macrophages in vesicular exosomes containing both host and bacterial constituents (101, 102). The exosomes are internalized by neighboring cells and are thought to represent a mechanism by which Mtb can extend its control over the host tissue beyond the boundaries of the infected macrophage. Such a mechanism could help explain how the relatively few bacteria present in human TB granulomas manage to exert such an extreme influence over the host infection site.
Fig. 4. Expression of the lipid droplet-associated protein Peripilin 2 in human tuberculosis granulomas.
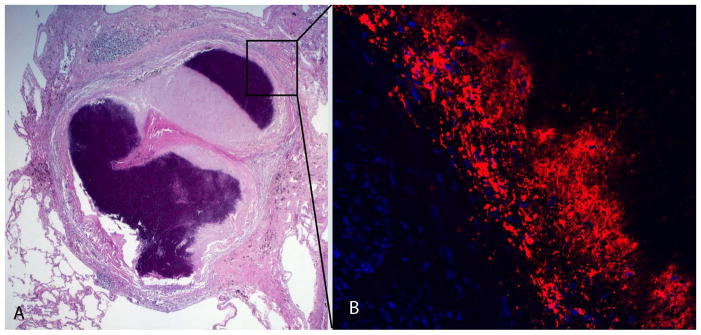
Immunofluorescence signals were obtained for each granuloma (B), and the corresponding region from a hematoxylin and eosin stained slide (A) is shown. Nuclei are shown in blue and antigens in red. The macrophages subtending the caseum of this fibrocaseous granuloma label strongly for peripilin 2 expression. This figure is reproduced from Kim et al. (94).
Mass spectrometry of caseum isolated from human TB granulomas revealed that the most abundant lipids present were triacylglycerol, cholesterol and cholesterol esters (94). The presence of cholesterol in its esterified form is particularly revealing. When cells such as macrophages internalize cholesterol on low-density lipoproteins they either actively excrete the cholesterol through ABC transporter-mediated efflux (103), or if retaining the cholesterol, they esterify it and sequester it in lipid droplets to minimize its toxicity (104). So the presence of cholesterol esters in the caseum implies that the lipid has come from dead foamy macrophages, and that death of these cells is what drives the expansion of the caseous center of the granuloma.
Mtb bacilli are frequently observed in the caseum of human and primate granulomas. In addition, it has been noted that bacteria recovered from the sputum of individuals with active TB are frequently loaded with lipid inclusions (105). All these observations indicate that Mtb has not only evolved to utilize cholesterol as its primary carbon source in its host cell but also to generate a chronic, proinflammatory response, similar to that observed in atherosclerotic lesions, that leads to retention of cholesterol and fatty acids at the site of infection.
Exploiting reporter strains to study Mtb infection in vitro and in vivo
The studies described above unmistakably demonstrate the critical role that sensing and responding to intraphagosomal environmental signals plays in Mtb’s successful colonization of its host. However, while genome-wide transcriptional studies, mutant analysis, and population-level assays such as CFU enumeration have yielded important insights into the environment that Mtb encounters inside its host and its response to it, there remain significant unanswered questions. Among others, further work is needed to understand (i) the interplay between the bacterium and the host’s response in the spatial context of the tissue environment, and (ii) the heterogeneity that may exist in the bacterial population’s response to its environment, and how this impacts on the progression of infection and disease treatment. Addressing questions such as these will require expanding our molecular ‘toolbox’ to allow examination of Mtb’s response at the level of the individual bacterium.
To this end, the use of fluorescent proteins has proven to be an especially productive tool in Mtb studies. Variants of green fluorescent protein (GFP) that are pH or redox sensitive (106, 107) have been successfully incorporated into Mtb to yield information about the intrabacterial pH and mycothiol redox potential during macrophage infection. Using a pH-sensitive GFP, Vandal and colleagues (50) demonstrated that wildtype Mtb is able to maintain its intrabacterial pH at or near neutral pH, even while residing in the phagosome of IFN-γ activated macrophages. This was not the case with a mutant deficient in rv3671c, a gene encoding a membrane-bound protease, with the majority of the mutant bacteria possessing an internal pH of less than 5.5 during infection of IFN-γ-activated macrophages (50). Bhaskar and colleagues (108) engineered a fusion of a mycoredoxin (Rv3198A) to the redox-sensitive roGFP2 to generate a mycothiol-specific intrabacterial fluorescent redox probe. Using Mtb carrying this probe, they found significant intrabacterial redox heterogeneity in Mtb-infecting macrophages, which was not observed in broth-grown Mtb. Through colocalization experiments with markers to approximate sub-vacuolar compartments, they determined that the heterogeneity reflected in part the presence of the bacteria in different compartments. Mtb in early endosomes (early endosome antigen and Rab5-positive) versus lysosomes (lysotracker and cathepsin D-positive) were more likely to possess reduced versus oxidized mycothiol redox potentials respectively (108). All the Mtb observed in autophagosomes (LC3-positive) had a highly oxidized mycothiol redox potential. Intrabacterial mycothiol redox potential was modulated by the host immune status, with immune activation leading to an oxidative shift. In accord with the concept that NO is a major source of redox stress in the Mtb phagosome, chemical inhibition of iNOS substantially decreased the oxidative shift that had been observed (108). Similarly, treatment with anti-TB drugs such as isoniazid, ethambutol, and rifampicin led to a shift towards an oxidized mycothiol redox potential, but only in the context of Mtb infection of host cells, suggesting that the drugs potentiate the host response to Mtb (108). Intriguingly, Bhaskar and colleagues (108) were able to correlate the observed mycothiol redox potential heterogeneity in the bacterial population during host cell colonization with differential susceptibility to anti-TB drugs. Specifically, they found that Mtb with a more oxidized mycothiol redox potential were more susceptible to drug killing, with increased membrane damage as reflected by increased propidium iodide (PI) staining (108). In contrast, Mtb with a more reduced mycothiol redox potential were more antibiotic tolerant, displaying no uptake of PI. These results demonstrate the utility of reporter strains for studying Mtb infection in the context of intact host cells, and provide a salient example of the biological relevance of Mtb’s population heterogeneity during infection.
