Abstract
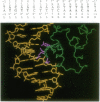
The cI repressor of bacteriophage 434, known as 434 repressor, binds to 14-bp operator sequences by means of a helix-turn-helix motif. To probe the requirements for selective DNA recognition by this class of DNA binding proteins, as well as to generate new proteins with altered specificities, a library of approximately 3 x 10(6) mutants was generated that contains all permutations of five residues in the recognition helix (helix 3) of the repressor. These mutants were then selected in vivo for their ability to bind both wild-type (WT) and mutant operator sequences. The results of the selection demonstrate that four of these residues--Gln28, Gln29, Ser30, and Gln33--play a critical role in recognition of the WT operator. A number of repressors with mutations at Thr27 showed altered DNA binding affinities and specificities. The approach described here may also prove useful in studies of DNA recognition by other classes of DNA binding proteins.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aggarwal A. K., Rodgers D. W., Drottar M., Ptashne M., Harrison S. C. Recognition of a DNA operator by the repressor of phage 434: a view at high resolution. Science. 1988 Nov 11;242(4880):899–907. doi: 10.1126/science.3187531. [DOI] [PubMed] [Google Scholar]
- Burnette W. N. "Western blotting": electrophoretic transfer of proteins from sodium dodecyl sulfate--polyacrylamide gels to unmodified nitrocellulose and radiographic detection with antibody and radioiodinated protein A. Anal Biochem. 1981 Apr;112(2):195–203. doi: 10.1016/0003-2697(81)90281-5. [DOI] [PubMed] [Google Scholar]
- Cwirla S. E., Peters E. A., Barrett R. W., Dower W. J. Peptides on phage: a vast library of peptides for identifying ligands. Proc Natl Acad Sci U S A. 1990 Aug;87(16):6378–6382. doi: 10.1073/pnas.87.16.6378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elledge S. J., Davis R. W. Position and density effects on repression by stationary and mobile DNA-binding proteins. Genes Dev. 1989 Feb;3(2):185–197. doi: 10.1101/gad.3.2.185. [DOI] [PubMed] [Google Scholar]
- Elledge S. J., Sugiono P., Guarente L., Davis R. W. Genetic selection for genes encoding sequence-specific DNA-binding proteins. Proc Natl Acad Sci U S A. 1989 May;86(10):3689–3693. doi: 10.1073/pnas.86.10.3689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorini L. Informational suppression. Annu Rev Genet. 1970;4:107–134. doi: 10.1146/annurev.ge.04.120170.000543. [DOI] [PubMed] [Google Scholar]
- Harrison S. C. A structural taxonomy of DNA-binding domains. Nature. 1991 Oct 24;353(6346):715–719. doi: 10.1038/353715a0. [DOI] [PubMed] [Google Scholar]
- Harrison S. C., Anderson J. E., Koudelka G. B., Mondragon A., Subbiah S., Wharton R. P., Wolberger C., Ptashne M. Recognition of DNA sequences by the repressor of bacteriophage 434. Biophys Chem. 1988 Feb;29(1-2):31–37. doi: 10.1016/0301-4622(88)87022-4. [DOI] [PubMed] [Google Scholar]
- Lim W. A., Sauer R. T. The role of internal packing interactions in determining the structure and stability of a protein. J Mol Biol. 1991 May 20;219(2):359–376. doi: 10.1016/0022-2836(91)90570-v. [DOI] [PubMed] [Google Scholar]
- Pabo C. O., Sauer R. T. Transcription factors: structural families and principles of DNA recognition. Annu Rev Biochem. 1992;61:1053–1095. doi: 10.1146/annurev.bi.61.070192.005201. [DOI] [PubMed] [Google Scholar]
- Wharton R. P., Ptashne M. A new-specificity mutant of 434 repressor that defines an amino acid-base pair contact. 1987 Apr 30-May 6Nature. 326(6116):888–891. doi: 10.1038/326888a0. [DOI] [PubMed] [Google Scholar]
- Wharton R. P., Ptashne M. Changing the binding specificity of a repressor by redesigning an alpha-helix. Nature. 1985 Aug 15;316(6029):601–605. doi: 10.1038/316601a0. [DOI] [PubMed] [Google Scholar]