Abstract
Background: Nuclear distribution E homolog 1 (NDE1), located within chromosome 16p13.11, plays an essential role in microtubule organization, mitosis, and neuronal migration and has been suggested by several studies of rare copy number variants to be a promising schizophrenia (SCZ) candidate gene. Recently, increasing attention has been paid to rare single-nucleotide variants (SNVs) discovered by deep sequencing of candidate genes, because such SNVs may have large effect sizes and their functional analysis may clarify etiopathology. Methods and Results: We conducted mutation screening of NDE1 coding exons using 433 SCZ and 145 pervasive developmental disorders samples in order to identify rare single nucleotide variants with a minor allele frequency ≤5%. We then performed genetic association analysis using a large number of unrelated individuals (3554 SCZ, 1041 bipolar disorder [BD], and 4746 controls). Among the discovered novel rare variants, we detected significant associations between SCZ and S214F (P = .039), and between BD and R234C (P = .032). Furthermore, functional assays showed that S214F affected axonal outgrowth and the interaction between NDE1 and YWHAE (14-3-3 epsilon; a neurodevelopmental regulator). Conclusions: This study strengthens the evidence for association between rare variants within NDE1 and SCZ, and may shed light into the molecular mechanisms underlying this severe psychiatric disorder.
Key words: YWHAE, SNV, target resequencing, rare variants, DISC1, protein-protein interaction
Introduction
Schizophrenia (SCZ), of which clinical features are characterized by hallucinations, delusions, and cognitive deficits, and cause enormous personal and societal burdens,1 is a severe psychiatric disorder with a lifetime risk of about 1%. The heritability of SCZ is as high as 80%,2,3 making this condition a target for human genetics research. Recent studies into the genetic architecture of SCZ have identified both common and rare variants.4 Several common variants with relatively small odds ratios (ORs) for SCZ have been identified in genome wide association studies (GWASs), and the findings indicate that common variants account for at least a third of the genetic contribution to SCZ risk3,5 and that genetic risk overlaps with bipolar disorders (BD).3 Additionally, studies of rare variations of SCZ and autism spectrum disorder (ASD) identified rare copy number variations (CNV) with large effect size,6 and such variants might be important to understand the etiopathology of SCZ and ASD. Despite their large effect size, the etiopathological roles of CNV remains largely unknown, partially because it remains unclear how the functional change of the genes within the regions of the CNVs lead to the pathogenesis of neuropsychiatric disorders.
On the other hand, there is growing evidence that rare SNVs, discovered from deep sequencing of candidate genes, may have large effect sizes and account for the missing heritability, as well as contribute to understanding of the pathogenesis of neuropsychiatric disease through functional analysis.7–10 Intriguingly, recently it has been suggested that some of genetic risk associated with SCZ11,12 converged on the similar set of genes in terms of their functions. For instance, the SNVs discovered from deep sequencing of ASD13,14 in the gene encoding the synaptic scaffolding protein SHANK3 (located within the CNVs at 22q13.3 which was associated with ASD),15 were found to affect dendritic spine morphology via an actin-dependent mechanism.10 Thus, sequencing the candidate genes from the large scale genome wide analyses may be a promising method for elucidating the pathophysiology of neuropsychiatric disorders such as SCZ and ASD.
There are multiple lines of evidence that deletions and reciprocal duplications at chromosome 16p13.11 are associated with disorders involving abnormal neurodevelopment, such as ASD,16 epilepsies,17–19 and SCZ.20–22 In some cases, microcephaly is a phenotypic manifestation of CNV deletions at 16p13.11.23,24 Given that the CNVs at 16p13.11 regions contain nuclear distribution E homolog 1 (NDE1), which are closely associated with neurodevelopmental phenotypes, NDE1 is a promising candidate gene. Furthermore, the NDE1 have been associated with SCZ25 and ASD26,27 risk by linkage analysis.
NDE1 plays a crucial role in the process of mammalian encephalization and human cerebral cortex growth because of its involvement in mitosis, neuronal migration, and microtubule organization during brain development.28 In an NDE1-deficient mouse model, in which the mice presented with microcephaly, it was found that NDE1-deficient mice had reduced progenitor cell division and alterations in mitotic spindle formation and in chromosome segregation.29 NDE1 protein has an N-terminal region with coiled-coil motif known to bind to lissencephaly 1,30 and a C-terminal region known to harbor several phosphorylation sites involved in subcellular localization, protein-protein interactions and the cell cycle.31,32 In humans, homozygous loss of function mutations at C-terminal regions have been associated with extreme microlissencephaly,28,32 suggesting that NDE1 is required for neurogenesis and neuronal migration. NDE1 was also identified as a SCZ susceptibility locus by a genome-wide linkage study on common risk variants of disrupted in schizophrenia 1 (DISC1),33 which is a scaffold protein indicated in neurodevelopment and synaptic regulation with several interacting proteins and is a suggested risk factor for SCZ, ASD, and BD.34 NDE1 and NDE-like 1 (NDEL1), which is a paralog of NDE1,35 are known to directly bind with DISC1.36,37 Furthermore, in the region of the YWHAE (14-3-3 epsilon), that binds to DISC1 and is involved in neural migration by recognizing the phosphorylation sites of NDEL1,38 we previously identified a SNP (rs28365859) that showed a significant association with Japanese SCZ.39
Considering that NDE1 is essential for neurodevelopment and is included in the SCZ and ASD susceptibility locus at 16p13.11, we hypothesized that mutations in NDE1 might confer susceptibility to SCZ, ASD, and BD pathogenesis. However, there have been no studies focusing only on NDE1 variants by deep sequencing of SCZ and ASD. Therefore, in this study, in order to discover rare variants with large effect size and explore the role in pathogenesis of the discovered rare variants, we performed mutation screening of NDE1 coding exons with SCZ and pervasive developmental disorder (PDD) samples, followed by association analysis and functional analysis of variants with putative large effects. In this study, we found that rare variants of NDE1 may have functional relevance for the pathophysiology of SCZ and BD.
