Abstract
Purpose
In proton MR spectra of the human brain, relatively broad macromolecule (MM) resonances underlie the narrower signals from metabolites. The purpose of this study was to quantify the MM profile in healthy human brain at 3T and 7T, both in gray matter (anterior cingulate cortex (ACC)) and white matter (centrum semiovale (CSO)).
Methods
A water-suppressed, inversion-recovery pulse sequence was used to null metabolite signals and acquire MM spectra in 20 healthy volunteers using very similar methodology at both field strengths (n=5 per region and field). The MM spectra were fitted with multiple Gaussian functions, and quantified relative to the unsuppressed water signal from the same volume.
Results
MM proton concentration values were in the range of 5 to 20 mmol/kg. No significant differences were found between the MM proton concentration measurements by region (p ≈ 0.8), nor by field strength (p ≈ 0.5). Linewidths of the well-resolved M1 peak were slightly more than double at 7T (43.0 ± 4.7 Hz in the ACC, 45.6 ± 4.1 Hz in the CSO) compared to 3T (19.8 ± 3.5 Hz in the ACC, 20.0 ± 4.3 Hz in the CSO).
Conclusion
The absence of differences in MM concentrations between white and gray matter implies that a single MM ‘baseline’ may be adequate for spectral fitting of multiple brain regions when determining metabolite concentrations. Visibility of MM signals is similar at 3 and 7T.
Keywords: In vivo 1H MRS, brain, macromolecules, macromolecular baseline, 7 Tesla
INTRODUCTION
In short-echo-time (TE) proton MR spectra of the human brain, a number of broad signals are observed which are believed to originate from ‘macromolecular’ (MM) compounds with relatively short T2 relaxation times (1). The biochemical origin of these signals is not completely certain, but it appears that they originate from cytosolic proteins, and perhaps also lipids (1). The different MM peaks (usually labeled M1 through M7) have been attributed to the various resonances of the protein amino acids, including leucine, isoleucine and valine (M1), threonine, and alanine (M2–3), lysine and arginine (M4 and M7), and glutamate and glutamine (M5–6). Resonances between 3.5 and 4.5 ppm have been assigned to α-methine protons also (1). These MM resonances overlap with the metabolite resonances; the accuracy of quantification of the smaller-molecular-weight metabolite signals is therefore quite strongly dependent on the estimation of the contribution of MM resonances to the spectrum. MM resonances may also be of interest in their own right, and have been reported to be age-dependent and to vary between brain regions in healthy subjects (2). For instance, higher MM concentrations have been reported (3) in the cerebellum and the motor cortex than in the pons and white matter (WM), which was attributed to be due to a higher relative proportion of gray matter (GM) in those regions. Another study at 3T suggested that the MM signal might vary among healthy subjects, which would then suggest the need for measurement of the MM signals in each subject in order to accurately quantify the metabolite concentrations (4). However, in a recent study performed in the occipital lobe at 7T, differences in MM signal between regions of WM and GM were reported to be marginal, suggesting that a common MM profile might be used for analysis of both tissue types (5).
MM resonances may also be affected under pathological conditions; for example, in acute multiple sclerosis (MS), elevated MM concentrations were reported (6) and interpreted as resulting from myelin protein breakdown, providing discrimination between acute and chronic MS lesions, as well as healthy controls. Degradation and liberation of the myelin membranes such as sphingomyelin might also be a potential cause for elevated MM concentrations in MS (7). Increased MM levels were also found in studies of stroke, and it has been suggested that they may result from the higher visibility of cytosolic proteins after cell death (8, 9). In brain tumors, MM levels were found to be higher than in normal brain and to increase with tumor grade, which was hypothesized to be consistent with progression from hypoxia to necrosis, as the tumor outgrows its blood supply (10, 11). Knowledge about the MM signal is therefore of interest both for an accurate estimation of metabolite concentrations, as well as providing information about tissue pathophysiology in various brain diseases.
Several strategies to assess MM signals have been explored. Direct spectral fitting of conventional short TE spectra is possible, but places high demands on the robustness of the fitting algorithm because of overlap with metabolite signals, and generally requires prior information for an accurate estimation of the MM concentrations. Alternatively, experimental methods based on the differences between MM and metabolite signal properties allow the separate MM measurement in vivo. In particular, MM resonances have significantly shorter T1 longitudinal relaxation times (~ 250 ms at 2.1T) compared to the mobile metabolite resonances (~ 1500 ms), due to their larger molecular weight (1, 12). MM spectra can therefore be acquired by inversion-nulling the metabolite signals (13).
