Abstract
Posttranslational modification of proteins often controls various aspects of their cellular function. Indeed, over the past decade or so, it has been discovered that posttranslational modification of lysine residues plays a major role in regulating translesion DNA synthesis (TLS) and perhaps the most appreciated lysine modification is that of ubiquitination. Much of the recent interest in ubiquitination stems from the fact that proliferating cell nuclear antigen (PCNA) was previously shown to be specifically ubiquitinated at K164 and that such ubiquitination plays a key role in regulating TLS. In addition, TLS polymerases themselves are now known to be ubiquitinated. In the case of human polymerase η, ubiquitination at four lysine residues in its C-terminus appears to regulate its ability to interact with PCNA and modulate TLS. Within the past few years, advances in global proteomic research has revealed that many proteins involved in TLS are, in fact, subject to a previously underappreciated number of lysine modifications. In this review, we will summarize the known lysine modifications of several key proteins involved in TLS; PCNA and Y-family polymerases η, ι, κ and Rev1 and we will discuss the potential regulatory effects of such modification in controlling TLS in vivo.
Keywords: translesion synthesis, ubiquitin, Y-family polymerase, PCNA
1. Introduction
Posttranslational modifications (PTMs) of proteins by attaching different functional groups to amino acids widens the target protein’s range of function and provides additional mechanisms by which the modified protein can be regulated. For example, PTMs can control a protein’s activity by influencing its ability to interact with protein-partners, alter its enzymatic activity, subcellular localization, and change the stability of the protein. Of all the experimentally identified PTMs in mammals, serine phosphorylation is the most frequent modification followed by lysine, which represents over 15% of all experimentally identified amino acid modifications (calculation based on data from [1]). Lysine can be modified in a variety of ways including, but not limited to: ubiquitination, ubiquitin-like protein (UBL) modification e.g. SUMOylation, ISGylation, neddylation, FATylation and other lysine modifications such as acetylation, methylation, butyrylation, crotonylation, glycation, malonylation, phosphoglycerylation, propionylation, succinylation, myristoylation [1–4].
Eukaryotic cells have evolved a plethora of mechanisms in order to protect genome stability by removing DNA lesions, or preventing their conversion into permanent mutations [5]. Importantly, due to partially overlapping functions of some of these pathways, or time and conditional cellular requirements, their actions need to be precisely controlled. Recent studies in the DNA repair field have accumulated evidence of an ever expanding role of ubiquitination in regulating diverse DNA repair mechanisms and pathways involved in genomic stability maintenance (reviewed in [6]). Ubiquitin- and ubiquitin-like-dependent signaling processes have an important function in controlling cellular responses to DNA damage by navigating through the range of DNA damage repair, or tolerance mechanisms (reviewed in [6–10]). The majority of DNA lesions are repaired by one of the specialized DNA repair pathways; however the repair processes can be slow and incomplete and as a consequence a number of DNA lesions remain in the template DNA. This causes a severe problem, especially during the S-phase of the cell cycle, when DNA is replicated, because efficient and accurate classical DNA polymerases are blocked at DNA lesions. At this critical juncture, distinct mechanisms are required to temporarily tolerate cellular DNA damage, thereby avoiding the permanent block to the replication fork and the threat of cell cycle arrest. Lesion tolerance can be achieved in two different ways; one via a damage avoidance pathway using the information from the undamaged sister chromatid as a template for replication of the damaged DNA region, or via translesion synthesis (TLS), which employs specialized DNA polymerases to synthesize past the lesion.
Over the past dozen years, it has become evident that modification of lysine residues through the covalent linkage of ubiquitin, or ubiquitin-like proteins, plays a central role in controlling both DNA damage avoidance mechanisms and TLS. This review will attempt to summarize the known sites and cellular effects of ubiquitination of several key proteins involved in TLS. We will recap the individually discovered and experimentally confirmed sites of ubiquitination and ubiquitin-like modifications of TLS proteins and combine them with recent data derived from multiple proteome-wide approaches that reveal a hitherto underappreciated extent of lysine ubiquitination of many of the TLS proteins.
