Abstract
Streptococcus pyogenes (group A streptococcus; GAS) is a strict human pathogen with a very high prevalence worldwide. This review highlights the genetic organization of the species and the important ecological considerations that impact its evolution. Recent advances are presented on the topics of molecular epidemiology, population biology, molecular basis for genetic change, genome structure and genetic flux, phylogenomics and closely related streptococcal species, and the long- and short-term evolution of GAS. The application of whole genome sequence data to addressing key biological questions is discussed.
Keywords: Group A streptococcus, Streptococcus pyogenes, Genomics, Population genetics, Population biology, Epidemiology, Evolution, Ecology, Streptococci
1. Taxonomy, habitats and disease
The importance of Streptococcus pyogenes as a human pathogen led to development of well-used clinical microbiology tools for its identification. Most notably, S. pyogenes forms large colonies and produces β-hemolysis following growth on blood agar, and is serologically distinguished from many other streptococcal species by its group carbohydrate that is covalently linked to the peptidoglycan cell wall. The term group A streptococci (or GAS) is commonly used as an alternative to S. pyogenes. Taxonomy based on 16S rRNA places S. pyogenes in what was historically referred to as the pyogenic (pus-forming) division of streptococci. The closest genetic relative of GAS is S. canis (group G streptococci; GGS) (Facklam, 2002). Other close genetic relatives, all of which lie within the pyogenic division and display β-hemolysis following growth on blood agar, include S. dysgalactiae subspecies equisimilis (SDE; largely group C streptococci [GCS] and GGS), S. dysgalactiae subspecies dysgalactiae (SDD), S. equi subspecies zooepidemicus (Sz) and equi (Se) (both are GCS), and S. agalactiae (group B streptococci [GBS]). Organisms that are (mostly) restricted to humans are GAS and SDE, whereas S. canis, SDD and S. equi subspecies Sz and Se primarily cause disease in other mammalian hosts. Environmental reservoirs appear to be non-existent or highly limited for the streptococcal species that cause human disease.
GAS is a human-specific pathogen that is highly prevalent worldwide, causing ~750 million infections per year (Carapetis, 2007; Carapetis et al., 2005), mostly at the throat (pharyngitis, tonsillitis) and skin (non-bullous impetigo). The epithelium of the throat and skin are the primary ecological niches of GAS and importantly, the tissue sites for most new GAS acquisitions and transmissions. Invasive disease is a relatively rare outcome of GAS infection, whereby the organism gains access to normally sterile tissue via the upper respiratory tract (URT) or breaks in the skin; from the bloodstream, GAS can disseminate to numerous deep tissues within the host. Although invasive GAS disease exacts a heavy toll in terms of morbidity and mortality, it is often an evolutionary dead end because an organism infecting deep tissue usually lacks an efficient means for transmission to a new host.
Symptomless throat infections are also known to occur, whereby the patient lacks overt clinical symptoms yet mounts a specific immune response to GAS antigens (i.e., clinically inapparent). Asymptomatic carriage of GAS at the URT - which is presumed to elicit little or no immune response due to the semi-quiescent state of the organism - can often achieve rates of >20% among school-aged populations (Kaplan, 1980). Although the organism is believed to be in a semi-quiescent, slow-growing state during carriage at the URT, transmission to new hosts can occur. Skin carriage appears to occur under endemic conditions (Anthony et al., 1976), but overall, it is less well documented. Thus, at least for GAS strains colonizing the oropharynx, a commensal-like state appears to be the predominant lifestyle.
The relative prevalence of pharyngitis and impetigo due to GAS varies in accordance with geographical location and season. Pharyngitis prevails in temperate regions and peaks in winter months, during which people spend extended time indoors and transmission occurs via a respiratory route. Impetigo is associated with warm, humid climates and is mostly observed in tropical and sub-tropical regions, or during summer months. Children are the primary targets of superficial GAS infections, and impetigo tends to afflict a slightly younger age cohort. Overall, there are spatial (geography) and temporal distances (winter versus summer seasons) that act to physically separate many organisms having a predilection for causing pharyngitis from those strains having a high tendency to cause impetigo. It is of great interest as to whether the spatial-temporal distances between GAS strains causing pharyngitis versus impetigo reduces the number of opportunities for horizontal gene transfer (HGT) which in turn, can shape the population genetic structure of a bacterial species (addressed in Section 3).
2. Molecular epidemiology
Pioneering work by Dr. Rebecca Lancefield aimed to understand the basis for protective immunity to GAS infection, and led to the development of a serological typing scheme based on the antiphagocytic M protein surface fibrils (Lancefield, 1962). More than 80 distinct M types were identified, whereby protective immunity to GAS is M type-specific. The M type-specific determinants map to the fibril tips, encoded by the 5' end of emm genes. More recently, a sequence-based emm typing scheme was implemented, based on extensive nt sequence differences at the 5' end of the emm gene, whereby a unique emm type is defined as having <92% sequence identity over the nt sequence corresponding to the first 30 codons of the mature M protein (Beall et al., 1996). Among the 234 emm types recognized to date are >1200 distinct allelic forms of the emm type-specific regions of emm genes, known as emm subtypes (Beall, 2014). Virtually all contemporary epidemiological studies define GAS isolates according to their emm type and therefore, emm type provides the primary framework for understanding the population biology and genetic structure of this species.
All GAS isolates harbor an emm gene. In addition, many GAS strains have paralogous emm-like genes lying immediately upstream and downstream of emm, and a few strains have only the downstream emm-like locus. Thus, a given GAS strain can have one or two emm-like genes, in addition to emm; the upstream emm-like gene is often referred to as mrp, and the downstream emm-like gene is often referred to as enn. The paralogous mrp and enn genes lack emm type-specific determinants, and are also distinct from the emm locus within their 3' end regions which encode a peptidoglycan-spanning domain. Based on the structure of the cell wall-spanning domain, there are four major forms or subfamilies (SF) of emm and emm-like genes, whereby emm is either SF-1 or SF-2, mrp is always SF-4, and enn is SF-1 or SF-3 (Hollingshead et al., 1994). Furthermore, there are five distinct chromosomal arrangements (Haanes et al., 1992; Hollingshead et al., 1993; Podbielski, 1993) of emm and emm-like genes and their SF forms, designated emm patterns A through E; emm patterns B and C are rare and grouped together with pattern A strains (referred to as emm pattern A–C), due to their structural similarities (i.e., all have an SF-1 emm gene and lack mrp). Pattern D and E strains have mrp and the SF-3 form of enn, but are distinct in that their emm genes are of the SF-1 and SF-2 forms, respectively. The SF-1 (58 aa residues) and SF2 (39 aa residues) forms of the peptidoglycan-spanning domain are markedly different in length, the functional significance of which has yet to be determined.
The emm pattern genotype has been determined for 170 of the 234 currently recognized emm types (McGregor et al., 2004; McMillan et al., 2013), including the most common ones. Thirty-six (21%) emm types are pattern A–C, 64 (38%) are emm pattern D, and 66 (39%) are emm pattern E; two emm types have a rearranged emm region (no pattern group is assigned), and only two (fairly rare) emm types are found in association with >1 emm pattern group. Importantly, the emm pattern group displays a statistically significant association with tissue site of infection, whereby GAS strains of the emm pattern A–C genotype show a statistically significant association with pharyngitis (throat specialists) and emm pattern D isolates show a statistically significant association with impetigo (skin specialists) (Bessen, 2009; Bessen et al., 2000b; Bessen and Lizano, 2010; Bessen et al., 1996).
For the vast majority of emm types examined (97%), multiple isolates sharing an emm type belong to the same emm pattern group (McGregor et al., 2004). Therefore, emm type is highly predictive of emm pattern grouping, and reasonable inferences can be made for emm pattern group based on knowledge of the emm type. The initial findings on the strong correlations between emm pattern grouping and preferred tissue site of infection is confirmed in a meta-analysis of 5,439 GAS isolates obtained from 23 pharyngitis and 6 impetigo population-based surveys conducted throughout the world (Bessen et al., 2011; Steer et al., 2009b), whereby emm pattern group is inferred from the emm type (Table 1). Table 1 data show that overall, pattern A–C strains represent 46.6% of pharyngitis isolates but only 8.2% of impetigo isolates. In sharp contrast, the emm pattern D strains represent 49.8% of impetigo isolates, but only 1.7% of pharyngitis isolates. Pattern A–C versus D fractional contents for each of the 29 surveys were compared by the paired t-test (2-tailed); data reveal highly significant differences for both pharyngitis (t = 7.135E-06) and impetigo (t = 8.73E-04) strain collections. In contrast, emm pattern E isolates account for almost equal fractions of throat and skin infections (51.7 and 42%, respectively); as a group, they are designated “generalists.” In conclusion, emm pattern is a genotypic marker for preferred tissue site of infection.
Table 1.
Inferred emm pattern group assignments based on emm type, for GAS recovered in 29 population- based surveys for pharyngitis or impetigo.
| Study | Country | Years | Disease | No. of isolates * |
No. pattern A–C |
No. pattern D |
No. pattern E |
% pattern A–C |
% pattern D |
% pattern E |
Simpson's D index # |
|---|---|---|---|---|---|---|---|---|---|---|---|
| (Mzoughi et al., 2004) | Tunisia | 2003 | pharyngitis | 29 | 4 | 0 | 25 | 13.8 | 0.0 | 86.2 | 0.6995 |
| (Tewodros and Kronvall, 2005) | Ethiopia | 1990 | pharyngitis | 53 | 15 | 18 | 20 | 28.3 | 34.0 | 37.7 | 0.9625 |
| (Majeed et al., 1992) | Kuwait | 1980–9 | pharyngitis | 120 | 76 | 3 | 41 | 63.3 | 2.5 | 34.2 | 0.8924 |
| (Dicuonzo et al., 2001) | Italy | 2000 | pharyngitis | 112 | 57 | 1 | 54 | 50.9 | 0.9 | 48.2 | 0.8977 |
| (Creti et al., 2004) | Italy | 1996 – 2001 | pharyngitis | 80 | 42 | 0 | 38 | 52.5 | 0.0 | 47.5 | 0.8623 |
| (Lorino et al., 2006) | Italy | 2001–2 | pharyngitis | 123 | 56 | 0 | 67 | 45.5 | 0.0 | 54.5 | 0.8847 |
| (Alberti et al., 2003) | Spain | 1996–9 | pharyngitis | 520 | 165 | 5 | 350 | 31.7 | 1.0 | 67.3 | 0.8791 |
| (Smeesters et al., 2006) | Belgium | 2004 | pharyngitis | 163 | 79 | 0 | 84 | 48.5 | 0.0 | 51.5 | 0.8918 |
| (Brandt et al., 2001) | Germany | 1997 | pharyngitis | 216 | 110 | 0 | 106 | 50.9 | 0.0 | 49.1 | 0.8843 |
| (Eisner et al., 2006) | Austria | 1996 – 2003 | pharyngitis | 69 | 27 | 0 | 42 | 39.1 | 0.0 | 60.9 | 0.8819 |
| (Dey et al., 2005) | India | 2000–1 | pharyngitis | 51 | 2 | 20 | 29 | 3.9 | 39.2 | 56.9 | 0.9639 |
| (Sagar et al., 2004) | India | 2003 | pharyngitis | 29 | 3 | 8 | 18 | 10.3 | 27.6 | 62.1 | 0.9507 |
| (Ma et al., 2002) | Japan | 2000–1 | pharyngitis | 66 | 19 | 0 | 47 | 28.8 | 0.0 | 71.2 | 0.8308 |
| (Wajima et al., 2008) | Japan | 2003–6 | pharyngitis | 302 | 140 | 0 | 162 | 46.4 | 0.0 | 53.6 | 0.8330 |
| (Rogers et al., 2007) | Australia | 2001–2 | pharyngitis | 33 | 12 | 0 | 21 | 36.4 | 0.0 | 63.6 | 0.8693 |
| (Dierksen et al., 2000) | New Zealand | 1997 | pharyngitis | 145 | 29 | 2 | 114 | 20.0 | 1.4 | 78.6 | 0.7982 |
| (Shulman et al., 2004) ^ | USA | 2000–2002 | pharyngitis | 1972 | 1045 | 6 | 921 | 53.0 | 0.3 | 46.7 | 0.8915 |
| (Richter et al., 2005) | USA | 2002–2003 | pharyngitis | 103 | 28 | 2 | 73 | 27.2 | 1.9 | 70.9 | 0.8736 |
| (Haukness et al., 2002) | USA | 1998 | pharyngitis | 61 | 36 | 0 | 25 | 59.0 | 0.0 | 41.0 | 0.9033 |
| (Kiska et al., 1997) | USA | 1993–4 | pharyngitis | 43 | 31 | 0 | 12 | 72.1 | 0.0 | 27.9 | 0.7342 |
| (Mejia et al., 1997) | Mexico | 1990 | pharyngitis | 51 | 40 | 1 | 10 | 78.4 | 2.0 | 19.6 | 0.8196 |
| (Espinosa et al., 2003) | Mexico | 1991–2000 | pharyngitis | 281 | 153 | 4 | 124 | 54.4 | 1.4 | 44.1 | 0.9076 |
| (Smeesters et al., 2006) | Brazil | 2004 | pharyngitis | 52 | 9 | 11 | 32 | 17.3 | 21.2 | 61.5 | 0.9495 |
| Total | All | pharyngitis | 4674 | 2178 | 81 | 2415 | 46.6 | 1.7 | 51.7 | 0.8722 | |
| (Tewodros and Kronvall, 2005) | Ethiopia | 1990 | impetigo | 43 | 1 | 18 | 24 | 2.3 | 41.9 | 55.8 | 0.9779 |
| (Sakota et al., 2006) | Nepal | 2000 | impetigo | 51 | 10 | 18 | 23 | 19.6 | 35.3 | 45.1 | 0.9443 |
| (Bessen et al., 2000a) ^ | Australia (tropical) | 1994–6 | impetigo | 123 | 16 | 55 | 52 | 13 | 44.7 | 42.3 | 0.9584 |
| (McDonald et al., 2007) | Australia (tropical) | 2003–5 | impetigo | 126 | 15 | 67 | 44 | 11.9 | 53.2 | 34.9 | 0.9497 |
| (Steer et al., 2009c) ^ | Fiji | 2006 | impetigo | 364 | 19 | 190 | 155 | 5.2 | 52.2 | 42.6 | 0.9570 |
| (Smeesters et al., 2006) | Brazil | 2004 | impetigo | 58 | 2 | 33 | 23 | 3.4 | 56.9 | 39.7 | 0.9685 |
| Total | All | impetigo | 765 | 63 | 381 | 321 | 8.2 | 49.8 | 42.0 | 0.9593 |
Number of GAS isolates of known or inferred emm pattern. Among the 23 pharyngitis studies, 108 GAS isolates could not be assigned an emm pattern; among the 6 impetigo studies, 3 isolates could not be assigned an emm pattern (overall, 98.0% of isolates were assigned to an emm pattern group). Of the 111 isolates with unassigned emm pattern groups, 92 (emm170, emm240) were restricted to a single study (Alberti et al., 2003).
All studies are derived from the original report by (Steer et al., 2009b), plus 3 additional studies (^). All studies meet the following criteria: (i), isolates are clearly defined as recovery from cases of pharyngitis (or tonsillitis) or impetigo; (ii), at least 25 isolates could be assigned to an emm pattern group based on emm type; and (iii), findings were published, as described (Bessen et al., 2011). Table 1 is an updated version of data reported in (Bessen et al., 2011), and incorporates recent changes to emm type assignments (Beall, 2014) and additional emm pattern determinations (McMillan et al., 2013).
Simpson's diversity index (D) and standard deviation (Grundmann et al., 2001) is based on emm types that could be assigned to an emm pattern group. For (Alberti et al., 2003), inclusion of the unassigned emm170 and emm240 isolates increases the D value slightly to 0.8972.
In four of the 23 collections of pharyngitis isolates, emm pattern D isolates outnumber emm pattern A–C isolates and comprise >20% of the GAS isolates (Table 1; bold); all four surveys were conducted in tropical or sub-tropical regions. Regardless of tissue site of isolation, data support the notion that emm pattern D and E strains tend to predominate in tropical regions whereas pattern A–C and E strains represent the vast majority of isolates in temperate regions. This regional dominance is reflected in invasive disease isolates, which are largely collected from temperate regions (Beall, 2014; Fiorentino et al., 1997; Haukness et al., 2002; Kiska et al., 1997; Rogers et al., 2007; Steer et al., 2009b). It is very important to emphasize that not all so-called "skin infections" have a similar clinical course or share the same set of host risk factors. Superficial non-bullous impetigo caused by GAS (as depicted in Table 1 data) is a self-limiting infection (i.e., can naturally resolve in ~two weeks), that is clearly distinct from wound and deep soft tissue infections or invasive disease in which GAS gain entry through breaks in the skin and requires antibiotics (and sometimes surgery) for treatment (Bisno and Stevens, 2009). The GAS strains causing impetigo (i.e., largely patterns D and E) are mostly sampled from tropical or subtropical regions (because that is where the disease is most prevalent), whereas GAS strains causing invasive skin infections (i.e., largely pattern A–C and E) are mostly sampled from temperate regions (because that is where the most resources are to support extensive epidemiologic sampling).
Another key feature revealed by Table 1 data is the relative degree of genetic diversity (D) amongst GAS derived from population-based collections of pharyngitis versus impetigo isolates. A D value equal to 1 signifies that all isolates are genetically distinct. Using emm type as the genetic marker, the mean average for D is 0.8722 and 0.9593 for the sets of pharyngitis and impetigo isolate studies, respectively. Importantly, the difference in D values for pharyngitis versus impetigo collections is highly significant (t = 2.34E-06; unpaired, 2-tailed); this finding is consistent with the observation that GAS pharyngitis tends to be dominated by a relatively small number of clones. An analysis that included all GAS, irrespective of clinical association, also shows fewer numbers of emm types accounting for a large proportion of isolates in resource-rich countries, where URT infections tend to predominate (Steer et al., 2009b). The biological basis for the difference in diversity among GAS in host communities experiencing pharyngitis versus impetigo may be tied to the mechanisms for transmission - respiratory droplets versus direct contact - with the respiratory route being highly efficient for at least a subset of strains (i.e., high basic reproductive rate, Ro) and resulting in a lower D value.
Despite differences in strain diversity among pharyngitis and impetigo isolates, within a well-circumscribed community over a narrow time frame, the number of distinct emm types circulating is quite large [for e.g., (Shulman et al., 2009; Steer et al., 2009a)]. This finding is indicative of high rates of GAS migration. The high level of GAS diversity holds true even for remote populations, such as an island community of aboriginal Australians at the Top End of the Northern Territory (Bessen et al., 2000a), where the D value for impetigo isolates over a 2-year surveillance period exceeds 0.95 (Table 1).
Although GAS strains most often recovered from cases of pharyngitis tend to belong to a more limited set of emm types, there is very high diversity among the classical throat strains (i.e., emm pattern A–C) in terms of the number of emm subtypes characterized (i.e., alleles based on the 5' end of the emm gene). The average mean number of emm subtypes identified for emm pattern A–C strains is 16.8 subtypes per emm type, compared to only 3.4 and 5.3 for emm pattern D and E strains, respectively, based on data available at (Beall, 2014). This 3- to 5-fold difference in the number of emm subtypes between throat specialists and the other two emm pattern groups may be partly due to sampling bias (i.e., skewed in favor of GAS invasive and pharyngitis isolates in resource-rich nations), or biological factors, or a combination of both.
Nucleotide sequence alignment of multiple emm subtype alleles for each emm type-specific region, collectively derived from >500 GAS isolates (N = 105 emm types), reveals an average ratio of non-synonymous substitutions per non-synonymous site (Ka) and synonymous substitutions per synonymous site (Ks) of 4.9, 1.5 and 1.3 for emm types of the emm pattern A–C, D and E groups, respectively (Bessen et al., 2008). Thus, diversifying (positive) selection acting on the emm type-specific region appears to be strongest for the classical throat strains (emm pattern A–C). Because emm types associated with GAS are largely restricted to this bacterial species, and each emm type-specific region has <92% nt sequence identity with all other emm types, the observed genetic changes giving rise to emm subtypes most likely originate as mutations arising among GAS, rather than emm gene acquisitions following HGT from another species, such as SDE which also have emm genes but they are (largely) of distinct emm types (Ahmad et al., 2009; Beall, 2014; McMillan et al., 2010).
A new cluster typing system based on GAS emm types was recently developed, using the portion of emm genes encoding the entire surface-exposed region of M proteins, for >1,000 emm genes corresponding to 175 emm types (Sanderson-Smith et al., 2014). Phylogenetic analysis revealed two main clades (X and Y), and 16 well-supported clusters accounting for 82% of emm types, with each cluster composed of multiple taxa. Of the pattern E emm types, 98% belong to clade X, whereas 92% of pattern A–C emm types fall into clade Y. Thus for these two emm pattern groups, the excluded cell-wall-spanning domain (SF-1 for pattern A–C, SF-2 for pattern E) is tightly linked to the phylogeny that is based on the surface-exposed portion of M proteins. In contrast, the pattern D emm types (SF-1) form three discrete groupings, and are present in both clades X and Y.
Collectively the M proteins bind numerous host factors such as IgA, IgG, fibrinogen and plasminogen. The M protein types assigned to an individual emm-cluster have high sequence similarity and as might be expected, share functional properties as well (Sanderson-Smith et al., 2014). Importantly, type-specific regions of M proteins belonging to the same cluster often elicit cross-protective antibodies; this may aid in the design of highly efficacious vaccines with broad coverage that are based on a limited selection of type-specific epitopes. Whereas 90% of pattern D and E emm types belong to one of the 16 emm-clusters, nearly half of pattern A–C emm types are standalone and do not cluster with any other emm type, highlighting once again, a distinct dynamic for the evolution of many pattern A–C emm types.
While emm typing has emerged as the dominant typing scheme for GAS, two other serology-based typing schemes have an important place in understanding the molecular epidemiology and genetic organization of this species. Serum opacity factor (SOF) is an LPXTG-anchored, multifunctional surface protein that also appears in a secreted form; SOF binds fibronectin and enzymatically disrupts the structure of high-density lipoproteins present in blood (Courtney and Pownall, 2010). The original SOF typing scheme was serologically based, wherein SOF type-specific serum neutralized enzymatic activity. More recently, sof sequence types have been defined based on a 450-bp region at the 5' end of the sof gene (Beall et al., 2000); the genome map position of sof lies ~16.5 kb from emm. All (or nearly all) emm pattern E strains have a sof gene and an enzymatically active SOF protein (Beall et al., 2000; Johnson et al., 2006; Kratovac et al., 2007). Several emm pattern D strains also have a sof gene; several of these (emm59, emm81, emm85) are assigned to emm-cluster E6 within clade X (Sanderson-Smith et al., 2014); interestingly, emm12 (pattern A–C) has a partial sof gene. Despite the close physical distance between sof and emm on the chromosome, several emm types are found in association with >1 sof type (and vice versa), indicative of HGT of emm or sof to new genetic backgrounds (Beall et al., 2000). Attempts to generate a phylogeny for the sof-specific determinants have been largely unsatisfactory (i.e., poorly supported trees) due to extensive intergenic recombination (Wertz et al., 2007). While there are currently >65 recognized emm types assigned to the emm pattern E group (McGregor et al., 2004; McMillan et al., 2013), the characterization of distinct sof types has not yet kept pace.
The third serological typing scheme for GAS is T-typing, based on the trypsin-resistant T-antigens that were more recently identified as surface pili which mediate adherence of GAS to host epithelium and promote biofilm formation (Manetti et al., 2007; Mora et al., 2005). The tee6 gene (Schneewind et al., 1990) maps to a genomic region containing genes known to encode microbial surface components recognizing adhesive matrix molecules (MSCRAMMs) (Hanski and Caparon, 1992; Podbielski et al., 1999), and has been designated the FCT region (FCT stands for Fibronectin- and Collagen-binding proteins and T-antigen) (Bessen and Kalia, 2002). Pilus structural components and enzymes (sortases) that mediate their assembly are encoded within the FCT region, whereby the pili correspond to T-antigens (Barnett and Scott, 2002; Lizano et al., 2007; Mora et al., 2005). Nine distinct FCT region forms are currently recognized, each containing between five and ten ORFs and differing in their content of genes encoding pilus structural components, enzymes involved in their assembly, other MSCRAMMs and transcriptional regulators (Bessen and Kalia, 2002; Falugi et al., 2008; Koller et al., 2010; Kratovac et al., 2007). The FCT region lies ~250 kb from the emm region on the genome. Despite their physical distance, there is strong linkage between FCT region form and emm pattern (Kratovac et al., 2007), suggesting that the FCT region gene products may play an adaptive role in establishing tissue tropisms.
3. Population genetics
MLST based on seven core housekeeping genes is routinely used to define clones of GAS (Enright et al., 2001). The MLST data posted at www.mlst.net currently lists 628 sequence types (ST) of GAS (i.e., S. pyogenes), based on allelic profiles at the seven loci (Aanensen, 2014). Numerous investigators from throughout the world have generously contributed to this rich data set. A population snapshot generated by the eBURST algorithm (Feil et al., 2004; Francisco et al., 2012) reveals 91 clonal complexes (CCs), in which the connected STs are single locus variants (SLVs) sharing 6 of the 7 housekeeping alleles. A high proportion of STs (40.1%) differ from all others by > 2 alleles (singletons), and the largest CC contains only 3.66% of the total STs. This places the population structure of GAS at the far end of the spectrum for bacteria, approaching that of the gut pathogen Helicobacter pylori; organisms falling within this part of the scale are characterized by high levels of genetic diversification due to both point mutation and homologous recombination (Turner et al., 2007).
