Abstract
Autonomous circadian oscillations arise from transcriptional-translational feedback loops of core clock components. The period of a circadian oscillator is relatively insensitive to changes in nutrients (e.g., glucose), which is referred to as “nutrient compensation”. Recently, a transcription repressor, CSP-1, was identified as a component of the circadian system in Neurospora crassa. The transcription of csp-1 is under the circadian regulation. Intriguingly, CSP-1 represses the circadian transcription factor, WC-1, forming a negative feedback loop that can influence the core oscillator. This feedback mechanism is suggested to maintain the circadian period in a wide range of glucose concentrations. In this report, we constructed a mathematical model of the Neurospora circadian clock incorporating the above WC-1/CSP-1 feedback loop, and investigated molecular mechanisms of glucose compensation. Our model shows that glucose compensation exists within a narrow range of parameter space where the activation rates of csp-1 and wc-1 are balanced with each other, and simulates loss of glucose compensation in csp-1 mutants. More importantly, we experimentally validated rhythmic oscillations of the wc-1 gene expression and loss of glucose compensation in the wc-1ov mutant as predicted in the model. Furthermore, our stochastic simulations demonstrate that the CSP-1-dependent negative feedback loop functions in glucose compensation, but does not enhance the overall robustness of oscillations against molecular noise. Our work highlights predictive modeling of circadian clock machinery and experimental validations employing Neurospora and brings a deeper understanding of molecular mechanisms of glucose compensation.
Introduction
Circadian rhythms play a vital role in an organism’s functions by anticipating daily changes in its environment. They influence various facets of organisms’ activity including physiology, cell cycle, and metabolism. In the last two decades, experiments revealed that the core clock components used to construct the circadian oscillator are assembled as transcription-translation feedback loops (1,2) whose activity is modified through posttranslational modifications of the core components (3,4).
In Neurospora crassa, circadian timekeeping is sustained through interlocked negative and positive feedback loops. The WHITE COLLAR complex (WCC), a heterodimer of the zinc-finger proteins WC-1 and WC-2, activates the transcription of the frequency (frq) gene (5–7). Then, translated FREQUENCY (FRQ) protein binds to FRQ-interacting RNA Helicase and CK1 (3,8), translocates into the nucleus, and suppresses its own transcription by inactivating WCC, closing the negative feedback loop (9–11). Simultaneously, FRQ promotes WC-1 accumulation in the cytoplasm, increasing the level of WCC forming a positive feedback loop (12,13).
Recently, the circadian clock-regulated developmental regulator, conidial separation-1 (csp-1) (14), was shown to be a global transcription repressor under the direct control of WCC, and was found to regulate expression of various metabolic and other genes (15). CSP-1 was shown to repress the expression of wc-1 in a glucose-dependent manner by binding at the respective promoter (16) and forming an additional negative feedback loop. These intriguing findings illuminate a novel function of CSP-1 as an important regulator of the Neurospora core clock oscillator resembling that of REV-ERB-α/β in the mammalian circadian clock (17–19). Further experiments suggested that negative feedback via CSP-1 maintains relatively similar abundance of WC-1 in a wide range of glucose concentrations in the growth media. This results in a relatively constant circadian period as a function of glucose concentrations, which is referred to as “glucose compensation”. In contrast, the glucose compensation was disrupted in csp-1ko with decreasing period with increasing glucose concentration (16).
In this article, we explore a Neurospora circadian clock model that includes CSP-1 as a newly identified element of the circadian system, and investigate molecular mechanisms for glucose compensation. Our model simulates glucose compensation of period by balancing the activation rates of csp-1 and wc-1 transcription, and loss of glucose compensation in csp-1 mutants. Moreover, we predict and experimentally validate loss of glucose compensation in the wc-1 overexpression (wc-1ov) mutant. The model also accurately reproduces changes in clock period observed in previous experiments, and differential expression of FRQ and WC-1 in the nucleus versus cytoplasm (4,20).
Finally, it has been shown that the interlocked dual negative feedback loop increases oscillatory domain and possibly enhances the robustness of autonomous oscillations (21). We investigate robustness of the system against molecular stochasticity in the presence or absence of CSP-1 and show that a CSP-1-mediated negative feedback loop does not enhance the overall robustness of the system against molecular noise.
Materials and Methods
Mathematical model of Neurospora circadian clock with CSP-1
The wiring diagram of the core oscillator and other ancillary components pertinent to this discussion is shown in Fig. 1. In the nucleus, nuclear WC-1 (WC-1n) activates frq transcription with the rate constant k1. FRQ protein is translated from frq mRNA in the cytosol and is translocated into the nucleus (processes with the rate constants k2 and k3, respectively). frq mRNA, cytosolic FRQ (FRQc), and nuclear FRQ (FRQn) degrade with the rate constants k4, k5, and k6, respectively. The expression of wc-1 is modeled with a maximum rate constant k7, and the wc-1 expression is modulated by the negative feedback from the global transcription repressor, CSP-1. Cytosolic WC-1 (WC-1c) is translated and degraded with the rate constants k8 and k10, respectively. We assume that the accumulation of WC-1c is positively regulated by FRQc (12). Then, WC-1c protein is either degraded (k11) or translocated into the nucleus (k9). We do not consider WC-2 in the model because WC-2 is present in excess of WC-1 and its concentration is relatively constant at all circadian times (6,7). Therefore, we consider the WCC complex to be represented in the model by WC-1n. Finally, WC-1n starts frq transcription and then either slowly degrades (k12) or is quickly inactivated (k13/k14) and degraded (k15) via complex-formation with FRQn closing the negative feedback loop.
Figure 1.
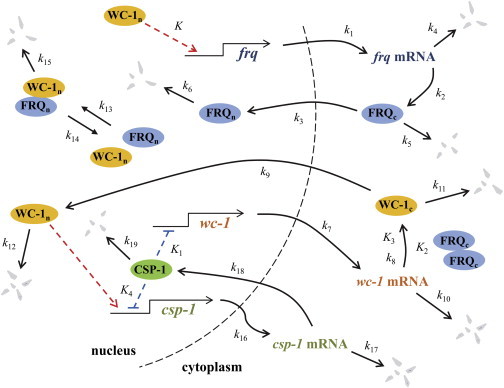
Neurospora clock model diagram with the global transcription repressor CSP-1. Transcriptional activator WC-1 starts transcription of the core clock gene frq whereas FRQ protein represses its own gene transcription in a negative feedback loop that promotes sustained circadian oscillations of core clock proteins. Transcriptional repressor CSP-1 inhibits wc-1 gene expression forming another negative feedback loop in the model. (Dashed arrows) Activation, (dashed lines with blunt ends) repression via binding at the respective promoters. To see this figure in color, go online.
