Abstract
microRNAs (miRNAs) are important modulators of development. Owing to their ability to simultaneously silence hundreds of target genes, they have key roles in large-scale transcriptomic changes that occur during cell fate transitions. In somatic stem and progenitor cells — such as those involved in myogenesis, haematopoiesis, skin and neural development — miRNA function is carefully regulated to promote and stabilize cell fate choice. miRNAs are integrated within networks that form both positive and negative feedback loops. Their function is regulated at multiple levels, including transcription, biogenesis, stability, availability and/or number of target sites, as well as their cooperation with other miRNAs and RNA-binding proteins. Together, these regulatory mechanisms result in a refined molecular response that enables proper cellular differentiation and function.
Mammalian development, which starts from the single-cell zygote, depends on the careful coordination of cell division, differentiation and cell death, to produce the complex tissues and organs of the adult organism. Trillions of cells cooperate to maintain their own fate in coordination with the fate of all other cells in the body. At the molecular level, development is governed by highly regulated activation and suppression of specific gene programmes through transcriptional, post-transcriptional and translational mechanisms. Furthermore, these mechanisms must communicate with one another to maintain robustness. Thus, the regulation of gene programmes depends on complex networks involving feedforward and feedback mechanisms, in which microRNAs (miRNAs) are key players.
miRNAs are short non-coding RNAs that function through the suppression of target genes. The production of miRNAs is a multistep process1. They are typically transcribed by RNA polymerase II (Pol II), and commonly arise from the introns of coding genes or from intergenic long non-coding RNAs called primary miRNAs (pri-miRNAs). pri-miRNAs contain one or more miRNAs within hairpins. These hairpins are cleaved from the pri-miRNA transcript in the nucleus by the Microprocessor complex, which consists of the RNA-binding protein (RBP) DGCR8 and the RNA endonuclease Drosha. The resulting pre-miRNA hairpins are transported to the cytoplasm where they are further processed into approximately 21-nucleotide-long double-stranded RNAs (dsRNAs) by the endonuclease Dicer. These processing steps represent the biogenesis of canonical miRNAs. Small numbers of non-canonical miRNAs are produced by alternative pathways2. Importantly, the existence of these crucial steps in the biogenesis of canonical miRNAs has enabled the study of global miRNA knockouts, by removing any one of the proteins involved in biogenesis. In mice, the knockout of any of these proteins results in early embryonic lethality, indicating that miRNAs are essential for mammalian development3,4. Numerous tissue-specific knockouts of these proteins have also been studied, examples of which are outlined in TABLE 1. In all the tissues that have been tested, global miRNA loss induces dramatic phenotypic changes, with one surprising exception: the maturing oocyte.
Table 1.
Examples of tissue-specific global microRNA knockouts and their effects
| Lineage | Defect | Refs |
|---|---|---|
| Limb | Morphogenesis defect and apoptosis | 94 |
| Skin | Hair follicle developmental defects | 62,63 |
| Pancreas | Increased apoptosis and β-cell differentiation defect | 95 |
| Skeletal muscle | Increased apoptosis and myofibre morphogenesis | 20 |
| B cells | Increased apoptosis | 49 |
| Neurons | Increased apoptosis and defective neuronal differentiation | 73 |
| Osteoblasts | Differentiation defect | 96 |
| Chondrocytes | Differentiation defect | 97 |
| Adipose tissue | Impaired differentiation of white adipose tissue | 98 |
| Kidney | Increased apoptosis, and defects in morphogenesis of nephrogenic and uretogenic compartments | 99 |
| Haematopoiesis | Increased apoptosis and differentiation skewed towards the myeloid lineage | 46 |
| Oligodendrocytes | Differentiation defect | 75,76 |
| Neural crest | Increased apoptosis | 100 |
| Tooth | Morphogenesis defects and impaired differentiation | 101 |
| Oocytes | No defects observed | 102 |
After processing of the pre-miRNA, one strand of the Dicer cleavage product enters the silencing complex, a ribonucleoprotein (RNP) complex that consists of the miRNA bound within a pocket of the argonaute proteins (AGO1–4 in mice and humans) along with associated proteins5. The silencing complex then recognizes targets by partial base pairing of the miRNA with, typically, the 3′ untranslated region (3′ UTR) of an mRNA, or, less commonly, the open reading frame. Binding inhibits translation and/or destabilizes the mRNA. Each miRNA targets hundreds of mRNAs, resulting in a broad effect on the mRNA content of cells even though each individual target is only partially suppressed6. Much effort has been put into identifying the targets of individual miRNAs and determining the extent to which modulation of any one target represents the functional effect of the miRNA. More recently, efforts have been made to determine how the many targets can function together to influence phenotypic outcomes7. Surprisingly, individual miRNA knockouts often lack dramatic phenotypic consequences8. For this reason, it has been proposed that miRNAs have a greater role in regulating the robustness of the cellular state than they do in driving cell fate choices9. However, the exogenous introduction of miRNAs can drive cells towards specific fates103. Therefore, it is likely that they are often part of redundant circuits that cannot be revealed by individual knockouts.
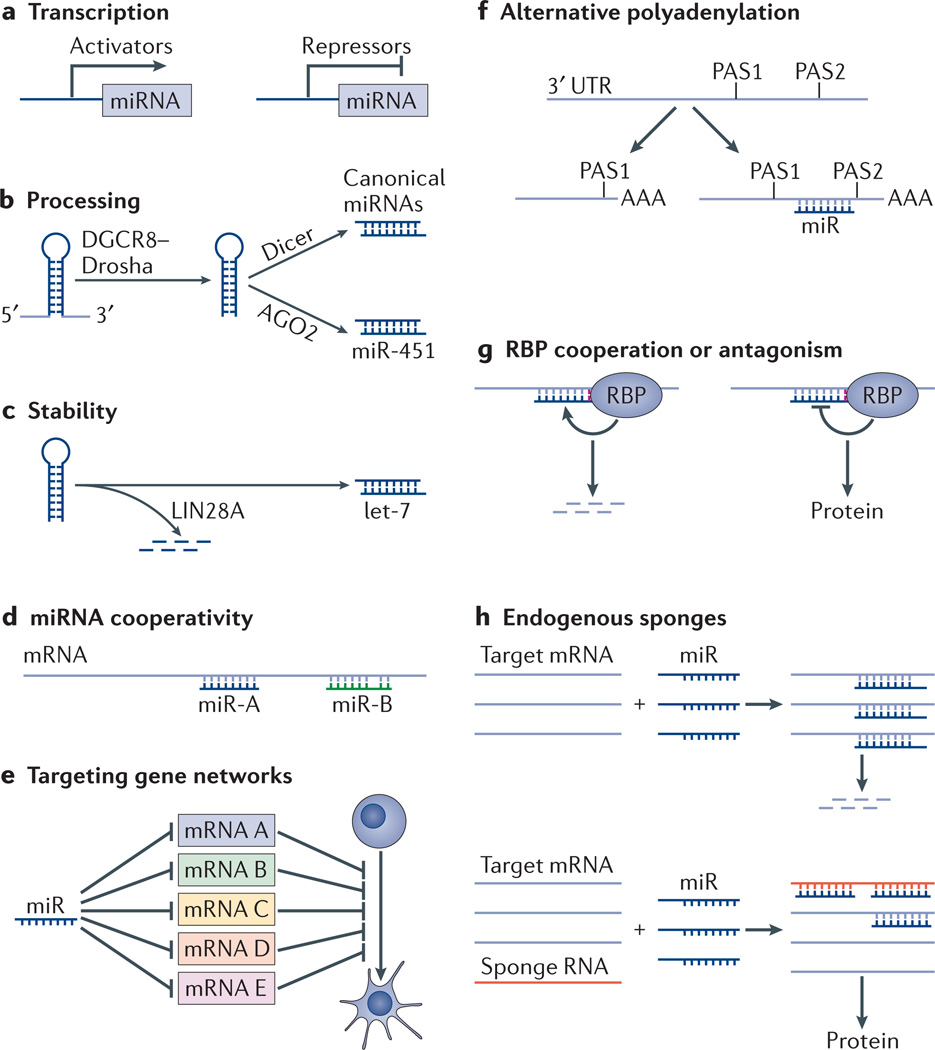
In addition to the focus on identifying miRNA-regulated targets and the downstream consequences of target suppression, there has been increasing attention given to the regulation of the miRNAs themselves (FIG. 1). miRNA levels are regulated by transcriptional and epigenetic mechanisms that are common to coding RNAs (FIG. 1a). miRNA abundance is further regulated through post-transcriptional mechanisms that affect their processing and stability (FIG. 1b,c). For example, the RBPs LIN28A and LIN28B inhibit the biogenesis of and destabilize the let-7 family of miRNAs, which are frequently used across tissue types to promote differentiation and maturation1,10,11. Beyond their levels, miRNA function is also modulated by regulating the interaction of miRNAs with their targets (FIG. 1d–h). For example, the suppression of target mRNAs is strongly influenced by the number of miRNA target sites in the 3′ UTR. There are often multiple target sites in an mRNA 3′ UTR, either for a single miRNA or for a combination of miRNAs, resulting in cooperative downregulation of a target mRNA and occasionally in redundancy among miRNAs. The availability of target sites themselves is also influenced by 3′ UTR length, which can be regulated through alternative polyadenylation (APA). The effect of the miRNA on individual targets can additionally be influenced by cooperation with RBPs that have their own binding specificities. Finally, recent work suggests that competition between targets can influence miRNA activity on any one target.
Figure 1. Regulation of miRNA biogenesis and function occurs at multiple levels.
a | microRNA (miRNA) expression is regulated by transcriptional and epigenetic activators and repressors. b | Post-transcriptional regulation can be modulated at the level of processing. A complex composed of DGCR8 and Drosha cleaves the primary transcript to generate a precursor miRNA hairpin, which is then processed by Dicer to generate the mature product. miR-451 is unique as it bypasses Dicer processing and is directly processed by argonaute 2 (AGO2). c | Mature miRNAs and their processing intermediates (pre-miRNAs) can be destabilized; for example, members of the let-7 miRNA family, which are destabilized by the RNA-binding protein (RBP) LIN28A. d | Multiple miRNAs can act on a single mRNA cooperating to destabilize the transcript. e | A single miRNA can target many mRNAs in a gene network that drives a specific cell fate transition. f | Alternative mRNA polyadenylation results in 3′ untranslated regions (3′ UTRs) of different lengths, which therefore may or may not contain miRNA target sites. g | RBPs can bind an mRNA and promote or inhibit miRNA activity at a neighbouring site, leading to mRNA degradation or translation, respectively. h | Expression of endogenous ‘sponges’ (with sequences partially complementary to miRNAs) has been proposed to interfere with miRNA-mediated inhibition of a target mRNA, thus promoting messenger stability and translation. PAS, polyadenylation site.