Methods
Samples
Two independent sample groups were used in this study. (1) For mutation screening, we used 433 SCZ (mean age ± SD = 52.3±14.4 years) and 145 PDD (15.6±8.3 years). (2) For genetic association analysis, we used a case control sample set consisting of 3554 SCZ (48.4±14.6 years), 1041 BD (51.0±14.1 years), and 4746 healthy control (HC) subjects (43.2±14.7 years). All subjects were unrelated, living in the mainland of Japan, and self-identified as Japanese. All of the cases were included if they met Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition-Text Revision (TR) criteria for SCZ, BD, and PDD (specifically, Autistic disorder, Asperger’s disorder, PDD not otherwise specified). In addition, the patients’ capacity to consent was confirmed by a family member when needed. Control subjects were selected from the general population and had no history of mental disorders based on questionnaire responses from the subjects themselves during the sample inclusion step. A general characterization and psychiatric assessment of the subjects is available elsewhere.40 Written informed consent was obtained from all participants. The Ethics Committees of the Nagoya University Graduate School of Medicine and associated institutes and hospitals approved this study.
Mutation Screening of NDE1 Coding Exons
For the purpose of mutation screening, we designed 8 primer sets (supplementary table S1) to cover the NDE1 coding exons (transcription ID: ENST00000396355 from ensemble database; Chr16: 15,737,124-15,820,210; human reference sequence NCBI (National Center for Biotechnology Information) build 37.1). Genomic DNA was extracted from peripheral blood using standard methods. Amplicons were generated using standard polymerase chain reaction (PCR) conditions. After PCR amplification, aliquots of PCR products were purified using Illustra Exonuclease I and Alkaline Phosphatase (GE Healthcare and Life Science). These were then sequenced using the Sanger method and a 3130XL Genetic Analyzer (Applied Biosystems). Mutation Surveyor (Softgenetics), which can detect sequencing variations within Sanger sequencing traces, was used to detect mutations in NDE1.41 We prioritized SNVs for follow-up association analysis as follows. (1) We included rare (minor allele frequency [MAF] ≤ 5%) missense and nonsense SNVs, canonical splicing junction mutations (within 2 bp of intronic sequence at the exon/intron boundaries), and indels. (2) We included only “novel” variants; those that were not registered in dbSNP version 136 (http://www.ncbi.nlm.nih.gov/projects/SNP/). Each rare mutation was reconfirmed using Sanger sequencing. After prioritization, all variants were evaluated in silico for possible structural and functional consequences using the following tools: (1) localization of protein domain and phosphorylation sites was based on the human protein reference database (http://www.hprd.org/index_html) (HPRD); (2) evolutionary conservation was assessed by the HomoloGene database (http://www.ncbi.nlm.nih.gov/homologene/); and (3) analysis of deleterious effects by amino acid substitution was performed by algorithms implemented in Polyphen-2,42 Sorting Tolerant from Intolerant (SIFT),43 and PMUT.44
Genetic Association Analysis
Genotyping was performed using Taqman (Applied Biosystems) custom probes (details about DNA sequences and PCR conditions are available upon request). Each 384-microtiter plate contained at least 2 non-template controls and the sample(s) in which novel variant was observed. Analysis was performed on an HT7900 instrument (Applied Biosystems) according to the standard protocol. All allele-wise association analyses were carried out by calculating the P values for each SNP using Fisher’s exact test (one-tail). Differences were considered significant when the P value was less than .05. Statistical calculations were done using SPSS v21 (SPSS Inc.) and Plink v1.07.45 Furthermore, we calculated adjusted P value (100 000 permutations) as implemented in Plink v1.07. Calculation of statistical power was performed by the Genetic Power Calculator (http://pngu.mgh.harvard.edu/purcell/gpc/) with the multiplicative model.
Materials and Chemicals
The cDNAs encoding NDE1 and YWHAE/14-3-3ε were isolated from a human fetal brain cDNA library (Clontech). Antibodies against green fluorescent protein (GFP) or glutathione S-transferase (GST) were purchased from Nacalai and Sigma, respectively. Other materials and chemicals were obtained from commercial sources.
Plasmid Constructs and Protein Purification
A full-length cDNA encoding human NDE1 (wild type [WT]) (amino acids 1–335) was inserted into pEGFP-C-terminal 1 (C1) (Clontech) and separately into pEFBOS-GST (provided by Dr S. Nagata, Osaka University). Additionally, cDNAs encoding NDE1-S214A (SA), NDE1-S214F (SF), NDE1-T215A (TA), or NDE1-R234C (RC) were generated by site-directed mutagenesis, and were then cloned into pEGFP-C1 and separately into pEFBOS-GST. For protein purification of GST-YWHAE, the cDNA encoding YWHAE was inserted into pGEX-4T1 (GE Healthcare).46 GST fusion proteins were produced in Escherichia coli BL21 (DE3), and purified on glutathione-Sepharose 4B beads according to the manufacturer’s instructions (GE Healthcare).
Cell Culture and Transfections
HEK293T cells were cultured in Dulbecco’s modified Eagle’s medium (Sigma-Aldrich) supplemented with 10% fetal bovine serum (Invitrogen Life Technologies) in a 5% CO2 atmosphere at 37oC. HEK293T cells were transfected with the indicated plasmids using Lipofectamine (Invitrogen Life Technologies). At 36 hours after transfection, the cultured cells were harvested for a GST pull-down assay. For the primary culture of rat hippocampal neurons, all animal studies were conducted in accordance with the Animal Care and Use Committee guidelines of Nagoya University in Japan. Hippocampal neurons were prepared from E16 embryos of WT (C57BL6J) mice as previously described.47 Briefly, neurons were seeded onto a poly-D-lysine-coated 35-mm glass-based dish and cultured in neurobasal medium (Invitrogen) supplemented with B-27 (Invitrogen) and 1mM L-glutamine.
