Summary
Programmed ribosomal frameshifting produces alternative proteins from a single transcript. -1-frameshifting occurs on Escherichia coli’s dnaX mRNA containing a slippery sequence AAAAAAG and peripheral mRNA structural barriers. Here we reveal hidden aspects of the frameshifting process, including its exact location on the mRNA and its timing within the translation cycle. Mass spectrometry of translated products shows that ribosomes enter the -1-frame from not one specific codon but various codons along the slippery sequence and slip by not just -1 but also -4, or +2-nucleotides. Single-ribosome translation trajectories detect distinctive codon-scale fluctuations in ribosome-mRNA displacement across the slippery sequence, representing multiple ribosomal translocation attempts during frameshifting. Flanking mRNA structural barriers mechanically stimulate the ribosome to undergo back-and-forth translocation excursions, broadly exploring reading frames. Both experiments reveal aborted translation around mutant slippery sequences, indicating that subsequent fidelity checks on newly adopted codon position base-pairings lead to either resumed translation or early termination.
Keywords: programmed frameshifting, ribosome, translocation, translation fidelity, optical tweezers, mass spectrometry
Introduction
During translation, the ribosome successively reads three nucleotides—one codon—at a time to produce the protein encoded in the messenger RNA (mRNA). This process involves base-pairing each codon with the anticodon of the cognate aminoacylated transfer RNA (aa-tRNA). As a result, each mRNA sequence specifies one unique polypeptide translated from start- to stop-codon in the so-called 0-frame. Crystals of such ribosomes in the decoding mode—complexed with mRNA and two tRNAs—have provided a detailed structural basis for reading frame maintenance (Jenner et al., 2010; Schmeing and Ramakrishnan, 2009; Selmer et al., 2006; Stahl et al., 2002; Yusupov et al., 2001). Specifically, the mRNA wrapping around the neck of the ribosome small subunit is kinked into segments, with the three consecutive codons positioned in the exit- (E), peptidyl- (P), and aminoacyl- (A) codon:anticodon binding sites. The anticodon stem-loop and the aminoacyl-acceptor end of each tRNA are accommodated on the small (30S) and large (50S) subunits, respectively, corresponding to the classical-state P/P- and A/A-tRNAs (capital letters denote sites on 30S/50S). After peptidyl-transfer between the tRNAs, the ribosome facilitates the two tRNAs to adopt hybrid states in which the anticodons remain in the 30S P- and A-sites, but the acceptor ends have advanced to the E- and P-sites on the 50S, denoted as P/E- and A/P-tRNAs (Bretscher, 1968; Moazed and Noller, 1989). Upon binding elongation factor EF-G in the A-site, the ribosome can proceed to translocate one codon forward (Rodnina and Wintermeyer, 2011; Savelsbergh et al., 2003). The strict reading frame configuration and the segregation of decoding and translocation help keep the translation accurate with an error rate of less than 0.1% (Drummond and Wilke, 2009).
However, ribosomes can be programmed to frameshift—accessing either of the two out-of-frames (-1 or +1-frame), thereby expanding gene coding capacity on a single transcript (Farabaugh, 1996). Such a mechanism is essential to the virulence of compact genomic systems such as HIV-1, where successive frameshifts occur on the mRNA to produce a retroviral polyprotein (Jacks et al., 1988).
Here we investigate frameshift-programming mRNAs derived from the Escherichia coli dnaX gene. Its -1-frameshift efficiency in vivo reaches 80%, yielding a 4:1 product ratio between the γ subunit and the τ subunit of DNA polymerase III (Tsuchihashi and Brown, 1992). Such translation regulation is achieved by three sequence elements in the mRNA: a heptanucleotide slippery sequence AAAAAAG that is flanked by an internal Shine-Dalgarno (SD) sequence located 10 nucleotides (nt) upstream, and an 11-basepair (bp) hairpin 6 nt downstream (Figure 1A). It has long been thought that while the ribosome decodes the 0-frame codons in the slippery sequence (A_AAA_AAG), the peripheral base-pairing structures on the mRNA serve as barriers to impede normal translation and promote backward frameshifting by 1 nt (Figure 1A) (Gesteland and Atkins, 1996). Specifically, the upstream SD sequence hybridizes with the complementary anti-SD sequence at the 3’-end of 16S ribosomal RNA (rRNA), thus forming a flexible yet mRNA-anchoring mini-helix (Jenner et al., 2007; Kaminishi et al., 2007; Korostelev et al., 2007; Yusupova et al., 2006). Downstream, the base-pairing junction of the hairpin acts as a roadblock situated at the mRNA entry site on the ribosome—a single-strand-permitting channel formed by three ribosomal proteins: S3 on the 30S head, and S4 and S5 from the 30S body (Yusupova et al., 2001).
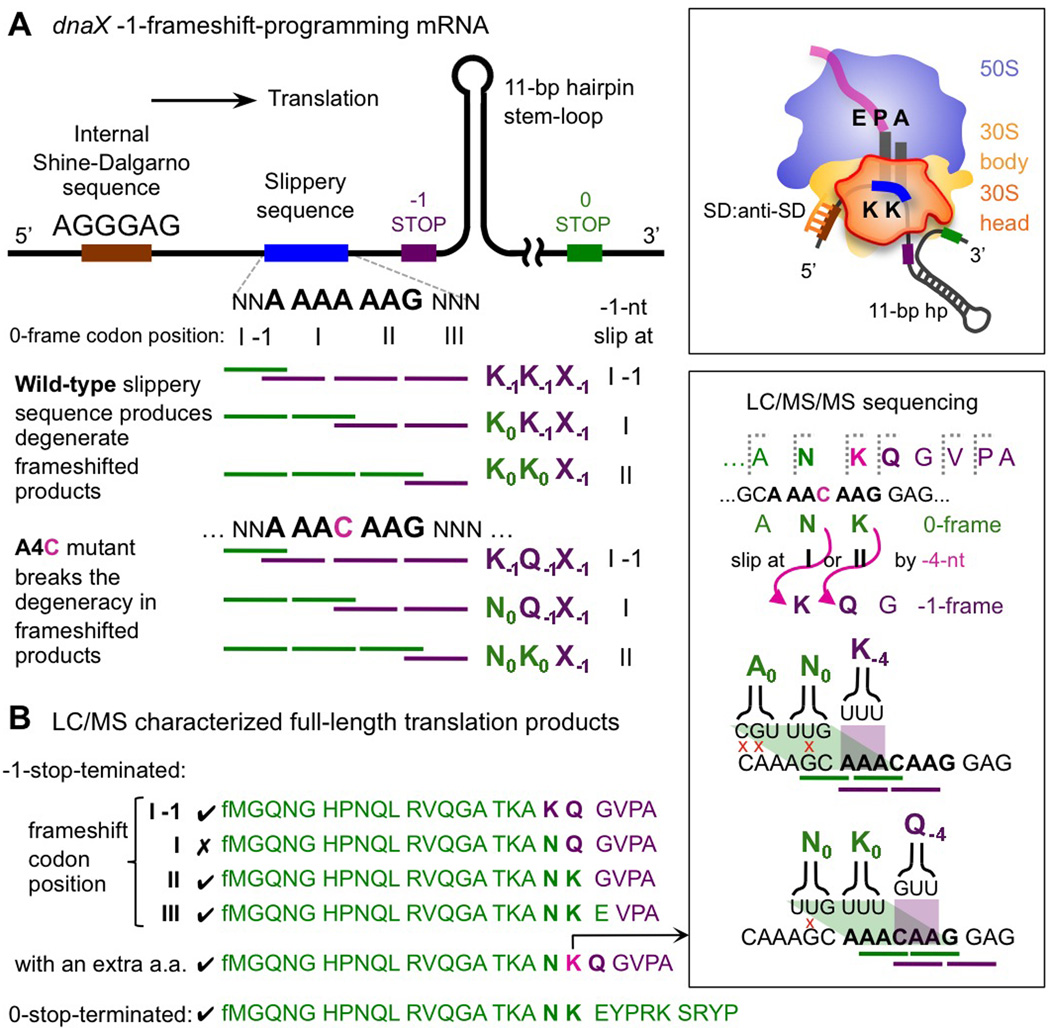
Figure 1. Resolving ribosomal frameshifting codon positions on dnaX-derived mRNAs.
