Abstract
A putative base-pairing interaction that determines the folding of the central region of 23S rRNA has been investigated by mutagenesis. Each of the possible base substitutions has been made at the phylogenetically covariant positions adenine-1262 (A1262) and U2017 in Escherichia coli 23S rRNA. Every substitution that disrupts the potential for Watson-Crick base pairing between these positions reduces or abolishes the participation of 23S rRNA in protein synthesis. All mutant 23S rRNAs are assembled into 50S subunits, but the mutant subunits are less able to stably interact with 30S subunits to form translationally active ribosomes. The function of 23S rRNA is largely reestablished by introduction of an alternative G1262.C2017 or U1262.A2017 pair, although neither of these supports polysome formation quite as effectively as the wild-type pair. A 23S rRNA with a C1262.G2017 pair is nonfunctional. In contrast to the considerable effect the mutations have on function, they impart only slight structural changes on the naked rRNA, and these are limited to the immediate vicinity of the mutations. The data show that positions 1262 and 2017 pair in a Watson-Crick manner, but the data also indicate that these nucleotides engage in additional interactions within the ribosome and that these interactions determine what base pairs are acceptable there.
Full text
PDF

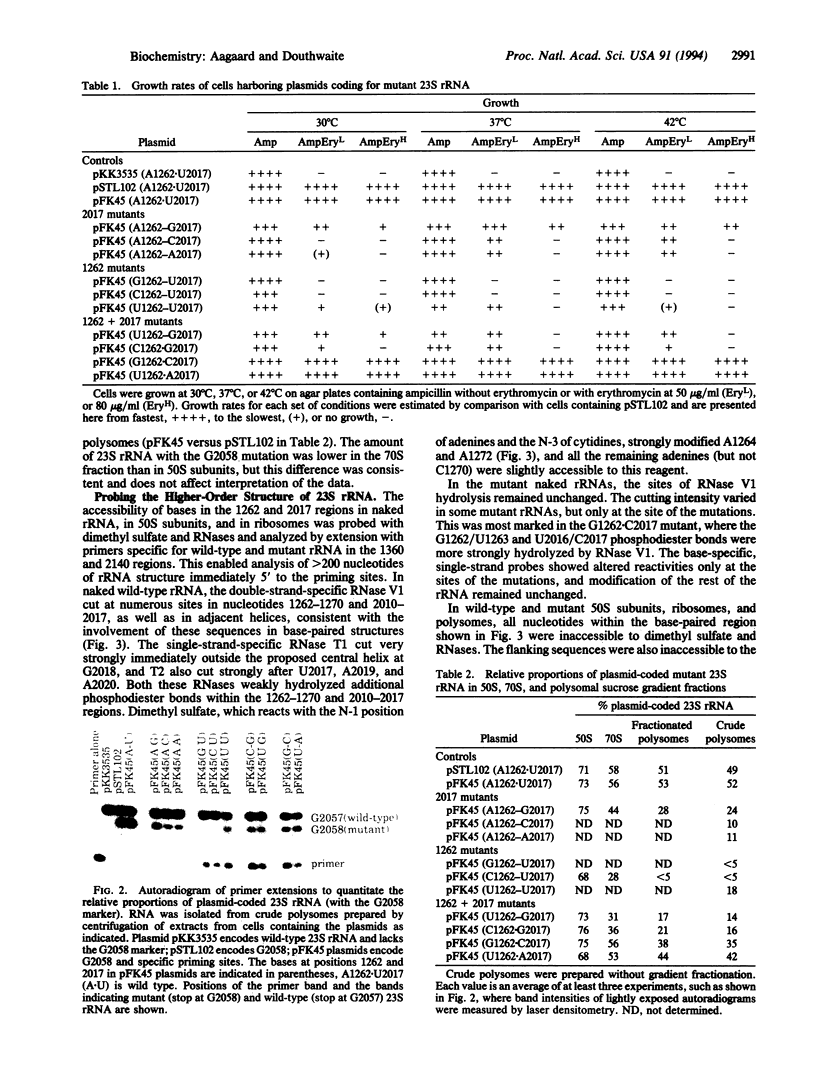


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aagaard C., Rosendahl G., Dam M., Powers T., Douthwaite S. Specific structural probing of plasmid-coded ribosomal RNAs from Escherichia coli. Biochimie. 1991 Dec;73(12):1439–1444. doi: 10.1016/0300-9084(91)90176-2. [DOI] [PubMed] [Google Scholar]
- Brosius J., Dull T. J., Sleeter D. D., Noller H. F. Gene organization and primary structure of a ribosomal RNA operon from Escherichia coli. J Mol Biol. 1981 May 15;148(2):107–127. doi: 10.1016/0022-2836(81)90508-8. [DOI] [PubMed] [Google Scholar]
- Brosius J., Ullrich A., Raker M. A., Gray A., Dull T. J., Gutell R. R., Noller H. F. Construction and fine mapping of recombinant plasmids containing the rrnB ribosomal RNA operon of E. coli. Plasmid. 1981 Jul;6(1):112–118. doi: 10.1016/0147-619x(81)90058-5. [DOI] [PubMed] [Google Scholar]
- Chen E. Y., Seeburg P. H. Supercoil sequencing: a fast and simple method for sequencing plasmid DNA. DNA. 1985 Apr;4(2):165–170. doi: 10.1089/dna.1985.4.165. [DOI] [PubMed] [Google Scholar]
- Dahlberg A. E. The functional role of ribosomal RNA in protein synthesis. Cell. 1989 May 19;57(4):525–529. doi: 10.1016/0092-8674(89)90122-0. [DOI] [PubMed] [Google Scholar]
- Douthwaite S., Powers T., Lee J. Y., Noller H. F. Defining the structural requirements for a helix in 23 S ribosomal RNA that confers erythromycin resistance. J Mol Biol. 1989 Oct 20;209(4):655–665. doi: 10.1016/0022-2836(89)93000-3. [DOI] [PubMed] [Google Scholar]
- Egebjerg J., Leffers H., Christensen A., Andersen H., Garrett R. A. Structure and accessibility of domain I of Escherichia coli 23 S RNA in free RNA, in the L24-RNA complex and in 50 S subunits. Implications for ribosomal assembly. J Mol Biol. 1987 Jul 5;196(1):125–136. doi: 10.1016/0022-2836(87)90515-8. [DOI] [PubMed] [Google Scholar]
