Abstract
Background
Spinocerebellar Ataxias (SCAs) belong to polyglutamine repeat disorders and are characterized by a predominant atrophy of the cerebellum and the pons.
Methods
Proton magnetic resonance spectroscopy (1H MRS) using an optimised semiadiabatic localization by adiabatic selective refocusing (semi-LASER) protocol was performed at 3 T to determine metabolite concentrations in the cerebellar vermis and pons of a cohort of patients with SCA1 (n = 16), SCA2 (n = 12), SCA3 (n = 21), SCA7 (n = 12) and healthy controls (n = 33).
Results
Compared to controls, patients displayed lower total N-acetylaspartate and, to a lesser extent, lower glutamate, reflecting neuronal loss/dysfunction, while the glial marker, myo-inositol, was elevated. Patients also showed higher total creatine as reported in Huntington disease, another polyglutamine repeat disorder. There was a strong correlation between the Scale for the Assessment and Rating of Ataxia and the neurometabolites in both affected regions of patients. Principal component analyses confirmed that neuronal metabolites (total N-acetylaspartate and glutamate) were inversely correlated in the vermis and the pons to glial (myo-inositol) and energetic (total creatine) metabolites, as well as to disease severity (motor scales). Neurochemical plots with selected metabolites also allowed the separation of SCA2 and SCA3 from controls.
Conclusion
The neurometabolic profiles detected in patients underlie cell-specific changes in neuronal and astrocytic compartments that cannot be assessed by other neuroimaging modalities. The inverse correlation between metabolites from these two compartments suggests a metabolic attempt to compensate for neuronal damage in SCAs. Because these biomarkers reflect dynamic aspects of cellular metabolism, they are good candidates for proof-of-concept therapeutic trials.
Keywords: Spinocerebellar ataxia, biomarker, movement disorders, NMR spectroscopy, neurochemical profile
INTRODUCTION
Spinocerebellar ataxias (SCA 1, 2, 3, 7) are polyglutamine repeat disorders inherited as an autosomal dominant trait. The cerebellum and the brainstem are mainly affected in SCAs,1 their atrophy being detectable several years before the predicted onset of motor symptoms.2 Progressive ataxia is the prominent symptom of all SCAs. In SCA1, SCA2, SCA3 and SCA7, ataxia is often accompanied by pyramidal signs, sensory disturbances, muscle wasting and brainstem oculomotor signs.3 SCA7 is clearly distinguished from all other SCAs by the invariable presence of pigmentary retinal dystrophy. Several scales exist to assess disease state but the Scale for the Assessment and Rating of Ataxia (SARA) is the best studied and validated so far.4 SARA is however of no use in presymptomatic individuals and is also not able to identify subtle differences that can serve as endpoints in future therapeutic trials.5 Since SCAs are rare, studying a large cohort with adequate power for clinical trials is a major hindrance.6 Magnetic resonance spectroscopy (MRS) offers a non-ionizing and non-invasive approach for quantitative information on the relationship between metabolism and clinical function in patients with neurodegenerative diseases.7 Moreover, MRS allows the identification of in vivo alterations in brain metabolite concentrations that are likely to occur prior to brain atrophy8 and may be amended by early therapeutic intervention.
Changes in brain metabolites in SCAs have been previously studied with MRS but mainly on a 1.5 Tesla MR system, with smaller cohorts and none included SCA7 patients.9-12 Most of these studies only reported metabolite ratios, which complicate data interpretation.13,14 A few MRS studies have also been implemented on a 4 Tesla MR system but only in SCA1, SCA2 and SCA6.15,16 Furthermore, 4 Tesla systems are uncommon, and MRS data obtained in patients with SCAs on clinical 3 Tesla scanners, such as those now widely available in hospitals, are still lacking. In this context, the purpose of our study was to identify in vivo metabolic biomarkers in a large cohort of patients with SCA1, SCA2, SCA3 and SCA7 on a 3 Tesla MR system commonly used in hospitals.
METHODS
Experimental procedures were approved by the local ethics committee (AOM10094, CPP Ile de France VI, Ref: 105-10). MRS data were acquired on a 3 Tesla whole-body Siemens Magnetom Trio scanner (Siemens Medical Solutions, Erlangen, Germany). All participants were over 18 years and signed a written informed consent before participating in the study.
Patients and Controls
We recruited 16 patients with SCA1, 12 patients with SCA2, 21 patients with SCA3 and 12 patients with SCA7 as part of the BIOSCA study (NCT01470729). All patients underwent neurological examinations. Thirty-three healthy volunteers with no history of neurologic diseases and with a median age, gender and BMI similar to the patient groups were also recruited (Table). SARA was used to evaluate the severity of the cerebellar ataxia.4 The scale ranges from 0 (no cerebellar symptoms) to 40 (most severe cerebellar symptoms).
Table.