GST Pull-down Assay
Cells were lysed with the lysis buffer (20mM Tris-HCl, 1mM EDTA, 250mM NaCl, 1mM DTT, 0.1% [wt/vol] Triton X-100, 10μM p-amidinophenyl methanesulfonyl fluoride hydrochloride, and 10 μg/ml leupeptin). The lysate was sonicated, and then clarified by centrifugation at 12 000g for 10 minutes at 4°C. The soluble supernatant was incubated with the glutathione–Sepharose 4B beads (GE Healthcare) coated with 50 pmol of GST alone or GST-YWHAE. Beads were washed with lysis buffer, and the bound GST-proteins were then eluted with a glutathione-containing buffer (10mM glutathione in Tris-based buffer [pH 7.4]). A portion of each eluate was subjected to SDS-PAGE, followed by immunoblot analyses with the indicated antibodies. We used cell lysates from COS7 that expressed GST-YWHAE or GFP-NDE1 for in vivo GST pull-down assays. The COS7 cell lysates were incubated with glutathione–Sepharose 4B beads. These beads were washed with lysis buffer, and the bound GST-proteins were then eluted with the glutathione-containing buffer. To quantify the effects of the NDE1 mutations on the interaction with YWHAE, we performed the GST pull-down assay in triplicate. Quantitative densitometric measurement of immunoblots was performed using Image J (http://rsb.info.nih.gov/ij/). The Student’s t test was used to assess statistical significance.
Neuromorphological Analysis
Hippocampal neurons (6 × l04 cells per well) were seeded onto 24-well plate. At one day in vitro (DIV1), a CalPhos mammalian transfection kit (Clontech) was used to introduce via transfection expression plasmids encoding the indicated cDNA (1.0 μg) into hippocampal neurons. At day three in vitro (DIV3), transfected and control hippocampal neurons were fixed with 3.7% formaldehyde in PBS for 10 minutes and treated with PBS that contained 0.05% Triton X-100 for 10 minutes. Neurons were incubated with anti-GFP and anti-Tau1 antibodies (Sigma) overnight at 4°C, washed, and incubated for 1 hour with secondary antibodies. Immunofluorescent signals were examined with a laser scanning confocal microscope (LSM510; Zeiss). Axon lengths were measured from the cell body to the tip of axonal processes. The Student’s t test was used to assess statistical significance between groups.
Results
Mutation Screening of NDE1 Coding Exons
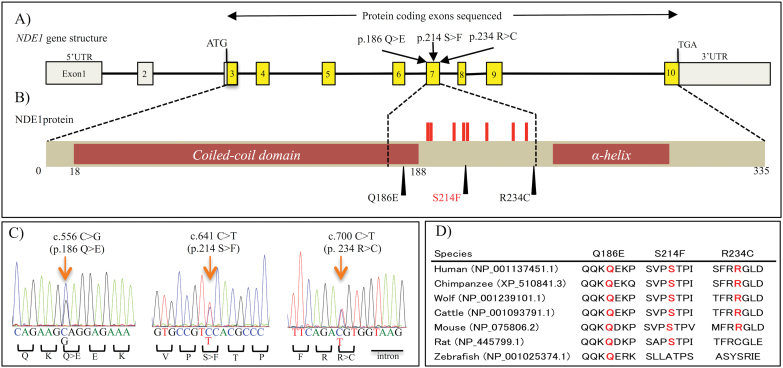
To clarify the genetic relationship of NDE1 in the pathology of SCZ and ASD, we performed a mutation screening of NDE1 coding exons using the 433 SCZ and 145 ASD samples. We identified 4 rare missense heterozygous mutations within NDE1 coding exons (table 1). No synonymous mutations, nonsense mutations, splicing site mutations, or indels were discovered. In one of the 145 ASD samples, we detected a known rare missense mutation (rs148118152) at exon4 within the N-terminal coiled-coil region. The other 3 rare missense mutations (Q186E, S214F, R234C) were novel and discovered within the 433 SCZ samples (MAF = 0.12%). All of the novel missense mutations were discovered within exon 7, where several phosphorylation sites are located (figure 1A); S214 is also located at a putative phosphorylation site according to the HPRD. S214F was predicted to be deleterious by all of the 3 algorithms (table 1). The result of evolutionary conservation analysis for the novel rare mutations is shown in figure 1D. Case summaries of patients with a novel rare missense mutation are provided in the supplementary results, supplementary tables S2 and S3, and supplementary figures S1 and S2.
Table 1.
Details of Discovered Rare Missense Mutations and In Silico Analysis
| Chromosome | Physical Positiona | Exon | Base Change Mb> mb | dbSNP Reference | Variantc | SCZ (n = 433) | PDD (n = 145) | In Silico Analysis for Amino Acid Substitution | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MACd | MAFe | MAC | MAF | SIFT | Polyphen-2 | PMut | ||||||
| 16 | 15761214 | 4 | C > T | Registered | T52M | 0 | 0 | 1 | 0.0035 | tolerated | benign | pathological |
| 16 | 15785033 | 7 | C > G | Not registered | Q186E | 1 | 0.0012 | 0 | 0 | tolerated | benign | neutral |
| 16 | 15785118 | 7 | C > T | Not registered | S214F | 1 | 0.0012 | 0 | 0 | damaging | possibly damaging | pathological |
| 16 | 15785177 | 7 | C > T | Not registered | R234C | 1 | 0.0012 | 0 | 0 | tolerated | benign | pathological |
Note: SCZ, schizophrenia; PDD, pervasive developmental disorder.
a Genomic position based on NCBI build 37.1(Transcript ID ENST00000396355).
b M, major allele; m, minor allele.
c Amino acid change based on NCBI reference sequence NP_060138.1.
d Minor allele count.
e Minor allele frequency.
