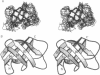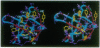Abstract
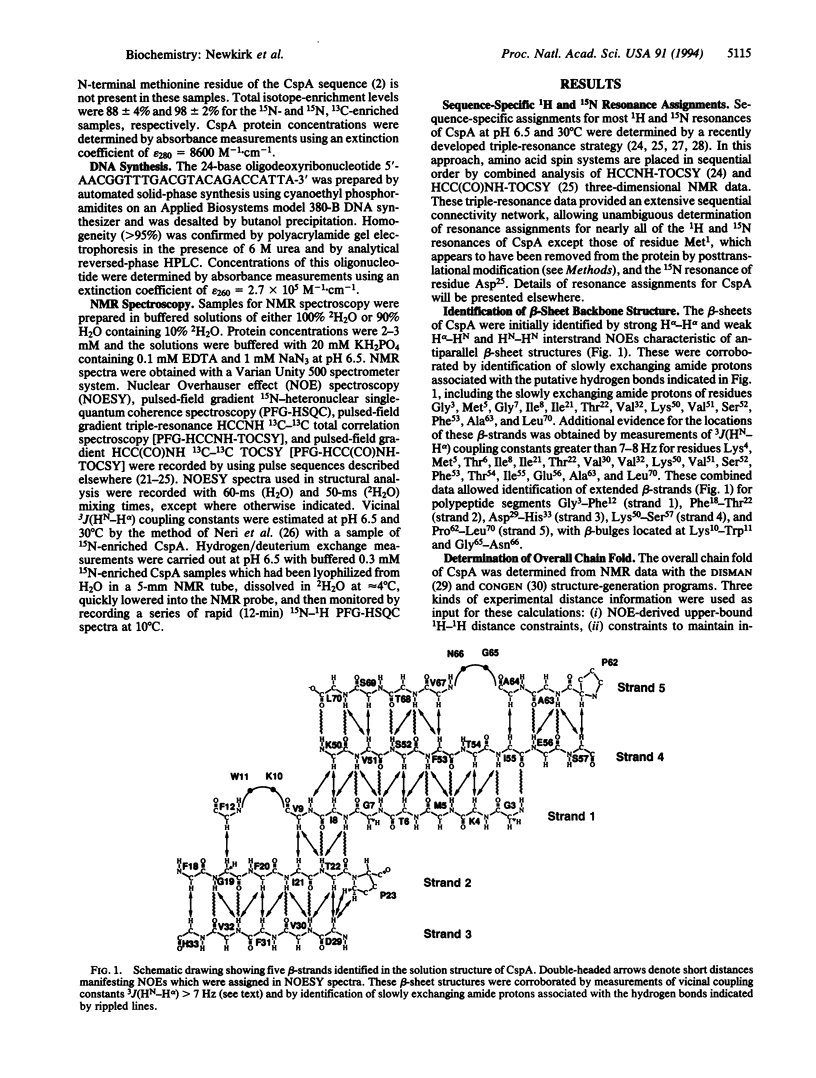
Sequence-specific 1H and 15N resonance assignments have been determined for the major cold shock protein (CspA) from Escherichia coli with recently developed three-dimensional triple-resonance NMR experiments. By use of these assignments, five antiparallel beta-strands were identified from analysis of NMR data. Strands 1-4 have a classical 3-2-1-4 Greek key beta-sheet topology and there are two beta-bulges, at positions Lys10-Trp11 and Gly65-Asn66. Three-dimensional structures of CspA were generated from NMR data by using simulated annealing with molecular dynamics. The overall chain fold of CspA is a beta-barrel structure, with a tightly packed hydrophobic core. Two-dimensional isotope-edited pulsed-field gradient 15N-1H heteronuclear single-quantum coherence spectroscopy was used to characterize the 15N-1H fingerprint spectrum with and without a 24-base oligodeoxyribonucleotide, 5'-AACGGTTTGACGTACAGACCATTA-3'. Protein-DNA complex formation perturbs a subset of the amide resonances that are located mostly on one face of the CspA molecule. This portion of the CspA molecular surface includes two putative RNA-binding sequence motifs which contribute to an unusual cluster of eight surface aromatic side chains: Trp11, Phe12, Phe18, Phe20, Phe31, His33, Phe34, and Tyr42. These surface aromatic groups, and also residues Lys16, Ser44, and Lys60 located on this same face of CspA, are highly conserved in the family of CspA homologues. These isotope-edited pulsed-field gradient NMR data provide a low-resolution mapping of a DNA-binding epitope on CspA.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adam S. A., Nakagawa T., Swanson M. S., Woodruff T. K., Dreyfuss G. mRNA polyadenylate-binding protein: gene isolation and sequencing and identification of a ribonucleoprotein consensus sequence. Mol Cell Biol. 1986 Aug;6(8):2932–2943. doi: 10.1128/mcb.6.8.2932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Braun W., Go N. Calculation of protein conformations by proton-proton distance constraints. A new efficient algorithm. J Mol Biol. 1985 Dec 5;186(3):611–626. doi: 10.1016/0022-2836(85)90134-2. [DOI] [PubMed] [Google Scholar]
- Bruccoleri R. E., Karplus M. Conformational sampling using high-temperature molecular dynamics. Biopolymers. 1990 Dec;29(14):1847–1862. doi: 10.1002/bip.360291415. [DOI] [PubMed] [Google Scholar]
- Chatterjee S., Jiang W., Emerson S. D., Inouye M. The backbone structure of the major cold-shock protein CS7.4 of Escherichia coli in solution includes extensive beta-sheet structure. J Biochem. 1993 Nov;114(5):663–669. doi: 10.1093/oxfordjournals.jbchem.a124234. [DOI] [PubMed] [Google Scholar]
