Abstract
The Eco29k I restriction endonuclease is a Sac II isoschizomer that recognizes the sequence 5’-CCGCGG-3’ and is encoded, along with the Eco29k I methylase, in the Escherichia coli strain 29k. We have expressed the Eco29k I restriction-methylation system (RM2) in E. coli strain TG1 to produce the strain AXE688. We have developed a directed molecular evolution (DME) mutagenesis method that uses Eco29k I to restrict incoming parental DNA in transformed cells. Using our DME method, we have demonstrated that our AXE688 strain results in mutated directed molecular evolution libraries with diversity greater than 107 from a single transformation and with greater than 90% recombinant clones.
1. Introduction
Directed evolution involves the generation of random genetic diversity followed by high-throughput screening to isolate antibodies or engineered proteins with desired affinity, stability, specificity, and in some cases, catalytic activity. Phage display is a commonly used method for directed evolution because there is a physical linkage between genotype and phenotype (Smith and Scott, 1993). A basic requirement for this technique is a diverse nucleotide library (Jackel et al., 2008). Numerous methods have been developed to introduce diversity, including random mutagenesis (Fromant et al., 1995), DNA shuffling (Stemmer, 1994), and site-specific recombination (Waterhouse et al., 1993).
Random mutagenesis methods use a single gene as a starting point and introduce mutations along the entire gene or in predefined sites or regions. The most common method of producing genetic variation is by error-prone PCR. This technique relies on the misincorporation of nucleotides by DNA polymerase to generate point mutations in a gene sequence (Martineau, 2002). The accuracy of the DNA polymerase can be adjusted by addition of manganese ions in the reaction mixture (Cadwell and Joyce, 1994), incorporation of nucleotide analogs (Zaccolo et al., 1996), or the use of E. coli mutator strains or mutagenic polymerases (Coia et al., 1997; Emond et al., 2008).
Diversity can also be introduced in a targeted fashion at specific functional sites, such as the active site region of an enzyme or antibody hypervariable domains (Foote and Winter, 1992). Randomization of a specific site is achieved by using synthetic oligonucleotides possessing randomized codon(s) flanked by wild type sequence that can anneal to the parent gene template. Several codons can be randomized simultaneously, but this method relies heavily on structural insights for determining which sites to mutate. Several site-specific mutagenesis procedures are available for efficient incorporation of mutagenized primers, such as the ‘QuikChange’ method (Weiner et al., 1994; Agilent Technologies).
A commonly used site-directed mutagenesis method in phage display is ‘Kunkel mutagenesis’ (Kunkel, 1985; Scholle et al., 2005; Fellouse et al., 2007; Huang et al., 2012), where a uracil-incorporated, circular, single-stranded DNA (ssDNA) serves as a template to synthesize double-stranded DNA (dsDNA) in vitro with an oligonucleotide primer that introduces a mutation. After dsDNA is introduced into bacteria, recombinant clones predominate due to cleavage and repairbias of the uracilated strand in vivo.
We previously developed a mutagenesis technique that combines the universal nature of error-prone PCR with the efficiency of Kunkel mutagenesis in order to generate libraries up to three orders of magnitude larger than the standard error-prone PCR and subcloning approach (Holland et al., 2013). Using our method, termed AXM mutagenesis, a large, mutated DNA fragment is produced using PCR conditions that promote nucleotide misincorporation into newly synthesized DNA. In the PCR reaction, one of the primers contains 3 or more phosphorothioate linkages at its 5’ end. Treatment of the error-prone generated PCR product with the 5’ to 3’ exonuclease of bacteriophage T7 removes the strand synthesized with the non-modified primer. The result of the exonuclease treatment is a single-stranded DNA segment, or ‘megaprimer’. This megaprimer is used in a Kunkel-like mutagenesis reaction that takes advantage of the E. coli DNA base excision repair pathway to bias nucleotide base-changes between the megaprimer and a complementary uracilated DNA sequence in favor of the in vitro synthesized megaprimer. This method facilitates the rapid generation of multiple directed evolution libraries in parallel.
Although site-directed mutagenesis methods can be very efficient, the efficiency is substantially reduced when multiple changes at distant sites are needed. The efficiency of incorporation of diversity ultimately impacts the overall library size, and the affinity of a selected clone from a phage display library is known to be strongly correlated with the size of the library (Ling, 2003). In addition to library size, the quality of the library, including the percentage of phage displaying recombinant proteins or peptides, significantly impacts the efficiency of the phage selections (Menendez and Scott, 2005). For a library containing several partially randomized positions, mutagenesis efficiency is best assessed by sequencing DNA from single clones. This library quality control method is time consuming and labor intensive, limiting the number of libraries that can be processed at one time.
The true library size could be directly estimated from the number of colonies obtained from an E. coli transformation (without sequencing analysis) if the parental (non-mutagenized) clones were eliminated. Restriction enzymes have been used in vitro as a means to eliminate parental clones (Huang et al., 2012). However, the in vitro method requires two rounds of E. coli transformations to obtain the final library and therefore does not save time or labor. Instead, in vivo expression of a restriction enzyme would enable elimination of parental clones without the requirement for additional in vitro processing steps.
It has now been established for over 50 years that some bacterial strains are able to resist the invasion of bacteriophage by evolving a mechanism to restrict the invading DNA (Dussoix and Arber, 1965; Meselson and Yuan, 1968; Arber and Linn, 1969). Since then, molecular biologists have identified hundreds of restriction endonucleases that are able to perform a similar protective function (Loenen et al., 2014). The Eco29k I restriction-modification system (RMS2) is derived from E. coli strain 29k and recognizes the sequence CCGCGG in DNA (Zakharova et al., 1998; Pertzev et al., 1992). The Eco29k I enzyme, which is an isoschizomer of Sac II, lacks thymine nucleotides in the recognition site. Since Kunkel mutagenesis involves the formation of a heteroduplex in which the template strand contains uracil instead of thymine, the presence of uracil in the parent template should not interfere with cleavage at this “GC”-rich site.
Here, we have engineered a vector to express the Eco29k I RMS2 in the E. coli strain TG1. Using this strain, we demonstrate that we are able to generate phage display libraries with high complexity and that incorporate diversity over the entire 800 bp fragment in >90% of library clones. Although we used Eco29k I for this study, the method can be extended to other restriction enzymes that can be expressed in E. coli.
2. Materials and methods
2.1. Bacterial strains and vectors
The E. coli strain CJ236 (Genotype: FΔ(HinDIII)::cat (Tra+, Pil+, CamR)/ ung-1, relA1, dut-1, thi-1, spoT1, mcrA) was purchased from New England BioLabs (NEB, Waverly, MA). Electrocompetent cells were made from this strain following NEB’s protocol. The E. coli strain TG1 [F' (traD36, proAB+ lacIq, lacZΔM15), supE, thi-1, Δ(lac-proAB), Δ(mcrB-hsdSM)5, (rK−mK−)] was purchased from Lucigen (Middleton, WI). AXE688 is E. coli strain TG1 transformed with pAX1492 containing the Eco29 I RM operon. Competent cells were made following the NEB online protocol [https://www.neb.com/protocols/2012/06/21/making-your-own-electrocompetent-cells].
