Abstract
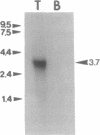
N-Arg dibasic convertase is a metalloendopeptidase from rat brain cortex and testis that cleaves peptide substrates on the N terminus of Arg residues in dibasic stretches. By using both an oligonucleotide and antibodies to screen a rat testis cDNA library, a full-length cDNA was isolated. The sequence contains an open reading frame of 1161 codons corresponding to a protein of 133 kDa that exhibits 35% and 48% similarity with Escherichia coli protease III (pitrilysin, EC 3.4.99.44) and rat or human insulinase (EC 3.4.99.45), respectively. Moreover, the presence of the HXXEH amino acid signature (XX = FL) clearly classifies N-Arg dibasic convertase as a member of the pitrilysin family of zinc-metalloendopeptidases. In addition, a Cys residue that may be responsible for the thiol sensitivity of the insulinase and N-Arg dibasic convertase was proposed. The protein sequence contains a distinctive additional feature consisting of a stretch of 71 acidic amino acids. We hypothesize that this metalloendopeptidase may be a member of a distinct class of processing enzymes.
Full text
PDF

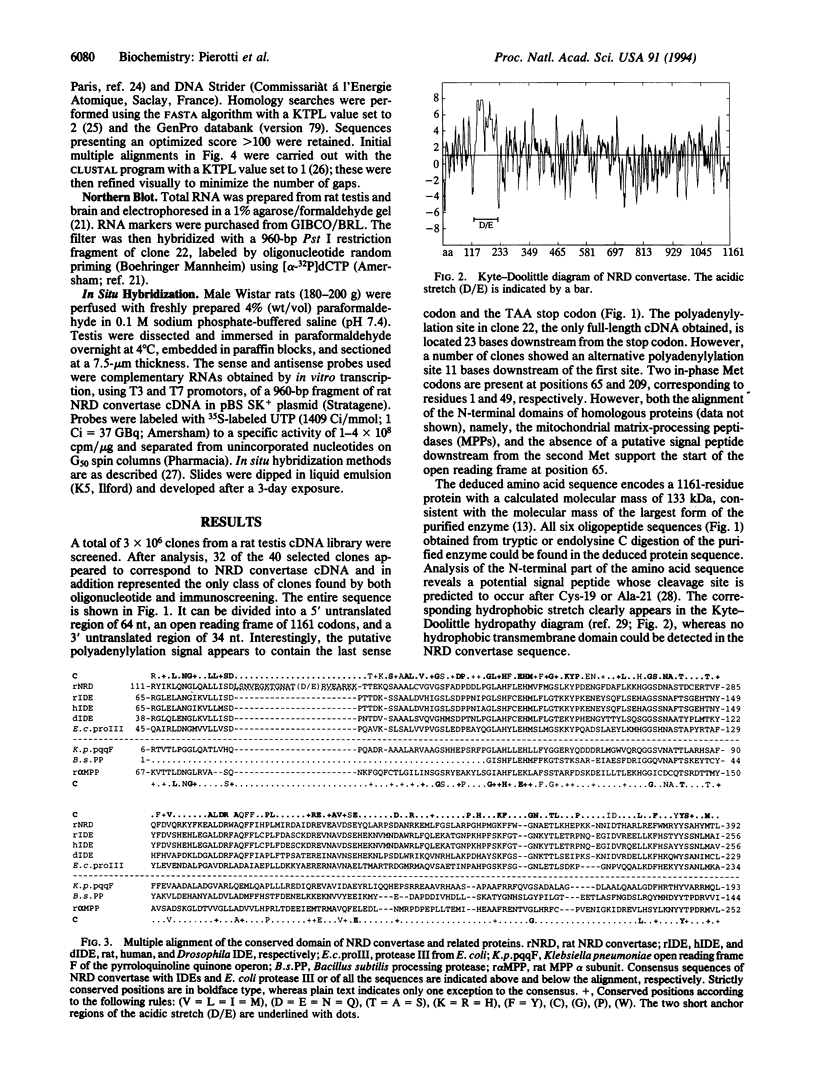


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Affholter J. A., Fried V. A., Roth R. A. Human insulin-degrading enzyme shares structural and functional homologies with E. coli protease III. Science. 1988 Dec 9;242(4884):1415–1418. doi: 10.1126/science.3059494. [DOI] [PubMed] [Google Scholar]
- Baumeister H., Müller D., Rehbein M., Richter D. The rat insulin-degrading enzyme. Molecular cloning and characterization of tissue-specific transcripts. FEBS Lett. 1993 Feb 15;317(3):250–254. doi: 10.1016/0014-5793(93)81286-9. [DOI] [PubMed] [Google Scholar]
- Bloomquist B. T., Eipper B. A., Mains R. E. Prohormone-converting enzymes: regulation and evaluation of function using antisense RNA. Mol Endocrinol. 1991 Dec;5(12):2014–2024. doi: 10.1210/mend-5-12-2014. [DOI] [PubMed] [Google Scholar]
- Chen N. Y., Jiang S. Q., Klein D. A., Paulus H. Organization and nucleotide sequence of the Bacillus subtilis diaminopimelate operon, a cluster of genes encoding the first three enzymes of diaminopimelate synthesis and dipicolinate synthase. J Biol Chem. 1993 May 5;268(13):9448–9465. [PubMed] [Google Scholar]
- Cheng Y. S., Zipser D. Purification and characterization of protease III from Escherichia coli. J Biol Chem. 1979 Jun 10;254(11):4698–4706. [PubMed] [Google Scholar]
- Chesneau V., Pierotti A. R., Barré N., Créminon C., Tougard C., Cohen P. Isolation and characterization of a dibasic selective metalloendopeptidase from rat testes that cleaves at the amino terminus of arginine residues. J Biol Chem. 1994 Jan 21;269(3):2056–2061. [PubMed] [Google Scholar]
- Darby N. J., Smyth D. G. Endopeptidases and prohormone processing. Biosci Rep. 1990 Feb;10(1):1–13. doi: 10.1007/BF01116845. [DOI] [PubMed] [Google Scholar]
- Dessen P., Fondrat C., Valencien C., Mugnier C. BISANCE: a French service for access to biomolecular sequence databases. Comput Appl Biosci. 1990 Oct;6(4):355–356. doi: 10.1093/bioinformatics/6.4.355. [DOI] [PubMed] [Google Scholar]