Demographic and spectroscopic parameters of all subjects.
| Variable | Control | SCA1 | SCA2 | SCA3 | SCA7 |
|---|---|---|---|---|---|
| No of participants | 33 | 16 | 12 | 21 | 12 |
| Gender (M/F) | 15/18 | 9/7 | 7/5 | 9/12 | 6/6 |
| Age (yrs) | 48±13 | 44±16 | 45±13 | 51±12 | 46±14 |
| BMI (kg/m2) | 25±4 | 24±6 | 26±5 | 24±4 | 23±3 |
| SARA score | 0.7±0.9 | 11.1±6.2 † | 12.6±6.0 † | 13.2±7.1 † | 9.2±7.2 † |
| CAG length | 47±7 | 40±3 | 69±6 | 42±5 | |
| Disease duration (yrs) | 7±7 | 10±6 | 9±5 | 9±5 | |
| Lw vermis (Hz) | 7.3±0.6 | 6.4±1.1 * | 5.6±1.1 † | 6.6±1.5 * | 6.7±1.2 |
| Lw pons (Hz) | 8.1±0.9 | 8.0±1.6 | 6.0±1.5 † | 7.6±1.1 | 7.7±0.8 |
| SNR vermis | 33±6 | 31±6 | 27±5 # | 31±4 | 29±4 |
| SNR pons | 21±3 | 19±3 | 16±3 # | 16±3 † | 18±4 |
| % CSF vermis | 9±4 | 20±8 † | 33±10 † | 22±8 † | 19±8 † |
| % CSF pons | 2±2 | 2±2 | 8±5 † | 3±2 | 2±1 |
SNR: signal-to-noise ratio; Lw: water linewidth; CSF: cerebrospinal fluid. Data are presented as mean ± SD and compared by one-way ANOVA with Dunnett correction.
p < 0.05
p < 0.01
p < 0.001 represent significant difference between SCAs and controls.
MR Protocol
A modified semiadiabatic localization by adiabatic selective refocusing (semi-LASER) sequence was used for 1H MRS.17 This sequence provides approximately twice the signal-to-noise ratio (SNR) compared to the stimulated-echo acquisition mode (STEAM) sequence, is less prone to motion artifacts compared to the spin echo full-intensity-acquired localization (SPECIAL) sequence and has the lowest chemical shift displacement artefact.18 3D T1-weighted volumetric images (TR = 2530 ms, TE = 3.65 ms, 1 mm isotropic, FOV = 256 × 176 mm2, matrix size = 256 × 256) were acquired for spatial normalization and localization of brain volumes. Shimming was performed on a 25 × 10 × 25 mm3 volume-of-interest (VOI) in the vermis, and a 16 × 16 × 16 mm3 VOI in the pons, using a fast automatic shimming technique with echo-planar signal trains utilizing mapping along projections, FAST(EST)MAP.19 The radiofrequency power for the 90° asymmetric pulse of the semi-LASER sequence and the variable pulse power and optimised relaxation delays (VAPOR) water suppression pulses were calibrated for each VOI. The power for the 90° excitation pulse also served as the basis for setting the power for the outer volume suppression (OVS) pulses. Siemens 32 channels head coil was used to collect signals from the VOI in the vermis and pons of participants using the semi-LASER sequence (TR = 5000 ms, TE = 28 ms, Averages = 64). VAPOR water suppression pulses in combination with 3D OVS pulses allowed for improved localization and water suppression performance. Two unsuppressed water spectra were acquired: one for eddy current correction (the radiofrequency pulses of the VAPOR scheme were turned off) and one for use as reference for metabolite quantification (VAPOR and OVS schemes turned off in order to eliminate magnetization transfer effects). Unsuppressed water spectra were also acquired at a series of TE values (TE = 28 – 4000 ms; TR = 15000 ms for full relaxation) to evaluate the cerebrospinal fluid (CSF) contribution to the VOI.20 The total acquisition time per voxel, including power calibrations, metabolite spectrum acquisition and water reference acquisition for quantification and CSF evaluation, was approximately 20 minutes.
Metabolite Quantification
Spectral processing and metabolite quantification were performed in the frequency domain using LCModel21 as described previously.22 The model spectra (basis set) were generated to include the following metabolites: alanine (Ala), ascorbate (Asc), aspartate (Asp), creatine (Cr), γ-aminobutyric acid (GABA), glycerophosphorylcholine (GPC), phosphorylcholine (PCho), phosphocreatine (PCr), glucose (Glc), glutamine (Gln), glutamate (Glu), glutathione (GSH), myo-inositol (myo-Ins), scyllo-inositol (sIns), lactate (Lac), N-acetylaspartate (NAA), N-acetylaspartylglutamate (NAAG), phosphorylethanolamine (PE), taurine (Tau) and experimentally measured macromolecules (Mac) as illustrated in Supplementary eFigure 1. Pre-processing steps included shot-to-shot phase and frequency correction as well as eddy-current compensation using the unsuppressed water spectra acquired from the same VOI. Metabolite concentrations were obtained using water as an internal concentration reference. Concentrations were corrected for CSF content determined by fitting the integrals of the unsuppressed water spectra at different TE values with a bi-exponential fit.20 Tissue water content was assumed at 82% for vermis and 72% for pons.23,24 T2 of CSF was fixed at 740 ms based on experimental measurement. The water signal at TE = 28 ms used as concentration reference was corrected for signal loss due to T2 relaxation. Since metabolites have longer T2 than water, the loss of signal due to T2 relaxation of metabolites at TE = 28 ms was neglected.22 Tissue concentrations of glucose and lactate were corrected for CSF contribution to the VOI assuming 3.2 mM of glucose and 1.8 mM of lactate in CSF.15 Only metabolites quantified with mean Cramér-Rao lower bounds (CRLB), which are estimated errors of metabolite quantification, ≤ 20% are reported. The 20% CRLB threshold allows the selection of the most reliably quantified metabolites, as outlined by the LCModel manual and the MRS Consensus Group.25,26 We calculated and reported the average of all concentration values of reliably quantified metabolites including values with high CRLB but excluding metabolites with CRLB = 999%. Only the sum of metabolites was reported if the correlation between two metabolites was consistently high (correlation coefficient < - 0.7) in a given region (e.g. total creatine, tCr = Cr + PCr). Similarly, when the correlation coefficient was in the range of -0.5 to -0.7, the sum of the two correlated metabolites was reported in addition to the individual metabolites (e.g. NAA, NAAG and tNAA = NAA + NAAG). Spectra with water linewidth > 10 Hz were excluded from data analysis. Approximately 3 minutes were required for preprocessing and quantification of each spectrum.