Several methods have been used to assess the relative amount of recombination and mutation that contributes to genetic change in GAS based on the seven core housekeeping genes. Statistical tests were used to measure the congruence between pair wise combinations of phylogenetic trees corresponding to each of the seven housekeeping genes, based on representative strains from each deep branch (Feil et al., 2001). Of the 42 possible pair wise tree comparisons, no significant congruence between trees was observed. The lack of congruence is indicative of relatively high levels of recombination within GAS. Using the multilocus infinite alleles model and an expanded set of isolates, high rates of recombination were again predicted for GAS (Hanage et al., 2006). The value for the estimated rate of occurrence of mutation (θ) was 7.1, which is similar to that calculated for S. pneumoniae (θ = 7.4); the estimated rate of recombination (ρ) was even higher for GAS (51.2) than for pneumococcus (ρ = 29.6). S. pneumoniae is similar to GAS in its lack of congruency among the topologies of phylogenetic trees based on housekeeping gene sequences (Feil et al., 2001). Using ClonalFrame, the ratio of nt changes as the result of recombination relative to point mutation (r/m) was calculated as 17.2 for GAS (Vos and Didelot, 2009). Together, the findings on housekeeping genes support the notion that GAS exhibit high levels of recombination that at least by some measures, is roughly comparable to that observed for the pneumococcus.
In a more extensive analysis of a highly diverse set of ~600 GAS isolates collected from >25 countries, representing 156 emm types and 259 STs, 56 SLVs were detected by eBURST (Bessen et al., 2008). The genetic mechanism of descent was estimated as recombination for ≥33 of the 56 SLVs, yielding a recombination to mutation ratio of ≥1.4, which is perhaps a bit lower than expected in light of the other types of measurements mentioned above. Interestingly, when stratified according to emm pattern group, the ratio of recombination to mutational events is 0.14, 4.0 and 3.0 for emm patterns A–C, D and E, respectively. In addition, emm pattern A–C strains show the highest level of congruence for housekeeping gene tree topologies, indicative of relatively lower levels of recombination, whereas emm pattern D strains are least congruent (Kalia et al., 2002). Therefore, different methodologies point to the same general trend, whereby the throat specialists have the highest tendency to diversify by mutation, and skin specialists and generalists have a greater tendency to diversify by recombination.
Molecular evolutionary analysis of emm type-specific determinants adds further support for a large role for mutation (relative to recombination) in genetic diversification among the classical throat strains. Antibodies to the M type-specific determinants are bactericidal and act by mediating opsonophagocytosis of the GAS organism (Cunningham, 2000; Lancefield, 1962); thus, strong diversifying selection acting on emm type may arise from host immune pressures. The average mean Ka to Ks ratio for pattern A–C emm type-specific regions exceeds the values observed for pattern D and E emm types by ~3- to 5-fold (described in Section 2) (Bessen et al., 2008). If indeed the respiratory route of transmission by classical throat strains leads to a high Ro, herd immunity may build quickly whereby selection pressure favors the emergence of immune escape mutants. Alternatively, the reduced level of diversifying selection among pattern D and E emm type-specific determinants, relative to pattern A–C emm types, may be the result of functional constraints. Human complement regulator C4b-binding protein (C4BP) binds to the N-terminal hypervariable region of many M proteins and confers resistance to phagocytosis; binding of C4BP was observed for 97 and 100% of the emm pattern D and E strains tested, respectively, compared to only 38% of the pattern A–C strains (Persson et al., 2006). Although there is no clear cut binding motif for C4BP, it seems plausible that purifying (negative) selection acting on pattern D and E emm type-specific regions may preserve functional activity, yielding both fewer emm subtypes (Section 2) and lower Ka/Ks values.
For the same set of >500 GAS isolates used to calculate Ka/Ks ratios, the average mean number of HGT events per emm type was measured, whereby an emm type found in association with >1 ST differing by ≥5 of the 7 housekeeping alleles is defined as a HGT event. Data indicate an average of 0.17, 0.60 and 0.75 HGT events per emm type for the pattern A–C, D and E groups, respectively (Bessen et al., 2008). Thus, as compared to the classical throat strains, immune escape by emm pattern D and E strains may be more often driven by serotype replacement mediated via HGT and homologous recombination of the emm gene onto a new genetic background. Also, the data supports the notion that for the classical throat strains, but not skin strains or generalists, emm type is a fairly good marker for clone or clonal complex as defined by MLST (Bessen et al., 2008; Enright et al., 2001). In summary, differences among the emm pattern-defined groups of GAS in terms of their mechanisms for genetic change point to different dynamics in shaping their population structures.
As described in Section 1, GAS strains may encounter some degree of ecological separation by virtue of their tissue site preferences for infection (throat versus skin), geographic partitioning (temperate versus tropical regions) and temporal distances (winter versus summer seasons). Ecological barriers can give rise to allopatric speciation and therefore, signatures of early stages of speciation within the GAS population were sought. Numerous analyses of housekeeping alleles - including phylogenetic trees of concatenated alleles, splits graphs, fixed nt differences, distribution of shared alleles - all provide strong support for ample recombination of housekeeping genes among all three emm pattern groupings (Kalia et al., 2002). These findings extend to isolates known to be recovered from the URT versus impetigo lesions, regardless of emm pattern group (Kalia et al., 2002). A phylogenetic tree based on concatenated nt sequences of the 7 housekeeping genes used in MLST, for 114 strains representing 114 STs and 113 distinct emm types, shows only a few branches with strong bootstrap support; nt diversity (π) is very low (Bessen, 2009). Strains assigned to emm pattern groups A–C, D and E are scattered throughout the tree and there is a lack of strong clustering. A SplitsTree graph of concatenated housekeeping alleles for 97 GAS isolates representing 95 emm types also shows a lack of clustering in accordance with emm pattern group (Bessen et al., 2011). Thus despite spatialtemporal distances between many GAS strains, there is no clear evidence for diminished opportunities for HGT of core housekeeping genes between strains of different emm pattern groups. Against a background of random genetic change in core housekeeping genes, loci exhibiting a high degree of linkage with emm pattern genotypes (Section 8), and which also map to distal positions on the genome, are strong candidates for playing a direct role in the adaptations leading to throat or skin infection.
4. Molecular mechanisms underlying genetic change
The footprints of past homologous recombination events are clearly evident within core housekeeping genes of GAS. Yet, one of the major unresolved puzzles of this species concerns the molecular mechanisms governing HGT that lead to high rates of homologous recombination. GAS are rich in bacteriophage (Section 6) and theoretically, non-specific packaging of chromosomal fragments into bacteriophage capsids might provide a means for HGT of bacterial DNA (i.e., generalized transduction). GAS produce numerous DNases, and GAS DNA is presumably protected during phage-mediated transfer.
Natural transformation is another plausible mechanism for HGT among GAS, yet it has been difficult to demonstrate experimentally and when it is achieved, transformation efficiencies are relatively low. Using a mouse model, in vivo transfer of a DNA marker between two GAS strains was demonstrated and the sil locus appears to be involved; in vitro transfer of DNA could also be demonstrated, albeit at a very low frequency (Hidalgo-Grass et al., 2002). The sil locus is regulated by quorum-sensing, and the SilCR autoinducing peptide can be sensed across multiple streptococcal species (i.e., SDE) (Belotserkovsky et al., 2009). However, any involvement of sil in DNA transfer involving GAS is unlikely to be a universal mechanism because the sil locus is present in only a minority of GAS strains (Plainvert et al., 2014).
Several Streptococcus species, such as S. pneumoniae and S. mutans in particular, readily undergo transformation and the genes that confer competence for the uptake of DNA uptake are well-characterized (Martin et al., 2006). For homologs present in GAS, activation of putative late competence genes by a small-peptide pheromone (XIP) and ComR, mediated via the alternative sigma factor and master competence regulator SigX/ComX, has been successfully demonstrated (Mashburn-Warren et al., 2012; Woodbury et al., 2006). However, uptake of DNA into the GAS cell was blocked; failure to import DNA could not be explained by production of DNases (Mashburn-Warren et al., 2012). Transformation at a low efficiency is detected for GAS grown as biofilms, whereby integration of a DNA marker is increased slightly following addition of XIP and is dependent on ComR, which is also required for biofilm formation (Marks et al., 2014). Although transformation rates in this study are ~100,000-fold less than those observed for the highly competent pneumococcus, natural transformation of GAS is enhanced by biofilm formation in mammalian cell culture and in nasal-associated lymphoid tissue (NALT) of mice.
Horizontal transfer of DNA between different GAS strains seems most likely to occur during co-colonization of the URT, or co-infection of impetigo lesions with multiple strains (Carapetis et al., 1995). GAS can grow as biofilms or form microcolonies during infection within both of these tissue niches (Akiyama et al., 2003; Roberts et al., 2012). Interspecies gene exchange is also known to occur (Section 7). Quorum sensing among GAS is mediated via transcriptional regulators of the Rgg-family and peptide pheromones (Chang et al., 2011). The short hydrophobic peptide (SHP) pheromones of GAS, and the Rgg-like regulators (Rgg2, Rgg3) that detect them, mediate cross-species signaling between GAS and other member species of the pyogenic division, specifically GBS and SDE (Cook et al., 2013). These bidirectional quorum sensing systems may influence numerous biological properties, and possibly promote interspecies exchange of DNA. Other ecological considerations that may affect HGT include bacteriocin production by GAS and related species (Wescombe et al., 2009; Wescombe and Tagg, 2003), leading to competition between GAS strains or between GAS and other species, and further modulation of the microbiome.
A mutator phenotype leading to high rates of mutation has been characterized for GAS. This defect in DNA mismatch repair (MMR) is controlled by excision and re-integration of a streptococcal phage-like chromosomal island (SpyCI) at a site between the mutS and mutL loci on the genome (Scott et al., 2008), which is discussed in detail in Section 6. The presence of SpyCI in GAS leads to a ~100-fold increase in the rate of spontaneous mutation, to as high as ~10−7 to 10−8 mutations per generation.
5. Whole genome sequences
The GAS genome is remarkable for its content of prophages, SpyCIs, and other mobile genetic elements (MGEs) such as integrative and conjugative elements (ICEs). An early analysis of 11 GAS genomes (of eight emm types) yielded a pan-genome of ~2,500 genes and a core-genome of 1,297 genes (Lefebure and Stanhope, 2007); the non-core genome contains a mix of prophages, and other MGEs and accessory gene regions (AGRs). Core-genome phylogenies based on concatenated sequences of the 1,297 genes were constructed for the 11 GAS strains, and non-core gene gain and loss was enumerated at the branch point for each strain. High levels of gene gain and gene loss along each branch were observed, even for GAS isolates sharing the same emm type. This study also estimated that up to ~37% of the genes in the GAS core-genome show evidence for recombination (Lefebure and Stanhope, 2007). A few core genes under positive selection were observed for some strain-specific branches, but some branches had no core genes under positive selection.
Estimation of prophage gain and loss in GAS is problematic due to multiple copies of phage genes having high similarity, their rapid evolution, and the difficulty in distinguishing vertical inheritance from horizontal transfer. By using a synteny-based method to sidestep these problems, 12 GAS whole genome sequences represented by nine emm types were assessed for genetic flux, which was found to be dominated by gain and loss of prophage genes (Didelot et al., 2008). Estimation of the rates of genomic flux indicates, at least for some strains, that prophage integration may have accelerated in recent time. This finding is consistent with another study (five GAS genomes of four emm types) showing that for GAS species-specific genes, those that underwent recent HGT have higher Ka/Ks ratios (by ~3-fold) and a faster rate of evolution; this finding may be the result of habitat-driven adaptation (Marri et al., 2006). Yet, in another phylogenomic analysis that used the same five GAS strains but considers a slightly larger (and possibly somewhat distinct) set of GAS genes, the proportion of genes with signatures of positive (diversifying) selection is about equal (~8%) for accessory and core genes (Anisimova et al., 2007).
As of June 2014, 20 gapless and annotated whole genome sequences have been reported for GAS (NCBI, 2014) (Table 2). Also included in Table 2 are new GAS draft genomes of strains of four additional emm types; each draft genome remains in multiple contigs separated by gaps. Together, these 24 GAS strains comprise a diverse group, represented by 13 emm types, 17 STs, all three emm pattern groups, and six FCT-region forms. Genome size ranges from 1.75 to 1.93 Mb among 20 the completed genomes, and the G+C content ranges from 38.4 to 38.7%. There are >200 incomplete GAS genomes with partial sequences available (NCBI, 2014), including published draft genomes of several additional emm types (Table 3), and thousands of other GAS genomes which were primarily sequenced in a search for SNPs, indels and recent events involving MGEs [e.g., (Beres et al., 2010; Nasser et al., 2014; Shea et al., 2011); Section 9]. Computational algorithms taking a genome wide approach have identified small regulatory RNAs (sRNA) in GAS, of which ~50 have been characterized to date (Patenge et al., 2012; Perez et al., 2009; Tesorero et al., 2013).
Table 2.
Assembled and annotated genomes of GAS
| Strain | Taxon label% |
emm subtype |
emm pattern |
emm clade: cluster * |
sof gene |
FCT region |
ST | No. of prophages # |
No. of SpyCIs @ |
CRISPR- 1 locus |
CRISPR- 2 locus |
No. of ICEs ^ |
Disease + | Origin | Genome size (bp) |
Chromosomal inversion |
Accession no. | Reference |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SF370 | M1_A-C_i | 1.6 | A-C | Y: A-C3 | no | 2 | 28 | 3 | 1 | + | + | 1 | wound | USA | 1852441 | no | NC_002737 | (Ferretti et al., 2001) |
| MGAS5005 | M1_A-C_ii | 1.0 | A-C | Y: A-C3 | no | 2 | 28 | 3 | 0 | + | + | 1 | invasive (CSF) | Canada | 1838554 | no | NC_007297 | (Sumby et al., 2005) |
| M1_476 | M1_A-C_iii | 1.0 | A-C | Y: A-C3 | no | 2 | 28 | 3 | 0 | + | + | 1 | STSS | Japan | 1831128 | remnant | NC_020540.2 | (Miyoshi-Akiyama et al., 2012) |
| A20 | M1_A-C_iv | 1.0 | A-C | Y: A-C3 | no | 2 | 28 | 3 | 0 | + | + | 1 | necrotizing fasciitis |
Taiwan | 1837281 | no | NC_018936.1 | (Zheng et al., 2013) |
| MGAS10270 | M2_E | 2.0 | E | X: E4 | yes | 6 | 55 | 4 | 1 | + | + | 2 | pharyngitis | USA | 1928252 | no | NC_008022 | (Beres and Musser, 2007) |
| MGAS315 | M3_A-C_i | 3.1 | A-C | Y: A-C5 | no | 3 | 15 | 6 | 0 | + | − | 0 | STSS | USA | 1900521 | no | NC_004070 | (Beres et al., 2002) |
| SSI-1 | M3_A-C_ii | 3.1 | A-C | Y: A-C5 | no | 3 | 15 | 6 | 0 | + | − | 0 | STSS | Japan | 1894275 | yes | NC_004606 | (Nakagawa et al., 2003) |
| MGAS10750 | M4_E | 4.0 | E | X: E1 | yes | 5 | 39 | 3 | 1 | + | + | 2 | pharyngitis | USA | 1937111 | no | NC_008024 | (Beres and Musser, 2007) |
| Manfredo | M5_A-C | 5.0 | A-C | Y: single | no | 3 | 99 | 4 | 1 | − | − | 0 | ARF | USA | 1841271 | yes | NC_009332 | (Holden et al., 2007) |
| MGAS10394 | M6_A-C | 6.4 | A-C | Y: single | no | 1 | 382 | 6 | 1 | − | − | 1 | pharyngitis | USA | 1899877 | no | NC_006086 | (Banks et al., 2003b) |
| MGAS2096 | M12_A-C_i | 12.0 | A-C | Y: A-C4 | partial | 4 | 36 | 3 | 0 | + | + | 2 | AGN | Trinidad | 1860355 | no | NC_008023 | (Beres and Musser, 2007) |
| MGAS9429 | M12_A-C_ii | 12.0 | A-C | Y: A-C4 | partial | 4 | 36 | 3 | 0 | + | + | 1 | pharyngitis | USA | 1836467 | no | NC_008021 | (Beres and Musser, 2007) |
| HKU16 | M12_A-C_iii | 12.0 | A-C | Y: A-C4 | partial | 4 | 36 | 3 | 0 | + | − | 0 | scarlet fever | Hong Kong | 1908100 | yes | AFRY0000001 | (Tse et al., 2012) |
| HSC5 | M14_A-C | 14.3 | A-C | Y: single | no | 4 | 680 | 3 | 0 | − | − | 0 | not known | ND | 1818351 | no | NC_021807.1 | (Port et al., 2013) |
| MGAS8232 | M18_A-C | 18.19 | A-C | Y: single | no | 3 | 42 | 5 | 0 | − | − | 0 | ARF | USA | 1895017 | no | NC_003485 | (Smoot et al., 2002) |
| MGAS6180 | M28_E | 28.4 | E | X: E4 | yes | 4 | 52 | 3 | 1 | + | + | 3 | puerperal sepsis |
USA | 1897573 | no | NC_007296 | (Green et al., 2005) |
| NZ131 | M49_E | 49.1 | E | X: E3 | yes | 3 | 30 | 3 | 0 | + | + | 0 | AGN | New Zealand |
1815785 | no | NC_011375 | (McShan et al., 2008) |
| Alab49 | M53_D | 53.0 | D | Y: D4 | no | 3 | 11 | 3 | 1 | − | − | 1 | impetigo | USA | 1827308 | no | NC_017596 | (Bessen et al., 2011) |
| AA472 | M56_D | 56.0 | D | Y: D4 | no | 3 | 115 | ND | ND | − | − | ND | impetigo | Australia | 1733605$ | no | JRLK00000000 | This report (gaps remain) |
| MGAS15252 | M59_D_i | 59.0 | D | X: E6 | yes | 4 | 172 | 0 | 1 | + | − | 0 | SSTI | Canada | 1750832 | no | NC_017040 | (Fittipaldi et al., 2012a) |
| MGAS1882 | M59_D_ii | 59.0 | D | X: E6 | yes | 4 | 172 | 1 | 1 | + | − | 0 | AGN, pyoderma |
USA | 1781029 | no | NC_017053 | (Fittipaldi et al., 2012a) |
| AA216 | M74_D | 74.0 | D | Y: single | no | 3 | 120 | ND | ND | − | − | ND | impetigo | Australia | 1839747$ | no | JRLJ00000000 | This report (gaps remain) |
| SS1447 | M85_D | 85.0 | D | X: E6 | partial | 4 | 109 | ND | ND | + | − | ND | ARF | UK | 1915158$ | no | JRLL00000000 | This report (gaps remain) |
| MGAS2111 | M95_D | 95.0 | D | Y: single | partial | 1 | 14 | ND | ND | + | − | ND | not known | USA | 2019649$ | no | JRLI00000000 | This report (gaps remain) |
emm clade and emm cluster assignments are predictions based on emm type (Sanderson-Smith et al., 2014).
Number of complete lambdoid prophages; please note that "Φ10394.4" is considered an ICE in Table 2, although it does have prophage-like features.
Number of streptococcal phage-like chromosomal islands (SpyCI)
Number of integrative and conjugative elements (ICE)
AGN, acute glomerulonephritis; ARF, acute rheumatic fever; CSF, cerebrospinal fluid; SSTI, skin or soft tissue infection; STSS, streptococcal toxic shock syndrome
Draft genome sizes are calculated as the sum of the length of all contigs ≥500 nt. As such, values can be inaccurate due to the presence of gaps as well as potentially redundant contigs that were not merged due to poor sequence quality, especially at contig edges where sequence read coverage is lower.
ND, not determined
Table 3.
Additional draft genomes of GAS that have been published.
| Strain |
emm type or subtype |
(predicted) emm pattern |
Disease/tissue ^ | Origin | Accession no. (series) | Reference |
|---|---|---|---|---|---|---|
| M49 591 | 49 | E | skin infection | Germany | AAFV00000000 | (Beyer-Sehlmeyer et al., 2005) |
| HKU30 | 12 | A–C | scarlet fever | Hong Kong | ERS046934 | (Tse et al., 2012) |
| BJCYGAS15 | 12 | A–C | scarlet fever | Beijing | ALKD00000000 | (You et al., 2012) |
| HLJGAS12011 | 12 | A–C | scarlet fever | Heilongjiang (China) | ALKE00000000 | (You et al., 2012) |
| PS001 | 28.8 | E | puerperal sepsis | New South Wales | ERS123195 | (Ben Zakour et al., 2012) |
| PS002 | 1.40 | A–C | puerperal sepsis | New South Wales | (Ben Zakour et al., 2012) | |
| PS003 | 77.0 | E | puerperal sepsis | New South Wales | (Ben Zakour et al., 2012) | |
| PS004 | 77.0 | E | puerperal sepsis | New South Wales | (Ben Zakour et al., 2012) | |
| PS005 | 28.8 | E | puerperal sepsis | New South Wales | ERS123196 | (Ben Zakour et al., 2012) |
| PS006 | 28.8 | E | puerperal sepsis | New South Wales | ERS123197 | (Ben Zakour et al., 2012) |
| PS007 | 28.8 | E | puerperal sepsis | New South Wales | ERS123198 | (Ben Zakour et al., 2012) |
| PS008 | 28.8 | E | puerperal sepsis | New South Wales | ERS123199 | (Ben Zakour et al., 2012) |
| PS009 | 89.0 | E | puerperal sepsis | New South Wales | (Ben Zakour et al., 2012) | |
| PS010 | 75.0 | E | puerperal sepsis | New South Wales | (Ben Zakour et al., 2012) | |
| PS011 | 89.9 | E | puerperal sepsis | New South Wales | (Ben Zakour et al., 2012) | |
| 06BA18369 | 41.2 | D | SSTI | Northern Canada | APMZ00000000 | (McDonald et al., 2013) |
| SP1 | 12 | A–C | pharyngitis | Beirut | AYPA00000000 | (Tokajian et al., 2014) |
| SP2 | 108 | D | skin | Beirut | AWOZ00000000 | (Tokajian et al., 2014) |
| SP3 | 89 | E | pharyngitis | Beirut | AWPA00000000 | (Tokajian et al., 2014) |
| SP4 | 28 | E | pharyngitis | Beirut | AWPB00000000 | (Tokajian et al., 2014) |
| SP5 | 1 | A–C | skin | Beirut | AWPC00000000 | (Tokajian et al., 2014) |
| SP6 | 89 | E | pharyngitis | Beirut | AWPD00000000 | (Tokajian et al., 2014) |
| SP7 | 22 | E | pharyngitis | Beirut | AWPE00000000 | (Tokajian et al., 2014) |
| SP8 | 85 | D | pharyngitis | Beirut | AWPF00000000 | (Tokajian et al., 2014) |
| SP10 | 118 | E | pharyngitis | Beirut | AWPG00000000 | (Tokajian et al., 2014) |
| ATCC 10728 * | 24.0 | A–C | ARF | New York | AEEO00000000 | Muzny et al., unpublished |
| NS88.2 | 98.1 | D | bacteremia | Northern Territory | PRJEA84331 | Muzny et al., unpublished |
ARF, acute rheumatic fever; SSTI, skin or soft tissue infection
Also known as the C98 typing strain, derived from 22RS72
The 24 strains listed in Table 2 comprise a genetically diverse set, and many were selected for study based on their disease association. A wide range of geographic sources is also represented. Nonetheless, it is important to bear in mind that these strains may not be representative of the species as a whole and therefore, meta-analyses may be somewhat biased in unknown ways. For example, strains of pattern D emm types in clade X (Sanderson-Smith et al., 2014), which include emm59 and emm85, have several features that are more typical of pattern E strains, such as sof genes and FCT-4 regions encoding pili (Bessen, 2012; Fittipaldi et al., 2012a). Several of the specific strains listed in Tables 2 and 3 are discussed in detail in Section 9.
The number of complete prophage genomes per strain ranges from none (MGAS15252) to six (Table 2). For those strains of the same emm type that have the same number of prophages (e.g., emm1, emm3 and emm12), the nature of the prophages and their loci of integration can differ (Section 6). Also, the number of prophages differs for the two emm59 isolates. About half of the GAS genomes have a SpyCI, and about half have one or more ICEs (Table 2). The genetic structures of prophages, SpyCIs, and ICEs are discussed in detail in Section 6.
The core-genome, which largely excludes prophages, other MGEs and non-phage AGRs (the latter is discussed in Section 8), can be defined by whole genome multiple alignment [e.g., Mugsy (Angiuoli and Salzberg, 2011)], combined with identification of regions of the alignment that are shared by all genomes. These core locally collinear blocks (LCBs) of the alignment were identified and ungapped segments concatenated using Phylomark (Sahl et al., 2012). Alignment of the 24 genomes of Table 2 yields a core-genome of 1,510,896 bp, whereby 2.4% of the sites are informative. The GAS core-genomes yield a star-like phylogeny (data not shown). A minimum evolution tree (Figure 1A) shows that only organisms sharing an emm type are very closely related and monophyletic, whereas the genetic distance between isolates of any two distinct emm types is relatively high. A SplitsTree graph of networked evolution adds further support for the highly recombinogenic nature of GAS, yet even here, long branches corresponding to distinct emm types predominate (Figure 1B). One of the interesting unresolved questions of GAS phylogenomics concerns the (seeming lack of) genetic intermediate or transitional forms among organisms of different emm types. The sequencing of genomes from strains of many more of the 234 recognized emm types may help to clarify this issue. Low levels of nt sequence diversity (with little/no phylogenetic signal) and high rates of recombination add to the challenge of tracing the ancestry of strains representing each emm type.