1A. Three mRNA sequence elements program the -1-nt ribosomal frameshift: a slippery sequence, AAAAAAG (region in blue); an internal Shine-Dalgarno sequence (SD, region in brown); and a downstream hairpin. The cartoon shows the position of these elements on the mRNA relative to the ribosome. Exact frameshift codon positions are indistinguishable due to the identical product sequence. A single mutation A4C in the slippery sequence (I and II denotes the two 0-frame codons) differentiates possible frameshift positions.
1B. Various -1-stop-terminated products (sequences ended in purple) from the A4C mutant, detected by liquid chromatography-mass spectrometry (LC/MS), show that ribosomes frameshift from different codons around the slippery sequence, including position I-1, II, and III. One major frameshifted product, sequenced by tandem mass spectrometry (LC/MS/MS), bears an extra amino acid in the slippery sequence region (Figure S1D and Table S2); thus, the ribosome has slipped by -4-nt to enter the -1-frame. Two degenerate frameshift pathways exist to translate such a product (right box): -4-slip at codon position I or II (green-shaded rhombus area); the latter imposes fewer codon:anticodon base-pair mismatches (red crosses).
Recently, perturbed ribosome translation dynamics on the slippery sequence has been confirmed and visualized in single-molecule fluorescence FRET experiments (Chen et al., 2014; Kim et al., 2014). However, exact details on how the programmed mRNA elements act on the ribosome to induce frameshifting dynamics remain unclear (Tinoco et al., 2013). For example, from which 0-frame codon does the ribosome frameshift? In which sub-step within the translation cycle does the frameshift take place? How does the frameshift-programming mRNA break the regular ribosome translation stepping—3 nt per codon—to promote efficient and apparently precise frameshifting? Answering these questions requires looking beyond ribosome conformational dynamics; we thus sought to examine the ribosome translation dynamics on the mRNA, and to characterize the synthesized polypeptides.
Here we use mass spectrometry (MS) to analyze the products of translation along dnaX-derived mRNAs containing wild-type or mutant slippery sequences. These analyses reveal that the ribosome can access a broad range of frameshift pathways—shifting from different codon positions and using various slipping sizes. We complement these studies by acquiring single-ribosome translation trajectories using optical tweezers (Wen et al., 2008) to follow in real-time the ribosome dynamics that accompanies the exploration of alternative frameshift pathways. These trajectories display distinctive fluctuations—larger than 1 nucleotide—in mRNA displacement during translocation as the ribosome attempts to overcome the mRNA structural barriers flanking the slippery sequence. We found that, after this dynamic exploration, the ribosome may frameshift, but it is sensitive to mismatches that result from the pairing between the frameshifted codons and anticodons. These mismatches likely trigger a fidelity check mechanism that results in the ribosome either to continue translation in a new frame or to prematurely abort translation.
Results
Frameshifts occur at various codon positions
Ribosomes are thought to back-shift on the mRNA slippery sequence—AAAAAAG for the dnaX gene—and translate to the -1-stop codon (Figure 1A) (Farabaugh, 1996). This conjecture is based on the fact that, here, both the 0-frame (A_AAA_AAG) and -1-frame (AAA_AAA_G) encode identical amino acids: a pair of lysines. Hence, a -1-nt slippage in this region involves minimal base-pairing difference between the lysine codons, AAA and AAG, and the UUU anticodon used in E. coli (Tsuchihashi and Brown, 1992). Note however that the resultant tandem lysines incorporated in the frameshifted product preclude identifying from the polypeptide sequence where exactly the 0-frame ends. As shown in Figure 1A, on the dnaX slippery sequence, there are three possible decoding routes, all translating the same amino acids but via a -1-frameshift at different codon positions. To differentiate those potential decoding routes, we introduced a slippery sequence variant: A_AAC_AAG, which retains ~5% frameshift efficiency in vivo (Tsuchihashi and Brown, 1992). The single mutation of the fourth adenine to a cytosine lifts the encoding degeneracy and yields different amino acid compositions depending on the last-read 0-frame codon position (Figure 1A, A4C mutant; the two 0-frame codons in the slippery sequence are denoted as positions I and II). Like the original dnaX mRNA, the mRNA variant contains an upstream internal SD sequence and a downstream 25-bp duplex equivalently 6 nt downstream from the slippery sequence (Figure S1A); we denote this design as the “25-bp” mRNA construct. To generate samples for frameshifting analysis, in vitro transcribed and gel-purified mRNAs, together with E. coli 70S ribosomes, were added to a reconstituted translation mixture (PURExpress® ΔRibosome kit, NEB) (Ohashi et al., 2010). The in vitro translation products were then collected and examined using liquid chromatography/mass spectrometry (LC/MS) intact polypeptide detection (Extended Experimental Procedures).
In addition to the non-frameshifted, 0-stop-terminated polypeptide (Figure 1B, bottom-most sequence in green), multiple frameshifted products terminating at the -1-stop were identified (Figure 1B; the last green-colored residue of each sequence show the last read 0-frame codon—from which the ribosome frameshifts). We found that ribosomes take two of the three possible -1-frameshift decoding routes (at codon position I-1 and II), respectively incorporating K−1Q−1 or N0K0 from the slippery sequence (subscripts denote the frame); or they switch to the -1-frame after the slippery sequence at codon position III (Figure 1B, fourth sequence from the top). The frameshifted polypeptide via slipping at codon position I was, however, not observed (Figure S1B). Whereas a recent search for -1-frameshifted products was limited to two codon positions on the slippery sequence of HIV-1 (Liao et al., 2011), here we observed that frameshifts in fact emerge from at least three positions.
We explored two more mRNA templates, bearing either the original frameshift-promoting slippery sequence, AAAAAAG (wild-type/25-bp; ~80% frameshift efficiency in vivo), or a frameshift-attenuating variant, AAAAGAG (A5G/25-bp; ~0% frameshift efficiency in vivo) (Tsuchihashi and Brown, 1992). In all templates examined, independently of the frameshift efficiencies attained, the ribosome undergoes -1-frameshifts from a broad range of codon positions spanning regions before, within, and beyond the slippery sequence (Table S1).
Ribosomes frameshift via various slip sizes
Intriguingly, one polypeptide ~100 Da heavier than the other identified -1-frameshifted products was consistently detected in the mass spectrum (N = 5) as a major species translated from the A4C mutant mRNA (Figure 2A top box, largest red bar: ~74% of all -1-stop-terminated products detected). To determine the sequence of this unexpected product, we employed tandem mass spectrometry (LC/MS/MS) to select and fragment the polypeptide (Figure 1B right box; Table S2). We found that not two but three amino acids were incorporated along the slippery sequence for this unusual -1-stop-terminated frameshifted product. To translate an extra amino acid while switching to the -1-frame the ribosome must slip by -4-nt during frameshifting.