In this Review, we discuss examples of these miRNA regulatory mechanisms that have roles in somatic stem and progenitor cells, both during development and in the adult. Somatic stem and progenitor cells are particularly appropriate for the study of miRNA control and function, as these cells are constantly undergoing either cell division, differentiation towards one or multiple cell lineages, or cell death. miRNAs have an important role in all these alternative fates. As the literature is much too extensive to provide a comprehensive discussion of this topic, we focus instead on four model lineages: skeletal muscle, blood, skin and neural lineages (FIG. 2). Within these lineages we attempt to provide examples of how specific miRNAs in stem and/or progenitor cells are integrated into the central gene networks that control cell fate transitions and cellular identity. The lineages chosen exemplify the variety of developmental paths taken in different tissues, and studies therein contain a rich body of work to illustrate the diversity of miRNA function. Our goal is not to comprehensively review the literature for each of these tissues, but to identify recurrent and emergent themes of miRNA biology in stem and/or progenitor cells during fetal organogenesis and adult homeostasis. For each tissue, we discuss the mechanisms that regulate miRNA expression and miRNA activity, along with the resulting downstream effects. Whereas some mechanisms span the four tissue types, others have so far been uncovered either in one tissue type only or in a subset of these tissues.
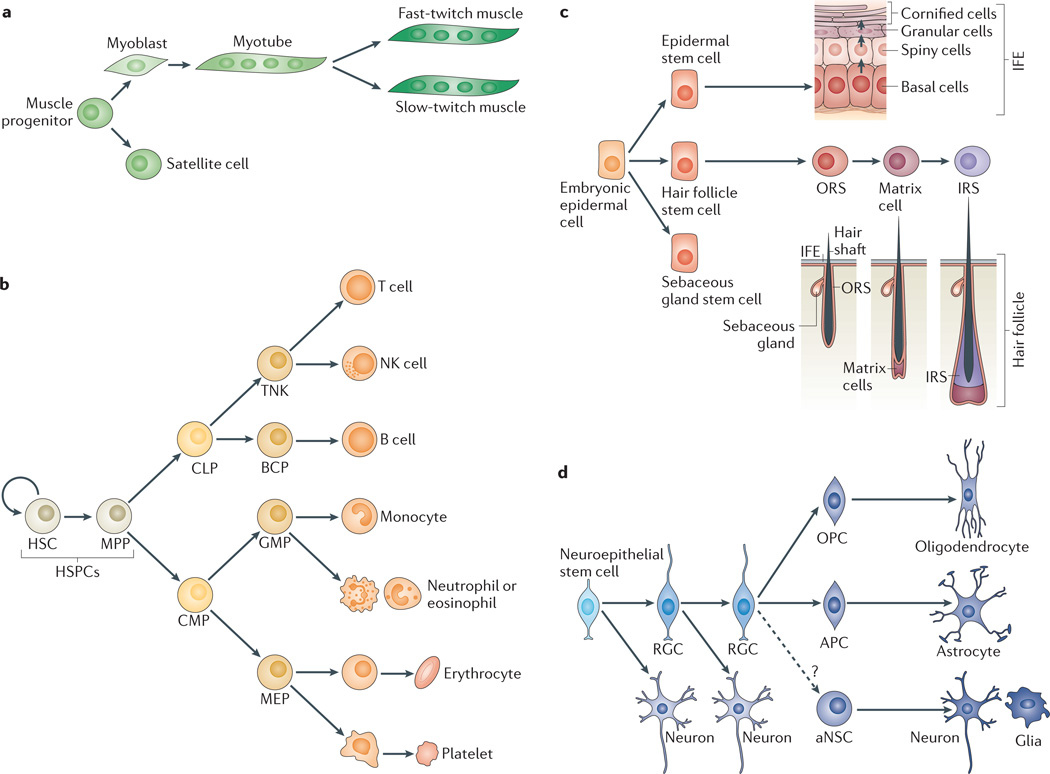
Figure 2. Lineage-restricted stem and progenitor cell differentiation during the development of four model lineages.
a | In skeletal muscle, progenitors that give rise to myoblasts arise from the paraxial mesoderm. Myoblasts terminally differentiate into muscle cells, which fuse to give rise to myotubes. The myotubes can produce fast-twitch or slow-twitch muscle fibres. Muscle progenitors also give rise to satellite cells, which serve as adult stem cells of the muscle. b | In the blood, self-renewing haematopoietic stem cells (HSCs) give rise to multipotent progenitors (MPPs), which further differentiate into the progenitors of lymphoid lineage (common lymphoid progenitor (CLP)) or myeloid lineage (common myeloid progenitor (CMP)). CLPs differentiate into progenitors of B cells (BCPs) and the progenitors of T cells and natural killer cells (TNKs). CMPs give rise to megakaryocyte–erythroid progenitors (MEPs; which differentiate into erythrocytes and platelets) and the granulocyte–macrophage progenitors (GMPs; which differentiate into monocytes, neutrophils and eosinophils). c | The epidermis is a stratified epithelium composed of progenitor cells in the innermost basal layer and successive layers of differentiating cells as one moves to the surface. The hair follicle develops when primitive epidermal cells form placodes that differentiate into hair follicle stem cells, which in turn undergo further sequential differentiation to produce the outer root sheath (ORS) progenitor cells, the matrix cells, the inner root sheath (IRS) cells, and finally the mature hair follicle. Epithelial stem cells at the junction of the epidermis and hair follicle give rise to the sebaceous gland. d | Neural — neuroepithelial cells expand and produce early neurons, as well as radial glial cells (RGCs). For most of embryogenesis, RGCs divide asymmetrically producing a neuron and a new RGC. During the end of embryogenesis and in early postnatal stages, RGCs acquire gliogenic competence (astrocyte precursor cell (APC) and oligodendrocyte progenitor cell (OPC) for the astrocyte and oligodendrocyte precursor cell, respectively), giving rise to astrocytes and oligodendrocytes. In later postnatal stages, adult neural stem cells (aNSCs) can give rise to neurons or glia. IFE, interfollicular epidermis. HSPCs, haematopoietic stem and progenitor cells.
miRNAs in myogenesis
During development, multipotent cells in the mesoderm produce progenitors of skeletal muscle called myoblasts, as well as quiescent adult muscle stem cells called satellite cells12 (FIG. 2a). Myoblasts proliferate, differentiate and then fuse to form the multinucleated myotubes of the muscle fibre. In vivo studies of muscle development are often complemented with in vitro approaches using a mouse myoblast cell line, C2C12. C2C12 cells can be maintained in culture and induced to terminally differentiate into myotubes13.
Regulation of miRNA levels
Extensive miRNA studies during myogenesis have led to a detailed understanding of how lineage-specific miRNAs are integrated within regulatory transcriptional and epigenetic networks. The presence of binding sites for myogenic transcription factors in miRNA promoters, as well as the locations of some miRNA loci within introns of myogenic genes, result in highly regulated expression. For example, myogenic transcription factors that are known drivers of skeletal muscle specification and differentiation, including MyoD, myogenin and MYF5, bind to and activate the promoters of miR-206, miR-486 and miR-499 and the bicistronic miRNAs miR-1 and miR-133 (REFS 14–18) (FIG. 3a). Additionally, miRNAs can be found in the introns of muscle-specific genes, such as miR-208 and miR-499, which are located in the introns of Myh7 (encoding β-MHC) and Myh7b, respectively19. Therefore, miRNA expression is transcriptionally regulated by the same upstream factors that drive the expression of the coding RNAs that are important for muscle development and maturation.
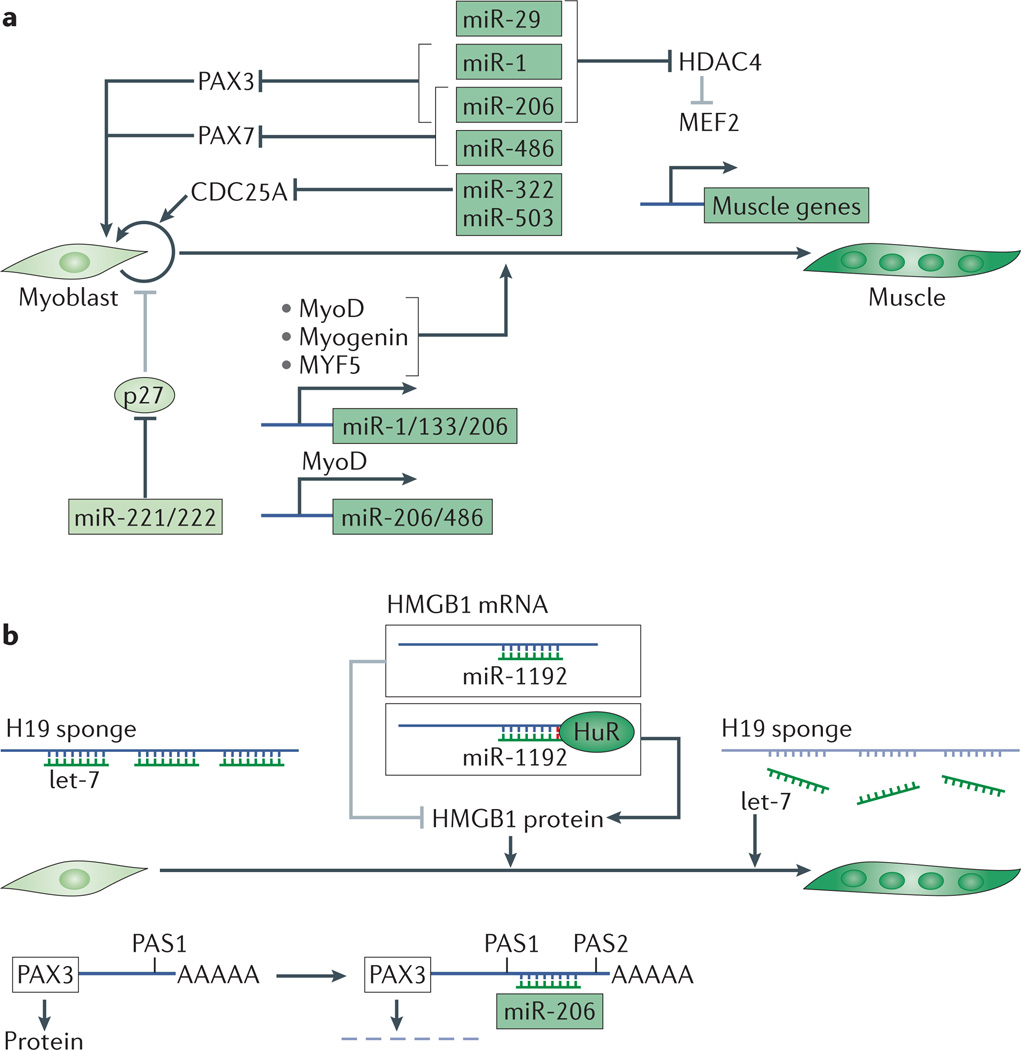
Figure 3. miRNA regulation during myogenesis.