Based on recent experimental findings (15,16), we extended the Neurospora circadian clock model (22) to include an additional negative feedback loop formed by the aforementioned transcriptional repressor, CSP-1. WC-1n starts transcription of csp-1 (k16), and the csp-1 mRNA is degraded with the rate k17. CSP-1 is translated and degraded with the rates k18 and k19, respectively. We used the rate of degradation of CSP-1 from the previous experimental measurements (15). In the nucleus, CSP-1 inhibits the expression of wc-1 as well as its own expression, as reported in Sancar et al. (15). In this study, we assume that the binding of CSP-1 to the wc-1 promoter can be described by the Hill-type function similar to the one used to model inhibition of its own transcription in Sancar et al. (15).
The model includes the following rate equations:
The parameter values are given in Table 1 (23–25). Parameters k01, k03, and k04 simulate additional transcription of corresponding genes to reproduce overexpression experiments with an inducible system, and have dimensions of a.u. h−1 and are initially set to zero. Concentrations in arbitrary units (a.u.) reflect the number of molecules per unit volume. Experimentally measured rate constants are indicated in Table 1. The rest of the rate constants have been chosen to fit the model to experimental data simulating the abundance of total and nuclear WC-1 and FRQ, and the period change observed in frq, wc-1, and csp-1 overexpression experiments in wild-type (WT) and mutant strains.
Table 1.
Parameter values for the Neurospora clock model
| Parameter | Value | Dimension | Parameter | Value | Dimension |
|---|---|---|---|---|---|
| k1 | 1.8 | a.u. h−1 | k13 | 50 | (a.u.)−1 h−1 |
| k2 | 1.8 | h−1 | k14 | 1 | h−1 |
| k3 | 0.05 | h−1 | k15 | 5 | h−1 |
| k4 (23) | 0.23 | h−1 | k16 | 0.12 | h−1 |
| k5 (24) | 0.27 | h−1 | k17 | 1.4 | h−1 |
| k6 (24) | 0.27 | h−1 | k18 | 50 | h−1 |
| k7 | 0.5 | a.u. h−1 | k19 (15) | 1.4 | h−1 |
| k8 | 1 | a.u. h−1 | K | 1.25 | (a.u.)6 |
| k9 | 40 | h−1 | K1 | 3 | a.u. |
| k10 (25) | 0.1 | h−1 | K2 | 1 | (a.u.)2 |
| k11 | 0.05 | h−1 | K3 | 10 | a.u. |
| k12 | 0.02 | h−1 | K4 | 3 | a.u. |
Experimentally measured rate constants are followed by a reference.
Justifications of the model
In this study, we use a relatively high cooperativity (n = 6) of WC-1n binding at the frq promoter. This high cooperativity is necessary to increase nonlinearity in our simple model to simulate the existing experimental data on changes in period in clock overexpression mutants (see Fig. 3 and Text S1 in the Supporting Material for more details). In our model, we do not include posttranslational modification such as progressive phosphorylation of FRQ (26) or the coiled-coil domain-mediated FRQ-FRQ interaction (4) to reduce the complexity of the model while simulating existing molecular phenotypes. If they are included in the model, they may increase overall nonlinearity of the system and allow for a lower Hill coefficient of WC-1n activation of frq transcription. However, this would make the model significantly more complex and, therefore, we do not include them in this simple model. See also Text S1 in the Supporting Material for further discussion. Also, we assume that the accumulation of WC-1c shows saturation kinetics with reference to wc-1 mRNA. This saturation behavior, although not reported experimentally, may occur due to FRQ-promoted accumulation of WC-1c as we showed before in Hong et al. (22) and allows the model to reproduce experimental data on wc-1 overexpression (see Fig. 3, B and C and Text S1 for more details).
Figure 3.
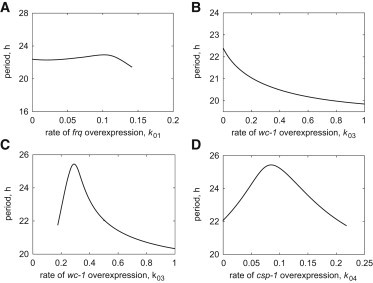
The model reproduces overexpression phenotypes of frq, wc-1, and csp-1. (A) Clock period as a function of the rate of frq overexpression, k01 in wild-type genetic background. (B) Clock period as a function of wc-1 overexpression rate, k03, in wild-type genetic background. (C) Clock period as a function of wc-1 overexpression rate, k03, in wc-1ko genetic background (the wc-1 expression rate constant k7 = 0). (D) Clock period as a function of k04, simulating the rate of csp-1 overexpression in qa-2 promoter knock-in experiment (the csp-1 expression rate constant k16 = 0).
Lastly, in our stochastic simulations, we used standard Michaelis-Menten and Hill-type functions (Table S1 in the Supporting Material) derived via quasi-steady-state approximation instead of a large number of elementary reactions. This approach, resulting in a compact model description, was studied before in a circadian clock model (27) and other systems (28) and was shown to produce satisfactory agreement between detailed and compact models. The compact models are favorable because the actual molecular mechanisms that comprise Hill-type and Michaelis-Menten functions are usually unknown. However, recent studies have found that stochastic simulations of the compact model are not always accurate (29–31). We confirm validity of our stochastic simulation results produced with the Gillespie algorithm by rewriting the model as Langevin-type equations with additive or multiplicative noise (32) and obtaining qualitatively similar results (see Figs. 7 and S9).
Figure 7.