To date, most studies of MM in the human brain have been performed at field strengths of 1.5 or 3T, although there has been one recent report of measurements at 7T (5), and to our knowledge, there have been no quantitative comparisons of MM signals between field strengths, although one non-quantitative study exists (39). The purpose of this study was therefore to use the inversion-recovery technique in the healthy human brain to compare MM spectra between 3T and 7T in the centrum semiovale (CSO), a region composed predominantly of WM, and in the anterior cingulate cortex (ACC), a region of mostly GM. Acquisition techniques were closely matched between field strengths, with the exception of differences in repetition (TR) and inversion (TI) times necessary to null the metabolite signals due to their different relaxation times at the two field strengths.
METHODS
Human subjects
20 normal volunteers (5 for each cerebral region and at each field strength, aged 33 ± 4 years, 9 male) participated in the study and gave informed consent under local Institutional Review Board approval. Because of lengthy scanning sessions, different volunteers were scanned at each field strength.
Data acquisition
All spectra were recorded on Philips ‘Achieva’ scanners using 32-channel receive head coils, operating at 3T and 7T. A water-suppressed, inversion-recovery scheme based on the semi-LASER pulse sequence (14, 15) was designed to acquire MM spectra free from longer T1 metabolite signals. The inversion-recovery module consisted of a 180° hyperbolic-secant RF pulse applied at a time TI before the excitation pulse of the semi-LASER localization sequence. Water suppression pulses were applied during TI. To avoid possible effects of the inversion pulse on the water suppression efficacy, the inversion pulse had a bandwidth of 4.69 ppm (600 Hz at 3T, and 1400 Hz at 7T) and was applied at a center frequency of 1.9 ppm, so the inversion range was approximately from 4.25 to −0.45 ppm. The semi-LASER sequence was chosen since it has good sensitivity, slice profiles and (when used with high bandwidth adiabatic refocusing pulses) relatively small chemical shift displacement artifacts even at high fields such as 7T (14, 15); however, it does have a longer minimum TE than other localization sequences such as STEAM or PRESS.
Water suppression was performed using the WET sequence (16), consisting of 4 chemical shift selective suppression (CHESS) pulses (17) applied after the inversion pulse but prior to the semi-LASER localization sequence. The WET sequence gives good suppression factors over a range of transmit B1 and water T1 values, and also has relatively short duration (329 ms in the current implementation), which defines the lower limit on TI that can be used. Suppression of subcutaneous lipid signals was performed with spatial outer-volume suppression (OVS) pulses (18) applied at the beginning of the sequence.
To determine the inversion time TInull with the smallest residual metabolite signals, an initial fast inversion-recovery (FIR) (19) experiment was first performed with a set of different inversion times; at 3T, TI ranged from 400 to 1000 ms with steps of 50 ms, while at 7T, TI ranged from 700 to 1000 ms with steps of 50 ms. The TR was 1559 ms at 3T and 3000 ms at 7T, while TE was 31 ms at both field strengths. The longer TR at 7T was necessary in order to keep the sequence specific absorption rate (SAR) under the FDA-approved limit for head imaging.
After determining optimal TInull values at both field strengths, spectra were recorded from the ACC and CSO in 5 subjects at each field strength. Three-dimensional anatomical T1-weighted images (MPRAGE, 1 mm isotropic resolution) were acquired prior to MRS in order to guide the voxel placements, and were also used to determine MRS voxel composition (% WM, GM and cerebrospinal fluid (CSF)). All first- and second-order shim currents were adjusted to optimize B0 homogeneity using the FASTMAP algorithm (20). The voxel volumes in the CSO and in the ACC were 3.0×2.5×2.5 cm3 and 2.5×2.5×2.5 cm3 respectively. The numbers of averages were 512 at 3 T and 256 at 7 T (because of the longer TR at 7T), corresponding to a scan time of approximately 13 minutes per voxel at both field strengths. The unsuppressed, fully-relaxed water signal was also recorded (TR was 3000 ms at 3T and 4000 ms at 7T) from the same regions of interest using 4 averages, as well as conventional water-suppressed sLASER spectra without inversion pulses. The TE was 31 ms at both field strengths whereas TR was 1559 ms at 3T and 3000 ms at 7T. Slice-selective excitation in sLASER was achieved by a frequency modulated 90° pulse (‘fremex05’) with a duration of 9.7 and 8.7 ms and a bandwidth of 4.2 and 4.7 kHz, respectively at 3T and 7T (21). Refocusing was achieved by two pairs of slice selective B1-insensitive adiabatic full passage (AFP) 180° pulses with durations of 4.3 and 5.2 ms, and bandwidths of 2 and 3 kHz, respectively at 3T and 7T (22).
