Abstract
miR-205 is an epithelial-specific miRNA and has been shown to orchestrate some cellular processes such as epithelial mesenchymal transition (EMT) and differentiation fate of stem cells in mammary gland. miR-205 play a part of a tumor suppressor in human cancers. However, the role of miR-205 in lung cancer is unclear. In this study, we detected the expression level of miR-205 in 46 cases clinical lung cancer specimens and adjacent normal tissues by stem-loop RT-PCR. We found that the expression of miR-205 was significantly increased in lung cancer specimens compared to adjacent normal tissues (P < 0.01). Furthermore, we observed the expressions of PTEN protein and mRNA in lung cancer tissues and adjacent normal tissues by methods of western blot and Real time PCR respectively. We found that the expressions of PTEN protein and mRNA was significantly decreased in lung cancer specimens compared to adjacent normal tissues. And then, we found there is a negative relationship between the expression of miR-205 and PTEN mRNA in lung cancer by analyzed. To validate whether PTEN was direct targets of miR-205, a dual-luciferase reporter assay was employed, the result showed that PTEN is a target gene of MiR-205. In subsequent experiments, we examined the expressions of PTEN protein and mRNA after transfection of miR-205 mimics or inhibitor into A549 cells, and A549 cell proliferation was measured by CCK-8 tests. We found that the expression of PTEN protein and mRNA in A549 cells were significantly down-regulated or up-regulated after miR-205 mimics and miR-205 inhibitors transfected into, and miR-205 could inhibits A549 cells proliferation. These results indicate that miR-205 might inhibitor the proliferation of A549 cells by regulating the expression of PTEN.
Keywords: Lung cancer, miR-205, PTEN, proliferation
Introduction
Lung cancer is a high incidence of cancer increasing year by year. With the development of science and technology, the treatment of lung cancer had penetrated the gene level. In recent years, miRNA has been paid more and more attention. MicroRNAs (miRNAs) are small noncoding RNAs that regulate gene expression through base-pairing with their target messenger RNAs. miR-205 is a highly conserved microRNA and it has been proved that miR-205 shares a similar expression pattern with miR-200 family [1]. It has been proved that miR-205 might associated with the occurrence, development and prognosis of kinds tumors, and many studies suggest that expression levels of miR-205 were evidently different in different tumors. For example, the expressions of miR205 was down-regulated in Breast cancer [2], the former Prostate cancer [3], however, up-regulate in Endometrial carcinoma [4], non small cell lung cancer [5], head and neck squamous cell carcinoma [6] and melanoma [7]. These results indicated that miR-205 might play a tumor suppressor in specific tissues. Therefore, further research is needed to understand the mechanism of action and function of miR-205 in different tumor tissues.
PTEN is a phosphatase that can affect the activity of Akt, a regulator involved in cell survival, which belongs to tumor suppressor gene family. Clinical studies have demonstrated that different degrees of PTEN gene mutation or loss exist in a variety of advanced tumor. Study found that the expression of PTEN in non small cell lung cancer (NSCLC) group was significantly lower than that of adjacent group, and irrelevant with gender, age and lymph node metastasis in patients [8]. There was low expression or no expression of PTEN in 74% regu/117) There was low expres [9]. The negative stained rate of PTEN inssion of PTEN in 74%regulator involved in cell survival, which belongs to tumor suppressor gene family. ssues, which indicate that PTENegative stained rate of PTEN inssion of PTEN in 74%regulator inv [10]. LimNegative stained rate of PTEN inssion of PTEN in 74%regulator involved in cell survival, which belongs to tumor suppressor gene family. ssues, which ies patients of PTEN positive group was significantly higher than in PTEN negative group in the period of IV. It suggested that the expression level of PTENof PTEN in 74%regulator involved in cell survival, which belongs to tumor suppressor gene family. ssue of PTEN could significantly shorten the patient’s survival. In conclusion,pression level of PTENof PTEN in 74%regulator involved in cell survival, which belwith occurrence, developmentn, preprognosis of lung cancer.
