SUMMARY
Growing evidence indicates that non-neuronal mutant huntingtin toxicity plays an important role in Huntington’s disease (HD); however, whether and how mutant huntingtin affects oligodendrocytes, which are vitally important for neural function and axonal integrity, remain unclear. We first verified the presence of mutant huntingtin in oligodendrocytes in HD140Q knock-in mice. We then established transgenic mice (PLP-150Q) that selectively express mutant huntingtin in oligodendrocytes. PLP-150Q mice show progressive neurological symptoms and early death, as well as age-dependent demyelination and reduced expression of myelin genes that are downstream of myelin regulatory factor (MYRF or MRF), a transcriptional regulator that specifically activates and maintains the expression of myelin genes in mature oligodendrocytes. Consistently, mutant huntingtin binds abnormally to MYRF and affects its transcription activity. Our findings suggest that dysfunction of mature oligodendrocytes is involved in HD pathogenesis and may also make a good therapeutic target.
INTRODUCTION
Huntington’s disease (HD) is caused by polyglutamine expansion in the N-terminal region of huntingtin (Htt), a large protein that consists of 3141 amino acids. Despite the ubiquitous expression of mutant Htt in the brain and peripheral tissues, the major pathological feature of HD is selective neurodegeneration (Vonsattel, & DiFiglia, 1998; Munoz-Sanjuan & Bates, 2001). Similarly, selective neurodegeneration is also seen in many other neurodegenerative diseases, among them Alzheimer’s and Parkinson’s diseases, which suggests multiple factors may contribute to selective neurodegeneration. Given the known genetic mutation in HD and its well-characterized neuropathology, HD makes an ideal model for investigating how selective neuropathology can be caused by a disease protein that is widely expressed.
Most previous studies focused on the effect of mutant Htt on neuronal cells and revealed that N-terminal fragments of mutant Htt are pathogenic and cause cell-autonomous or non-cell-autonomous disease processes in a variety of animal models (Heng et al., 2008; Li and Li, 2012; Lee et al., 2013). In the brain, over 90% of cells are non-neuronal cells that provide essential support to the survival and function of neuronal cells. These non-neuronal cells consist mainly of three types of glial cells: astrocytes, microglial cells and oligodendrocytes. Glial dysfunction has been well documented to contribute to a variety of neurodegenerative diseases. For example, oligodendrocyte dysfunction plays an important role in ALS (Fünfschilling et al., 2012; Phillips et al., 2013; Kang et al., 2013). In HD patient brains, glial degeneration and pathology have also been documented (Rosas et al., 2003; Fennema-Notestine et al., 2004; Bartzokis et al., 2007; Di Paola et al., 2012; 2014). For example, myelin damage and breakdown were found in presymptomatic HD patients (Bartzokis et al., 2007; Phillips et al., 2014), and white matter defects in HD patients were found to associate with motor and cognitive deficits (Bohanna et al., 2011). Recent studies show that mutant Htt is expressed in glial cells and affects the function of astrocytes (Shin et al., 2005; Bradford et al., 2009; Tong et al., 2014) and microglial cells (Crotti et al., 2014). For example, as in neuronal cells, mutant Htt in astrocytes can impact multiple targets, including GLT-1 to affect glutamate uptake (Shin et al., 2005; Bradford et al., 2009) and K channel function (Tong et al., 2014) to increase striatal neuronal excitability and vulnerability. Moreover, deficient myelination is seen in HD mouse models (Wade et al., 2008; Xiang et al., 2011). Nonetheless, since deficient myelination can be caused by multiple factors including neuronal and non-neuronal toxicity, whether and how mutant Htt affects the function of oligodendrocytes remain to be investigated.
The importance of investigating mutant Htt’s effects in oligodendrocytes is supported by the critical role of oligodendrocytes in maintaining axonal function and early pathological changes in HD (Li et al., 2001; Wang et al., 2008; Bankston et al., 2013). Oligodendrocytes produce myelin, which electrically insulates axons, allowing rapid and efficient propagation of nerve signals. Defective oligodendrocyte function and deficient myelination are found in numerous neurodegenerative diseases (Bankston et al 2013). In HD knock-in mice that do not show obvious neuronal loss, axonal degeneration is an early pathologic event (Li et al., 2001). In transgenic HD monkey brain, axonal degeneration is also seen in the absence of cell body degeneration (Wang et al., 2008). Such axonal degeneration could be caused by mutant Htt in axons, as well as defective oligodendrocyte function. Investigating the effect of mutant Htt in oligodendrocytes will help us both understand the mechanism behind early disease pathology and develop more effective treatments.
We have established a transgenic mouse model that selectively expresses mutant Htt in oligodendrocytes. The PLP-150Q mice show obvious axonal degeneration and an early-onset polyQ disease phenotype that includes impaired rotarod performance, body weight loss and early death, providing strong evidence that mutant Htt in oligodendrocytes contributes to neuronal toxicity. Furthermore, we found that mutant Htt affects a newly identified transcription factor, MYRF or MRF, to reduce myelin gene expression in mature oligodendrocytes, offering new mechanistic insight into the neural dysfunction and axonal degeneration seen in HD.
RESULTS
Expression of mutant huntingtin in oligodendrocytes in HD knock-in mouse
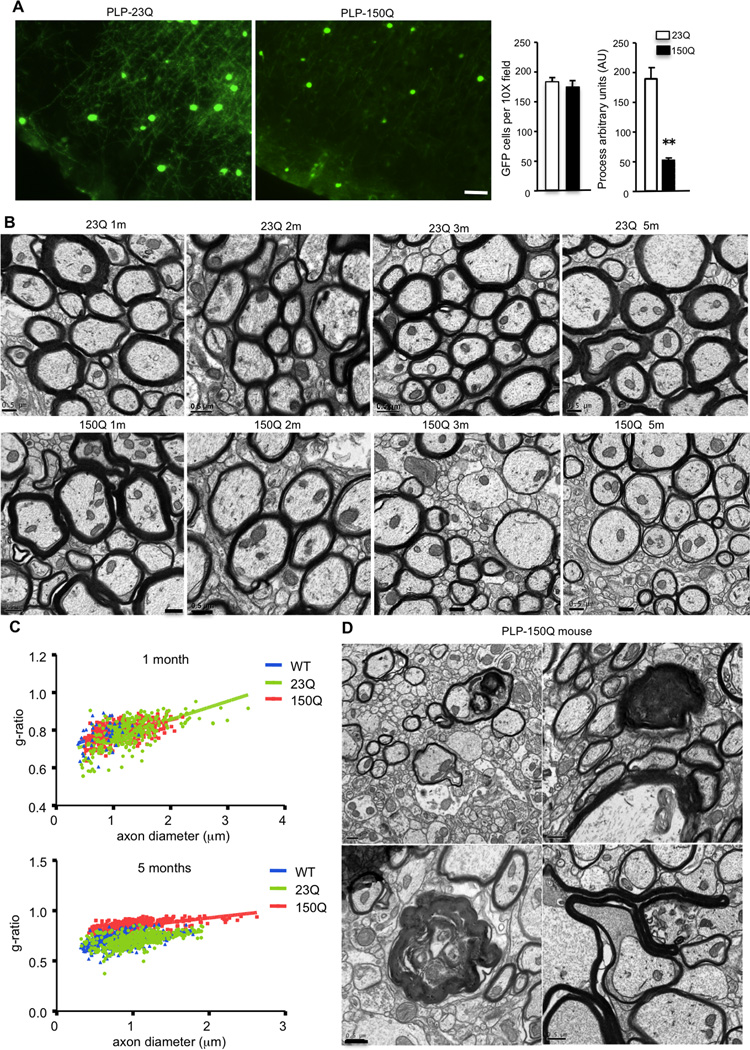
Although Htt is ubiquitously expressed in neuronal and non-neuronal cells, strong evidence for the presence of mutant Htt in oligodendrocytes is still lacking. To examine whether mutant Htt is expressed in oligodendrocytes at the endogenous level, we crossed HD 140CAG knock-in (KI) mice with transgenic mice that selectively express GFP in oligodendrocytes under the control of the proteolipid protein (PLP) gene promoter (Mallon et al., 2002), which drives transgene expression predominantly in oligodendrocytes in adult mouse brain (Michalski et al., 2011). The crossed mice would thus allow us to define whether full-length mutant Htt is expressed in GFP-positive oligodendrocytes. Immunofluorescent staining of the crossed mouse brains showed that GFP-positive oligodendrocytes (arrows in Figure 1A) do express mutant Htt, although their Htt staining is weaker than the neuronal staining of mutant Htt. Transgenic GFP expression in oligodendrocytes also allowed us to clearly visualize their processes. By comparing the GFP-positive processes in the cortex and striatum in wild-type and KI mice, we also saw the density of these processes reduced in the KI mouse brain (Figure 1B, supplemental Figure 1A), as well as short process length (Figure 1C). Since these GFP-positive processes represent myelinated axons and oligodendrocyte processes (Mallon et al., 2002), these results suggest that mutant Htt may affect the ability of oligodendrocytes to myelinate axons.
