Abstract
Genome-wide association studies (GWAS) of psychiatric disorders have identified multiple genetic associations with such disorders, but better methods are needed to derive the underlying biological mechanisms that these signals indicate. We sought to identify biological pathways in GWAS data from over 60,000 participants from the Psychiatric Genomics Consortium. We developed an analysis framework to rank pathways that requires only summary statistics. We combined this score across disorders to find common pathways across three adult psychiatric disorders: schizophrenia, major depression and bipolar disorder. Histone methylation processes showed the strongest association, and we also found statistically significant evidence for associations with multiple immune and neuronal signaling pathways and with the postsynaptic density. Our study indicates that risk variants for psychiatric disorders aggregate in particular biological pathways and that these pathways are frequently shared between disorders. Our results confirm known mechanisms and suggest several novel insights into the etiology of psychiatric disorders.
Psychiatric disorders account for a large proportion of global disease burden1. They are clinical syndromes with largely unknown etiology whose classification has been developed on the basis of their observable symptomatology and course of illness. However, there is considerable evidence for strong heritability of these disorders, and recent work by the Psychiatric Genomics Consortium (PGC) using genome-wide association study (GWAS) data has demonstrated that a considerable proportion of this heritability is attributable to common genetic variants2 and has also shown clear evidence of shared genetic risk at individual loci3.
To make further progress in the treatment and prevention of these disorders, there is an urgent need to clearly identify the biological mechanisms and pathways underlying risk. However, the analyses available to date have focused primarily on single disorders and on gene-expression approaches (for example, in schizophrenia4) and, although interesting, such approaches are subject to potential confounding by the downstream effects of disorders and their treatment. Genetic pathway analysis methods for GWAS data have been developed5,6 and aim to identify which biological pathways show an excess of etiological association. Though the power to detect pathway associations can be limited by lack of power in the original GWAS data, genome-wide ‘chip-heritability’ estimates3 demonstrate that the loci showing nominal significance, but with values below genome-wide significance cutoffs, contribute to a significant proportion of disease liability. Pathway analysis provides a way to separate the true signals among these loci from the noise. Furthermore, pathway analysis can translate GWAS signals into a level of understanding that is biochemical and/or system-wide and can provide successful replication in the presence of allelic and locus heterogeneity7,8.
In psychiatric genetics, several reports have found significant association with biological processes using GWAS. Analyses of bipolar disorder provided evidence of association with hormone action and adherens junctions7,9,10. Activity of voltage-gated calcium ions was also implicated in a pathway analysis of a bipolar disorder GWAS data set11. We hypothesized that combining pathway-based GWAS signals across multiple related disorders could be a powerful approach to identify pathways susceptible to genetic risk in neuropsychiatric disorders.
The PGC was established in 2007 (http://pgc.unc.edu)12 and has been conducting field-wide mega-analyses of genomic data for common and severe psychiatric disorders2,3,11,13–15. Summary data are now available for PGC phase 1 studies that comprise >60,000 study participants representing schizophrenia (SCZ), major depressive disorder (MDD), bipolar disorder (BIP), autism-spectrum disorder (ASD) and attention deficit–hyperactivity disorder (ADHD). We have previously reported cross-disorder analyses via a single-nucleotide polymorphism (SNP)/association-based approach3 and as estimated “chip-heritability” and genetic covariance via the genome-wide evidence2. We now extend this work, seeking to statistically identify the molecular pathways implicated by variants underlying genetic risk, the identification of which may have major impact on the understanding and future treatment of psychiatric disorders.
RESULTS
We summarize pathway data sets and provide gene membership in Supplementary Tables 1 and 2, respectively. From an initial compiled set of 19,752 pathways across five gene set databases (GO, KEGG, Panther, Reactome, TargetScan), we restricted downstream analyses to the 4,949 pathways of size 10–200 genes.
Comparisons among methods
We first obtained pathway-level P values for each pathway for the five disorders (SCZ, MDD, BIP, ASD and ADHD) across the five methods (SETSCREEN, MAGENTA, INRICH, FORGE and ALIGATOR). Overall, methods were significantly correlated with each other (see Supplementary Fig. 1 and Supplementary Table 3). Using both disorder data (SCZ) and null data (permuted phenotypes), we noted a statistically significant degree of overlap among methods (Supplementary Fig. 1).
Deriving a method-wise and disorder-wise joint statistic
Pathways that achieve strong association using all five methodologies would be expected to be more robustly associated with disorder, owing to the differences between methods. We estimated a combined P value for each pathway (within each disorder) by calculating the average rank of each pathway within each method (ranks were used to ensure comparability between methods) and then comparing to a null distribution of expected ranks, built by drawing from the uniform distribution, accounting for intermethod correlation (Fig. 1; see Online Methods).
Figure 1.
Overview of statistical approach for integrative pathway analysis of GWAS data. A summary of an analysis of one disorder is shown. Simulated data were generated by drawing from a null pathway P-value distribution for each method and for each disease that accounted for correlations between methods. Pathway results from all disorders were subsequently combined using Fisher’s method.
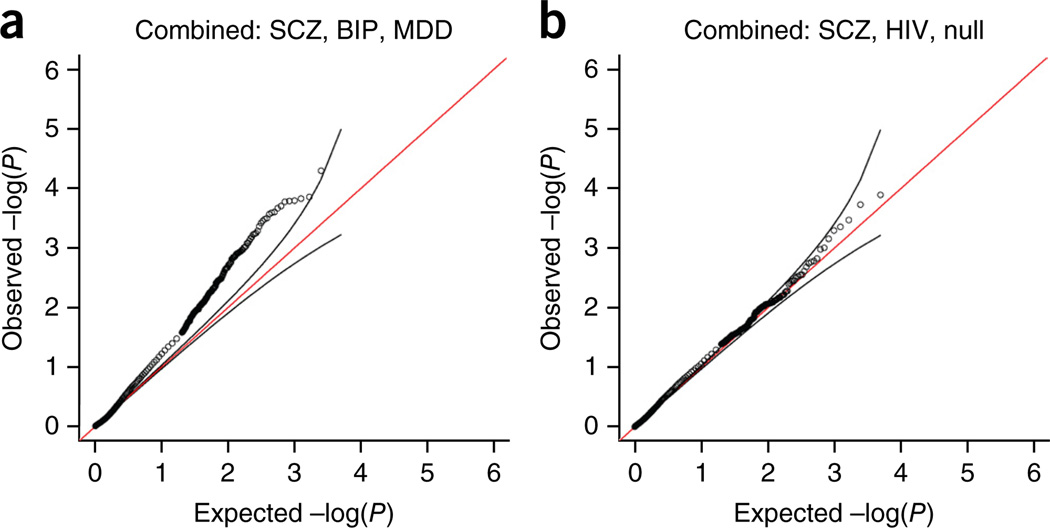
There was a significant degree of correlation of pathway-specific enrichment P values between disorders (Pearson correlation 0.2–0.3, Supplementary Table 4). Looking for pathways common to the adult disorders (SCZ, BIP, MDD), we then derived a combined statistic across disorders using Brown’s extension of Fisher’s combined P value16 (see Online Methods). Figure 2 shows quantile-quantile plots of P values from combining all methods across disorders. Supplementary Figure 2 shows quantile-quantile plots of combined P values for SCZ, BIP, MDD, ASD, ADHD and, for contrast, null and HIV acquisition data. The former plots show a marked enrichment of significant P values, indicative of shared disease biology captured by the pathways tested. The null and HIV plots shows no enrichment of P values, indicating both that there is a lack of shared biology (as expected) and that our analysis does not cause a systematic inflation of significance. Similarly, the HIV data set showed no inflation.
Figure 2.
Quantile-quantile plot showing P-value distribution for a combined analysis combining results from five pathway analysis methods and six pathway databases. (a,b) Data are shown for schizophrenia (SCZ), bipolar disorder (BIP) and major depressive disorder (MDD; a) and SCZ, HIV acquisition and a null simulated data set (b).
Top pathways in individual adult-onset psychiatric disorders
We present the results for the different psychiatric disorder data sets in Supplementary Table 5, obtained using our approach. We identified 10, 1 and 4 pathway(s) that are suggestively enriched (at an FDR q-value <0.1) with BIP, SCZ and MDD susceptibility alleles, respectively (Table 1). Note that the pathways showing enrichment may change as sample sizes increase.
Table 1.
Top pathways in schizophrenia, bipolar disorder and major depressive disordera
| No. Methods |
Avg. rank |
P rank | q-value | Pathway ID | Description |
|---|---|---|---|---|---|
| BIP | |||||
| 5 | 17 | 1.01 × 10−6 | 0.005 | GO:51568 | Histone H3-K4 methylation |
| 5 | 50.4 | 3.82 × 10−5 | 0.093 | path:hsa05218 | Melanoma |
| 5 | 79.2 | 1.16 × 10−4 | 0.093 | GO:7129 | (Chromosomal) synapsis |
| 5 | 81.8 | 1.27 × 10−4 | 0.093 | path:hsa05213 | Endometrial cancer |
| 5 | 83.3 | 1.34 × 10−4 | 0.093 | P00003 | Alzheimer disease–amyloid secretase pathway |
| 5 | 83.4 | 1.35 × 10−4 | 0.093 | path:hsa05215 | Prostate cancer |
| 5 | 87 | 1.50 × 10−4 | 0.093 | path:hsa05216 | Thyroid cancer |
| 4 | 89.5 | 1.59 × 10−4 | 0.093 | GO:90066 | Regulation of anatomical structure size |
| 5 | 95.6 | 1.81 × 10−4 | 0.093 | path:hsa05214 | Glioma |
| 5 | 96.9 | 1.87 × 10−4 | 0.093 | GO:70192 | Chromosome organization involved in meiosis |
| SCZ | |||||
| 5 | 38.4 | 1.58 × 10−5 | 0.078 | GO:14069 | Postsynaptic density |
| 5 | 68.6 | 7.15 × 10−5 | 0.160 | GO:45211 | Postsynaptic membrane |
| 5 | 76.8 | 9.67 × 10−5 | 0.160 | GO:43197 | Dendritic spine |
| 5 | 85.4 | 1.36 × 10−4 | 0.168 | GO:51568 | Histone H3-K4 methylation |
| 5 | 95.8 | 1.74 × 10−4 | 0.173 | GO:33267 | Axon part |
| MDD | |||||
| 5 | 25.4 | 2.63 × 10−6 | 0.012 | GO:8601 | Protein phosphatase type 2A regulator activity |
| 5 | 54.6 | 3.88 × 10−5 | 0.092 | GO:34330 | Cell junction organization |
| 5 | 68.8 | 7.70 × 10−5 | 0.094 | GO:43297 | Apical junction assembly |
| 5 | 70 | 7.92 × 10−5 | 0.094 | GO:45216 | Cell-cell junction organization |
| 5 | 99.8 | 1.97 × 10−4 | 0.186 | GO:31056 | Regulation of histone modification |
Top pathways with q < 0.1, for the Schizophrenia (SCZ), Bipolar Disorder (BIP) and Major Depressive Disorder (MDD) data sets. For disorders with fewer than five pathways with q < 0.1, the top five pathways are listed.
Top pathways shared across adult psychiatric disorders
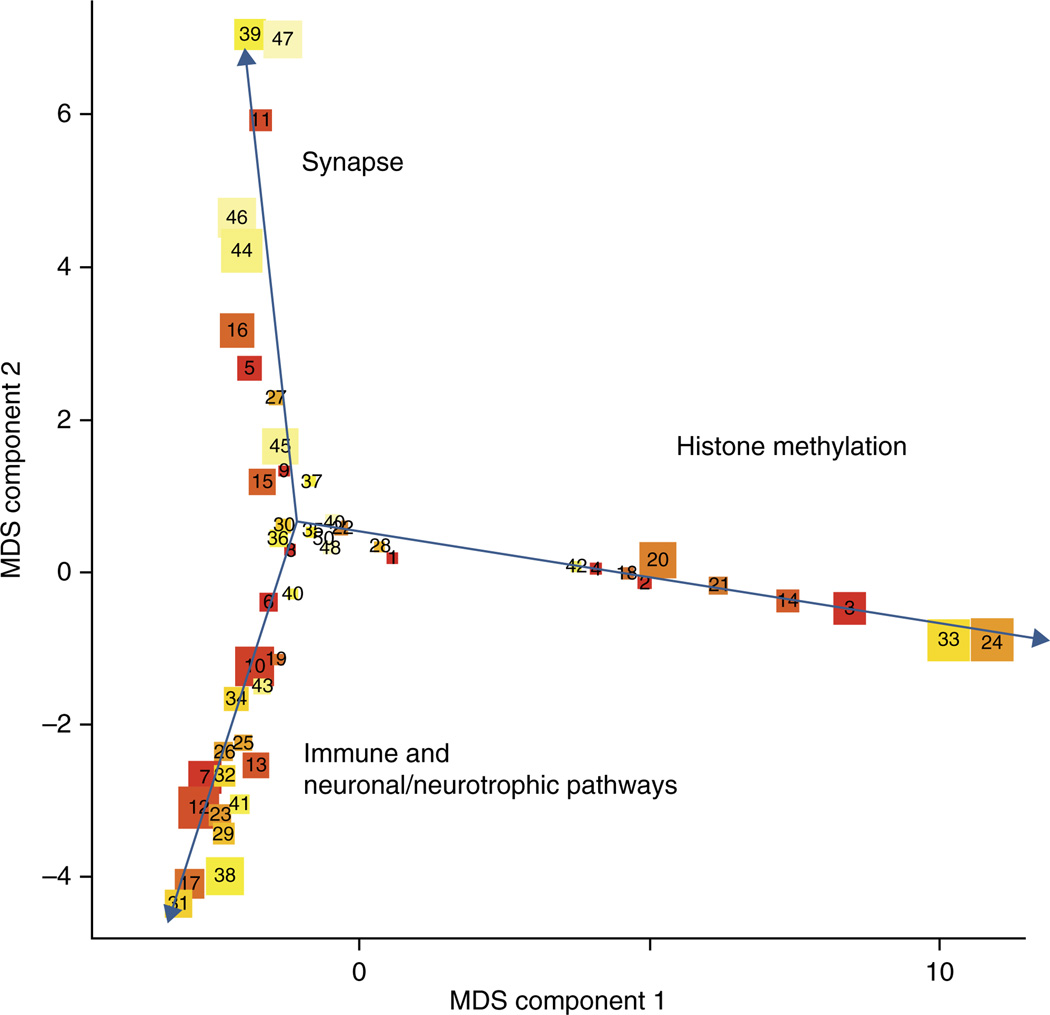
The degree of rank correlation between pathways across pairs of disorders was significant (Supplementary Tables 4 and 6), with 49 pathways with combined q-value <0.1 spanning the three adult disorders (Table 2, Supplementary Table 7). Of these, 16 are significant at q < 0.05, with the top pathway (GO:51568: histone H3-K4 methylation) having a q = 0.0003 (Table 2). These results are more significant than those observed in any of the disorders analyzed alone. We then use multidimensional scaling (MDS) to cluster these sets in terms of shared genes. Figure 3 shows a plot of every pathway with suggestive q < 0.1 on the first two MDS axes derived from the shared genes between these gene sets. The pathways separate in multidimensional space and reveal two distinct branches for neuronal synapse– and histone methylation– related gene sets, with a third branch containing pathways with genes that share membership in pathways with immune or neurotrophic functional annotation (Fig. 3). Although not a main focus of our analysis, the most significant pathway (GO:0005262, calcium channel activity) from the previous PGC Cross Disorder group SNP-based meta-analysis GWAS3 across the five disorders was found to show nominally significant association in our pathway level meta-analysis across all five disorders (P = 3.13 × 10−3) and also in SCZ, MDD and BIP (P = 8.07 × 10−3).