Figure 1. Phylogeny of the core-genome based on 24 GAS isolates.
A core-genome based on the whole genome sequences of the 24 GAS isolates listed in Table 2 was defined; there are 1,510,818 positions (36,362 informative characters) in the dataset and no gaps. (A), Evolutionary history was inferred using the minimum evolution method with default parameters, conducted in MEGA6 (Tamura et al., 2013). (B), SplitsTree graph (SplitsTree v. 4.10) employs uncorrected P distance, the equal angle method for splits transformation (no weights), and the neighbor net network (Huson and Bryant, 2006); 72 splits are evident; the phi test finds statistically significant evidence for recombination.
The size of the pan-genome of GAS, based on the 24 genomes listed in Table 2, can be estimated by tallying the cumulative lengths of all LCBs (i.e., core and non-core). Using default parameters for alignments, maximum gap size allowed for LCBs, and minimum length for LCBs (LCBs < 50 nt are excluded) (Angiuoli and Salzberg, 2011), the GAS pan-genome is estimated to be 3,113,976 bp in size. ORFs were queried for their distribution among the 24 genomes using Sybil (Riley et al., 2012); ORFs can be assigned to gene clusters and these are synteny-based assignments (Angiuoli et al., 2011). Based on whole genome sequences of the 24 GAS strains (Table 2), an estimated 1,200 core clusters of orthologous proteins, together with 1,503 non-core clusters (i.e., orthologs present in 2 to 23 genomes) and 1,240 singleton proteins (i.e., genome-specific, and not included in synteny-based clusters) comprise the GAS pan-genome, for a total of ~3,943 genes/proteins. Note that estimates for the size of the GAS pan-genome vary, in part, based on the methodologies and parameters employed, and the number and nature of the strains compared [e.g., (Beres et al., 2006)].
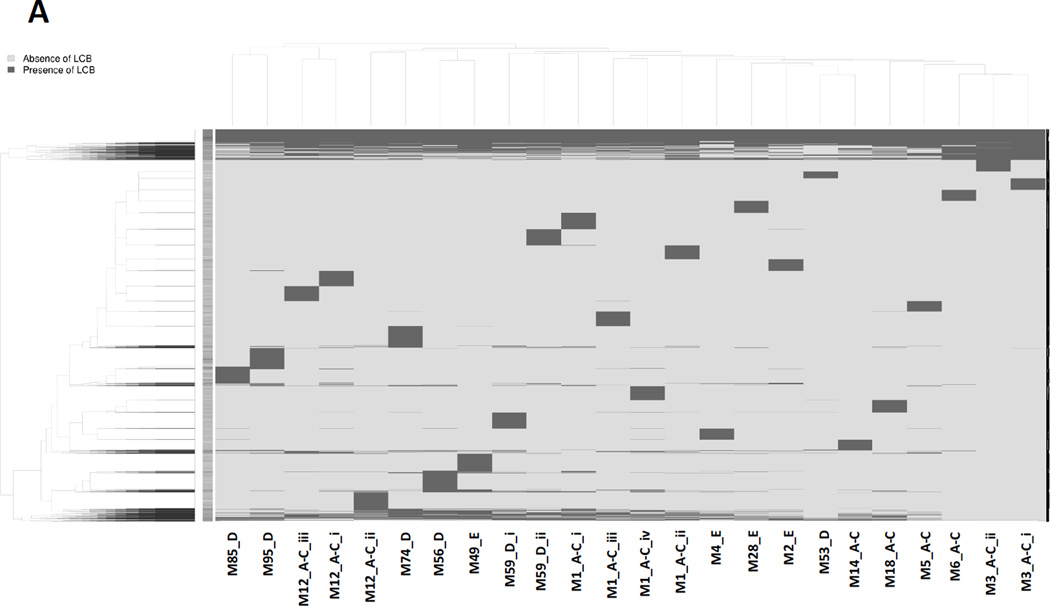
Figure 2 illustrates the distribution of whole genome alignment-derived LCBs based on the 24 GAS genomes listed in Table 2. Figure 2A includes all 5,210 individual LCBs. The alignment contains a relatively small number of large core LCBs shared by all strains (N = 159) and shared but non-core LCBs (N = 700). However, Figure 2A is dominated by the 4,351 strain-specific shorter LCBs that are approximately equally distributed across all 24 strains. The shared non-core LCBs are concentrated towards the top and bottom (tree on the left side) and are the primary drivers of strain clustering (tree on the top). Although the tree branches of the strain clusters (top tree) are fairly deep, clustering in accordance with emm type is still evident.
Figure 2. Distribution of whole genome alignment-derived locally collinear blocks (LCBs) based on 24 GAS isolates.
The 24 GAS isolates listed in Table 2 were aligned with Mugsy using default parameters. Each row of the plot depicts an LCB from the alignment; all rows are the same height and are not drawn to scale based on LCB length. Columns represent the 24 GAS strains. A dark grey line indicates that an LCB is present in a strain; light grey lines represent LCB absence. Strains (top phylogeny, with taxon labels at the bottom) and LCBs (phylogeny on left side) are clustered using the hierarchical clustering method and the associated trees are shown. The unlabeled vertical bar towards the left end of the figure represents a log heat map of LCB lengths ranging from longest (dark grey) to shortest (light grey). (A), All LCBs. (B), LCBs <1,000 nt are excluded.
In order to better visualize the distribution of shared non-core LCBs, all LCBs <1,000 nt were removed (Figure 2B). This step eliminates most, but not all strain-specific LCBs, as well as very short LCBs. The core LCBs are concentrated in the upper third of the panel. Strain clustering (tree on top) is largely driven by LCBs that are shared among a subset of strains, and many branch lengths are fairly short. For example, three emm1 strains (M1_A-C_ii to iv) uniquely share a relatively large number of LCBs depicted below the large block of core LCBs; strain M1_A-C_i (SF370; historic emm1; Section 9) also clusters with these three M1 strains but with a longer branch, reflected by the absence of some of the M1-specific LCBs in this strain. In addition to clustering in accordance with emm type, there is (partial) clustering in accordance with emm pattern group; possibly, this is related to linkage between AGRs and emm pattern group (discussed in Section 8). One striking feature is a fairly large number of strain-specific LCBs in strains M85_D and M95_D (Figure 2B); these are draft genomes and therefore, it is possible that at least some of the sequences correspond to redundant low quality contigs.
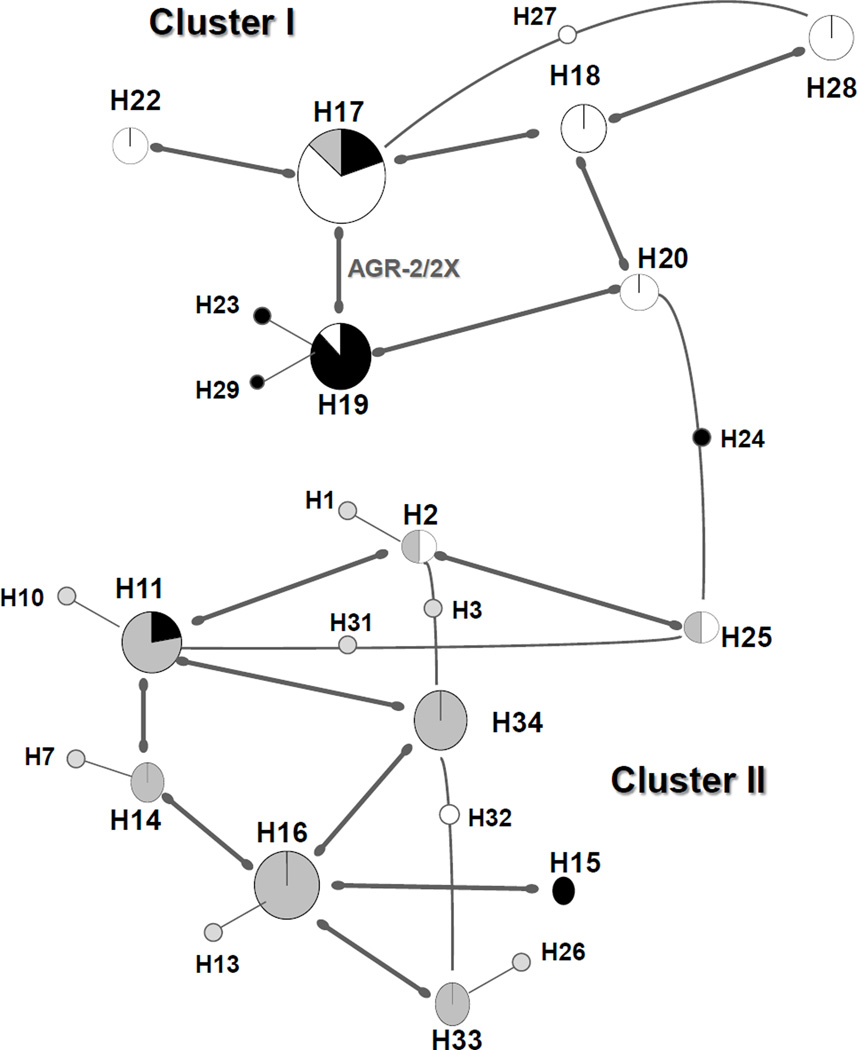
The presence or absence of genes among 97 GAS strains of 95 different emm types was assessed by comparative genome hybridization (CGH) using a GAS pan-genome microarray based on 14 whole genome sequences (Bessen et al., 2011). Of the >3,400 microarray targets, 393 were identified as having a differential distribution among 15 to 85% of the GAS strains. After excluding genes associated with prophage or SpyCIs, the differentially distributed target genes formed 22 AGRs, ranging from 1 to 43 genes per AGR. Fourteen of the 22 AGRs are represented by a single form, whereas the eight other AGRs have multiple (i.e., two or more) forms. Numerous AGRs are associated with transposons, suggesting that Tn-mediated insertions or deletions may have played a role in gene acquisition or loss. emm and sof lie within AGR-21/21X, whereas the FCT region lies within AGR-2/2X; both AGR-2 and AGR-21 have multiple forms. One of the AGRs (AGR-13) corresponds to an ICE-like region (Beres and Musser, 2007). The distribution of AGRs among GAS strains is discussed in detail in Section 8. Based on TA- and GC-skewing for 12 of the sequenced GAS genomes, it was concluded that emm lies within an ancient pathogenicity island of ~47 kb (Panchaud et al., 2009), which extensively overlaps with AGR-21/21X.
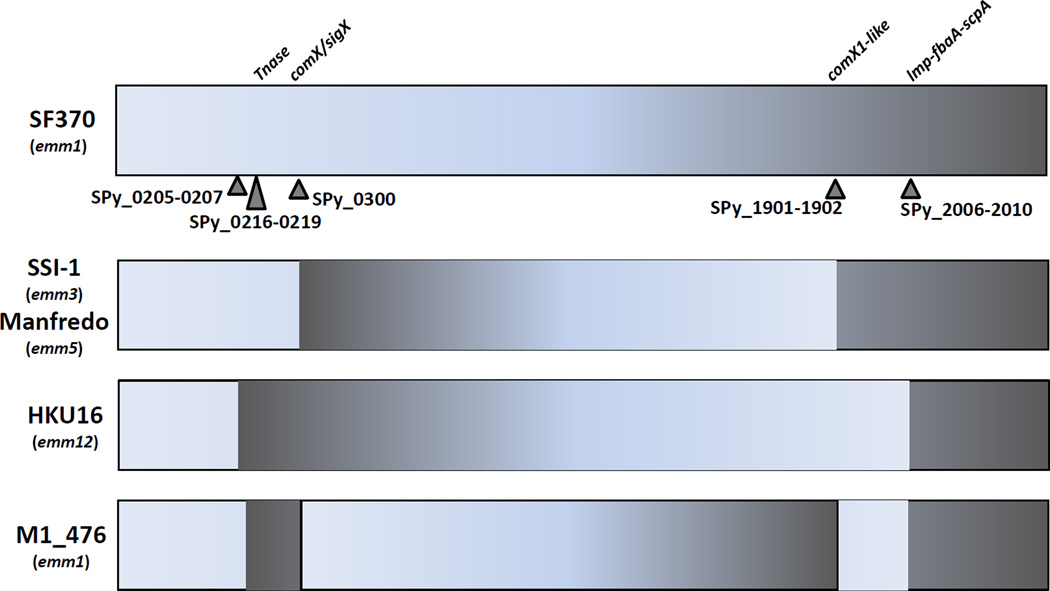
Of the 24 genomes listed in Table 2, four (16.7%) show evidence for chromosomal inversion relative to the majority of GAS genomes (Figure 3). For two of the strains - SSI-1 (emm3) and Manfredo (emm5) - a ~1.3 Mb inversion centered on the ori occurred following a recombinational event immediately downstream from the comX/sigX gene (corresponding to ORF SPy0300 in the SF370 M1 genome) and at a second comX1-like gene and glycerate kinase gene (corresponding to SPy1901–SPy1902 of the SF370 genome) (Holden et al., 2007; Nakagawa et al., 2003). Strain HKU16 (emm12) has a slightly larger chromosomal inversion (Tse et al., 2012), wherein the two crossover points lie in the vicinity of SPy0205 to SPy0207 and SPy2006 to SPy2010; the latter region encodes the virulence factors laminin-binding protein Lmp, the fibronectin-binding protein (FnBP) FbaA, and C5a peptidase (scpA) of the SF370 genome. Strain M1_476 has a remnant of a chromosomal inversion that partially reverted, and spans from a transposase corresponding to the vicinity of SF370 genome SPy0216–SPy0219 through to comX (on the right side of the ori) and SPy2006–2007 (encoding Lmp and FbaA) through to SPy1901–SPy1902 (on the left side of the ori). Thus, the comX region appears to be a hotspot for chromosomal inversion; the function of this region is discussed in Section 4. In addition, it may be noteworthy that fbaA (SPy2007) has a differential distribution among GAS strains, wherein fbaA is present in 100% of emm pattern D skin specialists but in only 25% of pattern A–C throat specialists evaluated (Bessen et al., 2011). Therefore, based on in silico analysis, this crossover hotspot (as well as the comX region) may directly impact GAS virulence or other key biological properties.
Figure 3. Chromosomal inversions.
Chromosomal crossover points are depicted for GAS genomes that have undergone an inversion, relative to the SF370 reference strain; all ORF assignments are based on the SF370 genome (Ferretti et al., 2001).
6. Mobile genetic elements
Beginning with the first sequenced GAS genome (Ferretti et al., 2001), and confirmed by each subsequent one (Table 2), it is clear that lambdoid prophages, S. pyogenes phage-like chromosomal islands (SpyCIs), integrative and conjugative elements (ICEs), and other mobile genetic elements (MGEs) are prominent features of the GAS genome. Among the completed and publicly available genomes, the number of endogenous prophages ranges from as few as zero to as many as six and together, can account for almost 10% of the total genome (Banks et al., 2004b). One GAS genome, MGAS15252, remarkably contains no intact prophages although it carries a SpyCI integrated into DNA mismatch repair gene mutL (Scott et al., 2012). The contribution of prophages and other MGEs to their host cell phenotype is evident through toxigenic conversion, antibiotic resistance, and modulation of host cell gene expression.
The earliest report that linked GAS phages to virulence showed that the “scarlatina toxin” of scarlet fever can be transmitted to a new GAS strain following exposure to a sterile filtrate from a toxigenic isolate (Frobisher and Brown, 1927). Numerous studies followed over the ensuing decades, but it was not until genome sequencing of multiple strains of GAS was complete that the true extent and diversity of endogenous prophages became more fully apparent. With few exceptions, GAS prophages follow the general structure of lambdoid phages, with some genes common to many distinct phages, whereas other genes are seemingly unique to one particular phage (Banks et al., 2002; Canchaya et al., 2002; Desiere et al., 2001). Associated with GAS prophages is a wide range of virulence-associated genes that encode exotoxins (including many superantigens) and enzymes such as DNases (i.e., streptodornases) (Table 4).
Table 4.
Prophages of GAS& and their integration sites and associated virulence genes
| Target gene | Locus code |
Integration side |
Associated phage# |
emm type of host cell |
Associated virulence genes^ |
|
|---|---|---|---|---|---|---|
| ssDNA binding protein recO | A | 3’ | 10394.1 | 6 | sdn | |
| RNA helicase snf | B | 3’ | 8232.1 | 18 | speA1 | |
| Promoter of hypothetical Spy49_0371 | C | 5’ | NZ131.1 | 49 | none identified | |
| Dipeptidase | D | 5’ | SF370.1 | 1 | speC-spd1 | |
| Man.4 | 5 | speC-spd1 | ||||
| 2096.1 | 12 | speC-spd1 | ||||
| 9429.1 | 12 | speC-spd1 | ||||
| 8232.2 | 18 | speC-spd1 | ||||
| 10270.1 | 2 | speC-spd1 | ||||
| 10750.1 | 4 | speC-spd1 | ||||
| tRNAarg | E | 3’ | 10394.2 | 6 | speA4 | |
| dTDP-glucose-4,6-dehydratase | F | 3’ | SF370.2 | 1 | speH-spel | |
| Man.3 | 5 | speH-spel | ||||
| 9429.2 | 12 | speH-spel | ||||
| HKU16.3 | 12 | speH-spel | ||||
| 10270.2 | 2 | spd3 | ||||
| 10750.2 | 4 | spd3 | ||||
| NZ131.2 | 49 | speH | ||||
| CRISPR type II system direct repeat sequence | G | 5’ | 315.1 | 3 | none identified | |
| SPsP6* | 3 | none identified | ||||
| tmRNA | H | 3’ | 5005.1 | 1 | speA2 | |
| A20.1 | 1 | speA2 | ||||
| M1_476.1 | 1 | speA2 | ||||
| 315.2 | 3 | ssa | ||||
| SPsP5 | 3 | ssa | ||||
| 10394.3 | 6 | speK-slaA | ||||
| HSC5.1 | 14 | speL-speM | ||||
| 8232.3 | 18 | speL-speM | ||||
| Alab49.1 | 53 | speL-speM | ||||
| 6180.1 | 28 | speC-spd1 | ||||
| comEC | I | 5’ | 10394.4 (ICE) | 6 | mefA | |
| DNA-binding protein HU | J | 5’ | 370.3 | 1 | spd3 | |
| 5005.2 | 1 | spd3 | ||||
| A20.2 | 1 | spd3 | ||||
| M1_476.2 | 1 | spd3 | ||||
| 315.3 | 3 | spd4 | ||||
| SPsP4 | 3 | ssa | ||||
| Man.2 | 5 | spd4 | ||||
| 10394.5 | 6 | speC-spd1 | ||||
| HSC5.2 | 14 | spd3 | ||||
| Alab49.2 | 53 | speC-spd1 | ||||
| 1882.1 | 59 | speK-slaA | ||||
| 10750.3 | 4 | ssa | ||||
| yesN promoter | K | 5’ | 315.4 | 3 | speK-slaA | |
| SPsP3 | 3 | speK-slaA | ||||
| 10270.3 | 2 | speK-slaA | ||||
| 6180.2 | 28 | speK-slaA | ||||
| recX | L | 5’ | 315.5 | 3 | speA3 | |
| SPsP2 | 3 | speA3 | ||||
| Man.1 | 5 | spd1 | ||||
| 10394.6 | 6 | sda | ||||
| Alab49.3 | 53 | spd3 | ||||
| putative gamma-glutamyl kinase | M | 5’ | 315.6 | 3 | sdn | |
| SPsP1 | 3 | sdn | ||||
| tRNAser | N | 3’ | 5005.3 | 1 | sda | |
| A20.3 | 1 | sdaD2 | ||||
| M1 476.3 | 1 | sdaD2 | ||||
| HKU16.2 | 12 | sdaD2 | ||||
| 2096.2 | 12 | sda | ||||
| 9429.3 | 12 | sda | ||||
| HAD-like hydrolase | O | 5’ | 8232.5 | 18 | sda | |
| Excinuclease subunit uvrA | P | 5’ | 10394.7 | 6 | spd3 | |
| HKU16.1 | 12 | speC | ||||
| HSC5.3 | 14 | spd3 | ||||
| Conserved hypothetical protein Spy49_1532 | Q | 5’ | NZ131.3 | 49 | spd3 | |
| SSU ribosomal protein S4P | R | 3’ | 10270.5 | 2 | none identified | |
| 6180.4 | 28 | none identified | ||||
Analysis includes the 20 completed and assembled GAS genomes listed in Table 2
SPsP1 through SPsP6 are the prophages in strain SSI-1
Gene abbreviations: speA, speC, speH, speI, speK, speL, speM and ssa and their alleles encode superantigens; sda, spn and spd alleles encode DNases; sla alleles encode phospholipase; mefA encodes a macrolide efflux pump.
The phage numbering scheme for many strains follows the assignments as defined in (Beres and Musser, 2007). Skips in numerical sequence reflect those elements that are presently defined as SpyCIs.
Botstein proposed that the product of phage evolution is a family of interchangeable genetic modules, each associated with a particular biological function (Botstein, 1980). This modular view is embodied by GAS prophages. For example, a given integrase gene and its targeted integration site may be associated with several different virulence genes (Table 4). In contrast, some modules are very strongly linked, such as the SF370.1-like integrase and the virulence genes speC-spd1 (Table 4; locus code D). Some prophages may be associated with a single emm type, as may be the case for ΦNZ131.3 and emm49 strains (McShan et al., 2008); host range restriction, which may or may not apply here, could potentially limit recombination opportunities between phages inhabiting GAS of other emm types. Highly conserved sequences within GAS prophage genomes, such as the paratox sequence (Aziz et al., 2005) or the hyaluronidase gene family, may help promote recombination between similar modules in other phages. In addition, the gene pool for GAS phages extends to other streptococcal species. Phages sharing highly similar genes or larger genetic blocks with GAS have been identified in S. equi subsp., S. agalactiae, S. equisimilis subsp., and Streptococcus thermophilus (Bai et al., 2013; Canchaya et al., 2002; Davies et al., 2007b; Desiere et al., 2001; Holden et al., 2009).
Although Escherichia coli phage lambda is the paradigm for site-specific integration (Hendrix et al., 1983), its use of an intergenic attachment site on the bacterial chromosome is somewhat atypical. An extensive analysis of bacterial genome prophages showed that tRNA genes, the tmRNA gene, and intragenic targets account for nearly 70% of prophage attachment sites (Fouts, 2006). Attachment sites within coding regions can be further subdivided according to whether the phage integrates into the 5’ or 3’ end of the gene (Canchaya et al., 2002). For tRNA genes, the sequence duplication between the host gene and phage attachment site ensures that gene function is preserved following integration at the 3’ end (Fouts, 2006). By contrast, integration into the 5’ end of a gene has the potential to disrupt gene function, as observed for the SpyCIs that integrate into the 5’ end of DNA mismatch repair gene mutL (Scott et al., 2012; Scott et al., 2008).
Numerous GAS genes are targeted by phages for site-specific integration, with integration occurring at either the 5’ and 3’ ends (Table 4). Some sites are frequently exploited by phages (e.g., the genes for tmRNA and DNA-binding protein hup; locus codes H and J, Table 4), whereas other integration sites are presently known by single representatives. The prophages that integrate into the promoters at the 5' ends of genes Spy49_0371 and yesN may interrupt the expression of these genes by separating the ORF from its transcriptional start site. GAS prophages integrate directly into the 5’ end of structural genes encoding a conserved dipeptidase, DNA translocation machinery channel protein ComEC, DNA-binding protein HU, RecX, and a gamma-glutamyl kinase (Table 4). Presently, the impact of these site-specific recombination events on the host cell phenotype is mostly unknown. The emm3 prophages Φ315.1 and ΦSPsP5 target a highly unusual chromosomal attachment site via integration into a direct repeat sequence for the CRISPR type II system (i.e., CRISPR-1 locus; Table 2). None of the spacers or direct repeats typically associated with this CRISPR in other GAS strains are evident in the emm3 isolates, and it is possible that the site-specific integration event led to inactivation of the CRISPR anti-phage system (Nozawa et al., 2011).