Statistical Analysis
Participant characteristics, spectral quality and metabolite concentrations – separately for the vermis and the pons – were analyzed using ANOVA with a Dunnett multiple comparison test to compare each SCA type to the controls. For each SCA type separately, using those regions and metabolites showing significant SCA versus control differences, Pearson correlation was performed between the metabolite and SARA score, CAG repeat length and disease duration; these p-values were corrected for multiple testing using the step-down Bonferroni procedure,27 separately for pons and vermis. Using patient data only, clinical characteristics and concentrations of metabolites that were significantly different in patients compared to controls were grouped into two principal components using principal component analysis (PCA). PCA was performed to investigate the global interaction between the selected metabolites and the clinical parameters across patient groups using XLSTAT. In addition, the metabolites that showed the largest difference between patients and controls were plotted against each other to find ratios that could separate patient groups from the control group. A partial least squares discriminant analysis (PLS-DA) was also performed on the entire set of metabolites to identify the variables with class separation information using XLSTAT.
RESULTS
Data quality control
Good quality spectra with a high SNR and excellent resolution were consistently obtained from both controls and patients. SNR was 6-24% lower and water linewidth 1-2 Hz narrower in some patient groups than controls, likely due to the higher CSF fraction in patients’ vermis and pons, especially in patients with SCA2 in whom cerebellar and brainstem atrophy was the most severe and % CSF highest (Table). Pons data were rejected in one patient with SCA1, one patient with SCA2, one patient with SCA7, 4 patients with SCA3 and 2 healthy control subjects due to broad water linewidth (> 10 Hz). Ala, Asp, Asc, GABA, Lac, PCho and PE did not meet the mean CRLB criterion of ≤ 20% in both the vermis and pons. In addition, NAAG did not meet this criterion in the vermis, whilst Cr, Glc, Gln, GSH, PCr and Tau did not meet the criterion in the pons. Our control data were also part of a two-site reproducibility study previously published in which the two participating centers obtained consistent spectral quality and similar neurochemical concentrations in controls by using the same MRS pulse sequence (semi-LASER) in conjunction with identical calibration and quantification procedures.22
Neurochemical alterations in patients with SCAs
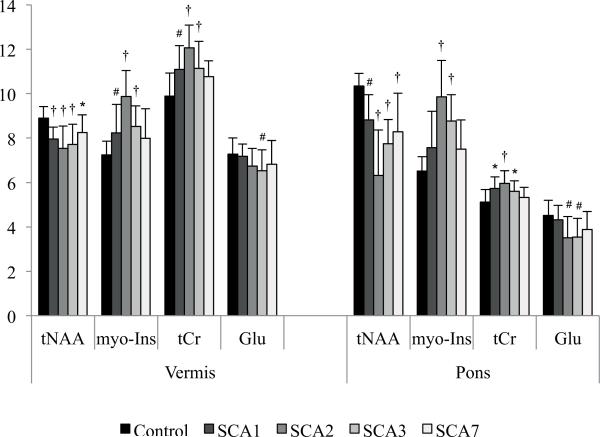
In patients with SCAs, NAA and tNAA were significantly lower than in controls in both the vermis and the pons (Figure 1). Glu was also significantly lower in the vermis of patients with SCA3 and the pons of patients with SCA2 and SCA3 (Figure 1). Lower NAA, a marker of neuronal loss/dysfunction, in the vermis and the pons of patients was associated with significantly higher concentrations of myo-Ins, a putative glial marker, and tCr, a marker of energy metabolism (Figure 1). Neurochemical alterations tended to be more pronounced in patients with SCA2 and SCA3 (Figure 1 and Supplementary eFigure 2 and eFigure 3).
Figure 1.
Mean concentrations of metabolites that showed significant differences in the vermis and the pons of patients with SCA1, 2, 3, 7 vs. controls. p values represent Dunnett-corrected statistically significant differences between patients and controls differences (* p < 0.05; # p < 0.01; † p < 0.001). Lower neuronal markers tNAA and Glu in patients are associated with higher glial marker myo-Ins and higher energy marker tCr. Error bars represent standard deviations (SDs).