Data analysis
All the spectra were pre-processed using in-house software (‘csx3’) and line broadened by 2 Hz. The MM spectra were baseline corrected using a cubic spline function. Further analysis was performed in MATLAB (version 8.1.0. Natick, Massachusetts: The MathWorks Inc., 2013); the MM peaks were fitted with a sum of 19 Gaussian functions between 0.50 ppm and 3.50 ppm and individual components integrated. Results are presented for groups of MM peaks assigned ‘M1’ through ‘M7’ according to the definition of Behar et al. (1). The unsuppressed water signal was integrated and the ratios between the MM integrals and the water integrals were calculated. The MM/water ratios were not corrected for either T1 or T2 relaxation time effects. The water concentrations in the ACC and in CSO voxels were computed after the voxel compositions in WM, GM and CSF were determined, taking into account the partial volume effects due to the CSF. The voxel water content was determined by segmenting the anatomical MPRAGE images into % WM, GM and CSF using the ‘SPM’ program, and assuming WM, GM and CSF water contents of 64.6, 78.0 and 100%, respectively (23). In this way, the unsuppressed water signal was used as an internal intensity standard for calibration of the MM concentrations (expressed as ‘proton concentration’, i.e. mmol proton/kg tissue), using a brain density of 1.05 g/cm3 (24).
A univariate ANOVA analysis was performed with field strength and voxel location as fixed factors, and MM proton concentration as dependent factor. Differences between regions and field strengths were subsequently tested for each MM peak; post-hoc Bonferroni-corrected parametric comparisons were performed.
RESULTS
The optimum inversion times TInull were found to be 600 ± 30 ms at 3T (TR 1559 ms), and 900 ± 50 ms at 7 T (TR 3000 ms), respectively. Examples of spectra with and without inversion pulses (applied at TInull) are shown in Figure 1 for the ACC and CSO for both 3 and 7T. The resonances of the main metabolites including total creatine (tCr at 3.90 ppm and 3.01 ppm), total choline (tCho at 3.19 ppm) and total N-acetylaspartate (tNAA at 2.01 ppm) are marked. The FIR TInull values determined experimentally can be used to estimate the metabolite T1 values at each field strength using the equation (25):
TInull = T1 ln [2 – exp ( – TR/T1)].
At 3 T the TInull of 600 ms with a TR value of 1559 ms would correspond to a T1 value of around 1100 ms, while at 7 T the TInull of 935 ms for a TR value of 3000 ms corresponds to an approximate metabolite T1 value of 1350 ms, assuming that inversion was perfect in both experiments. Some residual ‘out-of-voxel’ signal originating from unwanted coherence transfer pathways can be seen, e.g. in figure 1B, both around the water signal and between 3.3 to 3.9 ppm. This signal arises from insufficient gradient crusher strength and/or duration, and may be minimized by choosing the correct order of slice selection gradients within the pulse sequence (38). Note also that the 3.9 ppm Cr peak is observed in the metabolite-suppressed spectra, since it falls near the transition band of the frequency selective inversion pulse (nominal cutoff frequency 4.25 ppm) and is therefore not completely inverted. There may be some very small residual tNAA signals in some of the spectra also, due to imperfect nulling, which may cause some error in the M5 peak area measurement.
Figure 1.
Conventional water-suppressed in vivo spectra at (A) 3T and (B) 7T acquired with the semi-LASER pulse sequence from a 25×25×25 mm3 voxel located in the anterior cingulate cortex (ACC - inset). The major metabolite resonances include: tCr, total creatine (3.90 ppm and 3.01 ppm); tCho, total choline (3.19 ppm); tNAA, total N-acetylaspartate (2.01 ppm). The MM peak labeled M1 (0.9 ppm) can be clearly distinguished, whereas the other MM peaks are overlapped by the metabolite resonances. MM-spectra acquired with an inversion-recovery, water-suppressed semi-LASER pulse sequence from the same region as shown in (C) 3T (TR/TI 1559/600 ms) and (D) 7T (TR/TI 3000/900 ms). All spectra were line broadened by 2 Hz.
Conventional water-suppressed in vivo spectra at (E) 3T and (F) 7T acquired with the semi-LASER pulse sequence from a 30×25×25 mm3 voxel located in the centrum semiovale (CSO - inset). MM spectra acquired with an inversion-recovery, water-suppressed semi-LASER pulse sequence from the same region as shown in (G) 3T (TR/TI 1559/600 ms) and (H) 7T (TR/TI 3000/900 ms). All spectra were line broadened by 2 Hz.