a | microRNAs (miRNAs) that are expressed at high levels in myoblasts (light green box) promote cell cycle progression by suppressing the translation of the cell cycle inhibitor p27. miRNAs display increased expression during myogenesis (dark green box) and are transcriptionally activated by myogenic transcription factors such as MYF5, MyoD and myogenin. These miRNAs both repress genes that promote the progenitor fate and repress genes that silence the muscle programme. For example, some miRNAs inhibit the transcription factors PAX3 and PAX7 and repress the cell cycle regulator CDC25A. Others inhibit the epigenetic repressor histone deacetylase 4 (HDAC4), which in turn leads to derepression of the muscle transcription factor MEF2 and subsequent upregulation of muscle genes. b | Regulation of miRNA activity during myogenesis. miRNA activity during muscle differentiation is modulated by the RNA-binding protein HuR, which binds next to the miR-1192 binding site on the 3′ untranslated region (3′ UTR) of the transcript encoding the chromatin-binding protein HMGB1, inhibiting miRNA-mediated suppression of this gene. Alternative polyadenylation of the mRNA encoding PAX3 results in either a long form that contains a binding site for miR-206 in its 3′ UTR or a short form that does not contain the binding site and, therefore, is resistant to miRNA-mediated suppression. H19 is a long non-coding RNA with multiple let-7 binding sites, which has been proposed to titrate let-7 away from other targets such as Dicer and HMGA2, thus relieving the suppression of these targets. Dark lines represent activation and grey bars represent suppression.
miRNA function
Global loss of miRNAs using a conditional Dicer knockout in myogenic tissues results in widespread apoptosis and abnormal myofibre morphology20. Many studies over the past decade have identified roles for specific miRNAs in myoblast differentiation and fusion. These miRNAs are integrated into gene networks that regulate both proliferation and differentiation (FIG. 3a). For example, the myogenic miRNAs miR-1 and miR-206 target PAX3, and miR-206 and miR-486 target PAX7 during development21,22. PAX3 and PAX7 are transcription factors that promote myoblast fate and are downregulated during differentiation as the myogenic miRNAs are upregulated. miR-1 and miR-206 have analogous roles in the adult by targeting PAX3 and PAX7 during the differentiation of postnatal satellite cells23,24. Other miRNAs directly target the cell cycle machinery during myogenesis. For example, miR-221 and miR-222, which are downregulated during myogenesis, causing reciprocal upregulation of their target, the cell cycle inhibitor p27, and subsequent cell cycle exit25. In a study limited to the C2C12 model of myogenesis, miR-322 and miR-503 were shown to be upregulated during differentiation and to downregulate the CDC25A target to promote cell cycle exit26. These examples demonstrate various mechanisms by which increased or decreased expression of different miRNAs reinforces terminal differentiation by inhibiting cell cycle and transcriptional networks that are active in the progenitor state.
In addition to suppressing the progenitor state, miRNAs are also integrated into the regulatory circuits that establish a new gene expression programme during differentiation. In C2C12 cells, miR-1, miR-206 and miR-29 inhibit the expression of the histone deacetylase HDAC4, thus leading to the derepression of genes that are bound by its myogenic partner, the transcription factor MEF2 (REFS 27,28). Another example, miR-214, targets the Polycomb repressive complex protein EZH2, which leads to derepression and the expression of at least two muscle-specific genes, muscle creatine kinase and myosin29, thus aiding the establishment of the muscle programme. miRNAs are also involved in regulating the maturation of muscle cells into either fast-twitch or slow-twitch myotubes. Specifically, miR-499 and miR-208b target the transcription factor SOX6 to promote the slow-twitch fate19. Therefore, miRNAs have roles in all aspects of the myogenic programme, including exit from the cell cycle, silencing of the progenitor programme and activation of the terminal differentiation programme.
Regulation of miRNA activity
miRNA cooperativity, APA and RBPs have important roles in regulating miRNA activity during muscle development. As discussed above, there are multiple examples in the muscle of miRNAs cooperating to downregulate specific targets by binding distinct sites on the same mRNA (FIG. 3a). Conversely, APA suppresses miRNA activity by producing shortened 3′ UTRs that often lack miRNA sites that would exist in the longer 3′ UTR isoforms30. For example, in the satellite cells of limb and diaphragm muscles, miR-206 and its target PAX3 are co-expressed. However, the PAX3 transcript is resistant to miR-206 suppression because APA produces a short isoform of the transcript missing the miR-206 binding sites. Upon differentiation, the longer form of PAX3 is produced, which is then targeted by miR-206 (REF. 23) (FIG. 3b). The presence of miR-206 in the satellite cells brings up the interesting possibility that its expression had been co-opted for alternative targets specifically in these muscle stem cells and, therefore, that APA was required for the resistance of stemness genes such as Pax3 to miR-206 suppression. RBPs also regulate miRNA activity: they bind to the 3′ UTRs of mRNAs and modulate their levels and translation often through the regulation of neighbouring miRNA binding sites. For example, during C2C12 differentiation, the RBP HuR (also known as ELAVL1) has been shown to inhibit miR-1192 suppression of High mobility group protein B2 (HMGB2), which promotes differentiation31 (FIG. 3b). These examples highlight the complexity of miRNA control downstream of miRNA production.
A new level of miRNA regulation has been recently proposed based on the concept of competition between targets for an miRNA32. For example, during myoblast differentiation, it has been suggested that the long non-coding RNA, linc-MD1, sequesters miR-133 and miR-135, thus allowing the expression of their targets MAML1 and MEF2C, respectively, which promote muscle differentiation33. Such a model is surprising as this linc-MD1 has only one binding site for miR-133 and two binding sites for miR-135, thus making it difficult for this long non-coding RNA to compete with mRNAs for the binding to miRNAs, as target mRNAs are cumulatively much more highly expressed. Another reported example of this phenomenon in myogenesis is the imprinted long non-coding RNA H19, which has several binding sites for let-7, a family of miRNAs that promotes myoblast differentiation34. Future studies in which targeted mutations to the miRNA sites are made in the endogenously expressed forms of these RNAs are needed to determine to what extent these RNAs truly act as competitive sponges.
miRNAs in haematopoiesis
In contrast to muscle tissue where various subtypes are specified from distinct regions of the mesoderm, the diverse sets of cells that constitute the blood can be derived from a single stem cell. Furthermore, haematopoiesis is a well-characterized lineage-specification process that involves successive bifurcations that lead to increasingly specialized cells and, therefore, comprises several different mechanisms of cell fate control. During fetal development, primitive haematopoietic cells are first specified in the blood islands of the yolk sac and then differentiate into erythrocytes. Following cessation of primitive haematopoiesis, definitive haematopoietic stem cells (HSCs) are initially found in the fetal liver where they undergo rapid expansion and differentiation into multipotent progenitors that give rise to the lymphoid or myeloid lineage, which further differentiate to give rise to the diverse cell types found in the blood (reviewed in REF. 35) (FIG. 2b). The HSCs and multipotent progenitors are collectively referred to as haematopoietic stem and progenitor cells (HSPCs). Later in development, the site of haematopoiesis shifts from the liver to the bone marrow. Adult HSCs are generally quiescent and have greatly reduced proliferation in comparison to fetal HSCs. miRNAs have roles in all these developmental stages.
Regulation of miRNA levels
Similar to examples of the muscle lineage, miRNAs that are necessary for the proper development of the haematopoietic lineage are regulated by cell type-specific transcription factors (FIG. 4a). For example, during erythropoiesis, the key regulator GATA1 activates the transcription of miR-451 and miR-23, which promote terminal differentiation of erythroid progenitors36,37. Another interesting example is the transcription factor–co-repressor complex that consists of KRAB-ZFP and KAP1 (also known as TIF1β), known as KRAB-ZFP–KAP1, which suppresses several miRNAs during late-stage mouse erythroid differentiation38. In particular, the zinc finger proteins (ZFPs) ZFP689 and ZFP13 are upregulated at the late stages of erythroid differentiation and bind near the chromosomal region expressing miR-351. The ZFPs recruit KAP1, which in turn recruits epigenetic repressors that suppress miR-351 expression. miR-125a, which is a member of the same miRNA family, is regulated in an analogous manner in human cells. KRAB-ZFP–KAP1 complexes have generally been studied in the context of silencing retrosposons, but examples such as this one are beginning to uncover roles for these factors in cell fate regulation39.
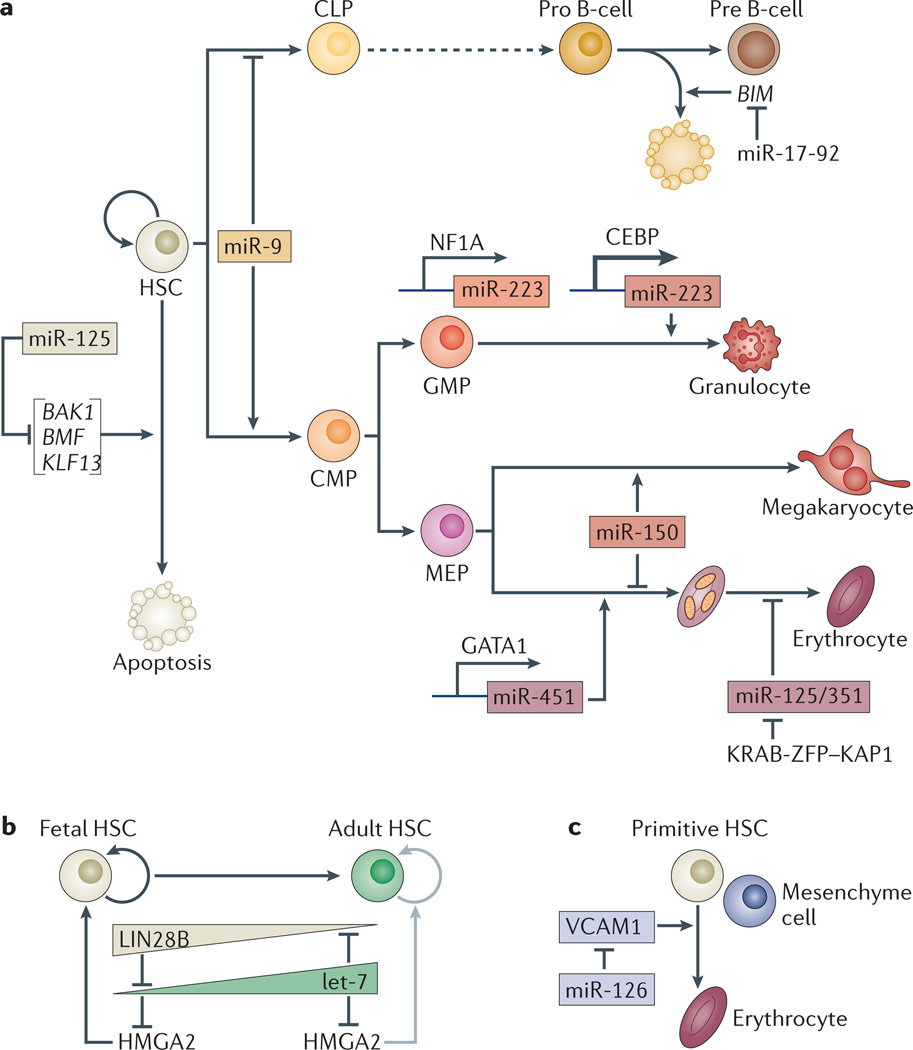
Figure 4. miRNAs regulate cell fate choices and differentiation during haematopoiesis.