The template plasmids for AXM mutagenesis (pAX1339 and pAX162) are derivatives of the phagemid, pAP-III6 (Haidaris et al., 2001) with the same single-chain variable fragment (scFv) antibody fused to the coat protein III of bacteriophage M13. Each of the 6 complementarity determining regions (CDRs) of the scFv in pAX1339 was modified to contain both opal (TGA) stop codons and Eco29k I restriction endonuclease recognition sites. In this template, non-recombinant clones are non-functional with respect to display of the scFv. The opal stop codons and Eco29k I restriction endonuclease sites are absent in pAX162. The template plasmid for standard Kunkel mutagenesis (pAX1392) is a derivative of pAP-III6 as above, but with a different scFv than either pAX1339 and pAX162. CDRs L2, L3, H2, and H3 contain both opal stop codons and Eco29k I endonuclease recognition sites.
The sequence for the Eco29k RM has been published (Zakharova et al., 1998). The gene sequence encoding the Eco29k I promoter, restriction endonuclease and DNA-methyltransferase was gene synthesized by Life Technologies. (Carlsbad, CA). The expression vector for Eco29k I (pAX1492) was made by cloning the synthesized Eco29k I operon between the EcoN I and Not I sites in pCDF-1b (Novagen, MA). By using these restriction sites, the T7 promoter was replaced by the natural promoter for Eco29k I. The pCDF-1b plasmid uses the CloDF13 origin of replication and is compatible with ColE1 plasmids.
2.2. Testing Eco29k I functionality
To test the functionality of the Eco29k I endonuclease, the expression vector (pAX1492) was transformed into TG1 cells to produce the strain designated as AXE688. Separately, the phagemids pAX1339 and pAX162 were packaged into phage particles from TG1(pAX1339) and TG1(pAX162) using the defective phage M13K07 and standard conditions (Soltes et al., 2007). The resulting phage supernatant containing the phagemids were stored at 4°C. 10 ml cultures of TG1 and AXE688 were grown at 37 °C with shaking at 225 rpm until OD600 of 0.4 was reached. To transduce the TG1 and AXE688 cells, 250 µl of each of the two strains at mid-log phase was added to 10 µl of phage for 30 min without shaking at 37°C. 200 µl of the phage and cell mixture dilutions were plated on LB plus ampicillin (100 µg ml−1) plates. Plates were incubated overnight at 37 °C and colonies were counted.
To test the functionality of the methylase gene, 3 ml of LB and 3 ml of LB plus spectinomycin (25 µg ml−1) were inoculated with 1 colony from the TG1 plate and 1 colony from AXE688 plate after the phage transduction, respectively. The cultures were grown with overnight shaking at 225 rpm at 37°C. The next day, the cultures were spun down and plasmid DNA was isolated using the Qiaprep Spin Mini prep kit (Qiagen, Valencia, CA) per manufacturer’s instructions. 10 µg of DNA was treated with 2 µl (20 U) of Sac II restriction endonuclease in 1× buffer 4 (NEB) for 3 hr at 37 °C followed by 65 °C for 10 min. 5 µl of the Sac II treated- and untreated- DNA were run on a 1% agarose gel at 140 volts for 40 min. The gel was stained with SYBR safe (Life Technologies) and imaged on a ChemiDoc MP system from Bio-rad (Hercules, CA).
2.3. Transformation of CJ236 strain
1 µl of the template plasmid (pAX1339 or pAX1392) DNA was added to 25 µl of electrocompetent CJ236 cells in a chilled 1 mm gap cuvette. The plasmid and cell mixture was electroporated in a Bio-Rad Gene Pulser electroporator at 1.6 kilovolts, 25 µF, and 200 Ω. 1ml of SOC media (2% tryptone, 0.5% yeast extract, 10 mM NaCl, 2.5 mM KCl, 10 mM MgCl2, 10 mM MgSO4, 20 mM glucose) was added to the cell mixture immediately after electroporation. The SOC plus cell mixture was then transferred to a 14 ml culture tube and incubated at 37 °C with shaking (~225 rpm). After 1 hr, 250 µl of the incubated mixture was spread onto a LB + amp (100 µg ml−1) + chloramphenicol (33 µg ml−1) petri plate. The plates were incubated overnight at 37 °C.
2.4. Production of uracilated, circular, single-stranded template DNA (dU-ssDNA)
This protocol for the production of uracilated template DNA was adapted from Fellouse and Sidhu (Fellouse et al., 2007). 1 ml of LB media plus ampicillin (60 µg ml−1), 100 µl of M13K07 helper phage (NEB, 1011 transducing forming units ml−1) and 5 colonies from an overnight transformation plate, picked with a single 10 µl tip were added to a 14 ml culture tube. The culture tube was then incubated at 37 °C at 225 rpm. After 2 hr, kanamycin was added (for a final concentration of 30 µg ml−1) to each culture tube and the tube was placed back at 37 °C shaking at 225 rpm for 6 hr (or until cloudy). The culture was then diluted into 30 ml of fresh LB medium containing ampicillin (60 µg ml−1), kanamycin (30 µg ml−1), and uridine (0.3 µg ml−1) in a 250 ml baffled flask. The flasks were then shaken at 225 rpm at 30 °C overnight (approximately 18 hr). The overnight culture was transferred to a round bottom centrifuge tube and spun at 15K rpm for 15 min in a chilled (4 °C) Sorvall RC-5B centrifuge using an SS34 rotor. The supernatant was passed through a 0.22 µm filter and transferred to a new tube containing 1/5 volume (6 ml for each 30 ml culture) of 20% PEG8000 + 2.5 M NaCl. After the 1hr incubation on ice, the tubes were spun for 20 min at 15K rpm in a 4 °C Sorvall RC-5B centrifuge using an SS34 rotor. The supernatant was decanted and the tubes were spun for an additional 5 min at 15K rpm in a 4°C Sorvall RC-5B centrifuge. The resulting phage particle pellet was re-suspended in 500 µl sterile 1× PBS (137 mM NaCl, 2.7 mM KCl, 4.3 mM Na2PO4, 1.47 mM KH2PO4, pH 7.4) and transferred to a 1.7 ml Eppendorf tube. The dU-ssDNA was purified using the M13 extraction kit from Qiagen. The resulting eluate contains uracilated-ssDNA. A 2 µl aliquot of the eluate was analyzed on a 1% agarose gel. dU-ssDNA appeared as a single predominant band.