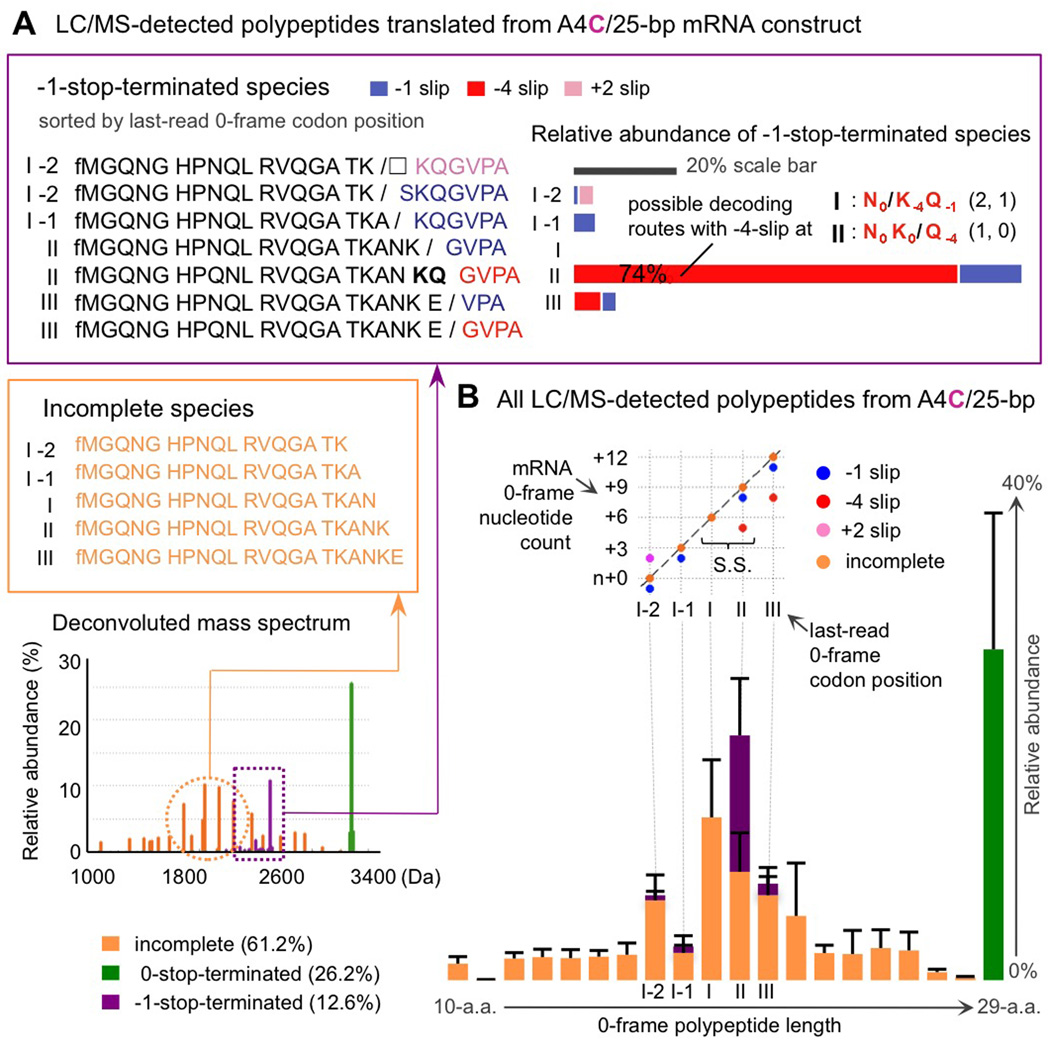
Figure 2. In addition to various stop-codon-terminated polypeptides, frameshift-programming mRNAs produce incomplete species.
2A. A4C slippery sequence variant (construct with a downstream 25-bp duplex) as an example: LC/MS detected a broad collection of -1-stop-terminated products frameshifted from codon positions around the slippery sequence; the top bar graph shows their relative abundance (x-axis). These frameshifted species were translated via -1-slips (blue), -4-slips (red), and +2-slips (pink); the latter two lead to polypeptides one amino acid longer or shorter (Figure S2). When degenerate decoding routes exist (as those shown in Figure 1B right box; numbers of base-pair mismatches for the last two 0-frame tRNAs are tabulated here in parenthesis; every non Watson-Crick base-pair scores a 1), we assigned the given product to frameshift codon positions with fewer mismatches. Incomplete polypeptides ended with 0-frame amino acids along the slippery sequence were also found (sequences in orange; orange peaks in the mass spectrum; Table S1).
2B. Bottom bar graph: all detected species in the MS spectrum are organized based on their last 0-frame amino acid incorporated, i.e. 0-frame polypeptide length. A 2D diagram, focusing on codon positions around the slippery sequence (S.S.) region, displays from where (x-axis) the ribosome frameshifts or leaves behind incomplete species. With the y-axis listing the mRNA nucleotide counts in reference to the 0-frame, incomplete species (orange dots) lie along the diagonal line; the frameshifted products distribute above and below—as located by the first nucleotide read in the -1-frame on the mRNA.
The resolved three amino acids, NKQ, can be translated via two possible decoding routes: a -4-slip either at codon position I or at position II on the slippery sequence (Figure 1B right box). In the first route the 0-frame ends at asparagine to yield N0K−1Q−1; thus, the last two 0-frame-specified tRNAs carrying the alanine and asparagine—after back-shifting on the mRNA by 4-nt—would encounter three mismatches (red crosses). In contrast, the second route would cause only one mismatch for the tRNAAsn, suggesting that this route involving a -4-slip from codon position II could be the more productive frameshift pathway.
For the other two templates (wild-type/25-bp and A5G/25-bp), several, though less abundant, frameshifted products bearing an extra amino acid were also detected (Figure S2A and Table S1; relative abundance ~5% and ~3% of overall -1-stop-terminated products). These species point to the general capability of the ribosome to conduct -4-nt slips on -1-frameshift-programming mRNAs. The presence of -4-slip products led us to expand the search for alternative slipping sizes entering the -1-frame (our template design is capable of detecting only slips into the -1-frame and thus precludes readout of potential +1-frameshifting; see Figure S1A). We found that ribosomes also take +2–slips and terminate at the -1-stop, producing frameshifted polypeptides one amino acid short (Figure 2A top box, sequences ended in pink; relative abundance ~3% for A4C).
Figure 2A top box summarizes the relative abundance of all -1-stop-terminated products detected by LC/MS for the A4C mutant template (wild-type and A5G are in Table S1; see Extended Experimental Procedures for explanation of abundance measurements). These findings show that ribosomes slip by -1, -4, or +2 nt at various codon positions around the slippery sequence region, producing a collection of -1-stop-terminated products. While an earlier work reported that specific slipping sizes, e.g. -2, -1, +2, +5, and +6 nt, can be individually programmed by different mRNA templates (Weiss et al., 1987), our study shows that various slipping sizes take place on a single naturally-occurring template. Note that the MS-resolved frameshifted polypeptides reported here would have appeared as a single protein band on electrophoresis gels, therefore being indistinguishable in earlier studies (Tsuchihashi and Brown, 1992).
The -1-stop and 0-stop-terminated full-length products, however, only account for a fraction of the polypeptides found in the mass spectrum, e.g., ~39% of the total intensity of all species detected from the A4C template (Figure 2A). We found the rest to be incomplete polypeptides ended at 0-frame codon positions around the slippery sequence, and particularly accumulating at positions where the ribosome tends to frameshift (Figure 2A left orange box). We hence sorted full-length and incomplete species by their last-incorporated 0-frame amino acids and rearranged the purely mass-based LC/MS spectrum into a bar graph ordered by 0-frame polypeptide length (i.e. x-axis of the bottom graph in Figure 2B); the bar heights depict the relative abundance of products detected (standard deviations from multiple measurements shown as error bars).
To provide a comprehensive view for the various ribosomal frameshifting translation events observed around the slippery sequence region, we construct a 2D diagram (Figure 2B upper-part) to visualize from which 0-frame codon and to which -1-frame codon the ribosome slips. Specifically, the x-axis marks the last-read 0-frame codon position along the mRNA, while the y-axis—counting mRNA nucleotides in the 0-frame by multiples of 3—indicates the first nucleotide read in the -1-frame. Therefore, incomplete species (orange dots) drop off along the diagonal, whereas -1-stop-terminated products frameshifted via plus slips (pink dots) and minus slips (blue and red dots) distribute above and below.
Ribosomes make distinctive translocation attempts
Having detected such diverse frameshift pathways via translation product analysis, we sought to unravel the molecular mechanisms that give rise to the broad range of ribosomal slippage observed. Specifically, how does the ribosome switch the mRNA reading frame—i.e. allowing the tRNAs to slip and to simultaneously base-pair across adjacent codons which are spatially kinked into nucleotide triplets by the intercalating 16S rRNA residues inside the 30S decoding groove (Yusupova et al., 2001)? This issue can be addressed by determining when—and how—within one translation cycle the ribosome frameshifts. To this end, we employed a real-time in vitro mRNA hairpin unwinding assay. Using optical tweezers we monitored codon-by-codon translation by a single ribosome along the entire frameshift-programming mRNA template embedded inside a 92-bp long hairpin (Figure 3; Figure S3A) (Qu et al., 2011; Wen et al., 2008).
Figure 3. Probing ribosomal frameshifting translation translocation dynamics using optical tweezers.