a | microRNAs (miRNAs) enriched in one cell type (outlined by the colour of the cell type) can promote the differentiation of progenitors (miR-223) or can inhibit the alternative lineages of oligopotent progenitors (miR-9 and miR-150). miRNAs also inhibit apoptosis in progenitors during haematopoiesis (miR-125 targets pro-apoptotic genes BAK1, KLF13 and BMF, and miR-17-92 targets BIM). The inhibition of miRNAs (miR-125/351) by a KRAB-ZFP transcription factor and its co-repressor KAP1 is necessary to enable mitophagy during erythrocyte maturation. miRNA levels are regulated at the transcriptional level. Levels of miR-223 are modulated by sequential binding of the transcription factors NF1A and CCAAT/enhancer-binding protein-β (CEBP). Binding of CEBP leads to maximal activation in granulocytes. The miR-451 locus is activated by binding of the erythrocyte-specific transcription factor GATA1. b | miRNAs that regulate developmental transitions in haematopoiesis from the embryo to the adult are indicated. The modulation of let-7 expression leads to a reduction in self-renewal in adult haematopoietic stem cells (HSCs) relative to fetal HSCs. In fetal HSCs, high levels of RNA-binding protein LIN28B repress let-7 biogenesis, which in turn leads to high levels of HMGA2. In adult HSCs, LIN28B levels are low, which leads to high let-7 activity and repression of the chromatin factor HMGA2. c | miRNAs expressed in niche cells can regulate stem cell proliferation and differentiation. For example, miR-126, expressed in neighbouring mesenchyme cells (outlined by a lilac box), represses the adhesion molecule vascular cell adhesion protein 1 (VCAM1) to promote maturation of primitive HSCs. CLP, common lymphoid progenitor; CMP, common myeloid progenitor; GMP, granulocyte–monocyte progenitor; MEP, megakaryocyte–erythroid progenitor. HSPC, haematopoietic stem and progenitor cell.
Granulocyte differentiation provides another model, in which expression of miRNAs is tightly controlled in a temporal manner. In particular, mouse miR-223 is regulated by two competing transcription factors: NF1A (also known as NF1X) binds to the miR-223 promoter in progenitors, resulting in low expression levels; however, during differentiation, the same sites are bound by CEBP (CCAAT/enhancer-binding protein-β), resulting in higher expression levels40 (FIG. 4a). Other miRNAs are regulated post-transcriptionally. For example, the let-7 miRNA family, which inhibits cell cycle progression, is regulated post-transcriptionally in HSCs. Mature let-7 miRNAs are highly expressed in adult mouse HSCs, but let-7 biogenesis is suppressed by LIN28B in fetal HSCs41 (FIG. 4b). Additionally, some members of the let-7 miRNAs are co-transcribed in a tricistron that is composed of miR-99/100 and miR-125 family miRNAs in human HSPCs and megakaryocytes. In haematopoietic cells, these clusters can be activated by the transcription factor HOXA10 (REF. 42). Thus, miRNA levels are tightly regulated by lineage-specific transcription factors during haematopoiesis.
miRNA function
The haematopoietic lineage provides excellent examples of context-dependent miRNA functions. As fetal HSCs mature they transition from rapidly proliferating fetal HSCs to quiescence in the adult, and let-7 plays an important part in this transition. As mentioned above, let-7 levels are suppressed post-transcriptionally by LIN28B in rapidly dividing fetal HSCs in mice. As LIN28B levels decrease and let-7 levels increase in the transition to adult HSCs, this promotes quiescence in part by suppressing HMGA2, thus relieving the inhibition of the HMGA2 targets the cyclin-dependent kinase inhibitors p16 and p19 (REF. 41). The let-7 to LIN28B switch in expression during HSC fetal–adult transition has also been reported in the lymphoid lineage43. Experimentally induced expression of LIN28B and the subsequent downregulation of let-7 result in a fetal-like differentiation programme. These studies exemplify the recurring roles for a number of miRNAs at different stages in a single lineage.
The sheer diversity of cell types generated from a single pool of HSCs during haematopoiesis requires that cell fate choices at each step be highly regulated. Studies have identified individual miRNAs that function at each of these steps (FIG. 4a). For example, expression of miR-9 promotes myelopoiesis and inhibits lymphopoiesis during differentiation of HSPCs44. Additionally, the expression of miR-150 promotes the differentiation of megakaryocyte–erythroid progenitor (MEP) cells into megakaryocytes and suppresses the erythroid lineage45. A number of studies have also shown that overexpression of miR-125 promotes the expansion of murine and human HSCs with a bias towards the development of myeloid lineages. Indeed, overexpression is associated with the development of a myeloproliferative disorder46,47. Strangely, one study has suggested that miR-125 induces a bias towards lymphoid lineages48. The basis of these presumably contradictory results will need to be resolved and could involve subtle differences in the cells targeted and/or the assays used as a readout for functional outcomes.
miR-125 and other miRNAs also have important roles in regulating apoptotic genes and hence cell death in both HSCs and downstream lineages (FIG. 4a). The loss of Dicer in HSPCs leads to their reduction owing to survival defects. Overexpression of the highly expressed miR-125 promotes the survival and expansion of wild-type HSPCs. miR-125 targets the pro-apoptotic genes BAK1, BMF and KLF13, and the overexpression of BAK prevents miR-125-mediated expansion of normal HSPCs46,48. Further down the haematopoietic lineage, miRNAs within the miR-17–92 cluster target the apoptotic gene BIM to regulate apoptosis in the pro-B cell to pre-B cell transition104. Loss of Dicer, and hence miR-17–92 miRNAs, during this transition leads to the upregulation of BIM and to increased cell death, which can be reversed by the removal of BIM49. These studies highlight the functional importance of miRNAs in regulating cell survival during stem cell growth and differentiation.
Individual miRNAs also drive differentiation within specific haematopoietic lineages (FIG. 4a). For example, miR-223 targets both NF1A and MEF2C to promote granulocyte differentiation40,50. The downregulation of NF1A leads to increased miR-223 levels, producing a positive feedback loop. Another crucial miRNA cluster that has more recently been the subject of extensive studies is the miR-144/451 cluster. The loss of this cluster leads to severe defects during erythropoiesis, attributable to the derepression of 14-3-3ζ, a regulator of cytokine signalling51. Inhibition of 14-3-3ζ is sufficient to reverse the effect of miR-451 loss. A particularly interesting aspect of this miRNA cluster is that, once transcribed and processed by the Drosha–DGCR8 complex, its biogenesis bypasses Dicer processing and instead requires the catalytic activity of AGO2 to generate a mature product52–54. Although most studies focus on miRNAs that are activated and promote a specific cell fate transition, a recent study has highlighted the necessity for the miR-351 cluster to be suppressed during erythroid differentiation38. The knockdown of genes expressing the KRAB-ZFP–KAP1 repressor complexes that normally downregulate these miRNAs at late stages of differentiation leads to diminished erythroid maturation and a resulting anaemia. These miRNAs target factors that are important in mitophagy, a crucial cellular event in erythrocyte maturation. Therefore, the increased expression of these targets is a key step in the final stages of erythropoiesis. Together, these studies show that temporally regulated activity of distinct miRNAs is necessary for establishing the unique characteristics of haematopoietic lineages.
In addition to their cell-autonomous roles, miRNAs in the cells of the niche can also regulate stem and progenitor cell function. Before the production of definitive HSCs, haematopoiesis occurs in the yolk sac of the early embryo. These primitive haematopoietic cells, called EryP-CFCs, are specified in the endothelium and exist for a defined developmental window. miR-126, which is expressed in the neighbouring mesenchyme cells, down-regulates the cell surface marker vascular cell adhesion protein 1 (VCAM1), thus promoting the maturation of EryP-CFCs and, subsequently, the cessation of primitive haematopoiesis55 (FIG. 4c). In the adult, the loss of miRNA in mouse osteoprogenitors, which form the HSC niche, leads to myelodysplasia, as well as to increased apoptosis and proliferation of haematopoietic progenitors56. The specific osteoprogenitor miRNAs that are responsible for these defects remain unknown. Future studies will need to elucidate whether the HSC defects are due to defective development of osteoblasts or due to direct regulation of signalling between the niche and HSC compartments by miRNAs.
These examples show that, analogous to miRNA function in myogenesis and consistent with the phenotypes seen with Dicer loss, miRNAs expressed during haematopoietic development promote the survival and differentiation of stem and progenitor cells. Additionally, they have a major role in regulating cell lineage choice and in promoting the development of the specific cell types that define the haematopoietic lineage.
miRNAs in skin development
As in the haematopoietic lineage, mammalian skin is composed of a diversity of cell types that give rise to the epidermis, hair follicle and sebaceous gland. All these cells initially arise from multipotent epithelial cells. However, once established, each skin appendage is sustained by a distinct stem cell population57 (FIG. 2c). The balance between stem cell maintenance and differentiation in each of these populations must be carefully coordinated to replenish the tissue during its constant turnover. In vivo studies in the skin are often complemented with work in vitro using keratinocyte cultures derived from the dissociation of the epidermis. Keratinocytes can be maintained in proliferative conditions and induced to differentiate into epidermal cells.
Regulation of miRNA levels
Purification of epidermal and follicular cells at different stages of differentiation by a combination of reporter genes (K14-RFP and SOX9-GFP) and cell surface markers (α-integrin) has made it possible to evaluate miRNA changes during these transitions58. Distinct patterns of miRNA expression have been found in the purified populations. For example, miR-125 and miR-205 are highly expressed in interfollicular epidermal stem cells, hair follicle stem cells and the outer root sheath (ORS) progenitors, but are downregulated in their differentiated progeny58,59. Furthermore, in situ studies have shown that miR-203 is specifically enriched in differentiating cells of the epidermis. The cell type-specific expression of these and other miRNAs suggests highly coordinated transcriptional regulation. Indeed, the miR-203 promoter has putative binding sites for transcription factors that are enriched in epidermal cells60. Furthermore, in primary mouse keratinocyte cultures, p63 binds to the promoter of miR-34 family miRNAs thus inhibiting their expression61. The loss of p63 in vivo in mouse epidermis is accompanied by increased expression of miR-34 and cell cycle arrest. Although these are important examples, much remains to be learned regarding the basis of the expression levels for the subsets of miRNAs enriched in the different subpopulations of the skin.
miRNA function
The loss of miRNA biogenesis proteins has been extensively studied in developing and postnatal skin compartments owing to the availability of specific Cre drivers in mice. During development, DGCR8 and Dicer loss results in increased apoptosis and in striking defects in the morphogenesis of the hair follicle. miRNA loss in the developing epidermis also promotes hyper-proliferation and apoptosis62,63. In postnatal skin, the loss of Dicer in the hair follicle leads to defects during the growth phase (anagen) and entry into the regression phase (catagen) but resting hair follicles are relatively unaffected64; by contrast, the interfollicular epidermis exhibits increased proliferation. These data suggest diverse roles of miRNAs in regulating the various skin lineages, consistent with findings associated with functional dissection of individual miRNAs (FIG. 5).