Mtb’s robust transcriptional response to particular environmental cues represents an additional opportunity for the construction of fluorescent reporter tools that can aid understanding of Mtb’s surroundings during in vivo infection. By carefully selecting Mtb gene promoters known to be induced in the presence of a given environmental signal and placing them upstream of GFP, reporter Mtb strains can be generated whose fluorescence varies in accord with the bacterium’s microenvironment. Combining this GFP reporter with a constitutively expressed second fluorophore (e.g. smyc′::mCherry) further enables visualization of all bacteria irrespective of GFP induction levels, allowing rigorous quantification of reporter GFP signal. Abramovitch and colleagues used this approach to study the expression of aprA in broth and during macrophage infection (aprA′::GFP reporter) (49). They found that aprA′::GFP fluorescence was induced to similar levels during Mtb infection of both resting and activated macrophages, with significant heterogeneity across the bacterial population.
Tan and colleagues have since generated a reporter Mtb strain responsive to pH and [Cl−] (rv2390c′::GFP) (54). GFP expression of this reporter strain was increased in activated versus resting macrophages, in accord with increased phagosomal maturation (54). They brought studies with the reporter Mtb strains a step further by using them for in vivo infections in a murine model of Mtb infection. By taking thick sections of lung tissue, imaging a 10 μm deep z-stack and reconstructing into 3D, they were able to quantify reporter induction in the context of the tissue environment. In agreement with the macrophage infection results, rv2390c′::GFP expression was lower in Mtb infecting IFN-γ−/− mice as compared to wildtype mice (54). Tan and colleagues (54) also constructed a second reporter Mtb strain responsive to hypoxia and NO (hspX′::GFP), and similarly, they examined its induction during infection of wildtype versus IFN-γ−/− mice. hspX′::GFP fluorescence was induced more strongly in wildtype as compared to IFN-γ−/− mice, and expression was significantly higher at 28 days versus 14 days post-infection, correlating with the time frame of the adaptive immune response and the production of iNOS (54, 109).
In addition to promoter-based reporters and reporters exploiting intrinsic properties of variant GFP molecules, Mtb protein-fluorophore fusions represent yet another method for the generation of informative reporter Mtb strains. Protein-fluorophore fusions have been extensively used for studying protein localization, and several studies have also employed this method in live-cell time-lapse imaging on the fast-growing saprophyte M. smegmatis, for example to track the dynamics of the cell wall synthetic machinery (110) and to examine KatG expression and its relation to killing by INH (111). However, protein-fluorophore fusions can potentially be exploited not just for the study of a given protein’s localization and dynamics but also in the generation of reporter strains that inform a broader phenotype/physiological state. A striking example of such a reporter Mtb strain was recently described by Sukumar and colleagues, consisting of a fusion of Mtb’s single stranded binding protein to GFP (SSB-GFP) (112). This fusion protein was driven by the native Mtb ssb promoter and was coupled to a constitutively expressed mCherry as with the other promoter-based reporter Mtb strains described above. In contrast to the other reporter Mtb strains that respond to particular environmental cues however, this reporter allows the tracking of Mtb replication by marking bacteria undergoing active DNA replication. Fluorescent tagging of components of the replisome, such as SSB, has been well-validated as a marker of active DNA replication in several organisms, including E. coli and Bacillus subtilis (113–115). Similar to these other bacteria, SSB-GFP foci were present during periods of DNA replication in both M. smegmatis and Mtb (112). While methods such as CFU enumeration and the use of a ‘clock’ plasmid lost at a fixed rate from replicating Mtb (44) provide information on the numbers of surviving Mtb and Mtb replication on a population level, the SSB-GFP reporter enables the study of Mtb replication at the level of the individual bacterium, in the context of its tissue environment.
By combining the use of these reporter Mtb strains with specific, immune-deficient mice and utilizing a heat-killed Mtb vaccination model, Sukumar and colleagues (112) were able to expand on the initial studies of Tan et al., probing the physiological and replication status of Mtb in heterogeneous tissue environments and examining the impact of the host immune status on the bacterium’s response. While Mtb infecting mock-treated mice showed initially high induction levels of the pH and Cl−-responsive rv2390c′::GFP reporter (to 28 days post-challenge), levels of this reporter fluorescence was significantly lower in Mtb infecting vaccinated mice at these early time points (112). The presence of a pre-existing immune response to Mtb stimulated by vaccination thus appears to either hasten adaptation of surviving Mtb to innate immune stresses, or to accelerate selection of the bacterial population best adapted for continued survival. As observed previously, the kinetics of the hypoxia and NO-responsive hspX′::GFP expression were different from those of the rv2390c′::GFP reporter, with low levels of fluorescence observed initially, followed by strong induction at later time points as the adaptive immune response was established (54, 112). As expected, vaccination accelerated the onset of the adaptive immune response and iNOS production, leading to high levels of hspX′::GFP induction in Mtb present in vaccinated mice even at 14 days post-challenge (112) (Fig. 5). Experiments with NOS2−/− mice confirmed that NO was the main driver of hspX expression in the murine model, as hspX′::GFP reporter fluorescence was ~3 logs lower in Mtb infecting NOS2−/− versus wildtype mice (Fig. 5).
Fig. 5. Impact of host immune response on dynamics of hspX′::GFP induction.
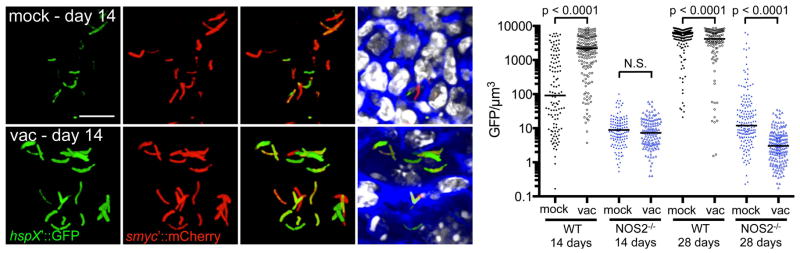
Erdman(hspX′::GFP, smyc′::mCherry) was inoculated into vaccinated or mock-treated C57BL/6J WT mice for up to 28 days. 3D confocal images from a 14-day infection are shown. All bacteria are marked in red (smyc′::mCherry), reporter signal is shown in green (hspX′::GFP), nuclei are marked in grayscale (DAPI), and phalloidin staining of f-actin is shown in blue. Scale bar 10 μm. Graph shows quantification of the GFP/μm3 signal for each bacterium measured from multiple 3D confocal images, at the indicated time points. Each point on the graph represents a bacterium or a tightly clustered group of bacteria (mock-treated – filled symbols, vaccinated – open symbols; WT mice infections – black, NOS2−/− mice infections - blue). Horizontal lines mark the median value for each group. p-values were obtained with a Mann-Whitney statistical test. This figure is reproduced from Sukumar et al. (112).