Fig. 1.
Locations of novel rare mutations and results of conservation analysis. (A) NDE1 gene structure based on ENST00000396355; yellow boxes indicate the protein-coding exons sequenced in this study. Gray boxes indicate the 5′ and 3′ untranslated regions (UTR). We discovered 3 rare novel mutations in exon7. (B) NDE1 protein structure (335 amino acids); S214F and R234C were located between the N-terminal coiled-coil region and C-terminal α-helix region.49 Several phosphorylation sites (red sticks) have been identified in the peptide regions encoded by exon7 of NDE1, and S214 is one of those phosphorylation sites. (C) Traces of Sanger sequence illustrating the novel rare variants. Arrows indicate the mutated sites. (D) Multiple alignments of amino acid sequences from NDE1 homologs; these alignments include the novel mutations.
Results of Genetic Association Analysis
The results of genetic association analysis of novel missense mutations are shown in table 2. We detected S214F in 6 SCZ cases and 1 HC, and we observed a significant association between SCZ and S214F (OR = 7.1, P = .039, adjusted P = .052). The clinical feature of 6 SCZ cases with S214F is in supplementary table S4. In addition, we discovered R234C in 1 SCZ case and 2 BD cases, but none in HC, and detected a significant association between BD and R234C (P = .032, adjusted P = .033). The MAF of Q186E in HC (0.24%) was higher than the MAF in SCZ (0.15%) and BD (0.098%).
Table 2.
Association Analysis of Novel Rare Variants
| Variant | Genomic Data | Schizophrenia | Bipolar Disorder | Healthy Control | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Position | Ma/ma | MAFb | Genotype Countc | P valued | Odds Ratio | MAF | Genotype Count | P value | Odds Ratio | MAF | Genotype Count | |
| Q186E | 16:15785033 | C/G | 0.0015 | 0/12/3948 | .15 | 0.65 | 0.0098 | 0/2/1025 | .17 | 0.41 | 0.0024 | 0/22/4679 |
| S214F | 16:15785118 | C/T | 0.00076 | 0/6/3954 | .039 | 7.1 | 0 | 0/0/1034 | .82 | NA | 0.00011 | 0/1/4693 |
| R234C | 16:15785177 | C/T | 0.00013 | 0/1/3960 | .46 | NAe | 0.0097 | 0/2/1029 | .032 | NA | 0 | 0/0/4693 |
Note: a M, major; m, minor.
b MAF, minor allele frequency.
c Genotype count; homozygote of minor allele/heterozygote/homozygote of major allele (Genotype count of SCZ was considering total sample number).
d P values were calculated by Fisher’s exact test (2×2 contingency table, one-tail)
e NA, not available
Functional Analysis of NDE1 Mutations
Of the 3 novel rare mutations, only S214F and R234C were predicted to be deleterious based on in silico analysis (table 1); they were also the two with a significant association with SCZ or BD (table 2). Notably, even adjusted P value (100 000 permutations) showed trend for statistical association. Therefore, we focused on these 2 rare missense mutations for further functional assays.
We considered that the S214F variant of NDE1 could contribute to the pathophysiology of SCZ by affecting NDE1 protein functions based on the following: (1) NDE1 and the homolog protein NDEL1 are highly similar35 and are centrosomal phosphoproteins with reported roles in cell proliferation and migration28,32; (2) YWHAE, which is one of the DISC1 interacting proteins and is closely related to neuronal migration and brain development, binds to phosphorylated NDEL1 and is required for normal NDEL1 localization and functions38; (3) the reported phosphorylation sites of NDEL1 for binding with YWHAE are conserved around S214 of NDE1 (figure 2A); and (4) the aromatic ring of NDE1 F214 might have steric effects on T215, which is suggested to be the phosphorylation site responsible for binding to YWHAE38 (supplementary figure S3).
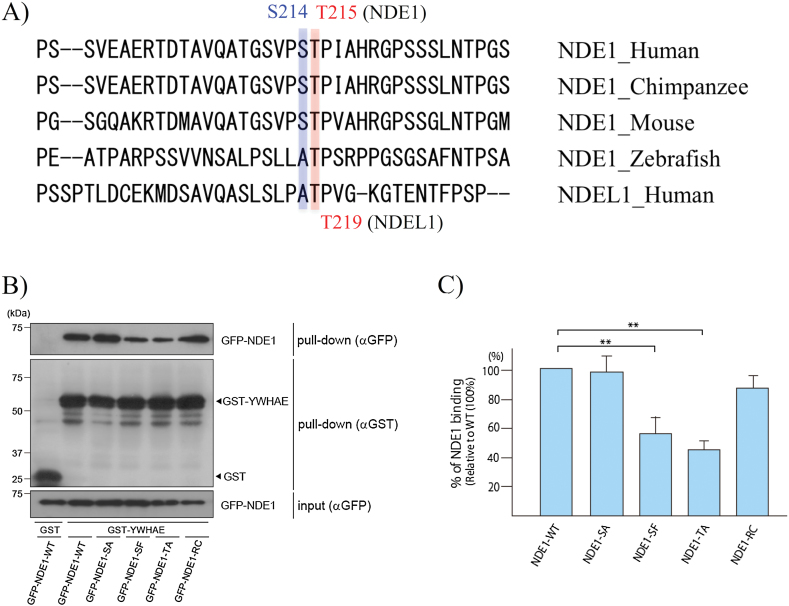
Fig. 2.
