Abstract
Directed cell migration requires a spatially polarized distribution of polymerized actin. We develop and treat a mechanical model of cell polarization based on polymerization and depolymerization of actin filaments at the two ends of a cell, modulated by forces at either end that are coupled by the cell membrane. We solve this model using both a simulation approach that treats filament nucleation, polymerization, and depolymerization stochastically, and a rate-equation approach based on key properties such as the number of filaments N and the number of polymerized subunits F at either end of the cell. The rate-equation approach agrees closely with the stochastic approach at steady state and, when appropriately generalized, also predicts the dynamic behavior accurately. The calculated transitions from symmetric to polarized states show that polarization is enhanced by a high free-actin concentration, a large pointed-end off-rate, a small barbed-end off-rate, and a small spontaneous nucleation rate. The rate-equation approach allows us to perform a linear-stability analysis to pin down the key interactions that drive the polarization. The polarization is driven by a positive-feedback loop having two interactions. First, an increase in F at one side of the cell lengthens the filaments and thus reduces the decay rate of N (increasing N); second, increasing N enhances F because the force per growing filament tip is reduced. We find that the transitions induced by changing system properties result from supercritical pitchfork bifurcations. The filament lifetime depends strongly on the average filament length, and this effect is crucial for obtaining polarization correctly.
Keywords: actin polymerization, symmetry breaking, bifurcation
1. Introduction
Many important functions of cells and microorganisms rely on directed migration based on polarization, including cancer metastasis and neutrophil tracking of bacteria. Polarization arises from an asymmetric actin distribution where a denser actin network grows at one side (leading edge) and a sparser network at the other side (trailing edge) of the cell [1]. Cells can polarize either spontaneously [2] or in response to extracellular signals such as chemoattractants [3, 4] and mechanical force [2]. After the stimulus is removed, cells sometimes maintain their polarity and keep migrating [2]. Thus under some conditions, the polarized state of a state is stable, while the unpolarized state is unstable or metastable. The connection between polarization and the direction of cell migration has recently been emphasized by experiments in which cells were polarized by micropatterns and subsequently allowed to migrate [5].
Several recent reviews [6, 7, 1, 8, 9, 10, 11] have treated the mathematical modeling of cell polarization. At this point the relative contributions of different interactions, and the conditions which favor the polarized state, are still not known in detail. Both chemical interactions and mechanical interactions play a role. Most models of cell polarization have employed the general framework of the local excitation and global inhibition (LEGI) mechanism [12], based on a slowly diffusing (local) activator and a fast diffusing (global) inhibitor. In the LEGI mechanism, the activator is a protein or other molecule, whose concentration grows autocatalytically, until stopped by growth of the inhibitor. If the ratio of inhibitor diffusion to activator diffusion is large enough, then a static buildup of activator will occur. Several treatments have identified the activator with signalling proteins such as Rac and phosphatidylinositol (3,4,5)-trisphosphate (PIP3) [3, 13], and the inhibitor with phosphatase and tensin homolog (PTEN).
However, a purely chemical picture is incomplete, and may not even capture the key ingredients. Recent measurements [3] have instead argued for mechanical tension as a key inhibitor driving cell polarization. It was shown that the inhibitory interactions propagate too quickly for diffusing signal proteins to play the role of the global inhibitor. Therefore, mechanical tension, which propagates almost instantaneously, was suggested as a candidate for the fast inhibitor. This is consistent with previous work in which polarization was induced by the application of mechanical force [2]. However, it is not known precisely how tension inhibits the key processes, nor how it interacts with the chemistry of actin polymerization.
Several theoretical treatments have aimed to clarify the role of mechanical force and actin polymerization in polarization, some of which are reviewed in Ref. [10]. Ref. [14] treated a simple model of “inside-out” polarization, in which actin filaments fixed at their nongrowing ends pushed on a bead from different sides. They found that this model led to directed motion of the bead via a force-dependent symmetry-breaking mechanism. However, it is not clear over what range of parameters symmetry breaking would occur, nor to what extent the same mechanism would operate in cells. Ref. [15] treated a model of autocatalytic actin network growth based on filament branching caused by Arp2/3 complex, in a simple cell geometry. The force was transmitted via flexible springs between the ends of the cell, which were treated as flexible polymers. It was found that autocatalytic branching nucleation of actin filaments could allow polymerized actin by itself to function as an activator molecule, and temporary polarization was found in the model. However, at large time scales, the model obtained Brownian motion rather than stable directed motion. In addition, transitions between polarized and unpolarized states with varying parameters were not explored. Finally, because the polarization relied on autocatalytic actin filament nucleation, it was not clear to what extent mechanical effects were important for polarization.
A simplified mechanical model of cell fragments treated the interplay between pushing force exerted by actin polymerization on the cell edges, contractile force powered by myosin II across the cell, and elastic tension in the cell membrane [16]. Forces from actin and myosin were treated as assumed force densities. In the energy as a function of deformation, two stable energy minima were found, one polarized and unpolarized. Polarization resulted from the active forces of myosin, and actin polymerization, as modulated by nonlinear mechanics. It was found that application of a force, as in Ref. [2], could induce a transition to a polarized state. Ref. [17] treated the interplay between membrane tension, filament force generation at the front of a cell, and filament breaking at the rear. Again using a distributed force density, they calculated a self-consistent polarized profile of polymerized actin. They found a correlation between front-to-rear distance of the cell and curvature of the leading edge, consistent with experiment.
The work of [18] treated a membrane that was deformed by actin polymerization and actomyosin contraction, based on continuous densities of actin and myosin. The front and back of the cell were coupled by the elasticity of the membrane. After perturbation of an initially circular cell by a pulse of F-actin at the front, the asymmetry in the actin distribution persisted, and the cell deformed, becoming longer in the direction transverse to its motion. Myosin moved to the rear of the cell. A major contribution to polarization in this case came from advection of actin and myosin due to active processes. The authors calculated a phase diagram giving cell shape as a function of cell-substrate adhesion strength and myosin activity. They found that cell asymmetry was most favored at a point where myosin activity and cell-substrate adhesion are intermediate.
In the models of Refs. [16], [17], and [18], the key mechanical interactions and their effect on actin polymerization are given explicitly. However, the treatment of actin polymerization in these models is simplified. Here we aim to grasp the key mechanical interactions in a more complete, stochastic model of actin polymerization. In order to focus on the mechanical interactions, we treat a model including only actin polymerization and mechanical force, and do not treat the upstream signaling pathways. The model treats actin networks present at the two sides of an idealized square cell fragment. The two sides serve as identical obstacles to actin polymerization, which slow the polymerization according to the the Brownian Ratchet mechanism [19]. The model is solved using both a stochastic simulation approach, and a rate equation approach that is systematically derived from the stochastic approach. The results of the two approaches agree closely, for both steady-state and dynamic properties of the system. Both treatments predict symmetry breaking over a broad range of parameter values. The origins of the symmetry breaking are explored by a linear stability analysis of the rate equations. We find that the symmetry breaking is caused by a feedback loop connecting fluctuations in the number of filaments on the two sides of the cell with fluctuations in the amount of polymerized actin. Increasing polymerized actin on one side favors growth of the number of filaments on that side, while the increased number of filaments can favor or inhibit actin polymerization depending on the parameter values. A key factor in obtaining accurate estimates of polarization in the rate-equation approach is the use of a state-dependent filament decay rate.
