Abstract
The deamination of adenine residues in DNA generates hypoxanthine, which is mutagenic since it gives rise to an A.T to G.C transition. Hypoxanthine is removed by hypoxanthine DNA glycosylase activity present in Escherichia coli and mammalian cells. Using polydeoxyribonucleotides or double-stranded synthetic oligonucleotides that contain dIMP residues, we show that this activity in E. coli is associated with the 3-methyladenine DNA glycosylase II coded for by the alkA gene. This conclusion is based on the following facts: (i) the two enzymatic activities have the same chromatographic behavior on various supports and they have the same molecular weight, (ii) both are induced during the adaptive response, (iii) a multicopy plasmid bearing the alkA gene overproduces both activities, (iv) homogeneous preparation of AlkA has both enzymatic activities, (v) the E. coli alkA- mutant does not show any detectable hypoxanthine DNA glycosylase activity. Under the same experimental conditions, but using different substrates, the same amount of AlkA protein liberates 1 pmol of 3-methyladenine from alkylated DNA and 1.2 fmol of hypoxanthine from dIMP-containing DNA. The Km for the latter substrate is 420 x 10(-9) M as compared to 5 x 10(-9) M for alkylated DNA. Hypoxanthine is released as a free base during the reaction. Duplex oligodeoxynucleotides containing hypoxanthine positioned opposite T, G, C, and A were cleaved efficiently. ANPG protein, APDG protein, and MAG protein--the 3-methyladenine DNA glycosylases of human, rat, and yeast origin, respectively--were also able to release hypoxanthine from various DNA substrates containing dIMP residues. The mammalian enzyme is by far the most efficient hypoxanthine DNA glycosylase of all the enzymes tested.
Full text
PDF

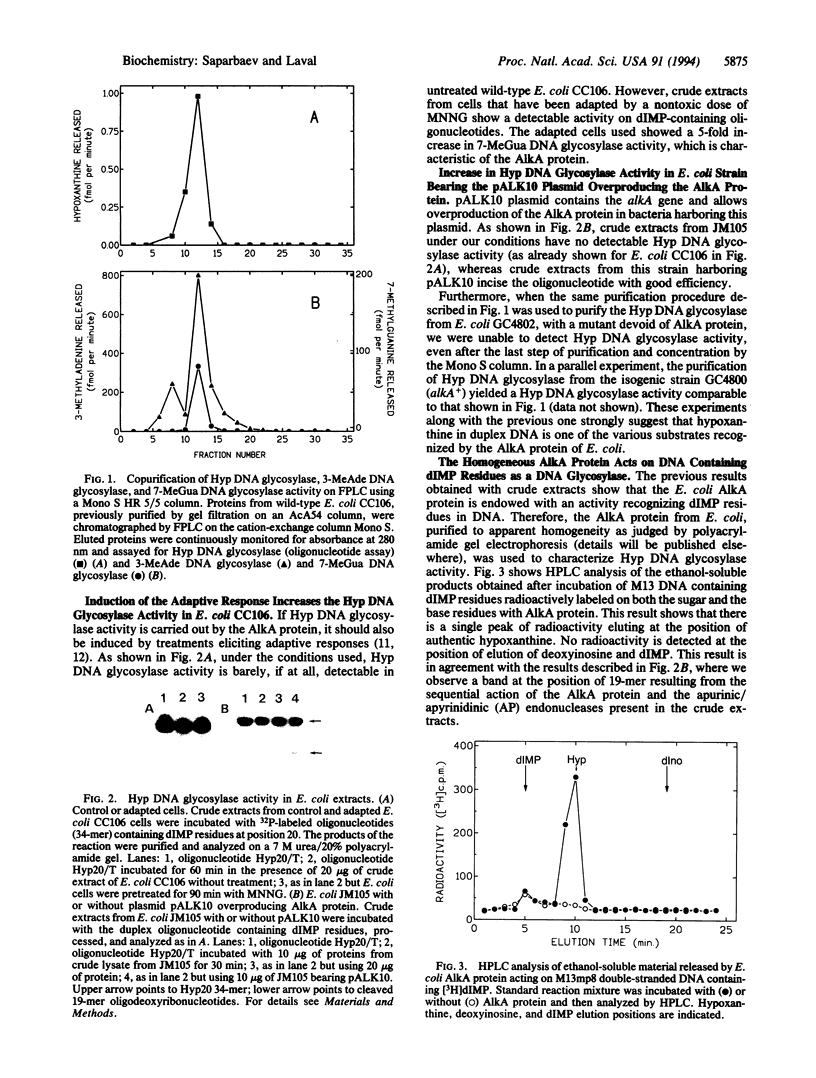
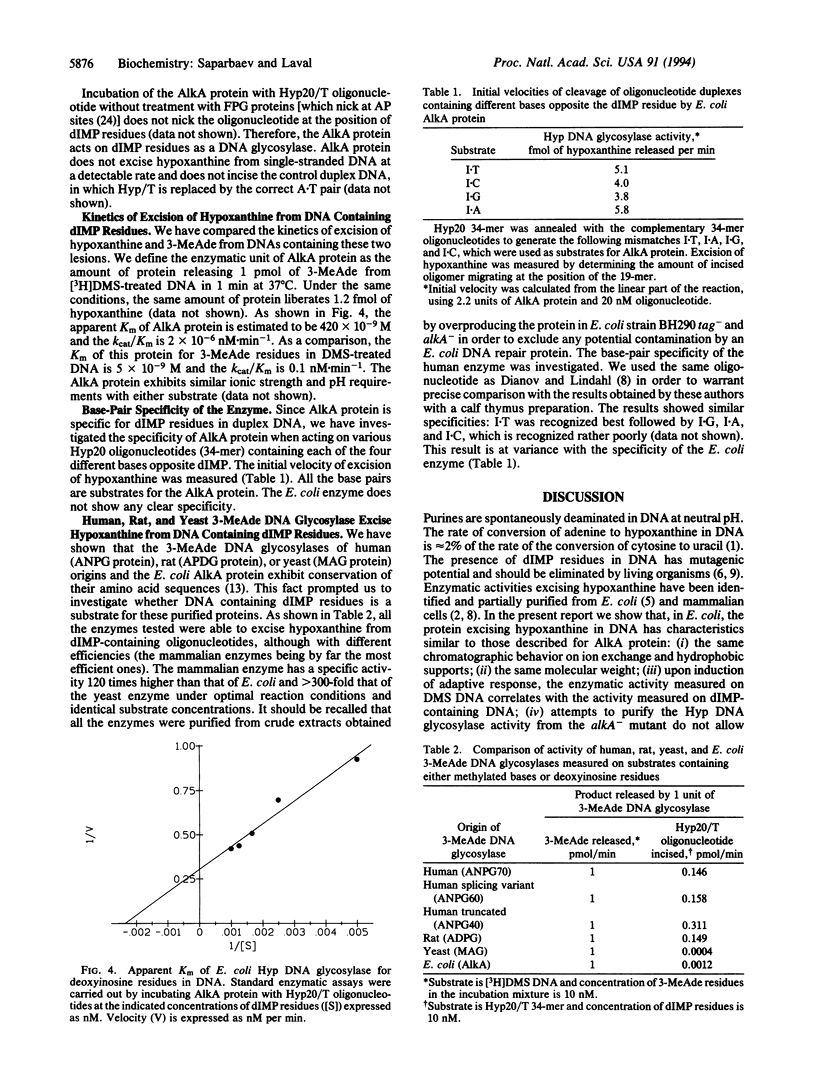

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Berdal K. G., Bjørås M., Bjelland S., Seeberg E. Cloning and expression in Escherichia coli of a gene for an alkylbase DNA glycosylase from Saccharomyces cerevisiae; a homologue to the bacterial alkA gene. EMBO J. 1990 Dec;9(13):4563–4568. doi: 10.1002/j.1460-2075.1990.tb07909.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boiteux S., Bichara M., Fuchs R. P., Laval J. Excision of the imidazole ring-opened form of N-2-aminofluorene-C(8)-guanine adduct in poly(dG-dC) by Escherichia coli formamidopyrimidine-DNA glycosylase. Carcinogenesis. 1989 Oct;10(10):1905–1909. doi: 10.1093/carcin/10.10.1905. [DOI] [PubMed] [Google Scholar]