In this study, we detected the expression of miR-205 and PTEN in lung cancer tissues. The functional role of miR-205 in lung cancer and its influence on PTEN protein and mRNA expression, and cell proliferation, was studied after transfection of miR-205 mimics or inhibitor into A549 cell line. The solution to these problems helped to illustrate that whether PTEN is a target of miR-205 in the pathological mechanism of lung cancer.
Materials and methods
Patients and tissue samples
46 specimens of lung cancer and adjacent normal tissue (The distance of 5-10 cm from tumor) were obtained from the Department of Pathology, Liaoning cancer hospital, between December 2010 and December 2012. This study was approved by our institution’s Research Ethics Committee; informed consent was obtained from each patient. The age of patients ranged from 42 to 75 years, mean 54.33 ± 7.79 years, and included 32 males and 14 females, 25 cases of squamous cell carcinoma and 21 cases adenocarcinoma.
Western blot analysis
The lung cancer specimens paired to adjacent normal tissues or approximately 2 × 108 cells was treated by 500 μl lysate, and then ultrasound in the ice bath, the supernatant was collected after low temperature centrifugation. The BAC method was chosen to detect the concentration of protein. Equal amounts of sample protein was separated by 12% polyacrylamide gel electrophoresis, and then transferred to the PVDF membrane. An anti-PTEN and anti-GAPDH were incubated overnight at 4 degrees after 5% BSA closed 2 h, subsequently incubated with secondary antibodies labeled with horseradish peroxidase for 1 h at room temperature, then ECL exposed and developed. Each sample was repeated 3 times. The ratio of target protein to the corresponding GAPDH protein optical density as the relative expression of the target protein.
Quantitative real time (RT)-PCR (qRT-PCR)
According to Takara Company SYBR® PrimeSCript® miRNA RT-PCR Kit instructions, the reaction begins after 1 μg total RNA and necessary reverse transcription reaction reagents were added, reaction conditions: 37°C, 60min, 85°C, 5 s. The expression of miR-205 which template was reverse transcription synthesized cDNA was observed by real-time PCR. According to Takara Company SYBR® PrimeSCript® miRNA RT-PCR Kit instructions, the reaction begins after SYBR® Premix Ex TaqTM II (2 ×), PCR Forward Primer (10 μmol L-1), Uni-miR qPCR Primer (10 μmol L-1), template (cDNA solution) were added, the reaction system was 25μL, reaction conditions: 95°C, 5 min; 95°C, 10 s, 60°C, 20 s, 40 cycles. U6B was used as a reference. RT-PCR was performed using the following primers: miR-205, 5’-CTTGTCCTTCATTCCACCGGA-3’ (forward) and 5’-TGCCGCCTGAACTTCACTCC-3’ (reverse) [12]; PTEN, 5’-TGTGGTCTGCCAGCTAAAGG-3’ (forward) and 5’-CGGCTGAGGGAACTCAAAGT-3’ (reverse) [12]; U6,5’-CTCGCTTCGGCAGCACA-3’/5’-AACGCTTCACGAATTTGCGT-3’ [13].
miR-205 target gene prediction
PicTar, TargetScan and miRanda were queried to Predict target gene of MiR-205, and then, a candidate target genes of the MiR-205 was Determined according to the result of target site combination.
Cell culture and transfection
A549 cells were seeded in 10% FBS + RPMI-1640 medium. Cultures were maintained in a humidified atmosphere of 5% CO2 at 37°C, and Conventional passaged. Cells were divided into negative control group, miR-205 mimics (miR-205), mimic NC, miR-205 inhibitor and inhibitor NC group [14]. Cells in logarithmic growth phase were transfected with negative control, miR-205 mimics or miR-205 inhibitor respectively according to Lipofectamin 2000 instructions.
Luciferase reporter assay
The target sequences of PTEN wild type 3’UTR were cloned into a luciferase vector which contained the Renilla luciferase gene. In addition, mutant 3’UTR was also cloned. HEK-293T cells were cotransfected with miR-NC or miR-205 mimics using Lipofectamine 2000 (Invitrogen). Cells were collected 48 h after transfection and analyzed using the Dual-Luciferase Reporter Assay System (Promega, Madison, WI), and luciferase activity values were normalized relative to that of the Renilla luciferase internal control.