Figure 1. Expression of mutant Htt in oligodendrocytes in HD 140Q knock-in mouse brain.
(A) PLP-GFP mice were crossed with HD 140Q KI mice, and the brains of the crossed mice (PLP-GFP/KI) at one year of age were examined via immunofluorescent staining with anti-Htt (red). Arrows: GFP-positive oligodendrocytes (green) also express mutant Htt (red), though neuronal cells show more abundant Htt staining. (B) In the brain cortex sections that reveal processes of GFP-positive oligodendrocytes, GFP-positive processes are shorter in KI mouse brains. Length is reported in arbitrary units (AU). (C) Quantitative analysis of processes of GFP-positive oligodendrocytes in WT and KI mouse brain cortex. *** p<0.001. (3 mice per group). Scale bars: (A), 10 µm; (B), 5 µm.
Generation of PLP-150Q transgenic mice that selectively express mutant Htt in oligodendrocytes
To determine whether mutant Htt in oligodendrocytes affects their function and contributes to HD neuropathology, we set out to establish transgenic mice that express mutant Htt under the control of the PLP promoter. We expressed N-terminal mutant Htt (the first 212 amino acids), including 150 glutamines in the repeat region (Htt-150Q), based on the facts that proteolysis of mutant Htt is crucial for HD pathogenesis and that N-terminal mutant Htt is misfolded and causes more severe pathology in animal models (Ehrnhoefer et al., 2011; Ross et al., 2011; Lee et al., 2013). For the control, we also expressed the same N-terminal Htt with 23Q (Htt-23Q) in transgenic mice (Figure 2A). We obtained 9 positive Htt-150Q founders and 7 positive Htt-23Q founders. Western blot analysis with mEM48 showed that Htt-150Q forms aggregated Htt in the stacking gel, though its soluble level is lower than Htt-23Q in the brains of transgenic mice (Figure 2B). Using 1C2 antibody that reacts selectively with expanded polyQ repeats, we found that PLP-150Q is more abundant in the brain stem, striatum and corpus callosum, which contain more oligodendrocytes than the cortex and cerebellum, in three lines of transgenic Htt-150Q mice (150Q-1, 150Q-4a and 150Q-5). Consistent with the selective expression revealed in oligodendrocytes by the PLP promoter, there was no Htt-150Q detected in the peripheral tissues, including heart, kidney and muscle (Figure 2C). Immunohistochemical analysis of the brains of 150Q-4a mice at 1,2, and 3 months of age showed that mutant Htt is accumulated in the corpus callosum and striatum in an age-dependent manner (Figure 2D). Because of the similar expression levels of transgenic Htt in different mouse lines, we focused on the 150Q-4a (simply named PLP-150Q) and 23Q-4 (named PLP-23Q) lines for more detailed characterization.
Figure 2. Generation of PLP-HD transgenic mice.
(A) DNA construct of the vector used to generate transgenic mice expressing N-terminal Htt (1–212 amino acids) with 23Q or 150Q under the control of the PLP promoter. (B) EM48 western blot analysis of transgenic mouse brain regions (brain stem and striatum) expressing transgenic mutant Htt (150Q4a line) and normal Htt (23Q-2 and 23Q-4). Aggregated Htt in the stacking gel is indicated. (C) 1C2 western blot analysis of multiple PLP-150Q lines showing mutant Htt expression in different brain tissues, but not in peripheral tissues. (D) mEM48 immunohistochemical staining of the striatum of PLP-150Q mice at 1, 2, and 3 months of age showing an increased accumulation of mutant Htt in the older mice. Right panels show high magnification micrographs. Scale bars: 10 µm.
Immunohistochemical studies showed that mutant Htt is restricted in areas that are enriched in oligodendrocytes, such as the corpus callosum, and appears to be localized in the nuclei (Supplemental Figure 1B, C). Compared with HD140Q KI mice, in which mutant Htt is ubiquitously expressed in neuronal and glial cells, PLP-150Q mice showed the selective distribution of mutant Htt in glial cells and less abundant aggregates (Supplemental Figure 1D). Using primers specific for human Htt, we performed quantitative RT-PCR to compare the mRNA expression levels of transgenic Htt. Compared with mutant Htt in HD140Q KI mice at the endogenous level, transgenic Htt mRNA was slightly higher (Supplemental Figure 2A). However, western blotting revealed that transgenic mutant Htt in the striatum and brain stem is expressed at a lower protein level than mutant Htt in HD140Q KI mouse brain (Supplemental Figure 2B).
To verify that transgenic mutant Htt is indeed localized in the nuclei of oligodendrocytes, we crossed PLP-150Q mice with PLP-GFP mice. As expected, transgenic mutant Htt is abundantly distributed in the white matter region (corpus callosum) between the cortex and striatum and is localized in the nuclei of GFP-positive oligodendrocytes (Figure 3A). Using anti-olig2 and anti-Htt antibodies for immunofluorescent double labeling, we confirmed that mutant Htt accumulates and forms aggregates in the nuclei of oligodendrocytes in adult PLP-150Q mouse brain (Figure 3B). Importantly, none of these aggregate-containing cells could be labeled by antibodies to the astrocyte-specific protein, GFAP, or the neuronal protein, NeuN (Figure 3C). These results taken together show that we have established transgenic mice that express transgenic Htt specifically in mature oligodendrocytes.
Figure 3. Selective expression of mutant Htt in oligodendrocytes in adult PLP-150Q mice.
(A) PLP-150Q mice were crossed with PLP-GFP mice, resulting in PLP-150Q/GFP mice in which GFP-positive oligodendrocytes (arrows) express mutant Htt. Immunostaining with mEM48 (red) showing that mutant Htt is enriched in the GFP-positive cells in the white matter (corpus callosum) and also localized in the nucleus. (B) Double immunocytochemical staining with antibodies to Htt (mEM48) and Olig2 showing the selective expression of mutant Htt in oligodendrocytes in in the striatum of 3-month-old PLP-150Q mouse. (C) Double staining with antibodies to GFAP and NeuN revealed that mutant Htt is expressed in GFP-positive oligodendrocytes, but not in GFAP-positive astrocytes or NeuN-positive neuronal cells. Scale bars: (A), 40 µm; (B, C): 10 µm.
Progressive neurological phenotypes of PLP-150Q mice
Transgenic mice (PLP-23Q and PLP-150Q) generated from F1 were born normally. They can be bred normally as wild type mice and do not show obvious behavioral abnormalities and motor deficits during postnatal developmental stage and before 2 months of age. However, PLP-150Q mice develop age-dependent neurological symptoms starting at 3 months. We found that old PLP-150Q mice at age of 5 months display a clasping phenotype and reduced body weight compared with PLP-23Q mice (Figure 4A). PLP-150Q mice at 5 months of age also have tremors and difficulty walking (Supplemental video). The body weight of PLP-150Q mice began to decrease at the age of 4 months and continued to decrease progressively (Figure 4B). As a result, PLP-150Q mice started to die at 5 months of age and could not live longer than 10 months (Figure 4C). Consistent with their severe body weight loss and early death, PLP-150Q mice showed a significant deficit in their motor function as measured by rotarod performance, and this motor dysfunction became progressively worse with age (Figure 4D). We also compared different PLP-150Q mouse lines at a young age (2 months) and found that all of these mouse lines had poor rotarod performance (Figure 4E). The locomotor activity of PLP-150Q mice is also lower than PLP-23Q and wild-type mice during their active times zone (7 pm-11 pm) (Figure 4F). Since mutant Htt is expressed in oligodendrocytes, it may cause dysfunction of these cells and affect axonal conductivity, which may alter the susceptibility to seizure insults; thus, we next measured the latency of mice to generalized seizure induced by flurothyl and found that PLP-150Q mice were more sensitive to seizure induction (Figure 4G).
Figure 4. Progressive neurological symptoms of PLP-150Q mice.
(A) Representative photos of PLP-23Q and PLP-150Q mice, 6 months old. (B) Age-dependent loss of body weight in PLP-150Q mice; at least 15 animals per group. (C) Early death of PLP-150Q mice. The survival plot shows that PLP-150Q mice die at the age of 5–10 months, n=15. (D) Age-dependent worsening of rotarod performance of PLP-150Q mice compared with wild-type and PLP-23Q control mouse lines. At least 15 animals per genotype were examined. (E) Motor deficits in different PLP-150Q mouse lines compared with the PLP-23Q mouse line, n=5. (F) Locomotor activity was recorded for 24 hours. Statistical analysis shows reduced activity of PLP-150Q mice during the dark cycle, n=8–9, 2–3 months of age. (G) Increased susceptibility of PLP-150Q mice to flurothyl-induced seizures. At least 10 animals per genotype, 3–6 months of age. In (A, D-G), data are mean ± SEM. (E) and (G), one-way ANOVA, p<0.05. (B) and (F), two-way ANOVA, p<0.05. In (B), * significant compared to PLP-23Q; ** significant compared to PLP-23Q and WT.
