Abstract
Background
The malaria situation has been worsening in Angola, partly due to armed conflict until the recent past and drug-resistant Plasmodium falciparum. Malaria transmission is heterogeneous within the country, and data on drug-resistant malaria in different parts of the country are incomplete. The aim of the present study was to evaluate resistance to 4-aminoquinolines and antifolate drugs in P. falciparum isolates collected in Benguela province, central Angola, using molecular markers.
Methods
Fingerprick capillary blood was collected from asymptomatic children aged less than 15 years old during a household survey in and around Balombo town in 2010–2011. Samples were screened for P. falciparum by nested PCR. Molecular markers (P. falciparum dihydrofolate reductase [pfdhfr], P. falciparum dihydropteroate synthase [pfdhps], P. falciparum chloroquine resistance transporter [pfcrt], and P. falciparum multidrug-resistance gene 1 [pfmdr1]) were sequenced to determine the key codons associated with drug resistance.
Results
A total of 60 blood samples were positive for P. falciparum. Most isolates with successful PCR amplification had mutant pfdhfr alleles, with either double mutant AICNI (69%) or triple mutant AIRNI (21%) haplotypes. A16V, S108T, and I164L substitutions were not found. Many of the isolates were carriers of either SGKAA (60%) or AGKAA (27%) pfdhps haplotype. K540E substitution was absent. There were only two pfcrt haplotypes: wild-type CVMNK (11%) and mutant CVIET (89%). Wild-type pfmdr1 NYSND haplotype was found in 19% of the isolates, whereas single mutant pfmdr1 YYSND and NFSND haplotypes occurred in 48% and 11%, respectively. Double mutant pfmdr1 haplotypes (YFSND and YYSNY) occurred rarely.
Conclusions
The results suggest that the high prevalence of mutant pfcrt CVIET haplotype is in agreement with low clinical efficacy of chloroquine observed in earlier studies and that the double pfdhfr mutant AICNI and single pfdhps mutant SGKAA are currently the predominant haplotypes associated with antifolate resistance in Benguela province. The hallmark of clinical resistance observed in East Africa, i.e. triple pfdhfr mutant haplotype (AIRNI) and double pfdhps mutant haplotype (SGEAA), was absent. These molecular findings need to be further evaluated in parallel with clinical studies, in particular with the efficacy of intermittent preventive treatment using sulphadoxine-pyrimethamine in pregnant women and artesunate-amodiaquine for uncomplicated malaria.
Keywords: Plasmodium falciparum, Drug resistance, Antifolate drugs, Chloroquine, Molecular markers
Background
Malaria is one of the major causes of morbidity and mortality in Angola. It is responsible for 35% of medical consultations, 20% of hospitalizations, and 25% of maternal deaths [1,2]. In 2010, the Angolan Ministry of Health reported that, of the estimated total population of 18–20 million, 3.7 million cases of malaria occurred, mostly due to Plasmodium falciparum [2].
Malaria is endemic throughout the country, but the transmission pattern is heterogeneous, varying from intense transmission and hyperendemicity in northern Angola to seasonal or epidemic, unstable malaria in the southern part of the country. In central Angola, including Benguela province, malaria transmission is characterized as meso-endemic and stable [2].
One of the causes of the worsening malaria situation in Angola was the civil war that ravaged the country and paralyzed the health system for three decades, until 2002. Another probable cause is drug-resistant malaria. The first chloroquine-resistant cases in the country were reported in non-immune expatriates returning to Europe from Angola in 1984 [3,4]. Low to moderate therapeutic efficacies of chloroquine, sulphadoxine-pyrimethamine, and amodiaquine monotherapies were confirmed in central Angola in 2002 [5]. Artemisinin-based combination therapy (ACT), using either artesunate-amodiaquine, artesunate-sulphadoxine-pyrimethamine, or artemether-lumefantrine, is highly effective [5,6]. Since 2006, the first-line drug for the treatment of uncomplicated malaria in Angola is artemether-lumefantrine, with artesunate-amodiaquine as an alternative drug combination.