Correlation between disease parameters and neurochemical concentrations
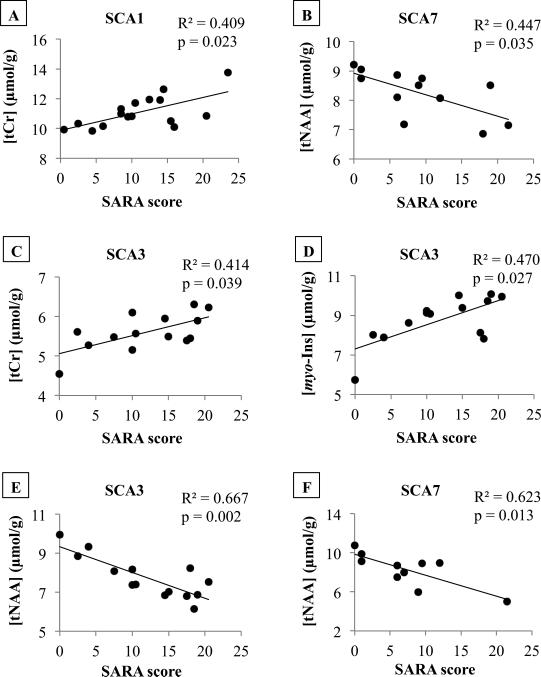
A strong negative correlation was found between SARA scores and tNAA in the vermis of patients with SCA7 and in the pons of patients with SCA3 and SCA7 (Figure 2). Total creatine also strongly correlated with SARA scores in the vermis of patients with SCA1 and in the pons of patients with SCA3 (Figure 2). Furthermore, SARA scores correlated with myo-Ins in the pons of patients with SCA3 (Figure 2). However, CAG repeat length and disease duration did not correlate with the concentration of any neurochemical (Supplementary eTable).
Figure 2.
Correlation between clinical scores and neurochemical concentrations in the vermis and the pons of patients with SCAs. SARA scores correlated with (A) tCr in SCA1 vermis, (B) tNAA in SCA7 vermis, (C) myo-Ins in SCA3 pons, (D) tCr in SCA3 pons, (E) tNAA in SCA3 pons, and (F) tNAA in SCA7 pons. Metabolites that showed Dunnett significance when each SCA type was compared to controls were included in the correlation analysis. p values of the correlations have been corrected for multiple testing with step-down Bonferroni method.
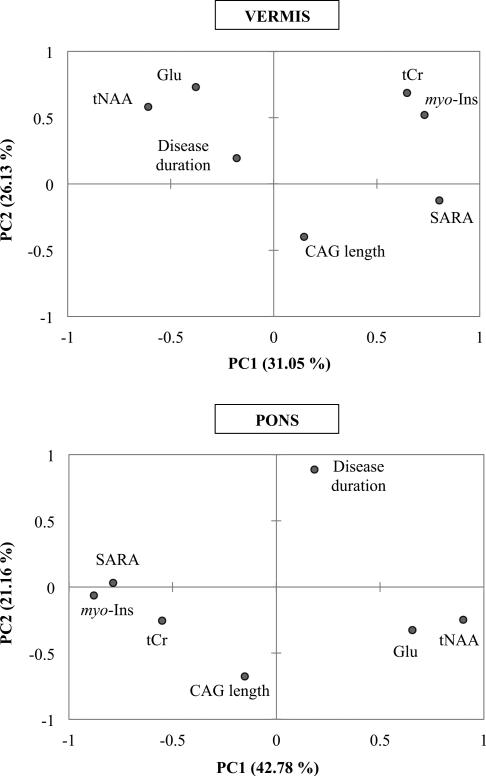
Using patient data only, PCA was used for metabolites with significant differences between patients and controls – tNAA, myo-Ins, tCr and Glu – as well as for disease parameters of interest in SCAs – SARA, CAG repeat length and disease duration. The first two principal components explained 57.18% and 63.94% of the variance in the vermis and the pons respectively (Figure 3). PCA separated the neuronal markers – tNAA and Glu – from the glial (myo-Ins) and energetic markers (tCr) in both vermis and pons (Figure 3). Among the disease parameters, PC1 further showed that the motor score (SARA) was the only one inversely correlated with the neuronal markers and correlated with the glial and energetic markers in both vermis and pons (Figure 3). Of note, PCA showed a negative correlation between CAG repeat length and disease duration (Figure 3).
Figure 3.
Principal component analysis of metabolites of interest and disease characteristics of patients with SCAs. The first two components accounted for 57.4% and 64% variation in the vermis and the pons respectively. PCA was able to separate the neuronal markers – tNAA and Glu – from the energetic marker (tCr), the glial marker (myo-Ins) and the SARA score.
