Abstract
In this study, a three step purification of alkaline phosphatase from non-pasteurized milk has been described. It included cream extraction, n-butanol treatment and acetone precipitation. Different parameters such as buffer concentration, temperature, pH, substrate concentration, acetone and n-butanol treatment were optimized to maximize the enzyme activity. The enzyme was fruitfully purified up to homogeneity from the milk, with percentage recovery and fold purification of 56.17 and 17.67 respectively. The kinetic parameters were determined to be 0.927 mM (Km) and 55.86 μM/min (Vmax), with specific activity of 11.31 U/mg. Other optimized parameters were estimated as a buffer concentration of 0.5 M with pH 9.0, temperature optima at 37 °C, with n-butanol and acetone concentration of 20 % (v/v) and 50 % (v/v) respectively. This approach provides a simple and effective method for the purification of alkaline phosphatase from non-pasteurized milk.
Keywords: Alkaline Phosphatase, Purification, Enzyme, Milk, n-butanol
Introduction
Alkaline phosphatase (orthophosphoric monoester phosphohydrolase, EC 3.1.3.1) is a non-specific monoesterases that catalyze the hydrolysis of various phosphate esters and anhydrides of phosphoric acid, under alkaline conditions (Junior et al. 2008). They are the non-specific phosphatases which are apparently ubiquitous in nature. They are widely used in recombinant DNA technology (Kopetzki et al. 1994) and DNA sequencing. It is also an important component of enzyme-linked immunosorbent assay (ELISA) based kits. So its purification on a large scale is required for different commercial and research purposes. The enzymes have been isolated from different sources, which include bacteria, fungi, organs of mammals and invertebrates, but a few have been reported in plants. Commercially available alkaline phosphatases (APase) are Bacterial and calf intestinal, which are used for clinical analysis and biological research work (Atlan and Portalier 1987). Among food, milk is the natural source of enzyme, where it is present in the form of a liopoprotein complex, known as microsomes. As the enzyme get inactivated at higher temperature, it is used as a sensitive method to determine the efficacy of pasteurization (Albillos et al. 2011). Previously, APase was isolated from cow milk by Morton, but it involved different purification techniques (Morton 1953a; Morton 1953b).
In this present study, alkaline phosphatase was extracted and purified up to homogeneity from buffalo milk. A three step approach comprising cream extraction, butanol treatment and acetone precipitation was employed. This method is facile, efficient and provides a cost effective process for APase purification from non-pasteurized milk, with maximum activity.
Materials and methods
Chemicals
Tris, n-butanol, and acetone were purchased from HiMedia, India, while p-nitrophenyl phosphate disodium salt (99 % purity) was obtained from Loba Chemie. Fresh buffalo milk was purchased from a local dairy. Deionized (DI) water from Millipore was used for the reagent preparation and throughout the process.
Enzyme extraction
Purification of the enzyme, APase was carried out in three different steps which are as follows. The activity of the enzyme was assayed at each purification step.
-
Step 1:
Extraction of cream
300 ml of non-pasteurized milk was equally diluted with water and warmed at 37 °C in a water bath for 30 min. Warmed milk was then centrifuged at 12,000 rpm, for 10 min at 37 °C to separate the cream and skimmed milk. The cream thus obtained, was washed twice with DI water, churned and suspended in DI to obtain a 25 % (w/v) cream solution.
-
Step 2:
Butanol treatment
Next, the cream solution was treated with n-butanol (20 % v/v) for 50 min, with constant stirring on a magnetic stirrer at RT. The aqueous phase was then separated out by centrifuging at 12,000 rpm for 15 min at 4 °C.
-
Step 3:
Acetone precipitation
The aqueous phase obtained after the butanol treatment, was finally treated with cold acetone to a final volume of 50 % (v/v), under constant stirring and left overnight (O/N) at 4 °C. Next day, the resulting precipitate was collected by centrifugation at 12,000 rpm for 10 min, and dissolved in Tris HCl buffer (0.5 M, pH 9).
Enzyme assay
The standard assay for APase was carried out in 1 mL reaction volume, containing 400 μL of Tris HCl buffer (0.5 M), 300 μL of p-nitrophenyl phosphate (PNPP, 5 mM) as substrate, and 300 μL of enzyme. The yellow colored p-nitrophenol produced after 30 min of incubation at 37 °C was measured spectrophotometrically at 405 nm in UV visible spectrophotometer (NanoDrop 1,000, v3.8.1, Thermoscientific). The concentration of liberated p-nitrophenol was then determined by Beer-Lambert equation using an extinction coefficient of 18.5 mM−1 cm−1. One unit of the alkaline phosphatase activity was defined as the amount of enzyme which liberated 1 μmol of p-nitrophenol per min, under the given assay conditions. For each assay, three readings were taken, and the average was used as a final reading. Protein estimation was done by Lowry method (Lowry et al. 1951), using a standard curve of bovine serum albumin (BSA) protein (1 mg/ml).