In the face of challenges posed by the malaria situation, several clinical and epidemiological studies have been undertaken in recent years, including malaria indicator surveys covering the entire Angolan territory [2,7]. However, many of these recent studies have been conducted in Luanda, the capital city, its neighbouring provinces, or in the northern provinces [8-14]. Data from Benguela province are scarce. The aim of the present study was to analyse the epidemiology of drug-resistant P. falciparum isolates using molecular markers in several localities in and near the town of Balombo.
Methods
Study sites
The study was conducted in 2010–2011 in the town of Balombo and its surrounding villages, situated in Benguela province in central Angola (Figure 1). Balombo (12°21’26” south, 14°46’16” east) is located about 150 km east of the coastal town of Lobito (12°19’11” south, 13°35’59” east) and 420 km to the south of Luanda (8°50’17” south, 13°14’3” east), the capital city of Angola. The estimated total population of Balombo and its surrounding villages was 48,465 inhabitants. The town is located in a valley surrounded by hills.
Figure 1.
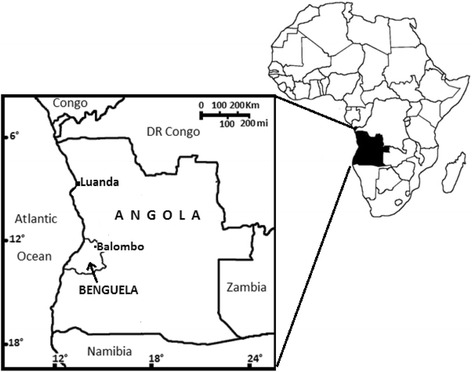
Map of Angola showing Benguela province and Balombo.
Benguela region has an equatorial climate characterized by a short dry season (June–September) and a long rainy season (October–May), with an average annual rainfall of 600–700 mm. Its geography is composed of savannah forests, with areas along the rivers covered by tropical forest. The major malaria vector in Benguela province is Anopheles gambiae s.l. [15,16].
Medical records in Balombo hospital indicate that presumptive malaria accounted for 46% of consultations and 62% of hospitalizations in 2005. However, since most malaria cases are diagnosed on a presumptive basis, the true prevalence of malaria is unknown in areas around Balombo. Nonetheless, a recent study has shown that malaria is highly prevalent in Benguela province and that severe and complicated malaria occurs in both children and adults [17].
Asymptomatic Plasmodium falciparum carriers
The present study was part of household surveys conducted during the rainy season by the joint teams of Malaria Control Programme of Société nationale de métallurgie (Sonamet) and Services Nationaux de Santé Publique (Angolan Ministry of Health) of Benguela province. Households were selected by cluster sampling to obtain 30 representative households in the study sites. A number was assigned to the households, which were randomly selected using a table of random numbers. After obtaining a written informed consent signed by the head of each household on behalf of all household members, fingerprick capillary blood was obtained from the target group, i.e. children aged less than 15 years old, to prepare thick smears and store blood on Whatman 3MM filter paper. In this household survey, clinical examination was not performed, and the enrolled children were asymptomatic. Blood smears were immediately stained using 10% Giemsa and examined in the field. Parasites were counted against 50 white blood cells for rapid screening in the field. As this household survey on asymptomatic children was part of a training session of inexperienced Angolan laboratory technicians, the gold standard for malaria diagnosis was PCR in this study. If a child was smear-positive and presented fever or other signs and symptoms suggestive of malaria, he or she was referred to the nearest health centre for further examination and appropriate treatment.
Ethical approval
This study was reviewed and approved by the National Malaria Control Programme of the Angolan Ministry of Health, the ethical authority that approves clinical studies on malaria research in Angola.
DNA extraction and PCR
DNA was extracted from Whatman 3MM filter papers using Chelex beads [18]. Briefly, a 2 mm diameter disc was cut from Whatman filter paper and incubated at 4°C overnight in 0.5 mL of phosphate-buffered saline (PBS) containing 0.5% saponin. The filter paper was rinsed twice in saponin-free PBS, placed in 100 μL of distilled water containing 10% Chelex, and incubated at 100°C for 20 min to elute DNA. The supernatant containing extracted DNA was conserved at -20°C. A diagnostic nested PCR was performed using 18S rRNA as the target gene to detect samples with P. falciparum [19].