Table 2.
Top results from integrative pathway analysis of three adult disorders
| Rank | BIP | MDD | SCZ | Combined P | q-value | Pathway ID | Description |
|---|---|---|---|---|---|---|---|
| 1 | 0.0000 | 0.0592 | 0.0001 | 5.75 × 10−8 | 0.0003 | GO:51568 | Histone H3-K4 methylation |
| 2 | 0.0004 | 0.0500 | 0.0006 | 1.46 × 10−5 | 0.0362 | GO:16571 | Histone methylation |
| 3 | 0.0004 | 0.1462 | 0.0011 | 4.73 × 10−5 | 0.0414 | GO:43414 | Macromolecule methylation |
| 4 | 0.0008 | 0.0630 | 0.0014 | 5.10 × 10−5 | 0.0414 | GO:34968 | Histone lysine methylation |
| 5 | 0.4200 | 0.0001 | 0.0023 | 5.58 × 10−5 | 0.0414 | GO:45216 | Cell-cell junction organization |
| 6 | 0.0001 | 0.0910 | 0.0064 | 5.69 × 10−5 | 0.0414 | P00003 | Alzheimer disease–amyloid secretase pathway |
| 7 | 0.0007 | 0.0495 | 0.0024 | 5.86 × 10−5 | 0.0414 | P04393 | Ras pathway |
| 8 | 0.3120 | 0.0000 | 0.1286 | 7.12 × 10−5 | 0.0422 | GO:8601 | Protein phosphatase type 2A regulator activity |
| 9 | 0.8980 | 0.0001 | 0.0017 | 7.83 × 10−5 | 0.0422 | GO:43297 | Apical junction assembly |
| 10 | 0.0013 | 0.0207 | 0.0055 | 9.25 × 10−5 | 0.0422 | P00052 | TGF-β signaling pathway |
| 11 | 0.4890 | 0.0203 | 0.0000 | 9.53 × 10−5 | 0.0422 | GO:14069 | Postsynaptic density |
| 12 | 0.0085 | 0.0009 | 0.0239 | 0.0001 | 0.0422 | GO:32869 | Cellular response to insulin stimulus |
| 13 | 0.0188 | 0.0054 | 0.0022 | 0.0001 | 0.0450 | P00010 | B cell activation |
| 14 | 0.0023 | 0.2988 | 0.0003 | 0.0001 | 0.0450 | GO:8757 | S-adenosylmethionine–dependent methyltransferase activity |
| 15 | 0.0073 | 0.0080 | 0.0044 | 0.0001 | 0.0454 | GO:23061 | Signal release |
| 16 | 0.4590 | 0.0000 | 0.0168 | 0.0002 | 0.0473 | GO:34330 | Cell junction organization |
Shown are the top 16 pathways with combined q < 0.05 spanning the three adult disorders. Full results for the three adult disorders are given in Supplementary Table 7 and for the five disorders in Supplementary Table 8. Supplementary Table 10 lists all pathways with q < 0.1 and Supplementary Table 11 the underlying SNP P values in all genes with these gene sets.
Figure 3.
Multidimensional scaling plot of top 50 pathways with suggestive (<0.1) q-values ranked across five methods and three disorders (schizophrenia, bipolar disorder and major depressive disorder). The number of genes in each pathway is listed in Table 2. Color reflects rank (red represents top-ranking sets with lowest P values). Sizes reflect the number of genes in the set (maximum of 200, minimum of 11). See Supplementary Data for source data.
Analysis of null and control disorder data
The enrichment P values of the pathways in Table 2 (and Supplementary Table 7), combined across SCZ, BIP and MDD, indicated that there are multiple significant pathways. To confirm that this was due to shared disorder biology, we repeated the rank-combining analysis on two further data sets as controls: a GWAS of HIV17, chosen because it is not thought to share significant disorder etiology with psychiatric disorders but might instead share real technical artifacts, and a null GWAS, simulated using the same SNPs as the psychiatric GWAS but with no significant loci. The most significant pathways for the HIV and null data sets analyzed separately are shown in Supplementary Table 4. As expected, no pathways showed significant enrichment in the null data set or HIV data set, and the SCZ, HIV and null data sets combined analysis (Supplementary Table 8) gave no significant pathways after multiple-testing correction (minimum q-value = 0.454), in contrast to the results for SCZ, MDD and BIP shown in Table 2. This gives further evidence that the significant pathway enrichments in Table 2 are due to shared biology across all three disorders, rather than being driven by enrichments in any single disorder.
Follow-up of brain gene expression of significant pathways
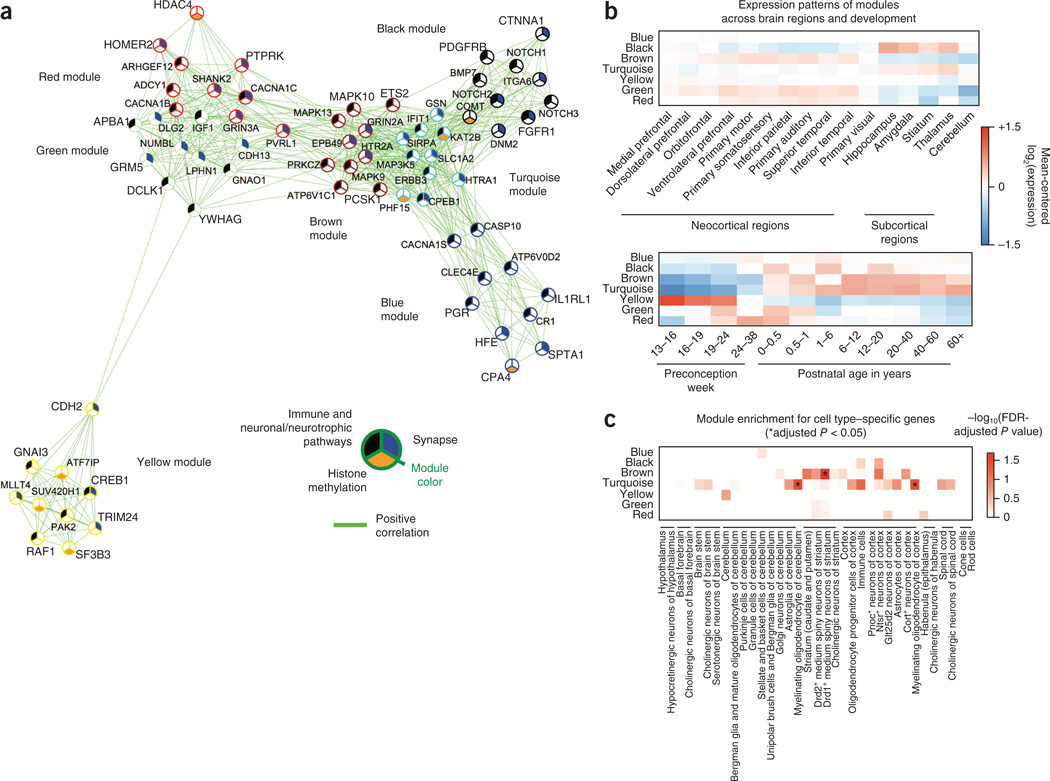
To place these pathways in a more specific neurobiological context, we analyzed coexpression relationships between the genes in pathways identified here at q < 0.1 using gene-expression data spanning brain regions and developmental time points18. Out of 797 genes assessed, 294 (32–37% of each branch from Fig. 3) were coexpressed in 7 modules. Figure 4a provides a network plot showing intramodular and intermodular connections between the top 10 hub genes in each module. Figure 4b summarizes the average expression level of genes in each module in different brain regions and temporal periods. These expression patterns are predominantly driven by temporal regulation (four modules have greater than twofold temporal change, while only one module, green, shows a twofold change between regions, Supplementary Table 9), suggesting that genetic risk across neuro-psychiatric disorders affects neurodevelopmentally regulated pathways.
Figure 4.
Gene coexpression networks across brain development and regions for genes in all pathways with FDR < 0.1. (a) Network plot of ten hubs genes from each module showing clustering across neuroanatomical regions and developmental epochs. The nodes (genes) are annotated by gene-set membership while the edges reflect positive correlations across brain regions and development. (b) Regional and temporal patterns of gene expression as summarized by the average expression level of genes in each module. (c) Enrichment for cell type–specific genes across multiple brain regions and cell types; asterisks highlight enrichments passing FDR-adjusted P < 0.05.
Of note, the yellow module is similarly expressed across regions and contains half (19/37) of the coexpressed histone methylation genes. Genes in this module exhibit over threefold higher average expression during early prenatal development (postconception week (PCW) 13–24) than during postnatal development or later aging, consistent with histone genes primarily functioning during neuronal differentiation and cell-fate commitment. Complete module membership and network details are available in Supplementary Table 9, which contains complete information about module-region associations and module-stage associations. Three other modules predominantly contain immunological and neuronal signaling and synapse genes whose expression increases in childhood and plateaus around adolescence to late adulthood (blue, brown, turquoise); three other modules are relatively highly expressed throughout life with ~75% maximum difference in expression between regions (green, black and red modules).
We then asked whether these modules exhibited a cell type–specific pattern by using gene lists from 35 genetically tagged and translationally profiled cell types in mouse19–21. Two of the modules containing immune-neuronal signaling and synapse genes that plateau in expression in maturing brain also exhibited cell type enrichment, with the brown module enriched for striatal neurons, particularly Drd1+ medium spiny neurons, and the turquoise module enriched for Cnp+ myelinating oligodendrocytes, suggesting its involvement in white matter maturation. Other modules did not exhibit strong cell type specificity, suggesting that either they affect multiple cell lineages (for example, the yellow module) or the matching cell type profile has yet to be defined.
DISCUSSION
Major advances have occurred in psychiatric genetics over recent years, largely driven by an order-of-magnitude increase in sample sizes. While the identification of specific loci is critical to moving the field forward, so too is developing an understanding of the underlying biology. In this study we address the latter, integrating data from the largest reported psychiatric genetics data sets with well-established tools for interrogating such data sets. We developed a novel rank-based method to combine pathway enrichment results across analysis methods and disorders in a manner that is not confounded by the biases or shortcomings of the methods, to maximize the informativeness of the results. This contrasts with the approach of the previous PGC Cross Disorder group GWAS4, which used a SNP-based meta-analysis. Pathway analyses based on such results will be powerful if the same SNP is implicated across disorders. Importantly, the pathway analyses described here do not require the same SNP (or, indeed, gene) within a pathway to be implicated across disorders, so they will be more robust to allelic heterogeneity within genes and within pathways across disorders, thereby providing a conceptually more powerful framework for conducting analyses, while losing power compared to single-SNP-level analyses if the same SNPs are driving the association across related disorders. We note that the most significant pathway identified from the PGC cross-disorder analysis3 (GO:0005262, calcium channel activity) also showed nominally significant enrichment in our analysis, confirming a role for calcium channel activity in these disorders.
Our analysis method uses GWAS summary statistics rather than using phenotype permutations on individual genotype data for two reasons. First, the PGC data sets are large and highly complex (>50 separate data sets) with a mixture of study designs and covariates (for example, those modeling ethnic stratification). For example, the PGC autism and ADHD genotype data are mixtures between trio and case-control studies. All of these complexities greatly increase the computational time necessary, so we implemented a more efficient method. For example, using a permutation-based approach that performed sufficient permutations across all the different data sets to generate disorder-level P values would have been computationally prohibitive. In addition, it is not always possible to obtain individual genotype data, particularly for large meta-analyses. It is therefore important that a pathway analysis method is applicable in such situations.
The primary strength of our integrated GWAS pathway analysis approach is the use of multiple analysis methods that differ in their assumptions and individual strengths. Methods that combine individual SNP P values across genes and pathways (FORGE, SETSCREEN) will pick up pathways containing genes with multiple, perhaps weak, independent association signals. Conversely, methods such as ALIGATOR or INRICH assign significance to genes based on the single most significant SNP in that gene, and will thus detect enrichment to pathways containing genes with individually, stronger associations. Notably, despite these differences, all of these methods yielded pathway rankings that were correlated with each other (Supplementary Table 5). As expected, the strongest correlations were observed between the most similar methods: FORGE and SETSCREEN, and ALIGATOR and INRICH.