The development of an accurate rate-equation approach for treating actin polymerization and depolymerization will have applications beyond the specific problem of cell polarization treated here. Simulations of whole-cell structure and function usually require simplifying approximations such as the description of actin and related proteins by continuous densities. A major difficulty in such calculations has been knowing how to treat actin network disassembly correctly, and the state-dependent decay rate developed here is a step in this direction.
2. Model
2.1. Stochastic Model
We treat a 2μm × 2μm square cell fragment with height 0.2μm, containing actin filaments. The lateral size is taken to correspond to a small cell fragment, while the height is chosen to correspond to a typical lamellipodium thickness. Changing the system size leads to changes in parameter values, but not in the qualitative behavior seen below. For example, doubling the length of the cell changes the critical values of the parameters required for polarization by about 50%. The model treats free actin (G-actin) and polymerized actin filaments (F-actin). The actin filaments are located at either end of the cell. The G-actin monomers diffuse freely in the cytoplasm of the cell, and are converted to F-actin by polymerization or nucleation events. Conversely, F-actin subunits convert to G-actin through depolymerization from filaments. Each individual filament has two chemically distinct ends, a “barbed” end where polymerization is favored, and a “pointed” end where depolymerization is favored. Force on the barbed ends slows actin polymerization according to the Brownian Ratchet mechanism [19]. In order to focus on the effects of mechanical interactions on cell polarization, we treat only the most basic unit processes: nucleation and growth of actin filaments. This model is similar to that of Ref. [15], but differs in that autocatalytic branching is not included, and that we treat a rigid membrane at each side of the cell. In both approaches to solving the model, the following assumptions are made:
Filaments nucleate only right at the front and back of the cell, as indicated in blue in Fig. 1. This assumption is based on the fact that actin filament nucleation requires Arp2/3 complex, which is activated by agents in the membrane [20]. The nucleated filament network is taken to be two dimensional because of the limited height of the simulation cell. Thus the height controls only the actin monomer supply and the rate of filament nucleation at the cell edges.
Arp2/3 complex is present in excess, so that we can ignore the effect of filament nucleation on the cytosolic Arp2/3 concentration. Thus chemical feedbacks involving the Arp2/3 concentration are ignored.
G-actin diffusion is infinitely fast. In the present model, actin assembly and disassembly occur at the same side of the cell, rather than assembly at one end and disassembly at the other end. Therefore, there is no net transport of monomers from one side of the cell to the other. Previous analysis [21] has shown that diffusion of G-actin in this case is not a significant constraint. The rate-limiting G-actin diffusion process is from the front to the rear of the actin network at either side of the cell. Typical diffusion coefficients of actin monomers in vivo are ~ 5×10−8cm2/s [22], which implies that the time scale for this process is on the order of a second, much smaller than the calculated time scale of polarization. Thus the steady state of the model is not affected strongly this assumption. However, diffusion rate limitations could affect the dynamics of polarization more strongly.
The actin filament barbed ends point out of the cell fragment, as found in many studies including Ref. [23]. Each filament has an angle α = ±35° with respect to the forward direction [23, 24]. In Fig. 1, the red filaments are growing and generating force against the membrane.
There is no pointed-end growth.
Filament capping and branching are ignored, in order to limit the number of variables. However, as discussed below, our model may in some ways mimic the nucleation-growth-capping-depolymerization cycle believed to control actin dynamics cells.
The actin filaments are rigid and tightly attached to an underlying substrate. The most pronounced departures from this scenario would result from retrograde flow at the leading edge, in which polymerized actin slides over the substrate. The extent of retrograde flow varies between cell types, and our results will be most relevant to those where retrograde flow is slow.
The positions X1 and X2 of the membrane at the two sides of the cell are determined by the average of the barbed-end coordinates x1,i and x2,j: , with a similar result for side 2, where where N1 and N2 are the numbers of filaments at the two sides.
-
Actin polymerization stretches the top and side membranes of the cell elastically, with a spring constant K, as indicated by the purple springs in Fig. 1. The interaction force between the membrane and the actin filament network at is equal to the tension
(1) where L0 is the original length of the cell and(2) is the length after being stretched by the filaments. We do not include the effect of volume changes during the simulation on the actin concentration. Our additional tests showed that including this effect changed the critical parameter values for polarization by less than 5%.
Figure 1.
Schematic of model. Actin filaments are represented by red lines and the two nucleation regions by blue dashed lines. Springs mimicking the mechanical function of the cell membrane along the sides and top of the cell are shown in purple. The schematic is compared with a polarized neutrophil, where red labels polymerized actin and green labels tubulin [29] (https://www.london-nano.com/cleanroom-and-facilities/facilities/confocal-microscopes).
The stochastic approach is implemented via Monte-Carlo simulation of the polymerization, depolymerization and nucleation processes. We use a time-driven evolution algorithm with a constant time step Δt, rather the event-driven Gillespie algorithm [25], because this simplifies the updating of the continuous variable T. During each time step, uniformly distributed random numbers ξ between 0 and 1 are generated for each process.
-
A free actin will polymerize onto a barbed end if ξ is less than , where
(3) is the polymerization rate, kon is the on-rate constant, δ is the filament length increment per added actin subunit, and β is the inverse temperature. The exponential behavior comes from the analysis of Ref. [19]. This process is repeated for each filament. When polymerization occurs, G is reduced appropriately.
Barbed-end depolymerization will occur if ξ is less than , where is the barbed-end depolymerization rate. A subunit from the a barbed end is removed and G is increased. Pointed end depolymerization is treated similarly. When the last subunit of a filament depolymerizes, the filament is removed from the simulation.
Filament nucleation occurs if ξ is less than knΔt, where kn is the random nucleation rate. A number l0 of G-actins are removed from solution to form a new filament of length l0.
The nucleation rate kn is taken proportional to G2 as suggested by Ref. [26]: 1
| (4) |
where kr is a constant, A is the frontal area of the leading edge, and A0 = 0.4μm2 is the reference value of the area. The area scaling is based on the assumption that actin nucleators are uniformly distributed across the front edge of the cell. During the course of a given simulation, the frontal area is kept fixed. The above scaling was used to compare simulations for cells with different volumes. The new filaments are placed at the average barbed-end locations X1 and X2 with equal probability, and distributed uniformly parallel to the leading and trailing edges of the cell.
In reality, some filaments will push farther into the membrane than others, and one might expect some filaments to be left behind the moving membrane. However, these differences are reduced by membrane flexibility. Our previous studies of filament-membrane interactions (Ref. [27]) showed that filaments can grow while attached to the membrane. Trailing filaments being pulled by the membrane grow faster, and the ones that penetrate into the membrane grow slower, tending to equalize the positions of the filaments. We account approximately for flexibility by assuming that the filaments realign after each time step so that their barbed ends are at the same location as the membrane, placing them at X1 and X2 (see Fig. 1).
We have also considered an alternative model with explicitly flexible front and back membranes. Each filament has an individual harmonic force acting on its barbed end, which is evaluated from its extension beyond the original position of the unextended membrane. This does not cause any qualitative changes in the results. The same type of bifurcation is found in this case, and the effects of the bifurcation parameters described have the same sign. However, it is hard to compare these two treatments quantitatively since the membrane spring constant K must be very different in the two cases.