Cell Counting Kit-8
Cell proliferation analysis was performed using the Cell Counting Kit-8 (CCK-8) (Tongren, Shanghai, China). The optimum reaction time of CCK-8 was determined to be 2.5 h. When 80% confluent, A549 cells (100 μL/well) were seeded into 96-well plates, and were left untransfected or transfected with miR-205-sense or miR-205-antisense, and further incubated for 12, 24, 48, and 72 hours using three replicates. Approximately 10 μL of tetrazolium salt, WST-8, was added to each well for 1 hour. The optical density (OD), at 450 nm, of each well was determined by microplate reader.
Statistical analysis
Statistical analysis of data was performed using SPSS19.0 software. Statistical evaluation was performed using one-way analysis of variance (ANOVA; P < 0.05) and Student’s t-test. All data from experiments were expressed as mean ± SD unless otherwise stated.
Results
Expression of MiR-205 was upregulated in lung cancer tissues
The expressions of miR-205 in 46 lung cancer specimens and their corresponding adjacent normal tissues (The distance of 5-10 cm from tumor) were detected by RT-PCR. We found that the expression of miR-205 was significantly higher in lung cancer specimens than in adjacent normal tissues (P < 0.01; Figure 1). The results suggested that the up-regulation of miR-205 might be involved in human lung cancer disease processes.
Figure 1.
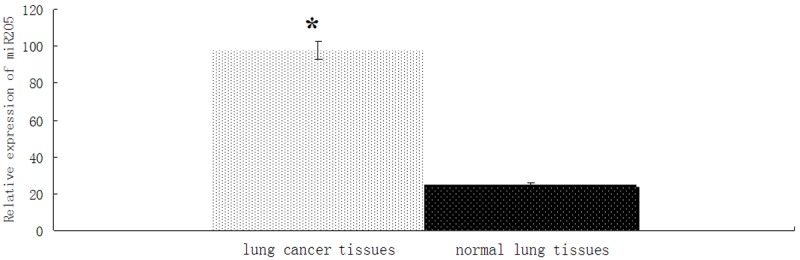
The expression of MiR-205 was up-regulated in lung cancer tissue. U6 RNA was used as an endogenous normalizer and the relative combined expression levels of miR-205 are shown. 2-ΔΔCt values ± SEM, n = 46, *P < 0.01.
Expression of PTEN mRNA was downregulated in lung cancer tissues
The expressions of PTEN mRNA in 46 lung cancer specimens and their corresponding adjacent normal tissues (The distance of 5-10 cm from tumor) were detected by RT-PCR. We found that the expression of PTEN mRNA was significantly lower in lung cancer specimens than in adjacent normal tissues (P < 0.01; Figure 2). The results suggested that the down-regulation of PTEN mRNA might be involved in human lung cancer disease processes.
Figure 2.
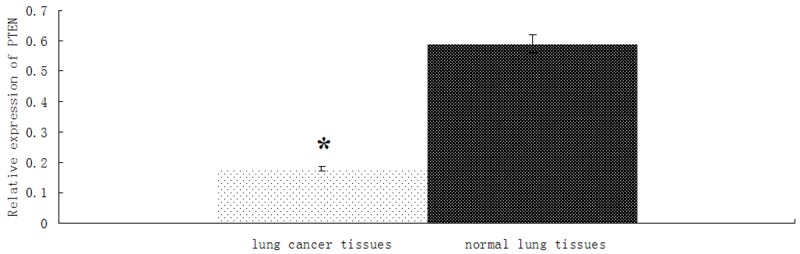
The expression of PTEN is downregulated in lung cancer tissue. U6 RNA was used as an endogenous normalizer and the relative combined expression levels of miR-205 are shown. 2-ΔΔCt values ± SEM, n = 46, *P < 0.01.
Expression of PTEN protein was downregulated in lung cancer tissues
The expression of PTEN in 46 lung cancer specimens and their corresponding adjacent normal tissues (The distance of 5-10 cm from tumor) was detected by western blot method. We found that the expressionof PTEN was significantly decreased in lung cancer specimens compared to adjacent normal tissues (P < 0.01; Figure 3A, 3B).
Figure 3.