- Boiteux S., Huisman O., Laval J. 3-Methyladenine residues in DNA induce the SOS function sfiA in Escherichia coli. EMBO J. 1984 Nov;3(11):2569–2573. doi: 10.1002/j.1460-2075.1984.tb02175.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boiteux S., O'Connor T. R., Lederer F., Gouyette A., Laval J. Homogeneous Escherichia coli FPG protein. A DNA glycosylase which excises imidazole ring-opened purines and nicks DNA at apurinic/apyrimidinic sites. J Biol Chem. 1990 Mar 5;265(7):3916–3922. [PubMed] [Google Scholar]
- Castaing B., Geiger A., Seliger H., Nehls P., Laval J., Zelwer C., Boiteux S. Cleavage and binding of a DNA fragment containing a single 8-oxoguanine by wild type and mutant FPG proteins. Nucleic Acids Res. 1993 Jun 25;21(12):2899–2905. doi: 10.1093/nar/21.12.2899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cupples C. G., Miller J. H. A set of lacZ mutations in Escherichia coli that allow rapid detection of each of the six base substitutions. Proc Natl Acad Sci U S A. 1989 Jul;86(14):5345–5349. doi: 10.1073/pnas.86.14.5345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dehazya P., Sirover M. A. Regulation of hypoxanthine DNA glycosylase in normal human and Bloom's syndrome fibroblasts. Cancer Res. 1986 Aug;46(8):3756–3761. [PubMed] [Google Scholar]
- Dianov G., Lindahl T. Preferential recognition of I.T base-pairs in the initiation of excision-repair by hypoxanthine-DNA glycosylase. Nucleic Acids Res. 1991 Jul 25;19(14):3829–3833. doi: 10.1093/nar/19.14.3829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evensen G., Seeberg E. Adaptation to alkylation resistance involves the induction of a DNA glycosylase. Nature. 1982 Apr 22;296(5859):773–775. doi: 10.1038/296773a0. [DOI] [PubMed] [Google Scholar]
- Graves R. J., Felzenszwalb I., Laval J., O'Connor T. R. Excision of 5'-terminal deoxyribose phosphate from damaged DNA is catalyzed by the Fpg protein of Escherichia coli. J Biol Chem. 1992 Jul 15;267(20):14429–14435. [PubMed] [Google Scholar]
- Habraken Y., Carter C. A., Sekiguchi M., Ludlum D. B. Release of N2,3-ethanoguanine from haloethylnitrosourea-treated DNA by Escherichia coli 3-methyladenine DNA glycosylase II. Carcinogenesis. 1991 Oct;12(10):1971–1973. doi: 10.1093/carcin/12.10.1971. [DOI] [PubMed] [Google Scholar]