Evaluation of interaction between YWHAE and NDE1-mutants. Legend: (A) Sequence alignment between the YWHAE-binding site of human NDEL1 and NDE1 orthologs of different species. The T215 residue is highly conserved among some species. T219 of NDEL1 (the counterpart of NDEL1 T219 is NDE1 T215) is also conserved in several species, and had been supposed to be the most important site for binding with YWHAE.38 (B) Representative immunoblots of total cell lysates and the proteins pulled down by GST-YWHAE. A portion (10%) of each eluate and the inputs (5%) were subjected to SDS-PAGE and subsequent immunoblot analyses with the indicated antibodies. (C) Graph of the relative amount of bound NDE1 normalized to the amount of pulled-down GST-YWHAE. To quantify the effects of each NDE1 mutation on the interaction with YWHAE, we performed the GST-pulldown assay in triplicate. The GST-pulldown assay revealed a reduced interaction of YWHAE with S214F and T215A compared with WT (double asterisks: P < .01). No difference was detected between S214A and WT, or between R234C and WT. Each bar represents the mean of the triplicate experiments, and the error bars indicate ±SE.
To investigate the effect of S214F and that of R234C on the interaction with YWHAE, we used HEK293T cells to perform GST pull-down assays. The results are shown in figures 2B and 2C. As expected from previous studies of NDEL1 (34), NDE1 interacted with YWHAE. We also discovered a reduced interaction between YWHAE and S214F or T215A mutants relative to WT (P < .01); the T215A mutant was used as a positive control because it cannot be phosphorylated at T215A.38 However, no significant difference was detected in the interaction of R234C and the S214A (constructed as a mutation with less steric effects than F214), compared with WT. To further examine in vivo interactions between NDE1-WT and the NDE1 mutants, we performed a GST pull-down assay using lysates for COS7 cells that co-expressed GST-YWHAE and GFP-NDE1 (supplementary figure S4). The decreased effects of S214F and T215A on the interaction with YWHAE was also observed in the assay.
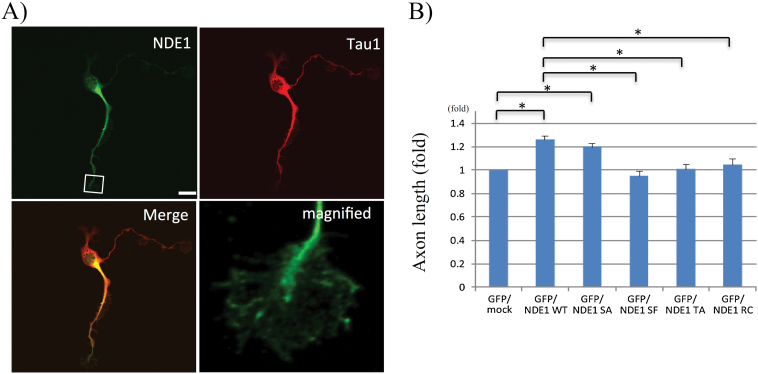
To examine the effect of S214F and that of R234C on neurodevelopment, we performed a morphological study of nascent hippocampal neurons, as NDE1 is reportedly known to contribute to neurite extension.48 We found that endogenous NDE1 localized at nascent axons in rat hippocampal neurons (figure 3A), and that overexpression of NDE1-WT or NDE1-S214A increased the axon length, while that of NDE1-T215A, NDE1-S214F or NDE1-R234C did not (figure 3B).
Fig. 3.
Effect of NDE1-mutants on axonal outgrowth in hippocampal neurons. (A) Colabeling of NDE1 (green) and Tau1 protein (red) in nascent neurons (DIV2). Tau1 is used as an axon marker. The image in the lower right panel is a magnified view of the tip of an axonal process (rectangle in the upper left panel). Scale bar, 15 μm. (B) Quantitative analysis of axon length in transfected neurons. Hippocampal neurons transfected with the indicated plasmids encoding EGFP and Myc-NDE1 at DIV1 were detected by EGFP fluorescence and staining of Tau-1, which is a maker of axons in DIV3 cells. Each bar shows the mean of axon length in the transfectants. Error bars represent ±SE. n = 60, data sets from 3 independent experiments. Asterisks indicate statistical significance (P < .05).
Discussion
In this study, we conducted exon sequencing of NDE1 as a candidate gene for SCZ, ASD, and BD. After sequencing more than 5.7 × 102 kb, we discovered 3 novel rare missense mutations in exon 7 of the NDE1 gene in SCZ cases. NDE1 exon 7 is located C-terminalα-helix where many phosphorylation sites are located (figures 1A and 1B), and predicted to regulate the NDE1 structure and interactions with other proteins.31,49 Moreover, it was suggested disease associated mutations cluster within protein interacting domains.50 In the current study, we could detect a significant association (OR = 7.1, P = .039) between SCZ and S214F in the sample comprising 8734 unrelated individuals. Post hoc calculations of statistical power showed that our total SCZ sample (n = 3987) had sufficient statistical power (1-β > 80%) for SNV with MAF (0.12%) detected within our SCZ resequencing samples (table 1) if the relative risk was >2.7. This result is consistent with the recent exome sequencing study of large SCZ samples, which suggested that rare mutations (MAF < 0.1%) with large effect are significantly enriched in SCZ candidate genes, especially synapse developing related genes.12 However, the 6 cases with S214F did not seem to share any specific clinical features (supplementary table S4). Considering a high OR, a part of S214F mutations might be a de novo (ie the parents of patients with NDE1 S214F did not suffer from SCZ).