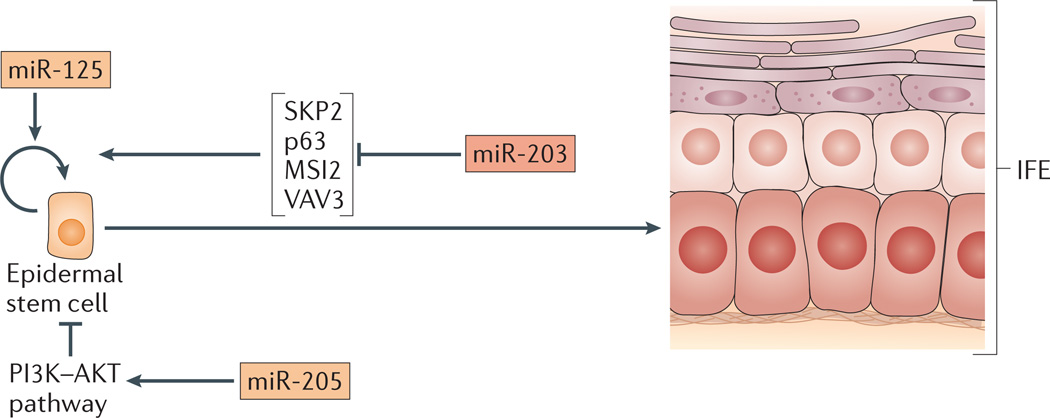
Figure 5. miRNAs regulate the development of the epidermis and hair follicle.
microRNAs (miRNAs) function during differentiation of epidermal stem cells into cells of the interfollicular epidermis (IFE). miR-205 and miR-125 promote self-renewal of epidermal stem cells but are downregulated during differentiation. miR-205 suppresses multiple inhibitors of the PI3K–AKT pathway, thereby promoting AKT signalling. By contrast, miR-203, upregulated during differentiation, targets genes that encode proteins that promote cell cycle progression (SKP2, p63, MSI2 and VAV3) thereby inducing cell cycle exit. miRNAs highly expressed in stem cells and differentiated progeny are outlined in orange and red, respectively.
Distinct miRNAs regulate the expansion and differentiation of epidermal and hair follicle stem cells. For example, miR-125, which is normally most highly expressed in hair follicle bulge cells and in basal cells of the epidermis, promotes the stemness of these populations59. Overexpression of miR-125 in the skin of mice increases proliferation of the hair bulge stem cells and ORS progenitors, and blocks their differentiation. Similarly, the loss of miR-205 results in a thinner epidermis and a shorter hair follicle, owing to the reduced proliferation of the stem cell populations, as assayed by Ki67 staining and BrdU incorporation58. miR-205 inhibits SHIP2 (also known as INPPL1), a negative regulator of AKT signalling, in mouse keratinocyte cultures65. The dampening of AKT signalling results in increased apoptosis. More careful examination has revealed that miR-205 actually targets multiple negative regulators of the PI3K–AKT pathway58. This later finding exemplifies how miRNAs often function by targeting a signalling pathway at multiple levels.
In contrast to miR-125 and miR-205, miRNAs such as miR-203 have active roles in terminally differentiating skin. In mice, the level of miR-203 rises dramatically during the differentiation of skin stem cells and miR-203 remains highly expressed in terminally differentiated cells60. The ectopic expression of miR-203 in primary human and mouse keratinocytes leads to reduced proliferation and inhibits colony formation60,66. Similar to miR-205, miR-203 functions through multiple targets60. In cells overexpressing miR-203, the forced expression of the target S phase kinase-associated protein 2 (SKP2) but not p63, MSI2 (also known as Musashi 2) or VAV3 blocked the ability of miR-203 to decrease the number of cells in the S phase of the cell cycle. However, all four targets were able to partially rescue the loss of colony formation induced by miR-203 overexpression. Together, these studies in mice exemplify how distinct miRNAs have opposing effects on self-renewal, proliferation and differentiation, thereby providing a bistable switch in cell fate choice. In a tissue such as the skin, which undergoes constant turnover, such regulatory switches are especially important.
miRNAs in neural development
The neural lineage presents an example of how stem and progenitor cells change in their developmental potential over time. The three main cell types of the nervous system — neurons, astrocytes and oligodendrocytes — are derived from progenitors in the neuroepithelium, which is specified by embryonic day 8.5 (E8.5) in mice (FIG. 2d). Neural progenitors undergo sequential differentiation into neurons and glia (astrocytes and oligodendrocytes) during development.
Regulation of miRNA levels
Several self-reinforcing feedback loops regulating miRNA expression have been uncovered during murine neural stem cell (NSC) differentiation (FIG. 6a). REST-SCP1, a protein complex that is known to silence neuronal genes in non-neuronal cells, represses the neuronal-specific miRNAs miR-9 and miR-124 in progenitors. However, miR-124 feeds back to the REST complex by inhibiting SCP1, thus suppressing its activity during neuronal differentiation67. The levels of another REST component, CTDSP2, are controlled by a negative feedback loop with miR-26. miR-26b is produced from one of the introns of CTDSP2 and targets the 3′ UTR of CTDSP2, forming a negative feedback loop. In addition, miR-26b is post-transcriptionally upregulated during neural differentiation, further silencing remaining REST activity68.
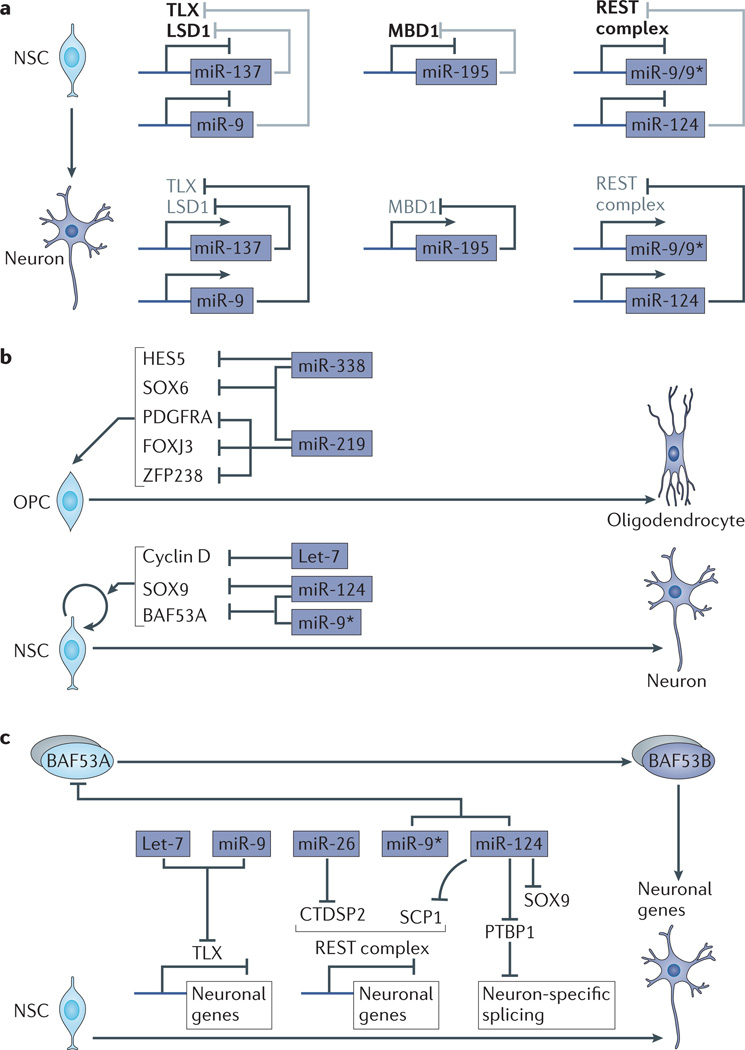
Figure 6. miRNAs regulate the development of neural cells.
a | microRNAs (miRNAs) that promote neurogenesis are often integrated into positive transcriptional and epigenetic feedback loops. The transcription factor TLX recruits the histone H3K4 demethylase LSD1 to inhibit miR-9 and miR-137 in neural stem cells (NSCs). During differentiation, miR-9 and miR-137 levels increase and target TLX and LSD1, respectively. Similarly, the methylated DNA binding protein MBD1 inhibits miR-195 in NSCs. During differentiation, miR-195 levels increase and this represses MBD1. Additionally, the REST complex, which represses neuronal gene expression, is highly expressed in NSCs and represses miR-9 and miR-124. During differentiation, REST levels decrease and miR-9 and miR-124 levels increase. miR-9 and miR-124 target components of the REST complex during differentiation. b | miRNAs enriched in oligodendrocytes (dark blue) promote oligodendrocyte differentiation by targeting transcription factors that are highly expressed in progenitors (HES5, SOX6, FOXJ3 and ZFP238) and signalling pathway components (platelet-derived growth factor receptor A (PDGFRA)). miRNAs upregulated during neuronal differentiation (let-7, miR-124 and miR-9*) (dark blue) promote cell cycle exit by inhibiting the cell cycle regulator cyclin D, chromatin remodeller BAF53A and transcription factors (such as SOX9) that are important for the progenitor state. c | miRNAs upregulated during neuronal differentiation (dark blue) promote derepression of neuronal gene expression (miR-26 and miR-124 targeting the REST complex, and miR-9/let-7 targeting TLX) and neuron-specific splicing (miR-124 targeting splicing regulator polypyrimidine tract-binding protein 1 (PTBP1)). miR-9* and miR-124 also promote the formation of a neuron-specific chromatin remodelling complex by inhibiting a progenitor-specific subunit (miR-9* and miR-124 targeting BAF53A) of the complex. OPC, oligodendrocyte progenitor cell.