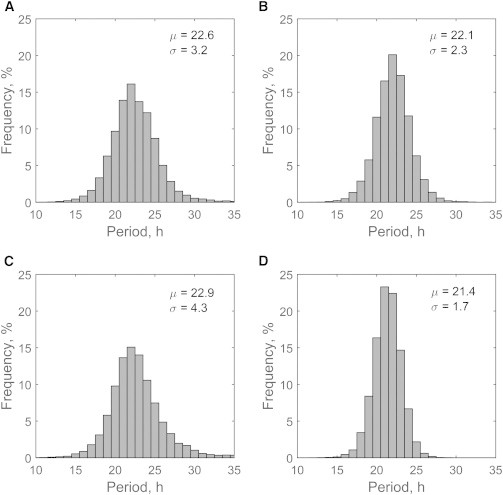
Histograms of period distributions obtained by stochastic simulations with strong noise. Period distribution in the presence of strong noise for the WT (A and C) and csp-1ko (B and D) in low and high glucose, correspondingly. The histograms of periods of 10,000 cycles were computed from 100 simulation runs with 100 successive cycles with the volume factor Ω = 100. The period was determined as the time interval separating two successive peaks of FRQc. The mean value (μ) and standard deviation (σ) of the period (in hours) is shown in each histogram. Parameter values are as described in Materials and Methods, except (B) k16 = 0, (C) k7 = 0.6, k16 = 0.57 and (D) k7 = 0.6, k16 = 0.
Computer simulations
We used XPPAUT software (Bard Ermentrout, University of Pittsburgh, http://www.math.pitt.edu/bard/xpp/xpp.html) to obtain numerical solutions, bifurcation, and period diagrams of the model equations.
Stochastic simulations
Stochastic simulations were carried out in MATLAB (The MathWorks, Natick, MA) by utilizing the Gillespie algorithm (33,34). We assigned the probability of occurrence of a particular reaction to each term of the kinetic model as shown in Table S1. The Gillespie algorithm then randomly chooses the reaction that occurs at each time step according to its probability and the time interval to the next reaction step. The number of molecules and the reaction probabilities are then updated at each time step according to Table S1. Parameter Ω in the Gillespie method allows modulation of the strength of noise in the stochastic model formulation by controlling the number of molecules that are present in the system. Each simulation was run for 3000 h and we removed the first 500 h to allow for transients to subside.
Strains
Strains used for the experiments are a clock wild-type ras-1bd (328-4), wc-1-luc::csr-1, and wc-1ov;frq-luc::his-3. The strain wc-1-luc::csr-1 was created by integrating the wc-1 promoter with a codon-optimized firefly luciferase into the csr-1 locus as previously described in Chen et al. (35) in a clock wild-type ras-1bd (328-4). The strain wc-1ov was created by integrating a wc-1 open reading frame with its endogenous promoter into the csr-1 locus as previously described in Chen et al. (35) in the strain containing frq-luciferase reporter (36).
Western blot
Neurospora were grown in liquid culture media containing Vogel’s medium (pH 5.8) with either 0.1% or 0.3% glucose, 0.5% arginine, 50 ng/mL biotin, and harvested at indicated time points. Immunoblot analysis was performed as previously described in Garceau et al. (26) except that we used 1) 20 μg of protein per lane, 2) 6.5% sodium dodecyl sulfate-polyacrylamide gel electrophoresis, 3) anti-WC-1 or anti-FRQ (1:300) as primary antibody, and 4) goat anti-rabbit IgG (H+L) (1:5000, No. 170-6515; Bio-Rad, Hercules, CA) as secondary antibody.
Race tube and bioluminescence assays
Race tube and bioluminescence assays were performed as previously described in Hong et al. (37) and Dunlap and Loros (38). We used standard race tubes containing Vogel’s medium (pH 5.8) with 0.17% arginine, 50 ng/mL biotin, 1.5% agar, and 12.5 μM of luciferin for bioluminescence assays, and varying concentrations of glucose (0.1–0.3% m/v).
Results
The proposed circadian clock model reproduces wild-type and mutant phenotypes
Fig. 2, A–C, shows oscillation profiles derived from the simulated WT circadian system and Fig. 2 D shows experimental data from the wc-1-luciferase reporter.
Figure 2.
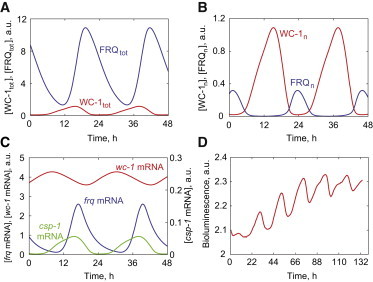
Oscillatory profiles for the wild-type parameterization. (A) Total WC-1 and FRQ levels show greater abundance of FRQ compared to WC-1. (B) The abundance of WC-1n is greater than FRQn for the most of circadian time. (C) All three transcripts of frq, wc-1, and csp-1 show sustained oscillations. (D) The wc-1-luciferase reporter shows robust circadian oscillations as predicted in the model. This data is a representative of three independent experiments. Arbitrary units (a.u.) are shown. To see this figure in color, go online.
The abundance of total FRQ is greater than the abundance of total WC-1 (20), and there is 4–10 h of phase difference between FRQ and WC-1 depending on whether we measure by the peak or trough of the protein concentrations (Fig. 2 A) (12). In contrast to total WC-1 and FRQ concentrations, WC-1n levels are higher than FRQn levels because FRQ is mainly localized in the cytoplasm (4,20), which is also simulated in our model (Fig. 2 B). Negative feedback from CSP-1 onto wc-1 expression is moderate, resulting in low-amplitude oscillations of wc-1 mRNA levels (Fig. 2 C) that were not reported previously. Oscillations of wc-1 mRNA may not have been observed in previous experiments (12) due to their low amplitude and the presence of molecular noise. We constructed a wc-1 bioluminescence reporter by fusing the promoter of wc-1 with a codon-optimized firefly luciferase, and validated that gene expression of wc-1 oscillates in a circadian manner (Fig. 2 D). Garceau et al. (26) observed a rapid rise in frq mRNA level and its subsequent rapid exponential decay. In contrast, the increase in FRQ protein concentration was followed by a slower, almost linear decay of FRQ protein level (26). In the model, the temporal profiles of frq mRNA (Fig. 2 C) and total FRQ protein (Fig. 2 A) accurately reproduce these distinct shapes of frq mRNA and FRQ protein oscillations. The expression of csp-1 shows circadian oscillations with peak expression aligned with the peak of WC-1 (WCC) because its expression is controlled by WCC (14,15) (Fig. 2 C).