2. Types of lysine modifications
2.1. Ubiquitination
In eukaryotic cells, ubiquitination is involved in the regulation of almost all cellular processes, including cell division, membrane transport, signal transduction, DNA repair, endocytosis, inflammatory signalling, apoptosis, etc [11–14]. It has been estimated that roughly 10% of human genes encode for proteins involved in ubiquitin metabolism [15]. The malfunction of ubiquitination processes and ubiquitin-mediated proteolysis has been implicated in various pathologies, including neurodegenerative disorders, inflammatory diseases and cancers [16–19]. Due to their important cellular functions, ubiquitination pathways are significant targets for therapeutics [20, 21].
Protein ubiquitination is a dynamic and reversible process where a three-step enzymatic cascade conjugates a small, regulatory protein, ubiquitin, to a specific lysine residue in a target protein [22]. Initially, one of the ubiquitin-activating enzymes (E1s) forms an ATP-dependent thioester bond with ubiquitin. The activated ubiquitin is then transferred from the E1 enzyme to one of multiple ubiquitin-conjugating enzymes (E2s). E2 then transfers the activated ubiquitin to a protein substrate, either by itself, or with the help of one of the many ubiquitin ligases (E3s). Ubiquitin is linked through its C-terminal glycine residue to a specific internal lysine residue of the target protein. It has been also shown that in some proteins, ubiquitin can be attached to the N-terminus of the protein and in rare cases to a serine, threonine or cysteine residue [23–25]. Monoubiquitinated substrates can undergo further ubiquitination [26–28]. There are seven lysine residues in ubiquitin; K6, K11, K27, K29, K33, K48 and K63; all of them can be involved in polyubiquitin chain assembly. Additionally linear N-terminal polyubiquitin chains can also be formed [29]. Depending on the type of ubiquitin-chain linkage, polyubiquitinated proteins might be destined for degradation by the 26S proteasome in an ATP-dependent manner or alternatively, polyubiquitination might provide a signal for distinct cellular processes such as the inflammatory response or DNA repair [10]. Chains that are linked through K48 are the principal signal for degradation by the proteasome [30, 31]. Recent studies, based on mass spectrometry have shown that homogeneous chains consisting of K29, K11, K27 and K6-linkages, heterogeneous chains with mixed lysine linkages, as well as multiple nearby monoubiquitination and, in cases of substrates up to 150 amino acids, even monoubiquitination can promote proteasomal degradation [32, 33]. Chain elongation of ubiquitinated substrates is mediated via another class of ubiquitin ligases, E4s [34, 35]. Ubiquitination can be reversed through the activity of deubiquitinating enzymes (DUBs), which primarily disassemble polyubiquitin chains before protein degradation, but will also cleave off a single ubiquitin moiety, or a polyubiquitin chain to regulate protein functionality [36].
2.2. Ubiquitin-like posttranslational modification
Besides ubiquitin, at least 10 different ubiquitin-like proteins (UBLs) exist in mammals (reviewed in [37, 38]) with SUMO, NEDD8 and ISG15 being the best known. UBL modifiers, similar to ubiquitin, form an isopeptide bond between their C-terminal glycine and lysine residues of the substrate [38]. UBLs often have low sequence homology, but share a similar three-dimensional structure [38]. Posttranslational modification with UBL proteins can alter cellular function, stability, interactions with protein partners, or subcellular localization of the target protein [37, 39]. Protein modification by UBLs follows the same three-step cascade similar to ubiquitination in that it is catalyzed by sets of analogous activation (E1), conjugation (E2s) and ligation (E3s) enzymes and can be reversed by deconjugating enzymes [40].
SUMO (Small Ubiquitin-like MOdifier) is the most studied UBL modifier and is expressed in all eukaryotes, mainly as a single variant. However in human cells there are four different paralogs (SUMO1–4), representing various homology, expression levels and substrate preferences. Many proteins interacting with a SUMOlyated substrate possess specific SIM domains (from SUMO- interaction motif) [41]. SUMOylation of a target protein can influence the protein degradation, signal transduction, localization, transcription activation, cell cycle, chromatin organization, DNA repair and other functions (reviewed in [42]). Dysfunction of SUMOylation can lead to neurodegenerative diseases, heart defects, diabetes or cancer [42–45].