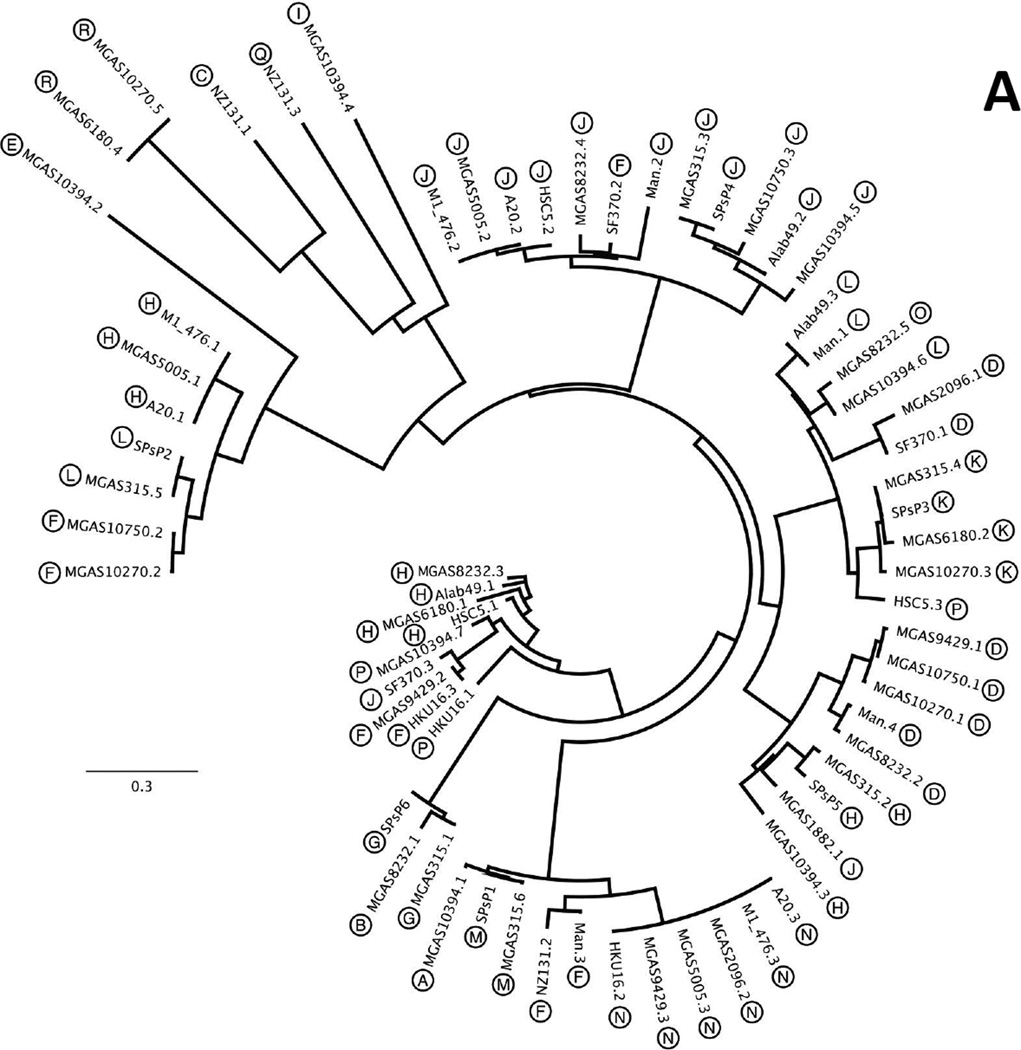
Phylogenetic analysis shows that the genome prophages are highly diverse, despite the frequent sharing of lysogeny modules, virulence genes, or both (Figure 4A). Some lysogeny modules are strongly linked to particular virulence genes, such as the integrases that target the dipeptidase gene paired with speC-spd1, or integrases targeting tRNAser paired with DNase genes (Table 4). Other modules are associated with multiple virulence genes, such as integrases targeting the tmRNA gene (locus code H, Table 4). However, these apparent linkages do not necessarily result from clonality: the prophages integrated into the tRNAser (locus code N) form a closely related branch of the tree, whereas those targeting the dipeptidase gene (locus code D) are somewhat diverse (Figure 4A). As mentioned, prophage integration at the dipeptidase gene has the potential alter its expression, and it may be that dipeptidase inhibition enhances the functional impact of the prophage-encoded speC-spd1 virulence cluster.
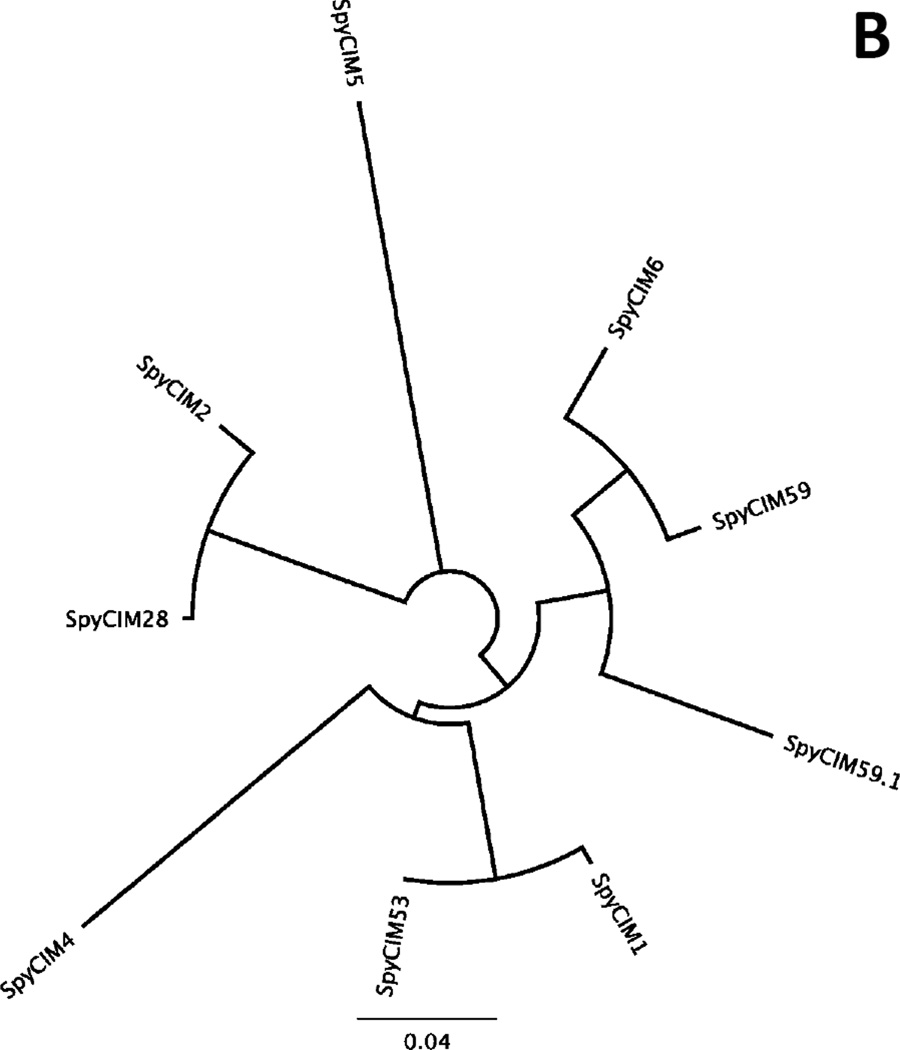
Figure 4. Prophage and SpyCI phylogenies.
Phylogenetic trees of the endogenous lambdoid prophages (A), and SpyCIs (B), from the 20 complete and publicly available GAS whole genome sequences (Table 2) is presented. When possible, the ends of each prophage genome were identified by the flanking duplications generated by site-specific recombination. For this analysis, all ICEs (except 10394.4, which has prophage features) and other MGEs are omitted. The encircled letters next to each phage taxon label (A) match the target gene attachment site identifier listed in Table 4. Trees were generated using the software package Geneious 6.1.7 (Biomatters Ltd.), employing the Tamura-Nei genetic distance model with neighbor joining and no outgroup.
The impact of phages and other MGEs upon their bacterial host cells can be divided into two broad categories: phage-encoded genes that contribute to the phenotype of the host cell (e.g., toxigenic conversion) and alterations of host cell gene expression due to integration of the phage. The impact of phage-encoded virulence genes that are mobilized by specialized transduction is well known, whereas the impact of site-specific integration on host cell gene expression is only beginning to be understood. An important class of phage-encoded GAS virulence genes includes the erythrogenic (pyrogenic) exotoxins that function as superantigens. Not all streptococcal superantigens are phage-associated (e.g., streptococcal mitogenic exotoxin Z [SmeZ]). However, the exotoxin genes that are phage-encoded are uniform in their association with phages, and they are not observed to occur as non-phage-associated genes, although a phage toxin gene may be part of a decaying prophage remnant that becomes fixed in the bacterial genome. Interestingly, some virulence genes often occur in genetically linked pairs that may enhance virulence through synergistic interactions (e.g., speC-spd1 and speH-speI). It has been proposed that the hyaluronidase that forms part of the phage tail fiber structure acts as a virulence factor, but such a role appears to be unresolved (Starr and Engleberg, 2006), and the function of hyaluronidase may be restricted to degrading the GAS capsular polysaccharide for the sole purpose of allowing phage attachment. Included among GAS prophages are genes with identifiable functions (Canchaya et al., 2002; Desiere et al., 2001), plus numerous ORFs encoding proteins of unknown function. Some of the uncharacterized genes may encode additional virulence factors as well as sRNAs and short peptides that participate in genetic regulatory networks within the bacterial host cell (Gottesman and Storz, 2011; Tesorero et al., 2013).
Several studies indicate that the expression of GAS prophage-encoded virulence genes is not autonomous, and may be linked to bacterial cell gene regulation. Expression of pyrogenic exotoxin C (speC) is up-regulated when GAS are co-cultured with human pharyngeal cells (Broudy et al., 2001). These growth conditions also stimulate prophage induction, which is apparently induced by a soluble factor released by the pharyngeal cells (Broudy and Fischetti, 2003; Gottesman and Storz, 2011; Holden et al., 2009). Similar findings were independently obtained when co-culture of GAS with pharyngeal cells lead to expression of prophage-encoded virulence genes for pyrogenic exotoxin K (speK), phospholipase A2 (sla), and streptodornase (sdn) (Banks et al., 2003a). GAS regulatory proteins can directly interact with phage genes. The GAS transcriptional regulator Rgg/RopB regulates expression of the prophage DNase Spd3 gene by binding its promoter (Anbalagan and Chaussee, 2013; Anbalagan et al., 2012; McShan et al., 2008), and the peroxide response regulator PerR enhances expression of the related prophage DNase Sda1 gene (Wang et al., 2013). The ability to selectively enhance the expression of DNases may facilitate survival of GAS when entangled in neutrophil extracellular traps (NETs) (Buchanan et al., 2006; de Buhr et al., 2014; Wartha et al., 2007). The interaction between GAS phage and non-phage genes and their products is an evolving field that is bound to shed important new light on the pathogenesis of GAS infections.
SpyCIs are a unique group of MGEs found frequently in GAS genomes. These phage-like elements integrate into the MMR gene mutL (Scott et al., 2008). SpyCIs have small genomes (12–17 kb) containing lysogeny and DNA replication modules, but they lack identifiable genes for DNA packaging or capsids. By integrating into the MMR operon, SpyCIs silence not only mutL but also the downstream genes lmrP (multidrug efflux pump), ruvA (Holliday junction helicase subunit), and tag (base excision repair). The consequence of this gene silencing is that the GAS cell adopts a complex mutator phenotype (Scott et al., 2012; Scott et al., 2008). Remarkably, SpyCI are dynamic elements. When GAS cells reach early logarithmic growth, SpyCI excises from the genome and allows transcription of mutL and the downstream genes to proceed. As the GAS cells approach stationary phase, SpyCI re-integrates into mutL and once again silences the downstream genes. This cycle of excision and re-integration causes the GAS cell to alternate between a wild type and mutator phenotype. One exception to this cycle is found in the emm5 strain (Manfredo), wherein a deletion in the SpyCIM5 integrase gene was compensated for by a novel promoter in the gene remnant that constitutively transcribes mutL and the other downstream genes. Outside of the highly conserved modules for lysogeny and replication, the remainder of SpyCI DNA is often quite divergent (Figure 4B), and includes many ORFs of unknown function, including potential sRNAs or peptides that may influence host cell function.
More than half of the GAS strains with completed genome sequences are host to a SpyCI (Scott et al., 2012; Scott et al., 2008) (Table 2). Some, but not all strains sharing an emm type, harbor a SpyCI (e.g., emm1 strain SF370 versus the three other emm1 strains; Table 2). Curiously, SpyCIM53 as defined by the primase C terminal 1 family protein target probe (based on strain Alab49), is present in only 20% of emm pattern A–C strains, but is detected in 70% of pattern D and 59% of pattern E strains evaluated by CGH (Bessen et al., 2011). Given the relatively low ratios for rates of recombination to mutation observed in housekeeping genes for emm pattern A–C organisms as compared to pattern D and E strains, and the large number of pattern A–C emm subtypes that presumably arise by mutation (Section 3), one might expect the throat specialists to have a higher concentration of mutator phenotypes. But that may not be the case if the distribution of SpyCIs is skewed in favor of skin specialist and generalist strains.
An efficient means for dissemination of SpyCIs must exist in spite of their lack of capsid genes. It may be that helper phages are tapped as a source of structural genes, as has been observed in the Staphylococcus aureus pathogenicity islands (SaPI) (Novick et al., 2010). The SpyCIs are not unique to GAS; genomes of Streptococcus anginosus, Streptococcus intermedius, SDE, S. canis, and Streptococcus parauberis display similar elements integrated into mutL (Nguyen and McShan, 2014). Although mutator phenotypes are common in wild populations of bacteria (Bucci et al., 1999; LeClerc et al., 1996; Oliver et al., 2000; Taddei et al., 1997), based on published reports to date, it appears that only in streptococci has such a remarkable molecular switch evolved that allows the bacterial host cell to cycle between mutator and wild type in response to growth.
Clustered, regularly interspaced short palindromic repeat (CRISPR) elements and CRISPR-associated (Cas) proteins constitute a microbial immune system that can protect the bacterial host cell from invasion by MGEs, including phage (Barrangou et al., 2007). CRISPR expression appears to generate anti-phage mRNA that interferes with the genetic program of the lytic cycle (Barrangou et al., 2007; Bolotin et al., 2005; Mojica et al., 2005). Examination of 13 GAS genomes reveals two CRISPR-Cas loci, wherein a given strain has both, one, or neither locus (Nozawa et al., 2011). A CRISPR locus is characterized by a succession of ~21- to 47-bp direct repeat sequences, separated by unique spacer sequences (~30- to 36-bp). Expression of repeat/spacer-derived, short CRISPR RNA (crRNA) leads to the silencing of foreign nucleic acids in a sequence-specific manner (Deltcheva et al., 2011). The CRISPR spacers in GAS genomes are homologs of known prophage sequences, or they may match yet undiscovered sequences (McShan et al., 2008); no sequence homologies have yet been found with other MGEs such as ICEs (Nozawa et al., 2011). The CRISPR elements provide a glimpse of previous encounters with bacteriophages in the evolutionary past of each GAS genome, and as more phage sequence information is acquired, they may provide insight into subpopulations of phages that interact with GAS which differ by emm type or disease association.
For 13 GAS genomes examined, a statistically significant inverse relationship between the numbers of CRISPR spacers versus prophages was observed (Nozawa et al., 2011). These findings support the notion that in GAS, CRISPRs inhibit prophage incorporation into host cell genomes and this action is dependent on the number of spacers. In addition, GAS strains lacking CRISPRs possess statistically significantly more prophages than CRISPR-harboring strains (Nozawa et al., 2011). However, based on Table 2 data for 20 complete and assembled GAS genomes, which include three emm pattern D strains and a slightly different accounting for the number of prophages in each strain (i.e., Table 2 counts only complete lambdoid phage genomes), the difference in the number of prophages for CRISPR-1-positive GAS versus CRISPR-1-negative GAS is not statistically significant (Student t-test, 2-tailed). Similarly, no significant difference is observed in the number of prophages for CRISPR-2-positive versus CRISPR-2-negative GAS. Of the 20 GAS strains considered, six lack both CRISPR elements (Table 2). Considering the number of prophages, or the combined number of prophages plus SpyCIs, and the presence versus absence of both CRISPR elements, no significant differences are detected; a lack of significance is also evident for the combined number of prophages plus SpyCIs versus either CRISPR-1 or CRISPR-2 (Table 2). Findings from CGH for 97 GAS strains of 95 distinct emm types reveal that the CRISPR-1 element is present in 40, 48 and 52% of emm pattern A–C, D and E strains, respectively (Bessen et al., 2011). However, there are highly significant differences in the distribution of the CRISPR-2 locus in accordance with emm pattern group, whereby none of the pattern D strains harbor CRISPR-2, in contrast to 35% and 66% of pattern A–C and E strains, respectively. The biological significance of the lack of the CRISPR-2 element among skin specialist strains remains to be established.
Among the other MGEs inhabiting the GAS genome are ICEs or ICE-like regions of difference (RD) [reviewed in (Banks et al., 2002; Beres and Musser, 2007; Green et al., 2005)]. Several ICE or RD forms have been well-described in GAS. An ~10–15 kb ICE/RD, which varies in size among GAS strains, harbors genes encoding the bacteriocin known as streptin; this RD is equivalent to AGR-13 and is present in >50% of GAS isolates of 95 different emm types (Bessen et al., 2011). Within a far more restricted set of GAS strains lies a ~37 kb element that encodes the R28 surface protein (Green et al., 2005) (discussed in Section 7); this ICE/RD is present in emm2 and emm28 strains and is nearly identical in nt sequence in both strains, indicative of a recent common ancestor (Beres and Musser, 2007).
Genes conferring resistance to antibiotics have been localized to ICEs within GAS. The mef(A) gene, which encodes a macrolide efflux pump and gives rise to the M phenotype for erythromycin resistance, is present within a ~59 kb chimeric element composed of a transposon inserted into a structure with prophage features, and has been associated with an emm6 outbreak clone (Φ10394.4) (Banks et al., 2003b). Also contained within this element is a type II restriction endonuclease-modification cassette, which may protect the host cell from subsequent infections by bacteriophage or other elements (Euler et al., 2007). In addition, mef(A) can be associated with other MGEs in GAS such as ΦM46.1, which also harbors the tet(O) gene (Brenciani et al., 2010). Numerous other ICEs (or plasmids) present in GAS harbor additional antibiotic resistance genes including tet(M), erm(A) and erm(B) (Banks et al., 2004a; Beres and Musser, 2007; Brenciani et al., 2012; Giovanetti et al., 2012; Tse et al., 2012; Varaldo et al., 2009; Willems et al., 2011; Woodbury et al., 2008). It was estimated that resistance to macrolides was acquired by GAS in at least 49 independent genetic events (Robinson et al., 2006), and tetracycline resistance has arisen ≥80 times (Ayer et al., 2007); both values may be gross underestimates.
7. Phylogenomics of GAS and closely related species
A very recent phylogeny was constructed using concatenated nt sequences of 136 core genes derived from whole genome sequences of 46 Streptococcus species (Richards et al., 2014). GAS is placed within the clade corresponding to the pyogenic division of streptococcal species. Its closest genetic relatives are S. canis and SDE, with SDD closely related to SDE; the S. equi subspecies Se and Sz lie on a closely related branch; S. agalactiae (GBS) also lies within the pyogenic division, but within a distinct sub-cluster. This finding (Richards et al., 2014) is generally consistent with the taxonomies derived from 16S rRNA sequences (Facklam, 2002), concatenates of inferred sequences of DNA replication and repair proteins (Beres et al., 2008), and a larger set of core genes from fewer pyogenic species (Lefebure et al., 2012), and phylogenetic reconstruction based on multilocus sequence analysis (MLSA) using seven housekeeping genes that are shared among numerous streptococcal species (Jensen and Kilian, 2012).
Gene families can arise and expand via intra-chromosomal gene duplication followed by neo- or sub-functionalization (i.e., paralogs) or alternatively, by xenologous gene displacement wherein a horizontally transferred ortholog from another lineage or species replaces the target gene, often via homologous recombination (Koonin et al., 2001). The latter is a form of replacing HGT. Analysis of 17 Streptococcus genomes, of which 12 are GAS, reveals that HGT of orthologs far outweighs the contribution of gene duplication to the expansion and diversification of gene families (Treangen and Rocha, 2011). Thus, replacing HGT involving orthologous genes provides an efficient means to introduce novel functions that may provide a selective advantage.
Interspecific HGT events among species of the pyogenic division are well-documented. Foreign DNA can be introduced into a recipient genome by additive or replacing HGT, and each mechanism results in a distinct phylogenetic signature (Choi et al., 2012). Whole genome sequence data allows for a comprehensive overview of gene gain and loss that occurred during species formation and evolution. Whole genome sequence data are presently available for many species within the pyogenic division including S. canis (Richards et al., 2012), SDE (Shimomura et al., 2011; Suzuki et al., 2011), SDD (Suzuki et al., 2011), Sz (Beres et al., 2008; Holden et al., 2009), Se (Holden et al., 2009), and GBS [e.g., (Glaser et al., 2002; Tettelin et al., 2005; Tettelin et al., 2002)].
HGT events that replace recipient DNA sequences with donor DNA via homologous recombination were inferred using the ClonalOrigin model-based approach, for GAS, SDE and SDD (Choi et al., 2012). Using one or two genomes per species, alignments yielded a shared core-genome of ~1.5 Mb (which is about the same size of the estimated GAS pan-genome based on 24 strains [Section 5], although the methodologies differ and the model-based method may be less conservative); although only one or two genomes of each species could be included in this analysis, other GAS genomes yielded similar results. Data indicate elevated rates of replacing HGT between GAS and SDE in both directions, but with marked asymmetry favoring the GAS → SDE direction by ~2-fold. Elevated rates of replacing HGT were also observed for SDE → SDD, but not for SDD → SDE or between GAS and SDD in either direction (Choi et al., 2012).
A potential drawback to the model-based approach used (Choi et al., 2012) is that only two SDE whole genome sequences are presently available for evaluation. An oligonucleotide-based microarray containing probes for >200 GAS genes (derived from GAS strains of three emm types) was used to examine the genomic content of 57 SDE strains by DNA-DNA hybridization (Davies et al., 2007a). Two major clusters of SDE strains could be distinguished by the presence or absence of up to 35 GAS gene homologs, many of whose products likely serve housekeeping functions. These data, much of which likely represent replacing HGT, provide further support for the GAS → SDE direction of HGT. Additional replacing HGTs between GAS and SDE, occurring in both directions, involve alleles of at least three of the housekeeping genes used for MLST (Ahmad et al., 2009; McMillan et al., 2010). There is also evidence for HGT between GAS and SDE of a FnBP gene within the FCT region (Towers et al., 2004) and emm types 12 and 57 (McMillan et al., 2010; Simpson et al., 1992).
Population surveys that evaluate individual gene sequences but include many more strains of both GAS and SDE, show that for at least two genes essential for virulence, there is strong support for interspecies gene flow that is consistent with skewing in the SDE → GAS direction. One possible SDE → GAS replacing HGT involves the pilus transcriptional regulatory rofA locus within the FCT region: rofA alleles present in only 73 (64%) of the 114 GAS isolates, but the highly homologous rofCG alleles are present in all 33 (100%) SDE strains examined (Bessen et al., 2005). The remaining GAS isolates have nra replacing rofA at the same locus, whereby alleles of the rofA and nra lineages exhibit ~33% nt sequence divergence. In contrast, the maximal nucleotide sequence divergence between rofA alleles of GAS and the rofCG alleles of SDE is only 2.0%, and the extent of sequence differences within both the rofA and rofCG sets of alleles are roughly of the same order as the differences between the two populations, indicative of a recent common ancestor. In another example suggestive of replacing HGT, ska alleles of the well-supported ska-2a subcluster are present in all 33 (100%) SDE strains examined, but in only 15 (17%) of 90 GAS strains (Kalia and Bessen, 2004); ska encodes streptokinase, a plasminogen activator having a well-established role in virulence. The presumptive replacing HGTs of rofA and ska-2a, from SDE → GAS, may represent xenologous gene displacements. Of biological relevance is the finding that the distributions of rofA/nra and ska subcluster alleles among GAS also show statistically significant correlations with emm pattern grouping, and the orthologous forms show signs of positive selection (Bessen et al., 2005; Kalia and Bessen, 2004). It is not known whether these likely HGT events involving virulence genes were detected by the ClonalOrigin analysis (Choi et al., 2012), and DNA-DNA hybridization may not provide sufficient sensitivity (Davies et al., 2007a). More extensive whole genome sequence analysis of numerous strains belonging to both streptococcal species, as well as other related species, should permit a more precise analysis of the direction of net gene flow, as well as aid in identifying species-specific genes.
Divergent (but homologous) regulatory genes and their targets can give rise to cross-regulation and add layers of complexity to transcriptional networks. RofA-like proteins (RALPs) are present in various combinations among the different GAS strains, and many of their target genes encode MSCRAMMs which also comprise a gene family (Kreikemeyer et al., 2007). The presence of RALPs and their target genes in GAS may have occurred by a mixture of replacing and additive HGT events (Section 8). Members of the Rgg family of transcriptional regulators (Rgg2 and Rgg3 [distinct from Rgg/RopB]) and their cognate pheromone peptides [Section 4; (Cook et al., 2013)], which are also present in multiple species of the pyogenic division, may have also evolved by similar mechanisms, leading to a complex interspecific communication network.
A parsimony-based analysis that considers the pan-genome was used to evaluate additive HGT events among GAS, SDE and SDD, however, findings failed to reveal a strong bias in directionality for additive transfers between GAS and SDE (Choi et al., 2012). Genome-wide phylogenetic analyses involving 15 Streptococcus species were used to ascertain the number and nature of genes that were gained on the lineage leading to GAS (13 strains); 113 genes were gained from a combination of sources that include other GAS strains, other Streptococcus species and other bacterial species, or they arose de novo among GAS; together the gained genes represent ~6% of the average GAS genome size and almost half of them are phage-associated (Lefebure et al., 2012). Gained genes with an established role in virulence in GAS include speB, plus its upstream transcriptional regulatory gene rgg/ropB. SpeB encodes a secreted cysteine protease that degrades numerous host proteins involved in defense, as well as extracellular proteins of GAS (Nelson et al., 2011); the ancestral source of the speB gene remains unknown.