Molecular markers for drug resistance analysed in the present study (P. falciparum dihydrofolate reductase [pfdhfr], P. falciparum dihydropteroate synthase [pfdhps], P. falciparum chloroquine resistance transporter [pfcrt], and P. falciparum multidrug-resistance gene 1 [pfmdr1]) were amplified by nested PCR, as described in previous studies [20-22]. PCR products were sequenced using the Sanger dideoxy method. Nucleotide sequences were analysed using Chromas 2.4.1 software for DNA sequencing (Technelysium Pty Ltd., South Brisbane, Australia) and aligned using MultAlin software to identify mutations [23].
Data interpretation and statistical calculations
A total of 576 children aged less than 15 years old were enrolled in the present household survey. The sample size was calculated on the basis of an expected smear-positive rate of 25% determined in an earlier pilot study in Balombo, risk of error of 5%, and 5% precision.
The sequences of the following key codons were determined: Ala16Val, Asn51Ile, Cys59Arg, Ser108Asn or Ser108Thr, and Ile164Leu for pfdhfr; Ser436Ala, Ala437Gly, Lys540Glu, Ala581Gly, and Ala613Thr or Ala613Ser for pfdhps; codons Cys72Ser, Val73 (invariable), Met74Ile, Asn75Glu, Lys76Thr for pfcrt; and Asn86Tyr, Tyr184Phe, Ser1034Cys, Asn1042Asp, and Asp1246Tyr for pfmdr1. Based on these codons, different haplotypes were determined and compared. Wild-type pfdhfr haplotype was defined as ANCSI. Double pfdhfr mutant alleles were defined as Ile-51/Asn-108 or Arg-59/Asn-108 (AICNI or ANRNI haplotypes). Triple pfdhfr mutant alleles were defined as Ile-51/Arg-59/Asn-108 (AIRNI haplotype). Wild-type pfdhps refers to SAKAA haplotype. Pfcrt haplotype was defined as wild-type CVMNK and chloroquine-resistant type CVIET. Wild-type pfmdr1 haplotype denotes NYSND.
Results and discussion
A total of 386 blood samples collected in and around Balombo town were screened for the presence of malaria parasites by microscopy and nested PCR. Sixty-one and 80 samples were positive for P. falciparum by microscopy and diagnostic nested PCR, respectively. Sixty PCR-positive samples were further analysed to determine the key codons of pfdhfr, pfdhps, pfcrt, and pfmdr1 genes.
Fifty-eight of 60 samples were successfully amplified to determine pfdhfr alleles. A large majority (69%) carried double Ile-51/Asn-108 mutations, while only 3 (5.2%) had the alternative Arg-59/Asn-108 double mutations (Table 1). Triple pfdhfr mutant alleles (AIRNI haplotype) were present in 20.7% of the field isolates. Both wild-type (n = 2; 3.4%) and single Asn-108 mutant allele (n = 1; 1.7%) were rarely observed. All isolates had the wild-type Ile-164 allele.
Table 1.
pfdhfr , pfdhps , pfcrt , and pfmdr1 haplotypes of Plasmodium falciparum isolates in Balombo area, Benguela province, central Angola
| Molecular marker (no. of isolates analysed) | No. of isolates (%) |
|---|---|
| dhfr (n = 58) | |
| Wild-type haplotype ANCSI | 2 (3.4) |
| Single mutant haplotype ANCNI | 1 (1.7) |
| Double mutant haplotype AICNI | 40 (69.0) |
| Double mutant haplotype ANRNI | 3 (5.2) |
| Triple mutant haplotype AIRNI | 12 (20.7) |
| dhps (n = 30) | |
| Wild-type haplotype SAKAA | 4 (13.3) |
| Single mutant haplotype SGKAA | 18 (60.0) |
| Double mutant haplotype AGKAA | 8 (26.7) |
| pfcrt (n = 36) | |
| Wild-type haplotype CVMNK | 4 (11.1) |
| Mutant haplotype CVIET | 32 (88.9) |
| pfmdr1 (n = 54) | |
| Wild-type haplotype, NYSND | 19 (35.2) |
| Single mutant haplotype, YYSND | 26 (48.1) |
| Single mutant haplotype, NFSND | 6 (11.1) |
| Double mutant haplotype, YFSND | 2 (3.7) |
| Double mutant haplotype, YYSNY | 1 (1.8) |
For each haplotype, mutant alleles are in bold.