Polyacrylamide gel electrophoresis (PAGE) analysis
Electrophoretic analysis of purified alkaline phosphatase was carried out in 10 % polyacrylamide gel (PAGE) under non-denaturing conditions. After the run, the protein bands were visualized by Coomassie Blue R-250 staining.
Parameters effect
Effect of buffer ionic strength
To determine the effect of buffer concentration on enzyme activity, Tris HCl buffer in the range from 0.1 M to 0.6 M were used for the enzyme assay, with 5 mM PNPP substrate and 300 μL of enzyme. The above experiment was performed in triplicates and ANOVA was used to analyze the results for their significance.
-
2.
Effect of substrate concentration
To examine the effect of substrate concentration on enzyme activity, the enzyme was incubated with different substrate concentrations in the range from 0.5 mM to 10 mM. After 30 min of incubation at 37 °C, the amount of p-nitrophenol was measured spectrophotometrically. The results were expressed in Lineweaver Burk plot in order to determine the Km and Vmax values.
-
3.
Effect of temperature and pH
The temperature dependent phosphatase activity was examined by assaying the activity of purified enzyme at 5 different temperatures (25, 30, 37, 40 and 60 °C), while the effect of pH on the activity of alkaline phosphatase was studied by altering the pH of Tris HCl buffer from 7 to 10. After 30 min of incubation, the amount of p-nitrophenol was measured spectrophotometrically at 405 nm. The above experiment was performed in triplicates and ANOVA was used to analyze the results for their significance.
-
4.
Effect of n-butanol treatment
To study the effect of n-butanol treatment on enzyme purification, its concentration was varied from 10–30 %. After regular intervals, the aqueous layer was separated by centrifugation and, assayed for activity and protein determination.
-
5.
Effect of acetone concentration
The acetone treatment was optimized in terms of concentration and incubation time, by precipitating the protein with acetone to a final concentration from 30–60 % (v/v) with different incubation period. At each concentration, one sample was incubated for 1 h, while another was left for overnight incubation, at the same temperature (4 °C). After the incubation, the precipitate was collected by centrifugation at 12,000 rpm for 15 min, and assayed.
Statistical analysis
For every parameters studied which were performed in triplicates (n = 3), the results were statistically analyzed by analysis of variance (ANOVA). A mean with standard deviation was calculated and reported using Origin Pro (Version 8).
Results and discussion
Alkaline phosphatase purification
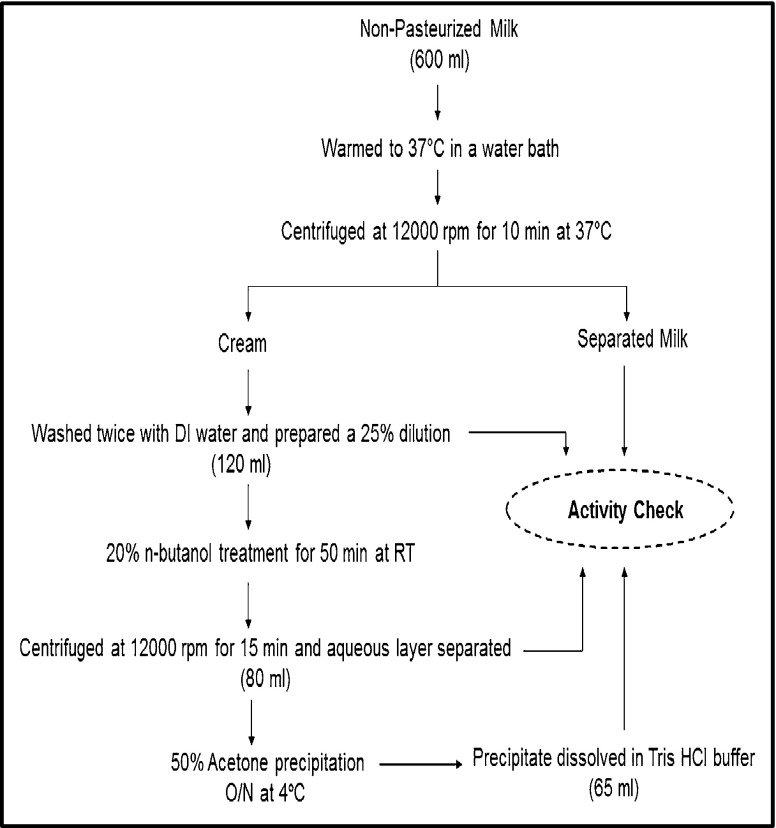
A three step approach was employed for the purification of enzyme from non-pasteurized milk (Fig. 1). In the first step, cream was extracted from the raw milk which contained most of the enzyme in the form of lipoprotein complex. It was then treated with n-butanol, which is the only solvent reported to be effective for the removal of lipid from lipoprotein complex and thus, released enzyme into the solution (Morton 1953a). This n-butanol treated solution was then centrifuged to obtain two layers, aqueous layer which contain the enzyme, and organic solvent (butanol) layer. This aqueous layer was carefully pipetted out, and the remaining solution was discarded. Next, this enzyme containing aqueous solution