Another example of a feedback loop involves the transcription factor TLX (also known as NR2E1). TLX represses miR-9 and miR-137 in NSCs, while miR-9 feeds back to TLX to inhibit its translation69,70. Interestingly, TLX suppresses miR-137 in part through the recruitment of the histone demethylase LSD1, which is a target of miR-137, forming an additional feedback loop. Similarly, in mouse adult NSCs, the DNA methyl-binding protein MBD1 represses the differentiation-promoting miRNAs miR-184 and miR-195. miR-195 targets MBD1 (REFS 71,72). The above examples demonstrate how miRNA expression during differentiation can be controlled by both positive and negative feedback loops involving transcriptional and epigenetic factors.
miRNA function
The first phenotype seen following conditional loss of Dicer in neurogenic compartments in the mouse cortex is a defect in neuronal differentiation, as evidenced by a decrease in the thickness of the neuronal layer73. At this stage, the progenitor compartments do not exhibit proliferation and apoptosis phenotypes probably owing to the persistence of miRNAs following Dicer deletion74. However, at later stages, increased apoptosis is a predominant phenotype in the progenitor compartments. Consistent with this observation, profiling of neural stem cells derived from Dicer-knockout mouse brains shows the dysregulation of apoptotic and survival genes74. Unlike the haematopoietic lineages, however, apoptosis has not been linked to direct targeting of apoptotic genes by expressed miRNAs. An alternative possibility is that apoptosis is a secondary effect of the widespread transcriptomic and proteomic dysregulation that occurs in the absence of miRNAs.
The nervous system provides excellent examples of how miRNAs can concomitantly drive cell cycle exit and silence the stem and progenitor cell fate (FIG. 6b). During oligodendrocyte differentiation, miR-219 and miR-338 increase and target the transcription factors SOX6 and HES5 that have been associated with progenitor proliferation and stemness75,76. During differentiation of adult NSCs, miR-124 promotes the transition of neurogenic precursors to the neuroblast stage by inhibiting the transcription factor SOX9 (REF. 77). Also in adult mouse NSCs, let-7 targets cyclin D1, and overexpression of let-7 in fetal neurogenic regions — where let-7 is not normally expressed — leads to increased cell cycle exit and differentiation78. Surprisingly, another study using human embryonic stem cell (hESC)-derived neural progenitors suggested that let-7 suppresses differentiation79. Additional studies will be needed to resolve these differences. Consistent with the important role of miRNAs in coordinating cell cycle exit during differentiation, the loss of miR-9 in neurogenic regions of the hindbrain leads to an increase in the number of proliferative cells, partially due to the derepression of a miR-9 target Hairy1 (also known as HES1), and the subsequent downregulation of the Hairy1 target p27, a cell cycle inhibitor80.
During differentiation, cell cycle exit is accompanied by the activation of a neuron-specific transcriptional programme (FIG. 6c). During this process, miRNAs are vital to enable the functional and morphological specialization of the differentiating neurons. This role is partially achieved by miRNAs targeting epigenetic factors and transcriptional repressors. For example, miR-9* and miR-124 target the BAF53A subunit of the SWI/SNF chromatin remodelling complex during neuronal differentiation81. BAF53A is required for the proliferation of NPCs, but when downregulated during differentiation, is replaced by the neuronal BAF53B subunit that drives expression of neuron-specific genes82. In addition, both let-7 and miR-9 target TLX, a repressor of neuronal genes, further promoting the activation of the neuronal gene expression programme during differentiation69,78. miRNAs also regulate alternative splicing during neuronal differentiation. miR-124 promotes the switch in alternative splicing patterns during differentiation by targeting polypyrimidine tract-binding protein 1 (PTBP1) mRNA, a repressor of neuron-specific pre-mRNA splicing83. Thus, miRNAs are situated in regulatory circuits whereby repression of the progenitor chromatin and splicing modulators can lead to activation of neuronal gene expression.
Recently, several studies have highlighted the roles that miRNAs have in regulating developmental timing during neural differentiation. Similar to the fetal to adult transition in haematopoiesis, increasing levels of the miRNA let-7 are partially responsible for the reduction in self-renewal capacity of neural stem cells in adult mice relative to fetal mice. In fetal neural stem and progenitor cells, LIN28A expression levels are high, suppressing let-7 biogenesis. Maintaining low let-7 expression enables increased levels of two important targets, the RBP IMP1 (also known as IGF2BP1 and IF2B1) and the chromatin factor HMGA2 (REF. 84). IMP1 suppresses the translation of a number of genes increased during neural differentiation and simultaneously promotes expression of HMGA2 and CCND2. HMGA2 suppresses the expression of the cell cycle inhibitors p16 and p19 thereby promoting cell division. Interestingly, in adult NSCs, IMP1 is no longer expressed, but HMGA2 is expressed. As adults age, let-7 increasingly suppresses HMGA2 thereby further diminishing adult NSC self-renewal as the mice get older85. Another miRNA, miR-125, which is often co-expressed with let-7 regulates the differentiation of adult, but not fetal interneurons of the olfactory bulb86. In particular, miR-125 regulates dendritic morphogenesis and synapse formation. It is quite remarkable how the two orthologues of the original heterochronic miRNAs let-7 and lin-4 (miR-125) found in Caenorhabditis elegans are used over and over again in evolution and across cell types to regulate cell cycle and differentiation87.
Although the miRNAs let-7 and miR-125 have been implicated in the suppression of neurogenesis, miR-17 and miR-106 seem to have the opposite effect88. In in vitro assays using mouse ESC-derived neurospheres, the overexpression of miR-17 and miR-106 miRNAs promotes the expansion of the neuronal progenitor cells while suppressing the transition to a gliogenic fate. They are thought to function at least in part by inhibiting p38, which leads to precocious expression of the astrocyte marker glial fibrillary acidic protein (GFAP) when overexpressed in vivo. Therefore, once again, it seems that distinct miRNA families are playing opposing parts in the regulation and stabilization of cell fate transitions.
Conclusion and perspectives
In this Review, we have provided examples of how miRNAs constitute an essential part of the gene networks that drive proliferation, viability, and differentiation of stem and progenitor cells across different somatic lineages. Upstream of miRNAs is a network of cell type-specific transcription factors that function together with epigenetic regulators to tightly regulate miRNA levels spatially and temporally. miRNA levels are further fine-tuned through post-transcriptional mechanisms that regulate their processing to the mature form, as well as their stability. Downstream of miRNAs are large networks of mRNA targets that influence cell fate choice. The ultimate effect of miRNAs on those targets is influenced by several factors. First, the number of miRNA binding sites within each target, which can be regulated by APA. Second, whether other miRNAs target the same transcript. Moreover, RBPs may bind along with miRNAs either synergizing or antagonizing the activity of the associated miRNAs. Third, multiple RNAs may be competing for the same miRNA, raising the possibility that a small number of these transcripts titrate away the miRNAs from other potential targets. The complexity of miRNA biology attests to the important regulatory roles of this class of small RNAs.
Surprisingly, knockouts of individual miRNAs often produce subtle or indiscernible phenotypes under homeostatic conditions8. Therefore, it has been proposed that miRNAs mostly exist to buffer fluctuations in gene expression, especially when under the influence of external stressors9. Such a model fits well with an evolutionary argument of miRNAs arising to provide robustness of cell fate stability essential for multicellularity. Interestingly, in contrast to miRNA knockout, miRNA overexpression often leads to specific and easily detectable phenotypes. Indeed, overexpression or misexpression of miRNAs can promote remarkable alterations in cell fate, including dedifferentiation of somatic cells to induced pluripotent stem cells, or transdifferentiation across somatic cell lineages such as fibroblasts to neurons and cardiomyocytes89–93. Such transitions require the precise silencing of one programme and the activation of a new one. Overexpression of miRNAs probably coordinates a complex set of targets to achieve a specific outcome. Therefore, dissecting the factors upstream and downstream of miRNAs should continue to provide fundamental insights into the networks of genes that regulate cell fate decisions.
Despite the large body of work described here, many unanswered questions remain. The miRNA repertoire of a cell consists of hundreds of miRNAs and the roles of only a handful have been elucidated. What are the roles of the lowly expressed miRNAs? What are the roles of other broadly expressed miRNAs? Additionally, how do miRNAs find their targets? How are the levels of miRNAs and argonaute proteins regulated? How are miRNAs degraded or turned over? In cases where there is not a cell type-specific miRNA, such as miR-124, does combinatorial expression of different miRNAs further define cellular identity? How are the miRNA-target gene networks perturbed in disease and/ or stress states?
Acknowledgements
The authors would like to thank members of the Blelloch laboratory, especially A. Chen, J. Freimer, R. Krishnakumar and M. Shveygert for helpful discussions and comments. This work was funded by a CIRM training grant TG2-01153 (A.S.) and grants to R.H.B. from the US National Institutes of Health (R01 NS057221 and GM101180) and the California Institute of Regenerative Medicine (New Faculty Award, RN2-00906-1).
Glossary
- Argonaute proteins
A family of proteins that function as effectors of microRNA (miRNA) function. Along with other proteins, such as GW182, they form a complex with miRNAs and bind target mRNAs to promote translational inhibition or mRNA degradation
- Mitophagy
A form of macroautophagy that degrades mitochondria. Autophagosomes enclose whole mitochondria and fuse with the lysosome, leading to degradation
Footnotes
Competing interests statement
The authors declare no competing interests.
References
- 1.Ha M, Kim VN. Regulation of microRNA biogenesis. Nature Rev. GMo. Cell Biol. 2014;15:509–524. doi: 10.1038/nrm3838. [DOI] [PubMed] [Google Scholar]
- 2.Babiarz JE, Blelloch R. Small RNAs – their biogenesis, regulation and function in embryonic stem cells. StemBook. 2009 [PubMed] [Google Scholar]
- 3.Bernstein E, et al. Dicer is essential for mouse development. Nature Genet. 2003;35:215–217. doi: 10.1038/ng1253. [DOI] [PubMed] [Google Scholar]
- 4.Wang Y, Medvid R, Melton C, Jaenisch R, Blelloch R. DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nature Genet. 2007;39:380–385. doi: 10.1038/ng1969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Huntzinger E, Izaurralde E. Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nature Rev. Genet. 2011;12:99–110. doi: 10.1038/nrg2936. [DOI] [PubMed] [Google Scholar]
- 6.Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136:215–233. doi: 10.1016/j.cell.2009.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Judson RL, Greve TS, Parchem RJ, Blelloch R. MicroRNA-based discovery of barriers to dedifferentiation of fibroblasts to pluripotent stem cells. Nature Struct. Mol. Biol. 2013;20:1227–1235. doi: 10.1038/nsmb.2665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Park CY, et al. A resource for the conditional ablation of microRNAs in the mouse. Cell Rep. 2012;1:385–391. doi: 10.1016/j.celrep.2012.02.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ebert MS, Sharp PA. Roles for microRNAs in conferring robustness to biological processes. Cell. 2012;149:515–524. doi: 10.1016/j.cell.2012.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Thornton JE, Gregory RI. How does Lin28 let-7 control development and disease? Trends Cell Biol. 2012;22:474–482. doi: 10.1016/j.tcb.2012.06.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Piskounova E, et al. Lin28A and Lin28B inhibit let-7 microRNA biogenesis by distinct mechanisms. Cell. 2011;147:1066–1079. doi: 10.1016/j.cell.2011.10.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Buckingham M, et al. The formation of skeletal muscle: from somite to limb. J. Anat. 2003;202:59–68. doi: 10.1046/j.1469-7580.2003.00139.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yaffe D, Saxel O. Serial passaging and differentiation of myogenic cells isolated from dystrophic mouse muscle. Nature. 1977;270:725–727. doi: 10.1038/270725a0. [DOI] [PubMed] [Google Scholar]
- 14.Rosenberg MI, Georges SA, Asawachaicharn A, Analau E, Tapscott SJ. MyoD inhibits Fstl1 and Utrn expression by inducing transcription of miR-206. J. Cell Biol. 2006;175:77–85. doi: 10.1083/jcb.200603039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Rao PK, Kumar RM, Farkhondeh M, Baskerville S, Lodish HF. Myogenic factors that regulate expression of muscle-specific microRNAs. Proc. Natl Acad. Sci. USA. 2006;103:8721–8726. doi: 10.1073/pnas.0602831103. Provides examples of how master transcriptional regulators of a specific cell fate act upstream of tissue-specific miRNAs.