Furthermore, we could find the probable biological effect of S214F, which was predicted to be deleterious from several bioinformatics tools, by investigating the interaction with YWHAE. In the pull-down assay using WT and mutant NDE1, the S214A mutation had no effect on the interaction between NDE1 and YWHAE, although the S214 residue is a phosphorylation site of NDE1 according to the PhosphoSitePlus database (http://www.phosphosite.org/homeAction.do). This result suggests that the interaction between NDE1 and YWHAE is independent of the phosphorylation state of NDE1 at S214. In contrast to S214A and R234C, the S214F and T215A mutations impaired the interaction between NDE1 and YWHAE. The mechanism of reduced interaction is probably a steric effect of F214 on T215 (supplementary figure S3), which is suggested to be the phosphorylation site responsible for binding to YWHAE.38 Furthermore, we found that S214F mutant did not promote axon elongation (figure 3). We believe that changes in axon length, associated with the S214F mutant, could be related to the reduction in the interaction with YWHAE. This hypothesis is based on several findings: (1) NDE1 binds to YWHAE and localizes at axon tips; (2) YWHAE is required for the proper localization of NDEL1, a dimeric partner of NDE1,38 and is involved in the axon elongation by conveying the protein complex including YWHAE and NDEL1.47 Importantly, NDE1 and YWHAE are expressed beginning in the early embryonic period (http://hbatlas.org) (supplementary figure S5).51 Furthermore, we previously identified a SNP (rs28365859) in YWHAE that is significantly associated with SCZ,39 and subjects carrying the minor allele (C) had increased mRNA transcription and protein expression compared to those with the major allele (G).39 Thus, in a future study, the epistatic effect of the YWHAE allele (rs28365859) with NDE1 S214F should also be investigated.
R234C missense mutation in NDE1 had an effect on axonal outgrowth (figure 3). Although the contribution of R234C mutation to neurodevelopment remains unclear, the change of the axon length by R234C might relate to NDE1 structure or protein-protein interactions. In fact, several proteins besides YWHAE bind to this region of NDE1, including the S214 and R234 resides; CENPF,52 Su48,31 and Katanin p60 are among these.53 Furthermore, in silico analysis with Splice Aid 254 indicates that the nucleotide change causing R234C may be located within an exon-splicing enhancer (ESE) motif54 and may decrease mRNA transcription because the alternative splicing might result in exon skipping during transcription. Indeed, 16%–20% of missense mutations may disrupt ESEs and change the splicing pattern of the gene.55 In a preliminary analysis of expression in lymphoblastoid cell lines, transcription levels of the R234C mRNA were between the first quartile and the minimum in 60 SCZ cases (supplementary methods and supplementary figure S6).
Interestingly, we discovered R234C in 2 BD patients and 1 SCZ patient. Therefore, the rare NDE1 variants may be common genetic factor not only for SCZ and BD. Mounting evidence from several types of studies, including GWASs56,57 and epidemiological family studies,3 indicates shared genetic components between SCZ and BD. In 1 GWAS, the authors combined SCZ and BD samples so that they could increase the statistical power and detect shared risk SNPs.56 Therefore, further replication studies for rare variants with putative large effect sizes should be performed in order to find shared pathophysiology between SCZ and BD.
Q186E was located in a highly conserved genomic region, but was discovered more frequently in HC and was judged to be neutral by several software packages for predicting mutation effects. Furthermore, in segregation analysis, we could obtain the DNA from the patient’s son who was also suffering from SCZ (supplementary results), but he did not carry the NDE1 Q186E variant.
There are several limitations to be discussed. First, several potentially valuable regions of NDE1, including intronic regions, the promoter, and 5′ and 3′ untranslated regions ends, were not sequenced. Second, we could not fully conduct segregation analyses for mutations due to limited access to the subjects’ family members. This made it impossible for us to accurately measure the inheritance of mutation in the pedigree. Furthermore, in the case summary of SCZ with novel rare variants, because we could not obtain detailed clinical information for the developmental period, the effect of discovered rare mutations could not be fully evaluated. Third, we could not perform structural analysis of S214F and R234C because these mutations are located at the C-terminal region, an area known to be difficult to predict the structure of.49,58 Future studies should focus on the structural effect of the mutations discovered in this study.
In conclusion, screening for mutations in NDE1, a candidate risk gene for SCZ and ASD based on CNV analysis, revealed novel rare missense mutations that might increase susceptibility to SCZ and BD. Importantly, we detected a significant association between SCZ and S214F, which was found to have the biological effects of changing the interaction between NDE1 and YWHAE and the axonal outgrowth. This study therefore strengthens the evidence for the role of rare variants in NDE1 in the etiopathological role of SCZ, and BD. The present results also suggest that deep sequencing of candidate genes will be a promising method, especially using current and developing sequencing technology for elucidating the missing heritability and pathogenesis of neuropsychiatric disorders.
Supplementary Material
Supplementary material is available at http://schizophreniabulletin.oxfordjournals.org.
Funding
This work was supported by research grants from the Ministry of Education, Culture, Sports, Science and Technology of Japan ; the Ministry of Health, Labor and Welfare of Japan; a Grant-in-Aid for “Integrated research on neuropsychiatric disorders” carried out under the Strategic Research Program for Brain Sciences by the Ministry of Education, Culture, Sports, Science and Technology of Japan; a Grant-in-Aid for Scientific Research on Innovative Areas, “Glial assembly: a new regulatory machinery of brain function and disorders”; Grant-in-Aid for Scientific Research on Innovative Areas (Comprehensive Brain Science Network) from the Ministry of Education, Science, Sports and Culture of Japan; and a Grant-in-Aid for Encouragement of young scientists Numbers 25933012.
Supplementary Material
Acknowledgment
The authors have no financial conflicts to declare.