Separation between patients and controls using neurochemical concentrations
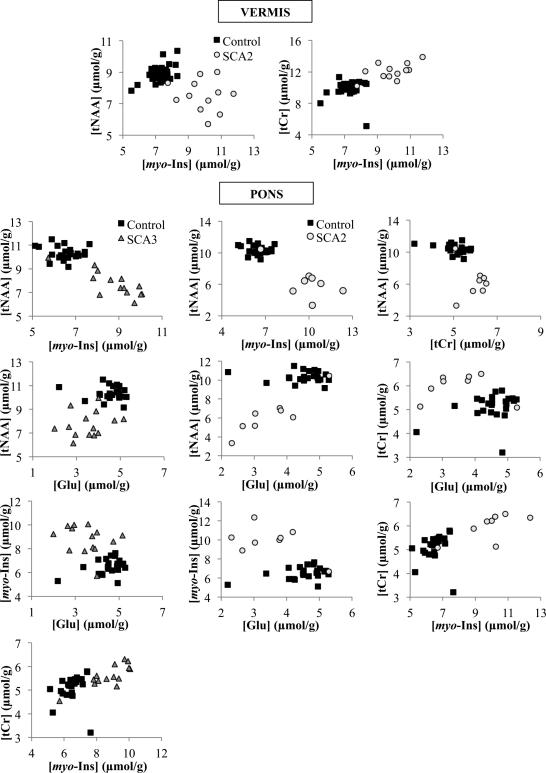
The concentrations of metabolites that showed the largest differences between patients and controls – tNAA, tCr, myo-Ins and Glu – were plotted against each other to determine the separation between patient and control groups. These neurochemical plots demonstrated the separation of patients with SCA2 and SCA3 from controls in both vermis and pons (Figure 4). However, we could not clearly separate patients with SCA1 and SCA7 from controls (data not shown). In addition, we used PLS-DA to identify the variables with class separation information but we were unable to obtain a good separation between the different SCAs (data not shown). Likewise, while MRS allowed discriminating metabolic profiles from patients and controls as well as correlating patients’ metabolic profile with clinical status, it did not reveal metabolic profiles characteristic to any of the SCA types.
Figure 4.
Separation between patients with SCAs and controls by plotting the concentrations of neurochemicals against each other. The concentrations of metabolites that showed significant differences in patients – tNAA, tCr, myo-Ins and Glu – were plotted against each other to determine the ratio that could separate subjects into patient and control groups, with almost no overlap. SCA2 has the best separation with at most one dataset overlapping with controls.
DISCUSSION
This is the first study reporting neurochemical profiling on a large cohort of patients with SCAs (SCA 1, 2, 3 and 7) using a 3 Tesla clinical MR system. In the vermis and the pons, patients displayed lower NAA, a neuronal marker, higher myo-Ins, a glial marker, and higher total Cr compared to controls. Glu, another neuronal marker, was also lower in patients’ vermis (SCA3) and pons (SCA2 and SCA3). Our control data has also been part of a two-site study to establish the reproducibility of our methods, which underscores the robustness of our findings.22 Although the CRLB ≤ 20% criterion is fairly conservative, quantification of some metabolites such as GSH from short-TE spectra still requires caution as their quantification may be sensitive to small changes in the baseline, even with the excellent spectral quality achieved in this study. However, all the main findings reported here focus on metabolites (NAA, myo-Ins, tCr, Glu) that have the most prominent resonances and are therefore visually apparent in spectra.
The significant decrease in NAA and Glu observed in patients with SCAs, likely reflecting neuronal loss/dysfunction in those structures,1 is in agreement with prior MRS studies in SCAs.28 We also observed the decrease of NAA in the pons to be more substantial than that in the vermis. This is similar to the observations in SCA1 studies15 and may be due to a more pronounced pathological involvement and atrophy in the pons.29 On the other hand, the increase in myo-Ins, a six-carbon sugar that serves as an intermediate in the metabolism of membrane and myelin phospholipids,30 can indicate damage to myelin sheaths as a result of neurodegeneration resulting in increased concentration of free myo-Ins. As a glial marker, the increase in myo-Ins levels in response to neuronal loss may also be an attempt to improve glial proliferation to modulate vascular and metabolic activities and therefore to compensate for neuronal loss.31 This pinpoints the important role of non-neuronal cells in SCAs, especially astrocytes that play key roles in brain homeostasis through their neurovascular and neurometabolic coupling with neurons – i.e. neurotransmitter recycling and provision of energy substrates.32
Furthermore, we observed a significant increase in tCr, an energetic marker often used in prior studies as a concentration reference in MRS,29 in both the vermis33 and the pons of patients. Due to the limited ability of the brain to store glucose, the creatine kinase/phosphocreatine (CK/PCr) system is important to keep up with the high energy demands of neurons.34 Since mitochondrial CK activity suppresses formation of free radicals within the mitochondria,35,36 the increase in tCr might be an attempt to increase Cr levels to suppress formation of free radicals and increase neuroprotection. Changes in Cr and PCr have been reported in Huntington disease (HD), another polyglutamine disease that shares pathophysiological commonalities with SCAs.37 Using microwave fixation techniques, which instantaneously inactivate brain enzymatic activities and preserve in vivo levels of analytes, increased levels of Cr and PCr were reported in HD mice brain. The increase of tCr preceded decreased ATP levels as early as 4-weeks of age in presymptomatic mice.38 Similarly, increased tCr39 and significant changes in [PCr] / [Cr] ratio40 have been reported in the brain of HD mouse models at an early age using 1H MRS techniques. Recently, we also described abnormal ratios of inorganic phosphate (Pi) / PCr using 31P MRS during brain activation in patients with HD at an early stage of the disease confirming altered brain bioenergetics in HD.41 Therefore, the increase of tCr in patients with SCAs is comparable to the dysregulation of the tCr pool that we observed in patients and animal models with HD. Accordingly, it would be interesting to determine whether the above-mentioned mechanisms are activated at a presymptomatic stage of SCAs.