Figure 2 shows the Gaussian curve-fits of the metabolite-nulled spectra in the ACC and CSO at 3T and at 7T. The MM peak labeled M1 (0.9 ppm), which presumably contains signals of MM methyl groups (1), was well-resolved in all spectra in both the ACC and in the CSO, at 3 and 7T, allowing the measurement of its line width. The M1 linewidth values were 19.8 ± 3.5 Hz in the ACC and 20.0 ± 4.3 Hz in the CSO at 3T, and 43.0 ± 4.7 Hz and 45.6 ± 4.1 Hz at 7T, respectively.
Figure 2.
Example results of the Gaussian curve-fitting of the MM spectra in the ACC and in the CSO at 3T (respectively A and B) and at 7T (respectively C and D). The MM peak clusters (M1–M7) were fitted using 19 Gaussian lineshapes and integrated. The full red lines represent the fits of the metabolite-nulled spectra.
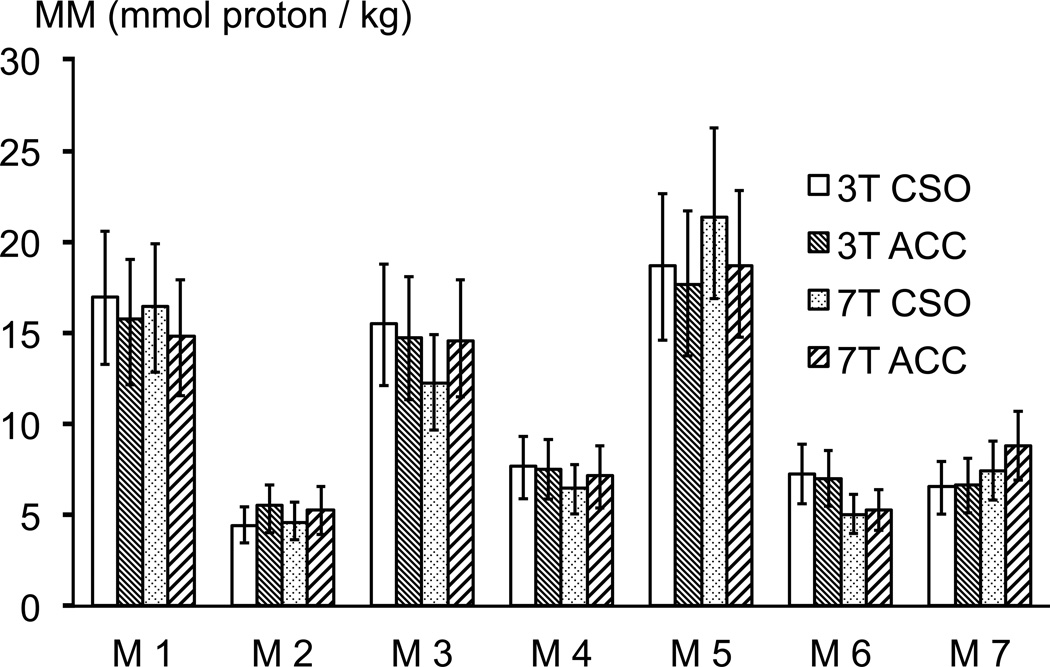
The MM proton concentration values (mmol/kg) are displayed in Figure 3 (mean ± standard deviation) for the M1–7 peaks in the CSO and in ACC at 3T and 7T, after the voxel water content was determined by segmenting the anatomical MPRAGE images into % WM, GM and CSF (3T ACC voxels: 9.4 ± 2.3 % CSF, 60.2 ± 4.5 % GM, 30.4 ± 3.8 % WM; 3T CSO voxels: 2.4 ± 1.1 % CSF, 19.8 ± 2.5 % GM, 77.8 ± 2.4 % WM; 7T ACC voxels: 8.6 ± 2.1 % CSF, 62.2 ± 3.9 % GM, 29.2 ± 3.8 % WM; 7T CSO voxels: 3.8 ± 0.9 % CSF, 22.8 ± 3.6 % GM, 73.4 ± 4.2 % WM). The MM peak M5 corresponds to the highest MM proton concentration, followed by MM peaks M1 and M3. The MM peak M2 corresponds to the lowest MM proton concentration. No significant differences were found between the MM proton concentrations by region (p ≈ 0.8), or by field strength (p ≈ 0.5).
Figure 3.
MM resonance proton concentrations for ACC and CSO at 3T and 7T (mmol/kg wet weight, mean ± st.dev). No corrections for relaxation time effects were applied. No significant differences are found between the MM proton concentrations by region (p ≈ 0.8), nor by field strength (p ≈ 0.5).
Figure 4 shows examples of 7T ACC spectra recorded at a range of inversion times, in order to determine optimum TI for metabolite nulling.
Figure 4.