A single-ribosome translation progression is reported by the step-wise unwinding of a 92-bp mRNA hairpin held on the optical tweezers (see also Experimental Procedures); 3 bp are unzipped per codon translocated at the hairpin junction, thus reflecting displacements between the ribosome and mRNA. When the first 0-frame codon in the slippery sequence (codon position I) resides in the ribosome 30S P-site, a 55-bp hairpin remains downstream. Hairpin portions not unwound by the ribosome were measured at the end of experiments; if the ribosome terminates at the -1-stop, it leaves a smaller residual hairpin, as compared to that for the 0-stop termination (Figure S3B). Both the wild-type slippery sequence and the frameshift-attenuating A5G mutant were examined.
Here, an mRNA hairpin molecule—bound with a single ribosome—is tethered through its two ends and held under tension by the optical tweezers (Figure 3). As the ribosome gradually translates the mRNA, it must unzip the hairpin by 3-bp per codon. As a result, the tether end-to-end distance extends by 6 nt (i.e. the gridline spacing: ~2.65 nm/codon on the left y-axis in Figure 4A and 5A trajectory plots), thus reporting the translocation movement of the ribosome from one codon to the next. As seen from the trajectory, translation occurs in alternating phases of translocations and dwells (seen as vertical extensions and horizontal segments). During each dwell, the ribosome decodes the A-site codon and catalyzes peptidyl-transfer between the P- and A-tRNAs (Rodnina et al., 2005; Wohlgemuth et al., 2008). The ribosome subsequently binds the GTPase EF-G, partially displacing the P- and A-tRNAs into the 50S E- and P-sites, and proceeds with mRNA forward translocation, which requires unzipping the transcript downstream (Qu et al., 2011; Rodnina and Wintermeyer, 2011; Savelsbergh et al., 2003; Wen et al., 2008). To confirm whether the ribosome terminated at the 0- or -1-frame stop, after translation ceased we applied force to unfold the remaining mRNA hairpin. Since the two stop codons result in different untranslated residual hairpin sizes—as measured by the mRNA extension gained from unfolding—we could verify where the ribosome ended in each trajectory (shown schematically in Figure S3B).
Figure 4. Characteristic fluctuations during ribosome translocation across the slippery sequence.
4A. A single-ribosome translation trajectory along the frameshift-promoting wild-type slippery sequence; recorded at 1 kHz and displayed at 20 Hz here. Upon each translocation step taken by the ribosome (vertical advances along y-axis, indicated by black arrow heads), the hairpin releases 6 nt per codon; this is seen as a 2.65-nm increment (spacing between gridlines of the same color) in mRNA end-to-end extension under a tension of 18 pN (Extended Experimental Procedures). Given the mRNA template, amino acids incorporated to the P-site tRNA after each translocation step are labeled next to the gridlines (in letter codes; green for 0-frame, purple for -1-frame). While the ribosome continually translocates against a hairpin, characteristic fluctuations in mRNA extension (zoom-in below) occur downstream from the internal SD sequence around the slippery sequence region (orange-shaded area; Figure S4B).
4B. The characteristic fluctuations were seen for both wild-type and A5G slippery sequence variants—and both in frameshifted and non-frameshifted trajectories, with an amplitude ≥1-codon (magenta double-headed arrow on the zoom-in trace in panel A) and an average excursion lifetime of ~0.5 second for one round of back-and-forth fluctuation (horizontal line segments in magenta; see also Figure S4C; N ≥ 10 trajectories analysed for each of the four categories).
Figure 5. Connecting frameshift translation dynamics and product distribution.
5A. The ≥1-codon translocation fluctuations (black-squared section on the blue trace; expanded underneath) persist in translation trajectory along the frameshift-attenuating A5G mutant, occurring around the slippery sequence (orange-shaded area). Meanwhile, LC/MS detected a wide range of frameshift translation species produced from the same A5G/55-bp construct, including frameshifted and incomplete species (purple and orange dots in 2D diagram; x-axis showing last read 0-frame codons, y-axis marking first read nucleotides in the -1-frame, relative to those counted in the 0-frame; Figure S5B). The accumulation of frameshifted and incomplete species at codon position II (the column of purple and orange dots indicated by red arrow), coincide with the locations on the mRNA slippery sequence region that were frequently explored by the back-and-forth fluctuating ribosome—as revealed by the trajectory zoom-in section.
5B. Relative abundances of LC/MS-detected translation products from the 55-bp mRNA constructs—each for the A5G, A4C, and wild-type slippery sequence variants representing low, medium, and high frameshift efficiency—are shown in bar graphs. Products are sorted by their last 0-frame amino acids incorporated along the mRNA (x-axis: increasing 0-frame polypeptide length), and their abundances shown in bar graphs (Table S1). Less efficient slippery sequence variants produce higher amount of incomplete species, particularly at codon positions I and II—from where most frameshifted species were also translated (purple bars). Numbers of base-pair mismatches for the frameshifted E- and P-tRNAs are tabulated in parenthesis for frameshift pathways at codon position I and II, via -1 or -4-slips (in subscripts). We count 1 for every non Watson-Crick interaction.
A ~55-bp hairpin remains ahead as the ribosome resides on the slippery sequence (Figure S3A) downstream from the internal SD sequence (with the first codon, i.e. position I, of the slippery sequence in the P-site) (Qu et al., 2011). We chose a “55-bp construct” for the tweezers experiments, instead of the previously discussed 25-bp construct, because the longer—and thus more stable—hairpin allows more accurate measurements of the termination codon positions. Both frameshift-promoting (wild-type/55-bp) and frameshift-attenuating (A5G/55-bp) slippery sequence variants were translated on the tweezers. The frameshift efficiencies for these two templates were 77% and 57%, respectively, showing a trend similar to that observed in vivo (Table 1) (Tsuchihashi and Brown, 1992).
Table 1. Apparent frameshift efficiency and overall frameshift slipping attempts.
In addition to the conventionally defined frameshift efficiency—accounting only the -1-stop and 0-stop-terminated products (row 1 (Tsuchihashi and Brown, 1992), 2 and 5), the overall slipping attempts made by the ribosome are estimated by including the incomplete, i.e. drop-off, species, or equivalently the prematurely stalled and aborted translation trajectories (row 3 and 6). For the two template variants examined (wild-type and A5G), the most probable aborted codon positions are consistent with the most probable drop-off codon positions resolved by LC/MS. Some differences in frameshift efficiency are seen between in vivo and in vitro translation conditions, which we attribute to known differences in overall translation rates.
 |
Significantly, ~90% of the trajectories exhibit distinct fluctuations in mRNA extension specifically around the slippery sequence region (orange-shaded area in Figure 4A and 5A; trajectories recorded at 1kHz and displayed at 20 Hz). These unique signals manifest back-and-forth movements of the ribosome on the mRNA ≥1 codon on average, distinctively above the noise level, and are not observed elsewhere in the trajectory (Figure 4A and 5A zoom-ins, a range of fluctuation amplitudes appear within each trace; Figure 4B table; more examples in Figure S4B). These fluctuations are noticeably different from the noisy sections recorded when the ribosome nears the end of the hairpin, where the tethered mRNA has mostly unwound into single strands and inevitably become much more elastic (the increase in noise is illustrated in Figure S4A, and corroborated by frequency analysis in Figure S4D). These large displacement fluctuations between the ribosome and the mRNA around the slippery sequence indicate that multiple mRNA translocation attempts occur at this region and that large slipping sizes such as -4 nt are indeed attainable. Interestingly, we find that fluctuations appear regardless of whether the ribosome ultimately frameshifts or not (Figure 4B table); also, they consistently occur even when the sequence is not slippery, e.g. on the A5G mutant mRNA (Figure 5A; Figure 4B table for lifetimes) whose frameshifting efficiency is reduced. Just as the slippery sequence is not the cause of fluctuations, neither is the hairpin: even though a hairpin barrier always remains in front of the ribosome throughout the entire trajectory, we detect these fluctuations only at the region downstream from the internal SD sequence. Thus, these observations indicate that a combination of flanking structural barriers—the upstream SD:antiSD mini-helix and the downstream hairpin junction—suffice to induce distinctive fluctuating ribosome translocation dynamics as the ribosome translates the region between the barriers. The barrier-induced, multiple translocation ribosomal excursions directly observed here have been indirectly detected in single-molecule fluorescence experiments (Chen et al., 2014; Kim et al., 2014).