Higher disease severity, reflected by higher SARA scores, was associated with lower tNAA concentration and higher concentrations of myo-Ins and tCr in both the vermis and the pons of several SCA types. Moreover, the PCA confirmed that neuronal metabolites – tNAA and Glu – varied inversely in the vermis and the pons to glial metabolites – myo-Ins –, energetic metabolites – tCr – and disease severity – SARA score – in patients with SCAs. This inverse correlation suggests that the metabolic attempt to compensate for neuronal damage is critical to SCAs pathophysiology, even more as the metabolic changes are associated with disease severity scores. The absence of correlations between neurochemical concentrations and CAG repeat length or disease duration may be partly explained by the limited dynamic range in CAG and duration.
Using neurochemical plots with our metabolites of interest – tNAA, myo-Ins, tCr and Glu –, we were able to separate patients with SCA2 and SCA3 from controls but not SCA1, unlike what was previously reported.15 Moreover, metabolic profiles could not discriminate the different SCAs from one another. Altogether, this study indicates that, rather than providing subtype-specific information about SCAs, MRS has the potential to unravel early metabolic/cellular changes in various SCAs, which are likely to occur prior to brain atrophy. Neurometabolic profiles also provide cell-specific information on neurons and astrocytes, which cannot be assessed in vivo by other neuroimaging modalities. Because these biomarkers reflect dynamic aspects of cellular metabolism and mirror disease severity, they may also be of special interest to establish proof-of-concepts for drugs prior to the evaluation of clinical outcome measures in phase III therapeutic trials. Our previous report of highly reproducible data in controls in a two-site study supports the claim that these biomarkers can be used in multicentric trials.22
Supplementary Material
ACKNOWLEDGEMENT
We are very grateful to the patients and volunteers who participated in this study. We will also like to thank the Centre d'Investigation Clinique Pitié Neurosciences, CIC-1422, Département des Maladies du Système Nerveux, Hôpital Pitié-Salpêtrière, Paris, France.
AUTHOR ROLES
Study design and conduct: Henry, Rinaldi, Jauffret, Öz, Durr, Mochel;
Collection, management, analysis and interpretation of the data: Adanyeguh, Henry, Nguyen, Valabregue, Emir, Deelchand, Brice, Eberly, Öz, Durr, Mochel;
Drafting/revision/approval of the manuscript: Adanyeguh, Henry, Nguyen, Rinaldi, Jauffret, Valabregue, Emir, Deelchand, Brice, Eberly, Öz, Durr, Mochel.
Statistical analysis: Adanyeguh, Eberly, Mochel.
Funding sources: The study was funded by the Programme Hospitalier de Recherche Clinique (PHRC, AOM10094, NCT01470729) and the program “Investissements d'avenir” ANR-10-IAIHU-06. The Center for Magnetic Resonance Research is funded by the National Institute of Biomedical Imaging and Bioengineering (NIBIB) grant P41 EB015894 and the Institutional Center Cores for Advanced Neuroimaging award P30 NS076408. Additional support was received from the National Institute of Neurological Diseases and Stroke grant R01 NS070815.
Footnotes
Financial disclosure/conflict of interest: None reported.
FULL FINANCIAL DISCLOSURE OF ALL AUTHORS
No disclosure: Adanyeguh, Nguyen, Rinaldi, Jauffret, Valabregue, Emir, Deelchand.
Henry: Research support by grants from NIH (P41 EB015894, P30 NS076408), Bob Allison Ataxia Research Center at the University of Minnesota and Friedreich's Ataxia Research Alliance.
Brice: Honoraria from the Wolfson Foundation for reviewing the scientific project and from Lundbeck for giving a talk. Research support by grants from the French Agency for Research and European Union.
Eberly: Research support by grants from NIH (R01 NR010731, R01 NS070815, R01 NS035192, R01 NS076665, R03 NS082541, R21 MH094558-01A1, P41 EB015894, P50 DA033942-01, R01 DA023651, R56 DK099137), Bob Allison Ataxia Research Center at the University of Minnesota, Minnesota Partnership for Biotechnology and Medical Genomics, Friedreich's Ataxia Research Alliance, Alzheimer's Association, Academic Health Center (FRD 11.12) and American Diabetes Association.
Öz: Salary and research support by grants from the NIH (R01 NS070815, R01 NS035192, P41 EB015894, P30 NS076408), Bob Allison Ataxia Research Center at the University of Minnesota, CHDI Foundation, Inc., Minnesota Partnership for Biotechnology and Medical Genomics and Friedrich's Ataxia Research Alliance.
Durr: Honoraria from Pfizer Inc. for participating in the Global TTR FAP Genetics Advisory Board. Research support by grants from UCL/High Q Foundation and the French Agency for Research and Pfizer Inc.
Mochel: Research support by grants from INSERM, Carnot Institutes, ASL Foundation, Ultragenyx Pharmaceutical and IPSEN.