2.5. Error Prone PCR for AXM mutagenesis
The Error-Prone PCR protocol of Cadwell and Joyce (Cadwell and Joyce, 1994) as modified by Holland et al. (Holland et al., 2013) was followed. In a 0.2 ml PCR tube, 10 µl of 10× mutagenic PCR buffer (70 mM MgCl2, 500 mM KCl, 100 mM Tris (pH 8.3), 0.1% (wt/vol) gelatin) was combined with 10 µL of 10× dNTP mix (2 mM dGTP, 2 mM dATP, 10 mM dCTP, 10 mM dTTP), 20 fmole of input DNA, 30 pmole each of forward and reverse primers, and H2O for a final volume of 88 µl. For amplifying the scFv genes the following two oligonucleotide primers were used: AXL6AS4-PCRR 5’-pGsTsCsGsACTGAGGAGACGGTGACC-3’ and AXL6KunkPCRF2, 5’-AAGCTTTCCTATGAGCTGACACAGCC-3’. where “s” corresponds to a phosphorothioate linkage between nucleotide bases and “p” indicates a 5’ phosphate. The primers were synthesized by Integrated DNA Technologies (Coralville, IA). 10 µl of 5 mM MnCl2 was then added, mixed well, and the absence of any precipitate visually verified. 2 µl of Taq DNA polymerase (NEB) was added to bring the final volume to 100 µL. Cycling conditions consisted of 30 cycles of: 94 °C 1 min, 60 °C 1 min, and 72 °C 1 min. After PCR was complete, cleanup was accomplished using the QIAquick PCR Purification Kit (Qiagen) according to the manufacturer’s protocol. The purified DNA was eluted in 40 µl of buffer EB. For scFvs, we expected a PCR amplification product of approximately 800 base pairs, which was verified by agarose gel electrophoresis.
2.6. T7 exonuclease treatment for AXM Mutagenesis
Forty microliters of the cleaned-up PCR product (20 pmole) was incubated with 10 µl T7 exonuclease (10,000 units per ml, NEB) in 1× buffer 4. The reaction was incubated at 25 °C for 1 hr, and the T7 exonuclease-treated sample was purified using the QIAquick PCR Purification Kit and eluted in 40 µl EB buffer.
2.7. DNA heteroduplex formation
For the AXM Mutagenesis libraries, 40 µl of T7 exonuclease treated- and cleaned-up reaction (~20 pmole) was mixed separately with 4 µl of pAX1339 ssDNA (4 pmole for a 5:1 ratio of megaprimer to dU-ssDNA, respectively), 25 µl of 10× TM buffer (0.1 M MgCl2, 0.5 M Tris, pH 7.5) buffer, and dH20 for a final reaction volume of 250 µl. For the naïve Kunkel mutagenic libraries, 600 ng of phosphorylated oligonucleotide was added to 20 µg of dU-containing pAX1392 ssDNA template in 1× TM buffer. The annealing reaction for both types of libraries was carried out by distributing the mixtures equally into five PCR tubes, then heating the mixture at 90 °C for 2 min, followed by a temperature decrease of 1 °C per minute to 25 °C in a thermal cycler. To the annealed product, 10 µl of 10 mM ATP, 10 µl of 100 mM dNTP mix, 15 µl of 100 mM DTT, 0.5 µl (30 U) T4 DNA ligase and 3 µl (30 U) T7 DNA polymerase was added. The mixture was distributed equally into five PCR tubes and incubated overnight (16 hr) at 20 °C.
The DNA was desalted and purified using a QIAquick DNA purification kit. 1 ml of buffer QG (Qiagen) was added. The sample was applied to two QIAquick spin columns placed in 1.7 ml microcentrifuge tubes. The columns were spun at 13K rpm for 1 min in an Eppendorf microcentrifuge, and the flow-through was discarded. To each column, 750 µl of buffer PE was added and the sample was spun again at 13K rpm in the microcentrifuge. The column was transferred to a clean 1.7 ml microcentrifuge tube, and DNA was eluted from each column in 35 µl of buffer EB (QIAquick DNA purification kit). The eluates from the two columns (70 µl final) were combined. 2 µl of the product was resolved by agarose gel electrophoresis, alongside 2 µl of dU-ssDNA. For the heteroduplex product, up to 3 bands are generally observed: the upper (and often most prevalent) band corresponds to strand-displaced DNA, the middle (usually faint) band corresponds to nicked DNA (i.e., not properly extended and ligated), and the lower band represents the extended and ligated product. The dU-ssDNA has a faster mobility than the correctly extended and ligated band.
2.8. Library transformation
Transformation reactions were set up as follows: 1 µl heteroduplex DNA was mixed with 25 µl of either TG1 electrocompetent cells (Lucigen, Middleton, WI) or AXE688 electrocompetent cells and added to a 0.1 mm gap cuvette. DNA was electroporated into bacterial cells using a Bio-Rad Gene Pulser electroporator at 1.6 kilovolts, 25 µF, and 200 Ω. 1 ml of Lucigen recovery media (Lucigen) was added and the electroporated cells were transferred to 14 ml culture tubes and shaken at 225 rpm at 37 °C. After 1 hr, serial dilutions up to 10−8 were plated onto LB plates containing ampicillin (100 µg ml−1), and the plates are incubated overnight at 37 °C. PCR products were sequenced (Quintara, CA) to determine the percentage of recombinants.
3. Results
3.1. Monitoring library diversity
To determine whether an in vivo restriction system would be beneficial for recombinant antibody library construction, we initially evaluated the efficiency of primer incorporation using both a standard Kunkel mutagenesis approach and our AXM mutagenesis approach.
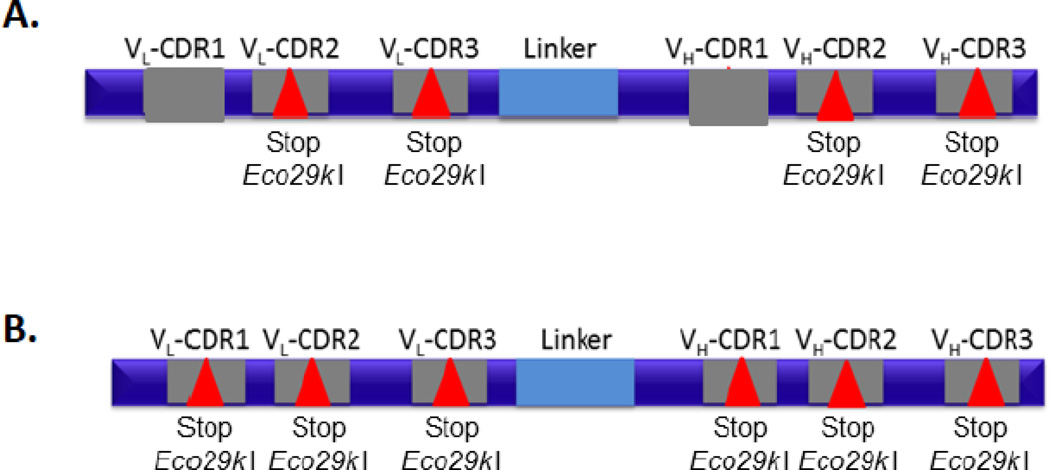
For the Kunkel mutagenesis library, a single framework scFv based on the human germline sequences VH5-3 and VL3–10 was used as a template based on its favorable expression in E. coli, yeast, and mammalian cells (data not shown). In order to monitor the efficiency of incorporation of randomized primers at each of 4 complementarity determining regions (CDRs), we introduced Eco29k I (CCGCGG) sites in CDRs L2, L3, H2, and H3 (Fig. 1A). A total of eighteen positions were randomized in these CDRs using an “NNK” strategy that introduces all 20 amino acids as well as one stop codon (Table 1). In addition to the sequence diversity, three different CDR H3 lengths (7, 12, or 16 amino acids) were generated (Table 1). Oligonucleotides incorporating the diversity at each of the 4 CDRs were annealed to a uracilated, circular, single-stranded DNA template containing Eco29k I sites in the corresponding CDRs. After extension by T7 DNA polymerase and ligation by T4 DNA ligase, the resulting heteroduplex was transformed into strain TG1. The efficiency of incorporating all 4 oligonucleotides was evaluated by DNA sequence analysis. Of 77 clones evaluated, 33 were parental (retaining all 4 Eco29k I sites), 25 contained 3 Eco29k I sites, 8 contained 2 Eco29k I sites, 5 contained 1 Eco29k I site, and 6 were fully recombinant (lacking Eco29k I sites). Based on this analysis, only 8 % of the library represented fully recombinant (and functional) clones (Table 2).