To probe the nature of these large and persistent fluctuations, we characterized their dynamics. The average excursion lifetimes—i.e. the time between a backward shift and a forward motion (Figure 4A and 5A zoom-ins)—is ~0.5 s, independent of slippery sequence variant and frameshifting outcome (Figure 4B table). The distribution of the pooled excursion lifetimes is not a single exponential, indicating that more than one rate-limiting stochastic event control their duration. Accordingly, from the mean lifetime and its standard deviation, we calculated a lower bound for the apparent number of rate-limiting steps, nmin, to be ~1.6 (Figure S4C) (Moffitt et al., 2010). This value indicates that at least two rate-limiting dynamic events, of similar time scales, are required to return from a translocation excursion.
In addition to the above ~2-Hz dynamics resolved from the real-time trajectory data displayed at 20 Hz, we identified faster dynamics using power spectrum density analysis (from the 1 kHz raw data; Figure S4D). Enhanced fluctuations of 30, 85, and 180 Hz take place exclusively in the slippery sequence region, as compared to elsewhere in the trajectory. These time scales are similar to those reported for the 30S body and head dynamics during regular translation—in particular the head forward rotation at 80 Hz and reverse rotation at ~4-5 Hz (Guo and Noller, 2012). It is thus possible that the fluctuations captured at the slippery sequence region in the tweezers data reflect the conformational excursions of the ribosome 30S head during multiple mRNA forward translocation attempts. These fluctuations—which are not present during regular unidirectional translation on a hairpin—are uniquely promoted by mechanical mRNA barriers flanking the slippery sequence in order to achieve frameshifting.
Translocation fluctuations allow reading frame sampling
The large-scale, multiple translocation excursions observed across the slippery sequence region provide direct real-time insight into the mechanical movements that may be required for ribosomes to access the broad range of frameshift pathways independently resolved by MS with the 25-bp mRNA constructs. To relate these two findings, we analyzed the LC/MS-detected polypeptides translated with the same 55-bp mRNA hairpin construct used in the tweezers experiments; we show the frameshift-attenuating A5G template as an example (Figure 5B top bar graph). Similarly to Figure 2B, we compiled the frameshift pathways identified for A5G/55-bp into a 2D diagram (Figure 5A bottom-center) and aligned the resultant 2D plot with the translocation dynamics observed in the single-ribosome translation trajectory (Figure 5A lower-left zoom-in). The two results show clear correspondence along the slippery sequence region (orange-shaded area).
Since the ribosome constantly moved back and forth over ≥1-codon around the slippery sequence (Figure 4A and 5A, black-squared sections), it is not possible to pinpoint when along the fluctuating trajectory the new frame is established on the mRNA sequence. Nonetheless, the zoom-in of the fluctuating dynamics does reveal the locations on the mRNA transiently visited by the translocating ribosome. The fluctuation magnitude, i.e., the slipping range spanned by the translocating ribosome, coincides with the range of protein products distributed in the 2D diagram, supporting the inference that the observed multiple translocation excursions reflect the ribosome sampling of different reading frames. The correlation between the two independently acquired data sets is further strengthened when the abundance of frameshift translation products—including incomplete species—is taken into account. Specifically, the locations around the slippery sequence that were frequently visited by the fluctuating ribosome—thus transiently establishing alternative codon:anticodon base-pairing—are also the places where the higher populated frameshift translation products are found (Figure 5A, red arrow).
Altogether our findings portray a dynamic frameshifting scheme via alternative reading frame sampling, which is accessed upon multiple mRNA translocation attempts by the ribosome. Since these distinctive translocation excursions are seen only when translating between mRNA structural barriers, the energetic cost of breaking codon:anticodon base-pairs to frameshift are likely partially balanced by the energy liberated at the peripheral base-pairing interactions. It is conceivable that multiple EF-G binding events (Chen et al., 2014) may also play a role in driving the excursions. Furthermore, such a broad browsing range is presumably permitted by the swiveling and rotating 30S head when the ribosome is in the translocating mode—as suggested by the fluctuation frequency analysis described above. Indeed, structure studies have shown that, first, head tilting rearranges the mRNA binding groove on the 30S neck by disengaging the 16S rRNA residues that intercalate on the mRNA (Pulk and Cate, 2013; Schuwirth et al., 2005; Tourigny et al., 2013; Zhou et al., 2013); this process should ease the spatial restriction that prevents the tRNAs from base-pairing across adjacent codons in the out-of-frame manner. Second, 30S head rotation is coupled with mRNA forward translocation (Dunkle et al., 2011; Ermolenko and Noller, 2011; Gao et al., 2009; Guo and Noller, 2012; Ramrath et al., 2013; Ramrath et al., 2012; Ratje et al., 2010; Zhang et al., 2009; Zhou et al., 2012); thus, the 30S head is likely the agent for achieving large mRNA displacements that facilitate ribosomal frameshifting. A similar mechanism was proposed in recent kinetic studies, where—averaging over an ensemble and presumably over multiple translocation attempts—prolonged 30S head rotation was observed on a frameshift-programming mRNA (Caliskan et al., 2014). Our results hence illustrate a mechanism by which the 30S head rotation is perturbed by the flanking mechanical barrier elements, leading to multiple translocation attempts that enable frameshifting.
Not every frameshift attempt succeeds
Although the distinctive translocation excursions observed in the single-ribosome translation trajectories occur at the slippery sequence region, they are independent of the detailed content encoded in that sequence. Hence, we wondered how the overall distribution of frameshift translation products varies as the ribosome translates different templates.
For all template variants examined (the “55-bp” mRNA construct series; Figure 5B bar graphs), as noted before, incomplete species (ended with 0-frame amino acids before reaching the 0-stop; orange bars) accumulate at codon positions along the slippery sequence where the ribosome frequently frameshifts. When the slippery sequence becomes “less slippery,” i.e., more likely to cause codon:anticodon base-pair mismatches upon a slip, the incomplete species also become more populated (Figure 5B; 55%, 71%, and 25% for A5G, A4C, and wild-type, respectively). The equivalents of these aborted translation products were also observed among the single-ribosome translation trajectories: some of the ribosomes aborted translation midway and prematurely stalled around the slippery sequence. Based on the residual hairpin sizes measured upon ribosome stalling (Figure S3B; position resolution: ±0.8 nm ~1/3 codon), we found that the most probable aborted codon positions for each slippery sequence variant agree with the most abundant drop-off species detected by MS (Table 1). The presence of prematurely stalled ribosomes and aborted polypeptides indicate that not every frameshift attempt succeeds in resuming translation. Accordingly, our data suggest that while translocation excursions induced around the slippery sequence cause the ribosome to slip out of the 0-frame, if no compatible codon:anticodon base-pairing is found, the ribosomes stall and fail to incorporate the next amino acid, leading to the generation of incomplete 0-frame polypeptides.
To learn why some frameshift attempts fail, we compared the relative abundances of incomplete species to their frameshifted, -1-stop-terminated counterparts. For the A4C mutant (Figure 5B middle bar graph), ~87% of the -1-stop-terminated products (purple bars) come from codon position II, via -1 and -4-slips. However, a comparable amount of incomplete polypeptides (orange bars)—and no frameshifted products—was detected at codon position I. Such biases over particular frameshift pathways can be explained by the number of codon:anticodon base-pair mismatches encountered by the two tRNAs on a translocating ribosome. Here we use a dyad notation: (x, y) to annotate the number of mismatches at the two frameshifted tRNAs, in the E- and P-sites respectively, before the subsequent -1-frame aa-tRNA; every non-canonical base-pairing is given a score of 1, and G:U wobbles are allowed only at the third nucleotide position. Specifically, both -1 and -4-slips at codon position II result in fewer mismatches for the two tRNAs, as compared to those at codon position I (Figure 5B middle bar graph, numbers of mismatches tabulated). Therefore, due to better alternative base-pairing options, codon position II on the A4C mutant becomes the productive frameshift pathway. This analysis indicates that, once the ribosome has attempted to frameshift, the nucleotide composition along the slippery sequence dictates the outcome. Whereas the accumulation of incomplete species reveals the frequent slippages attempted at codon position I, the specific A4C mutation rendered those events unsuccessful. Consequently, the overall frameshift attempts on a given mRNA template should be estimated by including the amount of both frameshifted products and incomplete species (Table 1).