Biological themes within and across disorders
Our primary aim was to combine pathway associations across disorders, as we hypothesized that this would be a more powerful approach, and we show a large increase in the evidence arising after meta-analyzing across disorders. The correlation of pathway-specific enrichment P values between SCZ and BIP was the highest among all pairs of disorders (0.29, Supplementary Table 5), consistent with the reported genetic correlation using common SNPs for these disorders4. Notably, combining the pathway-specific P values across the three ‘adult’ disorders in our primary meta-analysis (SCZ, BIP and MDD) resulted in greatly increased significance compared to the analyses of the separate disorders. This allowed us to identify biological themes spanning these disorders.
A secondary aim was to examine pathway themes within individual disorders, in order to show which disorders were contributing most to the cross-disorder findings. Among individual disorders, BIP, SCZ, MDD and ADHD gave significant results at FDR q < 10%, but ASD and HIV did not (Table 1 and Supplementary Table 4). Our results suggest that histone methylation appears to play a more prominent role in bipolar disorder and that synapse- and postsynapse-related processes are more strongly implicated in the etiology of schizophrenia (Table 1). We note that pathways involved in the methylation of other molecules, such as DNAs, were not highly enriched in this study, suggesting a degree of specificity in the nature of methylation gene sets implicated here. For schizophrenia, we note that “KEGG_DOPAMINERGIC_SYNAPSE” was reported as the third-highest-ranked pathway (of 9,016) in the latest GWAS data (analysis by ALIGATOR)22. Our top SCZ pathway—postsynaptic density—did not rank highly in either the ALIGATOR or INRICH analysis reported in that paper, but those pathway analyses were applied to a heavily restricted set of genes (those containing a SNP with P < 5 × 10−8) and used only two of our five analysis methods (ALIGATOR and INRICH), making that analysis not directly comparable with ours. We also include methods (FORGE, MAGENTA, SETSCREEN) that may favor more polygenic, complex patterns of association. Analyses presented here use a more relaxed significance criterion, thus picking up signals that do not reach genome-wide significance. We note that strong, independent support of our findings comes from the observation that analyses of rare variants have also implicated postsynaptic pathways in SCZ etiology23,24, suggesting that both rare and common variants are relevant.
Emerging landscape of psychiatric pathways
We show that synapse-related as well as newly implicated histone methylation and immune and neuronal signaling pathways are statistically significantly associated within and across SCZ, BIP and MDD. These pathways are core molecular processes, disruption of which may increase risk for multiple psychiatric disorders. Especially interesting in this regard is the identification of synaptic and immune dysfunction as the major pathways altered in postmortem brain in ASD25, as well as of emerging genetic overlap across many neurodevelopmental conditions. Our top results for schizophrenia, post-synaptic density, has been previously suggested by CNV findings and convergent lines of evidence23,26, as well as recent exome sequencing data27,28, suggesting a role for both rare and common variants in affecting SCZ-relevant changes in the postsynaptic membrane proteins. “Histone H3-K4 methylation” featured among top bipolar hits, achieving study-wide significance (q = 0.005). Variable H3-K4 methylation of synapsin genes has been shown to give rise to altered expression patterns in bipolar disorder and major depression29, which may suggest a role for epigenetic regulatory mechanisms in the etiology of mood disorders. Histone methylation mechanisms have roles in the coordination of complex cognitive processes such as long-term memory and roles in conditions from addiction to schizophrenia to neurodegeneration30.
Recent studies of de novo mutations have also implicated this pathway (often referred to by a broader term, chromatin regulation/ modification or transcriptional regulation) in ASD31–33. However, our ASD data set, based on GWAS, did not have sufficient power (owing to both the smaller sample size available and the combination of case-control and trio data sets) and was excluded from the primary pathway analysis. For MDD alone the top pathway was “protein phosphatase type 2A regulator activity,” which achieved a q-value of 0.012. Prior studies have implicated this pathway in serotonergic neurotransmission and in the mechanism of response to antidepressants34.
We excluded the schizophrenia-associated MHC region of 6p21–22 in our analysis to avoid confounding of methods by the very high levels of linkage disequilibrium at this locus. Despite this, key immune processes such as TGF-beta_signaling (P00052), B_cell_activation (P00010) and T_cell_activation (P00053) feature highly in our significant pathways. KEGG infectious disease pathways also featured at suggestive significance. Although these pathways contain immune processes, such as the Tuberculosis (hsa05152) and Hepatitis C (hsa05160) pathways, their role in the brain has been highlighted by, for example, studies finding that both hepatitis C infection and interferon alpha treatment for hepatitis C are associated with a range of additional neuropsychiatric symptoms35.
We also found pathway genome-wide association across SCZ, BIP and MDD with histone methylation (rather than DNA methylation) processes via Histone H3-K4 methylation (GO:51568), Histone methylation (GO:16571), Histone lysine methylation (GO:34968) and Macromolecule methylation (GO:43414). Replicated environmental risks for schizophrenia occur at critical periods early in development, for example, in the Dutch Hunger Winter and Chinese famine studies36, when the epigenome is known to be particularly labile. At this time rapid cell replication is occurring and the standard epigenetic signals, including histone H3-K4 and lysine methylation and other related processes, are driving development and tissue differentiation37. Given the role of this process in establishing active promoters38, it appears likely that the dysregulation of histone methylation may have downstream effects with the potential to disrupt neurodevelopment and coordinated gene expression, as animal studies have demonstrated39. Our results suggest that dysregulation of the genes in histone methylation pathways is a common etiological mechanism for adult psychiatric disorders.
Our gene network analysis of how the identified pathways are expressed in brain revealed modules of coexpressed genes that identify developmental time points and brain regions that may inform future experiments aimed at manipulating the identified pathways (Fig. 4). Additionally, enrichment for cell type–specific transcriptomic signatures identifies which cellular subtypes may be affected upon manipulation of specific genes. For example, the temporal trajectory of modules suggests which pathways may not be amenable to pharmacologic manipulation as a result of their prominence in early development (for example, the yellow module which contains histone methylation genes), but also pathways whose activity is potentially modifiable in adults (for example, the brown and turquoise modules, which were enriched for genes specific to striatal neurons and white matter, respectively). Additionally, the blue module’s expression pattern is similar across regions, and the hub genes suggest both intracellular and intercellular signaling processes. It shows an increase during late prenatal and early postnatal development, peaking before age 6, suggesting the genes in this module may be related to postnatal synaptic pruning, another potential target for developing interventions. We emphasize that these results should be considered preliminary, as they are a first-pass analysis using the results of a novel approach. Nevertheless, they do point a clear way forward for unbiased assessment of pathways affected by common genetic variation, which is expected to explain a majority of the genetic architecture of neuropsychiatric diseases. They suggest that the integration of gene expression with pathway-level analyses on larger GWAS can identify even greater specificity. Furthermore, future analyses can increase neurobiological specificity by searching across denser time point data or using cell type–specific transcriptomes acquired from single-cell sequencing in human brain, as they become available.
Our analyses have shown the general ability of pathway analyses to discover novel biology underlying complex human disorders. The immune-neuronal signaling and histone methylation findings illustrate how genetic risk aggregation in pathways may underlie vulnerability to environmental risk factors in the prenatal environmental, while strengthening the evidence for the role of synaptic pathways. Our results shed light on the biology underlying GWAS of psychiatric disorder and could suggest novel functional and drug discovery studies, as pathways make far larger and better drug targets than individual genes40. Our observation that the degree of correlation between pathways across disorders is higher than expected by chance builds on the observation of shared genetics between these disorders4 and, importantly, indicates that polygenic overlap is nonrandom at a molecular or pathway level.
ONLINE METHODS
Method rationale
The PGC (Psychiatric Genomics Consortium) has applied SNP association data3 and estimated “chip-heritability” estimate approaches2 to compare and contrast psychiatric disorders. Here we study the molecular pathways in which genetic risk for psychiatric disorders aggregate and examine whether or not these pathways are shared between related psychiatric disorders. The multiplicity of methods and parameters that can be used for such an undertaking led to the formation of a subgroup of the Psychiatric Genomics Consortium (http://pgc.unc.edu/) to develop a protocol and pipeline for five published methods of GWAS pathway analysis along with a methodology to combine results from different analytical methods to show the most robust pathway signals arising from GWAS data.
Samples and genotypes
The samples for these analyses (total N = 61,220) included cases, controls and family-based samples assembled for published genome-wide mega-analyses of individual-level data conducted by the PGC (see refs. 2,3,11,13–15 for details). To ensure comparability across samples, raw genotype and phenotype data for each study were uploaded to a central server and processed through the same quality control, imputation and analysis pipeline. This approach is detailed elsewhere11, but we describe it briefly here: to ensure independence of individual disorder analyses, only one of pair of related or duplicate individuals were retained, and in only one disorder case or control set, resulting in 61,220 cases and controls in total. Stringent and standardized quality control procedures were applied as previously described3. For the family-based samples, alleles transmitted to affected offspring (“trio cases”) were matched with untransmitted alleles (“pseudo-controls”). The disorder samples comprised ASD (4,788 trio cases, 4,788 trio pseudo-controls, 161 cases, 526 controls), ADHD (1,947 trio cases, 1,947 trio pseudo-controls, 840 cases, 688 controls,), BPD (6,990 cases, 4,820 controls), major depressive disorder (9,227 cases, 7,383 controls) and SCZ (9,379 cases, 7,736 controls). Identity-by-descent relationships were estimated for all pairs of individuals to identify any duplicate individuals across the component data sets. When duplicates were detected, one member of each set was retained. These individuals were then randomly apportioned to a single disorder case-control data set. All subjects were of European ancestry and met DSM-III-R or DSM-IV criteria for the primary disorder of interest.
Study sample numbers for individual disorders were: ASD (n = 4,949 affected/5,314 unaffected), ADHD (n = 2,787/2,635), BIP (n = 6,990/4,820), MDD (n = 9,227/7,383) and SCZ (n = 9,379/7,736). Imputation was conducted using HapMap III data as references, resulting in over 1.2M SNPs. SNPs with imputation quality scores less than 0.8 were filtered out. Single SNP-based association analyses were conducted using logistic regression on individual disorders with ancestry covariates and GC-corrected. The MHC region on chromosome 6 (25–35 Mbp) was excluded from further analyses to prevent potential impact of extensive linkage disequilibrium (LD) in the region.
For all analyses, five P-value sets were used: schizophrenia (SCZ), 1,227,336 SNPs; bipolar disorder (BIP), 1,223,695 SNPs; major depressive disorder (MDD), 1,220,925 SNPs; ADHD, 1,219,982 SNPs; autism (ASD), 1,232,050 SNPs. Our primary analysis included SCZ, BIP and MDD.
We also analyzed a null GWAS sample, to assess the degree of dependence between methods, as well as to ensure that our analysis did not generate an excesive amount of false positives. This was generated from unrelated CEU+TSI Hapmap3 data sets via random assignment of case/control phenotypes (100 cases and 100 controls). PGC1 data involves only European subjects (imputed using Hapmap3 CEU+TSI panels) and thus the use of Hapmap3 European data most closely matches the LD structure of the PGC1 disease samples. Since the null sample by definition contains no true effects, power is not an issue. Therefore, the relatively small sample size is not important.
Gene and pathway data
We used Ensembl41 gene definitions as the reference gene annotation and map. We combined gene set data from six sources: KEGG, GO, PANTHER, TARGETSCAN, REACTOME and OMIM. An overview of our methodology is shown in Figure 1. All gene sets were downloaded from their respective sources (11 August 2011). The parsing of these sets is summarized in Supplementary Table 1 and summary statistics are given in Supplementary Table 2.
To ensure the specificity of gene set association findings, further analyses were restricted to 4,949 gene sets of at most 200 genes and at least 10 genes (we limited our analysis to pathways containing 10–200 genes because statistics for smaller gene sets were over-dispersed and the few outlier sets with >200 genes were computationally inefficient to analyze and largely nonspecific due to the large number of genes (data not shown)). Identical pathways were removed. Overlap, measured by correlation between gene content across pathways, between the six gene set resources were minimal (R2 < 0.12). The different pathway sets were combined into one database and identical pathways merged.
Gene definitions
For all analyses, we used Ensembl identifiers as the master gene set, with −35 kb upstream and +10 kb downstream to define the gene boundaries, since transcriptional regulatory elements are likely to be contained within these intervals and that there is thus merit in capturing the variation within these regions42. Analysis was run both with and without the MHC region (chromosome 6, 25–35 Mb).
Standardization of pathway inputs
SNPs were assigned to genes based on human genome build 37 positions if they lay within 35 kb upstream or 10 kb downstream of the gene. In total, 739,373 SNPs were assigned to 18,689 genes. Note that if SNPs mapped within more than one gene, they were assigned to all such genes. SNPs were also filtered by imputation quality (INFO > 0.8), which resulted in 477,792–543,578 SNPs being assigned to 16,334–17,352 genes (these numbers vary slightly between disorders). We used a standardized framework of data input for our analyses. Empirical results (P values for individual SNPs) from PGC GWAS were gathered for each disorder, and all P values were GC-corrected. For all analyses, we used pathways with a minimum of 10 genes and a maximum of 200 genes.
Five published pathway analysis methods were used. These fell into two classes with differing approaches and assumptions regarding genomic architecture of risk variants in pathways as well as different methods for the correction of LD and gene size effects. FORGE43 and SET SCREEN TEST44 are meta-analysis methods that combine P values across all the SNPs in genes or pathways while adjusting for the confound of LD. INRICH45, ALIGATOR9 and MAGENTA46 are “best SNP per gene” methods that count the number of genes in a pathway where a number of independent SNPs exceed a predefined significance, and adjust for LD and genomic structure with corrected statistics derived by Monte-Carlo simulation. We describe these methods below.