Fig. 1.
Parent template for Kunkel mutagenesis. A constant single chain antibody fragment (scFv) framework was used for library construction. Diagrams of the circular, single-stranded template for standard Kunkel mutagenesis and AXM mutagenesis are indicated in (A) and (B) respectively. To eliminate parental clones from the library, Eco29k I endonuclease recognition sites as well as opal stop codons were introduced at all 6 of the complementarity determining regions (CDRs) in the single-stranded template for AXM mutagenesis (B) and in the four CDRs targeted for randomization in the naïve libraries (A). Grey boxes represent the CDR regions in the heavy and light chain variable genes. Red triangles represent stop codons and Eco29k I sites in the CDRs.
Table 1.
Sequence of oligonucleotides used for Kunkel mutagenesis
| Oligo Name | Oligo Sequence (Note: Oligos are reverse orientation to anneal to the phagemid single strand DNA) |
|---|---|
| L2 mutNNK HM | GAATCTCTCAGGGATCCCGGAMNNTCGTTTMNNGTCCTCATAGATGACCAGCAC |
| L3 mutNNK HM | CTCACCTAGGACGGTGAGCTTMNNCCCTCCGCCMNNMNNMNNMNNATTACCACTGCTGTCTGTTGA |
| H2 mutNNK HM | GTCAGCTGAGATGGTGACGTGGCCTTGGAAGGACGGGCCGTAMNNGGTMNNGGAMNNMNNMNNMNNAATMNNCCCCATCCACTCCAGGCCTTTCCC |
| H3 mut_4NNK HM | AGGAATAGTACCCCCAGATGTMNNMNNMNNMNNCCTCACACAGTAATACATGGCGGT |
| H3 mutR_7NNK HM | TTCAGGAATGGTGCCCGGGCCCCAMNNMNNMNNMNNMNNMNNMNNCCTCACACAGTAATACATGGCGGT |
| H3 mutRR_12 | CCAGGGTGCCCGGGCCCCARKNRKNRKNRKNRKNRKNRKNRKNRKNRKNRKNRKNCCTCACACAGTAATACATGG |
Table 2.
Efficiency of Kunkel Mutagenesis and AXM Mutagenesis
| Library Type | % Full Recombinants |
% Partial Recombinants | % Parental |
Transformation Efficiency (cfu/µg) |
||||
|---|---|---|---|---|---|---|---|---|
| (No Eco29k I sites) |
1 Eco29k I site |
2 Eco29k I sites |
3 Eco29k I sites |
4 Eco29k I sites |
5 Eco29k I sites |
6 Eco29k I sites |
||
| Standard Kunkel | 8 | 7 | 10 | 32 | 43 | NA | NA | 1.1 × 109 |
| AXM Mutagenesis | 17 | 8 | 12 | 21 | 17 | 8 | 17 | 1.5 × 109 |
To compare the efficiency of incorporation of a single megaprimer spanning the entire scFv gene, we have generated a library of variants by our previously described AXM mutagenesis method (Holland et al., 2013 and Fig. 2). In this method, an 800 bp PCR product was generated by error-prone PCR of an anti-EZH2 scFv using a reverse primer containing phosphorothioate linkages at its 5’ end. The product was treated with T7 exonuclease to selectively degrade the non-modified (unprotected) strand. The resultant megaprimer was annealed to a uracilated, circular, single-stranded template containing the same scFv framework sequence but with Eco29k I sites incorporated at each of the 6 CDRs (Fig. 1B). After extension and ligation, the heteroduplex product was transformed into strain TG1 as above. The transformation efficiency was determined by counting the number of colonies obtained from serial dilutions. Twenty-four individual clones were PCR-amplified and sequenced to evaluate the efficiency of incorporation of the mutated megaprimer. Of the twenty-four analyzed clones, only 4 (17%) were fully recombinant (containing no Eco29k I sites) (Table 2).
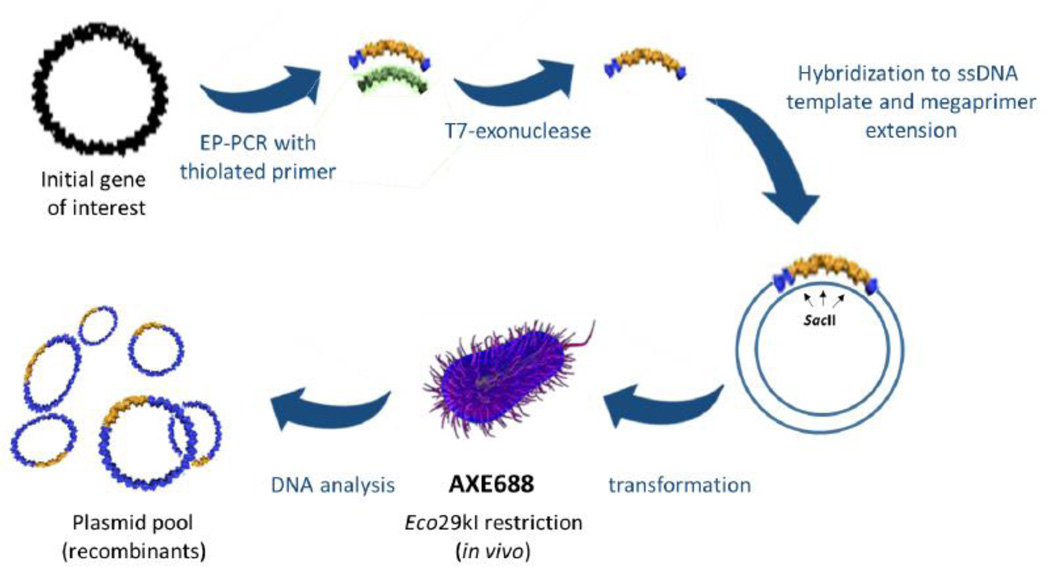
Fig. 2.