Our model not only reproduces wild-type profiles, but also simulates experimentally observed changes in period in different clock overexpression strains. In Neurospora, the quinic acid (QA)-inducible system is readily used to increase the expression of a target gene in a QA dosage-dependent manner (39,40). Use of this for frq overexpression (frq+;qa-2-frq) results in small changes in period at low induction, and abolishes oscillations at high induction (40). In Fig. 3 A, we show that overexpression of frq with parameter k01 that simulates additional transcription of the frq gene (see model equations in Materials and Methods) does not produce large changes in period, and the system transitions into a stable steady state with no oscillations when the value of k01 is >0.14. It is important to note that the amplitude of FRQn plays a critical role in determining the period of the system (Fig. S1 in the Supporting Material). The period is maintained in the case of frq overexpression, because there is only a minimal change in the amplitude of FRQn oscillations before the system transitions to a stable steady state crossing a Hopf bifurcation point (Fig. S2).
On the other hand, wc-1 overexpression (wc-1+;qa-2-wc-1) experiments reveal a monotonically decreasing period with increasing concentration of the inducer, QA (6). This decrease of period is observed as a function of the rate of wc-1 overexpression, k03 (Fig. 3 B). Cheng et al. (6) also performed overexpression of wc-1 in the wc-1ko background (wc-1ko;qa-2-wc-1), and observed no oscillations at low QA levels. However, the period decreased from ∼24.5 h to 22.8 h at higher QA levels (6). Our model reproduces experimentally observed shortening of the period with the increase in wc-1 overexpression rate k03 in a simulated wc-1ko strain (k7 = 0) (Fig. 3 C). The initial transient increase of the period in the model (Fig. 3 C) corresponds to the parameter range near the Hopf bifurcation that is characterized by low albeit increasing amplitude of FRQn oscillations (Fig. S3 and see Text S2 in the Supporting Material for the mechanism of period regulation), which are easily destroyed in the model in the presence of molecular noise (Fig. S4).
In contrast to the decrease of period with wc-1 overexpression, Sancar et al. (15) observed an increase of period by ∼2.5 h with a moderate overexpression of csp-1, and then a loss of circadian rhythms with higher induction of csp-1 in a csp-1ko background (csp-1ko;qa-2-csp-1) (15). Our model reproduces period lengthening of the circadian clock with the increase of csp-1 expression as modeled by increasing the rate of csp-1 overexpression, k04, in the csp-1ko background (k16 = 0) (Fig. 3 D). The increase of period coincides with the rapid decrease in FRQn levels (Fig. S5, A and B, and see Text S2 in the Supporting Material). Interestingly, FRQc levels decrease more gradually than FRQn (Fig. S5 B). Finally, a decrease in period occurs in the model for larger values of k04 (Fig. 3 D). This period decrease is characterized by low abundance of FRQn (Fig. S5 B) that is no longer able to effectively clear WC-1 from the nucleus so that the amplitude of WC-1n oscillations becomes small (Fig. S5 C). These low-amplitude oscillations are likely to be destroyed by molecular noise (see Fig. S4) and, therefore, may not be observed experimentally (16). This conclusion is indirectly suggested by a strongly damped amplitude of frq-luc bioluminescence as well as weakened conidiation rhythm observed at an intermediate QA level right before the loss of circadian oscillations in csp-1 overexpression experiments (16). It is important to note that the increase in csp-1 overexpression (k04) corresponds to the decrease of wc-1 overexpression rate, k03, because of the repression CSP-1 exerts on wc-1 transcription. Hence, the decrease of period observed in Fig. 3 D is equivalent to the increase of period shown in Fig. 3 C.
Point mutations of frq that alter the circadian period have been interpreted as changing the half-life of the FRQ protein (41,42). Experimental (24,43) and modeling (23,44) studies are consistent with this, showing a strong correlation between FRQ stability and the period of the clock, with slow FRQ degradation (frq7) seen with longer periods whereas fast degradation of FRQ (frq1) is seen with shorter periods. Our model simulates periods of frq1 and frq7 mutants by changing the FRQ degradation rate constant (Fig. S6), which is consistent with previous results.
The glucose compensation is achieved by balancing the rates of activation between frq and csp-1
The period of circadian rhythms is relatively insensitive to changes in nutrient conditions. As demonstrated in recent experiments, Neurospora crassa showed small changes in period over a wide range of glucose concentrations (16). In contrast, a significantly shorter period was observed in csp-1ko compared to WT with increasing glucose levels (16). In the same study, an increased abundance of wc-1 mRNA and WC-1 were observed in csp-1ko compared to WT in high-glucose (2%) but not in low-glucose (0.1%) conditions. On the other hand, a glucose-dependent increased level of CSP-1 was observed in wc-2ko (16), which suggested that glucose-induced expression of CSP-1 is independent of WCC. The glucose-dependent repression of wc-1 transcription by CSP-1 strongly suggests a central role of CSP-1 in a circadian glucose compensation mechanism. This led Sancar et al. (16) to hypothesize that glucose compensation of the circadian period is achieved by balancing increased abundance of WC-1 with the CSP-1-dependent repression of wc-1 transcription.
To begin with, we simulated the csp-1ko strain by setting the rate constant of csp-1 expression k16 = 0. We modeled different glucose levels by changing the rate constant of wc-1 expression, k7, according to the reported increase in WC-1 protein abundance at high glucose concentrations (16). Our model reproduces the decrease of period as a function of the rate of wc-1 transcription (k7) (Fig. 4 A). The decrease in period is accompanied by an increase in FRQn amplitude (Fig. 4 B and see Text S2 in the Supporting Material), whereas the saturation of FRQn and WC-1n amplitudes (Fig. 4, B and C) corresponds to the stabilization of the period at ∼20 h (Fig. 4 A). There exists a narrow region where the period sharply increases with increasing rate constant k7 (Fig. 4 A). This region corresponds to low-amplitude oscillations near Hopf bifurcation (Fig. 4, B and C) that are easily destroyed by molecular noise (see Fig. S4).
Figure 4.