was treated with cold acetone at 50 % concentration (v/v) for overnight to precipitate the enzyme/protein. The precipitate was collected by centrifugation at 12,000 rpm for 15 min, and dissolved in Tris HCl buffer (0.5 M, pH 9) to obtain the purified alkaline phosphatase solution. Table 1 shows the activity comparison of the enzyme at different purification stages. Here, we obtained a 17 fold purification of enzyme after acetone precipitation with a 56 % recovery. The specific activity was determined to be 6.55 U/mg after butanol treatment and 11.31 U/mg after acetone precipitation. For separated milk (after cream removal), a very low protein was obtained (0.34 mg/ml), with specific activity of 0.71 U/mg (data not shown).
Fig. 1.
Diagrammatic representation of different steps employed for alkaline phosphatase purification from non-pasteurized milk
Table 1.
Purification summary of alkaline phosphatase from non-pasteurized milk at each step
| Purification Steps | Volume (ml) | Activity (U) | Total Activity (U) | Total Protein (mg) | Specific Activity (U/mg) | Fold Purification | Recovery (%) |
|---|---|---|---|---|---|---|---|
| Whole Milk | 600 | 7.80 ± 0.72 | 4684.68 | 7,320 | 0.64 | – | 100 |
| Separated Cream | 120 | 25.58 ± 1.84 | 3070.27 | 1,032 | 2.98 | 4.65 | 65 |
| Butanol Treatment | 80 | 34.71 ± 2.75 | 2777.18 | 424 | 6.55 | 10.23 | 59 |
| Acetone Treatment | 65 | 40.48 ± 2.24 | 2631.23 | 232.7 | 11.31 | 17.67 | 56 |
Native PAGE analysis
Figure 2 shows the native PAGE profile of purified alkaline phosphatase (Lane B) with molecular weight markers to determine the molecular mass (Lane A). Two protein bands were observed in the purified enzyme sample, with approximate molecular weight of 75 kDa and 60 kDa. These two bands may correspond to the different isozyme of APase on non-denaturing gel. However, three to five bands were also reported to be observed in the native PAGE of APase (Munson and Fall 1978). This analysis revealed that the butanol and acetone treatment removed most of the unwanted protein from the cream solution, and purified the enzyme, alkaline phosphatase, up to homogeneity.
Fig. 2.
Native PAGE profile of milk alkaline phosphatase with molecular weight markers (Lane A), and purified alkaline phosphatase (Lane B)
Parameters effects
Effect of buffer ionic strength
The choice and concentration of buffer plays an important role in enzyme activity. For alkaline phosphatase, Tris HCl has been reported as a suitable buffer for maximum activity (Hethey et al. 2002). The effect of buffer concentration on enzyme activity was carried out by assaying the enzyme in Tris HCl buffer in the concentration ranges from 0.1 M to 0.8 M. As shown in Fig. 3, as the buffer concentration increases, the enzyme activity also increases up to 0.5 M, and remains constant at 0.6 M (p < 0.05). On further increasing the ionic strength, the activity decreases, so 500 mM Tris HCl buffer was chosen as an optimum concentration for further study. Buffers with same concentration has also been reported for APase assay (Neumann et al. 1971; Dean 2002). Table 2 shows the analysis of variance for the experimental data, and the results were found to be statistically significant at α = 0.05.
-
2.
Effect of substrate concentration
Fig. 3.
Effect of Tris HCl buffer concentration on the enzyme activity. Each observation is an average of three repeats (n = 3)
Table 2.