- Hill-Perkins M., Jones M. D., Karran P. Site-specific mutagenesis in vivo by single methylated or deaminated purine bases. Mutat Res. 1986 Sep;162(2):153–163. doi: 10.1016/0027-5107(86)90081-3. [DOI] [PubMed] [Google Scholar]
- Kamiya H., Miura H., Kato H., Nishimura S., Ohtsuka E. Induction of mutation of a synthetic c-Ha-ras gene containing hypoxanthine. Cancer Res. 1992 Apr 1;52(7):1836–1839. [PubMed] [Google Scholar]
- Karran P., Hjelmgren T., Lindahl T. Induction of a DNA glycosylase for N-methylated purines is part of the adaptive response to alkylating agents. Nature. 1982 Apr 22;296(5859):770–773. doi: 10.1038/296770a0. [DOI] [PubMed] [Google Scholar]
- Karran P., Lindahl T. Enzymatic excision of free hypoxanthine from polydeoxynucleotides and DNA containing deoxyinosine monophosphate residues. J Biol Chem. 1978 Sep 10;253(17):5877–5879. [PubMed] [Google Scholar]
- Karran P., Lindahl T. Hypoxanthine in deoxyribonucleic acid: generation by heat-induced hydrolysis of adenine residues and release in free form by a deoxyribonucleic acid glycosylase from calf thymus. Biochemistry. 1980 Dec 23;19(26):6005–6011. doi: 10.1021/bi00567a010. [DOI] [PubMed] [Google Scholar]
- Kawase Y., Iwai S., Inoue H., Miura K., Ohtsuka E. Studies on nucleic acid interactions. I. Stabilities of mini-duplexes (dG2A4XA4G2-dC2T4YT4C2) and self-complementary d(GGGAAXYTTCCC) containing deoxyinosine and other mismatched bases. Nucleic Acids Res. 1986 Oct 10;14(19):7727–7736. doi: 10.1093/nar/14.19.7727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laval J. Two enzymes are required from strand incision in repair of alkylated DNA. Nature. 1977 Oct 27;269(5631):829–832. doi: 10.1038/269829a0. [DOI] [PubMed] [Google Scholar]
- Lindahl T. Instability and decay of the primary structure of DNA. Nature. 1993 Apr 22;362(6422):709–715. doi: 10.1038/362709a0. [DOI] [PubMed] [Google Scholar]
- Martin F. H., Castro M. M., Aboul-ela F., Tinoco I., Jr Base pairing involving deoxyinosine: implications for probe design. Nucleic Acids Res. 1985 Dec 20;13(24):8927–8938. doi: 10.1093/nar/13.24.8927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Myrnes B., Guddal P. H., Krokan H. Metabolism of dITP in HeLa cell extracts, incorporation into DNA by isolated nuclei and release of hypoxanthine from DNA by a hypoxanthine-DNA glycosylase activity. Nucleic Acids Res. 1982 Jun 25;10(12):3693–3701. doi: 10.1093/nar/10.12.3693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakabeppu Y., Miyata T., Kondo H., Iwanaga S., Sekiguchi M. Structure and expression of the alkA gene of Escherichia coli involved in adaptive response to alkylating agents. J Biol Chem. 1984 Nov 25;259(22):13730–13736. [PubMed] [Google Scholar]