- Goldstein J., Pollitt N. S., Inouye M. Major cold shock protein of Escherichia coli. Proc Natl Acad Sci U S A. 1990 Jan;87(1):283–287. doi: 10.1073/pnas.87.1.283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Görlach M., Wittekind M., Beckman R. A., Mueller L., Dreyfuss G. Interaction of the RNA-binding domain of the hnRNP C proteins with RNA. EMBO J. 1992 Sep;11(9):3289–3295. doi: 10.1002/j.1460-2075.1992.tb05407.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones P. G., Inouye M. The cold-shock response--a hot topic. Mol Microbiol. 1994 Mar;11(5):811–818. doi: 10.1111/j.1365-2958.1994.tb00359.x. [DOI] [PubMed] [Google Scholar]
- Jones P. G., VanBogelen R. A., Neidhardt F. C. Induction of proteins in response to low temperature in Escherichia coli. J Bacteriol. 1987 May;169(5):2092–2095. doi: 10.1128/jb.169.5.2092-2095.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- La Teana A., Brandi A., Falconi M., Spurio R., Pon C. L., Gualerzi C. O. Identification of a cold shock transcriptional enhancer of the Escherichia coli gene encoding nucleoid protein H-NS. Proc Natl Acad Sci U S A. 1991 Dec 1;88(23):10907–10911. doi: 10.1073/pnas.88.23.10907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee S. J., Xie A., Jiang W., Etchegaray J. P., Jones P. G., Inouye M. Family of the major cold-shock protein, CspA (CS7.4), of Escherichia coli, whose members show a high sequence similarity with the eukaryotic Y-box binding proteins. Mol Microbiol. 1994 Mar;11(5):833–839. doi: 10.1111/j.1365-2958.1994.tb00361.x. [DOI] [PubMed] [Google Scholar]
- Logan T. M., Olejniczak E. T., Xu R. X., Fesik S. W. Side chain and backbone assignments in isotopically labeled proteins from two heteronuclear triple resonance experiments. FEBS Lett. 1992 Dec 21;314(3):413–418. doi: 10.1016/0014-5793(92)81517-p. [DOI] [PubMed] [Google Scholar]
- Lyons B. A., Tashiro M., Cedergren L., Nilsson B., Montelione G. T. An improved strategy for determining resonance assignments for isotopically enriched proteins and its application to an engineered domain of staphylococcal protein A. Biochemistry. 1993 Aug 10;32(31):7839–7845. doi: 10.1021/bi00082a001. [DOI] [PubMed] [Google Scholar]
- Mattaj I. W. RNA recognition: a family matter? Cell. 1993 Jun 4;73(5):837–840. doi: 10.1016/0092-8674(93)90265-r. [DOI] [PubMed] [Google Scholar]