AXM mutagenesis procedure. The coding region for the recombinant antibody (rAb) is amplified under error-prone PCR (EP-PCR) conditions, using a reverse primer containing phosphorothioate linkages on its 5’ end. The resulting double-stranded DNA is treated with T7 exonuclease to selectively degrade the unmodified strand of the dsDNA molecule. The resulting single-stranded DNA, or ‘megaprimer’, is then annealed to the uracilated, circular, single-stranded phagemid DNA and used to prime in vitro synthesis by DNA polymerase. The ligated, heteroduplex product is then transformed into E. coli AXE688 cells, where the uracilated strand is cleaved in vivo by uracil N-glycosylase, favoring survival of the newly synthesized, recombinant strand containing the megaprimer. The Eco29k I enzyme selectively degrades parental (template) DNA retaining Eco29k I recognition sites.
3.2. Construction of AXE688 strain
Due to the poor efficiency of generating clones incorporating the intended diversity in all CDRs, we sought an in vivo approach to eliminate template clones and simplify the library construction and analysis process. Eco29k I is an isoschizomer of Sac II that can be expressed well in E. coli (Zakharova et al., 1998; Pertzev et al., 1992). This enzyme was selected because the recognition site contains only guanine and cytosine base pairs. Since Kunkel mutagenesis uses a template containing uracil instead of thymine, restriction sites containing adenine and thymine base pairs might not be cleaved efficiently.
To generate an expression construct for Eco29k I, the Eco29k I RM operon encoding both the restriction endonuclease and methyltransferase genes was synthesized with the natural promoter and terminator sequences (Zakharova et al., 1998; Pertzev et al., 1992) and cloned into the pCDF-1b vector (Novagen) between the EcoN I and Not I restriction sites. This cloning strategy removed the T7 promoter and allowed the Eco29 RM to be expressed from its natural promoter. This plasmid encodes the spectinomycin resistance gene and contains the CloDF13 origin, making it compatible with both the ampicillin-resistant ColE1 phagemid and kanamycin-resistant helper phage required for phage display. E. coli TG1 were transformed with the Eco29k I expression vector to develop the strain, AXE688.
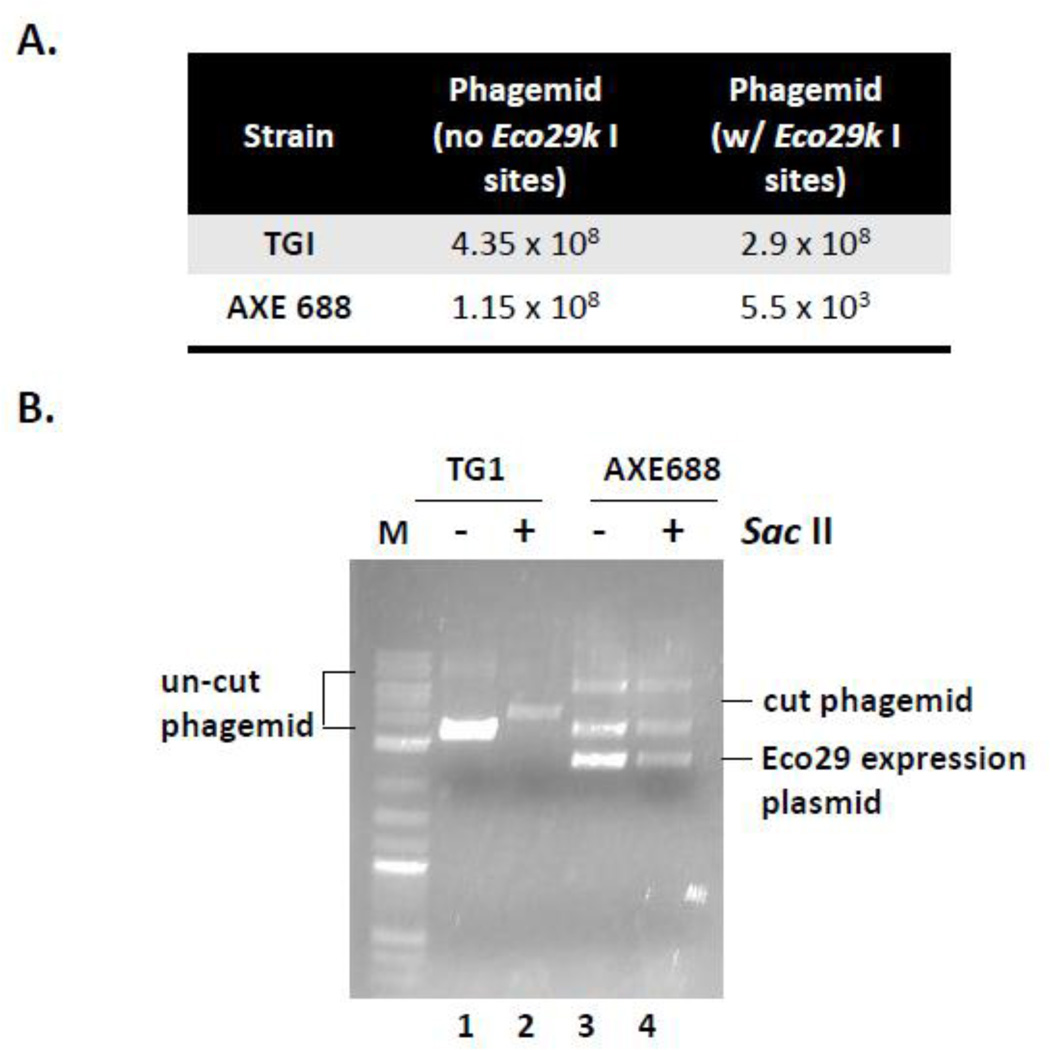
To test the function of the Eco29k I system, the AXE688 strain containing the Eco29k I expression construct, was transduced with the Eco29k I site-containing phagemid or a control phagemid lacking Eco29k I sites within the scFv gene. The standard TG1 strain was also transduced with these phagemids in parallel. The transductants were selected on LB plates containing ampicillin. An at least 10,000-fold reduction in transduction efficiency was observed for the Eco29k I-containing phagemid in AXE688 cells compared with the standard TG1 strain (Fig. 3A). No difference in transduction efficiency between the two strains was observed for the phagemid lacking Eco29k I sites, and both phagemids yielded similar numbers of transductants in TG1 cells that do not express Eco29k I (Fig. 3A). The lower transduction efficiency for the Eco29k I site-containing phagemid in the AXE688 strain is consistent with restriction of the incoming DNA by the Eco29k I endonuclease.
Fig. 3.