Using the SSB-GFP replication reporter Mtb, Sukumar and colleagues (112) found significant heterogeneity in the percentage of actively replicating Mtb at day 14 post-challenge in both mock-treated and vaccinated mice (Fig. 6). Strikingly, vaccinated mice displayed a much lower percentage of actively replicating Mtb at day 28 post-challenge as compared to mock-treated mice (Fig. 6). In accord with this growth profile, Mtb recovered from vaccinated mice were more tolerant to isoniazid, a drug with preferential activity against actively replicating bacteria, versus Mtb recovered from mock-treated mice (112). While it is thus clear that vaccination can reduce at least initial bacterial burden in the host, the study by Sukumar et al. (112) indicate that it also accelerates the surviving Mtb’s adaptation to immune pressure and its progression to a more persistent stage of colonization. These reporter Mtb-based studies also demonstrate the close link between the host’s immune status and the environmental signals that Mtb finds itself exposed to. While many of the environmental signals described above are manifestations of the host’s attempt to clear the infection, Mtb’s ability to resist the various defense mechanisms allows it to usurp these very signals as cues that inform it both of its location in the host and the immune status of the host, and to respond appropriately to enable its continued survival.
Fig. 6. SSB-GFP reporter tracks Mtb replication in vivo.
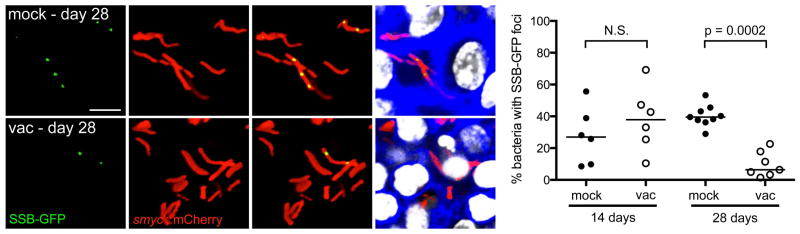
Erdman(SSB-GFP, smyc′::mCherry) was inoculated into vaccinated or mock-treated C57BL/6J WT mice for up to 28 days. 3D confocal images from a 28-day infection are shown with all bacteria marked in red (smyc′::mCherry), reporter signal shown in green (SSB-GFP), nuclei marked in grayscale (DAPI), and phalloidin staining of f-actin shown in blue. For clarity of foci visualization, SSB-GFP signal is shown in extended focus, overlaid on the 3D image. Scale bar 10 μm. Graph shows the percentage of Mtb displaying SSB-GFP foci for each mouse, measured from multiple 3D confocal images, at 14 and 28 days post-challenge. Each point on the graph represents a mouse (mock-treated – filled symbols, vaccinated – open symbols). Horizontal lines mark the median value for each group. p-values were obtained with a Mann-Whitney statistical test. This figure is reproduced from Sukumar et al. (112).
While the in vivo environment is certainly more complex than the defined in vitro conditions under which the reporter constructs are initially characterized, the studies by Tan et al. and Sukumar et al. illustrate how the use of multiple reporter strains, combined with genetic and immunological tools on the host side, allow for insightful discoveries into Mtb’s microenvironment in vivo and the impact of the host immune status on the bacterium’s response to environmental signals. Another important observation from studies with the various reporter Mtb strains is the heterogeneity that exists in the bacterial population during infection. Lesion to lesion heterogeneity within the lungs of Mtb-infected rabbit or macaques have been elegantly demonstrated in studies from Lin et al. and Via et al., with the progress and fate of each lesion being independent (18, 116). The data from the reporter Mtb strains further indicate that heterogeneity is present at the level of the individual bacilli. That this heterogeneity can have significant effects on Mtb biology and drug treatment efficacy is demonstrated by the data of Bhaskar and colleagues (108). We envision that continued expansion of the types of reporter Mtb strains available will significantly enhance the field’s ability to understand Mtb-host interactions in vivo.
Concluding remarks
Intracellular Mtb senses and responds to immune-mediated alterations in the physiological status of its host cell. These alterations tune bacterial transcriptional and metabolic responses to facilitate survival of the bacterium. Many of the current analyses have relied on population-based measurements that have been invaluable in the identification of the major themes and metabolic adjustments undergone by Mtb but do not address the issue of heterogeneity. We feel that the more recent studies exploiting reporter strains of Mtb that exhibit either conditional expression of fluorescent proteins, or express fluorescent sensor proteins, represent the latest generation of tools for mining the biology of mycobacterial infections. These are exciting new probes for understanding bacterial fitness in the context of the tissue environment of the host, with reference to experimental models of both vaccination and chemotherapy.
Acknowledgments
The research discussed in this article was supported in the most part by the National Institutes of Health awards HL055936 and AI067027 to DGR.
Footnotes
The authors have no conflicts of interest to declare.