References
- 1.Collins PY, Patel V, Joestl SS, et al. Grand challenges in global mental health. Nature. 2011;475:27–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Sullivan PF, Kendler KS, Neale MC. Schizophrenia as a complex trait: evidence from a meta-analysis of twin studies. Arch Gen Psychiatry. 2003;60:1187–1192. [DOI] [PubMed] [Google Scholar]
- 3.Lichtenstein P, Yip BH, Björk C, et al. Common genetic determinants of schizophrenia and bipolar disorder in Swedish families: a population-based study. Lancet. 2009;373:234–239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sullivan PF, Daly MJ, O’Donovan M. Genetic architectures of psychiatric disorders: the emerging picture and its implications. Nat Rev Genet. 2012;13:537–551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ripke S, O’Dushlaine C, Chambert K, et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat Genet. 2013;45:1150–1159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Malhotra D, Sebat J. CNVs: harbingers of a rare variant revolution in psychiatric genetics. Cell. 2012;148:1223–1241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Timms AE, Dorschner MO, Wechsler J, et al. Support for the N-methyl-D-aspartate receptor hypofunction hypothesis of schizophrenia from exome sequencing in multiplex families. JAMA Psychiatry. 2013;70:582–590. [DOI] [PubMed] [Google Scholar]
- 8.Hamilton PJ, Campbell NG, Sharma S, et al. De novo mutation in the dopamine transporter gene associates dopamine dysfunction with autism spectrum disorder. Mol Psychiatry. 2013;18:1315–1323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Mondal K, Ramachandran D, Patel VC, et al. Excess variants in AFF2 detected by massively parallel sequencing of males with autism spectrum disorder. Hum Mol Genet. 2012;21:4356–4364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Durand CM, Perroy J, Loll F, et al. SHANK3 mutations identified in autism lead to modification of dendritic spine morphology via an actin-dependent mechanism. Mol Psychiatry. 2012;17:71–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Szatkiewicz JP, O’Dushlaine C, Chen G, et al. Copy number variation in schizophrenia in Sweden. Mol Psychiatry. 2014;19:762–773. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Purcell SM, Moran JL, Fromer M, et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature. 2014;506:185–190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Moessner R, Marshall CR, Sutcliffe JS, et al. Contribution of SHANK3 mutations to autism spectrum disorder. Am J Hum Genet. 2007;81:1289–1297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Durand CM, Betancur C, Boeckers TM, et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet. 2007;39:25–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bonaglia MC, Giorda R, Borgatti R, et al. Disruption of the ProSAP2 gene in a t(12;22)(q24.1;q13.3) is associated with the 22q13.3 deletion syndrome. Am J Hum Genet. 2001;69:261–268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ullmann R, Turner G, Kirchhoff M, et al. Array CGH identifies reciprocal 16p13.1 duplications and deletions that predispose to autism and/or mental retardation. Hum Mutat. 2007;28:674–682. [DOI] [PubMed] [Google Scholar]
- 17.Mefford HC, Muhle H, Ostertag P, et al. Genome-wide copy number variation in epilepsy: novel susceptibility loci in idiopathic generalized and focal epilepsies. PLoS Genet. 2010;6:e1000962. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Heinzen EL, Radtke RA, Urban TJ, et al. Rare deletions at 16p13.11 predispose to a diverse spectrum of sporadic epilepsy syndromes. Am J Hum Genet. 2010;86:707–718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.de Kovel CG, Trucks H, Helbig I, et al. Recurrent microdeletions at 15q11.2 and 16p13.11 predispose to idiopathic generalized epilepsies. Brain. 2010;133:23–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ingason A, Rujescu D, Cichon S, et al. Copy number variations of chromosome 16p13.1 region associated with schizophrenia. Mol Psychiatry. 2011;16:17–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ikeda M, Aleksic B, Kirov G, et al. Copy number variation in schizophrenia in the Japanese population. Biol Psychiatry. 2010;67:283–286. [DOI] [PubMed] [Google Scholar]
- 22.Kirov G, Grozeva D, Norton N, et al. Support for the involvement of large copy number variants in the pathogenesis of schizophrenia. Hum Mol Genet. 2009;18:1497–1503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Nagamani SC, Erez A, Bader P, et al. Phenotypic manifestations of copy number variation in chromosome 16p13.11. Eur J Hum Genet. 2011;19:280–286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hannes FD, Sharp AJ, Mefford HC, et al. Recurrent reciprocal deletions and duplications of 16p13.11: the deletion is a risk factor for MR/MCA while the duplication may be a rare benign variant. J Med Genet. 2009;46:223–232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Williams NM, Norton N, Williams H, et al. A systematic genomewide linkage study in 353 sib pairs with schizophrenia. Am J Hum Genet. 2003;73:1355–1367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.International Molecular Genetic Study of Autism C. A genomewide screen for autism: strong evidence for linkage to chromosomes 2q, 7q, and 16p. Am J Hum Genet. 2001;69:570–581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lamb JA, Barnby G, Bonora E, et al. Analysis of IMGSAC autism susceptibility loci: evidence for sex limited and parent of origin specific effects. J Med Genet. 2005;42:132–137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Bakircioglu M, Carvalho OP, Khurshid M, et al. The essential role of centrosomal NDE1 in human cerebral cortex neurogenesis. Am J Hum Genet. 2011;88:523–535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Feng Y, Walsh CA. Mitotic spindle regulation by Nde1 controls cerebral cortical size. Neuron. 2004;44:279–293. [DOI] [PubMed] [Google Scholar]