Axonal degeneration and reduced myelin gene expression in PLP-150Q mouse brain
Next, we wanted to examine whether mutant Htt in PLP-150Q mice causes any myelination defect or axonal degeneration. The PLP-150Q/PLP-GFP crossed mice allowed us to examine the integrity of oligodendrocyte processes. We found that there is indeed reduced oligodendrocyte process length and no difference in the number of oligodendrocytes, in PLP-150Q mice (Figure 5A). Electron microscopy revealed that myelinated axons in the striatum of PLP-150Q and PLP-23Q mice at the age of 1 and 2 months are not distinguishable. However, at 3 months, there was a reduction of myelination in axons in PLP-150Q mice compared with PLP-23Q mice, and this reduction became more severe at 5 months (Figure 5B). Quantitative analysis of g-ratios (the inner axonal diameter to the total outer diameter) confirmed that mutant Htt significantly increased this ratio in PLP-150Q mice in an age-dependent manner (Figure 5C). It is well known that demyelination can lead to axonal degeneration (Rodriguez and Scheithauer, 1994; Sathornsumetee et al., 2000; Lee et al., 2014). Consistently, electron microscopy also revealed a number of degenerated axons, which appear swollen and dark, in PLP-150Q mice at 5 months of age (Figure 5D). The obvious axonal degeneration was not seen in 3-month-old PLP-150Q mice, suggesting that this degeneration occurs after demyelination (Supplemental Figure 2C). We also obtained HD patient brains that contain 45–53 CAG repeats in the HD gene (Supplemental Table-1). Although the ultrastructure of postmortem brain tissue was not well preserved, we were able to identify demyelinated axons in the HD patient brain compared with the control individual that did not die of neurodegenerative diseases (Supplemental Figure 2D).
Figure 5. Demyelination and axonal degeneration in PLP-150Q mice.
(A) Loss of oligodendrocyte processes in the brain cortex in PLP-150Q mouse. Quantitative analysis of the number of GFP-positive oligodendrocytes and process length is shown in the right panel. Oligodendrocytes in PLP-150Q mouse brain have significantly shorter processes than PLP-23Q. Process length is reported in arbitrary units (AU). Data are mean ± SEM. ** P<0.01. (B) Electron microscopic graphs of the striatum of PLP-23Q and PLP-150Q mice at 1, 2, 3, and 5 months of age. When PLP-150Q mice become old (3 and 5 months), they show demyelination compared with one-month-old PLP-150Q mice and the age-matched PLP-23Q mice. Scale bars: 0.5 µm. (C) G-ratios were calculated and plotted against axon diameter with linear regression. G-ratio is significantly increased in PLP-150Q striatum (g=0.7568±0.0068) compared to age-matched WT and PLP-23Q (g=0.6315±0.0099 and g=0.6083±.0095, respectively). One-way ANOVA, p<0.05. At least 327 axons per genotype were examined. (D) Axonal degeneration and demyelination were also seen in PLP-150Q mice at the age of 5 months. Scale bar: 0.5 µm.
Mutant Htt reduces the expression of myelin proteins
To understand how mutant Htt affects myelination, we performed microarrays aiming to identify molecules likely to be involved in Htt-mediated toxicity. Microarray analysis of gene expression in PLP-150Q-mouse spinal cord at 3 months revealed 998 significantly upregulated and 152 significantly downregulated genes in PLP-150Q compared to WT mice (Supplemental Figure 3A,B). Of these potentially altered genes, many are important for lipid metabolism, cell death and proliferation (Supplemental Figure 3C); their altered expression could be secondary responses to demyelination and axonal degeneration. Myelin synthesis requires cholesterol (Saher et al., 2011), and lipid metabolism is mediated by the transcription factor PGC1α(Camacho et al., 2013). Previous studies showed that the expression of PGC1a and the homeostasis of cholesterol are affected in HD mouse models (Xiang et al., 2011; Valenza & Cattaneo, 2011). Quantitative RT-PCR analysis of PGC1 α, cholesterol transport (StarD4) and sterol regulatory element binding transcription factor 2 (SREBF2) revealed no significant differences in their expression in the striatum of PLP-23Q and PLP-150Q mice (Supplemental Figure 3D). Further, using GFP-positive oligodendrocytes that were isolated via fluorescence-activated cell sorting from double transgenic mice expressing GFP and transgenic Htt in oligodendrocytes, we did not see significant differences in the expression levels of PGC1α, StarD4, and SREBF2 between WT, PLP-23Q, and PLP-150Q mice (Supplemental Figure 4A,B) It has been reported that MCT1, the monocarboxylate transporter 1 that is responsible for releasing lactate and ketone bodies from oligodendrocytes, is affected in ALS (Lee et al., 2012). We performed both quantitative RT-PCR and western blotting assays but could not see a significant defect in its expression in PLP-150Q mouse brains (Supplemental Figure 4C,D). Taken together, these results suggest that when mutant Htt is selectively expressed in oligodendrocytes, it may affect the expression of selective genes via different mechanisms.
Since there is a marked defect in myelination in PLP-150Q mice, we performed western blotting to detect the expression of myelin proteins, such as the oligodendrocyte-specific proteins 2', 3' cyclic nucleotide 3'-phosphodiesterase (CNP), myelin basic protein (MBP), myelin-associated oligodendrocytic basic protein (MOBP) and myelin oligodendrocyte glycoprotein (MOG), in the mouse brain and found that their expression is upregulated from postnatal day 14 (Figure 6 A). Importantly, mutant Htt does not affect the postnatal expression of myelin proteins at postnatal day 14. However, at 1 month, PLP-150Q mice showed decreases in the expression of these oligodendrocyte-specific proteins, in both the striatum and brain stem (Fig. 6B,C, Supplemental Figure 5A, B). Importantly, these decreases are age dependent and became more significant when PLP-150Q mice were 6 months old and were also seen in different PLP-150Q mouse lines (Figure 6, Supplemental Figure 5C). To further verify that the reduced expression of MBP is due to the effect of nuclear mutant Htt on gene transcription, we performed quantitative PCR. Results confirmed the reduction in MBP mRNA in PLP-150Q mouse brains (Figure 6D). To test if reduced MBP also occurs in the brains of HD patients, we performed both quantitative RT-PCR (Figure 6E) and western blot analyses (Figure 6 F , G). The results indicate that MBP is indeed decreased at both mRNA and protein levels in HD patient brains.
Figure 6. Reduced expression of myelin genes in PLP-150Q mice.
(A) Western blotting showing up-regulation of multiple myelin proteins (MBP, CNP, MOBP and MOG) in the mouse brains during postnatal days, which is not affected by mutant Htt. (B) Western blotting showing the age-dependent decrease of myelin proteins (MBP, CNP, MOBP and MOG) in the brain stem of adult PLP-150Q, but not PLP-23Q and WT, mice. Transcription factor MYRF for the myelin genes and oligodendroglial lineage marker olig-2 remain unchanged in PLP-150Q mouse brain. (C) Densitometric analysis of the relative levels of myelin proteins (ratio of myelin protein to GAPDH) in (B). (D) qPCR of MBP transcripts verified that MBP mRNA levels are reduced in PLP-150Q mouse brain (n=3/genotype). (A–D): n=3 per genotype and age. Data are mean ± SEM. Two-way ANOVA, p<0.05. (E, F) qPCR (E) and Western blot (F) analysis of the expression of MBP in the brain cortex of HD patients and control individuals (n=3 each group). (G) The relative levels of MBP were quantified by measuring the ratios of MBP to GAPDH on the western blots in (F). * p< 0.05; ** p<0.01.
Because there were no significant decreases in these proteins in the PLP-150Q mouse brains at 1 month, mutant Htt may not have an impact on these proteins during development, but may affect mature oligodendrocytes in adult mouse brains. In support of this idea, oligodendrocyte transcription factor (OLIG2), which is also expressed in undifferentiated oligodendrocytes and their precursors (Ligon et al., 2006; Meijer et al., 2012), is not affected significantly by mutant Htt. To further verify the selective effect of mutant Htt in adult mouse brains, we examined the expression of neuronal and glial marker proteins in postnatal mouse brains. Western blot results revealed no alteration of these proteins in PLP-150Q mouse brains at P1, P7, and P14 (Supplemental Figure 6A). Both Western blot and immunocytochemical analyses did not detect caspase-3 activation and increased GFAP either (Supplemental Figure 6A, B). Immunocytochemical analyses of NeuN and olig2 staining also support the western blot results, which demonstrate the unaltered expression of these neuronal and oligodendrocyte specific proteins in PLP-150Q mouse brains (Supplemental Figure 6C). By counting the number of Olig2 (WT: 268.000 ± 4.626, 23Q: 262.333 ± 8.939, 150Q: 272.333 ± 6.726, n=3 mice/per group) and NeuN (WT: 1263.333 ± 45.507, 23Q: 1292.667 ± 65.787, 150Q: 1201.333 ± 27.591, n=3 mice/per group) positive cells, we did not find any difference between WT, PLP-23Q, and PLP-150Q mice at P14. In addition, we did not find that mutant Htt affects proliferation of glial cells in the corpus callosum of PLP-150Q mice and differentiation of cultured oligodendrocytes from PLP-150Q mice (Supplemental Figure 7). All these results support the idea that mutant Htt affects the expression of myelin genes in an age-dependent manner, which is consistent with the aging-related and progressive symptoms of HD mice.
