Abstract
We present a tool for the prediction of conserved secondary structure elements of a family of homologous non-coding RNAs. Our method does not require any prior multiple sequence alignment. Thus, it successfully applies to datasets with low primary structure similarity. The functionality is demonstrated using three example datasets: sequences of RNase P RNAs, ciliate telomerases and enterovirus messenger RNAs. CARNAC has a web server that can be accessed at the URL http://bioinfo.lifl.fr/carnac.
INTRODUCTION
The past decade has been important for the study of RNA molecules. It is now well acknowledged that non-coding RNAs play an important role in many cellular processes, including translation regulation in messenger RNA and catalytic properties. Since there is a close interplay between function and structure, software tools that predict secondary structure from the base sequence are very helpful. In this paper, we focus our attention on the analysis of secondary structure for a group of related sequences. Structure conservation in spite of sequence variation suggests that structure is functionally important. It is possible in this context to take advantage of the evolutionary information contained in the sequences to recover the conserved structure. A number of methods have recently addressed this problem (1–4), but either they are computationally demanding or they use an initial full multiple sequence alignment to build the consensus structure. In the former case, the size of the dataset is a major limitation. In the latter, the correction of the predicted pairings is closely related to the quality of the input alignment. To circumvent this problem, we devised a novel tool, named CARNAC, that relies on a stepwise efficient strategy. CARNAC does not require any prior alignment between sequences. This implies that it can successfully handle sequences with reasonable nucleotide variations. The first application of CARNAC is to the prediction of secondary structures for a group of homologous sequences. CARNAC can handle two or more sequences. The size of the supported sequences ranges from a few dozen nucleotides to two thousand nucleotides. In this context, CARNAC is able to recover >60% of the real structure, with >80% of the predicted stems being correct. CARNAC can also be used to detect the existence of a conserved secondary structure. If the sequences do not actually share a common functional structure, then CARNAC will predict no stem, or only sparse isolated stems.
METHOD
Like most other RNA prediction tools, CARNAC combines three features: energy minimization, phylogenetic comparison and sequence conservation. The structures computed by CARNAC are composed of stable stems. Isolated pairings as well as pseudo-knots are banned. In this context, the difficulty is to extract a common secondary structure from the huge number of virtual stems. For that we devised a heuristic algorithm that tackles the problem in three steps. Figure 1 gives an overview of the algorithm.
Figure 1.
Example showing the three steps of the CARNAC algorithm. First, all potential stems of all sequences are identified. Then we extract the most promising stems by pairwise comparison. With these stems, we are able to build the stem graph. Each node of the graph corresponds to a stem for a sequence. In this example, we get eight different stems (A–H), which are organized into three connected components. The first one is the best one: it contains one stem per sequence, and it is fully connected. The two remaining connected components are less reliable. In the second one, one sequence is missing (sequence 2), and in the last one, one edge is missing (connecting sequences 1 and 3).
Step 1—potential stems. The first step consists of identifying all potential stems for all sequences. We select the stems with a low free energy level. The search is performed by dynamic programming using the thermodynamic model of (5).
Step 2—pairwise foldings. We analyse all possible pairs of sequences and build a pairwise folding for each such pair. For this, given two sequences, we first select all pairs of stems that are present and equivalent in both sequences. This means that a selected pair of stems should fulfil the following constraints: being compatible with locally well-conserved regions and containing at least one compensatory mutation. Then we extract the optimal common secondary structure using a dynamic programming algorithm. A full description of this algorithm is given in (6).
Step 3—extension to n sequences. After step 2, we get n−1 predicted structures for each sequence. CARNAC combines all these stems into a single putative structure using graph-theoretical techniques. The aim is to select the most reliable stems and to eliminate the others. For this, we study the topological relationships of the stems. We build a data structure that we call the stem graph. The set of vertices is the set of all stems predicted in all sequences. For each stem A in a sequence seq1 and each stem B in a sequence seq2, there is an edge between A and B if, and only if, the pair (A,B) belongs to the pairwise folding of seq1 with seq2. We enrich the graph by adding new ‘identity edges’ between stems that are perfectly identical. In this case, we know that this pair could not be selected by CARNAC in the pairwise folding step, since it does not contain any compensatory mutation. We then investigate the structure of each connected component of the stem graph. For each connected component, we compute an index that takes into account the following topological features: (i) the number of nodes, (ii) the number of stems for each sequence, (iii) the total number of edges and (iv) the number of identity edges. The index is maximal when each sequence has exactly one stem in the component, and when the component is fully connected. The last step is the construction of a final secondary structure for each sequence. For this, we sort stems according to the confidence index of the associated connected component, and we incorporate them greedily. For a stem to be valid, we require that it does not conflict with any previously selected stem. Overlapping stems and pseudo-knots are cases of conflicts.