- 16.Sweetman D, et al. Specific requirements of MRFs for the expression of muscle specific microRNAs miR-1, miR-206 and miR-133. Dev. Biol. 2008;321:491–499. doi: 10.1016/j.ydbio.2008.06.019. [DOI] [PubMed] [Google Scholar]
- 17.Small EM, et al. Regulation of PI3-kinase/Akt signaling by muscle-enriched microRNA-486. Proc. Natl Acad. Sci. USA. 2010;107:4218–4223. doi: 10.1073/pnas.1000300107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Yeung F, Chung E, Guess MG, Bell ML, Leinwand LA. Myh7b/miR-499 gene expression is transcriptionally regulated by MRFs and Eos. Nucleic Acids Res. 2012;40:7303–7318. doi: 10.1093/nar/gks466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. van Rooij E, et al. A family of microRNAs encoded by myosin genes governs myosin expression and muscle performance. Dev. Cell. 2009;17:662–673. doi: 10.1016/j.devcel.2009.10.013. Study showing how two miRNAs cooperate to promote further maturation of muscle fibres into slow fibres versus fast fibres.
- 20.O’Rourke JR, et al. Essential role for Dicer during skeletal muscle development. Dev. Biol. 2007;311:359–368. doi: 10.1016/j.ydbio.2007.08.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Goljanek-Whysall K, et al. MicroRNA regulation of the paired-box transcription factor Pax3 confers robustness to developmental timing of myogenesis. Proc. Natl Acad. Sci. USA. 2011;108:11936–11941. doi: 10.1073/pnas.1105362108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Dey BK, Gagan J, Dutta A. miR-206 and −486 induce myoblast differentiation by downregulating Pax7. Mol. Cell. Biol. 2011;31:203–214. doi: 10.1128/MCB.01009-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Boutet SC, et al. Alternative polyadenylation mediates microRNA regulation of muscle stem cell function. Cell Stem Cell. 2012;10:327–336. doi: 10.1016/j.stem.2012.01.017. Provides a specific example of how miRNA activity can be controlled by APA in skeletal muscle subsets.
- 24.Chen JF, et al. microRNA-1 and microRNA-206 regulate skeletal muscle satellite cell proliferation and differentiation by repressing Pax7. J. Cell Biol. 2010;190:867–879. doi: 10.1083/jcb.200911036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Cardinali B, et al. Microrna-221 and microrna-222 modulate differentiation and maturation of skeletal muscle cells. PLoS ONE. 2009;4:e7607. doi: 10.1371/journal.pone.0007607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Sarkar S, Dey BK, Dutta A. MiR-322/424 and −503 are induced during muscle differentiation and promote cell cycle quiescence and differentiation by down-regulation of Cdc25A. Mol. Biol. Cell. 2010;21:2138–2149. doi: 10.1091/mbc.E10-01-0062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Chen J-F, et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nature Genet. 2005;38:228–233. doi: 10.1038/ng1725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Winbanks CE, et al. TGF-beta regulates miR-206 and miR-29 to control myogenic differentiation through regulation of HDAC4. J. Biol. Chem. 2011;286:13805–13814. doi: 10.1074/jbc.M110.192625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Juan AH, Kumar RM, Marx JG, Young RA, Sartorelli V. Mir-214-dependent regulation of the polycomb protein Ezh2 in skeletal muscle and embryonic stem cells. Mol. Cell. 2009;36:61–74. doi: 10.1016/j.molcel.2009.08.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Di Giammartino DC, Nishida K, Manley JL. Mechanisms and consequences of alternative polyadenylation. Mol. Cell. 2011;43:853–866. doi: 10.1016/j.molcel.2011.08.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Dormoy-Raclet V, et al. HuR and miR-1192 regulate myogenesis by modulating the translation of HMGB1 mRNA mRNA. Nat Comms. 2013;4 doi: 10.1038/ncomms3388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Bak RO, Mikkelsen JG. miRNA sponges: soaking up miRNAs for regulation of gene expression. WIREs RNA. 2014;5:317–333. doi: 10.1002/wrna.1213. [DOI] [PubMed] [Google Scholar]
- 33.Cesana M, et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell. 2011;147:358–369. doi: 10.1016/j.cell.2011.09.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kallen AN, et al. The imprinted H19 LncRNA antagonizes Let-7 microRNAs. Mol. Cell. 2013;52:101–112. doi: 10.1016/j.molcel.2013.08.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Orkin SH. Diversification of haematopoietic stem cells to specific lineages. Nature Rev. Genet. 2000;1:57–64. doi: 10.1038/35049577. [DOI] [PubMed] [Google Scholar]
- 36.Dore LC, et al. A GATA-1-regulated microRNA locus essential for erythropoiesis. Proc. Natl Acad. Sci. USA. 2008;105:3333–3338. doi: 10.1073/pnas.0712312105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zhu Y, et al. A comprehensive analysis of GATA-1-regulated miRNAs reveals miR-23a to be a positive modulator of erythropoiesis. Nucleic Acids Res. 2013;41:4129–4143. doi: 10.1093/nar/gkt093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Barde I, et al. A KRAB/KAP1-miRNA cascade regulates erythropoiesis through stage-specific control of mitophagy. Science. 2013;340:350–353. doi: 10.1126/science.1232398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Crichton JH, Dunican DS, MacLennan M, Meehan RR, Adams IR. Defending the genome from the enemy within: mechanisms of retrotransposon suppression in the mouse germline. Cell. Mol. Life Sci. 2014;71:1581–1605. doi: 10.1007/s00018-013-1468-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Fazi F, et al. A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPα regulates human granulopoiesis. Cell. 2005;123:819–831. doi: 10.1016/j.cell.2005.09.023. Study showing how the interplay between an miRNA and two transcription factors can feed back to modulate levels of expression of the miRNA.
- 41.Copley MR, et al. The Lin28b-let-7-Hmga2 axis determines the higher self-renewal potential of fetal hematopoietic stem cells. Nature Cell Biol. 2013;15:916–925. doi: 10.1038/ncb2783. [DOI] [PubMed] [Google Scholar]
- 42.Emmrich S, et al. miR-99a/100~125b tricistrons regulate hematopoietic stem and progenitor cell homeostasis by shifting the balance between TGFβ and Wnt signaling. Genes Dev. 2014;28:858–874. doi: 10.1101/gad.233791.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Yuan J, Nguyen CK, Liu X, Kanellopoulou C, Muljo SA. Lin28b reprograms adult bone marrow hematopoietic progenitors to mediate fetal-like lymphopoiesis. Science. 2012;335:1195–1200. doi: 10.1126/science.1216557. Demonstrates that let-7 and LIN28 control the transition between fetal and adult haematopoiesis, a theme that is also seen in neurogenesis, as described in this Review.
- 44.Senyuk V, et al. Critical role of miR-9 in myelopoiesis and EVI1-induced leukemogenesis. Proc. Natl Acad. Sci. USA. 2013;110:5594–5599. doi: 10.1073/pnas.1302645110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Lu J, et al. MicroRNA-mediated control of cell fate in megakaryocyte-erythrocyte progenitors. Dev. Cell. 2008;14:843–853. doi: 10.1016/j.devcel.2008.03.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Guo S, et al. MicroRNA miR-125a controls hematopoietic stem cell number. Proc. Natl Acad. Sci. USA. 2010;107:14229–14234. doi: 10.1073/pnas.0913574107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.O’Connell RM, et al. MicroRNAs enriched in hematopoietic stem cells differentially regulate long-term hematopoietic output. Proc. Natl Acad. Sci. USA. 2010;107:14235–14240. doi: 10.1073/pnas.1009798107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Ooi AGL, et al. MicroRNA-125b expands hematopoietic stem cells and enriches for the lymphoid-balanced and lymphoid-biased subsets. Proc. Natl Acad. Sci. USA. 2010;107:21505–21510. doi: 10.1073/pnas.1016218107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Koralov SB, et al. Dicer ablation affects antibody diversity and cell survival in the B lymphocyte lineage. Cell. 2008;132:860–874. doi: 10.1016/j.cell.2008.02.020. [DOI] [PubMed] [Google Scholar]
- 50.Johnnidis JB, et al. Regulation of progenitor cell proliferation and granulocyte function by microRNA-223. Nature. 2008;451:1125–1129. doi: 10.1038/nature06607. [DOI] [PubMed] [Google Scholar]
- 51.Patrick DM, et al. Defective erythroid differentiation in miR-451 mutant mice mediated by 14-3-3zeta. Genes Dev. 2010;24:1614–1619. doi: 10.1101/gad.1942810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Cifuentes D, et al. A novel miRNA processing pathway independent of Dice. Science. 2010;328:1694–1698. doi: 10.1126/science.1190809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Cheloufi S, Santos , Dos CO, Chong MMW, Hannon GJ. A dicer-independent miRNA biogenesis pathway that requires Ago catalysis. Nature. 2010;465:584–589. doi: 10.1038/nature09092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Rasmussen KD, et al. The miR-144/451 locus is required for erythroid homeostasis. J. Exp. Med. 2010;207:1351–1358. doi: 10.1084/jem.20100458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Sturgeon CM, et al. Primitive erythropoiesis is regulated by miR-126 via nonhematopoietic Vcam-1+ cells. Dev. Cell. 2012;23:45–57. doi: 10.1016/j.devcel.2012.05.021. Example of miRNAs functioning in the niche cells indirectly affecting primitive HSC development.
- 56.Raaijmakers MHGP, et al. Bone progenitor dysfunction induces myelodysplasia and secondary leukaemia. Nature. 2010;464:852–857. doi: 10.1038/nature08851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Fuchs E. Scratching the surface of skin development. Nature. 2007;445:834–842. doi: 10.1038/nature05659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Wang D, et al. MicroRNA-205 controls neonatal expansion of skin stem cells by modulating the PI(3)K pathway. Nature Cell Biol. 2013;15:1153–1163. doi: 10.1038/ncb2827. Example of a single miRNA targeting multiple components of a signalling pathway synergizing to promote skin stem cell proliferation.