Mutant Htt binds N-terminal MYRF and affects its transcription activity
The decrease in several oligodendrocyte-specific proteins led us to focus on the transcription factors that mediate the expression of these proteins. Recently, myelin regulatory factor (MYRF) was found to regulate the expression of myelin-specific proteins (Emery et al., 2009; Hornig et al., 2013). MYRF is not expressed in oligodendrocyte precursors (McKenzie et al., 2014) but expressed only in postmitotic oligodendrocytes (Cahoy et al., 2008), and its depletion in oligodendrocytes causes failures in myelin gene expression and myelin formation (Emery, 2010). However, western blotting showed that MYRF is not decreased in the brains of PLP-150Q mice (Figure 6B). Because mutant N-terminal Htt is able to interact with a number of transcription factors in neuronal cells (Harjes et al., 2003; Li & Li, 2004), we wanted to know whether mutant Htt interacts with MYRF and affects myelin gene expression. To this end, we first generated the plasmid encoding mouse MYRF and expressed it in transfected HEK293 cells. We found that MYRF is unstable and cleaved to a smaller N-terminal fragment, which is labeled by an antibody against the N-terminal region of MYRF (Figure 7A). This finding is consistent with the recent report that MYRF can be self-cleaved (Bujalka et al., 2013; Li et al., 2013). Treatment of cells with the proteasome inhibitor MG132 significantly increased the expression levels of full-length MYRF and its cleaved products (Figure 7B). Using brain tissues of HD KI mice that express full-length mutant Htt at the endogenous level, we performed Htt immunoprecipitation and found that more N-terminal MYRF was also co-precipitated with mutant Htt (Figure 7C). To verify that this in vivo interaction also occurs in our transgenic Htt mouse brain tissues, we performed Htt immunoprecipitation and found that mutant Htt could co-precipitate more N-terminal MYRF (nMYRF) than full-length MYRF (Figure 7D).
Figure 7. Interaction of mutant Htt with MYRF.
(A) Comparison of in vitro synthesized and transfected MYRF showing that transfected full-length MYRF is cleaved to N-terminal MYRF (nMYRF) in cells. (B) Inhibition of MG132 markedly increased the levels of transfected full-length MYRF and nMYRF, suggesting that the ubiquitin-proteasome system degrades them. (C) Immunoprecipitation of mutant Htt from HD KI mouse brain selectively co-precipitates N-terminal MYRF (nMYRF). WT and IgG IP served as controls. (D) Immunoprecipitation of mutant Htt via mEM48 from PLP-150Q transgenic mouse brain also selectively co-precipitates N-terminal MYRF (nMYRF). Htt-150Q was preferentially labeled by 1C2 antibody on western blot (left panel). More nMYRF than full-length MYRF was precipitated with mutant Htt (right upper panel). Anti-GAPDH was also used to probe the immunoprecipitates (right low panel). (E) Expression of MYRF, nMYRF (1–635 aa) and cMYRF (616–1151 aa) in HEK293 cells. MYRF and nMYRF were detected with anti-myc, whereas cMYRF was detected by anti-flag. (F) Transfection of nMYRF and cMYRF into HEK293 cells revealed that only nMYRF is localized in the nucleus, supporting its role in gene transcription. Scale bars: 20 µm.
Although the above immunoprecipitations demonstrate the in vivo interaction of MYRF with mutant Htt, it remains unclear whether mutant Htt binds more N-terminal MYRF than normal Htt as the antibodies (mEM48 and 1C2) we used for immunoprecipitation and western blotting preferentially reacts with mutant Htt and precipitates more mutant Htt. To examine whether mutant Htt binds abnormally to MYRF, we used transfected cells to express N-terminal Htt and MYRF for studying their interactions. We generated N-terminal MYRF (nMYRF) tagged with the Myc epitope and C-terminal MYRF (cMYRF) tagged with the flag epitope (Figure 7E). Transfection of these two proteins in HEK293 cells confirmed that nMYRF is localized in the nucleus, and cMYRF is in the cytoplasm (Figure 7F). These results are in agreement with recent findings that nMYRF, once generated from the self-cleavage of full-length MYRF, is able to translocate into the nucleus to execute its transcription function (Bujalka et al., 2013; Li et al., 2013).
By co-expressing nMYRF or cMYRF with NLS-tagged N-terminal Htt (1–212 amino acids) containing 23Q or 150Q in HEK293 cells and performing immunoprecipitation of transfected MYRF, we found that more Htt-150Q was precipitated by nMYRF when the same levels of Htt-23Q and Htt-150Q were present in the input (Figure 8A). In contrast, immunoprecipitation of cMYRF failed to precipitate transfected Htt (Figure 8B). These results support the in vivo interaction of mutant Htt with N-terminal MYRF in HD mouse brains (Figure 7C,D). To further verify that mutant Htt can interact with MYRF directly, we performed an in vitro binding assay using in vitro synthesized Htt incubated with GST-nMYRF fusion proteins. The result also showed that more mutant Htt bound to GST-nMYRF than normal Htt (Figure 8C). By expressing different fragments of MYRF with mutant Htt in transfected cells and co-immunoprecipitation, we found that N-terminal MYRF containing the proline-rich domain interacts with Htt and that polyQ expansion could increase this interaction (Supplemental Figure 8A–D). Since N-terminal MYRF has transcriptional activity, the binding of mutant Htt to the N-terminal region of MYRF also supports the idea that this interaction affects the transcription activity of MYRF. Also, the interaction between mutant Htt and MYRF suggests that soluble mutant Htt binds MYRF, which is supported by immunofluorescent staining of transfected cells in which soluble MYRF is diffuse with mutant Htt in the nucleus, but is not colocalized with Htt aggregates (Supplemental Figure 8E).
Figure 8. Mutant Htt binds N-terminal MYRF and inhibits its transcription activity.
(A) Co-transfection of nMYRF (left panel) with N-terminal Htt containing 23Q or 150Q into HEK293 cells and immunoprecipitation of nMYRF. More Htt-150Q binds nMYRF than Htt-23Q. Co-transfection of cMYRF (right panel) with N-terminal Htt containing 23Q or 150Q into HEK293 cells showing no interaction between cMYRF and Htt. (B) The ratio of precipitated to input in (a) is shown. (C) GST-nMYRF and GST were expressed and purified to incubate with in vitro synthesized N-terminal Htt containing 23Q and 150Q also demonstrating that mutant Htt directly binds more MYRF. (D) MBP promoter was co-expressed with MYRF and Htt to assess its transcription activity via luciferase assay. MYRF markedly promotes MBP promoter activity. Co-expression of N-terminal Htt with 23Q did not significantly (p>0.05) affect the MBP promoter activity, whereas Htt-150Q significantly (p<0.001) reduces the MBP promoter activity. * p<0.05; *** p<0.001. (E) Expression of nMYRF, but not cMYRF, could rescue the inhibitory effect of mutant Htt on the MBP promoter activity. * p<0.05; ** p<0.01, *** p<0.001. (F) qPCR quantification of the MBP promoter DNAs associated with Myc-nMYRF that was immunoprecipitated by anti-Myc in ChIP assay. The results were obtained from three independent experiments. ** p=0.019. (G) A proposed model for the effect of mutant Htt in oligodendrocytes. According to recent studies (Bujalka et al., 2013; Li et al., 2013), full-length MYRF is self-cleaved to N-terminal MYRF (nMYRF), which is dissociated from ER and translocalized to the nucleus to activate the expression of myelin genes. In HD, the accumulation of N-terminal mutant Htt in the nucleus can lead to the abnormal binding of nMYRF and affects its transcription activity, leading to reduced expression of myelin genes and oligodendrocyte dysfunction.