Analysis of variance (ANOVA) for optimum parameters of purified alkaline phosphatase
| Parameters | SSa | DFb | MSc | F-Value | P-Value* | Remarks |
|---|---|---|---|---|---|---|
| Buffer Concentration | 0.1765 | 7 | 0.0252 | 16360.02 | <0.05 | Significant |
| Temperature | 0.7810 | 5 | 0.1562 | 49328.27 | <0.05 | Significant |
| pH | 1.1922 | 6 | 0.1987 | 9659.21 | <0.05 | Significant |
asum of square
bdegree of freedom
cmean square
*p-value is significant at α = 0.05
The effect of substrate concentration on the purified enzyme was studied using different concentrations of p-nitrophenyl phosphate, ranging from 0.5 mM to 10 mM. Fig. 4a indicates a classical Michaelis Menten behavior, with maxima obtained at 5 mM substrate concentration. On further increasing the substrate concentration, no significant changes were observed in the enzyme activity, which indicate that all the active sites of enzymes have been completely occupied. The Michaelis constant (Km) for p-nitrophenyl phosphate substrate was calculated from Lineweaver Burk plot (Fig. 4b). From the data, an apparent Km and Vmax values of 0.927 mM and 55.86 μM/min were obtained respectively, which is higher than the Vmax obtained from calf and rat intestinal APase, but with a similar Km for the same substrate (Chaudhuri et al. 2013; Njoku et al. 2013).
-
3.
Temperature and pH effect on APase activity
Fig. 4.
Effect of different substrate concentrations on enzyme activity (a), with Lineweaver Burk plot (b), to determine the Km and Vmax values for purified enzyme
The thermal stability of the purified APase was studied by assaying the enzyme at five different temperatures. The result showed that the enzyme activity increased with increase in temperature with maximum activity observed at 37 °C (Fig. 5a). Beyond that, a decrement in the enzyme activity was observed with 27 % and 56 % activity was lost at 50 and 60 °C respectively (p < 0.05). The complete denaturation of milk APase in 60–80 °C range has also been reported (Fadiloglu et al. 2004). The pH dependence activity was also carried out by assaying the activity at different pH. From Fig. 5b, the optimal pH for purified enzyme was found to be 9.0 (p < 0.05). Before and after that pH, a decrement in the enzyme activity was observed. Similar pH has also been reported for alkaline phosphatase assay (Thompson and Robert 1974; Mobley et al. 1984).
-
4.
Effect of n-butanol
Fig. 5.
Temperature (a) and pH profile (b) for the purified enzyme. Each observation is an average of three repeats (n = 3)
Fig. 6 represents the effect of n butanol concentration on APase extraction with respect to treatment time. For APase, a higher n-butanol concentration (30 %) results in the loss of activity, while a lower concentration (10 %) was found to be ineffective for the enzyme extraction from lipoprotein complex of cream. A 20 % butanol treatment for 50 min gives maximum yield of enzyme and thus, was considered as an optimum concentration of n-butanol. However, a higher concentration (Morton 1953a) and a lower concentration (Ahn and Snow 1993) value of butanol for APase purification from milk has also been reported earlier.
-
5.
Effect of acetone treatment
Fig. 6.
Effect of varying n-butanol concentrations for different incubation time on the enzyme activity. Each observation is an average of three repeats (n = 3)
Figure 7 shows the effect of different concentration of acetone (30–60 %) on the precipitation of protein, incubated for different time interval (1 h and O/N). It was observed that for O/N incubation, as the acetone concentration increased, more enzyme activity was observed up to 50 % (v/v) final concentration. After that, the enzyme activity declined, which may be due to enzyme denaturation at high solvent concentration. However, for 1 h incubation, the enzyme activity was observed to be directly proportional to the acetone concentration, but the activity was less as compared to O/N incubation. So O/N incubation of acetone to a final concentration of 50 % (v/v) was considered to be the optimum condition for the maximum enzyme activity.
Fig. 7.
Effect of varying acetone concentrations, and incubation time on the enzyme activity. Each observation is an average of three repeats (n = 3)
Conclusion
The present work described a facile method for the extraction and purification of the enzyme alkaline phosphatase from non-pasteurized milk. A three step approach was employed for enzyme purification, which includes cream extraction, n-butanol treatment and acetone precipitation. The purified enzyme showed 17.67 fold purification with a recovery of 56 %. The specific activity was calculated to be 11.31 U/mg with kmax and Vmax value of 0.927 mM and 55.86 μM/min, respectively. From the data, we can conclude that this strategy provides a rapid and an effective approach for the purification of alkaline phosphatase from non-pasteurized milk.