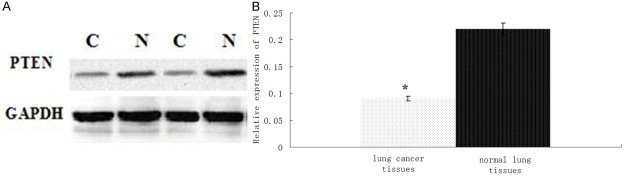
The expression of PTEN protein is downregulated in lung cancer tissue. (A) PTEN protein expression was determined by Western blot in lung cancer (C) and normal tissues (N). (B) Optical density values for combined PTEN protein relative to GAPDH in lung cancer tissues. N = 46. *P < 0.01.
A negative relationship between the expression of miR-205 and PTEN mRNA in lung cancer
The scatter diagram was obtained according to experimental data, and Spearman correlation analysis showed that miR-205 level were negatively correlated with the expression of PTEN (r = -0.594, P < 0.01) in lung cancer tissues (Figure 4).
Figure 4.
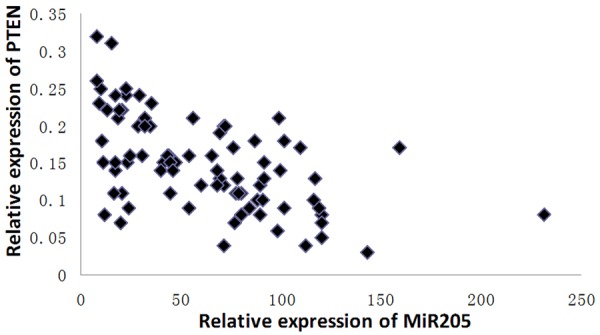
The scatter diagram of the expression of miR-205 and PTEN.
Luciferase reporter assay results
We employed the target gene prediction database to predict MiR 205 target genes, and we found that PTEN mRNA 3’UTR had multiple regions and 2~8 nucleotides of miR-205 Complete complementary (Figure 5A). Luciferase reporter assay results showed that the expression levels of fluorescent protein in PTEN 3’UTR and miR-205 mimics cotransfection group is lower than in PTEN 3’UTR and miR-205 control mimics cotransfection group (P < 0.01); However, the expression levels of fluorescent protein in Mut PTEN 3’UTR and miR 205 mimics cotransfection group is higher than in PTEN 3’UTR and miR-205 mimics cotransfection group (P < 0.01). These results indicated that PTEN is a target gene of MiR205 (Figure 5B).
Figure 5.
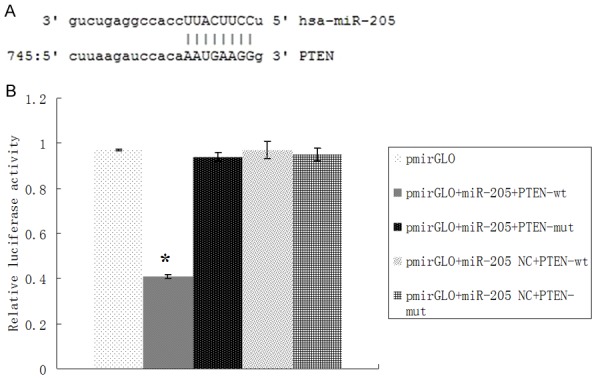
PTEN as a target gene of miR205. A. Schematic diagram of the predicted miR-205 binding site in the PTEN 3’-UTR. B. Luciferase reporter assays in untreated HEK-293T cells, or cotransfected with wild type (wt) or mutant (mut) PTEN 3’-UTR, and miR-133b or miR-133b negative control (NC). *P < 0.01.
Determination of miR-205 mimics transfection efficiency
The relative expression of miR-205 in A549 cells transfected by miR-205 mimics and normal control was determined by Real-time PCR. Compared with transfection of miR-205 normal control, the expression of miR-205 was increased significantly after transfection of miR-205 mimics (P < 0.01, Figure 6A), and decreased significantly after miR-205 mimics inhibitors (P < 0.01, Figure 6B).
Figure 6.