In a study on 21 isolates collected in Huambo, a neighbouring province to the east of Benguela province, all carried the mutant Asn-108 pfdhfr allele, and the double mutant haplotype Ile-51/Asn-108 was the most prevalent [24]. Overall, in that study, 25% of the isolates collected from different localities in the country presented the triple pfdhfr mutant alleles. In another study on isolates collected in Luanda [25], 31 of 61 (51%) samples were triple pfdhfr mutants, and 25 of 61 (41%) and 4 of 61 (6.6%) isolates were Ile-51/Asn-108 and Arg-59/Asn-108 double mutants, respectively. In the northern province of Uige, the majority (40 of 66, 61%) of clinical isolates were double Ile-51/Asn-108 pfdhfr mutants, followed by triple mutants (23 of 66, 35%) and the alternative double Arg-59/Asn-108 mutant (n = 1) and wild-type (n = 2) [26]. The results of several recent studies, including those of the present study, suggest that the double pfdhfr mutant AICNI is currently the predominant haplotype in several regions in Angola, with the possible exception of Luanda where triple pfdhfr mutants seem to predominate.
Thirty isolates were randomly selected for sequencing of pfdhps alleles. Most (60%) were characterized by the single mutant Gly-437, i.e. SGKAA haplotype (Table 1). The double mutant haplotype AGKAA and wild-type SAKAA were found in 8 (26.7%) and 4 (13.3%) isolates, respectively. Mutations were not observed in codons 540, 581, or 613.
Similarly, in Luanda, SGKAA was also the most frequent haplotype (18 of 30, 60%), followed by AGKAA (7 of 30, 23%) and the wild-type SAKAA (2 of 30, 7%) [25]. SGEAA and SGEGA haplotypes were found in 3 isolates in that study. The presence of Glu-540 pfdhps allele at a low frequency (14 of 221 samples, 6.3%) was also reported from an earlier study in Luanda [27]. In a study on P. falciparum isolates from different parts of the country [24], SGKAA haplotype largely predominated over other pfdhps haplotypes (339 of 372, 91%). In that study, there were 5 isolates with SGKAA, 1 SGEAA, and 2 SAKAA among 8 isolates collected from Huambo, the neighbouring province of Benguela. The wild-type SAKAA (21 of 372, 5.6%) and double mutant haplotype SGEAA (12 of 372, 3.2%) were found at low frequencies in different regions of Angola. In the northern province, most isolates were characterized by AGKAA (34 of 66, 51.5%), SGKAA (27 of 66, 40.9%), and SAKAA (5 of 66, 7.6%) [26]. Taken together, SGKAA is the major pfdhps haplotype in Angola, including Benguela province, and there appears to be a regional variation in the frequency of AGKAA haplotype. Both Glu-540 mutant allele and the wild-type SAKAA haplotype occur at low frequencies. These molecular data need to be interpreted in the light of clinical efficacy of sulphadoxine-pyrimethamine for intermittent preventive treatment in pregnancy.
Among 36 isolates randomly selected for PCR amplification and sequencing of pfcrt, a large majority (88.9%) were characterized by the mutant haplotype CVIET. The other isolates were of wild-type haplotype CVMNK. Ser-72 was not found in the present study.