- 30.Tarricone C, Perrina F, Monzani S, et al. Coupling PAF signaling to dynein regulation: structure of LIS1 in complex with PAF-acetylhydrolase. Neuron. 2004;44:809–821. [DOI] [PubMed] [Google Scholar]
- 31.Hirohashi Y, Wang Q, Liu Q, et al. Centrosomal proteins Nde1 and Su48 form a complex regulated by phosphorylation. Oncogene. 2006;25:6048–6055. [DOI] [PubMed] [Google Scholar]
- 32.Alkuraya FS, Cai X, Emery C, et al. Human mutations in NDE1 cause extreme microcephaly with lissencephaly [corrected]. Am J Hum Genet. 2011;88:536–547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Hennah W, Tomppo L, Hiekkalinna T, et al. Families with the risk allele of DISC1 reveal a link between schizophrenia and another component of the same molecular pathway, NDE1. Hum Mol Genet. 2007;16:453–462. [DOI] [PubMed] [Google Scholar]
- 34.Brandon NJ, Sawa A. Linking neurodevelopmental and synaptic theories of mental illness through DISC1. Nat Rev Neurosci. 2011;12:707–722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Efimov VP, Morris NR. The LIS1-related NUDF protein of Aspergillus nidulans interacts with the coiled-coil domain of the NUDE/RO11 protein. J Cell Biol. 2000;150:681–688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Bradshaw NJ, Christie S, Soares DC, et al. NDE1 and NDEL1: multimerisation, alternate splicing and DISC1 interaction. Neurosci Lett. 2009;449:228–233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Burdick KE, Kamiya A, Hodgkinson CA, et al. Elucidating the relationship between DISC1, NDEL1 and NDE1 and the risk for schizophrenia: evidence of epistasis and competitive binding. Hum Mol Genet. 2008;17:2462–2473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Toyo-oka K, Shionoya A, Gambello MJ, et al. 14-3-3epsilon is important for neuronal migration by binding to NUDEL: a molecular explanation for Miller-Dieker syndrome. Nat Genet. 2003;34:274–285. [DOI] [PubMed] [Google Scholar]
- 39.Ikeda M, Hikita T, Taya S, et al. Identification of YWHAE, a gene encoding 14-3-3epsilon, as a possible susceptibility gene for schizophrenia. Hum Mol Genet. 2008;17:3212–3222. [DOI] [PubMed] [Google Scholar]
- 40.Ikeda M, Aleksic B, Kinoshita Y, et al. Genome-wide association study of schizophrenia in a Japanese population. Biol Psychiatry. 2011;69:472–478. [DOI] [PubMed] [Google Scholar]
- 41.Dong C, Yu B. Mutation surveyor: an in silico tool for sequencing analysis. Methods Mol Biol. 2011;760:223–237. [DOI] [PubMed] [Google Scholar]
- 42.Adzhubei IA, Schmidt S, Peshkin L, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kumar P, Henikoff S, Ng PC. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc. 2009;4:1073–1081. [DOI] [PubMed] [Google Scholar]
- 44.Ferrer-Costa C, Gelpi JL, Zamakola L, et al. PMUT: a web-based tool for the annotation of pathological mutations on proteins. Bioinformatics. 2005;21:3176–3178. [DOI] [PubMed] [Google Scholar]
- 45.Purcell S, Neale B, Todd-Brown K, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Funahashi Y, Namba T, Fujisue S, et al. ERK2-mediated phosphorylation of Par3 regulates neuronal polarization. J Neurosci. 2013;33:13270–13285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Taya S, Shinoda T, Tsuboi D, et al. DISC1 regulates the transport of the NUDEL/LIS1/14-3-3epsilon complex through kinesin-1. J Neurosci. 2007;27:15–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Bradshaw NJ, Soares DC, Carlyle BC, et al. PKA phosphorylation of NDE1 is DISC1/PDE4 dependent and modulates its interaction with LIS1 and NDEL1. J Neurosci. 2011;31:9043–9054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Soares DC, Bradshaw NJ, Zou J, et al. The mitosis and neurodevelopment proteins NDE1 and NDEL1 form dimers, tetramers, and polymers with a folded back structure in solution. J Biol Chem. 2012;287:32381–32393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Wang X, Wei X, Thijssen B, et al. Three-dimensional reconstruction of protein networks provides insight into human genetic disease. Nat Biotechnol. 2012;30:159–164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kang HJ, Kawasawa YI, Cheng F, et al. Spatio-temporal transcriptome of the human brain. Nature. 2011;478:483–489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Soukoulis V, Reddy S, Pooley RD, et al. Cytoplasmic LEK1 is a regulator of microtubule function through its interaction with the LIS1 pathway. Proc Natl Acad Sci U S A. 2005;102:8549–8554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Toyo-Oka K, Sasaki S, Yano Y, et al. Recruitment of katanin p60 by phosphorylated NDEL1, an LIS1 interacting protein, is essential for mitotic cell division and neuronal migration. Hum Mol Genet. 2005;14:3113–3128. [DOI] [PubMed] [Google Scholar]
- 54.Piva F, Giulietti M, Burini AB, et al. SpliceAid 2: a database of human splicing factors expression data and RNA target motifs. Hum Mutat. 2012;33:81–85. [DOI] [PubMed] [Google Scholar]
- 55.Gorlov IP, Gorlova OY, Frazier ML, Amos CI. Missense mutations in hMLH1 and hMSH2 are associated with exonic splicing enhancers. Am J Hum Genet. 2003;73:1157–1161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Smoller JW, Craddock N, Kendler K, et al. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet. 2013;381:1371–1379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Williams HJ, Craddock N, Russo G, et al. Most genome-wide significant susceptibility loci for schizophrenia and bipolar disorder reported to date cross-traditional diagnostic boundaries. Hum Mol Genet. 2011;20:387–391. [DOI] [PubMed] [Google Scholar]
- 58.Bradshaw NJ, Ogawa F, Antolin-Fontes B, et al. DISC1, PDE4B, and NDE1 at the centrosome and synapse. Biochem Biophys Res Commun. 2008;377:1091–1096. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.