One ubiquitin-like molecule, ISG15 (the interferon-stimulated gene 15), has a primary sequence that consists of two domains with significant similarity to ubiquitin [46]. Interestingly, ISGylation shares some of the E2s and E3s enzymes used in ubiquitination and ISGylated proteins can also be targeted for degradation by the 20S proteasome [47–48]. ISG15 is only found in vertebrates. Type I interferon, viral infection, lipopolysaccharides and some types of genotoxic stress can rapidly induce ISG15 conjugation [49, 50] and it has been shown that enhanced ISGylation correlates with carcinogenesis [51].
Another example, NEDD8 (neural precursor cell-expressed developmentally downregulated-8), shares 60% identity and 80% homology with ubiquitin [52], and as a consequence, it can be incorporated into polyubiquitin chains by the E2 and E3 ubiquitin-conjugating enzymes [53]. The best characterized substrates known to be neddylated are cullins, scaffold proteins of SCF ubiquitin ligases (Skip-1, cullin, F-box) which regulate ubiquitination and proteasomal degradation of proteins involved in cell cycle control, transcriptional regulation, signal transduction [37, 54]. Other, non-cullin neddylation substrates include proteins involved in RNA splicing, DNA replication and repair and proteasomal degradation [55].
3. Identifying ubiquitination and UBL modification sites
The identification of lysine residue(s) to which ubiquitin, or UBL proteins are conjugated, is important for understanding its biological significance. Locating ubiquitination, or UBL sites, can be performed experimentally, using conventional approaches, such as site-directed mutagenesis of a potentially modified residue [56, 57], or by using antibodies against ubiquitin, or UBL proteins [58, 59]. Recently, however, high-throughput methods and mass-spectrometry have also been frequently employed [60–67]. A collection of experimentally determined proteins which can be ubiquitinated and/or UBL-modified proteins in which the modified residues have been verified have been assembled into several searchable databases, such as UbiProt (http://ubiprot.org.ru/) [68], SCUD (http://scud.kaist.ac.kr) [69], for ubiquitin modification and SysPTM (http://www.sysbio.ac.cn/SysPTM) [70] and CPLM (http://cplm.biocuckoo.org/index.php) [71] for general PTMs.
3.1. Prediction of ubiquitination and UBL sites
The experimental detection of the potential sites of modification is time- and labor-intensive, which is why multiple efforts have been undertaken to computationally predict protein ubiquitination and UBL sites [72–78]. Generally, they all are based on analysing features of experimentally verified ubiquitination sites. The datasets are used for training via various algorithms and tested for their prediction ability. Catic and co-workers discovered that ubiquitination sites are preferably exposed at the molecular surface and reside in loop regions [72]. Kim et al suggest the ubiquitination sites localize within a net negative charge [62]. The analysis of Radivojac et al calculated 586 sequence attributes for each lysine of the positive and negative datasets and demonstrated that the ubiquitination sites are often located in intrinsically disordered regions [74]. This is in contrast to predictions made based on the correlation of ubiquitination with protein structure performed by Walsh et al [78] and might be a result of different data sources (S. cerevisiae vs. H. sapiens). These findings generally indicate that ubiquitination site preferences seem to be poorly conserved across different species [61, 76, 77], therefore organism-specific predictors should be used for optimal results.
Several ubiquitin site predictors, such as UbiPred [73], UbPred [74] and CKSAAP_UbSite [76], trained on yeast datasets, have been developed. However, quite recently, some predictors have become available exclusively for mammalian sites, such as hCKSAAP_UbSite [77], UbiProber [79] and RUBI [78]. For SUMOylation, a consensus motif has been determined as ψKxD/E [80] (where ψ is a hydrophobic residue I, L, M, P, F, V or W and X is any residue). However, this motif is not very precise; about 40% of known SUMOylation sites do not match the consensus, and no detectible SUMOylation was found in some proteins having the ψKxD/E motif [81]. In the last decade, several methods have been used to generate SUMOylation site predictors, which include GPS, MotifX, SUMOsp, SUMOsp 2.0; seeSUMO, SUMOplot, SUMO-hydro and SUMmOn among others [81; 82–87]. SUMOylation frequently appears to be site-specific, thus prediction programs recognize SUMO-modified sites primary on amino acid sequence information.