In one of the studies conducted in Luanda in which only codon 76 was determined, Thr-76 was found in 230 of 241 (94%) isolates [27]. In the second study in Luanda, the mutant pfcrt haplotypes included the South American-type SVMNT (58 of 102, 57%), CVIET (13 of 102, 13%), and minor and mixed haplotypes [28]. The pfcrt SVMNT haplotype has been rarely reported from the African continent. This haplotype is more commonly found in South America and in some Asian countries and is associated with resistance to amodiaquine and chloroquine [29]. It has been hypothesized that the unusual predominance of SVMNT in Luanda may possibly be due to frequent travels and commercial exchanges between the Angolan capital city and Brazil [28]. However, other molecular studies conducted elsewhere in Angola, including the present study, have not reported the presence of SVMNT haplotype. In Bengo and Uige provinces in the north, only two pfcrt haplotypes were observed: CVIET (273 of 430, including mixed haplotypes, 63.5% in Bengo; 62 of 66 isolates, 94% in Uige) and CVMNK (157 of 430, excluding mixed haplotypes, 36.5% in Bengo; 4 of 66, 6% in Uige) [26,30]. With the exception of SVMNT haplotype found in Luanda, the results of the present study, as well as those performed earlier in other provinces, indicate that most isolates are carriers of mutant CVIET haplotype and are in agreement with the high prevalence of chloroquine-resistant P. falciparum in Angola.
Of 54 isolates analysed in the present study, most (29 of 54, 53.7%) carried the mutant pfmdr1 allele Tyr-86, but mutations occurred less frequently in codons 184 (8 of 54, 14.8%) and 1246 (1 of 54, 1.9%). All isolates had the wild-type Ser-1034 and Asn-1042 pfmdr1 codons.
In an early study, of 15 Angolan P. falciparum isolates (11 chloroquine-resistant and 4 chloroquine-sensitive in vitro), the majority (73%) of chloroquine-resistant isolates carried the mutant Tyr-86 [31]. In Luanda, pure Tyr-86 allele and Tyr- and Asn-86 mixed alleles were found in 142 of 199 (71%) isolates [27]. In the northern province of Uige, many isolates (45 of 66, including mixed Asn- and Tyr-86; 68%) also had the mutant Tyr-86 [26]. However, in Bengo province, the pure wild-type Asn-86 was predominant (94 of 138, 68%), mutant Phe-184 (pure or mixed alleles) occurred in 54 of 138 isolates (39.1%), and codons 1034, 1042, and 1246 were all wild-type [30]. These studies indicate that the majority of Angolan isolates carry the mutant Tyr-86 pfmdr1 allele.
Amodiaquine, in combination with artesunate, is widely used in sub-Saharan African countries. Previous studies have suggested that failure after amodiaquine treatment is associated with the selection of P. falciparum isolates carrying pfcrt Thr-76 allele and pfmdr1 haplotype Tyr-86, Tyr-184, and Tyr-1246 [32-34]. In the present study, most isolates had the mutant pfcrt Thr-76 allele but the pfmdr1 mutant haplotype associated with amodiaquine resistance was rare (1 of 54 isolates, 1.8%). These molecular findings suggest amodiaquine efficacy but require clinical validation studies in Benguela, Angola.
Conclusions
The results of the present study suggest that the high prevalence of mutant pfcrt CVIET haplotype is in agreement with low clinical efficacy of chloroquine observed in an earlier study conducted in the neighbouring province [5]. However, the pfmdr1 haplotype associated with amodiaquine failure was rare, suggesting amodiaquine efficacy. In Benguela province, the double pfdhfr mutant AICNI and single pfdhps mutant SGKAA are currently the predominant haplotypes associated with antifolate resistance. The hallmark of clinical resistance observed in East Africa, i.e. triple pfdhfr mutant haplotype (AIRNI) and double pfdhps mutant haplotype (SGEAA), was absent. These molecular findings need to be further evaluated in parallel with clinical studies, in particular with the efficacy of intermittent preventive treatment using sulphadoxine-pyrimethamine in pregnant women and therapeutic efficacy of artesunate-amodiaquine.
Acknowledgements
The authors wish to thank all households for taking part in this study and Dr Filomena Quinda (Angolan National Malaria Control Programme) for facilitating the malaria survey. This work was supported by the French Agence Nationale de la Recherche (grant number ANR-08-MIE-024).