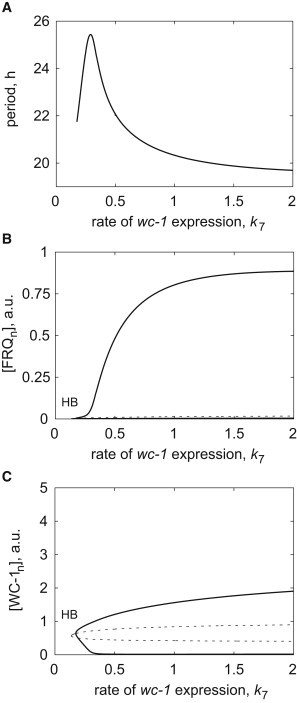
Simulated loss of glucose compensation of circadian clock in the csp-1ko strain. (A) Clock period as a function of parameter k7, simulating the increase in glucose concentration. One-parameter bifurcation diagrams show the envelopes (maximum and minimum) of (B) FRQn and (C) WC-1n oscillations as a function of k7. Oscillatory region is bounded by supercritical Hopf bifurcation (HB) at low values of k7 and is unbounded at high values of k7 due to saturation of WC-1 synthesis with respect to wc-1 mRNA (see Materials and Methods). (Thick solid curves) Stable oscillatory solutions; (thin solid curves) stable steady states; (dashed curves) unstable steady states. The rate of synthesis of csp-1 mRNA is set to zero (k16 = 0) for these simulations.
To investigate glucose compensation with our model, we computed the constant period curves in the two-parameter bifurcation plane of the rates of csp-1 (k16) and wc-1 (k7) transcription (Fig. 5). Our analysis demonstrates that fixed period curves have a remarkably constant positive slope around the wild-type period of ∼22.4 h. Hence, simultaneous increase of the above pair of parameters leads to a constant period in the model. The shorter period with an increase of wc-1 transcription rate as a function of glucose concentration is compensated near the WT period by the longer period with the increase of csp-1 transcription rate, which results in the repression of wc-1 expression by the increased amount of CSP-1 (16). Therefore, our model is consistent with the glucose compensation being achieved by balancing the synthesis of wc-1 and csp-1.
Figure 5.
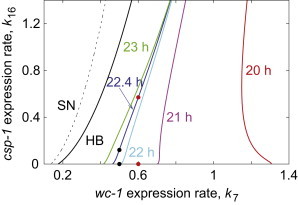
Simulated glucose compensation of the clock period in wild-type and its loss in wc-1ov mutant. The two-parameter bifurcation diagram in wc-1 and csp-1 transcription rate constants k7 and k16, correspondingly, shows curves of constant period, color-coded. Oscillatory solutions appear from the Hopf bifurcation (HB). Parameter sets in low-glucose (black dots) and high-glucose (red dots) conditions are used for stochastic simulations in the analysis of period robustness against molecular stochasticity. The csp-1ko corresponds to the x axis in the diagram (k16 = 0). The values of the model parameters are as described in Materials and Methods.
Intriguingly, the fixed period curve for a short period (T = 20 h) at higher values of k7 has a negative slope (Fig. 5). Near the period T = 20 h (k7 = 1.2), increase of csp-1 expression rate (k16) results in a shorter period, which is the opposite behavior to when the system is in the wild-type regime (k7 = 0.5). In other words, the balance between wc-1 and csp-1 transcription that we observe near the WT period T = 22.4 h is no longer present in a different parameter space with a period T = 20 h. Therefore, the wc-1 expression rate has to decrease to maintain constant period, which results in the negative slope of the fixed period curve T = 20 h (Fig. 5).
Based on this analysis, our model predicts a loss of glucose compensation in a wc-1 overexpression mutant (i.e., increased rate of wc-1 transcription) if we assume that both csp-1 and wc-1 synthesis rates increase as a function of glucose concentration in the growth media. To test this hypothesis, we constructed a wc-1 overexpression strain by transforming an additional wc-1 gene with its endogenous promoter at the csp-1 locus into a Neurospora strain that contains a frq-luc bioluminescence reporter (36). WC-1 protein abundance is increased in the wc-1ov strain compared to the wild-type (Fig. 6 A). More importantly, the wc-1ov strain demonstrates a decrease of period as a function of glucose concentration in both race tube (Fig. 6, B and C) and frq-luciferase bioluminescence assays (Fig. 6 D); these data indicate a loss of glucose compensation.
Figure 6.
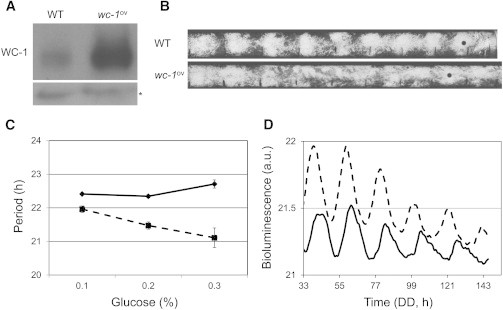
Loss of glucose compensation in the wc-1ov mutant. (A) Western blot shows increased abundance of WC-1 in the wc-1ov mutant. Nonspecific bands (∗) are shown as a loading control. This is a representative blot from three experiments. (B) Representative race tubes of wild-type and the wc-1ov mutant grown on a defined agar media (see Materials and Methods) with 0.3% glucose (m/v). (Small black bars) Markings of Neurospora growth front in every 24 h. (Black dots on the race tubes) Period of wc-1ov is shorter than the wild-type. (C) Graph showing free running periods from wild-type (solid line) and wc-1ov (dashed line) in the media containing different glucose concentrations. The period decreases in wc-1ov with increasing glucose concentration in the media. The periods of race tube assay of the wild-type are 22.41 ± 0.03 h at 0.1% glucose, 22.35 ± 0.04 h (n = 6) at 0.2% glucose, and 22.71 ± 0.12 h (n = 6) at 0.3% glucose. The quantified periods of race tube assay of the wc-1ov at different glucose concentrations are 21.96 ± 0.1 h (n = 6) at 0.1% glucose, 21.47 ± 0.11 h (n = 6) at 0.2% glucose, and 21.11 ± 0.3 h (n = 5) at 0.3% glucose. Error bars represent mean ± SE. (D) Graph showing bioluminescence activity from frq-luciferase reporter from wild-type (solid curve, 21.82 h) and wc-1ov (dashed curve, 20.87 h). Neurospora are grown in the media containing 0.3% glucose (m/v). wc-1ov demonstrates shorter period compared to wild-type, which is consistent with (B). The figure shows a representative data from three independent experiments.