- Finch P. W., Wilson R. E., Brown K., Hickson I. D., Emmerson P. T. Complete nucleotide sequence of the Escherichia coli ptr gene encoding protease III. Nucleic Acids Res. 1986 Oct 10;14(19):7695–7703. doi: 10.1093/nar/14.19.7695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuller R. S., Brake A. J., Thorner J. Intracellular targeting and structural conservation of a prohormone-processing endoprotease. Science. 1989 Oct 27;246(4929):482–486. doi: 10.1126/science.2683070. [DOI] [PubMed] [Google Scholar]
- Gaudoux F., Boileau G., Crine P. Localization of neprilysin (EC 3.4.24.11) mRNA in rat brain by in situ hybridization. J Neurosci Res. 1993 Mar 1;34(4):426–433. doi: 10.1002/jnr.490340407. [DOI] [PubMed] [Google Scholar]
- Gluschankof P., Gomez S., Morel A., Cohen P. Enzymes that process somatostatin precursors. A novel endoprotease that cleaves before the arginine-lysine doublet is involved in somatostatin-28 convertase activity of rat brain cortex. J Biol Chem. 1987 Jul 15;262(20):9615–9620. [PubMed] [Google Scholar]
- Gomez S., Gluschankof P., Lepage A., Cohen P. Relationship between endo- and exopeptidases in a processing enzyme system: activation of an endoprotease by the aminopeptidase B-like activity in somatostatin-28 convertase. Proc Natl Acad Sci U S A. 1988 Aug;85(15):5468–5472. doi: 10.1073/pnas.85.15.5468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Higgins D. G., Sharp P. M. CLUSTAL: a package for performing multiple sequence alignment on a microcomputer. Gene. 1988 Dec 15;73(1):237–244. doi: 10.1016/0378-1119(88)90330-7. [DOI] [PubMed] [Google Scholar]
- Julius D., Brake A., Blair L., Kunisawa R., Thorner J. Isolation of the putative structural gene for the lysine-arginine-cleaving endopeptidase required for processing of yeast prepro-alpha-factor. Cell. 1984 Jul;37(3):1075–1089. doi: 10.1016/0092-8674(84)90442-2. [DOI] [PubMed] [Google Scholar]
- Kleiber J., Kalousek F., Swaroop M., Rosenberg L. E. The general mitochondrial matrix processing protease from rat liver: structural characterization of the catalytic subunit. Proc Natl Acad Sci U S A. 1990 Oct;87(20):7978–7982. doi: 10.1073/pnas.87.20.7978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuo W. L., Gehm B. D., Rosner M. R. Cloning and expression of the cDNA for a Drosophila insulin-degrading enzyme. Mol Endocrinol. 1990 Oct;4(10):1580–1591. doi: 10.1210/mend-4-10-1580. [DOI] [PubMed] [Google Scholar]
- Kyte J., Doolittle R. F. A simple method for displaying the hydropathic character of a protein. J Mol Biol. 1982 May 5;157(1):105–132. doi: 10.1016/0022-2836(82)90515-0. [DOI] [PubMed] [Google Scholar]
- Lepage-Lezin A., Joseph-Bravo P., Devilliers G., Benedetti L., Launay J. M., Gomez S., Cohen P. Prosomatostatin is processed in the Golgi apparatus of rat neural cells. J Biol Chem. 1991 Jan 25;266(3):1679–1688. [PubMed] [Google Scholar]
- Meulenberg J. J., Sellink E., Riegman N. H., Postma P. W. Nucleotide sequence and structure of the Klebsiella pneumoniae pqq operon. Mol Gen Genet. 1992 Mar;232(2):284–294. doi: 10.1007/BF00280008. [DOI] [PubMed] [Google Scholar]
- Mizuno K., Nakamura T., Ohshima T., Tanaka S., Matsuo H. Yeast KEX2 genes encodes an endopeptidase homologous to subtilisin-like serine proteases. Biochem Biophys Res Commun. 1988 Oct 14;156(1):246–254. doi: 10.1016/s0006-291x(88)80832-5. [DOI] [PubMed] [Google Scholar]