Abbreviations
- ACT
Artemisinin-based combination therapy
- pfdhfr
Plasmodium falciparum dihydrofolate reductase
- pfdhps
Plasmodium falciparum dihydropteroate synthase
- pfcrt
Plasmodium falciparum chloroquine resistance transporter
- pfmdr1
Plasmodium falciparum multidrug-resistance gene 1
Footnotes
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
PC, PB, FF, and RT conceived the study and participated in study coordination. VFN implemented the study design in the field. CC, NP, and JDA performed laboratory analysis. RT and LKB performed data analysis and interpretation. LKB wrote the draft. All authors read and approved the final manuscript.
Contributor Information
Vincent Foumane Ngane, Email: vfoumane@yahoo.fr.
Joseph Allico Djaman, Email: djamanj@yahoo.fr.
Cécile Culeux, Email: cecile.culeux@etu.parisdescartes.fr.
Nathalie Piette, Email: piettenathalie.pn@gmail.com.
Pierre Carnevale, Email: pcarnevale2001@yahoo.fr.
Patrick Besnard, Email: patrick.besnard@subsea7.com.
Filomeno Fortes, Email: filomenofortes@gmail.com.
Leonardo K Basco, Email: lkbasco@yahoo.fr.
Rachida Tahar, Email: rachida.tahar@ird.fr.
References
- 1.WHO . World Malaria Report. Geneva: World Health Organization; 2013. [Google Scholar]
- 2.Cosep Consultoria, Consaúde, ICF Macro . Inquérito de Indicadores de Malária em Angola de 2011. Maryland: Calverton; 2011. [Google Scholar]
- 3.Kyronseppa H, Lumio J, Ukkonen R, Pettersson T. Chloroquine-resistant malaria from Angola. Lancet. 1984;i:1244. doi: 10.1016/S0140-6736(84)91731-8. [DOI] [PubMed] [Google Scholar]
- 4.Vestergaard Olsen V, Jensen T, Jørgensen M. Chloroquine-resistant Plasmodium falciparum malaria from Angola. Lancet. 1984;i:1462–3. doi: 10.1016/S0140-6736(84)91949-4. [DOI] [PubMed] [Google Scholar]
- 5.Guthmann JP, Ampuero J, Fortes F, van Overmeir C, Gaboulaud V, Tobback S, et al. Antimalarial efficacy of chloroquine, amodiaquine, sulfadoxine-pyrimethamine, and the combinations of amodiaquine + artesunate and sulfadoxine-pyrimethamine + artesunate in Huambo and Bié provinces, central Angola. Trans R Soc Trop Med Hyg. 2005;99:485–92. doi: 10.1016/j.trstmh.2004.11.010. [DOI] [PubMed] [Google Scholar]
- 6.Guthmann JP, Cohuet S, Rigutto C, Fortes F, Saraiva N, Kiguli J, et al. High efficacy of two artemisinin-based combinations (artesunate + amodiaquine and artemether + lumefantrine) in Caala, central Angola. Am J Trop Med Hyg. 2006;75:143–5. [PubMed] [Google Scholar]
- 7.Gosoniu L, Veta AM, Vounatsou P. Bayesian geostatistical modeling of malaria indicator survey data in Angola. PLoS One. 2010;5:e9322. doi: 10.1371/journal.pone.0009322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Thwing JI, Mihigo J, Fernandes AP, Saute F, Ferreira C, Fortes F, et al. How much malaria occurs in urban Luanda, Angola? A health facility-based assessment. Am J Trop Med Hyg. 2009;80:487–91. [PubMed] [Google Scholar]
- 9.Mendes C, Dias F, Figueiredo J, Mora VG, Cano J, de Sousa B, et al. Duffy negative antigen is no longer a barrier to Plasmodium vivax – molecular evidences from the African West coast (Angola and Equatorial Guinea) PLoS Negl Trop Dis. 2011;5:e1192. doi: 10.1371/journal.pntd.0001192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Valente B, Campos PA, Do Rosário VE, Varandas L, Silveira H. Prevalence and risk factors of Plasmodium falciparum infections in pregnant women of Luanda, Angola. Trop Med Int Health. 2011;16:1206–14. doi: 10.1111/j.1365-3156.2011.02830.x. [DOI] [PubMed] [Google Scholar]