Horizontal movement of bacteriophage between member species of the pyogenic division is well-established. Transfer of phage is likely to have occurred between GAS and SDE (Davies et al., 2007b; Shimomura et al., 2011) and between GAS and Se (Holden et al., 2009). Many GAS-derived prophage mobilize genes encoding pyrogenic exotoxins via specialized transduction, and exotoxin genes that are highly homologous to those found in GAS are also present in other streptococcal species (Holden et al., 2009; Shimomura et al., 2011); these exotoxins can function as superantigens and contribute to pathogenesis in toxic shock syndrome. Interestingly, the two sequenced genomes of Sz lack both prophage and ICEs (Beres et al., 2008; Holden et al., 2009).
Several of the genes gained on the S. pyogenes species-specific branch share sequence homology with GBS genes, suggesting that GBS is a potential donor, although most of these appear to be housekeeping genes (Lefebure et al., 2012). HGT likely occurred between GAS and GBS involving the gene encoding the virulence factor C5a peptidase (Chmouryguina et al., 1996), which is present in (nearly) all strains of both species. In GBS, the C5a peptidase gene and an adjacent gene encoding a laminin-binding protein (lmb) share 98% nt sequence identity to GAS orthologs and localize to a composite transposon (Franken et al., 2001). Of potential relevance is the finding that in GAS, the lmb gene lies adjacent to a hotspot for chromosomal inversion (SPy2006–2007; Figure 3).
A virulence factor that appears to originate from GBS and was acquired by a limited number of GAS strains is the R28 surface protein (SpyM28_1336). The R28 protein is a chimera of three GBS surface proteins (proteins Rib, α, and β); this mosaic structure is indicative of past recombinational events (Stalhammar-Carlemalm et al., 1999). R28 is encoded within an exogenous ICE/RD of ~37 kb in emm28 and emm2 strains (Beres and Musser, 2007; Green et al., 2005; Sitkiewicz et al., 2011). R28-expressing GAS have a strong epidemiologic association with puerperal sepsis, also known as childbed fever, which is a pelvic infection that occurs during or shortly after childbirth. The R28 protein mediates adherence of GAS to cultured epithelial cells derived from human cervical tissue (Stalhammar-Carlemalm et al., 1999); GBS naturally colonize the vaginal epithelium. Thus, puerperal fever caused by GAS and neonatal sepsis caused by GBS are probably initiated by bacteria colonizing the vaginal epithelium, and the possible acquisition of the R28 gene by GAS from GBS may have helped create disease-specialist clones of GAS (Green et al., 2005). Interestingly, the FCT region of an emm2 strain (FCT-6), which is rare among GAS of other emm types (Falugi et al., 2008; Kratovac et al., 2007), has high sequence similarity to the FCT-like region of a SDE strain (Shimomura et al., 2011) and to the pilus gene region of GBS (Beres and Musser, 2007), suggestive of acquisition of the FCT-6 region by GAS following HGT from another streptococcal species.
8. Long-term evolution: Population genomics and tissue tropisms
One of the striking features of GAS biology is the distinction in emm types among GAS causing disease in temperate versus tropical communities where pharyngitis and impetigo, respectively, prevail (Section 2). The spatial-temporal distances that act to separate certain strains of GAS from one another may limit HGT due to reduced opportunities in sharing the same niche. Yet, the emm pattern groupings do not correspond to discrete evolutionary lineages based on core housekeeping gene sequences (Section 3). Thus, there may be a common housekeeping gene pool that occupies both ecological niches, perhaps via HGT events involving emm pattern E generalist strains. The lack of clear pathways of evolutionary descent for throat versus skin specialist strains, based on core genes, prompted evaluation of the distribution of accessory genes that may play a critical role in establishing tissue site preferences for infection.
A population genomics approach was used to identify non-phage accessory gene regions (AGRs) of GAS, whereby 22 AGRs were identified by CGH of 97 GAS strains (95 emm types), defined by their presence/absence in 15 to 85% of strains (Section 5). A SplitsTree graph based on the presence or absence of 21 AGRs (excluding AGR-21/21X which contains the emm pattern genotype) reveals strong clustering for the vast majority of pattern D strains, and for the vast majority of pattern E strains, with pattern A–C strains sprinkled throughout both the pattern D- and E-dominant clusters (Bessen et al., 2011). The AGR-based SplitsTree finding was confirmed by population genetic structure analysis of GAS, inferred using a Bayesian nonhierarchical clustering method. A finding of pronounced clustering of AGRs in accordance with emm pattern genotype is indicative of co-inheritance and strong linkage among select combinations of AGRs. Maintaining linkage among AGRs, against a background of extensive recombination among core genes (which disrupts nonrandom associations), is highly consistent with the hypothesis that certain combinations of AGRs are subject to strong co-selection pressures as a result of having critical roles in adaptation, such as a tissue tropism for infection.
The distribution among 97 GAS strains of the 393 differential gene targets was further evaluated using the Fisher exact test for independence, for all possible pairwise combinations (>77,000 tests) (Bessen et al., 2011). Eight genes displaying the strongest linkage (BH-corrected p-values < 0.00001), and belonging to 8 different AGRs, were selected for construction of a simplified evolutionary network. Based on presence/absence of 8 genes, 34 unique haplotypes (H) were defined for the 97 GAS strains. A simplified evolutionary network, whereby every strain is connected to >1 other strain by gain/loss of a single AGR, was constructed and includes 26 of the 34 haplotypes (Figure 5). Two major clusters connected by a single weak link (H24) are evident, with 88% of emm pattern D (skin) strains in cluster I (upper) and 89% of emm pattern E (generalist) strains in cluster II (lower). Of the 18 emm-types representing emm pattern A–C strains (throat), 61% fall within cluster I, whereas 17% are in cluster II (including emm12); the remainder fall outside the main network (including two representatives of the highly prevalent emm1, strains SF370 and MGAS5005 [Table 2]).
Figure 5. Model of evolution for GAS based on highly linked accessory gene region (AGR) genes.
All possible pairwise comparisons between 393 differentially distributed GAS genes were made for 97 GAS strains of 95 emm types; eight AGRs having genes with the smallest p-values (Fisher's exact test), when paired with a gene from another AGR, were identified (AGRs 2X, 3, 4, 13, 14, 16B, 17A and 21X) (Bessen et al., 2011). The gene within each of the eight AGRs having the smallest p-value (i.e., strongest linkage disequilibrium) is used to define character states (0, absence; 1, presence) for their respective AGRs among the 97 GAS strains; 34 haplotypes (H) are defined according to the character states of these eight AGR genes (Bessen et al., 2011). Single locus differences among haplotypes were established using the eBURST clustering algorithm (www.mlst.net) and redrawn; 26 of the 34 haplotypes differed from at least one other haplotype by only a single AGR gene, and they are depicted in the haplotype network shown; not shown are eight haplotypes that differ from the main network by ≥two loci. Circles denote each haplotype (H), whereby the size/area of the circle reflects the number of assigned GAS strains; emm pattern genotypes are represented by black (pattern A–C), white (pattern D) and gray (pattern E), and their fractional content is displayed. Lines connect all haplotype pairs that differ by a single locus; the set of thicker lines connect two core haplotypes, which are defined as being represented by >1% of the 97 GAS strains. H24 provides a link between the two major sub-clusters (I, upper; II, lower). Adapted from (Bessen et al., 2011).
The emm pattern A–C (throat specialist) strains of cluster I belong to the connected haplotypes H17-H19-H23/H29 (Figure 5; upper). Assigned to these haplotypes are M-types associated with significant causes of pharyngitis and acute rheumatic fever (e.g., M-types 3, 5, 6, 14, 18 and 29) (Bisno, 1980; Johnson et al., 1992; Shulman et al., 2009; Steer et al., 2009b; Stollerman, 1991). Analysis of this major sub-cluster of emm pattern A–C strains within cluster I may lead to deeper insight on the key molecular determinants that distinguish many throat and impetiginous skin infections. The transition between H17 and H19 is marked by the loss/gain of a FnBP gene (prtF1) in AGR-2/2X. This finding sparked an even more detailed analysis of gain/loss of other differentially distributed FnBP genes among H17–H19 strains, and new extended haplotypes were defined (Bessen et al., 2011). The extended haplotypes reveal a key transition in the tendency for GAS strains to infect different host tissue, that lies between H17x and H17y (i.e., extended haplotypes that are subsets of H17), based on a meta-analysis of 29 surveillance studies (Table 1) normalized for equivalent numbers of pharyngitis and impetigo isolates. The percentage of pharyngitis and impetigo isolates sharing an emm type assigned to a given haplotype indicates that H19x and H17y represent classical throat strains, and H18 and H17x represent classical skin strains. For emm types associated with H18 and H17x, impetigo isolates outnumber pharyngitis isolates by ~20-fold (i.e., 95% are impetigo -derived). In sharp contrast, for isolates having emm types associated with H17y or H19x (N = 654 isolates), all (100%) were recovered from pharyngitis. Thus, the genetic transition between H18/H17x and H17y/H19x coincides with a profound switch in phenotype for the tendency to infect different host tissue.
Differences in gene content for H18, H17y and H19x GAS strains were assessed. The difference between H17x and H18 is loss/gain of AGR-4; AGR-4 lacks known virulence genes. Importantly, the transition from H18/H17x (impetigo) to H17y (pharyngitis) involves loss of fbaA. The transition from H17y to H19x involves loss of prtF2 and gain of prtF1. The fbaA locus belongs to AGR21/21X, whereas prtF1 and prtF2 belong to AGR-2/2X. Thus, the gain/loss of fbaA, prtF1 and/or prtF2 may impact virulence at the throat and skin. However, it remains possible that additional genetic changes contribute to the tissue-specific phenotype shift, such as SNPs in core or other accessory genes. Furthermore, other classical throat strains (e.g., M1, which lies outside the major haplotype network) might utilize alternative molecular pathways, as might also be the case for the emm pattern E generalists.
Gain/loss of other genes may also contribute to tissue site preferences for infection. Strong genetic linkage to emm pattern group is observed for several other genes having an established role in virulence or another important biological function (Table 5). Accessory genes that display linkage disequilibrium (i.e., p < 0.05) with the emm pattern marker for throat specialist versus skin specialist strains are strong candidates for conferring tissue-specific tropisms, particularly when they lie physically distant from AGR-21/21X on the chromosome. emm pattern-linked genes include numerous genes within AGR-2/2X and AGR-21/21X that encode transcriptional regulators (i.e., mga-1/mga-2, rofA/nra, msmR), and surface proteins having an established role in virulence (i.e., FnBP genes prtF1, prtF2 and fbaA), as well as the plasminogen activator streptokinase (ska) (Table 5). Accessory genes whose presence/absence are strongly linked to emm pattern E (e.g., sof, sfbX, spn/nga, sse, grab, ralp3; Table 5) might play a dual role in infection of both the skin and throat epithelial tissue sites.
Table 5.
Biologically important GAS genes having a strong correlation with emm pattern group
| Gene | AGR | Function # | Lineages ^ | % pattern A–C |
% pattern D |
% pattern E |
Significant differences (p < 0.05) * |
No. emm types in study |
Reference |
|---|---|---|---|---|---|---|---|---|---|
| rofA | 2/2X | transcriptional regulator | rofA; nra | 15 | 84 | 7 | D versus A–C or E |
112 | (Bessen et al., 2005; Kratovac et al., 2007) |
| msmR | 2/2X | transcriptional regulator | N/A | 50 | 91 | 68 | A–C versus D | 95 | (Bessen et al., 2011) |
| prtF1 | 2/2X | FnBP | N/A | 75 | 21 | 93 | D versus A–C or E |
112 | (Kratovac et al., 2007) |
| prtF2 | 2/2X | FnBP | N/A | 50 | 92 | 69 | A–C versus D | 112 | (Kratovac et al., 2007) |
| cpa | 2/2X | collagen-binding protein | N/A | 25 | 92 | 55 | A–C versus D or E |
112 | (Kratovac et al., 2007) |
| spn/nga | 3 | NAD+ glycohydrolase |
active; inactive | 10 | 14 | 94 | E versus A–C or D |
110 | (Riddle et al., 2010) |
| ralp3 | 9 | transcriptional regulator | N/A | 15 | 6 | 34 | D versus E | 95 | (Bessen et al., 2011) |
| srtT | 13 | lantibiotic (streptin) biosynthesis (ICE/RD) |
N/A | 45 | 24 | 82 | E versus A–C or D |
95 | (Bessen et al., 2011) |
| grab | 16A/B | α2-macroglobulin binding protein |
grab; AGR-16B | 85 | 94 | 55 | D versus E | 95 | (Bessen et al., 2011) |
| sse | 18A/B | esterase | sse-A; sse-B | 95 | 97 | 66 | D versus E | 95 | (Bessen et al., 2011) |
| mga | 21/21X | transcriptional regulator | mga-1; mga-2 | 95 | 0 | 0 | A–C versus D or E |
112 | (Bessen et al., 2005; Kratovac et al., 2007) |
| sof | 21/21X | serum opacity factor; FnBP | N/A | 5 | 11 | 100 | E versus A–C or D |
112 | (Kratovac et al., 2007) |
| sfbX | 21/21X | FnBP | N/A | 10 | 12 | 93 | E versus A–C or D |
95 | (Bessen et al., 2011) |
| fbaA | 21/21X | FnBP; antiphagocytic | N/A | 25 | 100 | 77 | D versus A–C or E |
95 | (Bessen et al., 2011) |
| ska | 21/21X | streptokinase; plasminogen activator |
ska-1; ska-2a; ska-2b | 0 | 67 | 4 | D versus A–C or E |
67 | (Kalia and Bessen, 2004) |
FnBP, fibronectin-binding protein
The lineage highlighted in bold corresponds to data for % distribution among emm pattern A–C, D and E groupings
Fisher exact test (2-tailed) with Benjamini-Hochberg corrected p-values
N/A, not applicable
9. Short-term evolution: Comparative genomics in epidemics and outbreaks
The monophyletic grouping of many GAS strains in accordance with emm type (Figure 1) readily allows for inferring ancestral-descendent pairs, particularly when coupled with epidemiological collections of strains with known dates of recovery. Whole genome sequencing of multiple isolates of GAS sharing the same emm type has provided important new insights on the genetic differences between organisms recovered from different hosts within the same community, genetic changes that are associated with different disease manifestations, the identity of genes subject to strong diversifying selection over time, and the evolutionary history underlying the emergence of new epidemic clones. Whole genome sequencing of GAS is also having increased utility in the clinical microbiology lab and in investigating nosocomial outbreaks.
GAS isolates of the emm3 type are among the most thoroughly studied by genomics. Organisms of the emm3 type are highly prevalent among cases of pharyngitis, invasive disease, and acute rheumatic fever. Since the first reports of whole genome sequences for two emm3 strains (Beres et al., 2002; Nakagawa et al., 2003) (Table 2), hundreds of genomes of emm3 isolates have been fully or partially sequenced via several methods (Beres et al., 2010; Beres et al., 2006; Beres et al., 2004; Shea et al., 2011). Ninety-five invasive disease isolates recovered from Ontario, Canada within an 11-year period revealed a total of 4,269 polymorphisms within the emm3 core-genome, with an average of 49 SNPs and 11 indels per isolate; ~65% of the polymorphisms are specific to a single isolate, and within a single isolate ~10% of the polymorphisms are specific to that isolate (Beres et al., 2010). Thus, the core-genomes of emm3 isolates from the Ontario study are closely related, yet each and every isolate is unique. Two epidemic waves of invasive emm3 infections (peaking in 1995 and 2000) show differences in emm3 subtype alleles, wherein isolates recovered in the second wave have a duplication of the first four amino acid residues of the mature M3 protein; this change led to increased resistance to opsonophagocytosis by the organism (Beres et al., 2004). However, a major source of genomic diversity among the invasive emm3 isolates involves the gain or loss of prophage.
A more precise model of invasive emm3 disease in Ontario emerged following inclusion of a third epidemic wave (peak in 2005) and analysis of 344 GAS isolates. This study yielded 280 informative core-genome polymorphisms that were used to construct a phylogenetic tree based on biallelic characters defining 97 distinct haplotypes (Beres et al., 2010). Ten major subclones were identified. Based on progenitor-descendent pairs, the rate of accumulation of core-genome polymorphisms was estimated at 1.7 SNPs per organism per year. Data show that each successive epidemic wave consists of organisms that are genetically distinct from those which preceded it. Although some subclones appear to have evolved from strains that were circulating during an earlier wave, new migrants also contributed to the overall level of emm3 invasive GAS disease.
The genome sequences of emm3 invasive isolates from Ontario were compared to emm3 organisms recovered from individuals with asymptomatic carriage in Texas (Beres et al., 2006). Despite differences in virulence in a mouse model for systemic infection, there were no SNPs in the core-genome that differentiated the two clinical groups, suggesting that the two phenotypes arose multiple times via independent evolutionary pathways. A similar conclusion was reached in a more recent study that compared genome sequences of 86 pharyngitis isolates to those of 100 invasive disease isolates, all from Ontario (Shea et al., 2011). In terms of genetic diversity in the emm3 core-genome, the pharyngitis and invasive isolates were highly similar, averaging 79 and 77 polymorphisms per isolate, respectively, relative to the MGAS315 reference strain. Data indicate that invasive isolates originate from all major lineages of pharyngitis isolates and they are more similar to the circulating pharyngitis strains from which they evolved as opposed to other invasive strains (Shea et al., 2011).
Despite the lack of "smoking gun" polymorphisms that clearly distinguish emm3 GAS isolates associated with different clinical syndromes, there are distinctions between disease groups in terms of the genes undergoing strong diversifying selection, which is indicative of ongoing adaptations. For example, in pharyngitis isolates there is an overrepresentation of indels in the hasA and hasB genes that are essential for biosynthesis of the hyaluronic acid capsule, a key virulence factor (Shea et al., 2011). The numbers of polymorphic sites in genes encoding the CovRS/CsrR and Rgg/RopB transcriptional regulators are also skewed in invasive disease versus pharyngitis isolates (Beres et al., 2010; Ikebe et al., 2010; Olsen et al., 2012; Shea et al., 2011). Many of the SNPs result in non-synonymous substitutions which in turn, lead to extensive diversity in phenotype among GAS isolates that are otherwise highly similar in their core-genomes (Carroll et al., 2011).
Prior to 2004, the worldwide recovery of emm59 GAS strains was extremely rare. However, by 2006 a hypervirulent emm59 clone emerged in western Canada and rapidly became a major cause of invasive GAS disease (Tyrrell et al., 2010). Genome sequencing of >100 isolates provided a magnified view of the unfolding emergence and epidemic spread of emm59 GAS (Fittipaldi et al., 2012a; Fittipaldi et al., 2012b). The epidemic isolates are virtually identical with an average of only 7.3 SNPs and 4.3 indels relative to the epidemic reference strain MGAS15249 (Table 2). Thus, the contemporary emm59 isolates descended from a recent common ancestor that underwent rapid geographic spread. When compared to a historic isolate recovered in the 1970s (MGAS1882; Table 2), the new epidemic emm59 clone differed by only 118 biallelic SNPs and 12 indels in the emm59 core-genome. Both isolates have a dearth of phage genes, whereby the older clone has only one prophage, which is absent from the epidemic clone, and the new clone has only a prophage remnant whose genes are absent from the historic clone. While the precise molecular steps and natural selection processes that led to the emergence of the new epidemic clone remain unresolved (Bessen, 2012), the contemporary and historic strains display distinct phenotypes in experimental models (Fittipaldi et al., 2012a).
The most common emm type recovered in association with GAS invasive disease in established market economy countries throughout the world is emm1; this emm type also ranks very high in prevalence for cases of pharyngitis (Beall, 2014; Shulman et al., 2009; Steer et al., 2009b). Studies on emm1 isolates that spread globally in the late 1980s and early 1990s had two prophage which were absent from emm1 isolates recovered prior to that period (Cleary et al., 1998). The very first sequenced genome of GAS is the emm1 strain SF370 (Table 2) (Ferretti et al., 2001), however, SF370 is more representative of the historic emm1 clones (Sumby et al., 2005). The evolution of emm1 isolates has been intensively studied by genome sequencing and other methods (Aziz et al., 2005; Aziz and Kotb, 2008; Maamary et al., 2012; Nasser et al., 2014; Sumby et al., 2005).
In a recent report based on >3,500 genome sequences, a detailed model for the most likely evolutionary history of contemporary emm1 isolates is put forth (Nasser et al., 2014). In this model, a phage encoding an extracellular DNase (sdaD2/sda1 gene) was acquired first, followed by acquisition of a second phage encoding the SpeA1 superantigen, whose gene subsequently underwent a non-synonymous point mutation to give rise to SpeA2 in the early 1970s; that mutational change confers an increased affinity of SpeA2 for HLA-DQ class II antigens (Kline and Collins, 1996). The final major change was acquisition of a 36-kb region from a probable emm12 donor strain in ~1983. The newly acquired 36-kb region encodes the secreted toxins NAD+-glycohydrolase (Spn/Nga) and streptolysin O; about half the polymorphisms in the 36-kb region are concentrated in a highly divergent 2.6-kb region between the slo and metB genes (Nasser et al., 2014). This narrow region roughly corresponds to AGR-3 (Bessen et al., 2011). In addition, the contemporary MGAS5005 strain lacks the G330D polymorphism in Spn/Nga and truncated ifs (endogenous inhibitor of Spn/Nga) that are accurate predictors of NADase inactivity (Riddle et al., 2010), whereas the historic SF370 isolate has both the G330D polymorphism and a truncated ifs pseudogene (data not shown); thus, a recent acquisition of NADase activity via the 36-kb region by contemporary emm1 strains may have contributed to its evolutionary success and/or hypervirulence. It is proposed that the 36-kb region was probably mobilized by generalized transduction and integrated into the emm1 genome following homologous recombination (Nasser et al., 2014; Sumby et al., 2005).
Comparison of the polymorphic sequences of genomes from invasive- versus pharyngitis-derived emm1 isolates failed to yield a clear disease association, supporting the idea that invasive emm1 organisms evolve repeatedly from the pool of pharyngitis strains (Nasser et al., 2014), similar to the conclusion reached for emm3 organisms. However, as previously observed for emm3 strains (Shea et al., 2011), the emm1 pharyngitis isolates have many more indels and SNPs in the capsular genes hasA and hasB as compared to emm1 invasive strains, indicative of strong diversifying selection in pharyngitis strains (Nasser et al., 2014). In addition, numerous mutations in the transcriptional regulatory genes covRS and rgg/ropB in emm1 strains and those of other emm types facilitate the transition between localized and invasive GAS infection (Cole et al., 2011; Garcia et al., 2010; Ikebe et al., 2010; Masuno et al., 2014). Mutations in covRS and rgg/ropB lead to the down-regulation of transcription of the gene encoding the broad-spectrum cysteine protease SpeB that, in turn, prevents the degradation of extracellular proteins such as DNase. In the absence of SpeB, the Sda1/SdaD2 DNase encoded by the phage recently acquired by contemporary emm1 strains is spared degradation, and is able to digest the DNA-rich neutrophil extracellular traps (NETs), facilitating the escape of trapped GAS and acting as a selective force that promotes the emergence and invasive spread of covRS mutants (Walker et al., 2007). A recent study that sequenced the genomes of GAS (emm1) recovered from several different body sites of a single patient throws into question the exact anatomical location where isolates with inactivating mutations in covRS or rgg/ropB begin to emerge (Flores et al., 2014). However, while the association of SpeB absence with invasive disease is very highly significant, it is an imperfect correspondence.
In 2011 there was an alarming increase in the number of cases (>1,000) of scarlet fever in Hong Kong, and many patients were hospitalized. The outbreak was largely attributable to emm12 GAS strains that were resistant to macrolides and tetracycline (Hsieh and Huang, 2011; Lau et al., 2012; Luk et al., 2012). A complete genome (HKU16; Table 2) and draft genome (HKU30; Table 3) of organisms recovered from scarlet fever patients (Tse et al., 2012) were compared to two previously sequenced emm12 genomes of non-scarlet-fever isolates (Beres and Musser, 2007). The pathognomonic scarlatina rash is triggered by superantigen activity (i.e., "erythrogenic toxin") and indeed, HKU16 and HKU30 have a novel prophage (ΦHKU.vir) harboring the superantigen genes speC and sse (Tse et al., 2012). Both strains also carry two prophage common to MGAS9429 that have speH and speI superantigen genes on one prophage, and the DNase sdaD2/sda1 gene on the other. However, the Hong Kong scarlet fever outbreak was multiclonal in origin. Strain HKU16 has a chromosomal inversion that is lacking in HKU30 (Table 2). HKU16 also harbors a 65-kb ICE (ICE-emm12) that contains erm(B) and tet(M) genes. HKU30 lacks ICE-emm12, but has related genes with an overall 67% similarity and positioned at a different chromosomal location; thus, the two genetic elements share a common but distant evolutionary ancestor. Despite the multi-clonal nature of the outbreak, most GAS recovered from scarlet fever patients had ΦHKU.vir and ICE-emm12 (Tse et al., 2012).