Based on the above findings, we went on to test whether mutant Htt affects the transcription activity of MYRF. For this assay, we isolated the promoter region of the mouse MBP gene based on an earlier finding (Mack et al., 2007) and linked this promoter with the luciferase reporter. An assay of luciferase reporter activity showed that MYRF significantly increased the MBP promoter activity, which was inhibited by mutant Htt (Figure 8D). Importantly, nMYRF, but not cMYRF, could also stimulate the MBP promoter activity and reverse the inhibition caused by mutant Htt (Figure 8E). Via chromatin immunoprecipitation (ChIP) assay, we immunoprecipitated nMYRF associated-DNAs in the presence of normal and mutant Htt and measured the associated MBP promoter DNAs using quantitative PCR. We found that mutant Htt significantly inhibited the association of nMYRF with the MBP promoter (Figure 8F). These results support the idea that mutant Htt binds nMYRF to affect its transcription activity by altering its association with DNAs. Based on the fact that MYRF is self-cleaved to nMYRF, which is then translocated into the nucleus to activate myelin gene expression (Bujalka et al., 2013; Li et al., 2013), we propose a model in which N-terminal mutant Htt accumulates in the nuclei of oligodendrocytes in an age-dependent manner. This age-related accumulation leads mutant Htt to bind nMYRF in mature oligodendrocytes and affects its transcription activity, resulting in decreased expression of myelin genes and an associated myelination deficiency and axonal degeneration in HD (Figure 8G).
DISCUSSION
By establishing transgenic mice that express mutant Htt selectively in oligodendrocytes, we provide strong evidence for the important contribution of mutant Htt in oligodendrocytes to HD neuropathology. We also show that mutant Htt can affect cell type-specific gene expression to reduce myelin gene expression, leading to axonal degeneration. In addition, we find that mutant Htt affects MYRF transcription activity via its abnormal association with MYRF, providing mechanistic insight into the specific effect of mutant Htt on a particular type of non-neuronal cell.
Although transgenic mice expressing N-terminal mutant Htt are valuable models for exploring the pathogenesis of HD, whether their pathological characteristics can be replicated in HD mouse models that express full-length mutant Htt merits discussion here. All full-length Htt HD mouse models, transgenic or knock-in, show milder disease progression than transgenic mice expressing N-terminal mutant Htt (Menalled & Chesselet, 2002; Heng et al., 2008). Because only N-terminal mutant Htt is able to misfold and form aggregates, proteolysis of full-length mutant Htt has been widely held to be an important process toward HD pathogenesis (Ehrnhoefer et al., 2011; Lee et al., 2013; Ross et al., 2014). In this regard, the expression of N-terminal mutant Htt can facilitate disease progression and yield robust phenotypes in mice, allowing us to identify pathogenic pathways and evaluate therapeutic targets (Menalled & Chesselet, 2002; Heng et al., 2008). One good example is that young R6/2 mice at 3–4 months of age, which express exon1 mutant Htt, recapitulate many pathological changes seen in the end stage of HD KI mice at 2 years of age (Woodman et al., 2007).
Mutant Htt affects the function of neuronal and glial cells in the brain. Since HD pathogenesis involves cell-autonomous and non-cell-autonomous toxicity of mutant Htt (Gu et al., 2005), generating animal models with cell type-specific expression of mutant Htt is important to dissect HD pathogenesis. In our PLP-150Q transgenic mice, we saw that mutant Htt in oligodendrocytes forms aggregates in their nuclei, interacts with the oligodendrocyte-specific transcription factor, MYRF, and reduces myelin gene expression. All these important pathogenic events were also seen in HD KI mouse brains. Moreover, the effect of mutant Htt on axons in PLP-150Q mice is consistent with our early findings that axonal degeneration is seen in the brains of a HD KI mouse model (Li et al., 2001) and transgenic monkey brains (Wang et al., 2008). Oligodendrocytes and myelin abnormalities can slow or stop fast axon transport, resulting in synaptic loss and axonal degeneration (Han et al., 2010). Our PLP-150Q mouse model supports the idea that axonal degeneration can also be caused by the non-cell-autonomous toxicity of mutant Htt in oligodendrocytes.
That said, however, PLP-150Q mice develop much more progressive and severe phenotypes, even when mutant Htt is selectively expressed in oligodendrocytes. Such severe phenotypes are unlikely to be due to the overexpression of N-terminal mutant Htt since the level of Htt-150Q is lower than the control transgenic Htt-23Q, which did not elicit any neurological phenotypes. Also, other transgenic HD mouse models expressing N-terminal mutant Htt at levels lower than endogenous mouse Htt in either astrocytes or axonal terminals show early death and progressive phenotypes (Bradford et al., 2009: Xu et al., 2013). All these findings suggest that, once N-terminal mutant Htt accumulates in oligodendrocytes, it can greatly facilitate disease progression.
Our identification of the effect mutant Htt has on MYRF-mediated myelin gene expression is consistent with the findings that N-terminal mutant Htt can interact with a number of transcription factors and affect gene transcription (Harjes & Wanker, 2003; Li & LI, 2004). A new insight from our study is that the transcription dysfunction in oligodendrocytes is both cell type-specific and protein context-dependent. Our findings also suggest that mutant Htt affects the function of mature oligodendrocytes. The accumulation of N-terminal mutant Htt in the nuclei of mature oligodendrocytes in adult mice is age-dependent, as evidenced by the following facts: (1) PLP-150Q mice show increased accumulation of mutant Htt in the adult brain with aging; (2) PLP-150Q mice display demyelination and reduced myelin gene expression in an age-dependent manner; (3) PLP-150Q mice develop age-dependent and progressive neurological symptoms. Perhaps in mature oligodendrocytes, mutant Htt is prone to accumulating in their nuclei, thereby affecting gene expression. Consistent with these facts, we also found that mutant Htt did not affect the survival of neuronal cells and oligodendrocyte proliferation during early postnatal development. Although the PLP promoter was found to drive gene expression in different types of cells during embryonic and early developmental stages, transgenic gene expression under this promoter becomes restricted to oligodendrocytes in adult mouse brain (Michalski et al., 2011). Mutant Htt-mediated toxicity in oligodendrocytes requires an age-dependent accumulation of misfolded Htt in cells. It is likely that mature oligodendrocytes are less capable of clearing misfolded N-terminal Htt than undifferentiated precursor cells. Thus, the expression of N-terminal mutant Htt, which is prone to misfolding, even at a low level could greatly facilitate the accumulation of the toxic form of mutant Htt in oligodendrocytes, whereas in HD KI mice, it takes a much longer time to accumulate N-terminal mutant Htt after being generated by the proteolysis of full-length mutant Htt.
The idea that mutant Htt affects the expression of selective genes in mature oligodendrocytes is also supported by the fact that mutant Htt does not reduce the expression of OLIG2, which is predominantly expressed in oligodendrocyte precursor cells and undifferentiated oligodendrocytes. Since OLIG2 is also present in mature oligodendrocytes and MYRF is restricted to mature oligodendrocytes (Cahoy et al., 2008; Emery 2009), the negative impact of mutant Htt on OLIG2 is consistent with the finding that OLIG2 expression is not mediated by MYRF (Emery 2009) and indicates a specific effect of mutant Htt on MYRF. Another line of evidence to support the specific effect of mutant Htt in oligodendrocytes is the negative impact of mutant Htt on the expression of MCT1. MCT1 transports lactate from oligodendrocytes to support neuronal survival and was found to be reduced in oligodendrocytes in ALS mouse brains (Lee et al., 2012). Since mutant SOD1 in ALS is largely expressed in the cytoplasm, the cytoplasmic effect of mutant SOD1 may reduce the expression of MCT1, which is different from the nuclear effect of mutant Htt, via a different mechanism.
Because oligodendrocytes are critical for axonal function and are also a desirable transplant population for remyelination strategies, the findings from our study provide a new therapeutic target for treating HD. Oligodendrocytes are generated mostly during the postnatal periods (P7-P14) in rodents. These cells differentiate from proliferative, migratory oligodendrocyte precursors (OPCs) and produce myelin proteins required for myelination. Although most myelination occurs early in life, myelination processes may exhibit substantial plasticity in adult brains and have evolved to combat a range of infectious, metabolic and immune insults, which can cause demyelination and lead to multiple sclerosis (Paus et al., 1999; Franklin et al., 2008; Emery 2010). A recent study demonstrates that depletion of MYRF prevents adult myelination, reduces the production of new oligodendrocytes during adulthood, and affects motor skill learning in mice (McKenzie et al., 2014). Newly differentiated oligodendrocytes are widely thought to generate additional myelin (Franklin et al., 2008; Emery 2010), and based on this idea, approaches to promote differentiation of OPCs into myelinating oligodendrocytes have been hotly pursued for the treatment of demyelination-related pathology (Rivers et al., 2008; Boulanger & Messier, 2014). Our studies indicate that mutant Htt affects mature oligodendrocytes to cause a decrease in myelin proteins and subsequent loss of myelin, suggesting that the production of myelin proteins in mature oligodendrocytes is important for the maintenance of myelin in adult brains, as well (Koenning et al., 2012). This idea is also supported by the finding that mutant Htt does not reduce Olig2, but rather affects the function of MYRF, which is expressed only in postmitotic oligodendrocytes (Cahoy et al., 2008; Emery et al., 2009; McKenzie et al., 2014). The findings from our study therefore suggest that improving the function of mature oligodendrocytes should at least be beneficial in HD, and perhaps other age-dependent neurodegenerative diseases that involve the dysfunction of mature oligodendrocytes and associated axonal dysfunction, as well.