Analysis of period robustness against molecular stochasticity
It has been shown that the interlocked dual negative feedback loop creates a wider oscillatory domain (21). We wondered if the additional negative feedback loop mediated by CSP-1 enhances the robustness of the system against molecular noise as a function of glucose concentration. To test this hypothesis, we performed stochastic simulations of the clock model to determine the effect of noise on the period of Neurospora circadian oscillator in both WT and csp-1ko as a function of glucose concentration (i.e., low and high values of k7 and k16). The period histograms of stochastic simulations in low-glucose conditions (black dots in Fig. 5) are plotted in Fig. 7. In the case of a strong noise, the WT parameter set shows larger variation of the periods (Fig. 7 A) when compared to the period variation in the case of the csp-1ko parameter set (Fig. 7 B). In the case of a weak noise, we observed similar behavior except period variation was smaller, as expected for weak noise (Fig. S7, A and B). Therefore, unexpectedly, csp-1ko appears to be slightly more robust than the WT with either strong or weak noise in the system.
We also investigated the effect of noise in the model in high glucose conditions (red dots in Fig. 5). Here, similar to low-glucose conditions, the WT had more variation in period compared to csp-1ko when strong noise was present (Fig. 7, C and D). Notably, the WT in high glucose had similar period sensitivity compared to the WT in low glucose (Fig. 7, C and A), whereas csp-1ko in high glucose had less variation in period compared to csp-1ko in low glucose (Fig. 7, D and B). In the case of weak noise, period variation in WT and csp-1ko in high glucose resembled the results in the case of strong noise (Fig. S7, C and D) as well as between high- and low-glucose parameter sets (Fig. S7), but variation of periods was smaller as expected for weak noise. The race tube and bioluminescence data show a robust circadian period with standard errors at ∼0.2 h or 10% even in the csp-1ko, which is consistent with our simulations (16).
It is important to note that the location of the WT parameter space is closer to the Hopf bifurcation compared to csp-1ko parameter space (Fig. 5), because it was previously shown that the proximity of a Hopf bifurcation leads to less robust oscillations against noise (45). Therefore, to make a fair comparison of robustness of csp-1ko and WT parameter sets against noise, we performed stochastic simulations of the csp-1ko parameter set in very low glucose (Fig. S8) that has a similar distance to Hopf bifurcation as the WT parameter set in low glucose. We observed period variation that was similar to the WT parameter set (Figs. S8 and 7), which indicates that the proximity to the Hopf bifurcation plays a defining role in determining the robustness of circadian oscillations against noise in the Neurospora clock.
Discussion
In this article, we constructed a mathematical model of the Neurospora circadian clock that incorporates a transcriptional repressor, CSP-1 (15), which resulted in a model with interlocked dual negative feedback loops (Fig. 1). This network topology resembles the mammalian circadian clock model in which REV-ERB-α/β forms a second negative feedback loop inhibiting the circadian clock transcription factor, CLOCK/BMAL1 (21). The model simulates the changes in clock period observed in different overexpression experiments (Fig. 3). Moreover, the model accurately reproduces compensation of clock period across glucose concentrations (Fig. 5) and the loss of this compensation in csp-1 mutants (Fig. 4 A). Glucose compensation in the model is achieved by balancing the rates of wc-1 and csp-1 transcription. This balance is lost in the csp-1 mutant and the period of the clock decreases at high glucose concentration in the media. Our model predicts the loss of glucose compensation in wc-1ov mutants (Fig. 5), which indicates that glucose compensation is achieved within a narrow range of parameter space. More importantly, we validate this modeling prediction in experiments (Fig. 6).
Based on our data and the similarities in the regulatory architecture of the fungal and animal circadian clocks (1,46), we hypothesize defective glucose compensation in Clk and Bmal1 overexpression mutant mice. The outcome of this hypothesis will depend on how glucose influences the master clock in the suprachiasmatic nucleus, which coordinates peripheral clocks (47). It has been shown that high-fat diet shifts patterns of mouse food intake and attenuates amplitudes of clock components (48). Similarly, it would be interesting to investigate glucose-dependent changes of phase and amplitude of core circadian clock components in Neurospora, and the function of CSP-1 in such responses. Furthermore, it will be critical to investigate the range of glucose compensation and interconnected molecular responses of circadian rhythms and glucose metabolism to determine the glucose metabolism-dependent physiological outcomes.
The robustness of an oscillatory system against molecular noise depends on several factors including the network topology, parameter space, and functions. Recently, several studies investigated the effect of the second negative feedback loop on robustness of the negative feedback oscillatory systems (21,49). Tsai et al. (49) reported no improvement from additional negative feedback in oscillatory range or operational frequency range for simple oscillator models in accord with our findings for the Neurospora clock. In contrast, Kim and Forger (21) showed that the additional slow negative feedback loop on the core clock oscillator increases the oscillatory range of the system. More importantly, they also demonstrated that the second negative feedback loop regulates stoichiometric balance of core clock repressors and activators that is critical for sustained oscillations. However, the second negative feedback loop formed by CSP-1 in Neurospora is fast relative to the core negative feedback loop (the ratio of csp-1 mRNA degradation rate to frq mRNA degradation rate δ ≈ 6). Both the Neurospora clock and the model of Tsai et al. (49) include Hill functions to describe transcription, whereas the model of Kim and Forger (21) uses protein sequestration. A recent study also showed how models behave differently depending on the choice of transcriptional mechanism between protein sequestration and Hill functions (50). Therefore, our study provides further confirmation for the hypothesis that the effect of an additional negative feedback loop on robustness of the system may depend on the mechanism of transcriptional regulation in the model.
Our stochastic simulations revealed that a CSP-1-mediated negative feedback loop does not enhance robustness of the oscillatory system against noise, because the robustness of the circadian oscillator strongly depends on the proximity of the system to the Hopf bifurcation (Figs. 5 and 7), which is in agreement with previous modeling results (21,45). In particular, in the model of Kim and Forger (21), the second negative feedback loop increases the oscillatory range and, thus, increases the distance of the system to the Hopf bifurcation. Moreover, our simulations indicate that the oscillations are quickly lost even in the presence of a weak molecular noise when the system is in immediate vicinity of the Hopf bifurcation (Fig. S4). These modeling results are confirmed by experimental data on wc-1 overexpression in wc-1ko background (6) and on csp-1 overexpression in csp-1ko background (16). These experimental data show weak and fragile conidiation rhythm, reduced amplitude of frq bioluminescence, and increased standard errors of circadian periods right after the appearance of oscillations or just before losing their rhythmicity, which strongly suggests loss of robustness of circadian clock against molecular noise near transitions to steady states. In this report, we demonstrate that iterative mathematical modeling and experimental validations advance our detailed understanding of the Neurospora circadian system, and continued development of such models will elucidate interconnected molecular responses between the circadian clock and glucose metabolism.