For l0, we consider both small and large values implementing different physical assumptions. The “small” value corresponds to the critical nucleus required to nucleate an actin filament. We choose l0 = 8 rather than commonly used values of 3–4, for practical reasons. If l0 is substantially smaller than 8, an unphysical scenario can occur in the stochastic simulations where all the filaments at the rear decay rapidly, so the extension vanishes. Then all the filaments will nucleate at the leading edge and experience zero opposing force. We avoid this scenario, in order to focus on the the interactions mediated by membrane force. For the “large” value, we choose l0 = 100. Here, the nucleation step in the model is viewed as including the first rapid steps in the filament life cycle in cells [20]. These include the creation of the filament and its growth until it is capped. The polymerization/depolymerization phase in our simulations corresponds to the last step in the life cycle, which is hydrolysis to ADP-actin and depolymerization.
This model is a first step in understanding the key interactions between mechanical force and actin polymerization. It is missing explicit treatment of several key features of importance in cells, including: i) barbed-end capping, which limits the length of filaments after they are nucleated, ii) ATP hydrolysis, which converts filament tips from growing to shrinking states (although hydrolysis it is included implicitly via the differences in critical concentrations between the barbed and pointed ends), and iii) autocatalytic filament nucleation [28], in which the rate of filament nucleation depends in the F-actin concentration. However, effects i) and ii) are included implicitly in our large-l0 model, and the effect of iii) is evaluated in the Discussion. The model is highly simplified, but this simplicity allows us to develop and test an analytic approach based on a combination of rate equations and bifurcation analysis. This provides a roadmap for the analysis and interpretation of more complex models in the future.
2.2. Rate-Equation Model
Our rate-equation approach attempts to mimic the stochastic simulations using three local variables at each end of the cell, and one global variable. Our derivation of the rate equations is based on a biased-diffusion description [30] of polymerization and depolymerization. A filament with l F-actin subunits can polymerize to length l + 1 or depolymerize to length l − 1. The length l is viewed as a continuous random variable, and its distribution function is described by the diffusion equation
| (5) |
where the index i denotes the region in the cell and the boundary conditions are ρi(l, t) = 0 for l = 0 or l → ∞. Here is the effective diffusion coefficient in region i and
| (6) |
is the effective convection coefficient. Eq. (5) differs from that given in Ref. [30] by the additional δ-function term, which describes the creation of new filaments with length l0.
We choose the local variables to be the zeroth, first and second moments of ρ1 and ρ2, as defined in Ref. [31], and the global variable to be the tension T. Note that the zeroth moment is just the number of filaments: Ni = ∫ρidl. Similarly, the first moment is the F-actin count Fi = ∫lρidl. Thus the free G-actin concentration is
| (7) |
where V is the volume of the cell and G0 is the initial free-actin concentration before polymerization begins.
The second moment Σi = ∫l2ρidl highlights a very important feature of the length distribution: its spread. We work with the standard deviation . The value of σi strongly affects the concentration of filaments near l = 0. Since these are the only filaments that can decay, this means that the decay rate of the filament number is strongly sensitive to σi.
Our equation set is:
| (8) |
| (9) |
| (10) |
| (11) |
| (12) |
| (13) |
| (14) |
where we have introduced the decay rate
| (15) |
and kn is given by Eq. (4).
Eq. (8) is derived as follows. The relations and , together with Eqs. (1) and (2), justify the first three terms. Pointed-end depolymerization does not appear in Eq. (8) because it does not affect the tension. The fourth term results from overcounting of barbed-end depolymerization in the first three terms. When a filament consisting of a single subunit depolymerizes, the filament disappears. In this case, the depolymerization does not affect the tension, which depends only on existing filaments (see Eq. (1)). Thus, the overcounted depolymerization contribution to Ṫ must be subtracted off. The overcounted depolymerization rate is . On the other hand, since ρi(0, t) = 0, ρi(1, t) ≃ ∂lρi(l, t) |l=0. Combining this with Eq. (15) gives the fourth term in Eq. (8).
The remaining equations are straightforwardly derived except for Eqs. (13), (14), and 15. Using Eq. (5), we obtain
| (16) |
Then Eq. (13) and Eq. (14) are obtained by integrating the first two terms in Eq. (16) by parts. Eq. 15 is obtained from the definition , by again taking ρ(1, t) ≃ ∂lρi(l, t) |l=0.
The contribution of nucleation to the dF/dt terms in Eqs. (11) and (12) gives corrections to the conventional formulas for the critical concentration Gc, where polymerization balances depolymerization [32]. For a bulk solution with no force acting on the filaments, and only one population N1 of filaments, Gc is determined by the conditions dF1/dt = dN1/dt = 0. Multiplying Eq. (9) by l0, and using Eq. (6), shows that , so that . The term lowers Gc, corresponding to the physical effect that nucleation removes free monomers from solution. When tension is are present, the polymerization rate is reduced and the critical concentration is increased. As we shall see later, the tension is higher when the cell is polarized, corresponding to a higher free-actin concentration.
Because our expressions involve ∂lρi(l, t) |l=0 in addition to the moments, it is necessary to estimate this quantity. For calculating steady-state solutions of the equations,∂lρi(l, t) can be obtained from the steady-state solution of Eq. (5). By matching the boundary conditions at l = 0, and l = ∞, and enforcing continuity at l = l0, we obtain
| (17) |
Note that the exponential in the second of Eqs. 17 is decaying because ui must be negative in steady state, to counter contributions to dFi/dt from nucleation.
From Eq. (17), we can calculate the three moments:
| (18) |
| (19) |
| (20) |
By solving for kn/ui and ui/Di in terms of Ni and Fi, we obtain
| (21) |
where li = Fi/Ni are the mean filament lengths (measured in subunits). Differentiating Eq. (21) and inserting into Eq. (15), we obtain the steady state decay rate:
| (22) |
We note that it is mathematically possible for the decay rate in Eq. (22) to become negative, if li becomes less than l0/2. This cannot occur in the stochastic treatment, because even in the extreme limit of no polymerization, li = l0/2. However, because the rate equations are approximate, it can happen in rare cases that li < l0/2. We avoid this by limiting ourselves to parameter values where the filaments are not too short.
The steady-state values of kd from Eq. (22), together with Eqs. (8–14), specify a dynamical system whose solution will give correct fixed points. However, it will not necessarily describe the dynamics of actin polymerization correctly, because the obtained by Eq. (22) are only guaranteed to be correct at steady state. Obtaining the exactly from Eq. (15) would require knowledge of all the moments of ρ, or another numerical approach to calculating ρ, both of which are impractical. Therefore, to obtain a practical approach, we truncate our description of ρ at the second moment. We devise a correction to Eq. (22) with a form based on the standard deviation σi and the average filament length li. We take the correction to have the functional form
| (23) |
where and are the steady state values of σi and li calculated from Eqs. (18), (19) and (20). This form is chosen for simplicity, and is guaranteed to give the correct values of at steady state. Fits to the decay rate obtained from the stochastic simulations for baseline values of the bifurcation parameters, plus an additional set using a value equal to 150% of its baseline value, obtained the best agreement for n = 3 and m = 1.6. Thus, the corrected decay rate reads:
| (24) |
The physical mechanism underlying the correction is shown in Fig. A1. The blue curve has a larger variance σ than the red one, although they have the same zeroth moment N and first moment F. The larger spread in the blue curve causes a larger value of ∂lρi(l, t) at the origin, increasing the filament decay rate according to Eq. (15).
In our implementation of this approach, we obtained n and m from a given parameter set, and tested the resulting method over a broad range of parameter values, from 50% to 150% of their baseline values. We found consistently improved agreement with the dynamic stochastic simulations (see Stochastic Results below), relative to using the steady-state values. The correction in Eq. (24) thus appears to be a broadly applicable approach. However, it is not essential for the the steady-state stability of the polarized vs. the unpolarized state.