REFERENCES
- 1.Yamada M, Sato T, Tsuji S, Takahashi H. CAG repeat disorder models and human neuropathology: similarities and differences. Acta Neuropathol. 2008;115:71–86. doi: 10.1007/s00401-007-0287-5. [DOI] [PubMed] [Google Scholar]
- 2.Jacobi H, Reetz K, du Montcel ST, et al. Biological and clinical characteristics of individuals at risk for spinocerebellar ataxia types 1, 2, 3, and 6 in the longitudinal RISCA study: analysis of baseline data. Lancet Neurol. 2013;12:650–658. doi: 10.1016/S1474-4422(13)70104-2. [DOI] [PubMed] [Google Scholar]
- 3.Durr A. Autosomal dominant cerebellar ataxias: polyglutamine expansions and beyond. Lancet Neurol. 2010;9:885–894. doi: 10.1016/S1474-4422(10)70183-6. [DOI] [PubMed] [Google Scholar]
- 4.Schmitz-Hübsch T, du Montcel ST, Baliko L, et al. Scale for the assessment and rating of ataxia: development of a new clinical scale. Neurology. 2006;66:1717–1720. doi: 10.1212/01.wnl.0000219042.60538.92. [DOI] [PubMed] [Google Scholar]
- 5.Saute JA, Donis KC, Serrano-Munuera C, et al. Ataxia rating scales – psychometric profiles, natural history and their application in clinical trials. Cerebellum. 2012;11:488–504. doi: 10.1007/s12311-011-0316-8. [DOI] [PubMed] [Google Scholar]
- 6.Underwood BR, Rubinsztein DC. Spinocerebellar ataxias caused by polyglutamine expansions: a review of therapeutic strategies. Cerebellum. 2008;7:215–221. doi: 10.1007/s12311-008-0026-z. [DOI] [PubMed] [Google Scholar]
- 7.Hall H, Cuellar-Baena S, Dahlberg C, In't Zandt R, Denisov V, Kirik D. Magnetic resonance spectroscopic methods for the assessment of metabolic functions in the diseased brain. Curr Top Behav Neurosci. 2012;11:169–198. doi: 10.1007/7854_2011_166. [DOI] [PubMed] [Google Scholar]
- 8.Öz G, Nelson CD, Koski DM, et al. Noninvasive detection of presymptomatic and progressive neurodegeneration in a mouse model of spinocerebellar ataxia type 1. J Neurosci. 2010;30:3831–3838. doi: 10.1523/JNEUROSCI.5612-09.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Guerrini L, Lolli F, Ginestroni A, et al. Brainstem neurodegeneration correlates with clinical dysfunction in SCA1 but not in SCA2. A quantitative volumetric, diffusion and proton spectroscopy MR study. Brain. 2004;127:1785–1795. doi: 10.1093/brain/awh201. [DOI] [PubMed] [Google Scholar]
- 10.Lirng JF, Wang PS, Chen HC, Soong BW, Guo WY, Wu HM, Chang CY. Differences between spinocerebellar ataxias and multiple system atrophy-cerebellar type on proton magnetic resonance spectroscopy. PLoS One. 2012;7:e47925. doi: 10.1371/journal.pone.0047925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Mascalchi M, Cosottini M, Lolli F, et al. Proton MR spectroscopy of the cerebellum and pons in patients with degenerative ataxia. Radiology. 2002;223:371–378. doi: 10.1148/radiol.2232010722. [DOI] [PubMed] [Google Scholar]
- 12.Wang PS, Chen HC, Wu HM, Lirng JF, Wu YT, Soong BW. Association between proton magnetic resonance spectroscopy measurements and CAG repeat number in patients with spinocerebellar ataxias 2, 3, or 6. PLoS One. 2012;7:e47479. doi: 10.1371/journal.pone.0047479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ferguson KJ, MacLullich AMJ, Marshall I, Deary IJ, Starr JM, Seckl JR, Wardlaw JM. Magnetic resonance spectroscopy and cognitive function in healthy elderly men. Brain. 2002;125:2743–2749. doi: 10.1093/brain/awf278. [DOI] [PubMed] [Google Scholar]
- 14.Henriksen O. In vivo quantitation of metabolite concentrations in the brain by means of proton MRS. NMR Biomed. 1995;8:139–148. doi: 10.1002/nbm.1940080402. [DOI] [PubMed] [Google Scholar]
- 15.Öz G, Hutter D, Tkáč I, et al. Neurochemical alterations in spinocerebellar ataxia type 1 and their correlations with clinical status. Mov Disord. 2010;25:1253–1261. doi: 10.1002/mds.23067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Öz G, Iltis I, Hutter D, Thomas W, Bushara KO, Gomez CM. Distinct neurochemical profiles of Spinocerebellar Ataxias 1,2,6, and Cerebellar Multiple System Atrophy. Cerebellum. 2011;10:208–217. doi: 10.1007/s12311-010-0213-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Öz G, Tkáč I. Short-echo, single-shot, full-intensity proton magnetic resonance spectroscopy for neurochemical profiling at 4 T: Validation in the cerebellum and brainstem. Magn Reson Med. 2011;65:901–910. doi: 10.1002/mrm.22708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Boer VO, van Lier AL, Hoogduin JM, Wijnen JP, Luijten PR, Klomp DW. 7-T 1H MRS with adiabatic refocusing at short TE using radiofrequency focusing with a dual-channel volume transmit coil. NMR Biomed. 2011;241:038–1046. doi: 10.1002/nbm.1641. [DOI] [PubMed] [Google Scholar]
- 19.Gruetter R, Tkac I. Field mapping without reference scan using asymmetric echo-planar techniques. Magn Reson Med. 2000;43:319–323. doi: 10.1002/(sici)1522-2594(200002)43:2<319::aid-mrm22>3.0.co;2-1. [DOI] [PubMed] [Google Scholar]
- 20.Ernst T, Kreis R, Ross BD. Absolute quantitation of water and metabolites in the human brain. I. Compartments and water. J Magn Reson. 1993;102:1–8. [Google Scholar]
- 21.Provencher SW. Estimation of metabolite concentrations from localized in vivo proton NMR spectra. Magn Reson Med. 1993;30:672–679. doi: 10.1002/mrm.1910300604. [DOI] [PubMed] [Google Scholar]
- 22.Deelchand DK, Adanyeguh IM, Emir UE, et al. Two-site reproducibility of cerebellar and brainstem neurochemical profiles with short-echo, single-voxel MRS at 3T. Magn Reson Med. 2014 doi: 10.1002/mrm.25295. doi: 10.1002/mrm.25295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Randall L. Chemical topography of the brain. J Biol Chem. 1938;124:481–488. [Google Scholar]
- 24.Siegel GJ, editor. Basic neurochemistry: Molecular, cellular and medical aspects. 6th ed. Lippincott-Raven Publishers; Philadelphia: 1999. [Google Scholar]
- 25.Oz G, Alger JR, Barker PB, et al. Clinical proton MR spectroscopy in central nervous system disorders. Radiology. 2014;270:658–679. doi: 10.1148/radiol.13130531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Provencher S. [Dec. 20, 2014];LCModel and LCMgui user's manual. 2014 http://s-provencher.com/pub/LCModel/manual/manual.pdf.