- Murray M. T., Schiller D. L., Franke W. W. Sequence analysis of cytoplasmic mRNA-binding proteins of Xenopus oocytes identifies a family of RNA-binding proteins. Proc Natl Acad Sci U S A. 1992 Jan 1;89(1):11–15. doi: 10.1073/pnas.89.1.11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagai K., Oubridge C., Jessen T. H., Li J., Evans P. R. Crystal structure of the RNA-binding domain of the U1 small nuclear ribonucleoprotein A. Nature. 1990 Dec 6;348(6301):515–520. doi: 10.1038/348515a0. [DOI] [PubMed] [Google Scholar]
- Sachs A. B., Bond M. W., Kornberg R. D. A single gene from yeast for both nuclear and cytoplasmic polyadenylate-binding proteins: domain structure and expression. Cell. 1986 Jun 20;45(6):827–835. doi: 10.1016/0092-8674(86)90557-x. [DOI] [PubMed] [Google Scholar]
- Schindelin H., Jiang W., Inouye M., Heinemann U. Crystal structure of CspA, the major cold shock protein of Escherichia coli. Proc Natl Acad Sci U S A. 1994 May 24;91(11):5119–5123. doi: 10.1073/pnas.91.11.5119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schindelin H., Marahiel M. A., Heinemann U. Universal nucleic acid-binding domain revealed by crystal structure of the B. subtilis major cold-shock protein. Nature. 1993 Jul 8;364(6433):164–168. doi: 10.1038/364164a0. [DOI] [PubMed] [Google Scholar]
- Schnuchel A., Wiltscheck R., Czisch M., Herrler M., Willimsky G., Graumann P., Marahiel M. A., Holak T. A. Structure in solution of the major cold-shock protein from Bacillus subtilis. Nature. 1993 Jul 8;364(6433):169–171. doi: 10.1038/364169a0. [DOI] [PubMed] [Google Scholar]
- Tafuri S. R., Wolffe A. P. DNA binding, multimerization, and transcription stimulation by the Xenopus Y box proteins in vitro. New Biol. 1992 Apr;4(4):349–359. [PubMed] [Google Scholar]
- Tafuri S. R., Wolffe A. P. Xenopus Y-box transcription factors: molecular cloning, functional analysis and developmental regulation. Proc Natl Acad Sci U S A. 1990 Nov;87(22):9028–9032. doi: 10.1073/pnas.87.22.9028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanabe H., Goldstein J., Yang M., Inouye M. Identification of the promoter region of the Escherichia coli major cold shock gene, cspA. J Bacteriol. 1992 Jun;174(12):3867–3873. doi: 10.1128/jb.174.12.3867-3873.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Willimsky G., Bang H., Fischer G., Marahiel M. A. Characterization of cspB, a Bacillus subtilis inducible cold shock gene affecting cell viability at low temperatures. J Bacteriol. 1992 Oct;174(20):6326–6335. doi: 10.1128/jb.174.20.6326-6335.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolffe A. P., Tafuri S., Ranjan M., Familari M. The Y-box factors: a family of nucleic acid binding proteins conserved from Escherichia coli to man. New Biol. 1992 Apr;4(4):290–298. [PubMed] [Google Scholar]