Eco29k I restriction endonuclease and methyltransferase are functionally expressed in AXE688. (A) AXE688 or the strain TG1 strain was transduced with a Sac II-containing phagemid or a control phagemid lacking Eco29k I sites within the scFv gene. The number of ampicillin-resistant transductants was determined. (B) Plasmid DNA was recovered from the AXE688 and TG1 strains transduced with the Eco29k I-containing phagemid. The DNA was either untreated (lanes 1 and 3) or digested with the Sac II endonuclease in vitro (lanes 2 and 4). DNA from the AXE688 strain is not cut by Sac II due to methylation of the Sac II site by the Eco29k I methylase.
The Eco29k I restriction methylation system is comprised of both a restriction endonuclease and a DNA methyltransferase. Eco29k I methyltransferase activity is required to protect host cell DNA from restriction by the Eco29k I endonuclease. Transformed DNA that is not cleaved by the endonuclease will become methylated. To confirm that the methyltransferase was functional in AXE688, plasmid DNA was isolated from ampicillin-resistant AXE688 cells from the transduction experiment above. The recovered DNA was not cleaved by the Sac II enzyme in vitro, suggesting that Eco29k I methylates the restriction sites in vivo, protecting the DNA from cleavage (Fig. 3B). The same plasmid isolated from the TG1 strain lacking Eco29k I was efficiently cleaved by Sac II in vitro (Figure 3B). Together, this data suggests that both the Eco29k I endonuclease and methyltransferase are expressed and are functional in AXE688 cells.
3.3. Recombinant rate increases with the use of AXE688 cells
To determine whether Eco29k I could enhance the efficiency of the AXM mutagenesis method, electrocompetent cells were generated from the AXE688 strain. The electroporation efficiencies of the AXE688 cells and standard TG1 cells using pUC19 DNA (which does not contain an Eco29k I site) are equivalent (Table 3).
Table 3.
Electroporation efficiency of AXE688
| Strain | Efficiency (cfu/µg)1 |
|---|---|
| TG1 | 2.5 × 1010 |
| AXE688 | 1.5 × 1010 |
Efficiencies were based on the number of ampicillin-resistant colonies after transformation with 0.01 ng of pUC19 DNA.
To evaluate the transformation efficiency of the AXE688 cells when one plasmid contains Eco29k I sites, we mixed the two plasmids described above, one lacking Eco29k I sites and one containing six Eco29k I sites, at various ratios. Equal amounts (100 ng) of the plasmid mixtures were transformed into either TG1 or AXE688 cells. The transformation efficiencies of TG1 and AXE688 cells were nearly equivalent (5.8 × 109 and 3.6 × 109 cfu/µg, respectively) when 100% of the input plasmid lacked Eco29k I sites (Table 4). As expected, the efficiency went down >10,000-fold in AXE688 when 100% of the input plasmid contained Eco29k I sites (Table 4). Although the transformation efficiency in TG1 remained constant with the different input plasmid mixtures, the frequency of clones lacking Eco29k I sites closely matched the frequency of the vector lacking Eco29k I sites in the input mixture (Table 4; 29% for the 50:50 mixture and 8% for the 90:10 mixture of Eco29k I site-containing vector to Eco29k I site-lacking vector). In contrast, although the transformation efficiency was slightly reduced in AXE688 when 90% of the input mixture contained Eco29k I sites, greater than 96% of the recovered clones lacked the Eco29k I site (Table 4).
Table 4.
Recovery of mixtures of plasmids with and without Eco29k I sites in TG1 and AXE688 cells
| Strain | Input Ratio Eco29k I +/ Eco29k I − |
Transformation efficiency (cfu/µg) |
Recovered clones (% Eco29k I −) |
|---|---|---|---|
| TG1 | 0/100 | 5.8 × 109 | 100 |
| TG1 | 50/50 | 1.1 × 1010 | 29 |
| TG1 | 90/10 | 1.1 × 1010 | 8 |
| TG1 | 100/0 | 1.25 × 1010 | 0 |
| AXE688 | 0/100 | 3.6 × 109 | 100 |
| AXE688 | 50/50 | 2.4 × 109 | >96 |
| AXE688 | 90/10 | 7 × 108 | >96 |
| AXE688 | 100/0 | 1.2 × 105 | NA |
These results suggested that the AXE688 cells could be used to create libraries of nearly 100% recombinant clones. A dsDNA heteroduplex library product generated using the AXM mutagenesis method on several different starting scFvs was transformed either into AXE688 or TG1 competent cells, and the number of recombinant clones was determined by DNA sequencing (Table 5). The number of recombinants, i.e., clones with fully incorporated error-prone PCR product varied from 6 to 40% when the heteroduplex product was transformed into standard TG1 cells. However, when the scFv libraries were transformed into the AXE688 strain expressing Eco29k I, 90% or more of recovered clones were non-parental (ie. containing no Eco29k I sites) (Table 5). Since the recombinant clones do not contain Eco29k I sites in the CDRs, they are not affected by the Eco29k I endonuclease activity. The increased frequency of recombinants in AXE688 is dependent on restriction of DNA containing Eco29k I sites because when the vector template used for Kunkel mutagenesis lacks Eco29k I sites, little difference in the number of recombinants is observed between the standard TG1 and AXE688 cells (Table 5). Sequence analysis confirms that the mutation rate in recombinant clones from the AXE688 library is comparable to the mutation rate observed in clones from standard TG1 cells (data not shown). The actual number of transformants more closely represents the true library size in the AXE688 strain (Table 5).
Table 5.
Use of restriction endonuclease in vivo reduces parental clones in affinity maturation libraries
| TG1 | AXE688 | ||||
|---|---|---|---|---|---|
| EP PCR Template |
Phagemid Template1 |
% Recombinants |
Transformation Efficiency (cfu/µg) |
% Recombinants |
Transformation Efficiency (cfu/µg) |
| scFv 1 | Eco29k I | 40 | 3.7 × 108 | 89 | 6.6 × 108 |
| scFv 2 | Eco29k I | 6 | 3.2 × 108 | 94 | 8.7 × 107 |
| scFv 3 | Eco29k I | 31 | 8 × 108 | 100 | 4.3 × 108 |
| scFv 3 | no RE sites | 30 | 5.7 × 108 | 29 | 8.7 × 108 |
The circular, single-stranded phagemid template either contains or lacks Eco29k I sites in the CDRs of the scFv.
Similar to the AXM mutagenesis libraries, when the standard Kunkel library (incorporating randomized oligonucleotides at 4 CDRs) was transformed into AXE688 cells, greater than 95% of the resultant clones were recombinant versus 8% in TG1 (Table 6). These results demonstrate that the AXE688 strain enables the construction of phage display libraries that have a higher (>90%) frequency of recombinant clones than typically achieved by Kunkel mutagenesis alone. These libraries with high diversity and functionality can be generated without the requirement for time consuming and labor intensive in vitro processing steps.
Table 6.