Example inversion recovery spectra from the anterior cingulate cortex recorded at 7 Tesla in one subject. (A) shows the conventional spectrum without inversion, while (B-E) show spectra as a function of inversion time (TI) ranging from (B) 900 to (E) 700 ms. Major metabolite peaks and MM peaks M1–M7 are indicated.
DISCUSSION
In this study of regional MM concentrations measured at both 3T and 7T in normal subjects, the primary observation was that no GM-WM differences or field-strength-dependent variations were found. M1–M7 proton concentration values were in the range of 5 to 20 mmol/kg, in general agreement with previous studies (2, 26). While several previous studies have suggested regional differences in MM concentrations (2, 4, 6), the current results are consistent with a more recent 7T study (5) which found only relatively minor differences between gray and white matter in the occipital lobe. The absence of differences in MM concentrations between the WM and GM regions studied implies that a general MM ‘baseline’ may be adequate for spectral fitting of multiple brain regions when determining metabolite concentrations, at least in subjects with normal brain. The lack of concentration differences between 3T and 7T (without any relaxation time corrections) also implies that the MM peaks have similar MR visibility (i.e. estimated concentrations) at both field strengths.
A number of technical issues need to be considered when performing MM spectroscopy in vivo. First of all, the ability to measure the MM peaks depends on the ability to accurately suppress the overlying metabolite signals, which depends on the correct choice of inversion time. In this study, TI was optimized experimentally, however, it cannot be completely ruled out that there are some minor contributions from residual metabolite peaks, such as tCho at 3.19 ppm, tNAA at 2.01 and 2.60 ppm, and myo-inositol at 3.60 ppm (27). In order to not interfere with the performance of the water suppression sequence, the inversion pulse was deliberately applied from approximately 4.25 to −0.45 ppm, however at the edges of its bandwidth, the inversion may not be perfect, particularly at 7T, where a residual peak from the CH2 of Cr was frequently observed at 3.9 ppm; this peak has also been seen in other MM studies where its presence was attributed to its shorter T1 relaxation time compared to the other metabolites (28–30). Some studies at 3T undertaken in the occipital lobe found either no significant difference in T1 relaxation time constants between WM and GM (31) or small differences for tCr (3.03 ppm) and tCho (3.22 ppm) (32). However, an increasing trend in T1 was observed for tNAA (CH3), tCr (CH3), and tCho at 7T as compared to 3T (33), which was consistent with the T1null relaxation times determined in the current study. At 7T, the T1 relaxation time was reported to be shorter in WM than in GM for tCho (−14%) whereas it was longer for tCr (3.03 ppm) (+2%) and tNAA (+4%) (33). The minimum TI which can be attained is governed by the length of the water suppression sequence used (if inversion is performed before rather than during the water suppression sequence); while sequences such as VAPOR (34) give excellent water suppression factors, the length of the VAPOR sequence was generally too long for the current application, so the shorter, alternative WET sequence (with 4 water suppression pulses) was used instead. VAPOR may still be used for short inversion times if the inversion pulse is interleaved within the VAPOR sequence, however, and water suppression performance will not be affected so long as the inversion pulse does not invert the water signal. Since it is also important to avoid lipid contamination of the MM spectrum, outer-volume suppression pulses were used to saturate any residual lipid signals originating from the scalp.
The M1 peak linewidth (as expressed in Hz) was approximately double at 7T compared to 3T. This is in contrast to the modeled MM spectra reported in reference (39), which show improving resolution of MM spectra (i.e. decreasing linewidths as measured in ppm) from 1.5 to 7.0T. However, the primary data in reference (39) was not shown, and different pulse sequences were used at different field strengths. Although only the well-resolved M1 peak linewidth was quantified in this study, the similar appearance of the MM spectra and 3 and 7T would suggest a similar increase for the other MM peaks also. It is well known that the smaller molecular weight metabolite signals show an approximately linear increase in linewidth in vivo with increasing B0 field strength (35), which is usually attributed to the increase in the microscopic field inhomogeneity associated with magnetic susceptibility differences between the various tissue compartments (36). Presumably a similar effect is also occurring for the larger molecular weight macromolecules, however the macromolecule linewidths are too large to be explained by this mechanism alone. For instance, at 7T, the field inhomogeneity component of the creatine linewidth has been estimated to be about 6 Hz (37), whereas in the current study the M1 linewidth at 7T was around 44 Hz. Therefore, there must also be additional mechanisms which shorten the MM T2 relaxation time as the magnetic field strength increases.