EXPERIMENTAL PROCEDURES
Animals
To generate PLP-Htt transgenic mice, cDNA encoding N-terminal human Htt (amino acids 1–212) containing 23Q or 150Q was subcloned into the pNEB193 vector using BamHI and XbaI. The plasmid was then digested with PmeI and AscI, and the Htt-containing fragment was inserted into the PLP promoter cassette. The pNEB193 vector and PLP promoter cassette were generated previously (Fuss et al., 2000). Pronuclear injection of the linearized construct was performed by the Transgenic Mouse and Gene Targeting Core at Emory University. Founder mice were obtained and maintained at the Emory mouse facility. Genomic DNA was isolated from mouse tails, and PCR genotyping with primers that flank the polyQ region was used to screen for transgenic mice. The sequences of the primers are as follows: forward: 5′-ATGAAGGCCTTCGAGTCCCTCAAGTCCTTC-3 ′ , and reverse, 5 ′ -AAACTCACGGTCGGTGCAGCGGCTCCTCAG-3′. We identified 9 positive PLP-150Q founders and 7 positive PLP-23Q founders. Transgene expression was determined via western blotting. PLP-GFP mice were obtained from Dr. Wendy Macklin (University of Colorado Denver) and bred with PLP-Htt mice to generate double-transgenic mice. Full-length mutant Htt CAG140Q (HD KI) mice were provided by Dr. Michael Levine (University of California, Los Angeles, CA).
Mouse behavior analysis
Mouse body weight was measured once every month, and survival was monitored regularly. The motor function of the mice was assessed with the rotarod test (Rotamex, Columbus Instruments). Mice were trained on the rotarod at 5 RPM for 5 min for 3 consecutive days. After training, the mice were tested for 3 consecutive days, 3 trials per day. The rotarod gradually accelerated to 40 RPM over a 5-min period. Latency to fall was recorded for each trial. At least 15 mice were analyzed per group when comparing PLP-150Q, PLP-23Q and WT. When comparing all PLP-150Q lines, at least 5 animals were analyzed per group.
The flurothyl-induced seizure experiment was performed as previously described44 Mice were placed in a clear Plexiglas chamber and exposed to flurothyl (2,2,2-trifluroethylether) (Sigma, 287571) at a rate of 20 µl/min. Latency to generalized seizure was recorded. Mice were at least 3 months old, and at least 10 animals were analyzed for each group. Locomotor activity was assessed using an automated system (PAS, San Diego Instruments) that records all photobeam interruptions. Each mouse was housed in a separate chamber with food and water. Activity was recorded once every 30 min for 30 h. The first 6 h were excluded from analysis. At least 8 animals were analyzed per group.
Immunofluorescence, immunohistochemistry and electron microscopy
Mice were anaesthetized with 5% chloral hydrate and perfused with 0.9% NaCl followed by 4% paraformaldehyde (PFA). The brains were removed and post-fixed in 4% PFA overnight at 4°C. The brains were transferred to 30% sucrose for 48 h, then cut to 20-or 40-µm sections with the cryostat (Leica CM1850) at −20°C. Sections were blocked in 4% donkey serum with 0.2% Triton X-100 and 3% BSA in PBS for 1 h. For immunofluorescent staining, 20-µm sections were incubated with primary antibodies in the same buffer at 4°C overnight. After washing with 1X PBS, the sections were incubated in fluorescent secondary antibodies and Hoechst stain. Fluorescent images were acquired with a Zeiss microscope (Carl Zeiss Imaging, Axiovert 200 MOT) and either a 40x or 63x lens (LD-Achroplan 40x/0.6 or 63x/0.75) with a digital camera (Hamamatsu, Orca-100) and Openlab software (Improvision). For immunohistochemistry with DAB staining, after blocking, 40-µm sections were incubated with mEM48 for at least 48 h at 4°C. A biotin/avidin immunoassay (Vector Laboratories) and DAB kit (Invitrogen, 00–2020) were used. Images were acquired with a Zeiss microscope (Carl Zeiss Imaging, Imager A.2) and either a 40x or 63x lens (Plan-Apochromat 40x/0.95 or 63x/1.4) with a digital camera (Carl Zeiss Imaging, AxioCam HRc) and AxioVision software.
For electro-microscope (EM) study, mice were perfused with 0.9% NaCl followed by 4% PFA containing 2.5% glutaraldehyde. After post-fixation, the brain was cut to 50-µm sections using the vibratome (Leica, VT1000). The sections were processed as previously described (Wang et al., 2008). Axon and myelin fiber diameters were measured using ImageJ (NIH). More than 300 axons were examined for each genotype.
Statistical analysis
Results are expressed as mean ± SEM. Prism6 (GraphPad Software) was used for statistical analysis. When only two experimental groups were compared, the Student’s t-test was used to calculate statistical significance. For all other experiments, statistical significance was calculated using one-way ANOVA or two-way ANOVA, followed by Tukey’s multiple comparisons test. A p-value <0.05 was considered significant.
Supplementary Material
ACKNOWLEDGEMENTS
This work was supported by grants from the National Institutes of Health (NS036232, NS041669 and NS045016, AG019206, AG31153) and the State Key Laboratory of Molecular Developmental Biology, China, We also thank Hong Yi of the Robert P. Apkarian Integrated Electron Microscopy Core at Emory University for her help with electron microscopy. Research reported in this publication was supported in part by the Emory Integrated Genomics Core (IEGC) Shared Resource of Winship Cancer Institute of Emory University and NIH/NCI under award number P30CA138292. Human tissue samples were provided by the Harvard Brain Tissue Resource Center, which is supported in part by PHS grant number MH/NS 31862, as well as Emory Alzheimer’s Disease Center Brain Bank (P50-AG025688). We thank Cheryl Strauss for critical reading of the manuscript.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
SUPPLEMENTAL INFORMATION
Supplemental Information includes eight figures, one table, one movie, and supplemental experimental procedures.
AUTHOR CONTRIBUTIONS
B.H. and W.W. contributed equally to this work. X-J.L. and S.L. designed experiments; B.H., W.W. and G.W. performed experiments and analyzed data; M.A.G. helped with maintaining mice; Y.F. provided advice and transgenic PLP-GFP mice and read the paper; B.H., W.W., S.L. and X-J.L. wrote the paper.