In comparison to A4C, the A5G mutant promoted -1-frameshift predominately at codon position I, while codon position II was a minor pathway (Figure 5B top bar graph). The former slipping route leads to mismatches (tabulated in the graph) only at the first tRNA, which moves to the E-site on the ribosome after translocation. In contrast, in the latter route, enduring only a P-site mismatch, ~90% of the ribosomes ceased translation (top bar graph position II). Thus, P-site mismatches appear to incur a higher penalty against continuing translation.
The correlation between the production of incomplete polypeptides and the occurrence of codon:anticodon mismatches is consistent with a retrospective fidelity check discovered in E. coli (Zaher and Green, 2009). After an amino acid mis-incorporation, the 30S subunit can recognize base-pair mismatches in its P- and E-sites as translation errors. The authors showed that post-translocation ribosomes—particularly with P-site mismatches—prematurely terminate translation by recruiting a release factor protein, e.g. RF2, into its A-site, in competition with the A-site codon specified aa-tRNA. Upon termination, incomplete polypeptides are released as premature drop-offs from the ribosomes. The correspondence between this retrospective quality control and our observations of prematurely aborted polypeptides, in the presence of both RF1 and RF2, indicate that a similar fidelity control operates during frameshifting along the slippery sequence.
By integrating all of our findings presented here with the current understanding of the bacterial ribosome translation mechanism, we arrive at the dynamic frameshifting scheme illustrated in Figure 6. In response to the flanking mRNA structural barriers acting as mechanical restoring devices, the ribosome stochastically makes multiple translocation attempts—i.e. excursions—promoted by the back-and-forth rotation of its 30S head. As each attempt has some probability of success, the succeeding translation may resume in a different frame, thereby mediating a widely branching set of translation pathways along the slippery sequence. The dynamic excursions observed in the single-ribosome translation trajectories not only corroborate the perturbed ribosome translocation revealed by ensemble kinetic studies (Caliskan et al., 2014), but also further refine their frameshifting model by showing that the barrier-hindered ribosome makes, in fact, multiple attempts to translocate through the frameshift-programming sequence region.
Figure 6. During mRNA translocation: a dynamic scheme for versatile ribosomal frameshifting.
Left: After the polypeptide chain (magenta curvy line) transfers from the P- to A-tRNA (blue and red vertical sticks), the elongation factor G (EF-G:GTP; complex in cyan and yellow star) catalyzes the P/E-, A/P-tRNAs translocation on the ribosome, along with mRNA (gray dashed line) forward translocation by one codon. This mRNA movement is brought by 30S head forward rotation (dark orange counter-clockwise arrow), displacing the E-, P-, and A-site codons (in green, blue, and red) to the left (gray downward arrow). To reset the ribosome for next round of translation, the head rotates back (middle cartoon; dark orange clockwise arrow). Multiple 30S head rotation—thus back-and-forth mRNA displacement—may be taken to achieve translocation between flanking mRNA structural barriers, e.g., SD:antiSD mini-helix and downstream hairpin, hence permitting the tRNAs to base-pair in alternative frames around the slippery sequence. When a new frame is adopted (top row)—at times with mismatches (black crosses)—both the new-frame specified aminoacyl-tRNA (delivered as EF-Tu:GTP:aa-tRNA: in orange, yellow star, and red) and the release factors (e.g. RF2, in green) can compete to bind with the mismatch-encountering, frameshifted ribosome. In the latter case, the ribosome ceases translation and releases an incomplete polypeptide.
We see no evidence for frameshifting pathways via schemes other than translocation excursions. Hence, while our data cannot rule out models in which A-site tRNA accommodation can also mediate frameshifting (Chen et al., 2014), we adopt the simpler translocation-mediated pathway branching mechanism.
Discussion
Versatile pathway branching regulates frameshifting
Previous studies have indicated that ribosomes are able to frameshift despite the creation of mismatches that would normally be perceived as translation errors (Atkins and Bjork, 2009; Farabaugh, 1996; Harger et al., 2002; Tsuchihashi and Brown, 1992). It is known that translation accuracy is primarily monitored and verified at steps prior to the irreversible peptidyl-transfer, ensuring proper charging and specific acceptance of the cognate aa-tRNA into the A-site on the ribosome (Gromadski and Rodnina, 2004; Guth and Francklyn, 2007). In the frameshifting scheme presented here, the potential conflict between frameshifting and fidelity is alleviated along the mRNA translocation step. Any mismatches upon codon:anticodon re-pairing during reading frame sampling would occur after the peptide bond formation (Figure 6) and, thus it is not susceptible to the fidelity controls governing proper mRNA decoding.
Instead, the retrospective fidelity check by the post-translocation ribosome very likely becomes the critical quality control in programmed frameshifting—a context that showcases its biological significance. This fidelity check subjects the mismatch-encountering ribosome to two competing routes: depending on the number and site of mismatches created upon a slip, the ribosome can either stop synthesis by recruiting a release factor, or proceed to the next round of amino acid incorporation in the new reading frame. Therefore, regardless of how the surrounding mRNA structural barriers may kinetically trap the ribosome and effectively promote frameshifting, only a fraction of ribosome slipping attempts during the dynamic translocation excursions succeeds to yield full-length frameshifted products. While mutant slippery sequences render a random slipping attempt risky, the dnaX slippery sequence—having evolved to offer optimal thermodynamic stability for alternative base-pairing—facilitates passing the fidelity check.
Our observations illustrate the indispensible role played by the slippery sequence—as well as adjacent codon positions before and after it—to assure high efficiency of “programmed” frameshifting. We can begin to parameterize and predict the apparent slipperiness of a given frameshift-programming mRNA by considering: (1) For each 0-frame codon within the sequence region flanked by the upstream and downstream barriers, how many alternative cognate and near-cognate base-pairing positions—i.e. slipping routes—exist in the out-of-frames; (2) For each of the slipping routes, how feasible is the required slipping size, given that it must be attained by ribosome 30S head rotation during mRNA translocation. Programmed mRNAs with greater totals of alternative codon:anticodon base-pairing options, weighted by the ease of the required slipping sizes, should exhibit greater frameshift efficiencies.
The fraction of completed frameshifted products—that as found here, can differ in length by a few amino acid residues—however, only reports half of the story for programmed frameshifting. The missing half is the previously unrecognized prematurely terminated polypeptides, whose significance is twofold. First, they likely reflect the retrospective fidelity control used by the ribosome; second, they represent the relics of unsuccessful slipping attempts induced by the surrounding secondary structures. Intriguingly, ribosome stalling and premature termination, as the aftermath of unsuccessful slipping attempts, have recently been shown to serve as characteristics of frameshift translation, which eukaryotic systems recognize to degrade exogenous programming mRNAs (Belew et al., 2014). Therefore, these impeded slipping attempts—ultimately leading to the “off-pathway” incomplete translation products—may have profound implications for the regulation both of translation and of mRNA abundance inside cells.
The overall frameshift attempts—composed of both the completed and the heretofore hidden aborted species—hence are the genuine measure for the frameshift-promoting strength of a programmed mRNA. In turn, this strength is determined by the mechanical properties of the mRNA structures that flank the slippery sequence and that interfere with the normal translation cycle. In this work, we have utilized these mechanical properties as a probe in situ: when the ribosome translates between the structural barriers, its intrinsic translocation dynamics are uniquely amplified, permitting a ribosomal slip and possibly engaging the retrospective fidelity check. Using optical tweezers we have captured the underlying dynamics of frameshifting translation in real-time, providing a glimpse of an unexpectedly versatile translation scheme.
Experimental Procedures
in vitro translation was performed using either the PURExpress® ΔRibosome kit (NEB) to synthesize polypeptide samples in large scale and analyze by mass spectrometry (MS), or a custom-made reconstituted reaction mixture for real-time single-ribosome translation in the optical tweezers experiments. Same preparations of purified E. coli MRE600 ribosomes and mRNA constructs were used in both experiments.