Whole genome sequencing was recently used to investigate an outbreak of puerperal fever (Section 7) in four hospitals in New South Wales, Australia during 2010 (Table 3) (Ben Zakour et al., 2012). Among the 11 GAS isolates from 9 patients were six different emm subtypes, indicative of independent sources of infection for at least five patients. The five emm28.8 isolates were recovered from four patients in three hospitals; among them were 72 SNPs, but all had the ICE/RD harboring the gene for the R28 protein that promotes vaginal colonization (Section 7). Data indicate that the two patients from hospital A were infected with identical clones, wherein the second infection was the likely result of patient-to-patient or staff-to-patient transfer. The genomic contents of emm28.8 isolates from patients in hospitals B and C were distinct from each other and from the hospital A isolates, indicative of an outbreak of a polyclonal nature (Ben Zakour et al., 2012). Whole genome sequencing of GAS is gradually becoming part of the clinical microbiology lab repertoire.
10. The shaping forces of natural selection
Although they are strictly human pathogens with no known environmental reservoir, the biological behavior of GAS is quite varied. The most common habitat for GAS is the non-diseased URT, wherein the organism assumes the commensal-like state of asymptomatic carriage. Next most often encountered by GAS is the purulent exudate (i.e., pus) triggered by infection at a superficial epithelial surface of the oropharynx or skin. For effective transmission to new hosts, it may be necessary for GAS to not only survive in this environment, but also produce new progeny. Once the protective barriers of the host are fully breached, and GAS gains access to normally sterile tissue such as the bloodstream, the local microenvironment shifts drastically once again. The more highly specialized disease conditions caused by GAS, such as scarlet fever and puerperal fever, may be due to disease-specialist clones that carry a unique array of virulence factors which are tailored to that specific disease. Even among invasive diseases, specialist clones appear to exist, such as those that cause toxic shock syndrome.
The greatest challenges to GAS survival are likely encountered within its human host. Among the most prominent threats to GAS survival and reproductive growth are antibiotics, lantibiotics produced by resident organisms of the local microbiome, and innate and acquired host defenses. The emerging picture for the strategy used by this species is one that includes an uncanny ability to acquire DNA from a multitude of diverse sources, while maintaining a core-genome that is very highly conserved. This operates in parallel with extensive changes in gene clusters that encode surface proteins (e.g., emm) and/or mutations in various regulatory genes, whereby genetic change is often subject to positive selection and results in immune escape and/or gain or loss of function. It is the highly diverse non-core genome that confers much of the complexity in biological behavior that characterizes GAS and defines it as a major problem pathogen.
Highlights.
-
*
The epidemiology of group A streptococci (GAS) is reviewed, highlighting emm genes
-
*
The genetic organization and population biology of GAS is discussed
-
*
The core and non-core genomes of 24 GAS isolates are evaluated, including prophage
-
*
Genes acquired by GAS genomes from related species are summarized
-
*
Genome changes underlying long- and short-term evolution of GAS are delineated
ACKNOWLEDGEMENTS
We thank Andrew Steer for providing detailed emm type data on published epidemiological studies, Sean C. Daugherty for help with annotation and data management, and our anonymous reviewers for many helpful suggestions. Work on Streptococcus pyogenes has received generous support from The National Institutes of Health (GM060793, AI053826, AI061454, AI065572 and AI072718) and the Oklahoma Center for the Advancement of Science and Technology (HR11-133).
Abbreviations
- AGR
accessory gene region
- C4BP
C4b-binding protein
- Cas
CRISPR-associated proteins
- CC
clonal complex
- CGH
comparative genomic hybridization
- CRISPR
clustered, regularly interspaced short palindromic repeat
- D
genetic diversity
- DNase
deoxyribonuclease
- emm
encodes M protein
- FCT region
encodes pili and T-antigen
- FnBP
fibronectin-binding protein
- GAS
group A streptococci
- GBS
group B streptococci
- GCS
group C streptococci
- GGS
group G streptococci
- H
haplotype
- HGT
horizontal gene transfer
- ICE
integrative and conjugative element
- LCB
locally collinear block
- MGE
mobile genetic element
- MLST
multilocus sequence typing
- MMR
mismatch repair
- MSCRAMMs
microbial surface components recognizing adhesive matrix molecules
- Ro
basic reproductive rate
- RD
region of difference
- Sda
streptococcal DNase (also, Sdn and Spd)
- SDD
Streptococcus dysgalactiae subspecies dysgalactiae
- SDE
Streptococcus dysgalactiae subspecies equisimilis
- Se
Streptococcus equi subspecies equi
- SF
subfamily
- SLV
single locus variant
- SOF
serum opacity factor
- Spe
streptococcal pyrogenic exotoxin
- Spn/Nga
NAD+-glycohydrolase
- SpyCI
streptococcal phage-like chromosomal island
- sRNA
small regulatory RNA
- ST
sequence type
- Sz
Streptococcus equi subspecies zooepidemicus
- URT
upper respiratory tract.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Contributor Information
W. Michael McShan, Email: William-McShan@ouhsc.edu.
Scott V. Nguyen, Email: Scott-Van-Nguyen@ouhsc.edu.
Amol Shetty, Email: AShetty@som.umaryland.edu.
Sonia Agrawal, Email: SAgrawal@som.umaryland.edu.
Hervé Tettelin, Email: tettelin@som.umaryland.edu.
REFERENCES
- Aanensen DM. London: Imperial College; 2014. http://spyogenes.mlst.net. [Google Scholar]
- Ahmad Y, Gertz RE, Jr, Li Z, Sakota V, Broyles LN, Van Beneden C, Facklam R, Shewmaker PL, Reingold A, Farley MM, Beall BW. Genetic relationships deduced from emm and multilocus sequence typing of invasive Streptococcus dysgalactiae subsp. equisimilis and S. canis recovered from isolates collected in the United States. Journal of clinical microbiology. 2009;47:2046–2054. doi: 10.1128/JCM.00246-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Akiyama H, Morizane S, Yamasaki O, Oono T, Iwatsuki K. Assessment of Streptococcus pyogenes microcolony formation in infected skin by confocal laser scanning microscopy. Journal of Dermatological Science. 2003;32:193–199. doi: 10.1016/s0923-1811(03)00096-3. [DOI] [PubMed] [Google Scholar]
- Alberti S, Garcia-Rey C, Dominguez MA, Aguilar L, Cercenado E, Gobernado M, Garcia-Perea A. Survey of emm gene sequences from pharyngeal Streptococcus pyogenes isolates collected in Spain and their relationship with erythromycin susceptibility. J. Clin. Microbiol. 2003;41:2385–2390. doi: 10.1128/JCM.41.6.2385-2390.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anbalagan S, Chaussee MS. Transcriptional regulation of a bacteriophage encoded extracellular DNase (Spd-3) by Rgg in Streptococcus pyogenes. PloS one. 2013;8:e61312. doi: 10.1371/journal.pone.0061312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anbalagan S, Dmitriev A, McShan WM, Dunman PM, Chaussee MS. Growth phase-dependent modulation of Rgg binding specificity in Streptococcus pyogenes. Journal of bacteriology. 2012;194:3961–3971. doi: 10.1128/JB.06709-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Angiuoli SV, Dunning Hotopp JC, Salzberg SL, Tettelin H. Improving pan-genome annotation using whole genome multiple alignment. BMC bioinformatics. 2011;12:272. doi: 10.1186/1471-2105-12-272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Angiuoli SV, Salzberg SL. Mugsy: fast multiple alignment of closely related whole genomes. Bioinformatics. 2011;27:334–342. doi: 10.1093/bioinformatics/btq665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anthony BF, Kaplan EL, Wannamaker LW, Chapman SS. The dynamics of streptococcal infections in a defined population of children: serotypes associated with skin and respiratory infections. Am.J.Epidemiol. 1976;104:652–666. doi: 10.1093/oxfordjournals.aje.a112344. [DOI] [PubMed] [Google Scholar]
- Ayer V, Tewodros W, Manoharan A, Skariah S, Luo F, Bessen DE. Tetracycline resistance in group a streptococci: emergence on a global scale and influence on multiple-drug resistance. Antimicrobial agents and chemotherapy. 2007;51:1865–1868. doi: 10.1128/AAC.01341-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aziz RK, Edwards RA, Taylor WW, Low DE, McGeer A, Kotb M. Mosaic prophages with horizontally acquired genes account for the emergence and diversification of the globally disseminated M1T1 clone of Streptococcus pyogenes. Journal of bacteriology. 2005;187:3311–3318. doi: 10.1128/JB.187.10.3311-3318.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aziz RK, Kotb M. Rise and persistence of global M1T1 clone of Streptococcus pyogenes. Emerging infectious diseases. 2008;14:1511–1517. doi: 10.3201/eid1410.071660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bai Q, Zhang W, Yang Y, Tang F, Nguyen X, Liu G, Lu C. Characterization and genome sequencing of a novel bacteriophage infecting Streptococcus agalactiae with high similarity to a phage from Streptococcus pyogenes. Archives of virology. 2013;158:1733–1741. doi: 10.1007/s00705-013-1667-x. [DOI] [PubMed] [Google Scholar]
- Banks DJ, Beres SB, Musser JM. The fundamental contribution of phages to GAS evolution, genome diversification and strain emergence. Trends in microbiology. 2002;10:515–521. doi: 10.1016/s0966-842x(02)02461-7. [DOI] [PubMed] [Google Scholar]
- Banks DJ, Lei B, Musser JM. Prophage induction and expression of prophage-encoded virulence factors in group A Streptococcus serotype M3 strain MGAS315. Infection and immunity. 2003a;71:7079–7086. doi: 10.1128/IAI.71.12.7079-7086.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Banks DJ, Porcella SF, Barbian KD, Beres SB, Philips LE, Voyich JM, DeLeo FR, Martin JM, Somerville GA, Musser JM. Progress toward characterization of the group A Streptococcus metagenome: complete genome sequence of a macrolide-resistant serotype M6 strain. The Journal of infectious diseases. 2004a;190:727–738. doi: 10.1086/422697. [DOI] [PubMed] [Google Scholar]
- Banks DJ, Porcella SF, Barbian KD, Beres SB, Philips LE, Voyich JM, DeLeo FR, Martin JM, Somerville GA, Musser JM. Progress toward characterization of the group A Streptococcus metagenome: complete genome sequence of a macrolide-resistant serotype M6 strain. Journal of Infectious Diseases. 2004b;190:727–738. doi: 10.1086/422697. [DOI] [PubMed] [Google Scholar]
- Banks DJ, Porcella SF, Barbian KD, Martin JM, Musser JM. Structure and distribution of an unusual chimeric genetic element encoding macrolide resistance in phylogenetically diverse clones of group A Streptococcus. Journal of Infectious Diseases. 2003b;188:1898–1908. doi: 10.1086/379897. [DOI] [PubMed] [Google Scholar]
- Barnett TC, Scott JR. Differential recognition of surface proteins in Streptococcus pyogenes by two sortase gene homologs. Journal of bacteriology. 2002;184:2181–2191. doi: 10.1128/JB.184.8.2181-2191.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007;315:1709–1712. doi: 10.1126/science.1138140. [DOI] [PubMed] [Google Scholar]
- Beall B. Atlanta: 2014. http://www.cdc.gov/ncidod/biotech/strep/strepindex.htm. [Google Scholar]
- Beall B, Facklam R, Thompson T. Sequencing emm-specific PCR products for routine and accurate typing of group A streptococci. J. Clin. Microbiol. 1996;34:953–958. doi: 10.1128/jcm.34.4.953-958.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beall B, Gherardi G, Lovgren M, Forwick B, Facklam R, Tyrrell G. Emm and sof gene sequence variation in relation to serological typing of opacity factor positive group A streptococci. Microbiology. 2000;146:1195–1209. doi: 10.1099/00221287-146-5-1195. [DOI] [PubMed] [Google Scholar]
- Belotserkovsky I, Baruch M, Peer A, Dov E, Ravins M, Mishalian I, Persky M, Smith Y, Hanski E. Functional analysis of the quorum-sensing streptococcal invasion locus (sil) PLoS pathogens. 2009;5:e1000651. doi: 10.1371/journal.ppat.1000651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ben Zakour NL, Venturini C, Beatson SA, Walker MJ. Analysis of a Streptococcus pyogenes puerperal sepsis cluster by use of whole-genome sequencing. Journal of clinical microbiology. 2012;50:2224–2228. doi: 10.1128/JCM.00675-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beres SB, Carroll RK, Shea PR, Sitkiewicz I, Martinez-Gutierrez JC, Low DE, McGeer A, Willey BM, Green K, Tyrrell GJ, Goldman TD, Feldgarden M, Birren BW, Fofanov Y, Boos J, Wheaton WD, Honisch C, Musser JM. Molecular complexity of successive bacterial epidemics deconvoluted by comparative pathogenomics. Proceedings of the National Academy of Sciences of the United States of America. 2010;107:4371–4376. doi: 10.1073/pnas.0911295107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beres SB, Musser JM. Contribution of exogenous genetic elements to the group A Streptococcus metagenome. PloS one. 2007;2:e800. doi: 10.1371/journal.pone.0000800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beres SB, Richter EW, Nagiec MJ, Sumby P, Porcella SF, Deleo FR, Musser JM. Molecular genetic anatomy of inter- and intraserotype variation in the human bacterial pathogen group A Streptococcus. Proceedings of the National Academy of Sciences of the United States of America. 2006;103:7059–7064. doi: 10.1073/pnas.0510279103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beres SB, Sesso R, Pinto SW, Hoe NP, Porcella SF, Deleo FR, Musser JM. Genome sequence of a Lancefield group C Streptococcus zooepidemicus strain causing epidemic nephritis: new information about an old disease. PloS one. 2008;3:e3026. doi: 10.1371/journal.pone.0003026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beres SB, Sylva GL, Barbian KD, Lei B, Hoff JS, Mammarella ND, Liu MY, Smoot JC, Porcella SF, Parkins LD, Campbell DS, Smith TM, McCormick JK, Leung DY, Schlievert PM, Musser JM. Genome sequence of a serotype M3 strain of group A Streptococcus: phage- encoded toxins, the high-virulence phenotype, and clone emergence. Proceedings of the National Academy of Sciences of the United States of America. 2002;99:10078–10083. doi: 10.1073/pnas.152298499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beres SB, Sylva GL, Sturdevant DE, Granville CN, Liu MY, Ricklefs SM, Whitney AR, Parkins LD, Hoe NP, Adams GJ, Low DE, DeLeo FR, McGeer A, Musser JM. Genome-wide molecular dissection of serotype M3 group A Streptococcus strains causing two epidemics of invasive infections. Proceedings of the National Academy of Sciences of the United States of America. 2004;101:11833–11838. doi: 10.1073/pnas.0404163101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bessen DE. Population biology of the human restricted pathogen, Streptococcus pyogenes. Infect Genet Evol. 2009;9:581–593. doi: 10.1016/j.meegid.2009.03.00. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bessen DE. Population genomics: an investigative tool for epidemics. The American journal of pathology. 2012;180:1358–1361. doi: 10.1016/j.ajpath.2012.01.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bessen DE, Carapetis JR, Beall B, Katz R, Hibble M, Currie BJ, Collingridge T, Izzo MW, Scaramuzzino DA, Sriprakash KS. Contrasting molecular epidemiology of group A streptococci causing tropical and non-tropical infections of the skin and throat. J. Infect. Dis. 2000a;182:1109–1116. doi: 10.1086/315842. [DOI] [PubMed] [Google Scholar]
- Bessen DE, Carapetis JR, Beall B, Katz R, Hibble M, Currie BJ, Collingridge T, Izzo MW, Scaramuzzino DA, Sriprakash KS. Contrasting molecular epidemiology of group A streptococci causing tropical and nontropical infections of the skin and throat. The Journal of infectious diseases. 2000b;182:1109–1116. doi: 10.1086/315842. [DOI] [PubMed] [Google Scholar]
- Bessen DE, Kalia A. Genomic localization of a T-serotype locus to a recombinatorial zone encoding extracellular matrix-binding proteins in Streptococcus pyogenes. Infect. Immun. 2002;70:1159–1167. doi: 10.1128/IAI.70.3.1159-1167.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bessen DE, Kumar N, Hall GS, Riley DR, Luo F, Lizano S, Ford CN, McShan WM, Nguyen SV, Dunning Hotopp JC, Tettelin H. Whole-genome association study on tissue tropism phenotypes in group A Streptococcus. Journal of bacteriology. 2011;193:6651–6663. doi: 10.1128/JB.05263-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bessen DE, Lizano S. Tissue Tropisms in Group A Streptococcus Infections. Future Microbiol. 2010;5:623–638. doi: 10.2217/fmb.10.28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bessen DE, Manoharan A, Luo F, Wertz JE, Robinson DA. Evolution of transcription regulatory genes is linked to niche specialization in the bacterial pathogen Streptococcus pyogenes. Journal of bacteriology. 2005;187:4163–4172. doi: 10.1128/JB.187.12.4163-4172.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bessen DE, McGregor KF, Whatmore AM. Relationships between emm and multilocus sequence types within a global collection of Streptococcus pyogenes. BMC Microbiol. 2008;8:59. doi: 10.1186/1471-2180-8-59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bessen DE, Sotir CM, Readdy TL, Hollingshead SK. Genetic correlates of throat and skin isolates of group A streptococci. J. Infect. Dis. 1996;173:896–900. doi: 10.1093/infdis/173.4.896. [DOI] [PubMed] [Google Scholar]
- Beyer-Sehlmeyer G, Kreikemeyer B, Horster A, Podbielski A. Analysis of the growth phase-associated transcriptome of Streptococcus pyogenes. International journal of medical microbiology : IJMM. 2005;295:161–177. doi: 10.1016/j.ijmm.2005.02.010. [DOI] [PubMed] [Google Scholar]
- Bisno AL. The concept of rheumatogenic and nonrheumatogenic group A streptococci. In: Read SE, Zabriskie JB, editors. Streptococcal diseases and the immune response. New York: Academic Press; 1980. pp. 789–803. [Google Scholar]
- Bisno AL, Stevens DL. Streptococcus pyogenes. In: Mandell GL, Douglas RG, Dolin R, editors. Principles and Practice of Infectious Diseases. Philadelphia: Churchill Livingstone; 2009. [Google Scholar]
- Bolotin A, Quinquis B, Sorokin A, Ehrlich SD. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology. 2005;151:2551–2561. doi: 10.1099/mic.0.28048-0. [DOI] [PubMed] [Google Scholar]
- Botstein D. A theory of modular evolution for bacteriophages. Ann N Y Acad Sci. 1980;354:484–491. doi: 10.1111/j.1749-6632.1980.tb27987.x. [DOI] [PubMed] [Google Scholar]
- Brandt CM, Spellerberg B, Honscha M, Truong ND, Hoevener B, Lutticken R. Typing of Streptococcus pyogenes strains isolated from throat infections in the region of Aachen, Germany. Infection. 2001;29:163–165. doi: 10.1007/s15010-001-1106-x. [DOI] [PubMed] [Google Scholar]
- Brenciani A, Bacciaglia A, Vignaroli C, Pugnaloni A, Varaldo PE, Giovanetti E. Phim46.1, the main Streptococcus pyogenes element carrying mef(A) and tet(O) genes. Antimicrobial agents and chemotherapy. 2010;54:221–229. doi: 10.1128/AAC.00499-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brenciani A, Tiberi E, Morici E, Oryasin E, Giovanetti E, Varaldo PE. ICESp1116, the genetic element responsible for erm(B)-mediated, inducible resistance to erythromycin in Streptococcus pyogenes. Antimicrobial agents and chemotherapy. 2012;56:6425–6429. doi: 10.1128/AAC.01494-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Broudy TB, Fischetti VA. In vivo lysogenic conversion of Tox(−) Streptococcus pyogenes to Tox(+) with Lysogenic Streptococci or free phage. Infection and immunity. 2003;71:3782–3786. doi: 10.1128/IAI.71.7.3782-3786.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Broudy TB, Pancholi V, Fischetti VA. Induction of lysogenic bacteriophage and phage-associated toxin from group A streptococci during coculture with human pharyngeal cells. Infection and immunity. 2001;69:1440–1443. doi: 10.1128/IAI.69.3.1440-1443.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bucci C, Lavitola A, Salvatore P, Del Giudice L, Massardo DR, Bruni CB, Alifano P. Hypermutation in pathogenic bacteria: frequent phase variation in meningococci is a phenotypic trait of a specialized mutator biotype. Mol Cell. 1999;3:435–445. doi: 10.1016/s1097-2765(00)80471-2. [DOI] [PubMed] [Google Scholar]
- Buchanan JT, Simpson AJ, Aziz RK, Liu GY, Kristian SA, Kotb M, Feramisco J, Nizet V. DNase expression allows the pathogen group A Streptococcus to escape killing in neutrophil extracellular traps. Curr Biol. 2006;16:396–400. doi: 10.1016/j.cub.2005.12.039. [DOI] [PubMed] [Google Scholar]
- Canchaya C, Desiere F, McShan WM, Ferretti JJ, Parkhill J, Brussow H. Genome analysis of an inducible prophage and prophage remnants integrated in the Streptococcus pyogenes strain SF370. Virology. 2002;302:245–258. doi: 10.1006/viro.2002.1570. [DOI] [PubMed] [Google Scholar]
- Carapetis J, Gardiner D, Currie B, Mathews JD. Multiple strains of Streptococcus pyogenes in skin sores of Aboriginal Australians. J. Clin. Microbiol. 1995;33:1471–1472. doi: 10.1128/jcm.33.6.1471-1472.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carapetis JR. Rheumatic heart disease in developing countries. N Engl J Med. 2007;357:439–441. doi: 10.1056/NEJMp078039. [DOI] [PubMed] [Google Scholar]
- Carapetis JR, Steer AC, Mulholland EK, Weber M. The global burden of group A streptococcal diseases. Lancet Infect Dis. 2005;5:685–694. doi: 10.1016/S1473-3099(05)70267-X. [DOI] [PubMed] [Google Scholar]
- Carroll RK, Beres SB, Sitkiewicz I, Peterson L, Matsunami RK, Engler DA, Flores AR, Sumby P, Musser JM. Evolution of diversity in epidemics revealed by analysis of the human bacterial pathogen group A Streptococcus. Epidemics. 2011;3:159–170. doi: 10.1016/j.epidem.2011.07.001. [DOI] [PubMed] [Google Scholar]
- Chang JC, LaSarre B, Jimenez JC, Aggarwal C, Federle MJ. Two group A streptococcal peptide pheromones act through opposing Rgg regulators to control biofilm development. PLoS pathogens. 2011;7:e1002190. doi: 10.1371/journal.ppat.1002190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chmouryguina I, Suvorov A, Ferrieri P, Cleary PP. Conservation of the C5a peptidase genes in group A and B streptococci. Infection and immunity. 1996;64:2387–2390. doi: 10.1128/iai.64.7.2387-2390.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi SC, Rasmussen MD, Hubisz MJ, Gronau I, Stanhope MJ, Siepel A. Replacing and additive horizontal gene transfer in Streptococcus. Molecular biology and evolution. 2012;29:3309–3320. doi: 10.1093/molbev/mss138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cleary PP, LaPenta D, Vessela R, Lam H, Cue D. A globally disseminated M1 subclone of group A streptococci differs from other subclones by 70 kilobases of prophage DNA and capacity for high-frequency intracellular invasion. Infection and immunity. 1998;66:5592–5597. doi: 10.1128/iai.66.11.5592-5597.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cole JN, Barnett TC, Nizet V, Walker MJ. Molecular insight into invasive group A streptococcal disease. Nat Rev Microbiol. 2011;9:724–736. doi: 10.1038/nrmicro2648. [DOI] [PubMed] [Google Scholar]
- Cook LC, LaSarre B, Federle MJ. Interspecies communication among commensal and pathogenic streptococci. mBio. 2013;4:e00382-00313. doi: 10.1128/mBio.00382-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Courtney HS, Pownall HJ. The structure and function of serum opacity factor: a unique streptococcal virulence determinant that targets high-density lipoproteins. Journal of biomedicine & biotechnology. 2010;2010:956071. doi: 10.1155/2010/956071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Creti R, Cardona F, Pataracchia M, Hunolstein CV, Cundari G, Romano A, Orefici G. Characterisation of group A streptococcal (GAS) isolates from children with tic disorders. The Indian journal of medical research. 2004;119(Suppl):174–178. [PubMed] [Google Scholar]
- Cunningham MW. Pathogenesis of group A streptococcal infections. Clin Microbiol Rev. 2000;13:470–511. doi: 10.1128/cmr.13.3.470-511.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davies MR, McMillan DJ, Beiko RG, Barroso V, Geffers R, Sriprakash KS, Chhatwal GS. Virulence profiling of Streptococcus dysgalactiae subspecies equisimilis isolated from infected humans reveals two distinct genetic lineages which do not segregate with their phenotypes or propensity to cause diseases. Clin. Infect. Dis. 2007a;44:1442–1454. doi: 10.1086/516780. [DOI] [PubMed] [Google Scholar]