FORGE method
FORGE is a software suite that implements a range of methods for the combination of P values for the individual genetic variants within a gene or genomic region while adjusting for linkage disequilibrium–induced correlations43. The software can be used with summary statistics (marker ids and P values) and accepts as input the result file formats of commonly used genetic association software. In addition, several utility programs are distributed with FORGE allowing users to (i) map SNP to genes using the Ensembl human genome annotation, (ii) parse different gene-set files, and (iii) calculate meta-analysis statistics for gene and gene-set analysis results when studies are carried out on multiple data sets. For each pathway, a nonparametric test yields a P value for enrichment of genes in a pathway given the entire set of pathways analyzed. FORGE can be freely accessed at https://github.com/inti/FORGE.
MAGENTA method
MAGENTA is an acronym for Meta-Analysis Gene-set Enrichment of variaNT Associations and is a program that takes as input summary P values from GWAS46. Testing for statistical significance of pathways using MAGENTA is a 3-step process. First, every gene is assigned the best GWAS P value that falls within that gene or the user-defined upstream and downstream regions of that gene. These P values are corrected, using multivariate linear regression, for known confounders of P values including gene size and linkage disequilibrium properties. Finally, for each pathway, the observed number of gene P values surpassing a certain user-specified threshold for P values (here 95%) is compared against expected number of gene P values surpassing that threshold for a given pathway size (i.e., number of genes). For each pathway, a nonparametric test yields a P value for enrichment of genes above the predetermined threshold.
Set screen test method
The set screen test is based on theoretical approximation of Fisher’s statistics such that the combination of P values at a gene or across a pathway is carried out in a manner that accounts for the correlation structure, or linkage disequilibrium, between single nucleotide polymorphisms. The approach is similar to that applied in FORGE (LD)43. The test is implemented in PLINK47. We applied this method to the PGC data, corrected for GC inflation, using CEU founders from HapMap to describe the LD structure. We used the same gene sets as described above which were filtered to contain no less than 2 and no more than 200 genes. For a given pathway or set, we assigned a P value to the set when at least one SNP was present. Where more than one SNP was present, the combined P value (accounting for LD) was given.
ALIGATOR method
ALIGATOR converts a list of significant and nominally significant SNPs into a list of significant genes, and, for each predefined gene set, tests whether this gene list contains more genes from the gene set than would be expected by chance9. This is done by comparing the gene list to 100,000 random gene lists of the same length generated by sampling SNPs (not genes) at random, correcting for variable numbers of SNPs per gene and variable gene size. Correction for the multiple testing of non-independent gene sets is performed using a bootstrap method repeated 5,000 times. Gene sets require at least two signals to be counted as enriched to remove the possibility of a small gene set being deemed significantly enriched based on one signal. An important modification to the original ALIGATOR method is that significant genes in the same gene set that mapped less than 1 Mb apart (and thus could be explained by the same association signal) are counted as a single signal. In this analysis, SNP-wise P value criteria for defining “significant” SNPs, and thus “significant” genes, were chosen so that the resulting list of significant genes contained the top 5% of all genes. When no filtering was performed, P value criteria varied from 8.4 × 10−4 to 3.31 × 10−4 when the −35 kb/+10 kb gene window was used. When SNPs were filtered by information score, the P values increased slightly, from 1.64 × 10−3 to 5.18 × 10−3.
INRICH method
INRICH45 takes a set of independent, nominally associated genomic intervals and then tests for the enrichment of predefined gene sets. An interval will typically correspond to a genomic region of SNP association defined by LD from a genome-wide scan, although intervals could also represent regions identified as homozygous-by-descent, for example, deletion or duplication events observed in cases. The INRICH analysis procedure comprises three major steps: (i) linkage disequilibrium (LD)-based interval data generation to identify unique regions of association; (ii) empirical enrichment calculation using an interval-based permutation strategy; and (iii) second-step permutation for multiple testing correction at the gene set level. INRICH also presents global enrichment statistics Gp, and the empirical significance of Gp is evaluated within a permutation procedure.
Pathway analysis strategy
Given that the results of the five analysis methods are correlated but not identical, pathways genuinely involved in disease susceptibility would be expected to show consistent enrichment for association signal across several methods. Therefore, we ranked the pathways in ascending order of enrichment P value for each method and calculated the average rank of each pathway across all five methods. This analysis was carried out for each disease separately. The lower the average rank of a pathway, the more consistent its evidence for enrichment of association signal across the methods, and thus the greater the likelihood of involvement in disease susceptibility. Ranks were used to control for differing power of each method.
Our general approach is outlined in Figure 1. Our primary analysis sets consisted of the samples for the adult disorders of schizophrenia (SCZ), bipolar disorder (BIP) and major depressive disorder (MDD) as these three have the highest genetic relationship in the recent pair-wise analysis of the five psychiatric disorders using GWAS data2. Supplemental analyses were performed on attention deficit hyperactivity disorder (ADHD) and autism (ASD) data sets3,13. We used a Monte-Carlo simulation approach, modeling the dependence between methods in terms of the observed pairwise correlations of pathway enrichment P values to calculate the average rank and significance of a pathway in a disorder across all methods (Supplementary Analysis and Supplementary Tables 12–15). However, the analysis method and results are robust to variation in these correlations (Supplementary Tables 15 and 16). The motivation for using ranks rather than P values was to ensure that all methods were treated equally. Specifically, FORGE and SET-SCREEN combine the P values of the SNPs, and it is therefore possible for them to achieve very small enrichment P values if the pathway contains strongly associated SNPs. This is not the case for ALIGATOR, INRICH and MAGENTA, which use simulation to give enrichment P values. Note also that INRICH and ALIGATOR require a pathway to contain two significant genes for an enrichment P value to be calculated. Thus, missing enrichment P values count as evidence against enrichment for these methods, and such pathways are assigned the joint bottom rank. It is possible that a pathway may rank relatively poorly on one method compared to the others, thus reducing the power of the average rank to detect enrichment. We therefore performed a secondary analysis based on the average of the best four ranks. However, this had little effect on the results (see Supplementary Analysis and Supplementary Tables 12 and 13).
We calculated observed and expected overlaps between all combinations of methods to assess the extent of concordance between methods. For the top 10% ranked pathways in each disorder, we then compared observed and expected overlaps between all combinations of the three adult disorders.
Combination of pathway ranks across methods
Finally, we combined pathway P values across disorders using Brown’s method (an extension of Fisher’s method for correlated data). To ensure that the observed pathway enrichments across the disorders were a result of shared biology, rather than artifacts of the analysis method, we also applied our rank-based method to a null data set (simulated to have no phenotype effects) and a GWAS of HIV-1 acquisition17. HIV infection acquisition can be assumed to share little biology with the psychiatric disorders studied here because, although two MHC associations have been discovered, we excluded the entire extended MHC region from our analyses. The null and HIV data sets were tested individually, as well as in combination with the SCZ data set.
The following procedure yields a single combined P value for each pathway in a given disease data set by merging results across the 5 methods, accounting for correlation. For each disease, do the following:
Determine average rank per pathway within each method. After ranking P values (ties receive the average rank), average the ranks across the five methods to yield 1 rank per pathway.
Determine the expected distribution of averaged ranks under the null. (a) Calculate the Pearson correlation statistics between pairs of methods. Null data (generated from unrelated European CEU+TSI Hapmap3 GWAS data set via random phenotype assignment (100 cases, 100 controls)) from each method was used for these calculations. (b) Generate 5 sets of null P values (1 for each pathway drawn without replacement from the uniform distribution [0,1]) such that the intercorrelation among methods is preserved, using the method described at http://comisef.wikidot.com/tutorial:correlateduniformvariates. Do this 10,000 times. (c) Transform P values into ranks for each permuted P value distribution. Note that in the permuted data, there will be few ties, if at all. Therefore, introduce ties by replacing the ranks for those pathways that tied in the real data with their average rank in the permuted data. (d) For each set of 5 permuted and ranked distributions, determine the average rank per pathway (as in Step 1).
Assign empirical P values to each pathway. , where i = 0 if the permuted rank is greater than the real rank, and i = 1 if the permuted rank is less than or equal to the real rank. Note that this procedure does not allow for dependence between pathways, so it cannot be used to test whether there is an excess of pathways with average rank achieving a given level of significance. However, it does give a valid test of significance for each pathway separately. To correct for multiple testing of pathways, q-values were calculated48. For each pathway, the q-value corresponds to the minimum value of the FDR at which that pathway would be declared significant.
Comparisons between methods: method overlap
To robustly test the significance of overlap in enriched pathways between methods, it is necessary to restrict the analysis to a set of independent pathways (by gene membership). This was done by pruning by Jaccard distance (see below). Note that such a restriction is not necessary and was not used for the pathway analysis combining methods, which uses the full set of pathways. To facilitate comparisons between methods, we use quantiles, not P values. Specifically, we focus on pathways that are in the top 25% for a particular method within a disease or null data set. This is because it is otherwise difficult to compare pathway P values from methods that have different statistical power. We use the following procedure to determine the extent to which methods overlap:
For a given data set, reduce the data set down to only pathways that are in the top 25% in ≥1 of the methods;
Reduce redundancies in this data set by removing smaller pathways for which there exists a larger pathway whose Jaccard distance (intersection divided by the union) is ≥0.2;
Using this reduced data set, which represents a set of independent pathways that are in the top 25% of ≥1 of the methods, calculate all overlaps among five methods (5-way, 4-way, 3-way and 2-way);
Calculate the expected overlap between pathways in the top 25% assuming top 25% of pathways for each method is random.
Testing pathway overlap between diseases
In order to enable testing to be carried out for correlation in pathway enrichment P values and overlap in top pathways among diseases, a subset of 1,918 pathways was selected such that no two pathways had a Jaccard similarity measure >0.2. Initially, Pearson correlation coefficients were calculated between the pathway-specific enrichment P values of the null data set and each of the five disease data sets in turn. These correlations lay between 0.111 and 0.156, with a mean of 0.132. This indicates that ranking pathways within methods and calculating the average rank across methods induces some correlation between pathway ranks between data sets. For example: ALIGATOR and INRICH only return a P value if a pathway contains at least two significant genes, otherwise the pathway was assigned equal bottom rank. Thus, small pathways are likely to have low ranks for these methods in most data sets. However, correlations between the diseases with respect to the pathway-specific enrichment P values were higher than those with the null or HIV (see Supplementary Tables 5 and 14), suggesting that the interdisease correlation is not simply a function of methodological correlation.
To test for significant correlation and overlap between the five disease data sets while allowing for correlations induced by the ranking method (as described above), we generated, for each pair of diseases, 1,000 random sets of 1,918 bivariate uniform variables with correlation of 0.132. The Pearson correlation of the two variables within each replicate data set was calculated and compared to that observed in the actual data. A similar method was used to compare the overlap in the top 10% of pathways between the two variables to that observed in the actual data. P values for these comparisons are shown in Supplementary Table 6. Correlation coefficients in the actual data were nominally significant for all pairs of the five diseases of interest (ADHD, ASD, BIP, MDD, SCZ), but not between these diseases and HIV or the null data set (with the exception of the HIV-MDD correlation, which was nominally significant but is likely to be an artifact of multiple testing).
Combining pathway enrichments across diseases
For each pathway, P values were combined across diseases using Brown’s method (an extension of Fisher’s method that accounts for correlation between data sets). This method is described49, but a brief description is provided here. To attain a joint test statistic for N tests that are not independent, the statistic has a mean m = 2N and a variance (σ2) where
and where pi and pj (i, j = 1,…, N) are the P values for each test and covariance (cov) is calculated as
for non-negative correlation coefficients ρij between the two P value distributions (here, set to be 0.132, which is the average correlation in P values between the disease data sets and the null, calculated in the previous section). Finally the overall significance of a set of non-independent tests is calculated using the statistic T which under the null hypothesis follows the central chi-square distribution T = T0/c, with 2N/c degrees of freedom, where
and T0 is the sum of −2ln(p), as used in Fisher’s method.
This analysis was primarily performed on the three adult diseases (SCZ, BIP, MDD), which have larger and more powerful samples than the other diseases (ADHD, ASD). A secondary analysis of all five diseases was also performed. Finally, an analysis was performed combining SCZ, HIV and the null data set to confirm that pathway enrichments observed in the analysis of SCZ+BIP+MDD were due to shared biology rather than artifacts of the analysis method.
When the diseases were tested separately, one pathway was significant after correction for multiple testing in BIP (GO:51568: histone H3-K4 methylation, q = 0.005) and MDD (GO:8601: protein phosphatase Type 2A regulator activity, q = 0.012). No pathway achieved q < 0.05 in the other diseases (or the null data set). When the three adult diseases were combined, 15 pathways were significant at q < 0.05, with the top pathway (GO:51568: histone H3-K4 methylation) having a q-value of 0.0005. These results are more significant than those observed in any of the diseases analyzed separately, and illustrate the power to be gained by combining pathway analyses across diseases with shared biology. When all five diseases were combined, GO:51568 was still significant (q = 0.045), although its significance was reduced due to lack of enrichment in ADHD or ASD. Finally, when SCZ was combined with HIV (not expected to share common biology with SCZ) and the null data set, no pathway was significant after multiple testing correction, as expected.
Supervised weighted coexpression network analysis
We performed a secondary analysis of 797 genes comprising all pathways with q < 0.1 from the primary cross disorder pathway enrichment analysis to explore how pathways relate to processes of in vivo brain development and aging. We asked how the genes cluster across brain regions and developmental time points using the BrainSpan exon array data18. We applied weighted gene coexpression network analysis50 to group genes into modules of coexpression. Modules were characterized for regional and temporal patterns as well as cell type specificity21.
Network analysis was performed using the WGCNA package in R. Gene-expression data were obtained from GSE25219 for 16,874 genes across 1,340 samples (75 individuals spanning 15 developmental stages with up to 16 brain regions per individual). Only regions with at least 10 samples were used, leaving 1,281 samples. After intersecting genes in the immune, synapse and methylation pathways, 797 genes were left for the network analysis, whose module parameters and membership are outlined in Supplementary Table 17. The specific parameters are made available in the supplemental R script for Figure 4 (Supplementary Software). Similar modules were found with variations on these parameters.