Exact composition and protocols of the PURExpress® system have been documented previously (Ohashi et al., 2010); see also Extended Experimental Procedures. All LC/MS experiments were performed on an LTQ Orbitrap XL mass spectrometer (Thermo Scientific), connected with an Agilent 1200 nanoflow HPLC system by means of nanoelectrospray. MS full scans were acquired in the Orbitrap analyzer (using internal lock mass recalibration in real-time), whereas tandem mass spectra were recorded in the linear ion trap. Each translation product was identified within 10-ppm deviation in mass (Da)—and with verification of its unique isotope pattern originated from specific amino acid composition.
Reconstituted reaction mixtures for tweezers experiments include: elongation factors (EF-G and EF-Tu), release factors (RF1 and RF2), GTP, etc., along with selectively charged aminoacyl-tRNAs to fuel translation inside a micro-fluidic chamber. We first tether one ribosome-mRNA complex (single ribosome initiated and halted on the long mRNA hairpin construct)—via a pair of terminal-modified dsDNA handles—between two 2.1-µm, surface-modified polystyrene beads in the tweezers setup (Figure 3A). Before beginning translation, we register the starting position of the ribosome on the mRNA based on the downstream hairpin size measured; this measurement is done by applying force to unfold and refold the hairpin. Upon flowing the above mixture into the chamber while holding the mRNA tether at constant force, we commence recording step-wise hairpin unwinding that reports single-ribosome translation in real-time. After the course of translation, we again measure the residual hairpin to verify the ribosome termination position. Detailed protocols for protein factor and material preparations, ribosome-mRNA complex formations, configurations of the single-trap optical miniTweezers (S.B. Smith, TweezersLAB), step-by-step instrumental operations, and single-ribosome translation trajectory data analysis have been described in the literature and our previous studies (Dincbas-Renqvist et al., 2000; Qu et al., 2011; Wen et al., 2008); see also Extended Experimental Procedures.
Supplementary Material
Acknowledgements
We thank Dr. S. B. Smith for help with the optical tweezers instrument, Dr. T. Iavarone (QB3/Chemistry Mass Spectrometry Facility) for assistance on MS data collection and analysis, Dr. L. Lancaster and Prof. H. Noller for expertise in sample preparations, Dr. C. L. Hetherington for comments on the manuscript, and members of the Tinoco and Bustamante labs for discussions. This work was supported by grants from the National Institutes of Health (GM10840 to I.T.).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Supplemental Information
Supplemental Information includes Extended Experimental Procedures, 5 figures, and 2 tables can be found with this article online at http://dx.doi.org/10.1016/j.cell.XXXX.XX.XXX.
Author contributions
S.Y. and J.-D.W. designed the experiments and prepared the materials; S.Y. performed the experiments and analyzed the data; all authors wrote the paper.
References
- Atkins JF, Bjork GR. A Gripping Tale of Ribosomal Frameshifting: Extragenic Suppressors of Frameshift Mutations Spotlight P-Site Realignment. Microbiol Mol Biol R. 2009;73 doi: 10.1128/MMBR.00010-08. 178-+ [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belew AT, Meskauskas A, Musalgaonkar S, Advani VM, Sulima SO, Kasprzak WK, Shapiro BA, Dinman JD. Ribosomal frameshifting in the CCR5 mRNA is regulated by miRNAs and the NMD pathway. Nature. 2014;512:265–269. doi: 10.1038/nature13429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bretscher MS. Translocation in protein synthesis: a hybrid structure model. Nature. 1968;218:675–677. doi: 10.1038/218675a0. [DOI] [PubMed] [Google Scholar]
- Caliskan N, Katunin VI, Belardinelli R, Peske F, Rodnina MV. Programmed-1 Frameshifting by Kinetic Partitioning during Impeded Translocation. Cell. 2014;157:1619–1631. doi: 10.1016/j.cell.2014.04.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J, Petrov A, Johansson M, Tsai A, O'Leary SE, Puglisi JD. Dynamic pathways of-1 translational frameshifting. Nature. 2014;512:328–332. doi: 10.1038/nature13428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dincbas-Renqvist V, Engstrom A, Mora L, Heurgue-Hamard V, Buckingham R, Ehrenberg M. A post-translational modification in the GGQ motif of RF2 from Escherichia coli stimulates termination of translation. Embo J. 2000;19:6900–6907. doi: 10.1093/emboj/19.24.6900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drummond DA, Wilke CO. The evolutionary consequences of erroneous protein synthesis. Nat Rev Genet. 2009;10:715–724. doi: 10.1038/nrg2662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunkle JA, Wang L, Feldman MB, Pulk A, Chen VB, Kapral GJ, Noeske J, Richardson JS, Blanchard SC, Cate JH. Structures of the bacterial ribosome in classical and hybrid states of tRNA binding. Science. 2011;332:981–984. doi: 10.1126/science.1202692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ermolenko DN, Noller HF. mRNA translocation occurs during the second step of ribosomal intersubunit rotation. Nat Struct Mol Biol. 2011;18 doi: 10.1038/nsmb.2011. 457-U492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farabaugh PJ. Programmed translational frameshifting. Microbiol Rev. 1996;60:103–134. doi: 10.1128/mr.60.1.103-134.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao YG, Selmer M, Dunham CM, Weixlbaumer A, Kelley AC, Ramakrishnan V. The structure of the ribosome with elongation factor G trapped in the posttranslocational state. Science. 2009;326:694–699. doi: 10.1126/science.1179709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gesteland RF, Atkins JF. Recoding: dynamic reprogramming of translation. Annu Rev Biochem. 1996;65:741–768. doi: 10.1146/annurev.bi.65.070196.003521. [DOI] [PubMed] [Google Scholar]
- Gromadski KB, Rodnina MV. Kinetic determinants of high-fidelity tRNA discrimination on the ribosome. Molecular Cell. 2004;13:191–200. doi: 10.1016/s1097-2765(04)00005-x. [DOI] [PubMed] [Google Scholar]
- Guo ZJ, Noller HF. Rotation of the head of the 30S ribosomal subunit during mRNA translocation. Proc Natl Acad Sci U S A. 2012;109:20391–20394. doi: 10.1073/pnas.1218999109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guth EC, Francklyn CS. Kinetic discrimination of tRNA identity by the conserved motif 2 loop of a class II aminoacyl-tRNA synthetase. Molecular Cell. 2007;25:531–542. doi: 10.1016/j.molcel.2007.01.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harger JW, Meskauskas A, Dinman JD. An 'integrated model' of programmed ribosomal frameshifting. Trends Biochem Sci. 2002;27:448–454. doi: 10.1016/s0968-0004(02)02149-7. [DOI] [PubMed] [Google Scholar]
- Jacks T, Power MD, Masiarz FR, Luciw PA, Barr PJ, Varmus HE. Characterization of Ribosomal Frameshifting in Hiv-1 Gag-Pol Expression. Nature. 1988;331:280–283. doi: 10.1038/331280a0. [DOI] [PubMed] [Google Scholar]
- Jenner L, Rees B, Yusupov M, Yusupova G. Messenger RNA conformations in the ribosomal E site revealed by X-ray crystallography. EMBO Rep. 2007;8:846–850. doi: 10.1038/sj.embor.7401044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jenner LB, Demeshkina N, Yusupova G, Yusupov M. Structural aspects of messenger RNA reading frame maintenance by the ribosome. Nat Struct Mol Biol. 2010;17:555–560. doi: 10.1038/nsmb.1790. [DOI] [PubMed] [Google Scholar]