- Müller D., Schulze C., Baumeister H., Buck F., Richter D. Rat insulin-degrading enzyme: cleavage pattern of the natriuretic peptide hormones ANP, BNP, and CNP revealed by HPLC and mass spectrometry. Biochemistry. 1992 Nov 17;31(45):11138–11143. doi: 10.1021/bi00160a026. [DOI] [PubMed] [Google Scholar]
- Pearson W. R., Lipman D. J. Improved tools for biological sequence comparison. Proc Natl Acad Sci U S A. 1988 Apr;85(8):2444–2448. doi: 10.1073/pnas.85.8.2444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perlman R. K., Gehm B. D., Kuo W. L., Rosner M. R. Functional analysis of conserved residues in the active site of insulin-degrading enzyme. J Biol Chem. 1993 Oct 15;268(29):21538–21544. [PubMed] [Google Scholar]
- Pfanner N., Neupert W. The mitochondrial protein import apparatus. Annu Rev Biochem. 1990;59:331–353. doi: 10.1146/annurev.bi.59.070190.001555. [DOI] [PubMed] [Google Scholar]
- Pierotti A., Dong K. W., Glucksman M. J., Orlowski M., Roberts J. L. Molecular cloning and primary structure of rat testes metalloendopeptidase EC 3.4.24.15. Biochemistry. 1990 Nov 13;29(45):10323–10329. doi: 10.1021/bi00497a006. [DOI] [PubMed] [Google Scholar]
- Rawlings N. D., Barrett A. J. Evolutionary families of peptidases. Biochem J. 1993 Feb 15;290(Pt 1):205–218. doi: 10.1042/bj2900205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rawlings N. D., Barrett A. J. Homologues of insulinase, a new superfamily of metalloendopeptidases. Biochem J. 1991 Apr 15;275(Pt 2):389–391. doi: 10.1042/bj2750389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rawlings N. D., Barrett A. J. Potential metal ligands in the insulinase superfamily of endopeptidases. Biochem Soc Trans. 1991 Aug;19(3):289S–289S. doi: 10.1042/bst019289s. [DOI] [PubMed] [Google Scholar]
- Rholam M., Nicolas P., Cohen P. Precursors for peptide hormones share common secondary structures forming features at the proteolytic processing sites. FEBS Lett. 1986 Oct 20;207(1):1–6. doi: 10.1016/0014-5793(86)80002-3. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schnabel E., Mains R. E., Farquhar M. G. Proteolytic processing of pro-ACTH/endorphin begins in the Golgi complex of pituitary corticotropes and AtT-20 cells. Mol Endocrinol. 1989 Aug;3(8):1223–1235. doi: 10.1210/mend-3-8-1223. [DOI] [PubMed] [Google Scholar]
- Sossin W. S., Fisher J. M., Scheller R. H. Sorting within the regulated secretory pathway occurs in the trans-Golgi network. J Cell Biol. 1990 Jan;110(1):1–12. doi: 10.1083/jcb.110.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steiner D. F., Smeekens S. P., Ohagi S., Chan S. J. The new enzymology of precursor processing endoproteases. J Biol Chem. 1992 Nov 25;267(33):23435–23438. [PubMed] [Google Scholar]
- Stentz F. B., Kitabchi A. E., Schilling J. W., Schronk L. R., Seyer J. M. Identification of insulin intermediates and sites of cleavage of native insulin by insulin protease from human fibroblasts. J Biol Chem. 1989 Dec 5;264(34):20275–20282. [PubMed] [Google Scholar]
- Van de Ven W. J., Roebroek A. J., Van Duijnhoven H. L. Structure and function of eukaryotic proprotein processing enzymes of the subtilisin family of serine proteases. Crit Rev Oncog. 1993;4(2):115–136. [PubMed] [Google Scholar]
- von Heijne G. A new method for predicting signal sequence cleavage sites. Nucleic Acids Res. 1986 Jun 11;14(11):4683–4690. doi: 10.1093/nar/14.11.4683. [DOI] [PMC free article] [PubMed] [Google Scholar]