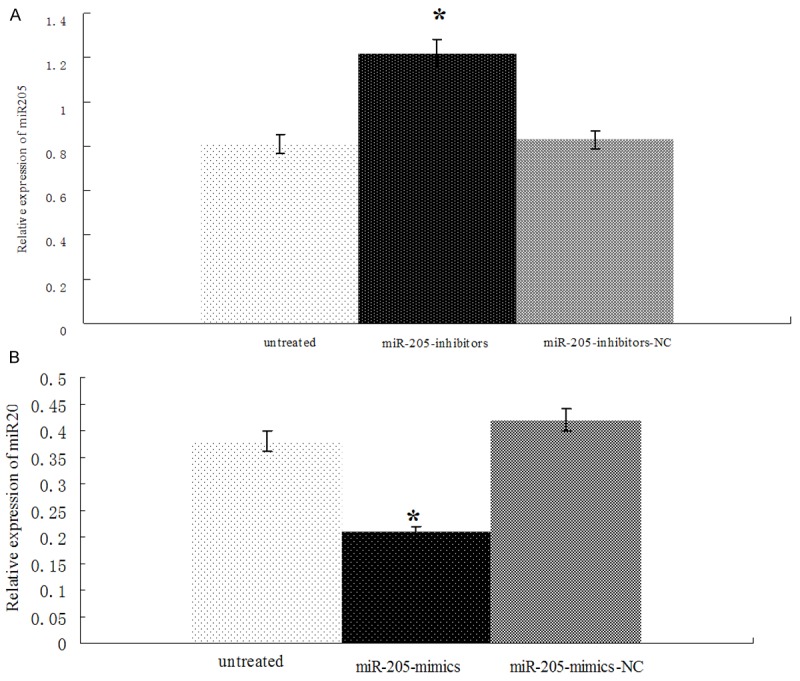
Determination of miR-205 mimics or miR-205 inhibitors transfection efficiency. A. The expression of miR-205 was significantly increased after miR-205 mimics’ transfection into A549 cells. B. The expression of miR-205 was significantly decreased after miR-205 inhibitors transfection into A549 cells. *P < 0.01.
Effect of miR-205 on the expression of PTEN protein and mRNA in A549 cells
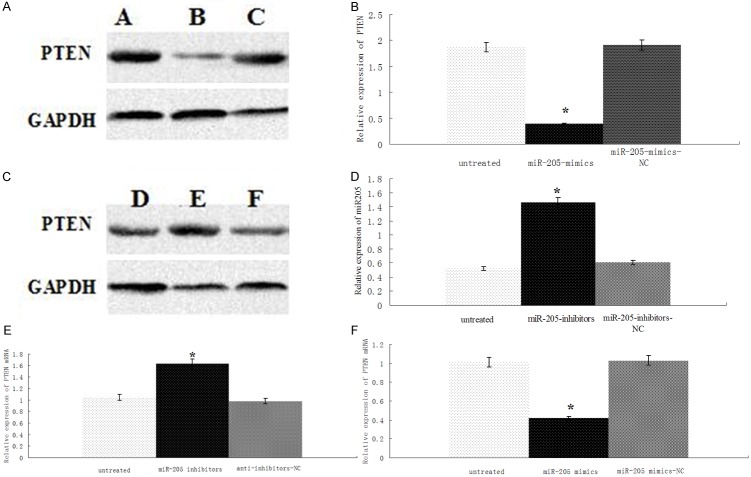
We employed western blot methods to study the effects of miR-205 mimics or inhibitor on the expression of PTEN in A549 cells. We found that the expression of PTEN in A549 cells was significantly down-regulated or up-regulated after miR-205 mimics (Figure 7A and 7B) or inhibitor (Figure 7C and 7D) was transfected into A549 cells respectively (P < 0.01 for both). We also observed the expression of PTEN mRNA in A549 cells by Realtime PCR, and we found that the expression level of PTEN in A549 cells was significantly down-regulated or up-regulated after miR-205 mimics (Figure 7E) or inhibitor (Figure 7F) was transfected into A549 cells respectively (P < 0.01 for both).
Figure 7.