- O'Connor T. R., Laval F. Isolation and structure of a cDNA expressing a mammalian 3-methyladenine-DNA glycosylase. EMBO J. 1990 Oct;9(10):3337–3342. doi: 10.1002/j.1460-2075.1990.tb07534.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Connor T. R., Laval J. Human cDNA expressing a functional DNA glycosylase excising 3-methyladenine and 7-methylguanine. Biochem Biophys Res Commun. 1991 May 15;176(3):1170–1177. doi: 10.1016/0006-291x(91)90408-y. [DOI] [PubMed] [Google Scholar]
- O'Connor T. R., Laval J. Physical association of the 2,6-diamino-4-hydroxy-5N-formamidopyrimidine-DNA glycosylase of Escherichia coli and an activity nicking DNA at apurinic/apyrimidinic sites. Proc Natl Acad Sci U S A. 1989 Jul;86(14):5222–5226. doi: 10.1073/pnas.86.14.5222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Connor T. R. Purification and characterization of human 3-methyladenine-DNA glycosylase. Nucleic Acids Res. 1993 Dec 11;21(24):5561–5569. doi: 10.1093/nar/21.24.5561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scicchitano D. A., Hanawalt P. C. Repair of N-methylpurines in specific DNA sequences in Chinese hamster ovary cells: absence of strand specificity in the dihydrofolate reductase gene. Proc Natl Acad Sci U S A. 1989 May;86(9):3050–3054. doi: 10.1073/pnas.86.9.3050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singer B., Antoccia A., Basu A. K., Dosanjh M. K., Fraenkel-Conrat H., Gallagher P. E., Kuśmierek J. T., Qiu Z. H., Rydberg B. Both purified human 1,N6-ethenoadenine-binding protein and purified human 3-methyladenine-DNA glycosylase act on 1,N6-ethenoadenine and 3-methyladenine. Proc Natl Acad Sci U S A. 1992 Oct 15;89(20):9386–9390. doi: 10.1073/pnas.89.20.9386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vickers M. A., Vyas P., Harris P. C., Simmons D. L., Higgs D. R. Structure of the human 3-methyladenine DNA glycosylase gene and localization close to the 16p telomere. Proc Natl Acad Sci U S A. 1993 Apr 15;90(8):3437–3441. doi: 10.1073/pnas.90.8.3437. [DOI] [PMC free article] [PubMed] [Google Scholar]