REFERENCES
- Bankston AN, Mandler MD, Feng Y. Oligodendroglia and neurotrophic factors in neurodegeneration. Neurosci Bull. 2013;29:216–228. doi: 10.1007/s12264-013-1321-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartzokis G, Lu PH, Tishler TA, Fong SM, Oluwadara B, Finn JP, Huang D, Bordelon Y, Mintz J, Perlman S. Myelin breakdown and iron changes in Huntington's disease: pathogenesis and treatment implications. Neurochem Res. 2007;32:1655–1664. doi: 10.1007/s11064-007-9352-7. [DOI] [PubMed] [Google Scholar]
- Bohanna I, Georgiou-Karistianis N, Sritharan A, Asadi H, Johnston L, Churchyard A, Egan G. Diffusion tensor imaging in Huntington's disease reveals distinct patterns of white matter degeneration associated with motor and cognitive deficits. Brain Imaging Behav. 2011;5:171–180. doi: 10.1007/s11682-011-9121-8. [DOI] [PubMed] [Google Scholar]
- Boulanger JJ, Messier C. From precursors to myelinating oligodendrocytes: Contribution of intrinsic and extrinsic factors to white matter plasticity in the adult brain. Neuroscience. 2014;269C:343–366. doi: 10.1016/j.neuroscience.2014.03.063. [DOI] [PubMed] [Google Scholar]
- Bradford J, Shin JY, Roberts M, Wang CE, Li XJ, Li S. Expression of mutant huntingtin in mouse brain astrocytes causes age-dependent neurological symptoms. Proc Natl Acad Sci U S A. 2009;106:22480–22485. doi: 10.1073/pnas.0911503106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bujalka H, Koenning M, Jackson S, Perreau VM, Pope B, Hay CM, Mitew S, Hill AF, Lu QR, Wegner M, Srinivasan R, Svaren J, Willingham M, Barres BA, Emery B. MYRF is a membrane-associated transcription factor that autoproteolytically cleaves to directly activate myelin genes. PLoS Biol. 2013;11:e1001625. doi: 10.1371/journal.pbio.1001625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cahoy JD, Emery B, Kaushal A, Foo LC, Zamanian JL, Christopherson KS, Xing Y, Lubischer JL, Krieg PA, Krupenko SA, Thompson WJ, Barres BA. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. The Journal of neuroscience : the official journal of the Society for Neuroscience. 2008;28:264–278. doi: 10.1523/JNEUROSCI.4178-07.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camacho A, Huang JK, Delint-Ramirez I, Yew Tan C, Fuller M, Lelliott CJ, Vidal-Puig A, Franklin RJ. Peroxisome proliferator-activated receptor gamma-coactivator-1 alpha coordinates sphingolipid metabolism, lipid raft composition and myelin protein synthesis. Eur J Neurosci. 2013;38:2672–2683. doi: 10.1111/ejn.12281. [DOI] [PubMed] [Google Scholar]
- Crotti A, Benner C, Kerman BE, Gosselin D, Lagier-Tourenne C, Zuccato C, Cattaneo E, Gage FH, Cleveland DW, Glass CK. Mutant Huntingtin promotes autonomous microglia activation via myeloid lineage-determining factors. Nat Neurosci. 2014;17:513–521. doi: 10.1038/nn.3668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Paola M, Luders E, Cherubini A, Sanchez-Castaneda C, Thompson PM, Toga AW, Caltagirone C, Orobello S, Elifani F, Squitieri F, Sabatini U. Multimodal MRI analysis of the corpus callosum reveals white matter differences in presymptomatic and early Huntington's disease. Cereb Cortex. 2012;22:2858–2866. doi: 10.1093/cercor/bhr360. [DOI] [PubMed] [Google Scholar]
- Di Paola M, Phillips OR, Sanchez-Castaneda C, Di Pardo A, Maglione V, Caltagirone C, Sabatini U, Squitieri F. MRI measures of corpus callosum iron and myelin in early Huntington's disease. Hum Brain Mapp. 2014;35:3143–3151. doi: 10.1002/hbm.22391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ehrnhoefer DE, Sutton L, Hayden MR. Small changes, big impact: posttranslational modifications and function of huntingtin in Huntington disease. Neuroscientist. 2011;17:475–492. doi: 10.1177/1073858410390378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emery B. Regulation of oligodendrocyte differentiation and myelination. Science. 2010;330:779–782. doi: 10.1126/science.1190927. [DOI] [PubMed] [Google Scholar]
- Emery B, Agalliu D, Cahoy JD, Watkins TA, Dugas JC, Mulinyawe SB, Ibrahim A, Ligon KL, Rowitch DH, Barres BA. Myelin gene regulatory factor is a critical transcriptional regulator required for CNS myelination. Cell. 2009;138:172–185. doi: 10.1016/j.cell.2009.04.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fennema-Notestine C, Archibald SL, Jacobson MW, Corey-Bloom J, Paulsen JS, Peavy GM, Gamst AC, Hamilton JM, Salmon DP, Jernigan TL. In vivo evidence of cerebellar atrophy and cerebral white matter loss in Huntington disease. Neurology. 2004;63:989–995. doi: 10.1212/01.wnl.0000138434.68093.67. [DOI] [PubMed] [Google Scholar]
- Franklin RJ, Ffrench-Constant C. Remyelination in the CNS: from biology to therapy. Nat Rev Neurosci. 2008;9:839–855. doi: 10.1038/nrn2480. [DOI] [PubMed] [Google Scholar]
- Funfschilling U, Supplie LM, Mahad D, Boretius S, Saab AS, Edgar J, Brinkmann BG, Kassmann CM, Tzvetanova ID, Mobius W, Diaz F, Meijer D, Suter U, Hamprecht B, Sereda MW, Moraes CT, Frahm J, Goebbels S, Nave KA. Glycolytic oligodendrocytes maintain myelin and long-term axonal integrity. Nature. 2012;485:517–521. doi: 10.1038/nature11007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuss B, Mallon B, Phan T, Ohlemeyer C, Kirchhoff F, Nishiyama A, Macklin WB. Purification and analysis of in vivo-differentiated oligodendrocytes expressing the green fluorescent protein. Dev Biol. 2000;218:259–274. doi: 10.1006/dbio.1999.9574. [DOI] [PubMed] [Google Scholar]
- Gu X, Li C, Wei W, Lo V, Gong S, Li SH, Iwasato T, Itohara S, Li XJ, Mody I, Heintz N, Yang XW. Pathological cell-cell interactions elicited by a neuropathogenic form of mutant Huntingtin contribute to cortical pathogenesis in HD mice. Neuron. 2005;46:433–444. doi: 10.1016/j.neuron.2005.03.025. [DOI] [PubMed] [Google Scholar]
- Han I, You Y, Kordower JH, Brady ST, Morfini GA. Differential vulnerability of neurons in Huntington's disease: the role of cell type-specific features. J Neurochem. 2010;113:1073–1091. doi: 10.1111/j.1471-4159.2010.06672.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harjes P, Wanker EE. The hunt for huntingtin function: interaction partners tell many different stories. Trends Biochem Sci. 2003;28:425–433. doi: 10.1016/S0968-0004(03)00168-3. [DOI] [PubMed] [Google Scholar]
- Havel LS, Wang CE, Wade B, Huang B, Li S, Li XJ. Preferential accumulation of N-terminal mutant huntingtin in the nuclei of striatal neurons is regulated by phosphorylation. Hum Mol Genet. 2011;20:1424–1437. doi: 10.1093/hmg/ddr023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heng MY, Detloff PJ, Albin RL. Rodent genetic models of Huntington disease. Neurobiol Dis. 2008;32:1–9. doi: 10.1016/j.nbd.2008.06.005. [DOI] [PubMed] [Google Scholar]
- Hornig J, Frob F, Vogl MR, Hermans-Borgmeyer I, Tamm ER, Wegner M. The transcription factors Sox10 and Myrf define an essential regulatory network module in differentiating oligodendrocytes. PLoS Genet. 2013;9:e1003907. doi: 10.1371/journal.pgen.1003907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kang SH, Li Y, Fukaya M, Lorenzini I, Cleveland DW, Ostrow LW, Rothstein JD, Bergles DE. Degeneration and impaired regeneration of gray matter oligodendrocytes in amyotrophic lateral sclerosis. Nat Neurosci. 2013;16:571–579. doi: 10.1038/nn.3357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koenning M, Jackson S, Hay CM, Faux C, Kilpatrick TJ, Willingham M, Emery B. Myelin gene regulatory factor is required for maintenance of myelin and mature oligodendrocyte identity in the adult CNS. J Neurosci. 2012;32:12528–12542. doi: 10.1523/JNEUROSCI.1069-12.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee CY, Cantle JP, Yang XW. Genetic manipulations of mutant huntingtin in mice: new insights into Huntington's disease pathogenesis. FEBS J. 2013;280:4382–4394. doi: 10.1111/febs.12418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee J, Taghian K, Petratos S. Axonal degeneration in multiple sclerosis: can we predict and prevent permanent disability? Acta Neuropathol Commun. 2014;2:97. doi: 10.1186/s40478-014-0097-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee Y, Morrison BM, Li Y, Lengacher S, Farah MH, Hoffman PN, Liu Y, Tsingalia A, Jin L, Zhang PW, Pellerin L, Magistretti PJ, Rothstein JD. Oligodendroglia metabolically support axons and contribute to neurodegeneration. Nature. 2012;487:443–448. doi: 10.1038/nature11314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Li SH, Yu ZX, Shelbourne P, Li XJ. Huntingtin aggregate-associated axonal degeneration is an early pathological event in Huntington's disease mice. The Journal of neuroscience : the official journal of the Society for Neuroscience. 2001;21:8473–8481. doi: 10.1523/JNEUROSCI.21-21-08473.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li SH, Cheng AL, Zhou H, Lam S, Rao M, Li H, Li XJ. Interaction of Huntington disease protein with transcriptional activator Sp1. Mol Cell Biol. 2002;22:1277–1287. doi: 10.1128/mcb.22.5.1277-1287.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li SH, Li XJ. Huntingtin-protein interactions and the pathogenesis of Huntington's disease. Trends Genet. 2004;20:146–154. doi: 10.1016/j.tig.2004.01.008. [DOI] [PubMed] [Google Scholar]