Effect of miR-205 mimics and inhibitor on PTEN mRNA and protein levels in A549 cells. A549 cells were untreated (A, D), treated with miR-205 mimics (B), treated with miR-205 NC (C), treated with miR-205 inhibitors (E), treated with miR-205 inhibitors NC(F). (A, C) The expression of PTEN protein was determined by Western blot. (B, D) Optical density values for combined PTEN protein relative to GAPDH. (E) The expression of PTEN mRNA was determined by Realtime PCR, (F) Optical density values for combined PTEN mRNA relative to U6, n = 46. *P < 0.01.
MiR-205 inhibits cell proliferation of A549 cells
We employed Cell Counting Kit-8 (CCK-8) methods to study the effects of miR-205 mimics or inhibitor on cell proliferation of A549 cells. Cell proliferation assays were performed at 12 h, 24 h, 48 h and 72 h after miR-205-mimics or miR-205-inhibitors transfection of A549 cells. Compared with untreated or miR-205-mimics-NC group, the cell proliferation of A549 cells in miR-205-mimics group was decreased significantly (P < 0.05; Figure 8A); compared with untreated or miR-205-inhibitors-NC group, the cell proliferation of A549 cells in miR-205-inhibitors group was increased significantly (P < 0.05; Figure 8B). These results indicate that miR-205 inhibits A549 cells proliferation.
Figure 8.
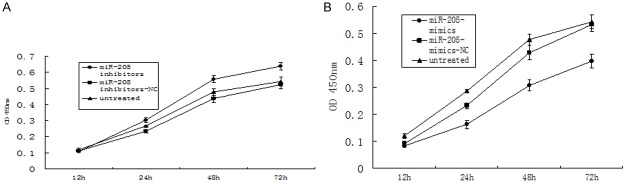
Effect of miR-205 on proliferation of A549 cells. A. A549 cells were untreated or transfected with miR-205-NC or miR-205-mimics for up to 72 h. B. A549 cells were untreated or transfected with miR-205-inhibitor-NC or miR-205- inhibitor for up to 72 h. Cell proliferation was measured by CCK-8. *P < 0.05.
Discussion
miR-205 conducts conflicting functions in human cancers, where it may functions either as a tumor suppressor or an oncogene in a single tumor type depending on targeted mRNA [15-17]. Then, what would miR-205 be like in human lung cancer? We detected the expression of miR-205 in 46 lung cancer specimens and their corresponding adjacent normal tissues by stem-loop RT-PCR. We found the expression of miR-205 was significantly increased in lung cancer specimens compared to adjacent normal tissues. The result implied that the down-regulation of miR-205 might be involved in human lung cancer disease processes, which consistent with previous research [5].
PTEN was the first discovered tumor suppressor gene with dual specificity phosphatase activity, and its mutation, inactivation and decreased expression were closely related with the occurrence, development and prognosis of patients with lung cancer, playing an important role in the tumor chemotherapy resistance [18-22]. We detected the expressions of PTEN mRNA and protein in 46 lung cancer specimens and their corresponding adjacent normal tissues, found the expressions of PTEN mRNA and protein was significantly decreased in lung cancer tissues than in adjacent normal tissues, which consistent with previous research [5]. However, it is a problem whether PTEN is a target of miR-205 in the pathogenesis of lung cancer.
To validate whether PTEN was direct targets of miR-205, at first, the statistical analyses of experimental data were done to draw a conclusion that miR-205 level were negatively correlated with the expression of PTEN in lung cancer tissues, and then we employed the target gene prediction database to predict miR-205 target genes, found that PTEN mRNA 3’UTR had multiple regions and 2~8 nucleotides of miR-205 complete complementary. At last, a dual-luciferase reporter assay was employed; the result showed that PTEN is a target gene of MiR205. But, it is a problem whether PTEN was regulated by miR-205 in lung cancer cells.
We designed miR-205 mimics and inhibitor for verifying the relationship between miR-205 and PTEN. We detected the expression change of PTEN mRNA and protein after miR-205 mimics or inhibitor were transfected into A549 cells. We found that the expression of PTEN protein and mRNA in A549 cells was down-regulated and up-regulated respectively after transfection of miR-205 mimics or inhibitor. These findings support the idea that PTEN might be a target of miR-205 in lung cancer. And then, CCK-8 tests were used to evaluate the effects of miR-205 on cell proliferation of A549 cells. We found that miR-205 overexpression caused A549 cell growth inhibition.