- 27.Holm S. A simple sequentially rejective multiple test procedure. Scand J Statist. 1979;6:65–70. [Google Scholar]
- 28.Viau M, Boulanger Y. Characterization of ataxias with magnetic resonance imaging and spectroscopy. Parkinsonism Relat Disord. 2004;10:335–351. doi: 10.1016/j.parkreldis.2004.02.006. [DOI] [PubMed] [Google Scholar]
- 29.Reetz K, Costa AS, Mirzazade S, et al. Genotype-specific patterns of atrophy progression are more sensitive than clinical decline in SCA1, SCA3 and SCA6. Brain. 2013;136:905–917. doi: 10.1093/brain/aws369. [DOI] [PubMed] [Google Scholar]
- 30.Maddock RJ, Buonocore MH. MR spectroscopic studies of the brain in psychiatric disorders. Curr Top Behav Neurosci. 2012;11:199–251. doi: 10.1007/7854_2011_197. [DOI] [PubMed] [Google Scholar]
- 31.Brand A, Richter-Landsberg C, Leibfritz D. Multinuclear NMR studies on the energy metabolism of glial and neuronal cells. Dev Neurosci. 1993;15:289–298. doi: 10.1159/000111347. [DOI] [PubMed] [Google Scholar]
- 32.Allaman I, Bélanger M, Magistretti PJ. Astrocyte-neuron metabolic relationships: for better and for worse. Trends Neurosci. 2011;34:76–87. doi: 10.1016/j.tins.2010.12.001. [DOI] [PubMed] [Google Scholar]
- 33.Guerrini L, Belli G, Mazzoni L, et al. Impact of cerebrospinal fluid contamination on brain metabolites evaluation with 1 H-MR Spectroscopy: A single voxel study of the cerebellar vermis in patients with degenerative ataxias. J Magn Reson Imaging. 2009;30(1):11–17. doi: 10.1002/jmri.21804. [DOI] [PubMed] [Google Scholar]
- 34.Brewer GJ, Wallimann TW. Protective effect of the energy precursor creatine against toxicity of glutamate and beta-amyloid in rat hippocampal neurons. J Neurochem. 2000;74:1968–1978. doi: 10.1046/j.1471-4159.2000.0741968.x. [DOI] [PubMed] [Google Scholar]
- 35.Genius J, Geiger J, Bender A, Moller HJ, Klopstock T, Rujescu D. Creatine protects against excitoxicity in an in vitro model of neurodegeneration. PLoS One. 2012;7:e30554. doi: 10.1371/journal.pone.0030554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Meyer LE, Machado LB, Santiago APSA, et al. Mitochondrial creatine kinase activity prevents reactive oxygen species generation. Antioxidant role of mitochondrial kinase-dependent ADP re-cycling activity. J Biol Chem. 2006;281:37361–37371. doi: 10.1074/jbc.M604123200. [DOI] [PubMed] [Google Scholar]
- 37.Gatchel JR, Zoghbi HY. Diseases of unstable repeat expansion: mechanisms and common principles. Nat Rev Genet. 2005;6:743–755. doi: 10.1038/nrg1691. [DOI] [PubMed] [Google Scholar]
- 38.Mochel F, Durant B, Meng X, et al. Early alterations of brain cellular energy homeostasis in Huntington disease models. J Biol Chem. 2012;287(2):1361–1370. doi: 10.1074/jbc.M111.309849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Zacharoff L, Tkáč I, Song Q, et al. Cortical metabolites as biomarkers in the R6/2 model of Huntington's disease. J Cereb Blood Flow Metab. 2012;32:502–514. doi: 10.1038/jcbfm.2011.157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Tkáč I, Henry PG, Zacharoff L, et al. Homeostatic adaptations in brain energy metabolism in mouse models of Huntington disease. J Cereb Blood Flow Metab. 2012;32:1977–1988. doi: 10.1038/jcbfm.2012.104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Mochel F, N'Guyen TM, Deelchand D, et al. Abnormal response to cortical activation in early stages of Huntington disease. Mov Disord. 2012;27:907–910. doi: 10.1002/mds.25009. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.