3. Stochastic Results
The simulation parameters are given in Tables 1 and 2. Some of these are kept constant for all of the runs, while other key parameters, which can be adjusted experimentally, are tuned to drive symmetry-breaking transitions. K is chosen such that the time for the cell to equilibrate is moderate. We also chose K large enough to eliminate excessive volume changes. Typical simulation snapshots of unpolarized and polarized states are given in Fig. 2. The tuned parameters G0, , and kr are given relative to baseline values (see Table 2), which define the threshold between polarized and unpolarized states. They are in the same general range as existing estimates, but some of them differ from the values typically used to fit in vitro experiments (see Ref. [26]). Relative to standard parameters, the main difference is that our value of is larger and is smaller. This choice is made in order to maintain a reasonable filament length in our simplified model, while keeping the tension continuous during the transition from the symmetric to asymmetric state. Note that in the four-dimensional parameter space there are many possible choices of baseline values at the threshold. Our results are fairly independent of the particular choice of baseline values. For example, we have increased by 35% and by 20 %, which kept the system at the threshold. The critical values of the other parameters did not change, and the shape of the bifurcation plots was the same.
Table 1.
Values of constant parameters. K is the spring constant, kon is the polymerization rate, and δ is the length increment per monomer.
| Parameter | Value (for both small and large l0) |
|---|---|
| Temperature | 300K |
| K | 5.7pN/nm |
| kon | 11.6Mμ−1s−1 (Ref. [33]) |
| δ | 2.7 nm |
Table 2.
Baseline values of key parameters that are varied.
| Parameter | Baseline value (small l0) | Baseline values (large l0) | |
|---|---|---|---|
|
|
12μM | 12μM | |
|
|
0.05s−1 | 0.05s−1 | |
|
|
2.9s−1 | 2.9s−1 | |
|
|
70μM−2s−1 | 2.5μM−2s−1 |
Figure 2.

Filament distribution from simulations, for parameters giving a symmetric steady state (frame a), and an polarized steady state (frame b). Color usage is as in Fig. 1.
Figs. 3, 4, and A2–A4 present typical time courses of the filament count N and the polymerized-actin count F at the two ends of the cell. Figure 3 shows results for the unpolarized regime. At the beginning of the simulation, actin filaments nucleate and polymerize very rapidly (within about a second). The nucleation is fast because of the high initial G-actin concentration and the G2-dependence of the nucleation rate. In the small-l0 case, the cell polarizes after the initial period. Then the polarization decays to symmetric steady state results. The time scale of the convergence to the final symmetric state is slow because the only disassembly mechanism included in the model is depolymerization; disassembly by severing would lead to faster convergence. The dynamics of N are fairly similar to those of F. In the large-l0 case, the initial polarization phase is absent.
Figure 3.
Dynamics of F and N before reaching symmetric steady states, for (see Table 2). Other parameters are as in Tables 1 and 2. Frames a) and b) are for small l0, while frames c) and d) are for large l0.
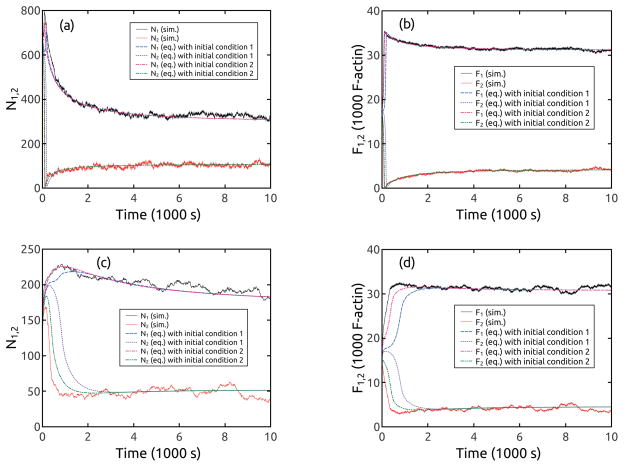
Figure 4.
Dynamics of F and N before reaching asymmetric steady states, for (see Table 2). Other parameters have the values given in Tables 1 and 2. Frames a and b are for small l0 and frames c and c are for large l0.
Increasing or decreasing the key parameters relative to the baseline values, while keeping the other parameters fixed, causes symmetry breaking. This is seen in Figs. 4 and (A2–A4), where individual parameters are varied from 50% to 150% of their baseline values while the others remain fixed. The polarization jumps rapidly over a period of a few hundred seconds, and then slowly drops to a finite value. The large-l0 results are similar to the small-l0 results. Polarization is favored by large G0, large , small , and small kr. Again, the dynamics of N are similar to those of F.
4. Rate-Equation Results
It is not possible for a deterministic rate-equation approach to precisely reproduce the dynamics leading to polarization from a symmetric starting point, since the polarization results from the growth of an initial stochastic fluctuation. Thus we use two sets of slightly asymmetric initial conditions to solve Eqs. (8–14). Set 1 has a minimal asymmetry: N1 = 2, N2 = 1, F1 = 2l0, F2 = l0, . Set 2 consists of stochastic-simulation values of T, N1,2, F1,2 and Σ1,2 evaluated (for the same parameter set) after 1 second of simulation time, so that the rate equations start at t = 1s. In this way the rate-equations treat the entire dynamics of polarization, except for its very initial stages. As seen in Figs. 3 and 4, the rate-equation results for small l0 match those from the stochastic simulation well for both the dynamic behavior and steady state values of F, independent of initial conditions. For large l0, the results are more sensitive to initial conditions. Using initial conditions 1 leads to noticeable delays in the polarization, while these delays are much smaller when using initial conditions 2.
To evaluate the necessity of the correction in Eq. (23) for accurate polymerization dynamics, Fig. 5 shows results for N1,2 with the correction (using Eq. (24)), and without the correction (using Eq. (22)), together with stochastic-simulation results for a typical polarized case. The rate-equation dynamics with the correction are much closer to the simulation dynamics. The curve with constant decay rate drops well below the stochastic-simulation curve at around 2000s, while the curve with the corrected decay state follows the stochastic-simulation curve closely. As a indicator of how methods typically used in the literature perform, we also show results for a constant decay rate set to be 50% greater than the average of the decay rates in the two regions at steady state. The polarization in both the initial period and steady state disappears completely. Other constant values of the decay rate also abolish the polarization. Thus the prediction of either steady-state or dynamic properties of polarization requires the use of a state-dependent decay rate.
Figure 5.
Dynamics of cell polarization, for different treatments of the filament decay rate kd. Parameters: (see table 2). Other parameters have the values given in Tables 1 and 2. Initial conditions are N1 = 2, N2 = 1.
5. Bifurcation Analysis of the Rate-Equation Results
Bifurcation analysis is an efficient method for understanding the qualitative behaviour of symmetry-breaking transitions, pinning down key interactions, and establishing possible long-term behaviors of a dynamical system [34]. By examining the stability of the solution of a dynamical system via diagonalization of a small matrix, one can find bifurcations and establish the key interactions responsible for the bifurcations. Such an instability analysis cannot be used directly for the stochastic simulations, but our development of a dynamical system in Eqs. (8–14) that mimics the simulation results allows us to perform a stability analysis of the unpolarized state. Because we seek only steady-state properties, we simplify the analysis by using the steady-state version of the decay rate from Eq. (22), so the variables Σ1,2 are not required. Fig. 6 shows the bifurcation diagram obtained from both the rate-equation calculation and the stochastic simulation. For each data point on the bifurcation diagram, both the simulation and rate equations were run for 50,000 seconds. The rate-equation result is the last data point, while the simulation result is the time average of the last 20,000–30,000 seconds’ data. The results from the two approaches match well, showing that the rate-equation method handles the bifurcation accurately. The continuous variation of the polarization as the various parameters are adjusted, together with the symmetry-breaking nature of the transition, suggests that the bifurcation is a supercritical pitchfork bifurcation [34]. Polarization is favored by increasing G0 and , and decreasing and kr.