Use of restriction endonuclease in vivo reduces parental clones in naïve library
| TG1 | AXE688 | ||||
|---|---|---|---|---|---|
| # mutagenic oligos1 |
ssDNA template2 |
% Non- parentals |
Transformation Efficiency (cfu/µg) |
% Non- parentals |
Transformation Efficiency (cfu/µg) |
| 4 | Eco29k I | 8 | 1.1 × 109 | 96 | 2.3 × 108 |
Oligonucleotides containing “NNK” nucleotides at variable positions in four of the CDRs of the scFv were generated.
The circular, single-stranded phagemid template contains Eco29k I sites in the 4 CDRs targeted for mutagenesis.
4. Discussion
We have expressed the Eco29k I restriction-methylation system (RM2) in E. coli strain TG1 to produce the strain AXE688. We have used AXE688 cells in a novel directed molecular evolution (DME) mutagenesis method to restrict incoming parental DNA in transformed cells. Using our method, we have demonstrated that the AXE688 strain enables the construction of phage display libraries that have a higher frequency of recombinant clones (>90%) than typically achieved by Kunkel mutagenesis alone.
We and others have previously shown that by incorporating restriction sites in the template plasmid used for Kunkel mutagenesis, we can eliminate parental (non-recombinant) clones from the library (Holland et al., 2013; Huang et al., 2012). However, the in vitro method requires two rounds of E. coli transformations to obtain the final library and adds labor and cost to the library construction process. In addition, it is difficult to measure the loss of diversity resulting from isolation of DNA from the initially transformed cells, in vitro digestion step, and second round of E. coli transformations. Sequence analysis is still required prior to the second round of transformations to determine the actual potential diversity of the final library. Hence, this approach does not improve the throughput of library construction and analysis.
We have shown here that competent cells engineered to express the restriction endonuclease in vivo are just as efficient in eliminating partial and parental clones as in vitro digestion of library DNA and subsequent re-transformation. This new method enables the generation of libraries with high diversity and functionality in a single round of transformations. In our system, the restriction enzyme is expressed from a separate plasmid maintained in the E. coli strain, but the expression cassette can also be incorporated in the E. coli genome (eg. GeneBridge).
Although we used Eco29k I for this study, the method can be extended to other restriction enzymes that can be expressed in E. coli. The Eco29k I enzyme, which is an isoschizomer of Sac II, was chosen here because it lacks thymine nucleotides in the recognition site. Since Kunkel mutagenesis involves the formation of a heteroduplex in which the template strand contains uracil instead of thymine, restriction enzymes recognizing thymine-containing sites might not be able to cleave the transforming DNA as efficiently. Since the M13 helper phage contains a Eco29k I recognition site in its genome, we produced the helper phage in the AXE688 strain so that its DNA is methyl-protected and cannot be cleaved by the Eco29k I endonuclease (Table 7).
Table 7.
Helper phage titer comparison in TG1 and AXE688 cells
| Cell Strain | Helper phage1 | pfu/ml2 |
|---|---|---|
| AXE688 | AXKM13 | 4.3 × 1014 |
| AXE688 | KM13 | 5.7 × 1013 |
| TG1 | AXKM13 | 2.2 × 1014 |
| TG1 | KM13 | 1.5 × 1014 |
KM13 helper phage was produced in TG1. AXKM13 helper phage was produced in AXE688.
Average phage titers from two replicates are represented.
Recombination-based cloning methods such as Gateway (Life Technologies) take advantage of the high efficiency and accuracy of natural recombinases to generate clones with the desired changes (Bubeck et al., 1993; Degryse, 1996; Oliner et al., 1993; Luckow et al., 1993; Mejia and Larin, 2000; Gruet et al., 2012; Park and Labaer, 2006). In many cases, a toxic gene such as ccdB or sacB (Lund et al., 2014; Muhl and Filloux, 2014) in the parent construct is used to increase the likelihood of recovering recombinant clones. These systems generally require specific hosts or selection schemes and can be limited in scope. Here, we show that simply introducing restriction enzyme cleavage sites within the parent construct and using an E. coli strain that is engineered to express the restriction enzyme in vivo can be as effective as these other methods for deriving a high frequency of recombinant clones.
Acknowledgements
This work was supported by the National Institutes of Health [1 U54 DK093444-01].
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Arber W, Linn S. DNA modification and restriction. Annu. Rev. Biochem. 1969;38:467–500. doi: 10.1146/annurev.bi.38.070169.002343. [DOI] [PubMed] [Google Scholar]
- Bubeck P, Winkler M, Bautsch W. Rapid cloning by homologous recombination in vivo. Nucleic Acids Res. 1993;21:3601–3602. doi: 10.1093/nar/21.15.3601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cadwell RC, Joyce GF. Mutagenic PCR. PCR Methods Appl. 1994;3:S136–S140. doi: 10.1101/gr.3.6.s136. [DOI] [PubMed] [Google Scholar]
- Coia G, Ayres A, Lilley GG, Hudson PJ, Irving RA. Use of mutator cells as a means for increasing production levels of a recombinant antibody directed against Hepatitis B. Gene. 1997;201:203–209. doi: 10.1016/s0378-1119(97)00452-6. [DOI] [PubMed] [Google Scholar]
- Degryse E. In vivo intermolecular recombination in Escherichia coli: application to plasmid constructions. Gene. 1996;170:45–50. doi: 10.1016/0378-1119(95)00858-6. [DOI] [PubMed] [Google Scholar]
- Dussoix D, Arber W. Host specificity of DNA produced by Escherichia Coli. IV. Host specificity of infectious DNA from bacteriophage lambda. J. Mol. Biol. 1965;11:238–246. doi: 10.1016/s0022-2836(65)80054-7. [DOI] [PubMed] [Google Scholar]
- Emond S, Mondon P, Pizzut-Serin S, Douchy L, Crozet F, Bouayadi K, Kharrat H, Potocki-Véronèse G, Monsan P, Remaud-Simeon M. A novel random mutagenesis approach using human mutagenic DNA polymerases to generate enzyme variant libraries. Protein Eng. Des. Sel. 2008;21:267–274. doi: 10.1093/protein/gzn004. [DOI] [PubMed] [Google Scholar]
- Fellouse FA, Esaki K, Birtalan S, Raptis D, Cancasci VJ, Koide A, Jhurani P, Vasser M, Wiesmann C, Kossiakoff AA, Koide S, Sidhu SS. High-throughput generation of synthetic antibodies from highly functional minimalist phage-displayed libraries. J. Mol Biol. 2007;373:924–940. doi: 10.1016/j.jmb.2007.08.005. [DOI] [PubMed] [Google Scholar]