- 11.Campos PA, Valente B, Campos RB, Gonçalves L, Rosário VE, Varandas L, et al. Plasmodium falciparum infection in pregnant women attending antenatal care in Luanda, Angola. Rev Soc Bras Med Trop. 2012;45:369–74. doi: 10.1590/S0037-86822012000300017. [DOI] [PubMed] [Google Scholar]
- 12.Soares Magalhães RJ, Langa A, Sousa-Figueiredo JC, Clements ACA, Nery SV. Finding malaria hot-spots in northern Angola: the role of individual, household and environmental factors within a meso-endemic area. Malar J. 2012;11:385. doi: 10.1186/1475-2875-11-385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Sousa-Figueiredo JC, Gamboa D, Pedro JM, Fançony C, Langa AJ, Magalhães RJS, et al. Epidemiology of malaria, schistosomiasis, geohelminths, anemia and malnutrition in the context of a demographic surveillance system in northern Angola. PLoS One. 2012;7:e33189. doi: 10.1371/journal.pone.0033189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Fançony C, Sabastiao YV, Pires JE, Gamboa D, Nery SV. Performance of microscopy and RDTs in the context of a malaria prevalence survey in Angola: a comparison using PCR as the gold standard. Malar J. 2013;12:284. doi: 10.1186/1475-2875-12-284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Cuamba N, Choi KS, Townson H. Malaria vectors in Angola: distribution of species and molecular forms of the Anopheles gambiae complex, their pyrethroid insecticide knockdown resistance (kdr) status and Plasmodium falciparum sporozoite rates. Malar J. 2006;5:2. doi: 10.1186/1475-2875-5-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Toto JC, Besnard P, Le Mire J, Almeida DSI, dos Santos MA, Fortes F, Foumane V, Simard F, Awono-Ambene HP, Carnevale P. Preliminary evaluation of the insecticide susceptibility in Anopheles gambiae and Culex quinquefasciatus from Lobito (Angola), using WHO standard assay. Bull Soc Pathol Exot. 2011;104:307–12. doi: 10.1007/s13149-010-0125-1. [DOI] [PubMed] [Google Scholar]
- 17.Peixoto B, Kalei I. Neurocognitive sequelae of cerebral malaria in adults: a pilot study in Benguela Central Hospital, Angola. Asian Pac J Trop Biomed. 2013;3:532–5. doi: 10.1016/S2221-1691(13)60108-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kain KC, Lanar DE. Determination of genetic variation within Plasmodium falciparum by using enzymatically amplified DNA from filter paper disks impregnated with whole blood. J Clin Microbiol. 1991;29:1171–4. doi: 10.1128/jcm.29.6.1171-1174.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Snounou G, Viriyakosol S, Zhu XP, Jarra W, Pinheiro L, Do Rosario VE, et al. High sensitivity of detection of human malaria parasites by the use of nested polymerase chain reaction. Mol Biochem Parasitol. 1993;61:315–20. doi: 10.1016/0166-6851(93)90077-B. [DOI] [PubMed] [Google Scholar]
- 20.Djimdé A, Doumbo OK, Cortese JF, Kayentao K, Doumbo S, Diourté Y, Dicko A, Su XZ, Nomura T, Fidock DA, Wellems TE, Plowe CV. A molecular marker for chloroquine-resistant falciparum malaria. N Eng J Med. 2001;344:257–63. doi: 10.1056/NEJM200101253440403. [DOI] [PubMed] [Google Scholar]
- 21.Basco LK, Ringwald P. Molecular epidemiology of malaria in Cameroon. X. Evaluation of pfmdr1 mutations as genetic markers for resistance to amino alcohols and artemisinin derivatives. Am J Trop Med Hyg. 2002;66:667–71. doi: 10.4269/ajtmh.2002.66.667. [DOI] [PubMed] [Google Scholar]
- 22.Tahar R, Basco LK. Molecular epidemiology of malaria in Cameroon. XXVII. Clinical and parasitological response to sulfadoxine-pyrimethamine treatment and Plasmodium falciparum dihydrofolate reductase and dihydropteroate synthase alleles in Cameroonian children. Acta Trop. 2007;103:81–9. doi: 10.1016/j.actatropica.2007.04.008. [DOI] [PubMed] [Google Scholar]