The top 10 genes in each module are plotted in Figure 4a using the igraph package in R. Only correlations with r > 0.2 are shown, and the Fruchterman-Reingold force-directed algorithm as implemented in igraph was used to layout nodes using default parameters. Module expression profiles were summarized by taking the mean expression level of all genes in each module for each different brain region (across all time points) and each neurodevelopmental epoch (across all regions). We focused on relatively large changes between regions or time points (>75%), though statistical analysis of spatial and temporal patterns with Kruskal-Wallis tests and pairwise Mann-Whitney U-tests demonstrates many smaller differences are significant (due to the large sample size). In general, alternative methods for evaluating regional and temporal differences (for example, correlations with regional indicator variables, ANOVA with regions as factors) yielded similar patterns as those seen in Figure 4b. For simplicity of interpretation and discussion, we therefore chose to focus on these larger fold changes in order to highlight the most salient neurodevelopmental changes at the pathway level.
Cell type–specific enrichment for modules was performed by using the cell specific expression analysis (CSEA) tool (http://genetics.wustl.edu/jdlab/csea-tool-2/). This tool contains cell type–specific genes that are derived from a translational profiling approach that isolates transcriptomes in mouse from specific, marker-defined cellular subpopulations21. We assessed each module for enrichment of lists for 35 broad and specific cell type gene sets (CSEA specificity threshold set to 0.05) across multiple brain regions in mouse, and corrected for multiple comparisons across 7 modules and the 35 cell type lists. We report enrichments at Benjamini-Hochberg corrected P < 0.05 in Figure 4c. The code underlying this analysis is included as Supplementary Software.
A Supplementary Methods Checklist is available.
Supplementary Material
ACKNOWLEDGMENTS
G.B. and S.N. acknowledge funding support for this work from the National Institute for Health Research (NIHR) Mental Health Biomedical Research Centre at South London and Maudsley NHS Foundation Trust and King’s College London. P.H.L. is supported by US National Institute of Mental Health (NIMH) grant K99MH101367. The PGC Cross-Disorder Group is supported by NIMH grant U01 MH085520. Statistical analyses were carried out on the Genetic Cluster Computer, which is financially supported by the Netherlands Scientific Organization (NOW; 480-05-003; principal investigator D.P.) along with a supplement from the Dutch Brain Foundation and VU University. Numerous (>100) grants from government agencies along with substantial private and foundation support worldwide enabled the collection of phenotype and genotype data, without which this research would not be possible.
Appendix
Colm O’Dushlaine1,2,257, Lizzy Rossin2,257, Phil H Lee3,257, Laramie Duncan2,4, Neelroop N Parikshak5, Stephen Newhouse6,7, Stephan Ripke2,4, Benjamin M Neale2,4, Shaun M Purcell2,4,8, Danielle Posthuma9–11, John I Nurnberger12,13, S Hong Lee14, Stephen V Faraone15,16, Roy H Perlis2,3, Bryan J Mowry14,17, Anita Thapar18,19, Michael E Goddard20,21, John S Witte22, Devin Absher23, Ingrid Agartz24,25, Huda Akil26, Farooq Amin27, Ole A Andreassen24,28, Adebayo Anjorin29, Richard Anney30, Verneri Anttila2, Dan E Arking31, Philip Asherson6, Maria H Azevedo32, Lena Backlund33, Judith A Badner34, Anthony J Bailey35, Tobias Banaschewski36, Jack D Barchas37, Michael R Barnes38, Thomas B Barrett39, Nicholas Bass29, Agatino Battaglia40, Michael Bauer41, Mònica Bayés42, Frank Bellivier43–46, Sarah E Bergen4,3,47, Wade Berrettini48, Catalina Betancur49–51, Thomas Bettecken52, Joseph Biederman53, Elisabeth B Binder52, Donald W Black54, Douglas H R Blackwood55, Cinnamon S Bloss56,57, Michael Boehnke58,59, Dorret I Boomsma60–62, René Breuer63, Richard Bruggeman64, Paul Cormican30, Nancy G Buccola65, Jan K Buitelaar66, William E Bunney67, Joseph D Buxbaum68, William F Byerley69,70, Enda M Byrne14, Sian Caesar71, Wiepke Cahn72, Rita M Cantor73, Miguel Casas74,75, Aravinda Chakravarti31, Kimberly Chambert4, Khalid Choudhury29, Sven Cichon76–79, Manuel Mattheisen78,80–82, C Robert Cloninger83, David A Collier6, Edwin H Cook84, Hilary Coon85, Bru Cormand86–88, Aiden Corvin30, William H Coryell54, David W Craig89, Ian W Craig6, Jennifer Crosbie90, Michael L Cuccaro91, David Curtis92, Darina Czamara52,93, Susmita Datta94, Geraldine Dawson95–97, Richard Day98, Eco J De Geus60–62, Franziska Degenhardt76,78, Srdjan Djurovic24,99, Gary J Donohoe30, Alysa E Doyle100, Jubao Duan101, Frank Dudbridge102, Eftichia Duketis103, Richard P Ebstein104, Howard J Edenberg105, Josephine Elia48,106, Sean Ennis107, Bruno Etain43,46,108, Ayman Fanous109,110, Anne E Farmer6, I Nicol Ferrier111, Matthew Flickinger58,59, Eric Fombonne112,113, Tatiana Foroud22, Josef Frank63, Barbara Franke66, Christine Fraser18,19, Robert Freedman114, Nelson B Freimer115, Christine M Freitag103, Marion Friedl116, Louise Frisén33, Louise Gallagher30, Pablo V Gejman101, Lyudmila Georgieva18,19, Elliot S Gershon34, Ina Giegling116, Michael Gill30, Scott D Gordon117, Katherine Gordon-Smith18,71, Elaine K Green118, Tiffany A Greenwood119, Dorothy E Grice120,121, Magdalena Gross122, Detelina Grozeva18, Weihua Guan58,59,123, Hugh Gurling29, Lieuwe De Haan124, Jonathan L Haines125, Hakon Hakonarson126,127, Joachim Hallmayer128, Steven P Hamilton69, Marian L Hamshere129, Thomas F Hansen80,130, Annette M Hartmann116, Martin Hautzinger129, Andrew C Heath83, Anjali K Henders117, Stefan Herms76,79, Ian B Hickie131, Maria Hipolito132, Susanne Hoefels122, Florian Holsboer52, Witte J Hoogendijk133, Jouke-Jan Hottenga60,62, Christina M Hultman47, Vanessa Hus134, Andrés Ingason80,130, Marcus Ising52, Stéphane Jamain43,46,108, Edward G Jones80,130, Ian Jones18,19, Lisa Jones71, Jung-Ying Tzeng135, Anna K Kähler47, René S Kahn72, Radhika Kandaswamy29, Matthew C Keller136, James L Kennedy137, Elaine Kenny30, Lindsey Kent138, Yunjung Kim139, George K Kirov18,19, Sabine M Klauck140, Lambertus Klei141, James A Knowles142, Martin A Kohli52, Daniel L Koller22, Bettina Konte116, Ania Korszun143, Lydia Krabbendam144, Robert Krasucki29, Jonna Kuntsi6, Phoenix Kwan58,59, Mikael Landén47,145, Niklas Längström47, Mark Lathrop146, Jacob Lawrence29, William B Lawson132, Marion Leboyer43,46,108, David H Ledbetter147, Todd Lencz148–150, Klaus-Peter Lesch151,152, Douglas F Levinson153, Cathryn M Lewis6, Jun Li154, Paul Lichtenstein47, Jeffrey A Lieberman155, Dan-Yu Lin156, Don H Linszen157, Chunyu Liu158, Falk W Lohoff48, Sandra K Loo115,159, Catherine Lord160, Jennifer K Lowe161,162, Susanne Lucae52, Donald J MacIntyre55, Pamela A F Madden83, Elena Maestrini163, Patrik K E Magnusson47, Pamela B Mahon164, Wolfgang Maier122, Anil K Malhotra148–150, Shrikant M Mane165, Christa L Martin147, Nicholas G Martin117, Keith Matthews98, Morten Mattingsdal24,166, Steven A McCarroll4, Kevin A McGhee55, James J McGough167, Patrick J McGrath155, Peter McGuffin6, Melvin G McInnis168, Andrew McIntosh55,169, Rebecca McKinney119, Alan W McLean55,169, Francis J McMahon170, William M McMahon85, Andrew McQuillin29, Helena Medeiros142, Sarah E Medland117, Sandra Meier63, Ingrid Melle24,28, Fan Meng26, Jobst Meyer171, Christel M Middeldorp60,62, Lefkos Middleton172, Vihra Milanova173, Ana Miranda174, Anthony P Monaco175,176, Grant W Montgomery117, Jennifer L Moran4, Daniel Moreno-De-Luca177, Gunnar Morken178,179, Derek W Morris30, Eric M Morrow180,181, Valentina Moskvina18,182, Pierandrea Muglia183, Thomas W Mühleisen76,78,184, Walter J Muir55,169,256, Bertram Müller-Myhsok52,93, Michael Murtha185, Richard M Myers23, Inez Myin-Germeys144, Michael C Neale110, Stan F Nelson115, Caroline M Nievergelt119, Ivan Nikolov18,19, Vishwajit Nimgaonkar186,187, Willem A Nolen188, Markus M Nöthen76,78, Evaristus A Nwulia132, Dale R Nyholt117, Robert D Oades189, Ann Olincy114, Guiomar Oliveira32,190, Line Olsen80,130, Roel A Ophoff115,191, Urban Osby33, Michael J Owen18,19, Aarno Palotie192, Jeremy R Parr111, Andrew D Paterson193,194, Carlos N Pato142, Michele T Pato142, Brenda W Penninx61,62,195, Michele L Pergadia83, Margaret A Pericak-Vance91, Benjamin S Pickard55,169, Jonathan Pimm29, Joseph Piven97, James B Potash54, Fritz Poustka103, Peter Propping78, Vinay Puri29, Digby J Quested196, Emma M Quinn30, Josep Antoni Ramos-Quiroga74,75, Henrik B Rasmussen80,130, Soumya Raychaudhuri2,4, Karola Rehnström192, Andreas Reif197, Marta Ribasés74,198, John P Rice199, Marcella Rietschel63, Kathryn Roeder200, Herbert Roeyers201, Aribert Rothenberger202, Guy Rouleau203, Douglas Ruderfer8, Dan Rujescu116, Alan R Sanders101, Stephan J Sanders177,186,204,205, Susan L Santangelo206,207, Joseph A Sergeant208, Russell Schachar90, Martin Schalling33, Alan F Schatzberg209, William A Scheftner210, Gerard D Schellenberg211, Stephen W Scherer212, Nicholas J Schork56,213, Thomas G Schulze164,214, Johannes Schumacher78, Markus Schwarz215, Edward Scolnick4, Laura J Scott58,59, Jianxin Shi216, Paul D Shilling119, Stanley I Shyn217, Jeremy M Silverman121, Susan L Slager218, Susan L Smalley115,159, Johannes H Smit61,195, Erin N Smith56,213, Edmund J S Sonuga-Barke201,219, David St. Clair220, Matthew State177,185,204, Michael Steffens221, Hans-Christoph Steinhausen222–224, John S Strauss225, Jana Strohmaier63, T Scott Stroup226, James S Sutcliffe227, Peter Szatmari228–230, Szabocls Szelinger89, Srinivasa Thirumalai231, Robert C Thompson26, Alexandre A Todorov83, Federica Tozzi183, Jens Treutlein63, Manfred Uhr52, Edwin J C G van den Oord232, Gerard Van Grootheest61,195, Jim Van Os144, Astrid M Vicente233–235, Veronica J Vieland236, John B Vincent226, Peter M Visscher30, Christopher A Walsh237–240, Thomas H Wassink54, Stanley J Watson26, Myrna M Weissman241, Thomas Werge130,80,242, Thomas F Wienker243, Ellen M Wijsman244,245, Gonneke Willemsen60,61, Nigel Williams18,19, A Jeremy Willsey185,204, Stephanie H Witt63, Wei Xu194, Allan H Young111,246, Timothy W Yu247, Stanley Zammit18,19, Peter P Zandi248, Peng Zhang58,59,168, Frans G Zitman249, Sebastian Zöllner58,59,168, International Inflammatory Bowel Disease Genetics Consortium (IIBDGC)250, Bernie Devlin141, John R Kelsoe119,251, Pamela Sklar19, Mark J Daly2,4, Michael C O’Donovan18,19, Nicholas Craddock18,19, Kenneth S Kendler110,252,253, Lauren A Weiss69, Naomi R Wray14, Zhaoming Zhao254,255, Daniel H Geschwind161,162, Patrick F Sullivan139, Jordan W Smoller3,4, Peter A Holmans18,182,257& Gerome Breen6,7,257
1Regeneron Genetics Center, Regeneron Pharmaceuticals Inc., Tarrytown, NY. 2Stanley Center for Psychiatric Research, Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA. 3Psychiatric and Neurodevelopmental Genetics Unit, Massachusetts General Hospital, Boston, Massachusetts, USA. 4Analytic and Translational Genetics Unit, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA. 5Program in Neurobehavioral Genetics, Semel Institute, David Geffen School of Medicine, UCLA, Los Angeles, California, USA. 6Medical Research Council (MRC) Social Genetic and Developmental Psychiatry (SGDP) Centre, King’s College London, The Institute of Psychiatry Psychology and Neuroscience, London, UK. 7National Institute of Health Research (NIHR) Biomedical Research Centre for Mental Health, South London and Maudsley NHS Trust and Insitute of Psychiatry, London, UK. 8Division of Psychiatric Genomics, Department of Psychiatry, Icahn School of Medicine at Mount Sinai, New York, New York, USA. 9Department of Functional Genomics, VU University, Amsterdam, the Netherlands. 10Department of Clinical Genetics, VU Medical Center, Amsterdam, the Netherlands. 11Department of Child and Adolescent Psychiatry, Erasmus University Medical Center, Rotterdam, the Netherlands. 12Institute of Psychiatric Research, Indiana University School of Medicine, Indianapolis, Indiana, USA. 13Department of Medical and Molecular Genetics, Indiana University School of Medicine, Indianapolis, Indiana, USA. 14The University of Queensland, Queensland Brain Institute, Brisbane, Queensland, Australia. 15Department of Psychiatry, State University of New York (SUNY) Upstate Medical University, Syracuse, New York, USA. 16Department of Neuroscience and Physiology, SUNY Upstate Medical University, Syracuse, New York, USA. 17Queensland Centre for Mental Health Research, Wacol, Queensland, Australia. 18MRC Centre for Neuropsychiatric Genetics and Genomics, Cardiff University School of Medicine, Cardiff, UK. 19Institute of Psychological Medicine and Clinical Neurosciences, Cardiff University School of Medicine, Cardiff, UK. 20Biosciences Research Division, Department of Environment and Primary Industries Victoria, Melbourne, Victoria, Australia. 21Faculty of Land and Environment, University of Melbourne, Melbourne, Victoria, Australia. 22Department of Epidemiology and Biostatistics, University of California, San Francisco, San Francisco, California, USA. 23HudsonAlpha Institute of Biotechnology, Huntsville, Alabama, USA. 24KG Jebsen Centre for Psychosis Research, Institute of Clinical Medicine, University of Oslo, Oslo, Norway. 25Department of Research, Diakonhjemmet Hospital, Oslo, Norway. 