- 59.Zhang L, Stokes N, Polak L, Fuchs E. Specific microRNAs are preferentially expressed by skin stem cells to balance self-renewal and early lineage commitment. Cell Stem Cell. 2011;8:294–308. doi: 10.1016/j.stem.2011.01.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Jackson SJ, et al. Rapid and widespread suppression of self-renewal by microRNA-203 during epidermal differentiation. Development. 2013;140:1882–1891. doi: 10.1242/dev.089649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Antonini D, et al. Transcriptional repression of miR-34 family contributes to p63-mediated cell cycle progression in epidermal cells. J. Investig. Dermatol. 2010;130:1249–1257. doi: 10.1038/jid.2009.438. [DOI] [PubMed] [Google Scholar]
- 62.Andl T, et al. The miRNA-processing enzyme dicer is essential for the morphogenesis and maintenance of hair follicles. Curr. Biol. 2006;16:1041–1049. doi: 10.1016/j.cub.2006.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Yi R, et al. Morphogenesis in skin is governed by discrete sets of differentially expressed microRNAs. Nature Genet. 2006;38:356–362. doi: 10.1038/ng1744. [DOI] [PubMed] [Google Scholar]
- 64.Teta M, et al. Inducible deletion of epidermal Dicer and Drosha reveals multiple functions for miRNAs in postnatal skin. Development. 2012;139:1405–1416. doi: 10.1242/dev.070920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Yu J, et al. MicroRNA-184 antagonizes microRNA-205 to maintain SHIP2 levels in epithelia. Proc. Natl Acad. Sci. USA. 2008;105:19300–19305. doi: 10.1073/pnas.0803992105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Lena AM, et al. miR-203 represses ‘stemness’ by repressing DeltaNp63. Cell Death Differ. 2008;15:1187–1195. doi: 10.1038/cdd.2008.69. [DOI] [PubMed] [Google Scholar]
- 67.Visvanathan J, Lee S, Lee B, Lee JW, Lee S-K. The microRNA miR-124 antagonizes the anti-neural REST/SCP1 pathway during embryonic CNS development. Genes Dev. 2007;21:744–749. doi: 10.1101/gad.1519107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Dill H, Linder B, Fehr A, Fischer U. Intronic miR-26b controls neuronal differentiation by repressing its host transcript, ctdsp2. Genes Dev. 2012;26:25–30. doi: 10.1101/gad.177774.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Zhao C, Sun G, Li S, Shi Y. A feedback regulatory loop involving microRNA-9 and nuclear receptor TLX in neural stem cell fate determination. Nature Struct. Mol. Biol. 2009;16:365–371. doi: 10.1038/nsmb.1576. Example of a positive feedback loop that leads to a switch from TLX repression of miR-9 in progenitors and miR-9 repression of TLX with differentiation.
- 70.Sun G, et al. miR-137 forms a regulatory loop with nuclear receptor TLX and LSD1 in neural stem cells. Nat. Comms. 2011;2:529. doi: 10.1038/ncomms1532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Liu C, et al. Epigenetic Regulation of miR-184 by MBD1 governs neural stem cell proliferation and differentiation. Cell Stem Cell. 2010;6:433–444. doi: 10.1016/j.stem.2010.02.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Liu C, et al. An epigenetic feedback regulatory loop involving microRNA-195 and MBD1 governs neural stem cell differentiation. PLoS ONE. 2013;8:e51436. doi: 10.1371/journal.pone.0051436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.De Pietri Tonelli D, et al. miRNAs are essential for survival and differentiation of newborn neurons but not for expansion of neural progenitors during early neurogenesis in the mouse embryonic neocortex. Development. 2008;135:3911–3921. doi: 10.1242/dev.025080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Kawase-Koga Y, Otaegi G, Sun T. Different timings of Dicer deletion affect neurogenesis and gliogenesis in the developing mouse central nervous system. Dev. Dyn. 2009;238:2800–2812. doi: 10.1002/dvdy.22109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Dugas JC, et al. Dicer1 and miR-219 Are required for normal oligodendrocyte differentiation and myelination. Neuron. 2010;65:597–611. doi: 10.1016/j.neuron.2010.01.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Zhao X, et al. MicroRNA-mediated control of oligodendrocyte differentiation. Neuron. 2010;65:612–626. doi: 10.1016/j.neuron.2010.02.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Cheng L-C, Pastrana E, Tavazoie M, Doetsch F. miR-124 regulates adult neurogenesis in the subventricular zone stem cell niche. Nature Neurosci. 2009;12:399–408. doi: 10.1038/nn.2294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Zhao C, et al. MicroRNA let-7b regulates neural stem cell proliferation and differentiation by targeting nuclear receptor TLX signaling. Proc. Natl Acad. Sci. USA. 2010;107:1876–1881. doi: 10.1073/pnas.0908750107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Cimadamore F, Amador-Arjona A, Chen C, Huang CT, Terskikh AV. SOX2-LIN28/let-7 pathway regulates proliferation and neurogenesis in neural precursors. Proc. Natl Acad. Sci. USA. 2013;110:E3017–E3026. doi: 10.1073/pnas.1220176110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Bonev B, Pisco A, Papalopulu N. MicroRNA-9 reveals regional diversity of neural progenitors along the anterior-posterior axis. Dev. Cell. 2011;20:19–32. doi: 10.1016/j.devcel.2010.11.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81. Yoo AS, Staahl BT, Chen L, Crabtree GR. MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature. 2009;460:642–646. doi: 10.1038/nature08139. Demonstrates how miRNAs can drive activation of the neuronal gene expression programme by repressing chromatin remodellers of the progenitor cells.
- 82.Lessard J, et al. An essential switch in subunit composition of a chromatin remodeling complex during neural development. Neuron. 2007;55:201–215. doi: 10.1016/j.neuron.2007.06.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Makeyev EV, Zhang J, Carrasco MA, Maniatis T. The microRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell. 2007;27:435–448. doi: 10.1016/j.molcel.2007.07.015. Demonstrates how miRNAs can affect global cell type-specific splicing by inhibiting a splicing regulator.
- 84.Nishino J, et al. A network of heterochronic genes including Imp1 regulates temporal changes in stem cell properties. eLife. 2013;2:e00924–e00924. doi: 10.7554/eLife.00924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Nishino J, Kim I, Chada K, Morrison SJ. Hmga2 promotes neural stem cell self-renewal in young but not old mice by reducing p16Ink4a and p19Arf expression. Cell. 2008;135:227–239. doi: 10.1016/j.cell.2008.09.01. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Åkerblom M, et al. microRNA-125 distinguishes developmentally generated and adult-born olfactory bulb interneurons. Development. 2014;141:1580–1588. doi: 10.1242/dev.101659. [DOI] [PubMed] [Google Scholar]
- 87.Reinhart BJ, et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature. 2000;403:901–906. doi: 10.1038/35002607. [DOI] [PubMed] [Google Scholar]
- 88.Naka-Kaneda H, et al. The miR-17/106-p38 axis is a key regulator of the neurogenic-to-gliogenic transition in developing neural stem/progenitor cells. Proc. Natl Acad. Sci. USA. 2014;111:1604–1609. doi: 10.1073/pnas.1315567111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Judson RL, Babiarz JE, Venere M, Blelloch R. Embryonic stem cell-specific microRNAs promote induced pluripotency. Nature Biotech. 2009;27:459–461. doi: 10.1038/nbt.1535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Subramanyam D, et al. Multiple targets of miR-302 and miR-372 promote reprogramming of human fibroblasts to induced pluripotent stem cells. Nature Biotech. 2011;29:443–448. doi: 10.1038/nbt.1862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Anokye-Danso F, et al. Highly efficient miRNA-mediated reprogramming of mouse and human somatic cells to pluripotency. Cell Stem Cell. 2011;8:376–388. doi: 10.1016/j.stem.2011.03.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Yoo AS, et al. MicroRNA-mediated conversion of human fibroblasts to neurons. Nature. 2011;476:228–231. doi: 10.1038/nature10323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Nam Y-J, et al. Reprogramming of human fibroblasts toward a cardiac fate. Proc. Natl Acad. Sci. USA. 2013;110:5588–5593. doi: 10.1073/pnas.1301019110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Harfe BD, McManus MT, Mansfield JH, Hornstein E, Tabin CJ. The RNaseIII enzyme Dicer is required for morphogenesis but not patterning of the vertebrate limb. Proc. Natl Acad. Sci. USA. 2005;102:10898–10903. doi: 10.1073/pnas.0504834102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Lynn FC, et al. MicroRNA expression is required for pancreatic islet cell genesis in the mouse. Diabetes. 2007;56:2938–2945. doi: 10.2337/db07-0175. [DOI] [PubMed] [Google Scholar]
- 96.Gaur T, et al. Dicer inactivation in osteoprogenitor cells compromises fetal survival and bone formation, while excision in differentiated osteoblasts increases bone mass in the adult mouse. Dev. Biol. 2010;340:10–21. doi: 10.1016/j.ydbio.2010.01.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Kobayashi T, et al. Dicer-dependent pathways regulate chondrocyte proliferation and differentiation. Proc. Natl Acad. Sci. USA. 2008;105:1949–1954. doi: 10.1073/pnas.0707900105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Mudhasani R, et al. Dicer is required for the formation of white but not brown adipose tissue. J. Cell. Physiol. 2011;226:1399–1406. doi: 10.1002/jcp.22475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Nagalakshmi VK, et al. Dicer regulates the development of nephrogenic and ureteric compartments in the mammalian kidney. Kidney Int. 2011;79:317–330. doi: 10.1038/ki.2010.385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Zehir A, Hua LL, Maska EL, Morikawa Y, Cserjesi P. Dicer is required for survival of differentiating neural crest cells. Dev. Biol. 2010;340:459–467. doi: 10.1016/j.ydbio.2010.01.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Michon F, Tummers M, Kyyrönen M, Frilander MJ, Thesleff I. Tooth morphogenesis and ameloblast differentiation are regulated by micro-RNAs. Dev. Biol. 2010;340:355–368. doi: 10.1016/j.ydbio.2010.01.019. [DOI] [PubMed] [Google Scholar]
- 102.Suh N, et al. MicroRNA function is globally suppressed in mouse oocytes and early embryos. Curr. Biol. 2010;20:271–277. doi: 10.1016/j.cub.2009.12.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Shenoy A, Blelloch R. microRNA induced transdifferentiation. F1000 Biol. Rep. 2012;4:2. doi: 10.3410/B4-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Ventura A, et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell. 2008;132:875–886. doi: 10.1016/j.cell.2008.02.019. [DOI] [PMC free article] [PubMed] [Google Scholar]