References
- 1.Sreevatsan S, et al. Restricted structural gene polymorphism in the Mycobacterium tuberculosis complex indicates evolutionarily recent global dissemination. Proc Natl Acad Sci U S A. 1997;94:9869–9874. doi: 10.1073/pnas.94.18.9869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Cole ST, et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature. 1998;393:537–544. doi: 10.1038/31159. [DOI] [PubMed] [Google Scholar]
- 3.Hershberg R, et al. High functional diversity in Mycobacterium tuberculosis driven by genetic drift and human demography. PLoS Biol. 2008;6:e311. doi: 10.1371/journal.pbio.0060311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Russell DG. Mycobacterium tuberculosis and the intimate discourse of a chronic infection. Immunological reviews. 2011;240:252–268. doi: 10.1111/j.1600-065X.2010.00984.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Russell DG, Vanderven BC, Glennie S, Mwandumba H, Heyderman RS. The macrophage marches on its phagosome: dynamic assays of phagosome function. Nature reviews Immunology. 2009;9:594–600. doi: 10.1038/nri2591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Yates RM, Hermetter A, Taylor GA, Russell DG. Macrophage activation downregulates the degradative capacity of the phagosome. Traffic. 2007;8:241–250. doi: 10.1111/j.1600-0854.2006.00528.x. [DOI] [PubMed] [Google Scholar]
- 7.VanderVen BC, Yates RM, Russell DG. Intraphagosomal measurement of the magnitude and duration of the oxidative burst. Traffic. 2009;10:372–378. doi: 10.1111/j.1600-0854.2009.00877.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Yates RM, Hermetter A, Russell DG. The kinetics of phagosome maturation as a function of phagosome/lysosome fusion and acquisition of hydrolytic activity. Traffic. 2005;6:413–420. doi: 10.1111/j.1600-0854.2005.00284.x. [DOI] [PubMed] [Google Scholar]
- 9.Pethe K, Swenson DL, Alonso S, Anderson J, Wang C, Russell DG. Isolation of Mycobacterium tuberculosis mutants defective in the arrest of phagosome maturation. Proc Natl Acad Sci U S A. 2004;101:13642–13647. doi: 10.1073/pnas.0401657101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sturgill-Koszycki S, et al. Lack of acidification in Mycobacterium phagosomes produced by exclusion of the vesicular proton-ATPase. Science. 1994;263:678–681. doi: 10.1126/science.8303277. [DOI] [PubMed] [Google Scholar]
- 11.Podinovskaia M, Lee W, Caldwell S, Russell DG. Infection of macrophages with Mycobacterium tuberculosis induces global modifications to phagosomal function. Cell Microbiol. 2013;15:843–859. doi: 10.1111/cmi.12092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yates RM, Hermetter A, Russell DG. Recording phagosome maturation through the real-time, spectrofluorometric measurement of hydrolytic activities. Methods in molecular biology. 2009;531:157–171. doi: 10.1007/978-1-59745-396-7_11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yates RM, Russell DG. Real-time spectrofluorometric assays for the lumenal environment of the maturing phagosome. Methods in molecular biology. 2008;445:311–325. doi: 10.1007/978-1-59745-157-4_20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Sturgill-Koszycki S, Schaible UE, Russell DG. Mycobacterium-containing phagosomes are accessible to early endosomes and reflect a transitional state in normal phagosome biogenesis. EMBO J. 1996;15:6960–6968. [PMC free article] [PubMed] [Google Scholar]
- 15.Ullrich HJ, Beatty WL, Russell DG. Direct delivery of procathepsin D to phagosomes: implications for phagosome biogenesis and parasitism by Mycobacterium. European journal of cell biology. 1999;78:739–748. doi: 10.1016/S0171-9335(99)80042-9. [DOI] [PubMed] [Google Scholar]
- 16.Russell DG, Dant J, Sturgill-Koszycki S. Mycobacterium avium- and Mycobacterium tuberculosis-containing vacuoles are dynamic, fusion-competent vesicles that are accessible to glycosphingolipids from the host cell plasmalemma. J Immunol. 1996;156:4764–4773. [PubMed] [Google Scholar]
- 17.Schaible UE, Sturgill-Koszycki S, Schlesinger PH, Russell DG. Cytokine activation leads to acidification and increases maturation of Mycobacterium avium-containing phagosomes in murine macrophages. J Immunol. 1998;160:1290–1296. [PubMed] [Google Scholar]
- 18.Via LE, Fratti RA, McFalone M, Pagan-Ramos E, Deretic D, Deretic V. Effects of cytokines on mycobacterial phagosome maturation. J Cell Sci. 1998;111(Pt 7):897–905. doi: 10.1242/jcs.111.7.897. [DOI] [PubMed] [Google Scholar]
- 19.MacMicking JD, North RJ, LaCourse R, Mudgett JS, Shah SK, Nathan CF. Identification of nitric oxide synthase as a protective locus against tuberculosis. Proc Natl Acad Sci U S A. 1997;94:5243–5248. doi: 10.1073/pnas.94.10.5243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Nozaki Y, Hasegawa Y, Ichiyama S, Nakashima I, Shimokata K. Mechanism of nitric oxide-dependent killing of Mycobacterium bovis BCG in human alveolar macrophages. Infect Immun. 1997;65:3644–3647. doi: 10.1128/iai.65.9.3644-3647.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Weinberg JB, et al. Human mononuclear phagocyte inducible nitric oxide synthase (iNOS): analysis of iNOS mRNA, iNOS protein, biopterin, and nitric oxide production by blood monocytes and peritoneal macrophages. Blood. 1995;86:1184–1195. [PubMed] [Google Scholar]
- 22.Schnappinger D, et al. Transcriptional Adaptation of Mycobacterium tuberculosis within Macrophages: Insights into the Phagosomal Environment. J Exp Med. 2003;198:693–704. doi: 10.1084/jem.20030846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Rohde KH, Abramovitch RB, Russell DG. Mycobacterium tuberculosis invasion of macrophages: linking bacterial gene expression to environmental cues. Cell Host Microbe. 2007;2:352–364. doi: 10.1016/j.chom.2007.09.006. [DOI] [PubMed] [Google Scholar]