Figure 6.
Bifurcation diagram of polarization as function of G0, , kr, using the small l0 value. The parameters are varied from to . In the simulation, each parameter is varied by 5% from dot to dot. In the rate equations, each parameter is varied by 1%, forming a smooth curve.
The symmetry breaking mechanism driving polarization results from an instability of the symmetric steady state to small perturbations. We perform a quantitative analysis of the instability, and identify the feedback loop that causes it. The instability is determined by the 5 × 5 Jacobian matrix [34] of the five variables T, N1,2 and F1,2, evaluated in a symmetric state. If all of the eigenvalues of this matrix are negative, the symmetric state is stable; if one or more is positive, the symmetric state is unstable. We force the system to be in a (potentially unstable) symmetric state by requiring that N1 = N2 and F1 = F2 as we solve the equations. If this restriction is lifted, for some parameter values the system will leave the symmetric steady state and polarize spontaneously. For clarity, we use symmetrized variables: N1, N2 → N = 1/2 (N1 + N2), ΔN = 1/2 (N1 − N2), and F1, F2 → F = 1/2 (F1 + F2), ΔF = 1/2 (F1 − F2). The Jacobian matrix is then
| (25) |
where the derivatives with respect to the new variables are
| (26) |
with similar expressions for ∂/∂F and ∂/∂ΔF.
The calculation is simplified by the decoupling of a 2 × 2 submatrix from the rest of the matrix. Since Eqs. (8–12) are invariant under the operation N1, F1 ↔ N2, F2 the variables can be chosen to be either symmetric or symmetric under this operation. Ṫ, Ṅ and Ḟ are symmetric under the operation, while ΔṄ and ΔḞ are asymmetric. Therefore the elements of  coupling ΔN and ΔF to the other variables vanish, so that
| (27) |
We define the lower right 2 × 2 sub-matrix of  to be
| (28) |
Numerical diagonalization of  reveals four eigenvalues that are always negative, and one that is positive for parameters that yield polarization. This eigenvalue is one of the eigenvalues of the â submatrix. Thus the symmetry-breaking instability is determined by â and independent of the other parts of Â. The interactions driving the instability can then be identified by examining the signs of the elements of â:
∂ΔṄ/∂ΔN < 0. This is derived in Appendix B.
∂ΔḞ/∂ΔF = 0. According to Eqs. (11) and (12), ΔḞ is not explicitly dependent on ΔF. The only dependence of these equations on F1 and F2 is via G, and G depends on the total F-actin number F, but not on ΔF.
∂ΔṄ/∂ΔF > 0. A positive ΔF corresponds to increasing F1 while keeping N1 constant. This causes the filaments in region 1 to be longer. The longer filaments will decrease the decay rate according to Eq. (22), and thus increase ΔṄ. As for case ii) above, G entering kn in Eqs. (9) and (10) does not depend on ΔF.
- Finally, two competing mechanisms determine ∂ΔḞ/∂ΔN, so it can be either positive or negative. Differentiation of Eqs. (11) and (12) shows that
(29) - The positive contribution comes from the first term in Eq. (29). Positive ΔN reduces the force per filament in region 1 while increasing the force per filament in region 2. Then the polymerization rate given by Eq. (3) in region 1 will be greater than that in region 2. The difference in the polymerization rates created by ΔN then increases ΔF, so the contribution to ∂ΔḞ/∂ΔN is positive.
- The negative contribution comes from the remaining terms in Eq. (29). Increasing ΔN increases the number of barbed and pointed ends in Region 1. At steady state, depolymerization exceeds polymerization because the sum of depolymerization and polymerization must cancel the actin subunits added by filament nucleation. Therefore, F-actin in region 1 will depolymerize faster than in region 2. Thus the contribution to ∂ΔḞ/∂ΔN is negative.
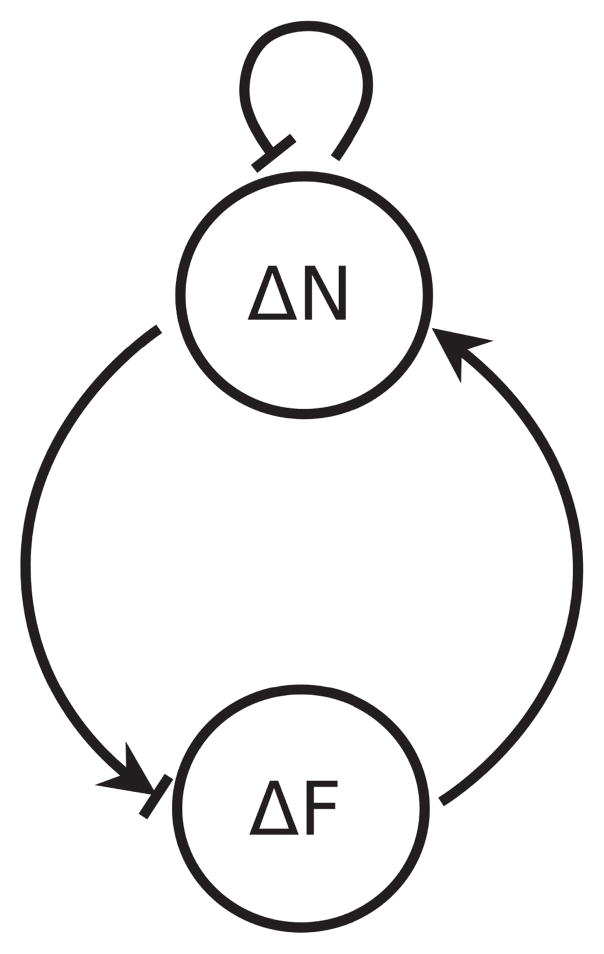
It is shown in Appendix B that polarization will occur only if ∂ΔḞ/∂ΔN > 0, so that the positive contribution exceeds the negative contribution. Thus the transition in the sign of this element drives the transition from a symmetric steady state to an asymmetric one. As shown under iv), this transition occurs when the interactions between N and F mediated by force become strong, which requires that the polymerization rate decreases with opposing force. The feedback loop driven by ∂ΔḞ/∂ΔN is illustrated in Fig. 7, where the arrow leading from ΔF to ΔN is always positive as discussed under iii), while the one from ΔN to ΔF can be either negative or positive depending on the magnitude of the tension T.
Figure 7.
Schematic of essential feedback loop that leads to polarization.
6. Discussion
In the previous section, we identified the key feedback loop in our model that destabilizes a symmetric steady state and thus causes polarization. This loop is based on three physical mechanisms relating ΔF to ΔN. Here we explore the relevance of the results obtained in this highly simplified model to the behavior of real cells, and analyze the parameter dependence in more depth.
First we compare the model predictions with the key experimental findings and previous theory. In Ref. [3], the authors laser-severed the body of a polarized cell from the leading edge and a new leading edge grew out of the cell body. To simulate this experiment, we cleaved a polarized cell in the model, cutting off the leading edge together with 70% of the original cell. Then the remaining cell body grew a new leading edge, consistent with Ref. [3]. We find that under this protocol, the new leading edge always is where the rear of the unsevered cell was previously. Of the two examples illustrated in Fig. 3 of Ref. [3] (one using laser severing and the other spontaneous severing), the leading edge in one case was at the old rear of the unsevered cell, and in the other case it was on the opposite side. If we use a different protocol, where the filaments in the cell “body” depolymerize after the cleavage, the new leading edge is randomly located, consistent with Ref. [3].