Experimental parameters were matched as closely as possible between field strengths, including using 32-channel head coil at both fields, the same localization sequence, and nearly identical TE. However, one difference was that the TR used was longer at 7T than at 3T, because of the higher sequence SAR at 7T; a longer TR was necessary in order for the 7T sequence to fall within the FDA approved SAR for head imaging. Since MM have short T1 relaxation times, we do not believe that the difference significantly influenced the MM concentration measurements; however, it did require that TI be optimized independently at both field strengths in order to null the metabolite signals, aside from any potential field dependent differences in metabolite T1 relaxation times.
Another limitation of the current study was that no attempt was made to measure MM T1 and T2 relaxation times, and the MM concentration values are presented without any relaxation time corrections. Considering that MM T2 relaxation times are short, the concentration values presented here are probably an underestimate of the true concentrations, given that appreciable MM T2 relaxation time will occur during the TE time period in these experiments. Finally, although MM peaks exist in the 3.8–4.3 ppm range of the spectrum (2), it was not possible to quantify them in the current study because of the limited bandwidth (cutoff ~ 4.25 ppm) of the inversion pulse used.
In conclusion, the current experiments found that normal human brain MM profiles and concentrations showed no significant differences, either between gray and white matter, or between 3T and 7T. MM resonance linewidths (measured in Hz) were broader at 7T than 3T, presumably reflecting decreasing MM T2 relaxation times. These results suggest that, in normal brain at least, a common MM profile may be used for quantitative analysis of spectra. However, in many diseases, MM peaks may often be different from normal, so flexibility in the MM profile is needed when fitting spectra from pathological brain tissue.
ACKNOWLEDGMENTS
We would like to thank Dr. Fei Gao for providing the brain segmentation method. This work was funded in part by NIH P41EB015909 and R01MH096263.
REFERENCES
- 1.Behar KL, Rothman DL, Spencer DD, Petroff OA. Analysis of macromolecule resonances in 1H NMR spectra of human brain. Magn Reson Med. 1994;32(3):294–302. doi: 10.1002/mrm.1910320304. [DOI] [PubMed] [Google Scholar]
- 2.Hofmann L, Slotboom J, Boesch C, Kreis R. Characterization of the macromolecule baseline in localized 1H-MR spectra of human brain. Magn Reson Med. 2001;46:855–863. doi: 10.1002/mrm.1269. [DOI] [PubMed] [Google Scholar]
- 3.Mader I, Seeger U, Weissert R, Klose U, Naegele T, Melms A, Grodd W. Proton MR spectroscopy with metabolite-nulling reveals elevated macromolecules in acute multiple sclerosis. Brain. 2001;124(Pt 5):953–961. doi: 10.1093/brain/124.5.953. [DOI] [PubMed] [Google Scholar]
- 4.Chong DG, Kreis R, Bolliger CS, Boesch C, Slotboom J. Two-dimensional linear-combination model fitting of magnetic resonance spectra to define the macromolecule baseline using FiTAID, a Fitting Tool for Arrays of Interrelated Datasets. MAGMA. 2011;24:147–164. doi: 10.1007/s10334-011-0246-y. [DOI] [PubMed] [Google Scholar]
- 5.Schaller B, Lijing X, Gruetter R. Is the macromolecule signal tissue-specific in healthy human brain? A 1H MRS study at 7 Tesla in the occipital lobe. Magn Reson Med. 2013 doi: 10.1002/mrm.24995. [DOI] [PubMed] [Google Scholar]
- 6.Mader I, Seeger U, Karitzky J, Erb M, Schick F, Klose U. Proton magnetic resonance spectroscopy with metabolite-nulling reveals regional differences of macromolecules in normal human brain. J Magn Reson Imaging. 2002;16:538–546. doi: 10.1002/jmri.10190. [DOI] [PubMed] [Google Scholar]