26Molecular Psychiatry Laboratory, Molecular and Behavioral Neuroscience Institute, University of Michigan, Ann Arbor, Michigan, USA. 27Department of Psychiatry and Behavioral Sciences, Atlanta Veterans Affairs Medical Center, Emory University, Atlanta, Georgia, USA. 28Division of Mental Health and Addiction, Oslo University Hospital, Oslo, Norway. 29Mental Health Sciences Unit, University College London, London, UK. 30Department of Psychiatry, Trinity College Dublin, Dublin, Ireland. 31McKusick-Nathans Institute of Genetic Medicine, Johns Hopkins University School of Medicine, Baltimore, Maryland, USA. 32Faculty of Medicine, University of Coimbra, Coimbra, Portugal. 33Department of Molecular Medicine and Surgery, Center for Molecular Medicine, Karolinska Institutet, Stockholm, Sweden. 34Department of Psychiatry, University of Chicago, Chicago, Illinois, USA. 35Department of Psychiatry, University of British Columbia, Vancouver, British Columbia, Canada. 36Department of Child and Adolescent Psychiatry and Psychotherapy, Central Institute of Mental Health, Medical Faculty Mannheim, University of Heidelberg, Mannheim, Germany. 37Department of Psychiatry, Weill Medical College, Cornell University, New York, New York, USA. 38GlaxoSmithKline, London, UK. 39Portland Veterans Affairs Medical Center, Portland, Oregon, USA. 40Stella Maris Institute for Child and Adolescent Neuropsychiatry, Calambrone, Pisa, Italy. 41Department of Psychiatry and Psychotherapy, Carl Gustav Carus University Hospital, Dresden, Germany. 42Centro Nacional de Análisis Genómico (CNAG), Parc Científic de Barcelona (PCB), Barcelona, Spain. 43Institut National de la Santé et de la Recherche Médicale (INSERM) U955, Psychiatrie Génétique, Créteil, France. 44Université Denis Diderot, Paris, France. 45Assistance Publique–Höpitaux de Paris (AP-HP), Groupe Hospitalier Saint-Louis, Lariboisiere, F Widal, Departement de Psychiatrie, Paris, France. 46ENBREC (European Network of Bipolar Research Expert Centres) Group, Fondation FondaMental, Créteil, France. 47Department of Medical Epidemiology and Biostatistics, Karolinska Institutet, Stockholm, Sweden. 48Department of Psychiatry, University of Pennsylvania, Philadelphia, Pennsylvania, USA. 49INSERM U952, Paris, France. 50Centre National de la Recherche Scientifique (CNRS) Unité Mixte de Recherche (UMR) 7224, Paris, France. 51Université Pierre et Marie Curie, Paris, France. 52Max Planck Institute of Psychiatry, Munich, Germany. 53Clinical and Research Programs in Pediatric Psychopharmacology and Adult ADHD, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA. 54Department of Psychiatry, University of Iowa, Iowa City, Iowa, USA. 55Division of Psychiatry, University of Edinburgh, Royal Edinburgh Hospital, Edinburgh, UK. 56The Scripps Translational Science Institute, La Jolla, California, USA. 57Scripps Health, La Jolla, California, USA. 58Department of Biostatistics, School of Public Health, University of Michigan, Ann Arbor, Michigan, USA. 59Center for Statistical Genetics, School of Public Health, University of Michigan, Ann Arbor, Michigan, USA. 60Department of Biological Psychology, VU University, Amsterdam, the Netherlands. 61EMGO+ (ExtraMuraalGeneeskundig Onderzoek) Institute for Health and Care Research, Amsterdam, the Netherlands. 62Neuroscience Campus Amsterdam, Amsterdam, the Netherlands. 63Department of Genetic Epidemiology in Psychiatry, Central Institute of Mental Health, Medical Faculty Mannheim, Heidelberg University, Mannheim, Germany. 64Department of Psychiatry, University Medical Center Groningen, University of Groningen, Groningen, the Netherlands. 65School of Nursing, Louisiana State University Health Sciences Center, New Orleans, Louisiana, USA. 66Department of Cognitive Neuroscience, Donders Institute for Brain, Cognition and Behavior, Radboud University Medical Centre, Nijmegen, the Netherlands. 67Department of Psychiatry and Human Behavior, University of California, Irvine, Irvine, California, USA. 68Seaver Autism Center for Research and Treatment, Department of Psychiatry, Icahn School of Medicine at Mount Sinai, New York, New York, USA. 69Department of Psychiatry, University of California, San Francisco, San Francisco, California, USA. 70NCIRE (Northern California Institute of Q Research and Education), San Francisco, California, USA. 71Department of Psychiatry, Birmingham University, Birmingham, UK. 72Department of Psychiatry, Rudolf Magnus Institute of Neuroscience, University Medical Center, Utrecht, the Netherlands. 73David Geffen School of Medicine, University of California, Los Angeles, Los Angeles, California, USA. 74Department of Psychiatry, Hospital Universitari Vall d’Hebron, CIBERSAM (Centro de Investigación Biomédica en el Area de Salud Mental), Barcelona, Spain. 75Department of Psychiatry and Legal Medicine, Universitat Autònoma de Barcelona, Barcelona, Spain. 76Department of Genomics, Life & Brain Center, University of Bonn, Bonn, Germany. 77Institute of Neuroscience and Medicine (INM-1), Research Center Jülich, Jülich, Germany. 78Institute of Human Genetics, University of Bonn, Bonn, Germany. 79Division of Medical Genetics, Department of Biomedicine, University of Basel, Basel, Switzerland. 80The Lundbeck Initiative for Integrative Psychiatric Research, iPSYCH, Roskilde, Denmark. 81Department of Biomedicine, Aarhus University, Aarhus, Denmark. 82Department of Genomic Mathematics, University of Bonn, Bonn, Germany. 83Department of Psychiatry, Washington University School of Medicine, St. Louis, Missouri, USA. 84Department of Psychiatry, Institute for Juvenile Research, University of Illinois, Chicago, Illinois, USA. 85Department of Psychiatry, University of Utah, Salt Lake City, Utah, USA. 86Departament de Genètica, Facultat de Biologia, Universitat de Barcelona, Barcelona, Spain. 87Biomedical Network Research Centre on Rare Diseases (CIBERER), Barcelona, Spain. 88Institut de Biomedicina de la Universitat de Barcelona (IBUB), Barcelona, Spain. 89The Translational Genomics Research Institute, Phoenix, Arizona, USA. 90Neurosciences and Mental Health Program, The Hospital for Sick Children, University of Toronto, Toronto, Ontario, Canada. 91John P. Hussman Institute for Human Genomics, University of Miami, Miami, Florida, USA. 92East London NHS Foundation Trust, Queen Mary, University of London, London, UK. 93Munich Cluster for Systems Neurology (SyNergy), Munich, Germany. 94Genetics Institute, University College London, London, UK. 95Autism Speaks, New York, New York, USA. 96Department of Psychiatry, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina, USA. 97Carolina Institute for Developmental Disabilities, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina, USA. 98Division of Neuroscience, Medical Research Institute, University of Dundee, Ninewells Hospital & Medical School, Dundee, UK. 99Department of Medical Genetics, Oslo University Hospital, Oslo, Norway. 100Psychiatric and Neurodevelopmental Genetics Unit, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA. 101Department of Psychiatry and Behavioral Sciences, NorthShore University Health System and University of Chicago, Evanston, Illinois, USA. 102Department of Non-Communicable Disease Epidemiology, London School of Hygiene and Tropical Medicine, London, UK. 103Department of Child and Adolescent Psychiatry, Psychosomatics and Psychotherapy, JW Goethe University Frankfurt, Frankfurt, Germany. 104Psychology Department, National University of Singapore, Singapore. 105Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis, Indiana, USA. 106AI Dupont Hospital for Children, University of Pennsylvania, Philadelphia, Pennsylvania, USA. 107School of Medicine, Medical Science University College, Dublin, Ireland. 108AP-HP, Höpital H Mondor–A Chenevier, Département de Psychiatrie, Créteil, France. 109Department of Psychiatry, Georgetown University School of Medicine, Washington, DC, USA. 110Virginia Institute of Psychiatric and Behavioral Genetics, Virginia Commonwealth University, Richmond, Virginia, USA. 111Institute of Neuroscience, Newcastle University, Newcastle upon Tyne, UK. 112Department of Psychiatry, Oregon Health & Science University, Portland, Oregon, USA. 113Institute for Development & Disability, Oregon Health & Science University, Portland, Oregon, USA. 114Department of Psychiatry, University of Colorado Denver, Aurora, Colorado, USA. 115Center for Neurobehavioral Genetics, University of California, Los Angeles, Los Angeles, California, USA. 116Department of Psychiatry, University of Halle, Halle, Germany. 117Queensland Institute of Medical Research, Brisbane, Queensland, Australia. 118Department of Biomedical and Biological Sciences, Plymouth University, Plymouth, UK. 119Department of Psychiatry, University of California, San Diego, La Jolla, California, USA. 120Division of Tics, OCD and Related Disorders, Icahn School of Medicine at Mount Sinai, New York, New York, USA. 121Department of Psychiatry, Icahn School of Medicine at Mount Sinai, New York, New York, USA. 122Department of Psychiatry, University of Bonn, Bonn, Germany. 123Division of Biostatistics, University of Minnesota, Minneapolis, Minnesota, USA. 124Department of Psychiatry, Academic Medical Centre, University of Amsterdam, Amsterdam, the Netherlands. 125Center for Human Genetics Research, Vanderbilt University Medical Center, Nashville, Tennessee, USA. 126The Center for Applied Genomics, Division of Human Genetics, The Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA. 127Department of Pediatrics, University of Pennsylvania School of Medicine, Philadelphia, Pennsylvania, USA. 128Department of Psychiatry, School of Medicine, Stanford University, Stanford, California, USA. 129Department of Clinical and Developmental Psychology, Eberhard Karls University of Tübingen, Tübingen, Germany. 130Institute of Biological Psychiatry, Copenhagen University Hospital, Roskilde, Denmark. 131Brain and Mind Research Institute, University of Sydney, Sydney, New South Wales, Australia. 132Department of Psychiatry and Behavioral Sciences, Howard University College of Medicine, Washington, DC, USA. 133Department of Psychiatry, Erasmus Medical Center, Rotterdam, the Netherlands. 134Department of Psychology, University of Michigan, Ann Arbor, Michigan, USA. 135Bioinformatics Research Center, North Carolina State University, Raleigh, North Carolina, USA. 136Department of Psychology, University of Colorado, Boulder, Colorado, USA. 137Psychiatric Neurogenetics Section, Centre for Addiction and Mental Health, Toronto, Ontario, Canada. 138School of Medicine, University of St Andrews, St Andrews, UK. 139Department of Genetics, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina, USA. 140Division of Molecular Genome Analysis, German Cancer Research Center (DKFZ), Heidelberg, Germany. 141Department of Psychiatry, University of Pittsburgh School of Medicine, Pittsburgh, Pennsylvania, USA. 142Department of Psychiatry, Zilkha Neurogenetic Institute, Keck School of Medicine, University of Southern California, Los Angeles, California, USA. 143Wolfson Institute of Preventative Medicine, Queen Mary University of London, London, UK. 144Department of Psychiatry and Neuropsychology, Maastricht University Medical Centre, South Limburg Mental Health Research and Teaching Network, Maastricht, the Netherlands. 145Institute of Neuroscience and Physiology, University of Gothenburg, Gothenburg, Sweden. 146Centre National de Genotypage, Evry, France. 147Geisinger Health System, Autism and Developmental Medicine Institute, Danville, Pennsylvania, USA. 148Department of Psychiatry, Division of Research, The Zucker Hillside Hospital Division of the North Shore, Long Island Jewish Health System, Glen Oaks, New York, USA. 149Center for Psychiatric Neuroscience, The Feinstein Institute of Medical Research, Manhasset, New York, USA. 150Department of Psychiatry and Behavioral Science, Albert Einstein College of Medicine of Yeshiva University, Bronx, New York, USA. 151Division of Molecular Psychiatry, ADHD Clinical Research Unit, Department of Psychiatry, Psychosomatics and Psychotherapy, University of Würzburg, Würzburg, Germany. 152Department of Psychiatry and Psychology, School for Mental Health and Neuroscience (MHENS), Maastricht University, Maastricht, the Netherlands. 153Department of Psychiatry and Behavioral Sciences, Stanford University, Stanford, California, USA. 154Department of Human Genetics, University of Michigan, Ann Arbor, Michigan, USA. 155New York State Psychiatric Institute, Columbia University, New York, New York, USA. 156Department of Biostatistics, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina, USA. 157Department of Psychiatry, Academic Medical Centre University of Amsterdam, Amsterdam, the Netherlands. 158Department of Psychiatry, Institute of Human Genetics, University of Illinois at Chicago, Chicago, Illinois, USA. 159Department of Psychiatry and Biobehavioral Science, University of California, Los Angeles, Los Angeles, California, USA. 160Center for Autism and the Developing Brain, Weill Cornell Medical College, White Plains, New York, USA. 161Department of Neurology, David Geffen School of Medicine, University of California, Los Angeles, Los Angeles, California, USA. 162Center for Autism Research and Treatment, Semel Institute, David Geffen School of Medicine, University of California, Los Angeles, Los Angeles, California, USA. 163Department of Pharmacy and Biotechnology, University of Bologna, Bologna, Italy. 