This stepwise strategy enables us to recover secondary structure elements that are present in only a fraction of the sequences. Moreover, it leads to a computationally efficient algorithm.
CARNAC WEB SERVER
CARNAC is written in ANSI C and all source files are freely available. CARNAC is also implemented as an online web-based tool, with a graphical user interface.
Submission form. The input of the web server is a set of unaligned RNA sequences in FASTA format. Each job is assigned a unique identifier (ID). The time required for computation of the putative foldings ranges from a few seconds for short sequences (<300 nt) to several minutes for longer sequences (up to 2000 nt). When the job is completed, the results are displayed on a new page and an alert email is sent to the user. This email contains a link to the page with the prediction results, which remains valid for 24 h. It is also possible to retrieve results directly using the ID.
Output pages. For each sequence, the predicted secondary structure is given in three formats: a Connect file (ct), which provides a textual description of the base pairings, a PostScript (ps) file and a JPEG (jpg) file, which provide graphical representations of the secondary structures. The JPEG and PostScript files are generated from the ct file using the freely distributed drawing tool Naview (7). If no structure is detected then the message ‘No structure found’ is displayed. It is also possible to display all foldings at once with RNAfamily, a home-made Java applet that is dedicated to the visualization of multiple RNA sequences. It creates a plot using linear backbone representation. This is a concise representation that makes it convenient to compare several related structures at a glance (see Figure 3). RNAfamily provides the usual graphical features such as zooming and scrolling.
Figure 3.
CARNAC results for enterovirus messenger RNAs. All stems predicted by CARNAC are located before the START codon. The representation is generated with the RNAfamily applet.
In the near future, we plan to support the RNAML standard (8), which is a universal XML exchange format for describing RNA structures, and to add other viewers for producing graphical files. It is also desirable to be able to post-process the foldings to incorporate the sequences into a multiple alignment. This alignment would extend the alignment induced by the conserved elements of the secondary structure.
RESULTS
We present three examples to provide an evaluation of the performance of CARNAC. All datasets and CARNAC results are available on the website.
RNase P RNA
The first example is composed of the Delta/Epsilon Purple Bacteria RNase P sequences available in the RNase P database (9). These RNAs present a common structure, in spite of their weak sequence conservation (60% identity on average). Sequences are ∼350 nt long. We kept only full and non-redundant sequences: Desulfovibrio desulfuricans, Desulfovibrio vulgaris, Geobacter sulfurreducens, Campylobacter jejuni and Helicobacter pylori. For the reference organism (D.desulfuricans), the real structure has around 15 stems, plus 2 pseudo-knots. Some stems are not present in the structure of the other organisms. We compare the secondary structure predicted by CARNAC with the reference structure provided by the database (Table 1). The standard thermodynamic folding programs applying to a single sequence, such as Mfold (10) and RNAfold from the Vienna Package (11), fail on this dataset. It appears that CARNAC recovers the major part of the structure (about two-thirds), with a low false positive rate. The runtime time is less than 10 s of CPU time for the whole dataset.
Table 1. CARNAC results for RNase P RNAs.
| Organism | Predicted pairings | |
|---|---|---|
| Number | Accuracy (%) | |
| D.desulfuricans | 77 | 96 |
| D.vulgaris | 70 | 97 |
| G.sulfurreducens | 85 | 92 |
| C.jejuni | 51 | 92 |
| H.pylori | 55 | 81 |
For each organism, we give the number of predicted pairings and the percentage accuracy for pairings. Most of the badly predicted pairings are local perturbations: bulge, small shift, extra pairing at the end of a stem. Each predicted structure contains 10 or 11 stems.