On the other hand, the model does not reproduce the phenomenology of Ref. [2], where imposition of force caused a transition from a symmetric state to a polarized state. In the model, initially symmetric cells always return to a symmetric state after imposition of force, as expected from the supercritical nature of the bifurcation. It may be that reproducing the observed behavior requires the inclusion of additional nonlinear mechanical terms as in Ref. [16].
As discussed in the Introduction, most previous theories of cell polarization have assumed a combination of diffusion and chemical reactions containing positive and negative feedbacks of proteins and lipds including Rac, PIP3, PTEN, and myosin. The model described here differs from these in that it does not assume explicit chemical feedback terms and requires no myosin activity. Rather, the positive-feedback terms that cause the polarization result from a combination of polymerization generating force, force regulating polymerization, and the effect of filament length on the filament decay rate. Of the models currently in the literature, the present model is closest in spirit to that of Ref. [15]. However, it differs from that model in that it does not require autocatalytic actin filament nucleation.
The mechanochemical feedback loop described here has some points in common with that discussed in Ref. [35] for focal adhesions. In that work, force was taken to regulate assembly of focal adhesions via a mechanism in which tensile stress lowers the chemical potential of assembled subunits relative to unassembled ones, enhancing assembly. In the present mechanism, force enters the feedback loop of Fig. 7 mainly via the effect of ΔN on ΔḞ. The reduction in the force per filament at one end of the cell that results from an increased ΔN causes ΔḞ to increase at that end as well. The main physics underyling this is the well-known Brownian-ratchet effect [19], in which compressive stress on the tip of a filament reduces the on-rate. We believe that this is a special case of the rigorous thermodynamic results described in Ref. [35].
Next we use our basic feedback loop to physically interpret the parameter dependences of polarization found in the simulation and rate-equation results. We found that polarization is favored by increased G0 and and reduced and kr. Eq. (29) shows that the key matrix element determining polarization is strongly dependent on T. Therefore, we based our analysis of the parameter dependence of polarization on T.
Increasing G0 increases the net barbed-end polymerization rate and thus T, favoring polarization.
Increasing increases G, which again increases T and favors polarization.
Decreasing increases the net polymerization rate of barbed ends, which increases T and thus favors polarization.
Decreasing kr causes more actin to be polymerized into filaments, and less to be used up in nucleation events. Because nucleation events by assumption do not generate tension (see Eq. (8)), focusing more of the free actin into polymerization increases T. This favors polarization.
Although the model is very simplified, some of these predicted trends might apply to cells where mechanical feedbacks are important. For example, G0 can be reduced by latrunculin treatment. Our finding that reduced G0 impairs cell polarization is thus consistent with the finding that latrunculin prevents polarization of HL-60 cells induced by a uniform attractant concentration [36]. We do not see how to vary and individually in a cell. However, actin disassembly is accelerated by cofilin, which might preferentially accelerate disassembly at one filament end relative to the other. Decreasing kr reduces the non-branching nucleation rate. Since formins are believed to cause non-branching nucleation, while Arp2/3 complex causes branching nucleation, downregulation of formin activity might correspond to reducing kr.
Finally, we explore the extent to which the basic picture embodied by Fig. 7, and criteria i) – iv) given above, will persist in more complete treatments of cell polarization:
-
∂ΔṄ/∂ΔN < 0. Since the decay rate Ṅ1 contains a term proportional to N1, this inequality will hold unless the nucleation rate on the “1”-side increases, or decreases, with increasing N1. An increase of the nucleation rate with N1 could result from autocatalytic nucleation by filament branching [28, 26]. It is not certain whether autocatalytic branching depends on N1 or on F1. In vitro, where actin nucleation factors are uniformly distributed in solution, branches occur along the length of the filament, suggesting a dependence on F1. However, in cells, branches are formed only near the membrane, so the filament length may not be an important factor. In this case the autocatalytic branching rate would depend on N1. If this causes ∂ΔṄ/∂ΔN to make a transition from negative to positive, and
 > 0, then Eq. (B.3) shows that the eigenvalues will make a transition from having negative real part with a nonzero imaginary part to having a positive real part with a nonzero imaginary part (a Hopf bifurcation [34]). Eigenvalues with positive real part and nonzero imaginary part correspond to a polarization that oscillates over time, as was observed in Ref. [37].
> 0, then Eq. (B.3) shows that the eigenvalues will make a transition from having negative real part with a nonzero imaginary part to having a positive real part with a nonzero imaginary part (a Hopf bifurcation [34]). Eigenvalues with positive real part and nonzero imaginary part correspond to a polarization that oscillates over time, as was observed in Ref. [37].A decrease in with increasing N1 could occur with a different decay mechanism. Refs. [10] and [17] suggested a “crushing” mechanism where filaments at the rear of the cell are broken by large force. Such a mechanism would be expected by have a decay rate roughly of the form , which decreases as a function of N1. This could also lead to a Hopf bifurcation to an oscillating state, under appropriate circumstances.
∂ΔḞ/∂ΔF = 0. The main correction to this result would come from steric crowding of actin, which would suppress polymerization where F is large and make ∂ΔḞ/∂ΔF negative. In this case, instability would still require a positive determinant for â, but now the determinant includes an extra contribution from the product of the diagonal terms. This means that ∂ΔḞ/∂ΔN would have to exceed a positive minimum critical value for polarization to occur.
∂ΔṄ/∂ΔF > 0. This should hold quite generally, since the filament lifetime should increase with its length. As mentioned above, another positive contribution to ∂ΔṄ/∂ΔF is present if side branching dominates. An additional possibility is the generation of new filaments by severing of existing filaments, which would provide a positive contribution provided that the severed fragments remain attached to the network [38, 39].
∂ΔḞ/∂ΔN has two competing contributions, a positive one from the force dependence of the polymerization rate, and the second one from the shorter lifetime associated with shorter filaments. These factors should also be present in real cells. However, some studies [40] have indicated that the polymerization velocity can have a force-independent plateau, and the positive term would be absent in this regime, abolishing polarization unless other positive-feedback effects become prominent.
We emphasize that obtaining criterion iii) depends crucially on having a state-dependent filament decay rate. If the decay rate is taken to be constant, then ∂ΔṄ/∂ΔF = 0, and there is no polarization unless autocatalytic branching or severing come into play.
The actual matrix of interactions describing a real cell is larger than that considered here. At the linear level asymmetric variables like ΔN and ΔF are still decoupled from symmetric variables. But other asymmetric variables are present, including those describing the distributions of upstream signaling proteins. In addition, beyond the linear level cell polarization might occur by a discontinuous change in several variables. In this case, changes in the symmetric variables could be coupled to changes in the asymmetric variables. However, the present analysis based on ΔN and ΔF provides a starting point for understanding the mechanical interactions driving cell polarization.