164Department of Psychiatry & Behavioral Sciences, Johns Hopkins University, Baltimore, Maryland, USA. 165Yale Center for Genome Analysis, Orange, Connecticut, USA. 166Sørlandet Hospital, Kristiansand, Norway. 167Child and Adolescent Psychiatry, Semel Institute, David Geffen School of Medicine, University of California, Los Angeles, Los Angeles, California, USA. 168Department of Psychiatry, University of Michigan, Ann Arbor, Michigan, USA. 169Molecular Medicine Centre, University of Edinburgh, Edinburgh, UK. 170National Institute of Mental Health, US National Institutes of Health, Bethesda, Maryland, USA. 171Department of Neurobehavioral Genetics, Trier University, Trier, Germany. 172Neuroepidemiology and Ageing Research, School of Public Health, Imperial College London, London, UK. 173Department of Psychiatry, First Psychiatric Clinic, Alexander University Hospital, Sofia, Bulgaria. 174Department of Developmental and Educational Psychology, University of Valencia, Valencia, Spain. 175Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford, UK. 176Office of the President, Tufts University, Medford, Massachusetts, USA. 177Department of Psychiatry, Yale University, New Haven, Connecticut, USA. 178Department of Psychiatry, St. Olavs Hospital, Trondheim, Norway. 179Department of Neuroscience, Norwegian University of Science and Technology, Trondheim, Norway. 180Department of Molecular Biology, Cell Biology and Biochemistry, Brown University, Providence, Rhode Island, USA. 181Department of Psychiatry and Human Behavior, Brown University, Providence, Rhode Island, USA. 182Biostatistics and Bioinformatics Unit, Cardiff University, Cardiff, UK. 183Neurosciences Centre of Excellence in Drug Discovery, GlaxoSmithKline Research and Development, Verona, Italy. 184Life & Brain Center, University of Bonn, Bonn, Germany. 185Child Study Center, Yale University, New Haven, Connecticut, USA. 186Department of Psychiatry, University of Pittsburgh, Pittsburgh, Pennsylvania, USA. 187Department of Human Genetics, University of Pittsburgh, Pittsburgh, Pennsylvania, USA. 188Department of Psychiatry, Groningen University Medical Center, Groningen, the Netherlands. 189Clinic for Child and Adolescent Psychiatry and Psychotherapy, University of Duisburg-Essen, Essen, Germany. 190Research and Clinical Training Department, Pediatric Hospital, Centro Hospitalar e Universitário Coimbra, Coimbra, Portugal. 191Department of Psychiatry, University Medical Center Utrecht, Utrecht, the Netherlands. 192Sanger Institute, Hinxton, Cambridge, UK. 193Program in Genetics and Genomic Biology, The Hospital for Sick Children, Toronto, Ontario, Canada. 194Dalla Lana School of Public Health, University of Toronto, Toronto, Ontario, Canada. 195Department of Psychiatry, VU University Medical Center, Amsterdam, the Netherlands. 196Academic Department of Psychiatry, University of Oxford, Oxford, UK. 197Department of Psychiatry, University of Würzburg, Würzburg, Germany. 198Psychiatric Genetics Unit, Vall d’Hebron Research Institute, Barcelona, Spain. 199Division of Biostatistics, Washington University School of Medicine, St. Louis, Missouri, USA. 200Department of Statistics, Carnegie Mellon University, Pittsburgh, Pennsylvania, USA. 201Department of Experimental Clinical & Health Psychology, Ghent University, Ghent, Belgium. 202Child and Adolescent Psychiatry, University Medicine Göttingen, Göttingen, Germany. 203Department of Neurology and Neurosurgery, McGill University, Montreal, Quebec, Canada. 204Department of Genetics, Yale University, New Haven, Connecticut, USA. 205Program on Neurogenetics, Yale University, New Haven, Connecticut, USA. 206Department of Psychiatry, Maine Medical Center, Portland, Maine, USA. 207Department of Psychiatry, Harvard Medical School, Boston, Massachusetts, USA. 208Department of Clinical Neuropsychology, VU University, Amsterdam, the Netherlands. 209Department of Psychiatry and Behavioral Science, Stanford University School of Medicine, Palo Alto, California, USA. 210Rush Ambulatory Behavioral Health, Rush University Medical Center, Chicago, Illinois, USA. 211Department of Pathology and Laboratory Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA. 212The Centre for Applied Genomics, The Hospital for Sick Children, Toronto, Ontario, Canada. 213The Scripps Research Institute, La Jolla, California, USA. 214Department of Psychiatry & Psychotherapy, University of Göttingen, Göttingen, Germany. 215Psychiatric Center Nordbaden, Wiesloch, Germany. 216Division of Cancer Epidemiology and Genetics, National Cancer Institute, Bethesda, Maryland, USA. 217Department of Psychiatry and Behavioral Sciences, University of Washington, Seattle, Washington, USA. 218Mayo Clinic, Rochester, Minnesota, USA. 219Developmental Brain & Behaviour Laboratory, Academic Unit of Psychology, University of Southampton, Southampton, UK. 220Institute of Medical Sciences, University of Aberdeen, Foresterhill, Aberdeen, UK. 221Research Department, Federal Institute for Drugs and Medical Devices (BfArM), Bonn, Germany. 222Research Unit of Child and Adolescent Psychiatry, Aalborg University Hospital, Aalborg, Denmark. 223Clinical Psychology and Epidemiology, University of Basel, Basel, Switzerland. 224Department of Child and Adolescent Psychiatry, University of Zurich, Zurich, Switzerland. 225Molecular Neuropsychiatry and Development Laboratory, Centre for Addiction and Mental Health, Toronto, Ontario, Canada. 226Department of Psychiatry, Columbia University, New York, New York, USA. 227Vanderbilt Brain Institute, Vanderbilt University, Nashville, Tennessee, USA. 228Department of Psychiatry, University of Toronto, Toronto, Ontario, Canada. 229Neurosciences and Mental Health Program, Hospital for Sick Children, Toronto, Ontario, Canada. 230Centre for Addiction and Mental Health, Toronto, Ontario, Canada. 231Oxford Health NHS Foundation Trust, Marlborough House Secure Unit, Milton Keynes, UK. 232Center for Biomarker Research and Personalized Medicine, Virginia Commonwealth University, Richmond, Virginia, USA. 233Instituto Nacional de Saude Dr Ricardo Jorge, Lisbon, Portugal. 234BioFIG—Center for Biodiversity, Functional and Integrative Genomics, Campus da FCUL, Campo Grande, Lisbon, Portugal. 235Instituto Gulbenkian de Cîencia, Lisbon, Portugal. 236Battelle Center for Mathematical Medicine, Nationwide Children’s Hospital, Columbus, Ohio, USA. 237Howard Hughes Medical Institute, Children’s Hospital Boston, Boston, Massachusetts, USA. 238Division of Genetics, Children’s Hospital Boston, Boston, Massachusetts, USA. 239Department of Neurology, Harvard Medical School Center for Life Sciences, Boston, Massachusetts, USA. 240Department of Pediatrics, Harvard Medical School Center for Life Sciences, Boston, Massachusetts, USA. 241Columbia University College of Physicians and Surgeons, New York, New York, USA. 242Faculty of Health and Medical Science, University of Copenhagen, Copenhagen, Denmark. 243Institute of Medical Biometry, University of Bonn, Bonn, Germany. 244Department of Biostatistics, University of Washington, Seattle, Washington, USA. 245Department of Medicine, University of Washington, Seattle, Washington, USA. 246Centre for Affective Disorders, Institute of Psychiatry, King’s College London, London, UK. 247Division of Genetics, Children’s Hospital Boston, Harvard Medical School, Boston, Massachusetts, USA. 248Department of Mental Health, Johns Hopkins University, Baltimore, Maryland, USA. 249Department of Psychiatry, Leiden University Medical Center, Leiden, the Netherlands. 250International Inflammatory Bowel Disease Genetics Consortium (IIBDGC). 251Department of Psychiatry, Special Treatment and Evaluation Program (STEP), Veterans Affairs San Diego Healthcare System, San Diego, California, USA. 252Department of Human and Molecular Genetics, Virginia Commonwealth University, Richmond, Virginia, USA. 253Department of Psychiatry, Virginia Commonwealth University, Richmond, Virginia, USA. 254Department of Biomedical Informatics, Vanderbilt University School of Medicine, Nashville, Tennessee, USA. 255Department of Psychiatry, Vanderbilt University School of Medicine, Nashville, Tennessee, USA. 256Deceased. 257These authors contributed equally to this work. Correspondence should be addressed to G.B. (gerome.breen@kcl.ac.uk) or P.A.H. (holmanspa@cardiff.ac.uk).
Footnotes
Note: Any Supplementary Information and Source Data files are available in the online version of the paper.
AUTHOR CONTRIBUTIONS
Project conception: G.B., P.F.S., P.A.H. Analysis: C.O’D., P.H.L., P.A.H., G.B., S.R., L.R., L.D., S.N. Writing of the manuscript: G.B., C.O’D. P.A.H., P.H.L., L.R., L.D., N.P. Quality control for PGC data: S. Ripke and B.M.N. Revisions to the manuscript: G.B., D.H.G., C.O’D., L.R., P.H.L., N.P. PGC Network and Pathway Analysis Workgroup: S.R., B.M.N., S.M.P., D.H.G., P.A.H., P.H.L., M.M., C.O’D., D.P., L.R., L.D., P.F.S., J.W.S., N.R.W., Z.Z. PGC Workgroup Chairs: M.J.D. (analysis), S.V.F. (ADHD), M.J.D. and B.D. (co-chairs ASD), G.B. and P.A.H. (Network and Pathway Analysis subgroup), J.K. and P. Sklar (co-chairs bipolar disorder), P.F.S. (major depressive disorder), M.C.O’D. (schizophrenia) and J.W.S. and K.S.K. (co-chairs cross-disorder group). Collection, genotyping and analysis for PGC Working Groups. PGC ADHD Working Group: B.M.N., S.V.F., A.T., R.A., P.A., T. Banaschewski, M. Bayés, J.B., J.K.B., M.C., B.C., J.C., A.E.D., R.P.E., J.E., B.F., C.M.F., L. Kent, J.K., K.-P.L., S.K.L., J.M., J.J.M., S.E.M., J.M.S., A. Miranda, S.F.N., R.D.O., J.A.R.-Q., A. Reif, M. Ribasés, H.R., A. Rothenberger, J.A.S., R.S., S.L. Smalley, E.J.S.S.-B., H.-C.S., A.A.T. and N.W. PGC ASD Working Group: R.A., D.E.A., A.J.B., A.B., C.B., J.D. Buxbaum, A. Chakravarti, E.H.C., H.C., M.L.C., G.D., E.D., S.E., E.F., C.M.F., L. Gallagher, D.H.G., M. Gill, D.E.G., J.L.H., H.H., J.H., V.H., S.M.K., L. Klei, D.H. Ledbetter, C. Lord, J.K.L., E.M., S.M.M., C.L.M., W.M.M., A.P.M., D.M.-D.-L., E.M.M., M. Murtha, G.O., A.P., J.R.P., A.D.P., M.A.P.-V., J. Piven, F.P., K. Rehnström, K. Roeder, G.R., S.J.S., S.L. Santangelo, G.D.S., S.W.S., M. State, J.S. Sutcliffe, P. Szatmari, A.M.V., V.J.V., C.A.W., T.H.W., E.M.W., A.J.W., T.W.Y., B.D. and M.J.D. PGC BPD Working Group: S.M.P., D.A., H.A., O.A.A., A.A., L.B., J.A.B., J.D. Barchas, T.B.B., N.B., M. Bauer, F.B., S.E.B., W.B., D.H.R.B., C.S.B., M. Boehnke, G.B., R. Breuer, W.E.B., W.F.B., S. Caesar, K. Chambert, S. Cichon, D.A.C., A. Corvin, W.H.C., D.W.C., R.D., F. Degenhardt, S. Djurovic, F. Dudbridge, H.J.E., B.E., A.E.F., I.N.F., M. Flickinger, T.F., J.F., C.F., L.F., E.S.G., M. Gill, K.G.-S., E.K.G., T.A.G., D.G., W.G., H.G., M.L.H., M. Hautzinger, S. Herms, M. Hipolito, P.A.H., C.M.H., S.J., E.G.J., I.J., L.J., R. Kandaswamy, J.L.K., G.K.K., D.L.K., P.K., M. Landén, N.L., M. Lathrop, J. Lawrence, W.B.L., M. Leboyer, P.H.L., J. Li, P.L., D.-Y.L., C. Liu, F.W.L., S.L., P.B.M., W.M., N.G.M., M. Mattheisen, K.M., M. Mattingsdal, K.A.M., P.M., M.G.M., A. McIntosh, R.M., A.W.M., F.J.M., A. McQuillin, S.M., I.M., F.M., G.W.M., J.L.M., G.M., D.W.M., V. Moskvina, P.M., T.W.M., W.J.M., B.M.-M., R.M.M., C.M.N., I.N., V.N., M.M.N., J.I.N., E.A.N., C.O., U.O., M.J.O., B.S.P., J.B.P., P.P., E.M.Q., S. Raychaudhuri, A. Reif, J.P.R., M. Rietschel, D. Ruderfer, M. Schalling, A.F.S., W.A.S., N.J.S., T.G.S., J. Schumacher, M. Schwarz, E.S., L.J.S., P.D.S., E.N.S., D.S.C., M. Steffens, J.S. Strauss, J. Strohmaier, S.S., R.C.T., F.T., J.T., J.B.V., S.J.W., T.F.W., S.H.W., W.X., A.H.Y., P.P.Z., P.Z., S. Zöllner, J.R.K., P. Sklar, M.J.D., M.C.O. and N.C. PGC MDD Working Group: M.R.B., T. Bettecken, E.B.B., D.H.R.B., D.I.B., G.B., R. Breuer, S. Cichon, W.H.C., I.W.C., D. Czamara, E.J.D.G., F. Degenhardt, A.E.F., J.F., S.D.G., M. Gross, S.P.H., A.C.H., A.K.H., S. Herms, I.B.H., F.H., W.J.H., S. Hoefels, J.-J.H., M.I., I.J., L.J., J.-Y.T., J.A.K., M.A.K., A.K., W.B.L., D.F.L., C.M.L., D.-Y.L., S.L., D.J.M., P.A.F.M., W.M., N.G.M., M. Mattheisen, P.J.M., P.M., A. McIntosh, A.W.M., C.M.M., L.M., G.W.M., P.M., B.M.-M., W.A.N., M.M.N., D.R.N., B.W.P., M.L.P., J.B.P., M. Rietschel, W.A.S., T.G.S., J. Shi, S.I.S., S.L. Slager, J.H.S., M. Steffens, F.T., J.T., M.U., E.J.C.G.v.d.O., G.V.G., M.M.W., G.W., F.G.Z., P.F.S. and N.R.W. PGC SCZ Working Group: S. Ripke, B.M.N., S.M.P., B.J.M., I.A., F.A., O.A.A., M.H.A., N.B., D.W.B., D.H.R.B., R. Bruggeman, N.G.B., W.F.B., W.C., R.M.C., K. Choudhury, S. Cichon, C.R.C., P.C., A. Corvin, D. Curtis, S. Datta, S. Djurovic, G.J.D., J.D., F. Dudbridge, A.F., R.F., N.B.F., M. Friedl, P.V.G., L. Georgieva, I.G., M. Gill, H.G., L.D.H., M.L.H., T.F.H., A.M.H., P.A.H., C.M.H., A.I., A.K.K., R.S.K., M.C.K., E.K., Y.K., G.K.K., B.K., L. Krabbendam, R. Krasucki, J. Lawrence, P.H.L., T.L., D.F.L., J.A.L., D.-Y.L., D.H. Linszen, P.K.E.M., W.M., A.K.M., M. Mattheisen, M. Mattingsdal, S.M., S.A.M., A. McIntosh, A. McQuillin, H.M., I.M., V. Milanova, D.W.M., V. Moskvina, I.M.-G., M.M.N., C.O., A.O., L.O., R.A.O., M.J.O., C.N.P., M.T.P., B.S.P., J. Pimm, D.P., V.P., D.J.Q., H.B.R., M. Rietschel, L.R., D. Ruderfer, D. Rujescu, A.R.S., T.G.S., J. Shi, J.M.S., D.S.C., T.S.S., S.T., J.V.O., P.M.V., T.W., S. Zammit, P. Sklar, M.J.D., M.C.O., N.C., P.F.S. and K.S.K. PGC Cross-Disorder Group Working Group: S.H.L., S. Ripke, B.M.N., S.M.P., R.H.P., A.T., A.F., M.C.N., J.I.N., B.W.P., M. Rietschel, T.G.S., N.C., S.L. Santangelo, P.F.S., J.W.S., K.S.K. and N.R.W. PGC Analysis Working Group: S.H.L., S. Ripke, B.M.N., S.M.P., V.A., E.M.B., P.H.L., S.E.M., M.C.N., D.P., G.B., M.J.D. and N.R.W.