Thus, we hypothesized that miR-205 may function as a tumor suppressor, and the abnormal expression of miR-205 might be an important factor in lung cancer incidence.
Acknowledgements
This study was financially supported (project 2010225032, and 2013225021) through the Shenyang Science and Technology Program. We give special thanks to all the teachers at the Department of Intensive Care Unit and Department of Pathology of Liaoning cancer hospital for their help and support.
Disclosure of conflict of interest
None.
References
- 1.Gregory PA, Bracken CP, Bert AG, Goodall GJ. MicroRNAs as regulators of epithelial-mesenchymal transition. Cell Cycle. 2008;7:3112–8. doi: 10.4161/cc.7.20.6851. [DOI] [PubMed] [Google Scholar]
- 2.Sempere LF, Christensen M, Silahtaroglu A, Bak M, Heath CV, Schwartz G, Wells W, Kauppinen S, Cole CN. Altered MicroRNA expression confined to specific epithelial cell subpopulations in breast cancer. Cancer Res. 2007;67:11612–20. doi: 10.1158/0008-5472.CAN-07-5019. [DOI] [PubMed] [Google Scholar]
- 3.Schaefer A, Jung M, Mollenkopf HJ, Wagner I, Stephan C, Jentzmik F, Miller K, Lein M, Kristiansen G, Jung K. Diagnostic and prognostic implications of microRNA profiling in prostate carcinoma. Int J Cancer. 2010;126:1166–76. doi: 10.1002/ijc.24827. [DOI] [PubMed] [Google Scholar]
- 4.Chung TK, Cheung TH, Huen NY, Wong KW, Lo KW, Yim SF, Siu NS, Wong YM, Tsang PT, Pang MW, Yu MY, To KF, Mok SC, Wang VW, Li C, Cheung AY, Doran G, Birrer MJ, Smith DI, Wong YF. Dysregulated microRNAs and their predicted targets associated with endometrioid endometrial adenocarcinoma in Hong Kong women. Int J Cancer. 2009;124:1358–65. doi: 10.1002/ijc.24071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Markou A, Tsaroucha EG, Kaklamanis I, Fotinou M, Georgoulias V, Lianidou ES. Prognostic value of mature micmRNA-21 and microRNA 205 overexpression in non-small cell lung cancer by quantitative real-time RT-PCR. Clin Chem. 2008;54:1696–704. doi: 10.1373/clinchem.2007.101741. [DOI] [PubMed] [Google Scholar]
- 6.Fletcher AM, Heaford AC, Trask DK. Detection of metastatic head and neck squamous cell carcinoma using the relative expression of tissue-specific mir-205. Transl Oncol. 2008;1:202–208. doi: 10.1593/tlo.08163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Dar AA, Majid S, de Semir D, Nosrati M, Bezrookove V, Kashani-Sabet M. miRNA-205 suppresses melanoma cell proliferation and induces senescence via regulation of E2FI protein. J Biol Chem. 2011;286:16606–14. doi: 10.1074/jbc.M111.227611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Wang L, Xu S, Yue W, Zhao X, Zhang L, Wang Y. Expression and clinical significance of mTOR and PTEN in non-small cell lung cancer. Zhong Guo Fei Ai Za Zhi. 2010;13:717–21. doi: 10.3779/j.issn.1009-3419.2010.07.11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Marsit CJ, Zheng S, Aldape K, Hinds PW, Nelson HH, Wiencke JK, Kelsey KT. PTEN expression in non-small-cell lung cancer: evaluating its relation to tumor characteristics, allelic loss, and epigenetic alteration. Hum Pathol. 2005;36:768–76. doi: 10.1016/j.humpath.2005.05.006. [DOI] [PubMed] [Google Scholar]
- 10.Yanagawa N, Leduc C, Kohler D, Saieg MA, John T, Sykes J, Yoshimoto M, Pintilie M, Squire J, Shepherd FA, Tsao MS. Loss of phosphatase and tensin homolog protein expression is an independent poor prognostic marker in lung adenocarcinoma. J Thorac Oncol. 2012;7:1513–21. doi: 10.1097/JTO.0b013e3182641d4f. [DOI] [PubMed] [Google Scholar]