- 7.Davie CA, Barker GJ, Tofts PS, Hawkins CP, Brennan A, Miller DH, McDonald WI. Detection of myelin breakdown products by proton magnetic resonance spectroscopy. The Lancet. 1993;341:630–631. doi: 10.1016/0140-6736(93)90390-3. [DOI] [PubMed] [Google Scholar]
- 8.Graham GD, Hwang J-H, Rothman DL, Prichard JW. Spectroscopic assessment of alterations in macromolecule and small-molecule metabolites in human brain after stroke. Stroke. 2001;32:2797–2802. doi: 10.1161/hs1201.099414. [DOI] [PubMed] [Google Scholar]
- 9.Saunders DE, Howe FA, van den Boogaart A, Griffiths JR, Brown MM. Discrimination of metabolite from lipid and macromolecule resonances in cerebral infarction in humans using short echo proton spectroscopy. J Magn Reson Imaging. 1997;7:1116–1121. doi: 10.1002/jmri.1880070626. [DOI] [PubMed] [Google Scholar]
- 10.Seeger U, Klose U, Mader I, Grodd W, Naegele T. Parameterized evaluation of macromolecules and lipids in proton MR spectroscopy of brain diseases. Magn Reson Med. 2003;49:19–28. doi: 10.1002/mrm.10332. [DOI] [PubMed] [Google Scholar]
- 11.Howe FA, Barton SJ, Cudlip SA, Stubbs M, Saunders DE, Murphy M, Wilkins P, Opstad KS, Doyle VL, McLean MA, Bell BA, Griffiths JR. Metabolic profiles of human brain tumors using quantitative in vivo 1H magnetic resonance spectroscopy. Magn Reson Med. 2003;49:223–232. doi: 10.1002/mrm.10367. [DOI] [PubMed] [Google Scholar]
- 12.Xin L, Schaller B, Mlynarik V, Lu H, Gruetter R. Proton T1 relaxation times of metabolites in human occipital white and gray matter at 7 T. Magn Reson Med. 2013;69(4):931–936. doi: 10.1002/mrm.24352. [DOI] [PubMed] [Google Scholar]
- 13.Behar KL, Ogino T. Characterization of macromolecule resonances in the 1H NMR spectrum of rat brain. Magn Reson Med. 1993;30(1):38–44. doi: 10.1002/mrm.1910300107. [DOI] [PubMed] [Google Scholar]
- 14.Scheenen TWJ, Klomp DWJ, Wijnen JP, Heerschap A. Short echo time 1H-MRSI of the human brain at 3T with minimal chemical shift displacement errors using adiabatic refocusing pulses. Magn Reson Med. 2008;59(1):1–6. doi: 10.1002/mrm.21302. [DOI] [PubMed] [Google Scholar]
- 15.Boer VO, van Lier ALHMW, Hoogduin JM, Wijnen JP, Luijten PR, Klomp DWJ. 7-T 1H MRS with adiabatic refocusing at short TE using radiofrequency focusing with a dual-channel volume transmit coil. NMR Biomed. 2011;24(9):1038–1046. doi: 10.1002/nbm.1641. [DOI] [PubMed] [Google Scholar]
- 16.Ogg RJ, Kingsley PB, Taylor JS. WET, a T1- and B1-insensitive water-suppression method for in vivo localized 1H NMR spectroscopy. The experimental approach to accurate carbon-13 spin-lattice relaxation measurements. J Magn Reson B. 1994;104:1–10. doi: 10.1006/jmrb.1994.1048. [DOI] [PubMed] [Google Scholar]
- 17.Haase A, Frahm J, Hanicke W, Matthaei D. 1H NMR chemical shift selective (CHESS) imaging. Phys Med Biol. 1985;30(4):341–344. doi: 10.1088/0031-9155/30/4/008. [DOI] [PubMed] [Google Scholar]
- 18.Duyn JH, Gillen J, Sobering G, van Zijl PC, Moonen CT. Multisection proton MR spectroscopic imaging of the brain. Radiology. 1993;188(1):277–282. doi: 10.1148/radiology.188.1.8511313. [DOI] [PubMed] [Google Scholar]
- 19.Levy GC, Peat IR. The experimental approach to accurate carbon-13 spin-lattice relaxation measurements. J Magn Reson. 1975;18:500–521. [Google Scholar]
- 20.Gruetter R. Automatic, localized in vivo adjustment of all first- and second-order shim coils. Magn Reson Med. 1993;29:804–811. doi: 10.1002/mrm.1910290613. [DOI] [PubMed] [Google Scholar]
- 21.Murdoch JB, editor. Still iterating… and iterating… to solve pulse design problems; 10th ISMRM Scientific Meeting; Hawai’i. 2002. p. 923. [Google Scholar]
- 22.Rosenfeld D, Zur Y. A new adiabatic inversion pulse. Magn Reson Med. 1996;36:124–138. doi: 10.1002/mrm.1910360121. [DOI] [PubMed] [Google Scholar]
- 23.Ernst T, Kreis R, Ross BD. Absolute quantitation of water and metabolites in the human brain. I. Compartments and water. J Magn Reson B. 1993;102:1–8. [Google Scholar]