- Kaminishi T, Wilson DN, Takemoto C, Harms JM, Kawazoe M, Schluenzen F, Hanawa-Suetsugu K, Shirouzu M, Fucini P, Yokoyama S. A snapshot of the 30S ribosomal subunit capturing mRNA via the Shine-Dalgarno interaction. Structure. 2007;15:289–297. doi: 10.1016/j.str.2006.12.008. [DOI] [PubMed] [Google Scholar]
- Kim HK, Liu F, Fei JY, Bustamante C, Gonzalez RL, Tinoco I. A frameshifting stimulatory stem loop destabilizes the hybrid state and impedes ribosomal translocation. Proc Natl Acad Sci U S A. 2014;111:5538–5543. doi: 10.1073/pnas.1403457111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korostelev A, Trakhanov S, Asahara H, Laurberg M, Lancaster L, Noller HF. Interactions and dynamics of the Shine Dalgarno helix in the 70S ribosome. Proc Natl Acad Sci U S A. 2007;104:16840–16843. doi: 10.1073/pnas.0707850104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liao PY, Choi YS, Dinman JD, Lee KH. The many paths to frameshifting: kinetic modelling and analysis of the effects of different elongation steps on programmed-1 ribosomal frameshifting. Nucleic Acids Res. 2011;39:300–312. doi: 10.1093/nar/gkq761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moazed D, Noller HF. Intermediate states in the movement of transfer RNA in the ribosome. Nature. 1989;342:142–148. doi: 10.1038/342142a0. [DOI] [PubMed] [Google Scholar]
- Moffitt JR, Chemla YR, Bustamante C. Mechanistic constraints from the substrate concentration dependence of enzymatic fluctuations. Proc Natl Acad Sci U S A. 2010;107:15739–15744. doi: 10.1073/pnas.1006997107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohashi H, Kanamori T, Shimizu Y, Ueda T. A highly controllable reconstituted cell-free system--a breakthrough in protein synthesis research. Curr Pharm Biotechnol. 2010;11:267–271. doi: 10.2174/138920110791111889. [DOI] [PubMed] [Google Scholar]
- Pulk A, Cate JH. Control of ribosomal subunit rotation by elongation factor G. Science. 2013;340:1235970. doi: 10.1126/science.1235970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qu X, Wen JD, Lancaster L, Noller HF, Bustamante C, Tinoco I., Jr The ribosome uses two active mechanisms to unwind messenger RNA during translation. Nature. 2011;475:118–121. doi: 10.1038/nature10126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramrath DJ, Lancaster L, Sprink T, Mielke T, Loerke J, Noller HF, Spahn CM. Visualization of two transfer RNAs trapped in transit during elongation factor G-mediated translocation. Proc Natl Acad Sci U S A. 2013;110:20964–20969. doi: 10.1073/pnas.1320387110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramrath DJF, Yamamoto H, Rother K, Wittek D, Pech M, Mielke T, Loerke J, Scheerer P, Ivanov P, Teraoka Y, et al. The complex of tmRNA-SmpB and EF-G on translocating ribosomes. Nature. 2012;485 doi: 10.1038/nature11006. 526-U140. [DOI] [PubMed] [Google Scholar]
- Ratje AH, Loerke J, Mikolajka A, Brunner M, Hildebrand PW, Starosta AL, Donhofer A, Connell SR, Fucini P, Mielke T, et al. Head swivel on the ribosome facilitates translocation by means of intra-subunit tRNA hybrid sites. Nature. 2010;468:713–716. doi: 10.1038/nature09547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodnina MV, Gromadski KB, Kothe U, Wieden HJ. Recognition and selection of tRNA in translation. FEBS Lett. 2005;579:938–942. doi: 10.1016/j.febslet.2004.11.048. [DOI] [PubMed] [Google Scholar]
- Rodnina MV, Wintermeyer W. The ribosome as a molecular machine: the mechanism of tRNA-mRNA movement in translocation. Biochem Soc Trans. 2011;39:658–662. doi: 10.1042/BST0390658. [DOI] [PubMed] [Google Scholar]
- Savelsbergh A, Katunin VI, Mohr D, Peske F, Rodnina MV, Wintermeyer W. An Elongation Factor G-Induced Ribosome Rearrangement Precedes tRNAmRNA Translocation. Molecular Cell. 2003;11:1517–1523. doi: 10.1016/s1097-2765(03)00230-2. [DOI] [PubMed] [Google Scholar]
- Schmeing TM, Ramakrishnan V. What recent ribosome structures have revealed about the mechanism of translation. Nature. 2009;461:1234–1242. doi: 10.1038/nature08403. [DOI] [PubMed] [Google Scholar]
- Schuwirth BS, Borovinskaya MA, Hau CW, Zhang W, Vila-Sanjurjo A, Holton JM, Cate JH. Structures of the bacterial ribosome at 3.5 A resolution. Science. 2005;310:827–834. doi: 10.1126/science.1117230. [DOI] [PubMed] [Google Scholar]
- Selmer M, Dunham CM, Murphy FV, Weixlbaumer A, Petry S, Kelley AC, Weir JR, Ramakrishnan V. Structure of the 70S ribosome complexed with mRNA and tRNA. Science. 2006;313:1935–1942. doi: 10.1126/science.1131127. [DOI] [PubMed] [Google Scholar]
- Stahl G, McCarty GP, Farabaugh PJ. Ribosome structure: revisiting the connection between translational accuracy and unconventional decoding. Trends Biochem Sci. 2002;27:178–183. doi: 10.1016/S0968-0004(02)02064-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tinoco I, Jr, Kim HK, Yan S. Frameshifting dynamics. Biopolymers. 2013;99:1147–1166. doi: 10.1002/bip.22293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tourigny DS, Fernandez IS, Kelley AC, Ramakrishnan V. Elongation factor G bound to the ribosome in an intermediate state of translocation. Science. 2013;340:1235490. doi: 10.1126/science.1235490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsuchihashi Z, Brown PO. Sequence requirements for efficient translational frameshifting in the Escherichia coli dnaX gene and the role of an unstable interaction between tRNA(Lys) and an AAG lysine codon. Genes & Development. 1992;6:511–519. doi: 10.1101/gad.6.3.511. [DOI] [PubMed] [Google Scholar]
- Weiss RB, Dunn DM, Atkins JF, Gesteland RF. Slippery runs, shifty stops, backward steps, and forward hops: −2,−1, +1, +2, +5, and +6 ribosomal frameshifting. Cold Spring Harb Symp Quant Biol. 1987;52:687–693. doi: 10.1101/sqb.1987.052.01.078. [DOI] [PubMed] [Google Scholar]
- Wen JD, Lancaster L, Hodges C, Zeri AC, Yoshimura SH, Noller HF, Bustamante C, Tinoco I. Following translation by single ribosomes one codon at a time. Nature. 2008;452:598–603. doi: 10.1038/nature06716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wohlgemuth I, Brenner S, Beringer M, Rodnina MV. Modulation of the rate of peptidyl transfer on the ribosome by the nature of substrates. J Biol Chem. 2008;283:32229–32235. doi: 10.1074/jbc.M805316200. [DOI] [PubMed] [Google Scholar]
- Yusupov MM, Yusupova GZ, Baucom A, Lieberman K, Earnest TN, Cate JH, Noller HF. Crystal structure of the ribosome at 5.5 A resolution. Science. 2001;292:883–896. doi: 10.1126/science.1060089. [DOI] [PubMed] [Google Scholar]
- Yusupova G, Jenner L, Rees B, Moras D, Yusupov M. Structural basis for messenger RNA movement on the ribosome. Nature. 2006;444:391–394. doi: 10.1038/nature05281. [DOI] [PubMed] [Google Scholar]
- Yusupova GZ, Yusupov MM, Cate JH, Noller HF. The path of messenger RNA through the ribosome. Cell. 2001;106:233–241. doi: 10.1016/s0092-8674(01)00435-4. [DOI] [PubMed] [Google Scholar]
- Zaher HS, Green R. Quality control by the ribosome following peptide bond formation. Nature. 2009;457:161–166. doi: 10.1038/nature07582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang W, Dunkle JA, Cate JH. Structures of the ribosome in intermediate states of ratcheting. Science. 2009;325:1014–1017. doi: 10.1126/science.1175275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou J, Lancaster L, Donohue JP, Noller HF. Crystal structures of EFG- ribosome complexes trapped in intermediate states of translocation. Science. 2013;340:1236086. doi: 10.1126/science.1236086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou J, Lancaster L, Trakhanov S, Noller HF. Crystal structure of release factor RF3 trapped in the GTP state on a rotated conformation of the ribosome. Rna. 2012;18:230–240. doi: 10.1261/rna.031187.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.