- Foote J, Winter G. Antibody framework residues affecting the conformation of the hypervariable loops. J. Mol. Biol. 1992;224:487–499. doi: 10.1016/0022-2836(92)91010-m. [DOI] [PubMed] [Google Scholar]
- Fromant M, Blanquet S, Plateau P. Direct random mutagenesis of gene-sized DNA fragments using polymerase chain reaction. Anal. Biochem. 1995;224:347–353. doi: 10.1006/abio.1995.1050. [DOI] [PubMed] [Google Scholar]
- Gruet A, Longhi S, Bignon C. One-step generation of error-prone PCR libraries using Gateway® technology. Microb. Cell Fact. 2012;11:14. doi: 10.1186/1475-2859-11-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haidaris CG, Malone J, Sherrill LA, Bliss JM, Gaspari AA, Insel RA, Sullivan MA. Recombinant human antibody single chain variable fragments reactive with Candida albicans surface antigens. J. Immunol. Methods. 2001;257:185–202. doi: 10.1016/s0022-1759(01)00463-x. [DOI] [PubMed] [Google Scholar]
- Holland EG, Buhr DL, Acca FE, Alderman D, Bovat K, Busygina V, Kay BK, Weiner MP, Kiss MM. AXM mutagenesis: an efficient means for the production of libraries for directed evolution of proteins. J Immunol Methods. 2013;394:55–61. doi: 10.1016/j.jim.2013.05.003. [DOI] [PubMed] [Google Scholar]
- Huang R, Fang P, Kay BK. Improvements to the Kunkel mutagenesis protocol for constructing primary and secondary phage-display libraries. Methods. 2012;58:10–17. doi: 10.1016/j.ymeth.2012.08.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jäckel C, Kast P, Hilvert D. Protein design by directed evolution. Annu. Rev. Biophys. 2008;37:153–173. doi: 10.1146/annurev.biophys.37.032807.125832. [DOI] [PubMed] [Google Scholar]
- Kunkel TA. Rapid and efficient site-specific mutagenesis without phenotypic selection. Proc. Natl. Acad. Sci. USA. 1985;82:488–492. doi: 10.1073/pnas.82.2.488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ling MM. Large antibody display libraries for isolation of high-affinity antibodies. Comb. Chem. High Throughput Screen. 2003;6:421–432. doi: 10.2174/138620703106298608. [DOI] [PubMed] [Google Scholar]
- Loenen WA, Dryden DT, Raleigh EA, Wilson GG, Murray NE. Highlights of the DNA cutters: a short history of the restriction enzymes. Nucleic Acids Res. 2014;42:3–19. doi: 10.1093/nar/gkt990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luckow VA, Lee SC, Barry GF, Olins PO. Efficient generation of infectious recombinant baculoviruses by site-specific transposon-mediated insertion of foreign genes into a baculovirus genome propagated in Escherichia coli. J Virol. 1993;67:4566–4579. doi: 10.1128/jvi.67.8.4566-4579.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lund BA, Leiros HK, Bjerga GE. A high-throughput, restriction-free cloning and screening strategy based on ccdB-gene replacement. Microb. Cell Fact. 2014;13:38. doi: 10.1186/1475-2859-13-38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martineau P. Error-prone polymerase chain reaction for modification of scFvs. Methods Mol. Biol. 2002;178:287–294. doi: 10.1385/1-59259-240-6:287. [DOI] [PubMed] [Google Scholar]
- Mejía JE, Larin Z. The assembly of large BACs by in vivo recombination. Genomics. 2000;70:165–170. doi: 10.1006/geno.2000.6372. [DOI] [PubMed] [Google Scholar]
- Menendez A, Scott JK. The nature of target-unrelated peptides recovered in the screening of phage-displayed random peptide libraries with antibodies. Anal. Biochem. 2005;336:145–157. doi: 10.1016/j.ab.2004.09.048. [DOI] [PubMed] [Google Scholar]
- Meselson M, Yuan R. DNA restriction enzyme from E. coli. Nature. 1968;217:1110–1114. doi: 10.1038/2171110a0. [DOI] [PubMed] [Google Scholar]
- Muhl D, Filloux A. Site-directed mutagenesis and gene deletion using reverse genetics. Methods Mol Biol. 2014;1149:521–539. doi: 10.1007/978-1-4939-0473-0_40. [DOI] [PubMed] [Google Scholar]
- Oliner JD, Kinzler KW, Vogelstein B. In vivo cloning of PCR products in E. coli. Nucleic Acids Res. 1993;21:5192–5197. doi: 10.1093/nar/21.22.5192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park J, Labaer J. Recombinational cloning. Curr. Protoc. Mol. Biol. 2006;Chapter 3(Unit 3.20) doi: 10.1002/0471142727.mb0320s74. [DOI] [PubMed] [Google Scholar]
- Pertzev AV, Ruban NM, Zakharova MV, Beletzkaja IV, Petrov SI, Kravetz AN, Solonin AS. Eco29kI, a novel plasmid encoded restriction endonuclease from Escherichia coli. Nucleic Acids Res. 1992;20:1991. doi: 10.1093/nar/20.8.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scholle MD, Kehoe JW, Kay BK. Efficient construction of a large collection of phage-displayed combinatorial peptide libraries. Comb. Chem. High Throughput Screen. 2005;8:545–551. doi: 10.2174/1386207054867337. [DOI] [PubMed] [Google Scholar]
- Smith GP, Scott JK. Libraries of peptides and proteins displayed on filamentous phage. Methods Enzymol. 1993;217:228–257. doi: 10.1016/0076-6879(93)17065-d. [DOI] [PubMed] [Google Scholar]
- Soltes G, Hust M, Ng KK, Bansal A, Field J, Stewart DI, Dübel S, Cha S, Wiersma EJ. On the influence of vector design on antibody phage display. J. Biotechnol. 2007;127:626–637. doi: 10.1016/j.jbiotec.2006.08.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stemmer WP. Rapid evolution of a protein in vitro by DNA shuffling. Nature. 1994;370:389–391. doi: 10.1038/370389a0. [DOI] [PubMed] [Google Scholar]
- Waterhouse P, Griffiths AD, Johnson KS, Winter G. Combinatorial infection and in vivo recombination: a strategy for making large phage antibody repertoires. Nucleic Acids Res. 1993;21:2265–2266. doi: 10.1093/nar/21.9.2265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiner MP, Costa GL, Schoettlin W, Cline J, Mathur E, Bauer JC. Site-directed mutagenesis of double-stranded DNA by the polymerase chain reaction. Gene. 1994;151:119–123. doi: 10.1016/0378-1119(94)90641-6. [DOI] [PubMed] [Google Scholar]
- Zaccolo M, Williams DM, Brown DM, Gherardi E. An approach to random mutagenesis of DNA using mixtures of triphosphate derivatives of nucleoside analogues. J. Mol. Biol. 1996;255:589–603. doi: 10.1006/jmbi.1996.0049. [DOI] [PubMed] [Google Scholar]
- Zakharova MV, Beletskaya IV, Kravetz AN, Pertzev AV, Mayorov SG, Shlyapnikov MG, Solonin AS. Cloning and sequence analysis of the plasmid-borne genes encoding the Eco29kI restriction and modification enzymes. Gene. 1998;208:177–182. doi: 10.1016/s0378-1119(97)00637-9. [DOI] [PubMed] [Google Scholar]