- 24.Fontan P, Aris V, Ghanny S, Soteropoulos P, Smith I. Global transcriptional profile of Mycobacterium tuberculosis during THP-1 human macrophage infection. Infect Immun. 2008;76:717–725. doi: 10.1128/IAI.00974-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Talaat AM, Lyons R, Howard ST, Johnston SA. The temporal expression profile of Mycobacterium tuberculosis infection in mice. Proc Natl Acad Sci U S A. 2004;101:4602–4607. doi: 10.1073/pnas.0306023101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rachman H, et al. Unique transcriptome signature of Mycobacterium tuberculosis in pulmonary tuberculosis. Infect Immun. 2006;74:1233–1242. doi: 10.1128/IAI.74.2.1233-1242.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pettersson J, et al. Modulation of virulence factor expression by pathogen target cell contact. Science. 1996;273:1231–1233. doi: 10.1126/science.273.5279.1231. [DOI] [PubMed] [Google Scholar]
- 28.Zierler MK, Galan JE. Contact with cultured epithelial cells stimulates secretion of Salmonella typhimurium invasion protein InvJ. Infect Immun. 1995;63:4024–4028. doi: 10.1128/iai.63.10.4024-4028.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Watarai M, Tobe T, Yoshikawa M, Sasakawa C. Contact of Shigella with host cells triggers release of Ipa invasins and is an essential function of invasiveness. Embo J. 1995;14:2461–2470. doi: 10.1002/j.1460-2075.1995.tb07243.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Storey JD, Xiao W, Leek JT, Tompkins RG, Davis RW. Significance analysis of time course microarray experiments. Proc Natl Acad Sci U S A. 2005;102:12837–12842. doi: 10.1073/pnas.0504609102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Martin-Orozco N, et al. Visualization of vacuolar acidification-induced transcription of genes of pathogens inside macrophages. Mol Biol Cell. 2006;17:498–510. doi: 10.1091/mbc.E04-12-1096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Roberts DM, Liao RP, Wisedchaisri G, Hol WG, Sherman DR. Two sensor kinases contribute to the hypoxic response of Mycobacterium tuberculosis. J Biol Chem. 2004;279:23082–23087. doi: 10.1074/jbc.M401230200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Voskuil MI, et al. Inhibition of respiration by nitric oxide induces a Mycobacterium tuberculosis dormancy program. J Exp Med. 2003;198:705–713. doi: 10.1084/jem.20030205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Sherman DR, Voskuil M, Schnappinger D, Liao R, Harrell MI, Schoolnik GK. Regulation of the Mycobacterium tuberculosis hypoxic response gene encoding alpha -crystallin. Proc Natl Acad Sci U S A. 2001;98:7534–7539. doi: 10.1073/pnas.121172498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ohno H, et al. The effects of reactive nitrogen intermediates on gene expression in Mycobacterium tuberculosis. Cell Microbiol. 2003;5:637–648. doi: 10.1046/j.1462-5822.2003.00307.x. [DOI] [PubMed] [Google Scholar]
- 36.Park HD, et al. Rv3133c/dosR is a transcription factor that mediates the hypoxic response of Mycobacterium tuberculosis. Mol Microbiol. 2003;48:833–843. doi: 10.1046/j.1365-2958.2003.03474.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kumar A, Toledo JC, Patel RP, Lancaster JR, Jr, Steyn AJ. Mycobacterium tuberculosis DosS is a redox sensor and DosT is a hypoxia sensor. Proc Natl Acad Sci U S A. 2007;104:11568–11573. doi: 10.1073/pnas.0705054104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Voskuil MI, Visconti KC, Schoolnik GK. Mycobacterium tuberculosis gene expression during adaptation to stationary phase and low-oxygen dormancy. Tuberculosis (Edinb) 2004;84:218–227. doi: 10.1016/j.tube.2004.02.003. [DOI] [PubMed] [Google Scholar]
- 39.Green J, Paget MS. Bacterial redox sensors. Nat Rev Microbiol. 2004;2:954–966. doi: 10.1038/nrmicro1022. [DOI] [PubMed] [Google Scholar]
- 40.Singh A, et al. Mycobacterium tuberculosis WhiB3 responds to O2 and nitric oxide via its [4Fe-4S] cluster and is essential for nutrient starvation survival. Proc Natl Acad Sci U S A. 2007;104:11562–11567. doi: 10.1073/pnas.0700490104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Crack JC, Stapleton MR, Green J, Thomson AJ, Le Brun NE. Mechanism of [4Fe-4S](Cys)4 cluster nitrosylation is conserved among NO-responsive regulators. J Biol Chem. 2013;288:11492–11502. doi: 10.1074/jbc.M112.439901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kiley PJ, Beinert H. Oxygen sensing by the global regulator, FNR: the role of the iron-sulfur cluster. FEMS Microbiol Rev. 1998;22:341–352. doi: 10.1111/j.1574-6976.1998.tb00375.x. [DOI] [PubMed] [Google Scholar]
- 43.Rohde KH, Veiga DF, Caldwell S, Balazsi G, Russell DG. Linking the transcriptional profiles and the physiological states of Mycobacterium tuberculosis during an extended intracellular infection. PLoS Pathog. 2012;8:e1002769. doi: 10.1371/journal.ppat.1002769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Gill WP, Harik NS, Whiddon MR, Liao RP, Mittler JE, Sherman DR. A replication clock for Mycobacterium tuberculosis. Nat Med. 2009;15:211–214. doi: 10.1038/nm.1915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Nicol MP, Wilkinson RJ. The clinical consequences of strain diversity in Mycobacterium tuberculosis. Trans R Soc Trop Med Hyg. 2008;102:955–965. doi: 10.1016/j.trstmh.2008.03.025. [DOI] [PubMed] [Google Scholar]
- 46.Homolka S, Niemann S, Russell DG, Rohde KH. Functional genetic diversity among Mycobacterium tuberculosis complex clinical isolates: delineation of conserved core and lineage-specific transcriptomes during intracellular survival. PLoS Pathog. 2010;6:e1000988. doi: 10.1371/journal.ppat.1000988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Golby P, et al. Comparative transcriptomics reveals key gene expression differences between the human and bovine pathogens of the Mycobacterium tuberculosis complex. Microbiology. 2007;153:3323–3336. doi: 10.1099/mic.0.2007/009894-0. [DOI] [PubMed] [Google Scholar]