- 11.Lim WT, Zhang WH, Miller CR, Watters JW, Gao F, Viswanathan A, Govindan R, McLeod HL. PTEN and phosphorylated AKT expression and prognosis in early-and late-stage non small cell lung cancer. Oncol Rep. 2007;17:853–857. [PubMed] [Google Scholar]
- 12.Zhang G, Hou X, Li Y, Zhao M. MiR-205 inhibits cell apoptosis by targeting phosphatase and tensin homolog deleted on chromosome ten in endometrial cancer ishikawa cells. BMC Cancer. 2014;14:440–8. doi: 10.1186/1471-2407-14-440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chen X, Wang K, Xu Z, Li SJ, Liu Q, Fu DH, Wang X, Wu B. MiR-133b regulates bladder cancer cell proliferation and apoptosis by targeting Bcl-w and Akt1. Cancer Cell Int. 2014;14:70. doi: 10.1186/s12935-014-0070-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wu J, Meng Z, Jiang M, Trippler M, Broering R, Bucchi A, Sowa JP, Dittmer U, Yang D, Roggendorf M, Gerken G, Lu M, Schlaak JF. Hepatitis B virus suppresses toll-like receptor-mediated innate immune responses in murine parenchymal and nonparenchymal liver cells. Hepatology. 2009;49:1132–1140. doi: 10.1002/hep.22751. [DOI] [PubMed] [Google Scholar]
- 15.Savad S, Mehdipour P, Miryounesi M, Shirkoohi R, Fereidooni F, Mansouri F, Modarressi MH. Expression analysis of MiR-21, MiR-205, and MiR-342 in breast cancer in Iran. Asian Pac J Cancer Prev. 2012;13:873–7. doi: 10.7314/apjcp.2012.13.3.873. [DOI] [PubMed] [Google Scholar]
- 16.Zhao BS, Liu SG, Wang TY, Ji YH, Qi B, Tao YP, Li HC, Wu XN. Screening of microRNA in patients with esophageal cancer at same tumor node metastasis stage with different prognoses. Asian Pac J Cancer Prev. 2012;13:139–43. doi: 10.7314/apjcp.2013.14.1.139. [DOI] [PubMed] [Google Scholar]
- 17.Qin AY, Zhang XW, Liu L, Yu JP, Li H, Wang SZ, Ren XB, Cao S. MiR-205 in cancer: an angel or a devil? Eur J Cell Biol. 2013;92:54–60. doi: 10.1016/j.ejcb.2012.11.002. [DOI] [PubMed] [Google Scholar]
- 18.Hollander MC, Blumenthal GM, Dennis PA. PTEN loss in the continuum of common cancers, rare syndromes and mouse models. Nat Rev Cancer. 2011;11:289–301. doi: 10.1038/nrc3037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Shi YJ, Benjamin E. PTEN at a glance. J Cell Sci. 2012;125:4687–92. doi: 10.1242/jcs.093765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kim EJ, Jeong JH, Bae S, Kang S, Kim CH, Lim YB. mTOR inhibitors radio sensitize PTEN-deficient non-small-cell lung cancer cells harboring an EGFR activating mutation by inducing autophagy. J Cell Biochem. 2013;114:1248–56. doi: 10.1002/jcb.24465. [DOI] [PubMed] [Google Scholar]
- 21.Yamamoto C, Basaki Y, Kawahara A, Nakashima K, Kage M, Izumi H, Kohno K, Uramoto H, Yasumoto K, Kuwano M, Ono M. Loss of PTEN expression by blocking nuclear translocation of EGR1 ingefitinib-resistant lung cancer cells harboring epidermal growth factor receptor-activating mutations. Cancer Res. 2010;70:8715–25. doi: 10.1158/0008-5472.CAN-10-0043. [DOI] [PubMed] [Google Scholar]
- 22.Bidkhori G, Moeini A, Masoudi-Nejad A. Modeling of tumor progression in NSCLC and intrinsic resistance to TKI in loss of PTEN expression. PLoS One. 2012;7:e48004. doi: 10.1371/journal.pone.0048004. [DOI] [PMC free article] [PubMed] [Google Scholar]