Contributor Information
Lata Sheo Bachan Upadhyay, Email: contactlataupadhyay@gmail.com.
Nishant Verma, Email: nishantbiotech1985@gmail.com.
References
- Ahn YS, Snow LD. Selective extraction of alkaline phosphatase and 5′-nucleotidase from milk fat globule membranes by a single phase n-butanol procedure. Prep Biochem. 1993;23:409–419. doi: 10.1080/10826069308544565. [DOI] [PubMed] [Google Scholar]
- Albillos SM, Reddy R, Salter R. Evaluation of alkaline phosphatase detection in dairy products using a modified rapid chemiluminescent method and official methods. J Food Prot. 2011;74:1144–1154. doi: 10.4315/0362-028X.JFP-10-422. [DOI] [PubMed] [Google Scholar]
- Atlan D, Portalier R. Purification of extracellular alkaline phosphatase released by Escherichia coli excretory mutants. Appl Microbiol Biotechnol. 1987 [Google Scholar]
- Chaudhuri G, Chatterjee S, Venu-Babu P, et al. Kinetic behaviour of calf intestinal alkaline phosphatase with pNPP. Indian J Biochem Biophys. 2013;50:64–71. [PubMed] [Google Scholar]
- Dean RL. Kinetic studies with alkaline phosphatase in the presence and absence of inhibitors and divalent cations. Biochem Mol Biol Educ. 2002;30:401–407. doi: 10.1002/bmb.2002.494030060138. [DOI] [Google Scholar]
- Fadiloglu S, Erkmen O, Sekeroglu G. Thermal and carbon dioxide inactivation of alkaline phosphatase in buffer and milk. Food Technol Biotechnol. 2004;42:27–32. [Google Scholar]
- Hethey J, Lai J, Loutet S, et al. Effects of tricine, glycine and tris buffers on alkaline phosphatase activity. J Exp Microbiol Immunol. 2002;2:33–38. [Google Scholar]
- Junior AB, GuimarAes LHS, Terenzi HF, et al. Purification and biochemical characterization of thermostable alkaline phosphatases produced by Rhizopus microsporus var. rhizopodiformis. Folia Microbiol (Praha) 2008;53:509–516. doi: 10.1007/s12223-008-0080-4. [DOI] [PubMed] [Google Scholar]
- Kopetzki E, Lehnert K, Buckel P. Enzymes in diagnostics: achievements and possibilities of recombinant DNA technology. Clin Chem. 1994;40:688–704. [PubMed] [Google Scholar]
- Lowry OH, Rosebrough NJ, Farr AL, Randall RJ. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951;193:265–275. [PubMed] [Google Scholar]
- Mobley DM, Chengappa MM, Kadel WL, Stuart JG. Effect of pH, temperature and media on acid and alkaline phosphatase activity in “clinical” and “nonclinical” isolates of Bordetella bronchiseptica. Can J Comp Med. 1984;48:175–178. [PMC free article] [PubMed] [Google Scholar]
- Morton RK. Alkaline phosphatase of milk. 1. Association of the enzyme with a particulate lipoprotein complex. Biochem J. 1953;55:786–795. doi: 10.1042/bj0550786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morton RK. Alkaline phosphatase of milk. 2. Purification of the enzyme. Biochem J. 1953;55:795–800. doi: 10.1042/bj0550795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Munson L, Fall RR. Purification and characterisation of Escherichia coli alkaline phosphatase. A biochemical experiment. Biochem Educ. 1978;6:53–56. doi: 10.1016/0307-4412(78)90059-6. [DOI] [Google Scholar]
- Neumann H, Wilson KJ, Hauck-Granoth R, Haran-Ghera N. A Comparative Biochemical Study of Alkaline Phosphatases in Normal and Leukemic Mice. Cancer Res. 1971;31:1695–1701. [PubMed] [Google Scholar]
- Njoku V, Chikezie P, Kaoje A. Kinetic studies of alkaline phosphatase extracted from rabbit (Lepus townsendii) liver. African J Biotechnol. 2013;10:3157–3162. [Google Scholar]
- Thompson LMM, Robert A. Factors affecting the activity and stability of alkaline phosphatase in a marine pseudomonad factors affecting the activity and stability of alkaline phosphatase in a marine pseudomonad. J Bacteriol. 1974;117:813–818. doi: 10.1128/jb.117.2.813-818.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]