- 48.Walters SB, Dubnau E, Kolesnikova I, Laval F, Daffe M, Smith I. The Mycobacterium tuberculosis PhoPR two-component system regulates genes essential for virulence and complex lipid biosynthesis. Mol Microbiol. 2006;60:312–330. doi: 10.1111/j.1365-2958.2006.05102.x. [DOI] [PubMed] [Google Scholar]
- 49.Abramovitch RB, Rohde KH, Hsu FF, Russell DG. aprABC: a Mycobacterium tuberculosis complex-specific locus that modulates pH-driven adaptation to the macrophage phagosome. Mol Microbiol. 2011;80:678–694. doi: 10.1111/j.1365-2958.2011.07601.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Vandal OH, Pierini LM, Schnappinger D, Nathan CF, Ehrt S. A membrane protein preserves intrabacterial pH in intraphagosomal Mycobacterium tuberculosis. Nat Med. 2008;14:849–854. doi: 10.1038/nmXXXX. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Faundez V, Hartzell HC. Intracellular chloride channels: determinants of function in the endosomal pathway. Sci STKE. 2004;2004:re8. doi: 10.1126/stke.2332004re8. [DOI] [PubMed] [Google Scholar]
- 52.Sonawane ND, Thiagarajah JR, Verkman AS. Chloride concentration in endosomes measured using a ratioable fluorescent Cl- indicator: evidence for chloride accumulation during acidification. J Biol Chem. 2002;277:5506–5513. doi: 10.1074/jbc.M110818200. [DOI] [PubMed] [Google Scholar]
- 53.Sonawane ND, Verkman AS. Determinants of [Cl-] in recycling and late endosomes and Golgi complex measured using fluorescent ligands. J Cell Biol. 2003;160:1129–1138. doi: 10.1083/jcb.200211098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Tan S, Sukumar N, Abramovitch RB, Parish T, Russell DG. Mycobacterium tuberculosis responds to chloride and pH as synergistic cues to the immune status of its host cell. PLoS Pathog. 2013;9:e1003282. doi: 10.1371/journal.ppat.1003282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Wagner D, et al. Elemental analysis of Mycobacterium avium-, Mycobacterium tuberculosis-, and Mycobacterium smegmatis-containing phagosomes indicates pathogen-induced microenvironments within the host cell’s endosomal system. J Immunol. 2005;174:1491–1500. doi: 10.4049/jimmunol.174.3.1491. [DOI] [PubMed] [Google Scholar]
- 56.Botella H, et al. Mycobacterial p(1)-type ATPases mediate resistance to zinc poisoning in human macrophages. Cell Host Microbe. 2011;10:248–259. doi: 10.1016/j.chom.2011.08.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Tailleux L, et al. Probing host pathogen cross-talk by transcriptional profiling of both Mycobacterium tuberculosis and infected human dendritic cells and macrophages. PLoS One. 2008;3:e1403. doi: 10.1371/journal.pone.0001403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Liu T, et al. CsoR is a novel Mycobacterium tuberculosis copper-sensing transcriptional regulator. Nat Chem Biol. 2007;3:60–68. doi: 10.1038/nchembio844. [DOI] [PubMed] [Google Scholar]
- 59.Ward SK, Abomoelak B, Hoye EA, Steinberg H, Talaat AM. CtpV: a putative copper exporter required for full virulence of Mycobacterium tuberculosis. Mol Microbiol. 2010;77:1096–1110. doi: 10.1111/j.1365-2958.2010.07273.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Wolschendorf F, et al. Copper resistance is essential for virulence of Mycobacterium tuberculosis. Proc Natl Acad Sci U S A. 2011;108:1621–1626. doi: 10.1073/pnas.1009261108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Maynes JT, et al. The crystal structure of 1-D-myo-inosityl 2-acetamido-2-deoxy-alpha-D-glucopyranoside deacetylase (MshB) from Mycobacterium tuberculosis reveals a zinc hydrolase with a lactate dehydrogenase fold. J Biol Chem. 2003;278:47166–47170. doi: 10.1074/jbc.M308914200. [DOI] [PubMed] [Google Scholar]
- 62.Spagnolo L, et al. Unique features of the sodC-encoded superoxide dismutase from Mycobacterium tuberculosis, a fully functional copper-containing enzyme lacking zinc in the active site. J Biol Chem. 2004;279:33447–33455. doi: 10.1074/jbc.M404699200. [DOI] [PubMed] [Google Scholar]
- 63.Cooper AM, Segal BH, Frank AA, Holland SM, Orme IM. Transient loss of resistance to pulmonary tuberculosis in p47(phox-/-) mice. Infect Immun. 2000;68:1231–1234. doi: 10.1128/iai.68.3.1231-1234.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Voskuil MI, Bartek IL, Visconti K, Schoolnik GK. The response of mycobacterium tuberculosis to reactive oxygen and nitrogen species. Front Microbiol. 2011;2:105. doi: 10.3389/fmicb.2011.00105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Piddington DL, Fang FC, Laessig T, Cooper AM, Orme IM, Buchmeier NA. Cu,Zn superoxide dismutase of Mycobacterium tuberculosis contributes to survival in activated macrophages that are generating an oxidative burst. Infect Immun. 2001;69:4980–4987. doi: 10.1128/IAI.69.8.4980-4987.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Wu CH, Tsai-Wu JJ, Huang YT, Lin CY, Lioua GG, Lee FJ. Identification and subcellular localization of a novel Cu,Zn superoxide dismutase of Mycobacterium tuberculosis. FEBS Lett. 1998;439:192–196. doi: 10.1016/s0014-5793(98)01373-8. [DOI] [PubMed] [Google Scholar]
- 67.Edwards KM, et al. Iron-cofactored superoxide dismutase inhibits host responses to Mycobacterium tuberculosis. Am J Respir Crit Care Med. 2001;164:2213–2219. doi: 10.1164/ajrccm.164.12.2106093. [DOI] [PubMed] [Google Scholar]
- 68.Bryk R, Lima CD, Erdjument-Bromage H, Tempst P, Nathan C. Metabolic enzymes of mycobacteria linked to antioxidant defense by a thioredoxin-like protein. Science. 2002;295:1073–1077. doi: 10.1126/science.1067798. [DOI] [PubMed] [Google Scholar]
- 69.Ng VH, Cox JS, Sousa AO, MacMicking JD, McKinney JD. Role of KatG catalase-peroxidase in mycobacterial pathogenesis: countering the phagocyte oxidative burst. Mol Microbiol. 2004;52:1291–1302. doi: 10.1111/j.1365-2958.2004.04078.x. [DOI] [PubMed] [Google Scholar]