- 23.Corpet F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 1988;16:10881–90. doi: 10.1093/nar/16.22.10881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fortes F, Dimbu R, Figueiredo P, Neto Z, Do Rosário VE, Lopes D. Evaluation of prevalences of pfdhfr and pfdhps mutations in Angola. Malar J. 2011;10:22. doi: 10.1186/1475-2875-10-22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gama BE, Pereira-Carvalho GAL, Lutucuta Kosi FJI, de Oliveira NK A, Fortes F, Rosenthal PJ, et al. Molecular markers of antifolate resistance in Plasmodium falciparum isolates from Luanda, Angola. Malar J. 2011;10:248. doi: 10.1186/1475-2875-10-248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Menegon M, Pearce RJ, Inojosa WO, Pisani V, Abel PM, Matondo A, et al. Monitoring for multidrug-resistant Plasmodium falciparum isolates and analysis of pyrimethamine resistance evolution in Uige province, Angola. Trop Med Int Health. 2009;14:1251–7. doi: 10.1111/j.1365-3156.2009.02369.x. [DOI] [PubMed] [Google Scholar]
- 27.Figueiredo P, Benchimol C, Lopes D, Bernardino L, Do Rosário VE, Varandas L, et al. Prevalence of pfmdr1, pfcrt, pfdhfr and pfdhps mutations associated with drug resistance, in Luanda, Angola. Malar J. 2008;7:236. doi: 10.1186/1475-2875-7-236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Gama BE, Pereira-Carvalho GAL, Lutucuta Kosi FJI, de Oliveira NK A, Fortes F, Rosenthal PJ, et al. Plasmodium falciparum isolates from Angola show the StctVMNT haplotype in the pfcrt gene. Malar J. 2010;9:174. doi: 10.1186/1475-2875-9-174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sa JM, Twu O. Protecting the malaria drug arsenal: halting the rise and spread of amodiaquine resistance by monitoring the PfCRT SVMNT type. Malar J. 2010;9:374. doi: 10.1186/1475-2875-9-374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Fançony C, Gamboa D, Sebastião Y, Hallett R, Sutherland C, Sousa-Figueiredo JC, et al. Various pfcrt and pfmdr1 genotypes of Plasmodium falciparum cocirculate with P. malariae, P. ovale spp., and P. vivax in northern Angola. Antimicrob Agents Chemother. 2012;56:5271–7. doi: 10.1128/AAC.00559-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Pinheiro L, Franco S, Adagu IS, Rosa R, Do Rosario VE, Warhurst DC. Detecção da mutação dupla 86Tyr e 1246Tyr no gene pfmdr1 em clones de uma amostra de Plasmodium falciparum da África Ocidental, resistente à cloroquina. Acta Méd Portug. 2003;16:229–33. [PubMed] [Google Scholar]
- 32.Duraisingh MT, Drakeley CJ, Muller O, Bailey R, Snounou G, Targett GA, et al. Evidence for selection for the tyrosine-86 allele of the pfmdr1 gene of Plasmodium falciparum by chloroquine and amodiaquine. Parasitology. 1997;114:205–11. doi: 10.1017/S0031182096008487. [DOI] [PubMed] [Google Scholar]
- 33.Holmgren G, Gil JP, Ferreira PM, Veiga MI, Obonyo CO, Björkman A. Amodiaquine resistant Plasmodium falciparum malaria in vivo is associated with selection of pfcrt76T and pfmdr1 86Y. Infect Genet Evol. 2006;6:309–14. doi: 10.1016/j.meegid.2005.09.001. [DOI] [PubMed] [Google Scholar]
- 34.Humphreys GS, Merinopoulos I, Ahmed J, Whitty CJ, Mutabingwa TK, Sutherland CJ, et al. Amodiaquine and artemether-lumefantrine select distinct alleles of the Plasmodium falciparum mdr1 gene in Tanzanian children treated for uncomplicated malaria. Antimicrob Agents Chemother. 2007;51:991–7. doi: 10.1128/AAC.00875-06. [DOI] [PMC free article] [PubMed] [Google Scholar]