The model studied here treats only two networks, one at the front and one at the back of the cell, interacting with rigid boundaries. Extending the analysis to more general cell shapes with multiple networks, and deformable boundaries, could reveal several new types of phenomemena. For example, in a circular cell shape the orientation of the wave could gradually shift over time, rather than being constrained to discrete jumps of 180°. This could lead to motion that over long times is like a correlated random walk. Including membrane flexibility might allow traveling waves of polymerized actin to form along the edge of the cell, although such waves probably would require additional negative feedback mechanisms [41]. Treating a circular cell shape could be accomplished by two modifications. First, the main system variables, F and N, could be transformed into continuously varying functions along the cell periphery. Similarly, the tension would be transformed into a force density varying continuously along the membrane. It is likely that some terms penalizing excessive variations along the periphery would be required, so that the system would be described by partial differential equations. Achieving more realistic cell shapes requires the inclusion of membrane flexibility. This has been a challenge for cell migration modelers, but a promising class of approaches is based on the “phase-field” method [18].
7. Conclusion
We have systematically derived a rate-equation model of force-dependent actin polymerization from known biochemical processes and their force dependence. The polarization predicted by steady state solution of the rate equations agrees with our stochastic-simulation results to within 5%. A modification of the model was shown to treat dynamic effects accurately as well. The model results show that a polarized F-actin distribution can result from a a combination of force generated by polymerization, the slowing of polymerization by opposing force, and the dependence of the filament decay rate on the filament length. The feedback loop driving polarization results from reciprocal interactions between the asymmetry ΔN in the number of filaments between the two ends of the cell, and the corresponding asymmetry ΔF of the amount of polymerized actin. For appropriate parameter values, ΔN and ΔF grow synergistically, so that ΔN feeds the growth of ΔF, and vice versa. Polarization is favored by large values of the pointed-end off-rate and the free-monomer concentration, and small values of the barbed-end off-rate and random-nucleation rate. Obtaining correct polarization dynamics requires the use of information about the effect of the variance of the filament length on the filament decay rate. The improvements in rate-equation descriptions of actin polymerization described here should be useful in future efforts to model whole-cell structure and function, using methods such as the “Virtual Cell” approach [42].
Acknowledgments
This work was supported by the National Institutes of Health under Grant Number R01 GM107667.
Appendix A. Additional Figures
Below is the schematic of the length distribution, mentioned in the Model section.
Below we show the time courses leading to polarized steady states, for variations in the three key parameters and kr. For each parameter, both the small l0 and large l0 cases are considered.
Figure A1.
Schematic of two possible filament-length distributions, with the blue line having a larger standard deviation.
Figure A2.
Approach to polarized steady state for (see Table 2). Other parameters have the values given in Tables 1 and 2. Frames a) and b) are for small l0, while frames c) and d) are for large l0.
Figure A3.
Approach to polarized steady state for (see Table 2). Other parameters have the values given in Tables 1 and 2. Frames a) and b) are for small l0; frames c) and d) are for large l0. We do not plot the rate-equation result for the large l0 value, because in this case the average filament length in the rate equations becomes less than 0.5l0 at one end of the cell, so Eq. 22 for the decay rate breaks down.
Figure A4.
Approach to polarized steady state for (see Table 2). Other parameters have the values given in Tables 1 and 2. Frames a) and b) are for small l0, while frames c) and d) are for large l0.
Appendix B. Mathematical Derivations
Relation of polarization instability to the signs of the terms in â (Eq. 28)
We note that the eigenvalues of â are determined [34] by the trace
| (B.1) |
and the determinant
| (B.2) |
as:
| (B.3) |
The fact that ∂ΔṄ/∂ΔN < 0 and ∂ΔḞ/∂ΔF = 0, as proved below and under Discussion respectively, imply that τ < 0. Then Eq. (B.3) implies that â will have a positive eigenvalue, leading to instability, if and only if
 < 0, which occurs only if ∂ΔḞ/∂ΔN > 0, since ∂ΔṄ/∂ΔN < 0 and ∂ΔḞ/∂ΔF = 0.
< 0, which occurs only if ∂ΔḞ/∂ΔN > 0, since ∂ΔṄ/∂ΔN < 0 and ∂ΔḞ/∂ΔF = 0.
Proof that ∂ΔṄ/∂ΔN < 0. This follows from inserting Eq. (22) into Eqs. (9) and (10). Differentiating Eq. (9) minus Eq. (10) with respect to ΔN gives
| (B.4) |
Since N1 = N + ΔN and N2 = N − ΔN, it is clear that ∂N1/∂ΔN > 0 and ∂N2/∂ΔN < 0. Also, when Eq. (22) is written in terms of N1 and N2,
| (B.5) |
Similarly, . Combining the above inequalities shows that ∂ΔṄ/∂ΔN < 0.
References
- 1.Dyche Mullins R. Cytoskeletal mechanisms for breaking cellular symmetry. Cold Spring Harbor perspectives in biology. 2010;2(1):a003392. doi: 10.1101/cshperspect.a003392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Verkhovsky Alexander B, Svitkina Tatyana M, Borisy Gary G. Self-polarization and directional motility of cytoplasm. Current Biology. 1999;9(1):11–S1. doi: 10.1016/s0960-9822(99)80042-6. [DOI] [PubMed] [Google Scholar]
- 3.Houk Andrew R, Jilkine Alexandra, Mejean Cecile O, Boltyanskiy Rostislav, Dufresne Eric R, Angenent Sigurd B, Altschuler Steven J, Wu Lani F, Weiner Orion D. Membrane tension maintains cell polarity by confining signals to the leading edge during neutrophil migration. Cell. 2012;148(1):175–188. doi: 10.1016/j.cell.2011.10.050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Xiong Yuan, Huang Chuan-Hsiang, Iglesias Pablo A, Devreotes Peter N. Cells navigate with a local-excitation, global-inhibition-biased excitable network. Proceedings of the National Academy of Sciences. 2010;107(40):17079–17086. doi: 10.1073/pnas.1011271107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jiang Xingyu, Bruzewicz Derek A, Wong Amy P, Piel Matthieu, Whitesides George M. Directing cell migration with asymmetric micropatterns. Proceedings of the National Academy of Sciences of the United States of America. 2005;102(4):975–978. doi: 10.1073/pnas.0408954102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Iglesias Pablo A, Devreotes Peter N. Navigating through models of chemotaxis. Current opinion in cell biology. 2008;20(1):35–40. doi: 10.1016/j.ceb.2007.11.011. [DOI] [PubMed] [Google Scholar]
- 7.Onsum Matthew D, Rao Christopher V. Calling heads from tails: the role of mathematical modeling in understanding cell polarization. Current opinion in cell biology. 2009;21(1):74–81. doi: 10.1016/j.ceb.2009.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Jilkine Alexandra, Edelstein-Keshet Leah. A comparison of mathematical models for polarization of single eukaryotic cells in response to guided cues. PLoS computational biology. 2011;7(4):e1001121. doi: 10.1371/journal.pcbi.1001121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Mogilner Alex, Allard Jun, Wollman Roy. Cell polarity: quantitative modeling as a tool in cell biology. Science. 2012;336(6078):175–179. doi: 10.1126/science.1216380. [DOI] [PubMed] [Google Scholar]