- 70.Newton GL, Fahey RC. Mycothiol biochemistry. Arch Microbiol. 2002;178:388–394. doi: 10.1007/s00203-002-0469-4. [DOI] [PubMed] [Google Scholar]
- 71.Buchmeier NA, Newton GL, Koledin T, Fahey RC. Association of mycothiol with protection of Mycobacterium tuberculosis from toxic oxidants and antibiotics. Mol Microbiol. 2003;47:1723–1732. doi: 10.1046/j.1365-2958.2003.03416.x. [DOI] [PubMed] [Google Scholar]
- 72.Lee WL, et al. Mycobacterium tuberculosis expresses methionine sulphoxide reductases A and B that protect from killing by nitrite and hypochlorite. Mol Microbiol. 2009;71:583–593. doi: 10.1111/j.1365-2958.2008.06548.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Cooper AM, Dalton DK, Stewart TA, Griffin JP, Russell DG, Orme IM. Disseminated tuberculosis in interferon gamma gene-disrupted mice. J Exp Med. 1993;178:2243–2247. doi: 10.1084/jem.178.6.2243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Flynn JL, Chan J, Triebold KJ, Dalton DK, Stewart TA, Bloom BR. An essential role for interferon gamma in resistance to Mycobacterium tuberculosis infection. J Exp Med. 1993;178:2249–2254. doi: 10.1084/jem.178.6.2249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Mattila JT, et al. Microenvironments in tuberculous granulomas are delineated by distinct populations of macrophage subsets and expression of nitric oxide synthase and arginase isoforms. J Immunol. 2013;191:773–784. doi: 10.4049/jimmunol.1300113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Nicholson S, et al. Inducible nitric oxide synthase in pulmonary alveolar macrophages from patients with tuberculosis. J Exp Med. 1996;183:2293–2302. doi: 10.1084/jem.183.5.2293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Choi HS, Rai PR, Chu HW, Cool C, Chan ED. Analysis of nitric oxide synthase and nitrotyrosine expression in human pulmonary tuberculosis. Am J Respir Crit Care Med. 2002;166:178–186. doi: 10.1164/rccm.2201023. [DOI] [PubMed] [Google Scholar]
- 78.Garbe TR, Hibler NS, Deretic V. Response to reactive nitrogen intermediates in Mycobacterium tuberculosis: induction of the 16-kilodalton alpha-crystallin homolog by exposure to nitric oxide donors. Infect Immun. 1999;67:460–465. doi: 10.1128/iai.67.1.460-465.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Darwin KH, Ehrt S, Gutierrez-Ramos JC, Weich N, Nathan CF. The proteasome of Mycobacterium tuberculosis is required for resistance to nitric oxide. Science. 2003;302:1963–1966. doi: 10.1126/science.1091176. [DOI] [PubMed] [Google Scholar]
- 80.Darwin KH, Lin G, Chen Z, Li H, Nathan CF. Characterization of a Mycobacterium tuberculosis proteasomal ATPase homologue. Mol Microbiol. 2005;55:561–571. doi: 10.1111/j.1365-2958.2004.04403.x. [DOI] [PubMed] [Google Scholar]
- 81.McKinney JD, et al. Persistence of Mycobacterium tuberculosis in macrophages and mice requires the glyoxylate shunt enzyme isocitrate lyase. Nature. 2000;406:735–738. doi: 10.1038/35021074. [DOI] [PubMed] [Google Scholar]
- 82.Sassetti CM, Rubin EJ. Genetic requirements for mycobacterial survival during infection. Proc Natl Acad Sci U S A. 2003;100:12989–12994. doi: 10.1073/pnas.2134250100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Pandey AK, Sassetti CM. Mycobacterial persistence requires the utilization of host cholesterol. Proc Natl Acad Sci U S A. 2008;105:4376–4380. doi: 10.1073/pnas.0711159105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Casabon I, Zhu SH, Otani H, Liu J, Mohn WW, Eltis LD. Regulation of the KstR2 regulon of Mycobacterium tuberculosis by a cholesterol catabolite. Mol Microbiol. 2013;89:1201–1212. doi: 10.1111/mmi.12340. [DOI] [PubMed] [Google Scholar]
- 85.Klink M, et al. Cholesterol oxidase is indispensable in the pathogenesis of Mycobacterium tuberculosis. PLoS One. 2013;8:e73333. doi: 10.1371/journal.pone.0073333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Miner MD, Chang JC, Pandey AK, Sassetti CM, Sherman DR. Role of cholesterol in Mycobacterium tuberculosis infection. Indian journal of experimental biology. 2009;47:407–411. [PubMed] [Google Scholar]
- 87.Van der Geize R, et al. A gene cluster encoding cholesterol catabolism in a soil actinomycete provides insight into Mycobacterium tuberculosis survival in macrophages. Proc Natl Acad Sci U S A. 2007;104:1947–1952. doi: 10.1073/pnas.0605728104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Wipperman MF, Sampson NS, Thomas ST. Pathogen roid rage: cholesterol utilization by Mycobacterium tuberculosis. Critical reviews in biochemistry and molecular biology. 2014;49:269–293. doi: 10.3109/10409238.2014.895700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Savvi S, Warner DF, Kana BD, McKinney JD, Mizrahi V, Dawes SS. Functional characterization of a vitamin B12-dependent methylmalonyl pathway in Mycobacterium tuberculosis: implications for propionate metabolism during growth on fatty acids. J Bacteriol. 2008;190:3886–3895. doi: 10.1128/JB.01767-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Gould TA, van de Langemheen H, Munoz-Elias EJ, McKinney JD, Sacchettini JC. Dual role of isocitrate lyase 1 in the glyoxylate and methylcitrate cycles in Mycobacterium tuberculosis. Mol Microbiol. 2006;61:940–947. doi: 10.1111/j.1365-2958.2006.05297.x. [DOI] [PubMed] [Google Scholar]
- 91.Munoz-Elias EJ, McKinney JD. Mycobacterium tuberculosis isocitrate lyases 1 and 2 are jointly required for in vivo growth and virulence. Nat Med. 2005;11:638–644. doi: 10.1038/nm1252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Munoz-Elias EJ, Upton AM, Cherian J, McKinney JD. Role of the methylcitrate cycle in Mycobacterium tuberculosis metabolism, intracellular growth, and virulence. Mol Microbiol. 2006;60:1109–1122. doi: 10.1111/j.1365-2958.2006.05155.x. [DOI] [PubMed] [Google Scholar]