Acknowledgments
We thank H. Kang for her input in our stochastic simulations, and J. Bellman and K. Noh for technical assistance.
This work was supported by Department of Interior grant No. D12AP00005 and the Charles Phelps Taft Research Center, University of Cincinnati.
Supporting Material
References
- 1.Dunlap J.C. Molecular bases for circadian clocks. Cell. 1999;96:271–290. doi: 10.1016/s0092-8674(00)80566-8. [DOI] [PubMed] [Google Scholar]
- 2.Dunlap J.C., Loros J.J., Lambreghts R. A circadian clock in Neurospora: how genes and proteins cooperate to produce a sustained, entrainable, and compensated biological oscillator with a period of about a day. Cold Spring Harb. Symp. Quant. Biol. 2007;72:57–68. doi: 10.1101/sqb.2007.72.072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Baker C.L., Kettenbach A.N., Dunlap J.C. Quantitative proteomics reveals a dynamic interactome and phase-specific phosphorylation in the Neurospora circadian clock. Mol. Cell. 2009;34:354–363. doi: 10.1016/j.molcel.2009.04.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Cheng P., Yang Y., Liu Y. Coiled-coil domain-mediated FRQ-FRQ interaction is essential for its circadian clock function in Neurospora. EMBO J. 2001;20:101–108. doi: 10.1093/emboj/20.1.101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Crosthwaite S.K., Dunlap J.C., Loros J.J. Neurospora wc-1 and wc-2: transcription, photoresponses, and the origins of circadian rhythmicity. Science. 1997;276:763–769. doi: 10.1126/science.276.5313.763. [DOI] [PubMed] [Google Scholar]
- 6.Cheng P., Yang Y., Liu Y. Interlocked feedback loops contribute to the robustness of the Neurospora circadian clock. Proc. Natl. Acad. Sci. USA. 2001;98:7408–7413. doi: 10.1073/pnas.121170298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Denault D.L., Loros J.J., Dunlap J.C. WC-2 mediates WC-1-FRQ interaction within the PAS protein-linked circadian feedback loop of Neurospora. EMBO J. 2001;20:109–117. doi: 10.1093/emboj/20.1.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cheng P., He Q., Liu Y. Regulation of the Neurospora circadian clock by an RNA helicase. Genes Dev. 2005;19:234–241. doi: 10.1101/gad.1266805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Cha J., Chang S.S., Liu Y. Control of WHITE COLLAR localization by phosphorylation is a critical step in the circadian negative feedback process. EMBO J. 2008;27:3246–3255. doi: 10.1038/emboj.2008.245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hong C.I., Ruoff P., Dunlap J.C. Closing the circadian negative feedback loop: FRQ-dependent clearance of WC-1 from the nucleus. Genes Dev. 2008;22:3196–3204. doi: 10.1101/gad.1706908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Schafmeier T., Diernfellner A., Brunner M. Circadian activity and abundance rhythms of the Neurospora clock transcription factor WCC associated with rapid nucleo-cytoplasmic shuttling. Genes Dev. 2008;22:3397–3402. doi: 10.1101/gad.507408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lee K., Loros J.J., Dunlap J.C. Interconnected feedback loops in the Neurospora circadian system. Science. 2000;289:107–110. doi: 10.1126/science.289.5476.107. [DOI] [PubMed] [Google Scholar]
- 13.Shi M., Collett M., Dunlap J.C. FRQ-interacting RNA helicase mediates negative and positive feedback in the Neurospora circadian clock. Genetics. 2010;184:351–361. doi: 10.1534/genetics.109.111393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lambreghts R., Shi M., Loros J.J. A high-density single nucleotide polymorphism map for Neurospora crassa. Genetics. 2009;181:767–781. doi: 10.1534/genetics.108.089292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Sancar G., Sancar C., Brunner M. A global circadian repressor controls antiphasic expression of metabolic genes in Neurospora. Mol. Cell. 2011;44:687–697. doi: 10.1016/j.molcel.2011.10.019. [DOI] [PubMed] [Google Scholar]
- 16.Sancar G., Sancar C., Brunner M. Metabolic compensation of the Neurospora clock by a glucose-dependent feedback of the circadian repressor CSP1 on the core oscillator. Genes Dev. 2012;26:2435–2442. doi: 10.1101/gad.199547.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bugge A., Feng D., Lazar M.A. Rev-erbα and Rev-erbβ coordinately protect the circadian clock and normal metabolic function. Genes Dev. 2012;26:657–667. doi: 10.1101/gad.186858.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Cho H., Zhao X., Evans R.M. Regulation of circadian behavior and metabolism by REV-ERB-α and REV-ERB-β. Nature. 2012;485:123–127. doi: 10.1038/nature11048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Solt L.A., Wang Y., Burris T.P. Regulation of circadian behavior and metabolism by synthetic REV-ERB agonists. Nature. 2012;485:62–68. doi: 10.1038/nature11030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Schafmeier T., Haase A., Brunner M. Transcriptional feedback of Neurospora circadian clock gene by phosphorylation-dependent inactivation of its transcription factor. Cell. 2005;122:235–246. doi: 10.1016/j.cell.2005.05.032. [DOI] [PubMed] [Google Scholar]
- 21.Kim J.K., Forger D.B. A mechanism for robust circadian timekeeping via stoichiometric balance. Mol. Syst. Biol. 2012;8:630. doi: 10.1038/msb.2012.62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hong C.I., Jolma I.W., Ruoff P. Simulating dark expressions and interactions of frq and wc-1 in the Neurospora circadian clock. Biophys. J. 2008;94:1221–1232. doi: 10.1529/biophysj.107.115154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ruoff P., Vinsjevik M., Rensing L. The Goodwin oscillator: on the importance of degradation reactions in the circadian clock. J. Biol. Rhythms. 1999;14:469–479. doi: 10.1177/074873099129001037. [DOI] [PubMed] [Google Scholar]