COMPETING FINANCIAL INTERESTS
The authors declare competing financial interests: details are available in the online version of the paper.
References
- 1.Vos T, et al. Years lived with disability (YLDs) for 1160 sequelae of 289 diseases and injuries 1990–2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet. 2012;380:2163–2196. doi: 10.1016/S0140-6736(12)61729-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Lee SH, et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 2013;45:984–994. doi: 10.1038/ng.2711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Cross-Disorder Group of the Psychiatric Genomics Consortium et al. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet. 2013;381:1371–1379. doi: 10.1016/S0140-6736(12)62129-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Mirnics K, Middleton FA, Marquez A, Lewis DA, Levitt P. Molecular characterization of schizophrenia viewed by microarray analysis of gene expression in prefrontal cortex. Neuron. 2000;28:53–67. doi: 10.1016/s0896-6273(00)00085-4. [DOI] [PubMed] [Google Scholar]
- 5.Nam D, Kim SY. Gene-set approach for expression pattern analysis. Brief. Bioinform. 2008;9:189–197. doi: 10.1093/bib/bbn001. [DOI] [PubMed] [Google Scholar]
- 6.Ackermann M, Strimmer K. A general modular framework for gene set enrichment analysis. BMC Bioinformatics. 2009;10:47. doi: 10.1186/1471-2105-10-47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Baranzini SE, et al. Pathway and network-based analysis of genome-wide association studies in multiple sclerosis. Hum. Mol. Genet. 2009;18:2078–2090. doi: 10.1093/hmg/ddp120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Wang K, et al. Diverse genome-wide association studies associate the IL12/IL23 pathway with Crohn Disease. Am. J. Hum. Genet. 2009;84:399–405. doi: 10.1016/j.ajhg.2009.01.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Holmans P, et al. Gene ontology analysis of GWA study data sets provides insights into the biology of bipolar disorder. Am. J. Hum. Genet. 2009;85:13–24. doi: 10.1016/j.ajhg.2009.05.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.O’Dushlaine C, et al. Molecular pathways involved in neuronal cell adhesion and membrane scaffolding contribute to schizophrenia and bipolar disorder susceptibility. Mol. Psychiatry. 2011;16:286–292. doi: 10.1038/mp.2010.7. [DOI] [PubMed] [Google Scholar]
- 11.Psychiatric GWAS Consortium Bipolar Disorder Working Group. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4 . Nat. Genet. 2011;43:977–983. doi: 10.1038/ng.943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Sullivan PF. The psychiatric GWAS consortium: big science comes to psychiatry. Neuron. 2010;68:182–186. doi: 10.1016/j.neuron.2010.10.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Neale BM, et al. Meta-analysis of genome-wide association studies of attention-deficit/hyperactivity disorder. J. Am. Acad. Child Adolesc. Psychiatry. 2010;49:884–897. doi: 10.1016/j.jaac.2010.06.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Schizophrenia Psychiatric Genome-Wide Association Study Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 2011;43:969–976. doi: 10.1038/ng.940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Major Depressive Disorder Working Group of the PGC et al. A mega-analysis of genome-wide association studies for major depressive disorder. Mol. Psychiatry. 2013;18:497–511. doi: 10.1038/mp.2012.21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Brown MB. A method for combining non-independent one-sided tests of significance. Biometrics. 1975;31:987–992. [Google Scholar]
- 17.McLaren PJ, et al. Association study of common genetic variants and HIV-1 acquisition in 6,300 infected cases and 7,200 controls. PLoS Pathog. 2013;9:e1003515. doi: 10.1371/journal.ppat.1003515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kang HJ, et al. Spatio-temporal transcriptome of the human brain. Nature. 2011;478:483–489. doi: 10.1038/nature10523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Gong S, et al. A gene expression atlas of the central nervous system based on bacterial artificial chromosomes. Nature. 2003;425:917–925. doi: 10.1038/nature02033. [DOI] [PubMed] [Google Scholar]
- 20.Xu X, Wells AB, O′Brien DR, Nehorai A, Dougherty JD. Cell type-specific expression analysis to identify putative cellular mechanisms for neurogenetic disorders. J. Neurosci. 2014;34:1420–1431. doi: 10.1523/JNEUROSCI.4488-13.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Doyle JP, et al. Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell. 2008;135:749–762. doi: 10.1016/j.cell.2008.10.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Schizophrenia Working Group of the Psychiatric Genomics. C. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–427. doi: 10.1038/nature13595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kirov G, et al. De novo CNV analysis implicates specific abnormalities of postsynaptic signalling complexes in the pathogenesis of schizophrenia. Mol. Psychiatry. 2012;17:142–153. doi: 10.1038/mp.2011.154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Purcell SM, et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature. 2014;506:185–190. doi: 10.1038/nature12975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Voineagu I, et al. Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature. 2011;474:380–384. doi: 10.1038/nature10110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ting JT, Peça J, Feng G. Functional consequences of mutations in postsynaptic scaffolding proteins and relevance to psychiatric disorders. Annu. Rev. Neurosci. 2012;35:49–71. doi: 10.1146/annurev-neuro-062111-150442. [DOI] [PubMed] [Google Scholar]
- 27.Purcell SM, et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature. 2014;506:185–190. doi: 10.1038/nature12975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Fromer M, et al. De novo mutations in schizophrenia implicate synaptic networks. Nature. 2014;506:179–184. doi: 10.1038/nature12929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Cruceanu C, et al. H3K4 tri-methylation in synapsin genes leads to different expression patterns in bipolar disorder and major depression. Int. J. Neuropsychopharmacol. 2013;16:289–99. doi: 10.1017/S1461145712000363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jarome TJ, Lubin FD. Histone lysine methylation: critical regulator of memory and behavior. Rev. Neurosci. 2013;24:375–387. doi: 10.1515/revneuro-2013-0008. [DOI] [PubMed] [Google Scholar]
- 31.Ben-David E, Shifman S. Combined analysis of exome sequencing points toward a major role for transcription regulation during brain development in autism. Mol. Psychiatry. 2013;18:1054–1056. doi: 10.1038/mp.2012.148. [DOI] [PubMed] [Google Scholar]
- 32.Parikshak NN, et al. Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell. 2013;155:1008–1021. doi: 10.1016/j.cell.2013.10.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ronemus M, Iossifov I, Levy D, Wigler M. The role of de novo mutations in the genetics of autism spectrum disorders. Nat. Rev. Genet. 2014;15:133–141. doi: 10.1038/nrg3585. [DOI] [PubMed] [Google Scholar]
- 34.Bauman AL, et al. Cocaine and antidepressant-sensitive biogenic amine transporters exist in regulated complexes with protein phosphatase 2A. J. Neurosci. 2000;20:7571–7578. doi: 10.1523/JNEUROSCI.20-20-07571.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Dieperink E, Willenbring M, Ho SB. Neuropsychiatric symptoms associated with hepatitis C and interferon alpha: a review. Am. J. Psychiatry. 2000;157:867–876. doi: 10.1176/appi.ajp.157.6.867. [DOI] [PubMed] [Google Scholar]
- 36.Susser E, St Clair D, He L. Latent effects of prenatal malnutrition on adult health: the example of schizophrenia. Ann. NY Acad. Sci. 2008;1136:185–192. doi: 10.1196/annals.1425.024. [DOI] [PubMed] [Google Scholar]
- 37.Heijmans BT, Tobi EW, Lumey LH, Slagboom PE. The epigenome: archive of the prenatal environment. Epigenetics. 2009;4:526–531. doi: 10.4161/epi.4.8.10265. [DOI] [PubMed] [Google Scholar]
- 38.Brykczynska U, et al. Repressive and active histone methylation mark distinct promoters in human and mouse spermatozoa. Nat. Struct. Mol. Biol. 2010;17:679–687. doi: 10.1038/nsmb.1821. [DOI] [PubMed] [Google Scholar]
- 39.Huang HS, et al. Prefrontal dysfunction in schizophrenia involves mixed-lineage leukemia 1-regulated histone methylation at GABAergic gene promoters. J. Neurosci. 2007;27:11254–62. doi: 10.1523/JNEUROSCI.3272-07.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yauch RL, Settleman J. Recent advances in pathway-targeted cancer drug therapies emerging from cancer genome analysis. Curr. Opin. Genet. Dev. 2012;22:45–49. doi: 10.1016/j.gde.2012.01.003. [DOI] [PubMed] [Google Scholar]
- 41.Flicek P, et al. Ensembl 2013. Nucleic Acids Res. 2013;41:D48–D55. doi: 10.1093/nar/gks1236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Maston GA, Evans SK, Green MR. Transcriptional regulatory elements in the human genome. Annu. Rev. Genomics Hum. Genet. 2006;7:29–59. doi: 10.1146/annurev.genom.7.080505.115623. [DOI] [PubMed] [Google Scholar]
- 43.Pedroso I, et al. Common genetic variants and gene-expression changes associated with bipolar disorder are over-represented in brain signaling pathway genes. Biol. Psychiatry. 2012;72:311–317. doi: 10.1016/j.biopsych.2011.12.031. [DOI] [PubMed] [Google Scholar]
- 44.Moskvina V, et al. Evaluation of an approximation method for assessment of overall significance of multiple-dependent tests in a genomewide association study. Genet. Epidemiol. 2011;35:861–866. doi: 10.1002/gepi.20636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Lee PH, O’Dushlaine C, Thomas B, Purcell SM. INRICH: interval-based enrichment analysis for genome-wide association studies. Bioinformatics. 2012;28:1797–1799. doi: 10.1093/bioinformatics/bts191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Segrè AV, Groop L, Mootha VK, Daly MJ, Altshuler D. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 2010;6:e1001058. doi: 10.1371/journal.pgen.1001058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Purcell S, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007;81:559–575. doi: 10.1086/519795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Storey JD, Tibshirani R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA. 2003;100:9440–9445. doi: 10.1073/pnas.1530509100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Brown MB. A method for combining non-independent, one-sided tests of significance. Biometrics. 1975;31:978–992. [Google Scholar]
- 50.Zhang B, Horvath S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. 2005;4 doi: 10.2202/1544-6115.1128. Article17. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.