- Davies MR, McMillan DJ, Van Domselaar GH, Jones MK, Sriprakash KS. Phage 3396 from a Streptococcus dysgalactiae subsp. equisimilis pathovar may have its origins in streptococcus pyogenes. Journal of bacteriology. 2007b;189:2646–2652. doi: 10.1128/JB.01590-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Buhr N, Neumann A, Jerjomiceva N, von Kockritz-Blickwede M, Baums CG. Streptococcus suis DNase SsnA contributes to degradation of neutrophil extracellular traps (NETs) and evasion of NET-mediated antimicrobial activity. Microbiology. 2014;160:385–395. doi: 10.1099/mic.0.072199-0. [DOI] [PubMed] [Google Scholar]
- Deltcheva E, Chylinski K, Sharma CM, Gonzales K, Chao Y, Pirzada ZA, Eckert MR, Vogel J, Charpentier E. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature. 2011;471:602–607. doi: 10.1038/nature09886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Desiere F, McShan WM, van Sinderen D, Ferretti JJ, Brussow H. Comparative genomics reveals close genetic relationships between phages from dairy bacteria and pathogenic Streptococci: evolutionary implications for prophage-host interactions. Virology. 2001;288:325–341. doi: 10.1006/viro.2001.1085. [DOI] [PubMed] [Google Scholar]
- Dey N, McMillan DJ, Yarwood PJ, Joshi RM, Kumar R, Good MF, Sriprakash KS, Vohra H. High diversity of group A Streptococcal emm types in an Indian community: the need to tailor multivalent vaccines. Clinical infectious diseases : an official publication of the Infectious Diseases Society of America. 2005;40:46–51. doi: 10.1086/426443. [DOI] [PubMed] [Google Scholar]
- Dicuonzo G, Gherardi G, Lorino G, Angeletti S, DeCesaris M, Fiscarelli E, Bessen DE, Beall B. Group A streptococcal genotypes from pediatric throat isolates in Rome, Italy. J. Clin. Microbiol. 2001;39:1687–1690. doi: 10.1128/JCM.39.5.1687-1690.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dierksen KP, Inglis M, Tagg JR. High pharyngeal carriage rates of Streptococcus pyogenes in Dunedin school children with a low incidence of rheumatic fever. N Z Med J. 2000;113:496–499. [PubMed] [Google Scholar]
- Eisner A, Leitner E, Feierl G, Kessler HH, Marth E. Prevalence of emm types and antibiotic resistance of group A streptococci in Austria. Diagnostic microbiology and infectious disease. 2006;55:347–350. doi: 10.1016/j.diagmicrobio.2006.01.025. [DOI] [PubMed] [Google Scholar]
- Enright MC, Spratt BG, Kalia A, Cross JH, Bessen DE. Multilocus sequence typing of Streptococcus pyogenes and the relationship between emm-type and clone. Infect. Immun. 2001;69:2416–2427. doi: 10.1128/IAI.69.4.2416-2427.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Espinosa LE, Li ZY, Barreto DG, Jaimes EC, Rodriguez RS, Sakota V, Facklam RR, Beall B. M protein gene type distribution among group A streptococcal clinical isolates recovered in Mexico City, Mexico, from 1991 to 2000, and Durango, Mexico, from 1998 to 1999: Overlap with type distribution within the United States. Journal of clinical microbiology. 2003;41:373–378. doi: 10.1128/JCM.41.1.373-378.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Euler CW, Ryan PA, Martin JM, Fischetti VA. M.SpyI, a DNA methyltransferase encoded on a mefA chimeric element, modifies the genome of Streptococcus pyogenes. Journal of bacteriology. 2007;189:1044–1054. doi: 10.1128/JB.01411-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Facklam R. What happened to the streptococci: Overview of taxonomic and nomenclature changes [Review] Clin Microbiol Rev. 2002;15:613–630. doi: 10.1128/CMR.15.4.613-630.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Falugi F, Zingaretti C, Pinto V, Mariani M, Amodeo L, Manetti AG, Capo S, Musser JM, Orefici G, Margarit I, Telford JL, Grandi G, Mora M. Sequence Variation in Group A Streptococcus Pili and Association of Pilus Backbone Types with Lancefield T Serotypes. The Journal of infectious diseases. 2008;198:1834–1841. doi: 10.1086/593176. [DOI] [PubMed] [Google Scholar]
- Feil EJ, Holmes EC, Bessen DE, Chan M-S, Day NPJ, Enright MC, Goldstein R, Hood D, Kalia A, Moore CE, Zhou J, Spratt BG. Recombination within natural populations of pathogenic bacteria: short-term empirical estimates and long-term phylogenetic consequences. Proc. Natl. Acad. Sci., USA. 2001;98:182–187. doi: 10.1073/pnas.98.1.182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feil EJ, Li BC, Aanensen DM, Hanage WP, Spratt BG. eBURST: Inferring patterns of evolutionary descent among clusters of related bacterial genotypes from multilocus sequence typing data. J. Bacteriol. 2004;186:1518–1530. doi: 10.1128/JB.186.5.1518-1530.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferretti JJ, McShan WM, Ajdic D, Savic DJ, Savic G, Lyon K, Primeaux C, Sezate S, Suvorov AN, Kenton S, Lai HS, Lin SP, Qian Y, Jia HG, Najar FZ, Ren Q, Zhu H, Song L, White J, Yuan X, Clifton SW, Roe BA, McLaughlin R. Complete genome sequence of an M1 strain of Streptococcus pyogenes. Proc. Natl. Acad. Sci. U.S.A. 2001;98:4658–4663. doi: 10.1073/pnas.071559398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fiorentino TR, Beall B, Mshar P, Bessen DE. A genetic-based evaluation of principal tissue reservoir for group A streptococci isolated from normally sterile sites. J.Infect.Dis. 1997;176:177–182. doi: 10.1086/514020. [DOI] [PubMed] [Google Scholar]
- Fittipaldi N, Beres SB, Olsen RJ, Kapur V, Shea PR, Watkins ME, Cantu CC, Laucirica DR, Jenkins L, Flores AR, Lovgren M, Ardanuy C, Linares J, Low DE, Tyrrell GJ, Musser JM. Full-genome dissection of an epidemic of severe invasive disease caused by a hypervirulent, recently emerged clone of group A Streptococcus. The American journal of pathology. 2012a;180:1522–1534. doi: 10.1016/j.ajpath.2011.12.037. [DOI] [PubMed] [Google Scholar]
- Fittipaldi N, Olsen RJ, Beres SB, Van Beneden C, Musser JM. Genomic analysis of emm59 group A Streptococcus invasive strains, United States. Emerging infectious diseases. 2012b;18:650–652. doi: 10.3201/eid1804.111803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flores AR, Sahasrabhojane P, Saldana M, Galloway-Pena J, Olsen RJ, Musser JM, Shelburne SA. Molecular characterization of an invasive phenotype of group A Streptococcus arising during human infection using whole genome sequencing of multiple isolates from the same patient. The Journal of infectious diseases. 2014;209:1520–1523. doi: 10.1093/infdis/jit674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fouts DE. Phage_Finder: automated identification and classification of prophage regions in complete bacterial genome sequences. Nucleic acids research. 2006;34:5839–5851. doi: 10.1093/nar/gkl732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Francisco AP, Vaz C, Monteiro PT, Melo-Cristino J, Ramirez M, Carrico JA. PHYLOViZ: phylogenetic inference and data visualization for sequence based typing methods. BMC bioinformatics. 2012;13:87. doi: 10.1186/1471-2105-13-87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franken C, Haase G, Brandt C, Weber-Heynemann J, Martin S, Lammler C, Podbielski A, Lutticken R, Spellerberg B. Horizontal gene transfer and host specificity of beta-haemolytic streptococci: the role of a putative composite transposon containing scpB and lmb. Molecular microbiology. 2001;41:925–935. doi: 10.1046/j.1365-2958.2001.02563.x. [DOI] [PubMed] [Google Scholar]
- Frobisher M, Brown JH. Transmissible toxicogenicity of streptococci. Bulletin of Johns Hopkins Hospital. 1927;41:167–173. [Google Scholar]
- Garcia AF, Abe LM, Erdem G, Cortez CL, Kurahara D, Yamaga K. An insert in the covS gene distinguishes a pharyngeal and a blood isolate of Streptococcus pyogenes found in the same individual. Microbiology. 2010;156:3085–3095. doi: 10.1099/mic.0.042614-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giovanetti E, Brenciani A, Tiberi E, Bacciaglia A, Varaldo PE. ICESp2905, the erm(TR)-tet(O) element of Streptococcus pyogenes, is formed by two independent integrative and conjugative elements. Antimicrobial agents and chemotherapy. 2012;56:591–594. doi: 10.1128/AAC.05352-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Glaser P, Rusniok C, Buchrieser C, Chevalier F, Frangeul L, Msadek T, Zouine M, Couve E, Lalioui L, Poyart C, Trieu-Cuot P, Kunst F. Genome sequence of Streptococcus agalactiae a pathogen causing invasive neonatal disease. Molecular microbiology. 2002;45:1499–1513. doi: 10.1046/j.1365-2958.2002.03126.x. [DOI] [PubMed] [Google Scholar]
- Gottesman S, Storz G. Bacterial small RNA regulators: versatile roles and rapidly evolving variations. Cold Spring Harbor perspectives in biology. 2011;3:a003798. doi: 10.1101/cshperspect.a003798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Green NM, Zhang S, Porcella SF, Nagiec MJ, Barbian KD, Beres SB, LeFebvre RB, Musser JM. Genome sequence of a serotype M28 strain of group a streptococcus: potential new insights into puerperal sepsis and bacterial disease specificity. Journal of Infectious Diseases. 2005;192:760–770. doi: 10.1086/430618. [DOI] [PubMed] [Google Scholar]
- Grundmann H, Hori S, Tanner G. Determining confidence intervals when measuring genetic diversity and the discriminatory abilities of typing methods for microorganisms. Journal of clinical microbiology. 2001;39:4190–4192. doi: 10.1128/JCM.39.11.4190-4192.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haanes EJ, Heath DG, Cleary PP. Architecture of the vir regulons of group A streptococci parallel opacity factor phenotype and M protein class. J.Bacteriol. 1992;171:4967–4976. doi: 10.1128/jb.174.15.4967-4976.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanage WP, Fraser C, Spratt BG. The impact of homologous recombination on the generation of diversity in bacteria. J Theor Biol. 2006;239:210–219. doi: 10.1016/j.jtbi.2005.08.035. [DOI] [PubMed] [Google Scholar]
- Hanski E, Caparon M. Protein F, a fibronectin-binding protein, is an adhesin of the group A streptococcus Streptococcus pyogenes. Proc.Natl.Acad.Sci.U.S.A. 1992;89:6172–6176. doi: 10.1073/pnas.89.13.6172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haukness HA, Tanz RR, Thomson RB, Jr, Pierry DK, Kaplan EL, Beall B, Johnson D, Hoe NP, Musser JM, Shulman ST. The heterogeneity of endemic community pediatric group a streptococcal pharyngeal isolates and their relationship to invasive isolates. The Journal of infectious diseases. 2002;185:915–920. doi: 10.1086/339407. [DOI] [PubMed] [Google Scholar]
- Hendrix RW, Roberts JW, Stahl FW, Weisberg RA, editors. Lambda II. Cold Spring Harbor, NY: Cold Spring Harbor Press; 1983. [Google Scholar]
- Hidalgo-Grass C, Ravins M, Dan-Goor M, Jaffe J, Moses AE, Hanski E. A locus of group A Streptococcus involved in invasive disease and DNA transfer. Molecular microbiology. 2002;46:87–99. doi: 10.1046/j.1365-2958.2002.03127.x. [DOI] [PubMed] [Google Scholar]
- Holden MT, Heather Z, Paillot R, Steward KF, Webb K, Ainslie F, Jourdan T, Bason NC, Holroyd NE, Mungall K, Quail MA, Sanders M, Simmonds M, Willey D, Brooks K, Aanensen DM, Spratt BG, Jolley KA, Maiden MC, Kehoe M, Chanter N, Bentley SD, Robinson C, Maskell DJ, Parkhill J, Waller AS. Genomic evidence for the evolution of Streptococcus equi: host restriction, increased virulence, and genetic exchange with human pathogens. PLoS pathogens. 2009;5:e1000346. doi: 10.1371/journal.ppat.1000346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holden MT, Scott A, Cherevach I, Chillingworth T, Churcher C, Cronin A, Dowd L, Feltwell T, Hamlin N, Holroyd S, Jagels K, Moule S, Mungall K, Quail MA, Price C, Rabbinowitsch E, Sharp S, Skelton J, Whitehead S, Barrell BG, Kehoe M, Parkhill J. Complete genome of acute rheumatic fever-associated serotype M5 Streptococcus pyogenes strain manfredo. Journal of bacteriology. 2007;189:1473–1477. doi: 10.1128/JB.01227-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hollingshead SK, Arnold J, Readdy TL, Bessen DE. Molecular evolution of a multigene family in group A streptococci. Molecular biology and evolution. 1994;11:208–219. doi: 10.1093/oxfordjournals.molbev.a040103. [DOI] [PubMed] [Google Scholar]
- Hollingshead SK, Readdy TL, Yung DL, Bessen DE. Structural heterogeneity of the emm gene cluster in group A streptococci. Molecular microbiology. 1993;8:707–717. doi: 10.1111/j.1365-2958.1993.tb01614.x. [DOI] [PubMed] [Google Scholar]
- Hsieh YC, Huang YC. Scarlet fever outbreak in Hong Kong, 2011. Journal of microbiology, immunology, and infection = Wei mian yu gan ran za zhi. 2011;44:409–411. doi: 10.1016/j.jmii.2011.07.003. [DOI] [PubMed] [Google Scholar]
- Huson DH, Bryant D. Application of phylogenetic networks in evolutionary studies. Molecular biology and evolution. 2006;23:254–267. doi: 10.1093/molbev/msj030. [DOI] [PubMed] [Google Scholar]
- Ikebe T, Ato M, Matsumura T, Hasegawa H, Sata T, Kobayashi K, Watanabe H. Highly frequent mutations in negative regulators of multiple virulence genes in group A streptococcal toxic shock syndrome isolates. PLoS pathogens. 2010;6:e1000832. doi: 10.1371/journal.ppat.1000832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jensen A, Kilian M. Delineation of Streptococcus dysgalactiae, its subspecies, and its clinical and phylogenetic relationship to Streptococcus pyogenes. Journal of clinical microbiology. 2012;50:113–126. doi: 10.1128/JCM.05900-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson DR, Kaplan EL, VanGheem A, Facklam RR, Beall B. Characterization of group A streptococci (Streptococcus pyogenes): correlation of M-protein and emm-gene type with T-protein agglutination pattern and serum opacity factor. J Med Microbiol. 2006;55:157–164. doi: 10.1099/jmm.0.46224-0. [DOI] [PubMed] [Google Scholar]
- Johnson DR, Stevens DL, Kaplan EL. Epidemiological analysis of group A streptococcal serotypes associated with severe systemic infections, rheumatic fever, or uncomplicated pharyngitis. J. Infect. Dis. 1992;166:374–382. doi: 10.1093/infdis/166.2.374. [DOI] [PubMed] [Google Scholar]
- Kalia A, Bessen DE. Natural Selection and Evolution of Streptococcal Virulence Genes Involved in Tissue-Specific Adaptations. J. Bacteriol. 2004;186:110–121. doi: 10.1128/JB.186.1.110-121.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalia A, Spratt BG, Enright MC, Bessen DE. Influence of recombination and niche separation on the population genetic structure of the pathogen Streptococcus pyogenes. Infect. Immun. 2002;70:1971–1983. doi: 10.1128/IAI.70.4.1971-1983.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaplan EL. The group A streptococcal upper respiratory tract carrier state: An enigma. J. Pediatr. 1980;97:337–345. doi: 10.1016/s0022-3476(80)80178-8. [DOI] [PubMed] [Google Scholar]
- Kiska DL, Thiede B, Caracciolo J, Jordan M, Johnson D, Kaplan EL, Gruninger RP, Lohr JA, Gilligan PH, Denny FW., Jr Invasive group A streptococcal infections in North Carolina: epidemiology, clinical features, and genetic and serotype analysis of causative organisms. The Journal of infectious diseases. 1997;176:992–1000. doi: 10.1086/516540. [DOI] [PubMed] [Google Scholar]
- Kline JB, Collins CM. Analysis of the superantigenic activity of mutant and allelic forms of streptococcal pyrogenic exotoxin A. Infect.Immun. 1996;64:861–869. doi: 10.1128/iai.64.3.861-869.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koller T, Manetti AG, Kreikemeyer B, Lembke C, Margarit I, Grandi G, Podbielski A. Typing of the pilus-protein-encoding FCT region and biofilm formation as novel parameters in epidemiological investigations of Streptococcus pyogenes isolates from various infection sites. J Med Microbiol. 2010;59:442–452. doi: 10.1099/jmm.0.013581-0. [DOI] [PubMed] [Google Scholar]
- Koonin EV, Makarova KS, Aravind L. Horizontal gene transfer in prokaryotes: quantification and classification. Annual review of microbiology. 2001;55:709–742. doi: 10.1146/annurev.micro.55.1.709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kratovac Z, Manoharan A, Luo F, Lizano S, Bessen DE. Population genetics and linkage analysis of loci within the FCT region of Streptococcus pyogenes. J. Bacteriol. 2007;189:1299–1310. doi: 10.1128/JB.01301-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kreikemeyer B, Nakata M, Koller T, Hildisch H, Kourakos V, Standar K, Kawabata S, Glocker MO, Podbielski A. The Streptococcus pyogenes serotype M49 Nra-Ralp3 transcriptional regulatory network and its control on virulence factor expression from the novel ERES pathogenicity region. Infection and immunity. 2007;75:5698–5710. doi: 10.1128/IAI.00175-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lancefield RC. Current knowledge of the type specific M antigens of group A streptococci. J. Immunol. 1962;89:307–313. [PubMed] [Google Scholar]
- Lau EH, Nishiura H, Cowling BJ, Ip DK, Wu JT. Scarlet fever outbreak, Hong Kong, 2011. Emerging infectious diseases. 2012;18:1700–1702. doi: 10.3201/eid1810.120062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LeClerc JE, Li B, Payne WL, Cebula TA. High mutation frequencies among Escherichia coli and Salmonella pathogens. Science. 1996;274:1208–1211. doi: 10.1126/science.274.5290.1208. [DOI] [PubMed] [Google Scholar]
- Lefebure T, Richards VP, Lang P, Pavinski-Bitar P, Stanhope MJ. Gene Repertoire Evolution of Streptococcus pyogenes Inferred from Phylogenomic Analysis with Streptococcus canis and Streptococcus dysgalactiae. PloS one. 2012;7:e37607. doi: 10.1371/journal.pone.0037607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lefebure T, Stanhope MJ. Evolution of the core and pan-genome of Streptococcus: positive selection, recombination, and genome composition. Genome biology. 2007;8:R71. doi: 10.1186/gb-2007-8-5-r71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lizano S, Luo F, Bessen DE. Role of streptococcal T-antigens in superficial skin infection. J. Bacteriol. 2007;189:1426–1434. doi: 10.1128/JB.01179-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lorino G, Gherardi G, Angeletti S, De Cesaris M, Graziano N, Maringhini S, Merlino F, Di Bernardo F, Dicuonzo G. Molecular characterisation and clonal analysis of group A streptococci causing pharyngitis among paediatric patients in Palermo, Italy. Clinical microbiology and infection : the official publication of the European Society of Clinical Microbiology and Infectious Diseases. 2006;12:189–192. doi: 10.1111/j.1469-0691.2005.01329.x. [DOI] [PubMed] [Google Scholar]
- Luk EY, Lo JY, Li AZ, Lau MC, Cheung TK, Wong AY, Wong MM, Wong CW, Chuang SK, Tsang T. Scarlet fever epidemic, Hong Kong 2011. Emerging infectious diseases. 2012;18:1658–1661. doi: 10.3201/eid1810.111900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma X, Kikuta H, Ishiguro N, Yoshioka M, Ebihara T, Murai T, Kobayashi I, Kobayashi K. Association of the prtF1 gene (encoding fibronectin-binding protein F1) and the sic gene (encoding the streptococcal inhibitor of complement) with emm types of group A streptococci isolated from Japanese children with pharyngitis. Journal of clinical microbiology. 2002;40:3835–3837. doi: 10.1128/JCM.40.10.3835-3837.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maamary PG, Ben Zakour NL, Cole JN, Hollands A, Aziz RK, Barnett TC, Cork AJ, Henningham A, Sanderson-Smith M, McArthur JD, Venturini C, Gillen CM, Kirk JK, Johnson DR, Taylor WL, Kaplan EL, Kotb M, Nizet V, Beatson SA, Walker MJ. Tracing the evolutionary history of the pandemic group A streptococcal M1T1 clone. FASEB journal : official publication of the Federation of American Societies for Experimental Biology. 2012;26:4675–4684. doi: 10.1096/fj.12-212142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Majeed HA, Yousof AM, Rotta J, Havlickpva H, Bahar G, Bahbahani K. Group A streptococcal strains in Kuwait: a nine-year prospective study of prevalence and associations. The Pediatric infectious disease journal. 1992;11:295–300. discussion 300-293. [PubMed] [Google Scholar]
- Manetti AG, Zingaretti C, Falugi F, Capo S, Bombaci M, Bagnoli F, Gambellini G, Bensi G, Mora M, Edwards AM, Musser JM, Graviss EA, Telford JL, Grandi G, Margarit I. Streptococcus pyogenes pili promote pharyngeal cell adhesion and biofilm formation. Molecular microbiology. 2007;64:968–983. doi: 10.1111/j.1365-2958.2007.05704.x. [DOI] [PubMed] [Google Scholar]
- Marks LR, Mashburn-Warren L, Federle MJ, Hakansson AP. Streptococcus pyogenes Biofilm Growth In Vitro and In Vivo and Its Role in Colonization, Virulence, and Genetic Exchange. The Journal of infectious diseases. 2014 doi: 10.1093/infdis/jiu058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marri PR, Hao W, Golding GB. Gene gain and gene loss in streptococcus: is it driven by habitat? Molecular biology and evolution. 2006;23:2379–2391. doi: 10.1093/molbev/msl115. [DOI] [PubMed] [Google Scholar]
- Martin B, Quentin Y, Fichant G, Claverys JP. Independent evolution of competence regulatory cascades in streptococci? Trends in microbiology. 2006;14:339–345. doi: 10.1016/j.tim.2006.06.007. [DOI] [PubMed] [Google Scholar]
- Mashburn-Warren L, Morrison DA, Federle MJ. The cryptic competence pathway in Streptococcus pyogenes is controlled by a peptide pheromone. Journal of bacteriology. 2012;194:4589–4600. doi: 10.1128/JB.00830-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masuno K, Okada R, Zhang Y, Isaka M, Tatsuno I, Shibata S, Hasegawa T. Simultaneous isolation of emm89-type Streptococcus pyogenes strains with a wild-type or mutated covS gene from a single streptococcal toxic shock syndrome patient. J Med Microbiol. 2014;63:504–507. doi: 10.1099/jmm.0.070300-0. [DOI] [PubMed] [Google Scholar]
- McDonald MI, Towers RJ, Fagan P, Carapetis JR, Currie BJ. Molecular typing of Streptococcus pyogenes from remote Aboriginal communities where rheumatic fever is common and pyoderma is the predominant streptococcal infection. Epidemiology and infection. 2007;135:1398–1405. doi: 10.1017/S0950268807008023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McDonald RR, Golding GR, Irvine J, Graham MR, Tyler S, Mulvey MR, Levett PN. Draft Genome Sequence of Streptococcus pyogenes Strain 06BA18369, a Human Pathogen Associated with Skin and Soft Tissue Infections in Northern Canada. Genome announcements. 2013;1:e00389–e00313. doi: 10.1128/genomeA.00387-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McGregor KF, Spratt BG, Kalia A, Bennett A, Bilek N, Beall B, Bessen DE. Multilocus sequence typing of Streptococcus pyogenes representing most known emm types and distinctions among subpopulation genetic structures. Journal of bacteriology. 2004;186:4285–4294. doi: 10.1128/JB.186.13.4285-4294.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McMillan DJ, Bessen DE, Pinho M, Ford C, Hall GS, Melo-Cristino J, Ramirez M. Population genetics of Streptococcus dysgalactiae subspecies equisimilis reveals widely dispersed clones and extensive recombination. PloS one. 2010;5:e11741. doi: 10.1371/journal.pone.0011741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McMillan DJ, Dreze PA, Vu T, Bessen DE, Guglielmini J, Steer AC, Carapetis JR, Van Melderen L, Sriprakash KS, Smeesters PR. Updated model of group A Streptococcus M proteins based on a comprehensive worldwide study. Clinical microbiology and infection : the official publication of the European Society of Clinical Microbiology and Infectious Diseases. 2013;19:E222–E229. doi: 10.1111/1469-0691.12134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McShan WM, Ferretti JJ, Karasawa T, Suvorov AN, Lin S, Qin B, Jia H, Kenton S, Najar F, Wu H, Scott J, Roe BA, Savic DJ. Genome sequence of a nephritogenic and highly transformable M49 strain of Streptococcus pyogenes. Journal of bacteriology. 2008;190:7773–7785. doi: 10.1128/JB.00672-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mejia LM, Stockbauer KE, Pan X, Cravioto A, Musser JM. Characterization of group A Streptococcus strains recovered from Mexican children with pharyngitis by automated DNA sequencing of virulence-related genes: unexpectedly large variation in the gene (sic) encoding a complement-inhibiting protein. Journal of clinical microbiology. 1997;35:3220–3224. doi: 10.1128/jcm.35.12.3220-3224.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyoshi-Akiyama T, Watanabe S, Kirikae T. Complete genome sequence of Streptococcus pyogenes M1 476, isolated from a patient with streptococcal toxic shock syndrome. Journal of bacteriology. 2012;194:5466. doi: 10.1128/JB.01265-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mojica FJ, Diez-Villasenor C, Garcia-Martinez J, Soria E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. Journal of molecular evolution. 2005;60:174–182. doi: 10.1007/s00239-004-0046-3. [DOI] [PubMed] [Google Scholar]