- Gutell R. R., Gray M. W., Schnare M. N. A compilation of large subunit (23S and 23S-like) ribosomal RNA structures: 1993. Nucleic Acids Res. 1993 Jul 1;21(13):3055–3074. doi: 10.1093/nar/21.13.3055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gutell R. R., Woese C. R. Higher order structural elements in ribosomal RNAs: pseudo-knots and the use of noncanonical pairs. Proc Natl Acad Sci U S A. 1990 Jan;87(2):663–667. doi: 10.1073/pnas.87.2.663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haselman T., Gutell R. R., Jurka J., Fox G. E. Additional Watson-Crick interactions suggest a structural core in large subunit ribosomal RNA. J Biomol Struct Dyn. 1989 Aug;7(1):181–186. doi: 10.1080/07391102.1989.10507759. [DOI] [PubMed] [Google Scholar]
- Höpfl P., Ludwig W., Schleifer K. H., Larsen N. The 23S ribosomal RNA higher-order structure of Pseudomonas cepacia and other prokaryotes. Eur J Biochem. 1989 Nov 6;185(2):355–364. doi: 10.1111/j.1432-1033.1989.tb15123.x. [DOI] [PubMed] [Google Scholar]
- Larsen N. Higher order interactions in 23s rRNA. Proc Natl Acad Sci U S A. 1992 Jun 1;89(11):5044–5048. doi: 10.1073/pnas.89.11.5044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsen N., Olsen G. J., Maidak B. L., McCaughey M. J., Overbeek R., Macke T. J., Marsh T. L., Woese C. R. The ribosomal database project. Nucleic Acids Res. 1993 Jul 1;21(13):3021–3023. doi: 10.1093/nar/21.13.3021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leffers H., Kjems J., Ostergaard L., Larsen N., Garrett R. A. Evolutionary relationships amongst archaebacteria. A comparative study of 23 S ribosomal RNAs of a sulphur-dependent extreme thermophile, an extreme halophile and a thermophilic methanogen. J Mol Biol. 1987 May 5;195(1):43–61. doi: 10.1016/0022-2836(87)90326-3. [DOI] [PubMed] [Google Scholar]
- Moazed D., Stern S., Noller H. F. Rapid chemical probing of conformation in 16 S ribosomal RNA and 30 S ribosomal subunits using primer extension. J Mol Biol. 1986 Feb 5;187(3):399–416. doi: 10.1016/0022-2836(86)90441-9. [DOI] [PubMed] [Google Scholar]
- Noller H. F., Kop J., Wheaton V., Brosius J., Gutell R. R., Kopylov A. M., Dohme F., Herr W., Stahl D. A., Gupta R. Secondary structure model for 23S ribosomal RNA. Nucleic Acids Res. 1981 Nov 25;9(22):6167–6189. doi: 10.1093/nar/9.22.6167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noller H. F. Ribosomal RNA and translation. Annu Rev Biochem. 1991;60:191–227. doi: 10.1146/annurev.bi.60.070191.001203. [DOI] [PubMed] [Google Scholar]
- Noller H. F. Structure of ribosomal RNA. Annu Rev Biochem. 1984;53:119–162. doi: 10.1146/annurev.bi.53.070184.001003. [DOI] [PubMed] [Google Scholar]
- Ron E. Z., Kohler R. E., Davis B. D. Polysomes extracted from Escherichia coli by freeze-thaw-lysozyme lysis. Science. 1966 Sep 2;153(3740):1119–1120. doi: 10.1126/science.153.3740.1119. [DOI] [PubMed] [Google Scholar]
- Sigmund C. D., Ettayebi M., Borden A., Morgan E. A. Antibiotic resistance mutations in ribosomal RNA genes of Escherichia coli. Methods Enzymol. 1988;164:673–690. doi: 10.1016/s0076-6879(88)64077-8. [DOI] [PubMed] [Google Scholar]
- Taylor J. W., Ott J., Eckstein F. The rapid generation of oligonucleotide-directed mutations at high frequency using phosphorothioate-modified DNA. Nucleic Acids Res. 1985 Dec 20;13(24):8765–8785. doi: 10.1093/nar/13.24.8765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R., Magrum L. J., Gupta R., Siegel R. B., Stahl D. A., Kop J., Crawford N., Brosius J., Gutell R., Hogan J. J. Secondary structure model for bacterial 16S ribosomal RNA: phylogenetic, enzymatic and chemical evidence. Nucleic Acids Res. 1980 May 24;8(10):2275–2293. doi: 10.1093/nar/8.10.2275. [DOI] [PMC free article] [PubMed] [Google Scholar]