- 24.Hofmann L, Slotboom J, Jung B, Maloca P, Boesch C, Kreis R. Quantitative 1H-magnetic resonance spectroscopy of human brain: influence of composition and parameterization of the basis set in linear combination model-fitting. Magn Reson Med. 2002;48:440–453. doi: 10.1002/mrm.10246. [DOI] [PubMed] [Google Scholar]
- 25.Canet D, Levy GC, Peat IR. Time saving in 13C spin-lattice relaxation measurements by inversion-recovery. J Magn Reson. 1975;18:199–204. [Google Scholar]
- 26.Gottschalk M, Lamalle L, Segebarth C. Short-TE localized 1H MRS of the human brain at 3 T: quantification of the metabolite signals using two approaches to account for macromolecular signal contributions. NMR Biomed. 2008;21(5):507–517. doi: 10.1002/nbm.1219. [DOI] [PubMed] [Google Scholar]
- 27.Mlynarik V, Cudalbu C, Xin L, Gruetter R. 1H NMR spectroscopy of rat brain in vivo at 14.1Tesla: improvements in quantification of the neurochemical profile. J Magn Reson. 2008;194:163–168. doi: 10.1016/j.jmr.2008.06.019. [DOI] [PubMed] [Google Scholar]
- 28.Pfeuffer J, Tkac I, Provencher SW, Gruetter R. Toward an in vivo neurochemical profile: quantification of 18 metabolites in short-echotime 1H NMR spectra of the rat brain. J Magn Reson. 1999;141:104–120. doi: 10.1006/jmre.1999.1895. [DOI] [PubMed] [Google Scholar]
- 29.Cudalbu C, Mlynarik V, Xin L, Gruetter R. Comparison of T1 relaxation times of the neurochemical profile in rat brain at 9.4T and 14.1T. Magn Reson Med. 2009;62:862–867. doi: 10.1002/mrm.22022. [DOI] [PubMed] [Google Scholar]
- 30.de Graaf RA, Brown PB, McIntyre S, Nixon TW, Behar KL, Rothman DL. High magnetic field water and metabolite proton T1 and T2 relaxation in rat brain in vivo. Magn Reson Med. 2006;56:386–394. doi: 10.1002/mrm.20946. [DOI] [PubMed] [Google Scholar]
- 31.Ethofer T, Mader I, Seeger U, Helms G, Erb M, Grodd W, Ludolph A, Klose U. Comparison of longitudinal metabolite relaxation times in different regions of the human brain at 1.5 and 3 Tesla. Magn Reson Med. 2003;50:1296–1301. doi: 10.1002/mrm.10640. [DOI] [PubMed] [Google Scholar]
- 32.Mlynarik V, Gruber S, Moser E. Proton T1 and T2 relaxation times of human brain metabolites at 3 Tesla. NMR Biomed. 2001;14:325–331. doi: 10.1002/nbm.713. [DOI] [PubMed] [Google Scholar]
- 33.Xin L, Schaller B, Mlynarik V, Lu H, Gruetter R. Proton T1 relaxation times of metabolites in human occipital white and gray matter at 7 T. Magn Reson Med. 2013;69:931–936. doi: 10.1002/mrm.24352. [DOI] [PubMed] [Google Scholar]
- 34.Tkac I, Starcuk Z, Choi IY, Gruetter R. In vivo 1H NMR spectroscopy of rat brain at 1 ms echo time. Magn Reson Med. 1999;41:649–656. doi: 10.1002/(sici)1522-2594(199904)41:4<649::aid-mrm2>3.0.co;2-g. [DOI] [PubMed] [Google Scholar]
- 35.Tkac I, Öz G, Adriany G, Ugurbil K, Gruetter R. In vivo 1H NMR spectroscopy of the human brain at high magnetic fields: metabolite quantification at 4T vs. 7T. Magn Reson Med. 2009;62:868–879. doi: 10.1002/mrm.22086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Michaeli S, Garwood M, Zhu X-H, DelaBarre L, Andersen P, Adriany G, Merkle H, Ugurbil K, Chen W. Proton T2 relaxation study of water, N-acetylaspartate, and creatine in human brain using Hahn and Carr-Purcell spin echoes at 4T and 7T. Magn Reson Med. 2002;47:629–633. doi: 10.1002/mrm.10135. [DOI] [PubMed] [Google Scholar]
- 37.Tkac I, Andersen P, Adriany G, Merkle H, Ugurbil K, Gruetter R. In vivo 1H NMR spectroscopy of the human brain at 7T. Magn Reson Med. 2001;46:451–456. doi: 10.1002/mrm.1213. [DOI] [PubMed] [Google Scholar]
- 38.Ernst T, Chang L. Elimination of artifacts in short echo time H MR spectroscopy of the frontal lobe. Magn Reson Med. 1996;36:462–468. doi: 10.1002/mrm.1910360320. [DOI] [PubMed] [Google Scholar]
- 39.Otazo R, Bueller B, Ugurbil K, Wald L, Posse S. Signal-to-Noise Ratio and Spectral Linewidth Improvements Between 1.5 and 7 Tesla in Proton Echo- Planar Spectroscopic Imaging. Magn Reson Med. 2006;56:1200–1210. doi: 10.1002/mrm.21067. [DOI] [PubMed] [Google Scholar]