- 24.Ruoff P., Loros J.J., Dunlap J.C. The relationship between FRQ-protein stability and temperature compensation in the Neurospora circadian clock. Proc. Natl. Acad. Sci. USA. 2005;102:17681–17686. doi: 10.1073/pnas.0505137102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yu Y., Dong W., Schüttler H.B. A genetic network for the clock of Neurospora crassa. Proc. Natl. Acad. Sci. USA. 2007;104:2809–2814. doi: 10.1073/pnas.0611005104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Garceau N.Y., Liu Y., Dunlap J.C. Alternative initiation of translation and time-specific phosphorylation yield multiple forms of the essential clock protein FREQUENCY. Cell. 1997;89:469–476. doi: 10.1016/s0092-8674(00)80227-5. [DOI] [PubMed] [Google Scholar]
- 27.Gonze D., Halloy J., Goldbeter A. Deterministic versus stochastic models for circadian rhythms. J. Biol. Phys. 2002;28:637–653. doi: 10.1023/A:1021286607354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Gonze D., Abou-Jaoudé W., Halloy J. How molecular should your molecular model be? On the level of molecular detail required to simulate biological networks in systems and synthetic biology. Methods Enzymol. 2011;487:171–215. doi: 10.1016/B978-0-12-381270-4.00007-X. [DOI] [PubMed] [Google Scholar]
- 29.Kim J.K., Josić K., Bennett M.R. The validity of quasi-steady-state approximations in discrete stochastic simulations. Biophys. J. 2014;107:783–793. doi: 10.1016/j.bpj.2014.06.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Agarwal A., Adams R., Shouval H.Z. On the precision of quasi steady state assumptions in stochastic dynamics. J. Chem. Phys. 2012;137:044105. doi: 10.1063/1.4731754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Thomas P., Straube A.V., Grima R. The slow-scale linear noise approximation: an accurate, reduced stochastic description of biochemical networks under timescale separation conditions. BMC Syst. Biol. 2012;6:39. doi: 10.1186/1752-0509-6-39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kampen N.G.v. North-Holland; Amsterdam, Netherlands: 1992. Stochastic Processes in Physics and Chemistry. [Google Scholar]
- 33.Gillespie D.T. General method for numerically simulating stochastic time evolution of coupled chemical-reactions. J. Comput. Phys. 1976;22:403–434. [Google Scholar]
- 34.Gillespie D.T. Stochastic simulation of chemical kinetics. Annu. Rev. Phys. Chem. 2007;58:35–55. doi: 10.1146/annurev.physchem.58.032806.104637. [DOI] [PubMed] [Google Scholar]
- 35.Chen C.H., DeMay B.S., Loros J.J. Physical interaction between VIVID and white collar complex regulates photoadaptation in Neurospora. Proc. Natl. Acad. Sci. USA. 2010;107:16715–16720. doi: 10.1073/pnas.1011190107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Gooch V.D., Mehra A., Dunlap J.C. Fully codon-optimized luciferase uncovers novel temperature characteristics of the Neurospora clock. Eukaryot. Cell. 2008;7:28–37. doi: 10.1128/EC.00257-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Hong C.I., Zámborszky J., Csikász-Nagy A. Circadian rhythms synchronize mitosis in Neurospora crassa. Proc. Natl. Acad. Sci. USA. 2014;111:1397–1402. doi: 10.1073/pnas.1319399111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Dunlap J.C., Loros J.J. Analysis of circadian rhythms in Neurospora: overview of assays and genetic and molecular biological manipulation. Methods Enzymol. 2005;393:3–22. doi: 10.1016/S0076-6879(05)93001-2. [DOI] [PubMed] [Google Scholar]
- 39.Giles N.H., Case M.E., Tyler B. Gene organization and regulation in the QA (quinic acid) gene cluster of Neurospora crassa. Microbiol. Rev. 1985;49:338–358. doi: 10.1128/mr.49.3.338-358.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Aronson B.D., Johnson K.A., Dunlap J.C. Negative feedback defining a circadian clock: autoregulation of the clock gene frequency. Science. 1994;263:1578–1584. doi: 10.1126/science.8128244. [DOI] [PubMed] [Google Scholar]
- 41.Gardner G.F., Feldman J.F. Temperature compensation of circadian period length in clock mutants of Neurospora crassa. Plant Physiol. 1981;68:1244–1248. doi: 10.1104/pp.68.6.1244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Aronson B.D., Johnson K.A., Dunlap J.C. Circadian clock locus frequency: protein encoded by a single open reading frame defines period length and temperature compensation. Proc. Natl. Acad. Sci. USA. 1994;91:7683–7687. doi: 10.1073/pnas.91.16.7683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Liu Y., Loros J., Dunlap J.C. Phosphorylation of the Neurospora clock protein FREQUENCY determines its degradation rate and strongly influences the period length of the circadian clock. Proc. Natl. Acad. Sci. USA. 2000;97:234–239. doi: 10.1073/pnas.97.1.234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Ruoff P., Mohsenzadeh S., Rensing L. Circadian rhythms and protein turnover: the effect of temperature on the period lengths of clock mutants simulated by the Goodwin oscillator. Naturwissenschaften. 1996;83:514–517. doi: 10.1007/BF01141953. [DOI] [PubMed] [Google Scholar]
- 45.Gonze D., Goldbeter A. Circadian rhythms and molecular noise. Chaos. 2006;16:026110. doi: 10.1063/1.2211767. [DOI] [PubMed] [Google Scholar]
- 46.Bell-Pedersen D., Cassone V.M., Zoran M.J. Circadian rhythms from multiple oscillators: lessons from diverse organisms. Nat. Rev. Genet. 2005;6:544–556. doi: 10.1038/nrg1633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Stetson M.H., Watson-Whitmyre M. Nucleus suprachiasmaticus: the biological clock in the hamster? Science. 1976;191:197–199. doi: 10.1126/science.942799. [DOI] [PubMed] [Google Scholar]
- 48.Kohsaka A., Laposky A.D., Bass J. High-fat diet disrupts behavioral and molecular circadian rhythms in mice. Cell Metab. 2007;6:414–421. doi: 10.1016/j.cmet.2007.09.006. [DOI] [PubMed] [Google Scholar]
- 49.Tsai T.Y., Choi Y.S., Ferrell J.E., Jr. Robust, tunable biological oscillations from interlinked positive and negative feedback loops. Science. 2008;321:126–129. doi: 10.1126/science.1156951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Kim J.K., Kilpatrick Z.P., Josić K. Molecular mechanisms that regulate the coupled period of the mammalian circadian clock. Biophys. J. 2014;106:2071–2081. doi: 10.1016/j.bpj.2014.02.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