- Li X-J, Li SH. Influence of species differences on the neuropathology of transgenic Huntington's disease animal models. J Genet Genomics. 2012;39:239–45. doi: 10.1016/j.jgg.2012.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Z, Park Y, Marcotte EM. A Bacteriophage tailspike domain promotes self-cleavage of a human membrane-bound transcription factor, the myelin regulatory factor MYRF. PLoS Biol. 2013;11:e1001624. doi: 10.1371/journal.pbio.1001624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ligon KL, Fancy SP, Franklin RJ, Rowitch DH. Olig gene function in CNS development and disease. Glia. 2006;54:1–10. doi: 10.1002/glia.20273. [DOI] [PubMed] [Google Scholar]
- Longair MH, Baker DA, Armstrong JD. Simple Neurite Tracer: open source software for reconstruction, visualization and analysis of neuronal processes. Bioinformatics. 2011;27:2453–2454. doi: 10.1093/bioinformatics/btr390. [DOI] [PubMed] [Google Scholar]
- Mack JT, Beljanski V, Soulika AM, Townsend DM, Brown CB, Davis W, Tew KD. "Skittish" Abca2 knockout mice display tremor, hyperactivity, and abnormal myelin ultrastructure in the central nervous system. Mol Cell Biol. 2007;27:44–53. doi: 10.1128/MCB.01824-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makinson CD, Tanaka BS, Lamar T, Goldin AL, Escayg A. Role of the hippocampus in Na1.6 (Scn8a) mediated seizure resistance. Neurobiol Dis. 2014 doi: 10.1016/j.nbd.2014.03.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mallon BS, Shick HE, Kidd GJ, Macklin WB. Proteolipid promoter activity distinguishes two populations of NG2-positive cells throughout neonatal cortical development. The Journal of neuroscience : the official journal of the Society for Neuroscience. 2002;22:876–885. doi: 10.1523/JNEUROSCI.22-03-00876.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKenzie IA, Ohayon D, Li H, de Faria JP, Emery B, Tohyama K, Richardson WD. Motor skill learning requires active central myelination. Science. 2014;346:318–322. doi: 10.1126/science.1254960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meijer DH, Kane MF, Mehta S, Liu H, Harrington E, Taylor CM, Stiles CD, Rowitch DH. Separated at birth? The functional and molecular divergence of OLIG1 and OLIG2. Nat Rev Neurosci. 2012;13:819–831. doi: 10.1038/nrn3386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Menalled LB, Chesselet MF. Mouse models of Huntington's disease. Trends Pharmacol Sci. 2002;23:32–39. doi: 10.1016/s0165-6147(00)01884-8. [DOI] [PubMed] [Google Scholar]
- Michalski JP, Anderson C, Beauvais A, De Repentigny Y, Kothary R. The proteolipid protein promoter drives expression outside of the oligodendrocyte lineage during embryonic and early postnatal development. PLoS One. 2011;6:e19772. doi: 10.1371/journal.pone.0019772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Munoz-Sanjuan I, Bates GP. The importance of integrating basic and clinical research toward the development of new therapies for Huntington disease. J Clin Invest. 2011;121:476–483. doi: 10.1172/JCI45364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paus T, Zijdenbos A, Worsley K, Collins DL, Blumenthal J, Giedd JN, Rapoport JL, Evans AC. Structural maturation of neural pathways in children and adolescents: in vivo study. Science. 1999;283:1908–1911. doi: 10.1126/science.283.5409.1908. [DOI] [PubMed] [Google Scholar]
- Phillips O, Squitieri F, Sanchez-Castaneda C, Elifani F, Caltagirone C, Sabatini U, Di Paola M. Deep White Matter in Huntington's Disease. PLoS One. 2014;9:e109676. doi: 10.1371/journal.pone.0109676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Philips T, Bento-Abreu A, Nonneman A, Haeck W, Staats K, Geelen V, Hersmus N, Kusters B, Van Den Bosch L, Van Damme P, Richardson WD, Robberecht W. Oligodendrocyte dysfunction in the pathogenesis of amyotrophic lateral sclerosis. Brain. 2013;136:471–482. doi: 10.1093/brain/aws339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rivers LE, Young KM, Rizzi M, Jamen F, Psachoulia K, Wade A, Kessaris N, Richardson WD. PDGFRA/NG2 glia generate myelinating oligodendrocytes and piriform projection neurons in adult mice. Nat Neurosci. 2008;11:1392–1401. doi: 10.1038/nn.2220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodriguez M, Scheithauer B. Ultrastructure of multiple sclerosis. Ultrastruct Pathol. 1994;18:3–13. doi: 10.3109/01913129409016267. [DOI] [PubMed] [Google Scholar]
- Rosas HD, Koroshetz WJ, Chen YI, Skeuse C, Vangel M, Cudkowicz ME, Caplan K, Marek K, Seidman LJ, Makris N, Jenkins BG, Goldstein JM. Evidence for more widespread cerebral pathology in early HD: an MRI-based morphometric analysis. Neurology. 2003;60:1615–1620. doi: 10.1212/01.wnl.0000065888.88988.6e. [DOI] [PubMed] [Google Scholar]
- Ross CA, Aylward EH, Wild EJ, Langbehn DR, Long JD, Warner JH, Scahill RI, Leavitt BR, Stout JC, Paulsen JS, Reilmann R, Unschuld PG, Wexler A, Margolis RL, Tabrizi SJ. Huntington disease: natural history, biomarkers and prospects for therapeutics. Nat Rev Neurol. 2014;10:204–216. doi: 10.1038/nrneurol.2014.24. [DOI] [PubMed] [Google Scholar]
- Ross CA, Tabrizi SJ. Huntington's disease: from molecular pathogenesis to clinical treatment. Lancet Neurol. 2011;10:83–98. doi: 10.1016/S1474-4422(10)70245-3. [DOI] [PubMed] [Google Scholar]
- Saher G, Quintes S, Nave KA. Cholesterol: a novel regulatory role in myelin formation. Neuroscientist. 2011;17:79–93. doi: 10.1177/1073858410373835. [DOI] [PubMed] [Google Scholar]
- Sathornsumetee S, McGavern DB, Ure DR, Rodriguez M. Quantitative ultrastructural analysis of a single spinal cord demyelinated lesion predicts total lesion load, axonal loss, and neurological dysfunction in a murine model of multiple sclerosis. Am J Pathol. 2000;157:1365–1376. doi: 10.1016/S0002-9440(10)64650-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shin JY, Fang ZH, Yu ZX, Wang CE, Li SH, Li XJ. Expression of mutant huntingtin in glial cells contributes to neuronal excitotoxicity. J Cell Biol. 2005;171:1001–1012. doi: 10.1083/jcb.200508072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tong X, Ao Y, Faas GC, Nwaobi SE, Xu J, Haustein MD, Anderson MA, Mody I, Olsen ML, Sofroniew MV, Khakh BS. Astrocyte Kir4.1 ion channel deficits contribute to neuronal dysfunction in Huntington's disease model mice. Nat Neurosci. 2014;17:694–703. doi: 10.1038/nn.3691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Valenza M, Cattaneo E. Emerging roles for cholesterol in Huntington's disease. Trends Neurosci. 2011;34:474–486. doi: 10.1016/j.tins.2011.06.005. [DOI] [PubMed] [Google Scholar]
- Vonsattel JP, DiFiglia M. Huntington disease. J Neuropathol Exp Neurol. 1998;57:369–384. doi: 10.1097/00005072-199805000-00001. [DOI] [PubMed] [Google Scholar]
- Wade A, Jacobs P, Morton AJ. Atrophy and degeneration in sciatic nerve of presymptomatic mice carrying the Huntington's disease mutation. Brain Res. 2008;1188:61–68. doi: 10.1016/j.brainres.2007.06.059. [DOI] [PubMed] [Google Scholar]
- Wang CE, Tydlacka S, Orr AL, Yang SH, Graham RK, Hayden MR, Li S, Chan AW, Li XJ. Accumulation of N-terminal mutant huntingtin in mouse and monkey models implicated as a pathogenic mechanism in Huntington's disease. Hum Mol Genet. 2008;17:2738–2751. doi: 10.1093/hmg/ddn175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woodman B, Butler R, Landles C, Lupton MK, Tse J, Hockly E, Moffitt H, Sathasivam K, Bates GP. The Hdh(Q150/Q150) knock-in mouse model of HD and the R6/2 exon 1 model develop comparable and widespread molecular phenotypes. Brain Res Bull. 2007;72:83–97. doi: 10.1016/j.brainresbull.2006.11.004. [DOI] [PubMed] [Google Scholar]
- Xiang Z, Valenza M, Cui L, Leoni V, Jeong HK, Brilli E, Zhang J, Peng Q, Duan W, Reeves SA, Cattaneo E, Krainc D. Peroxisome-proliferator-activated receptor gamma coactivator 1 alpha contributes to dysmyelination in experimental models of Huntington's disease. The Journal of neuroscience : the official journal of the Society for Neuroscience. 2011;31:9544–9553. doi: 10.1523/JNEUROSCI.1291-11.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu Q, Huang S, Song M, Wang CE, Yan S, Liu X, Gaertig MA, Yu SP, Li H, Li S, Li XJ. Synaptic mutant huntingtin inhibits synapsin-1 phosphorylation and causes neurological symptoms. J Cell Biol. 2013;202:1123–1138. doi: 10.1083/jcb.201303146. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.