- Mora M, Bensi G, Capo S, Falugi F, Zingaretti C, Manetti AG, Maggi T, Taddei AR, Grandi G, Telford JL. Group A Streptococcus produce pilus-like structures containing protective antigens and Lancefield T antigens. Proceedings of the National Academy of Sciences of the United States of America. 2005;102:15641–15646. doi: 10.1073/pnas.0507808102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mzoughi R, Bouallegue O, Selmi H, Ben Said H, Essoussi AS, Jeddi M. Group A streptococci in children with acute pharyngitis in Sousse, Tunisia. Eastern Mediterranean health journal = La revue de sante de la Mediterranee orientale = al-Majallah alsihhiyah li-sharq al-mutawassit. 2004;10:488–493. [PubMed] [Google Scholar]
- Nakagawa I, Kurokawa K, Yamashita A, Nakata M, Tomiyasu Y, Okahashi N, Kawabata S, Yamazaki K, Shiba T, Yasunaga T, Hayashi H, Hattori M, Hamada S. Genome sequence of an M3 strain of Streptococcus pyogenes reveals a large-scale genomic rearrangement in invasive strains and new insights into phage evolution. Genome Res. 2003;13:1042–1055. doi: 10.1101/gr.1096703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nasser W, Beres SB, Olsen RJ, Dean MA, Rice KA, Long SW, Kristinsson KG, Gottfredsson M, Vuopio J, Raisanen K, Caugant DA, Steinbakk M, Low DE, McGeer A, Darenberg J, Henriques-Normark B, Van Beneden CA, Hoffmann S, Musser JM. Evolutionary pathway to increased virulence and epidemic group A Streptococcus disease derived from 3,615 genome sequences. Proceedings of the National Academy of Sciences of the United States of America. 2014;111:E1768–E1776. doi: 10.1073/pnas.1403138111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- NCBI. Genome Assembly and Annotation report. 2014 In. [Google Scholar]
- Nelson DC, Garbe J, Collin M. Cysteine proteinase SpeB from Streptococcus pyogenes - a potent modifier of immunologically important host and bacterial proteins. Biol Chem. 2011;392:1077–1088. doi: 10.1515/BC.2011.208. [DOI] [PubMed] [Google Scholar]
- Nguyen SV, McShan WM. Chromosomal Islands of Streptococcus pyogenes and related streptococci: Molecular Switches for Survival and Virulence. Front Microbiol. 2014 doi: 10.3389/fcimb.2014.00109. Submitted: [DOI] [PMC free article] [PubMed] [Google Scholar]
- Novick RP, Christie GE, Penades JR. The phage-related chromosomal islands of Gram-positive bacteria. Nat Rev Microbiol. 2010;8:541–551. doi: 10.1038/nrmicro2393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nozawa T, Furukawa N, Aikawa C, Watanabe T, Haobam B, Kurokawa K, Maruyama F, Nakagawa I. CRISPR inhibition of prophage acquisition in Streptococcus pyogenes. PloS one. 2011;6:e19543. doi: 10.1371/journal.pone.0019543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oliver A, Canton R, Campo P, Baquero F, Blazquez J. High frequency of hypermutable Pseudomonas aeruginosa in cystic fibrosis lung infection. Science. 2000;288:1251–1254. doi: 10.1126/science.288.5469.1251. [DOI] [PubMed] [Google Scholar]
- Olsen RJ, Laucirica DR, Watkins ME, Feske ML, Garcia-Bustillos JR, Vu C, Cantu C, Shelburne SA, 3rd, Fittipaldi N, Kumaraswami M, Shea PR, Flores AR, Beres SB, Lovgren M, Tyrrell GJ, Efstratiou A, Low DE, Van Beneden CA, Musser JM. Polymorphisms in regulator of protease B (RopB) alter disease phenotype and strain virulence of serotype M3 group A Streptococcus. The Journal of infectious diseases. 2012;205:1719–1729. doi: 10.1093/infdis/jir825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Panchaud A, Guy L, Collyn F, Haenni M, Nakata M, Podbielski A, Moreillon P, Roten CA. M-protein and other intrinsic virulence factors of Streptococcus pyogenes are encoded on an ancient pathogenicity island. BMC genomics. 2009;10:198. doi: 10.1186/1471-2164-10-198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patenge N, Billion A, Raasch P, Normann J, Wisniewska-Kucper A, Retey J, Boisguerin V, Hartsch T, Hain T, Kreikemeyer B. Identification of novel growth phase- and media-dependent small non-coding RNAs in Streptococcus pyogenes M49 using intergenic tiling arrays. BMC genomics. 2012;13:550. doi: 10.1186/1471-2164-13-550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perez N, Trevino J, Liu Z, Ho SC, Babitzke P, Sumby P. A genome-wide analysis of small regulatory RNAs in the human pathogen group A Streptococcus. PloS one. 2009;4:e7668. doi: 10.1371/journal.pone.0007668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Persson J, Beall B, Linse S, Lindahl G. Extreme sequence divergence but conserved ligand-binding specificity in Streptococcus pyogenes M protein. PLoS pathogens. 2006;2:e47. doi: 10.1371/journal.ppat.0020047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Plainvert C, Dinis M, Ravins M, Hanski E, Touak G, Dmytruk N, Fouet A, Poyart C. Molecular epidemiology of sil locus in clinical Streptococcus pyogenes strains. Journal of clinical microbiology. 2014;52:2003–2010. doi: 10.1128/JCM.00290-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Podbielski A. Three different types of organization of the vir regulon in group A streptococci. Molecular & general genetics : MGG. 1993;237:287–300. doi: 10.1007/BF00282810. [DOI] [PubMed] [Google Scholar]
- Podbielski A, Woischnik M, Leonard BAB, Schmidt KH. Characterization of nra, a global negative regulator gene in group A streptococci. Molecular microbiology. 1999;31:1051–1064. doi: 10.1046/j.1365-2958.1999.01241.x. [DOI] [PubMed] [Google Scholar]
- Port GC, Paluscio E, Caparon MG. Complete Genome Sequence of emm Type 14 Streptococcus pyogenes Strain HSC5. Genome announcements. 2013;1:e00612–e00613. doi: 10.1128/genomeA.00612-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richards VP, Palmer SR, Pavinski Bitar PD, Qin X, Weinstock GM, Highlander SK, Town CD, Burne RA, Stanhope MJ. Phylogenomics and the dynamic genome evolution of the genus Streptococcus. Genome biology and evolution. 2014;6:741–753. doi: 10.1093/gbe/evu048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richards VP, Zadoks RN, Pavinski Bitar PD, Lefebure T, Lang P, Werner B, Tikofsky L, Moroni P, Stanhope MJ. Genome characterization and population genetic structure of the zoonotic pathogen, Streptococcus canis. BMC Microbiol. 2012;12:293. doi: 10.1186/1471-2180-12-293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richter SS, Heilmann KP, Beekmann SE, Miller NJ, Miller AL, Rice CL, Doern CD, Reid SD, Doern GV. Macrolide-resistant Streptococcus pyogenes in the United States, 2002–2003. Clinical infectious diseases : an official publication of the Infectious Diseases Society of America. 2005;41:599–608. doi: 10.1086/432473. [DOI] [PubMed] [Google Scholar]
- Riddle DJ, Bessen DE, Caparon MG. Variation in Streptococcus pyogenes NAD+ glycohydrolase is associated with tissue tropism. Journal of bacteriology. 2010;192:3735–3746. doi: 10.1128/JB.00234-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riley DR, Angiuoli SV, Crabtree J, Dunning Hotopp JC, Tettelin H. Using Sybil for interactive comparative genomics of microbes on the web. Bioinformatics. 2012;28:160–166. doi: 10.1093/bioinformatics/btr652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts AL, Connolly KL, Kirse DJ, Evans AK, Poehling KA, Peters TR, Reid SD. Detection of group A Streptococcus in tonsils from pediatric patients reveals high rate of asymptomatic streptococcal carriage. BMC pediatrics. 2012;12:3. doi: 10.1186/1471-2431-12-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson D, Sutcliffe J, Tewodros W, Manoharan A, Bessen D. Evolution and global dissemination of macrolide resistant group A streptococci. Antimicrob. Agents & Chemo. 2006;50:2903–2911. doi: 10.1128/AAC.00325-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rogers S, Commons R, Danchin MH, Selvaraj G, Kelpie L, Curtis N, Robins-Browne R, Carapetis JR. Strain prevalence, rather than innate virulence potential, is the major factor responsible for an increase in serious group A streptococcus infections. The Journal of infectious diseases. 2007;195:1625–1633. doi: 10.1086/513875. [DOI] [PubMed] [Google Scholar]
- Sagar V, Bakshi DK, Nandi S, Ganguly NK, Kumar R, Chakraborti A. Molecular heterogeneity among north Indian isolates of Group A Streptococcus. Letters in applied microbiology. 2004;39:84–88. doi: 10.1111/j.1472-765X.2004.01545.x. [DOI] [PubMed] [Google Scholar]
- Sahl JW, Matalka MN, Rasko DA. Phylomark, a tool to identify conserved phylogenetic markers from whole-genome alignments. Applied and environmental microbiology. 2012;78:4884–4892. doi: 10.1128/AEM.00929-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakota V, Fry AM, Lietman TM, Facklam RR, Li ZY, Beall B. Genetically diverse group A streptococci from children in Far-Western Nepal share high genetic relatedness with isolates from other countries. Journal of clinical microbiology. 2006;44:2160–2166. doi: 10.1128/JCM.02456-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanderson-Smith M, De Oliveira DM, Guglielmini J, McMillan DJ, Vu T, Holien JK, Henningham A, Steer AC, Bessen DE, Dale JB, Curtis N, Beall BW, Walker MJ, Parker MW, Carapetis JR, Van Melderen L, Sriprakash KS, Smeesters PR. A systematic and functional classification of Streptococcus pyogenes that serves as a new tool for molecular typing and vaccine development. The Journal of infectious diseases. 2014 doi: 10.1093/infdis/jiu260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schneewind O, Jones KF, Fischetti VA. Sequence and structural characterization of the trypsin-resistant T6 surface protein of group A streptococci. J. Bacteriol. 1990;172:3310–3317. doi: 10.1128/jb.172.6.3310-3317.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scott J, Nguyen S, King CJ, Hendrickson C, McShan WM. Mutator Phenotype Prophages in the Genome Strains of Streptococcus pyogenes: Control by Growth State and by a Cryptic Prophage-Encoded Promoter. Front Microbiol. 2012;3 doi: 10.3389/fmicb.2012.00317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scott J, Thompson-Mayberry P, Lahmamsi S, King CJ, McShan WM. Phage-associated mutator phenotype in group A Streptococcus. Journal of bacteriology. 2008;190:6290–6301. doi: 10.1128/JB.01569-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shea PR, Beres SB, Flores AR, Ewbank AL, Gonzalez-Lugo JH, Martagon-Rosado AJ, Martinez-Gutierrez JC, Rehman HA, Serrano-Gonzalez M, Fittipaldi N, Ayers SD, Webb P, Willey BM, Low DE, Musser JM. Distinct signatures of diversifying selection revealed by genome analysis of respiratory tract and invasive bacterial populations. Proceedings of the National Academy of Sciences of the United States of America. 2011;108:5039–5044. doi: 10.1073/pnas.1016282108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimomura Y, Okumura K, Murayama SY, Yagi J, Ubukata K, Kirikae T, Miyoshi-Akiyama T. Complete genome sequencing and analysis of a Lancefield group G Streptococcus dysgalactiae subsp. equisimilis strain causing streptococcal toxic shock syndrome (STSS) BMC genomics. 2011;12:17. doi: 10.1186/1471-2164-12-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shulman ST, Tanz RR, Dale JB, Beall B, Kabat W, Kabat K, Cederlund E, Patel D, Rippe J, Li Z, Sakota V. Seven-year surveillance of north american pediatric group a streptococcal pharyngitis isolates. Clinical infectious diseases : an official publication of the Infectious Diseases Society of America. 2009;49:78–84. doi: 10.1086/599344. [DOI] [PubMed] [Google Scholar]
- Shulman ST, Tanz RR, Kabat W, Kabat K, Cederlund E, Patel D, Li Z, Sakota V, Dale JB, Beall B. Group A streptococcal pharyngitis serotype surveillance in North America, 2000–2002. Clinical infectious diseases : an official publication of the Infectious Diseases Society of America. 2004;39:325–332. doi: 10.1086/421949. [DOI] [PubMed] [Google Scholar]
- Simpson WJ, Musser JM, Cleary PP. Evidence consistent with horizontal transfer of the gene (emm12) encoding serotype M12 protein between group A and group G pathogenic streptococci. Infect. Immun. 1992;60:1890–1893. doi: 10.1128/iai.60.5.1890-1893.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sitkiewicz I, Green NM, Guo N, Mereghetti L, Musser JM. Lateral gene transfer of streptococcal ICE element RD2 (region of difference 2) encoding secreted proteins. BMC Microbiol. 2011;11:65. doi: 10.1186/1471-2180-11-65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smeesters PR, Vergison A, Campos D, de Aguiar E, Deyi VY, Van Melderen L. Differences between Belgian and Brazilian Group A Streptococcus Epidemiologic Landscape. PloS one. 2006;1:e10. doi: 10.1371/journal.pone.0000010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smoot JC, Barbian KD, Van Gompel JJ, Smoot LM, Chaussee MS, Sylva GL, Sturdevant DE, Ricklefs SM, Porcella SF, Parkins LD, Beres SB, Campbell DS, Smith TM, Zhang Q, Kapur V, Daly JA, Veasy LG, Musser JM. Genome sequence and comparative microarray analysis of serotype M18 group A Streptococcus strains associated with acute rheumatic fever outbreaks. Proceedings of the National Academy of Sciences of the United States of America. 2002;99:4668–4673. doi: 10.1073/pnas.062526099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stalhammar-Carlemalm M, Areschoug T, Larsson C, Lindahl G. The R28 protein of Streptococcus pyogenes is related to several group B streptococcal surface proteins, confers protective immunity and promotes binding to human epithelial cells. Molecular microbiology. 1999;33:208–219. doi: 10.1046/j.1365-2958.1999.01470.x. [DOI] [PubMed] [Google Scholar]
- Starr CR, Engleberg NC. Role of hyaluronidase in subcutaneous spread and growth of group A streptococcus. Infection and immunity. 2006;74:40–48. doi: 10.1128/IAI.74.1.40-48.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steer AC, Jenney AW, Kado J, Good MF, Batzloff M, Magor G, Ritika R, Mulholland KE, Carapetis JR. Prospective surveillance of streptococcal sore throat in a tropical country. The Pediatric infectious disease journal. 2009a;28:477–482. doi: 10.1097/INF.0b013e318194b2af. [DOI] [PubMed] [Google Scholar]
- Steer AC, Law I, Matatolu L, Beall BW, Carapetis JR. Global emm type distribution of group A streptococci: systematic review and implications for vaccine development. Lancet Infect Dis. 2009b;9:611–616. doi: 10.1016/S1473-3099(09)70178-1. [DOI] [PubMed] [Google Scholar]
- Steer AC, Magor G, Jenney AW, Kado J, Good MF, McMillan D, Batzloff M, Carapetis JR. emm and C-repeat region molecular typing of beta-hemolytic Streptococci in a tropical country: implications for vaccine development. Journal of clinical microbiology. 2009c;47:2502–2509. doi: 10.1128/JCM.00312-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stollerman GH. Rheumatogenic streptococci and autoimmunity. Clin Immunol Immunopathol. 1991;61:131–142. doi: 10.1016/s0090-1229(05)80019-4. [DOI] [PubMed] [Google Scholar]
- Sumby P, Porcella SF, Madrigal AG, Barbian KD, Virtaneva K, Ricklefs SM, Sturdevant DE, Graham MR, Vuopio-Varkila J, Hoe NP, Musser JM. Evolutionary origin and emergence of a highly successful clone of serotype M1 group a Streptococcus involved multiple horizontal gene transfer events. The Journal of infectious diseases. 2005;192:771–782. doi: 10.1086/432514. [DOI] [PubMed] [Google Scholar]
- Suzuki H, Lefebure T, Hubisz MJ, Pavinski Bitar P, Lang P, Siepel A, Stanhope MJ. Comparative genomic analysis of the Streptococcus dysgalactiae species group: gene content, molecular adaptation, and promoter evolution. Genome biology and evolution. 2011;3:168–185. doi: 10.1093/gbe/evr006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taddei F, Radman M, Maynard-Smith J, Toupance B, Gouyon PH, Godelle B. Role of mutator alleles in adaptive evolution. Nature. 1997;387:700–702. doi: 10.1038/42696. [DOI] [PubMed] [Google Scholar]
- Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Molecular biology and evolution. 2013;30:2725–2729. doi: 10.1093/molbev/mst197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tesorero RA, Yu N, Wright JO, Svencionis JP, Cheng Q, Kim JH, Cho KH. Novel regulatory small RNAs in Streptococcus pyogenes. PloS one. 2013;8:e64021. doi: 10.1371/journal.pone.0064021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tettelin H, Masignani V, Cieslewicz MJ, Donati C, Medini D, Ward NL, Angiuoli SV, Crabtree J, Jones AL, Durkin AS, Deboy RT, Davidsen TM, Mora M, Scarselli M, Margarit y Ros I, Peterson JD, Hauser CR, Sundaram JP, Nelson WC, Madupu R, Brinkac LM, Dodson RJ, Rosovitz MJ, Sullivan SA, Daugherty SC, Haft DH, Selengut J, Gwinn ML, Zhou L, Zafar N, Khouri H, Radune D, Dimitrov G, Watkins K, O'Connor KJ, Smith S, Utterback TR, White O, Rubens CE, Grandi G, Madoff LC, Kasper DL, Telford JL, Wessels MR, Rappuoli R, Fraser CM. Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial "pan-genome". Proceedings of the National Academy of Sciences of the United States of America. 2005;102:13950–13955. doi: 10.1073/pnas.0506758102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tettelin H, Masignani V, Cieslewicz MJ, Eisen JA, Peterson S, Wessels MR, Paulsen IT, Nelson KE, Margarit I, Read TD, Madoff LC, Wolf AM, Beanan MJ, Brinkac LM, Daugherty SC, DeBoy RT, Durkin AS, Kolonay JF, Madupu R, Lewis MR, Radune D, Fedorova NB, Scanlan D, Khouri H, Mulligan S, Carty HA, Cline RT, Van Aken SE, Gill J, Scarselli M, Mora M, Iacobini ET, Brettoni C, Galli G, Mariani M, Vegni F, Maione D, Rinaudo D, Rappuoli R, Telford JL, Kasper DL, Grandi G, Fraser CM. Complete genome sequence and comparative genomic analysis of an emerging human pathogen, serotype V Streptococcus agalactiae. PNAS. 2002;99:12391–12396. doi: 10.1073/pnas.182380799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tewodros W, Kronvall G. M protein gene (emm type) analysis of group a betahemolytic streptococci from Ethiopia reveals unique patterns. Journal of clinical microbiology. 2005;43:4369–4376. doi: 10.1128/JCM.43.9.4369-4376.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tokajian S, Eisen JA, Jospin G, Coil DA. Draft Genome Sequences of Streptococcus pyogenes Strains Associated with Throat and Skin Infections in Lebanon. Genome announcements. 2014;2 doi: 10.1128/genomeA.00358-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Towers RJ, Gal D, McMillan D, Sriprakash KS, Currie BJ, Walker MJ, Chhatwal GS, Fagan PK. Fibronectin-Binding Protein Gene Recombination and Horizontal Transfer between Group A and G Streptococci. J. Clin. Microbiol. 2004;42:5357–5361. doi: 10.1128/JCM.42.11.5357-5361.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Treangen TJ, Rocha EP. Horizontal transfer, not duplication, drives the expansion of protein families in prokaryotes. PLoS genetics. 2011;7:e1001284. doi: 10.1371/journal.pgen.1001284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tse H, Bao JY, Davies MR, Maamary P, Tsoi HW, Tong AH, Ho TC, Lin CH, Gillen CM, Barnett TC, Chen JH, Lee M, Yam WC, Wong CK, Ong CL, Chan YW, Wu CW, Ng T, Lim WW, Tsang TH, Tse CW, Dougan G, Walker MJ, Lok S, Yuen KY. Molecular characterization of the 2011 Hong Kong scarlet fever outbreak. The Journal of infectious diseases. 2012;206:341–351. doi: 10.1093/infdis/jis362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turner KM, Hanage WP, Fraser C, Connor TR, Spratt BG. Assessing the reliability of eBURST using simulated populations with known ancestry. BMC Microbiol. 2007;7:30. doi: 10.1186/1471-2180-7-30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tyrrell GJ, Lovgren M, St Jean T, Hoang L, Patrick DM, Horsman G, Van Caeseele P, Sieswerda LE, McGeer A, Laurence RA, Bourgault AM, Low DE. Epidemic of group A Streptococcus M/emm59 causing invasive disease in Canada. Clinical infectious diseases : an official publication of the Infectious Diseases Society of America. 2010;51:1290–1297. doi: 10.1086/657068. [DOI] [PubMed] [Google Scholar]
- Varaldo PE, Montanari MP, Giovanetti E. Genetic elements responsible for erythromycin resistance in streptococci. Antimicrobial agents and chemotherapy. 2009;53:343–353. doi: 10.1128/AAC.00781-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vos M, Didelot X. A comparison of homologous recombination rates in bacteria and archaea. The ISME journal. 2009;3:199–208. doi: 10.1038/ismej.2008.93. [DOI] [PubMed] [Google Scholar]
- Wajima T, Murayama SY, Sunaoshi K, Nakayama E, Sunakawa K, Ubukata K. Distribution of emm type and antibiotic susceptibility of group A streptococci causing invasive and noninvasive disease. J Med Microbiol. 2008;57:1383–1388. doi: 10.1099/jmm.0.2008/002642-0. [DOI] [PubMed] [Google Scholar]
- Walker MJ, Hollands A, Sanderson-Smith ML, Cole JN, Kirk JK, Henningham A, McArthur JD, Dinkla K, Aziz RK, Kansal RG, Simpson AJ, Buchanan JT, Chhatwal GS, Kotb M, Nizet V. DNase Sda1 provides selection pressure for a switch to invasive group A streptococcal infection. Nature medicine. 2007;13:981–985. doi: 10.1038/nm1612. [DOI] [PubMed] [Google Scholar]
- Wang CH, Chiang-Ni C, Kuo HT, Zheng PX, Tsou CC, Wang S, Tsai PJ, Chuang WJ, Lin YS, Liu CC, Wu JJ. Peroxide responsive regulator PerR of group A Streptococcus is required for the expression of phage-associated DNase Sda1 under oxidative stress. PloS one. 2013;8:e81882. doi: 10.1371/journal.pone.0081882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wartha F, Beiter K, Normark S, Henriques-Normark B. Neutrophil extracellular traps: casting the NET over pathogenesis. Current opinion in microbiology. 2007;10:52–56. doi: 10.1016/j.mib.2006.12.005. [DOI] [PubMed] [Google Scholar]
- Wertz JE, McGregor KF, Bessen DE. Detecting key structural features within highly recombined genes. PLoS computational biology. 2007;3:e14. doi: 10.1371/journal.pcbi.0030014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wescombe PA, Heng NC, Burton JP, Chilcott CN, Tagg JR. Streptococcal bacteriocins and the case for Streptococcus salivarius as model oral probiotics. Future Microbiol. 2009;4:819–835. doi: 10.2217/fmb.09.61. [DOI] [PubMed] [Google Scholar]
- Wescombe PA, Tagg JR. Purification and characterization of streptin, a type A1 lantibiotic produced by Streptococcus pyogenes. Applied and environmental microbiology. 2003;69:2737–2747. doi: 10.1128/AEM.69.5.2737-2747.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Willems RJ, Hanage WP, Bessen DE, Feil EJ. Population biology of Gram-positive pathogens: high-risk clones for dissemination of antibiotic resistance. FEMS microbiology reviews. 2011;35:872–900. doi: 10.1111/j.1574-6976.2011.00284.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woodbury RL, Klammer KA, Xiong Y, Bailiff T, Glennen A, Bartkus JM, Lynfield R, Van Beneden C, Beall BW. Plasmid-Borne erm(T) from invasive, macrolide-resistant Streptococcus pyogenes strains. Antimicrobial agents and chemotherapy. 2008;52:1140–1143. doi: 10.1128/AAC.01352-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woodbury RL, Wang X, Moran CP., Jr Sigma X induces competence gene expression in Streptococcus pyogenes. Research in microbiology. 2006;157:851–856. doi: 10.1016/j.resmic.2006.07.002. [DOI] [PubMed] [Google Scholar]
- You Y, Yang X, Song Y, Yan X, Yuan Y, Li D, Yan Y, Wang H, Tao X, Li L, Jiang X, Zhou H, Xiao D, Jin L, Feng Z, Yang R, Luo F, Cui Y, Zhang J. Draft genome sequences of two Streptococcus pyogenes strains involved in abnormal sharp raised scarlet fever in China 2011. Journal of bacteriology. 2012;194:5983–5984. doi: 10.1128/JB.01474-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng PX, Chung KT, Chiang-Ni C, Wang SY, Tsai PJ, Chuang WJ, Lin YS, Liu CC, Wu JJ. Complete Genome Sequence of emm1 Streptococcus pyogenes A20, a Strain with an Intact Two-Component System, CovRS, Isolated from a Patient with Necrotizing Fasciitis. Genome announcements. 2013;1:e00149-00112. doi: 10.1128/genomeA.00149-12. [DOI] [PMC free article] [PubMed] [Google Scholar]