- 93.Lee W, VanderVen BC, Fahey RJ, Russell DG. Intracellular Mycobacterium tuberculosis exploits host-derived fatty acids to limit metabolic stress. J Biol Chem. 2013;288:6788–6800. doi: 10.1074/jbc.M112.445056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Kim MJ, et al. Caseation of human tuberculosis granulomas correlates with elevated host lipid metabolism. EMBO molecular medicine. 2010;2:258–274. doi: 10.1002/emmm.201000079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Peyron P, et al. Foamy macrophages from tuberculous patients’ granulomas constitute a nutrient-rich reservoir for M. tuberculosis persistence. PLoS Pathog. 2008;4:e1000204. doi: 10.1371/journal.ppat.1000204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Russell DG, Cardona PJ, Kim MJ, Allain S, Altare F. Foamy macrophages and the progression of the human tuberculosis granuloma. Nature immunology. 2009;10:943–948. doi: 10.1038/ni.1781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Flynn JL, Chan J, Lin PL. Macrophages and control of granulomatous inflammation in tuberculosis. Mucosal immunology. 2011;4:271–278. doi: 10.1038/mi.2011.14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Rhoades ER, Geisel RE, Butcher BA, McDonough S, Russell DG. Cell wall lipids from Mycobacterium bovis BCG are inflammatory when inoculated within a gel matrix: characterization of a new model of the granulomatous response to mycobacterial components. Tuberculosis (Edinb) 2005;85:159–176. doi: 10.1016/j.tube.2004.10.001. [DOI] [PubMed] [Google Scholar]
- 99.Bowdish DM, et al. MARCO, TLR2, and CD14 are required for macrophage cytokine responses to mycobacterial trehalose dimycolate and Mycobacterium tuberculosis. PLoS Pathog. 2009;5:e1000474. doi: 10.1371/journal.ppat.1000474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Ishikawa E, et al. Direct recognition of the mycobacterial glycolipid, trehalose dimycolate, by C-type lectin Mincle. J Exp Med. 2009;206:2879–2888. doi: 10.1084/jem.20091750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Beatty WL, Rhoades ER, Ullrich HJ, Chatterjee D, Heuser JE, Russell DG. Trafficking and release of mycobacterial lipids from infected macrophages. Traffic. 2000;1:235–247. doi: 10.1034/j.1600-0854.2000.010306.x. [DOI] [PubMed] [Google Scholar]
- 102.Rhoades E, Hsu F, Torrelles JB, Turk J, Chatterjee D, Russell DG. Identification and macrophage-activating activity of glycolipids released from intracellular Mycobacterium bovis BCG. Mol Microbiol. 2003;48:875–888. doi: 10.1046/j.1365-2958.2003.03473.x. [DOI] [PubMed] [Google Scholar]
- 103.Baldan A, Bojanic DD, Edwards PA. The ABCs of sterol transport. Journal of lipid research. 2009;50(Suppl):S80–85. doi: 10.1194/jlr.R800044-JLR200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Hutchins PM, Murphy RC. Cholesteryl ester acyl oxidation and remodeling in murine macrophages: formation of oxidized phosphatidylcholine. Journal of lipid research. 2012;53:1588–1597. doi: 10.1194/jlr.M026799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Garton NJ, et al. Cytological and transcript analyses reveal fat and lazy persister-like bacilli in tuberculous sputum. PLoS medicine. 2008;5:e75. doi: 10.1371/journal.pmed.0050075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Miesenbock G, De Angelis DA, Rothman JE. Visualizing secretion and synaptic transmission with pH-sensitive green fluorescent proteins. Nature. 1998;394:192–195. doi: 10.1038/28190. [DOI] [PubMed] [Google Scholar]
- 107.Hanson GT, et al. Investigating mitochondrial redox potential with redox-sensitive green fluorescent protein indicators. J Biol Chem. 2004;279:13044–13053. doi: 10.1074/jbc.M312846200. [DOI] [PubMed] [Google Scholar]
- 108.Bhaskar A, et al. Reengineering redox sensitive GFP to measure mycothiol redox potential of Mycobacterium tuberculosis during infection. PLoS Pathog. 2014;10:e1003902. doi: 10.1371/journal.ppat.1003902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.North RJ, Jung YJ. Immunity to tuberculosis. Annu Rev Immunol. 2004;22:599–623. doi: 10.1146/annurev.immunol.22.012703.104635. [DOI] [PubMed] [Google Scholar]
- 110.Meniche X, et al. Subpolar addition of new cell wall is directed by DivIVA in mycobacteria. Proc Natl Acad Sci U S A. 2014;111:E3243–3251. doi: 10.1073/pnas.1402158111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Wakamoto Y, et al. Dynamic persistence of antibiotic-stressed mycobacteria. Science. 2013;339:91–95. doi: 10.1126/science.1229858. [DOI] [PubMed] [Google Scholar]
- 112.Sukumar N, Tan S, Aldridge BB, Russell DG. Exploitation of Mycobacterium tuberculosis reporter strains to probe the impact of vaccination at sites of infection. PLoS Pathog. 2014 doi: 10.1371/journal.ppat.1004394. In press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Reyes-Lamothe R, Possoz C, Danilova O, Sherratt DJ. Independent positioning and action of Escherichia coli replisomes in live cells. Cell. 2008;133:90–102. doi: 10.1016/j.cell.2008.01.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Costes A, Lecointe F, McGovern S, Quevillon-Cheruel S, Polard P. The C-terminal domain of the bacterial SSB protein acts as a DNA maintenance hub at active chromosome replication forks. PLoS Genet. 2010;6:e1001238. doi: 10.1371/journal.pgen.1001238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Berkmen MB, Grossman AD. Spatial and temporal organization of the Bacillus subtilis replication cycle. Mol Microbiol. 2006;62:57–71. doi: 10.1111/j.1365-2958.2006.05356.x. [DOI] [PubMed] [Google Scholar]
- 116.Lin PL, et al. Sterilization of granulomas is common in active and latent tuberculosis despite within-host variability in bacterial killing. Nat Med. 2013;20:75–79. doi: 10.1038/nm.3412. [DOI] [PMC free article] [PubMed] [Google Scholar]