- 10.Mogilner Alex, Zhu Jie. Cell polarity: tension quenches the rear. Current Biology. 2012;22(2):R48–R51. doi: 10.1016/j.cub.2011.12.013. [DOI] [PubMed] [Google Scholar]
- 11.Goehring Nathan W, Grill Stephan W. Cell polarity: mechanochemical patterning. Trends in cell biology. 2013;23(2):72–80. doi: 10.1016/j.tcb.2012.10.009. [DOI] [PubMed] [Google Scholar]
- 12.Turing AM. The chemical basis of morphogenesis. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 1952;237(641):37–72. [Google Scholar]
- 13.Dawes Adriana T, Edelstein-Keshet Leah. Phosphoinositides and rho proteins spatially regulate actin polymerization to initiate and maintain directed movement in a one-dimensional model of a motile cell. Biophysical journal. 2007;92(3):744–768. doi: 10.1529/biophysj.106.090514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.van Oudenaarden Alexander, Theriot Julie A. Cooperative symmetry-breaking by actin polymerization in a model for cell motility. Nature cell biology. 1999;1(8):493–499. doi: 10.1038/70281. [DOI] [PubMed] [Google Scholar]
- 15.Sambeth R, Baumgaertner A. Autocatalytic polymerization generates persistent random walk of crawling cells. Physical review letters. 2001;86(22):5196. doi: 10.1103/PhysRevLett.86.5196. [DOI] [PubMed] [Google Scholar]
- 16.Kozlov Michael M, Mogilner Alex. Model of polarization and bistability of cell fragments. Biophysical journal. 2007;93(11):3811–3819. doi: 10.1529/biophysj.107.110411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ofer Noa, Mogilner Alexander, Keren Kinneret. Actin disassembly clock determines shape and speed of lamellipodial fragments. Proceedings of the National Academy of Sciences. 2011;108(51):20394–20399. doi: 10.1073/pnas.1105333108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Shao Danying, Levine Herbert, Rappel Wouter-Jan. Coupling actin flow, adhesion, and morphology in a computational cell motility model. Proceedings of the National Academy of Sciences. 2012;109(18):6851–6856. doi: 10.1073/pnas.1203252109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Peskin Charles S, Odell Garrett M, Oster George F. Cellular motions and thermal fluctuations: the brownian ratchet. Biophysical Journal. 1993;65(1):316–324. doi: 10.1016/S0006-3495(93)81035-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Pollard Thomas D, Borisy Gary G. Cellular motility driven by assembly and disassembly of actin filaments. Cell. 2003;112(4):453–465. doi: 10.1016/s0092-8674(03)00120-x. [DOI] [PubMed] [Google Scholar]
- 21.Novak Igor L, Slepchenko Boris M, Mogilner Alex. Quantitative analysis of g-actin transport in motile cells. Biophysical journal. 2008;95(4):1627–1638. doi: 10.1529/biophysj.108.130096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.McGrath JL, Tardy Y, Dewey CF, Jr, Meister JJ, Hartwig JH. Simultaneous measurements of actin filament turnover, filament fraction, and monomer diffusion in endothelial cells. Biophysical journal. 1998;75(4):2070–2078. doi: 10.1016/S0006-3495(98)77649-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Svitkina Tatyana M, Borisy Gary G. Arp2/3 complex and actin depolymerizing factor/cofilin in dendritic organization and treadmilling of actin filament array in lamellipodia. The Journal of cell biology. 1999;145(5):1009–1026. doi: 10.1083/jcb.145.5.1009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Svitkina Tatyana M, Verkhovsky Alexander B, McQuade Kyle M, Borisy Gary G. Analysis of the actin–myosin ii system in fish epidermal keratocytes: mechanism of cell body translocation. The Journal of cell biology. 1997;139(2):397–415. doi: 10.1083/jcb.139.2.397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gillespie Daniel T. Exact stochastic simulation of coupled chemical reactions. The journal of physical chemistry. 1977;81(25):2340–2361. [Google Scholar]
- 26.Carlsson AE, Wear MA, Cooper JA. End versus side branching by arp2/3 complex. Biophysical journal. 2004;86(2):1074–1081. doi: 10.1016/S0006-3495(04)74182-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhu Jie, Carlsson AE. Growth of attached actin filaments. The European Physical Journal E. 2006;21(3):209–222. doi: 10.1140/epje/i2006-10061-9. [DOI] [PubMed] [Google Scholar]
- 28.Carlsson Anders E. Growth of branched actin networks against obstacles. Biophysical journal. 2001;81(4):1907–1923. doi: 10.1016/S0006-3495(01)75842-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Irimia Daniel, Charras Guillaume, Agrawal Nitin, Mitchison Timothy, Toner Mehmet. Polar stimulation and constrained cell migration in microfluidic channels. Lab on a Chip. 2007;7(12):1783–1790. doi: 10.1039/b710524j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Fujiwara Ikuko, Takahashi Shin, Tadakuma Hisashi, Funatsu Takashi, Ishiwata Shin’ichi. Microscopic analysis of polymerization dynamics with individual actin filaments. Nature cell biology. 2002;4(9):666–673. doi: 10.1038/ncb841. [DOI] [PubMed] [Google Scholar]
- 31.Doubrovinski K, Kruse K. Self-organization of treadmilling filaments. Physical review letters. 2007;99(22):228104. doi: 10.1103/PhysRevLett.99.228104. [DOI] [PubMed] [Google Scholar]
- 32.Oosawa Fumio, Asakura Sho. Thermodynamics of the Polymerization of Protein. Academic Press; New York: 1975. [Google Scholar]
- 33.Pollard Thomas D. Rate constants for the reactions of ATP-and ADP-actin with the ends of actin filaments. The Journal of cell biology. 1986;103(6):2747–2754. doi: 10.1083/jcb.103.6.2747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Strogatz Steven H. Nonlinear dynamics and chaos (with applications to physics, biology, chemistry and engineering) Perseus Publishing; Cambridge, Massachusetts: 2006. [Google Scholar]
- 35.Shemesh Tom, Geiger Benjamin, Bershadsky Alexander D, Kozlov Michael M. Focal adhesions as mechanosensors: a physical mechanism. Proceedings of the National Academy of Sciences of the United States of America. 2005;102(35):12383–12388. doi: 10.1073/pnas.0500254102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Wang Fei, Herzmark Paul, Weiner Orion D, Srinivasan Supriya, Servant Guy, Bourne Henry R. Lipid products of PI(3)Ks maintain persistent cell polarity and directed motility in neutrophils. Nature cell biology. 2002;4(7):513–518. doi: 10.1038/ncb810. [DOI] [PubMed] [Google Scholar]
- 37.Westendorf Christian, Negrete Jose, Bae Albert J, Sandmann Rabea, Bodenschatz Eberhard, Beta Carsten. Actin cytoskeleton of chemotactic amoebae operates close to the onset of oscillations. Proceedings of the National Academy of Sciences. 2013;110(10):3853–3858. doi: 10.1073/pnas.1216629110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Condeelis John. How is actin polymerization nucleated in vivo? Trends in cell biology. 2001;11(7):288–293. doi: 10.1016/s0962-8924(01)02008-6. [DOI] [PubMed] [Google Scholar]
- 39.Carlsson AE. Stimulation of actin polymerization by filament severing. Biophysical journal. 2006;90(2):413–422. doi: 10.1529/biophysj.105.069765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Heinemann Fabian, Doschke Holger, Radmacher Manfred. Keratocyte lamellipodial protrusion is characterized by a concave force-velocity relation. Biophysical journal. 2011;100(6):1420–1427. doi: 10.1016/j.bpj.2011.01.063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Allard Jun, Mogilner Alex. Traveling waves in actin dynamics and cell motility. Current opinion in cell biology. 2013;25(1):107–115. doi: 10.1016/j.ceb.2012.08.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ditlev Jonathon A, Vacanti Nathaniel M, Novak Igor L, Loew Leslie M. An open model of actin dendritic nucleation. Biophysical journal. 2009;96(9):3529–3542. doi: 10.1016/j.bpj.2009.01.037. [DOI] [PMC free article] [PubMed] [Google Scholar]