Ciliate telomerase RNA
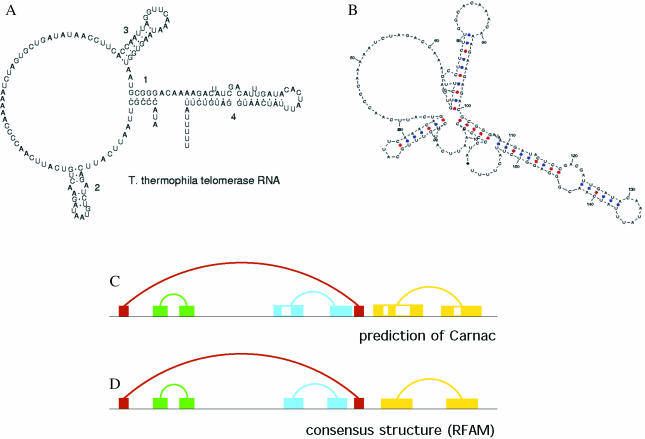
Telomerase is a ribonucleoprotein reverse transcriptase that synthesizes telomeric DNA. Sequences are available from the RFAM database (12), accession number RF00025. We selected three sequences (AF417611/283-441, U10565/50238 and AF417612/231392) with poor primary structure conservation. These sequences cannot be correctly aligned using the usual multiple alignment methods. Figure 2 shows that the structure predicted by CARNAC is complete and consistent with the model available in RFAM. For this small dataset, the computation time is less than 1 s.
Figure 2.
CARNAC prediction for ciliate telomerase RNA. (A) The reference structure proposed by RFAM and (B) the prediction of CARNAC for the sequence AF417611/283-44. We give a schematic backbone representation of the predicted structure (C), and we compare it with the consensus structure (D). Results for U10565/50238 and AF417612/231392 are identical.
Enterovirus
The program can also be used to analyse RNA sequences that do not share a common structure, or share only a partial common structure. We used CARNAC on a set of messenger RNA sequences of enterovirus, coding for a polyprotein. Each sequence is 1800 nt long and is composed of the 5′-untranslated region (5′-UTR) (∼700 nt) and the beginning of the Open reading frame (∼1100 nt). The 5′-UTR sequences are believed to share a common structure, but not the coding region (13). For this dataset, Figure 3 shows that all stems predicted by CARNAC are located in the 5′-UTR, before the START codon, and no conserved stem is found after the START codon. The putative positions of the conserved stems of the 5′-UTR are not known, so we cannot check the accuracy of the position.
This example also demonstrates that CARNAC is able to deal with long sequences (1800 nt). The user should, however, be patient: the overall computation takes >15 min. Regular users with such intensive computation needs should instead download and install CARNAC.
REFERENCES
- 1.Knudsen B. and Hein,J. (2003) Pfold: RNA secondary structure prediction using stochastic context-free grammars. Nucleic Acids Res., 31, 3423–3428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Tahi F., Gouy,M. and Regnier,M. (2002) Automatic RNA secondary structure prediction with a comparative approach. Comput. Chem., 26, 521–530. [DOI] [PubMed] [Google Scholar]
- 3.Gorodkin J., Heyer,L.J. and Stormo,G.D. (1997) Finding the most significant common sequence and structure motifs in a set of RNA sequences. Nucleic Acids Res., 25, 3724–3732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Chen J.H., Le,S.Y. and Maize,J.V. (2000) Prediction of common secondary structures of RNAs: a genetic algorithm approach. Nucleic Acids Res., 28, 991–999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jaeger J.A., Turner,D.H. and Zuker,M. (1989) Improved predictions of secondary structures for RNA. Proc. Natl Acad. Sci., USA, 86, 7706–7710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Perriquet O., Touzet,H. and Dauchet,M. (2003) Finding the common structure shared by two homologous RNAs. Bioinformatics, 19, 108–116. [DOI] [PubMed] [Google Scholar]
- 7.Bruccoleri R. and Heinrich,G. (1988) An improved algorithm for nucleic acid secondary structure display. Comput. Appl. Biosci., 4, 167–173. [DOI] [PubMed] [Google Scholar]
- 8.Waught A., Gendron,P., Altman,R., Brown,J.W., Case,D., Gautheret,D., Hartvey,S.C., Leontis,N., Westbrook,J., Westhof,E. et al. (2002) RNAML: a standard syntax for exchanging RNA information. RNA, 8, 707–717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Brown J.W. (1999) The Ribonuclease P Database. Nucleic Acids Res., 27, 314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zuker M. (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res., 31, 3406–3415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hofacker I.L. (2003) Vienna RNA secondary structure server. Nucleic Acids Res., 31, 3429–3431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Griffiths-Jones S., Bateman,A., Marshall,M., Khanna,A. and Eddy,S.R. (2003) RFAM: an RNA family database. Nucleic Acids Res., 31, 439–441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Le S.Y. and Zuker,M. (1990) Common structures of the 5′ non-coding RNA in enteroviruses and rhinoviruses. Thermodynamical stability and statistical significance. J. Mol. Biol., 216, 729–742. [DOI] [PubMed] [Google Scholar]