The high specificity and sensitivity desired in the computational determination of ubiquitination and UBL sites remains a challenge. In a recent paper from Schwartz, that assessed the different available predictors, it was pointed out that 9 out of 11 predictors perform no better than random, or unseen data [88], demonstrating that at the present time, bioinformatic analysis alone appears to be insufficient in confidently identifying bona fide sites of ubiquitination or SUMOylation. In short, ubiquitin and UBL sites still need to be determined experimentally.
3.2. Proteome-wide profiling of ubiquitin and UBL modification
The need for large-scale detection of modified proteins has been recognized for a long time and within the past several years the scientific community has witnessed an explosion in the global identification of ubiquitinated proteins and sites and to a lesser extent, UBL modification [61–67, 89–96]. Major progress has been made possible due to the development of new affinity purification tools and the improvement of mass spectrometers with increased speed, resolution and accuracy, together with decrease of false-positive recognition [97–99].
The abundance of modified proteins is usually too low to be directly detected by mass spectrometry, therefore a variety of methods that enrich modified proteins has been employed. The strategies for enriching substrates for high-throughput ubiquitination site recognition include the usage of epitope-tagged ubiquitin expression system [61, 91, 94, 100, 101] and the employment of specific ubiquitin binding domains and antibodies against ubiquitin or peptides containing a K-ε-GG remnant that is created by tryptic digestion of a ubiquitinated protein [93, 102–104]. All of the enrichment methods, however, carry some limitations that might lead to artifacts that one cannot exclude during analysis. For example, in a system using epitope-tagged ubiquitin, modified proteins can be purified under denaturing conditions to help reduce false positive identification. On the other hand, exogenously overexpressed tagged ubiquitin might hinder the kinetics of ubiquitination reactions, as well as hamper the formation of linear ubiquitin chains and consequently interfere with the cellular functions of modified proteins [105, 106]. The employment of tandem UBDs (Ubiquitin Binding Domains) and ubiquitin antibodies circumvents this problem, but there is an increased risk of co-purifying contaminant proteins or a bias towards a specifically modified substrate (linkage specific polyubiquitinated or monoubiquitinated proteins) [90, 91, 107, 108]. Both problems are circumvented by using the K-ε-GG remnant antibodies developed by Xu et al. [102], as they do not isolate the entire substrate, but rather a short peptide with the glycine-glycine signature obtained after tryptic digestion of a ubiquitinated substrate. This method is highly efficient and has been widely used for ubiquitination site profiling [62–65, 89, 92, 109]. It is noteworthy that less abundant ubiquitin-like modifications, such as neddylation and ISGylation, also generate the same di-GG signature after trypsinization and the di-GG antibodies do not discriminate them from ubiquitination [62, 110]. The COFRADIC technology, recently developed by Stes et al provides an alternative strategy for ubiquitination studies and does not require overexpression, epitope tags or specific antibodies [110].
To date, literally hundreds of mammalian proteins have been described as SUMOylation targets, however precise identification of SUMOylated sites remains challenging due to low abundance of modified proteins in vivo and the dynamic character of SUMOylation [95, 111–113]. Several attempts towards large-scale identification of SUMOylation sites have been undertaken. Most methods employ recombinant SUMO in in vitro or in vivo SUMOylation assays followed by antibody- or epitope-based purification, digestion by proteases and mass-spectrometry analysis of the peptide [94–96, 112, 114–117]. The remnant of SUMO proteins left on the modified lysine residue after digestion with trypsin is relatively long (19–32 amino acids depending on the SUMO isoform) leading to a complex fragmentation pattern which obstructs precise identification of the SUMOylated site. To overcome this problem and generate shorter SUMO isopeptide fragments, convenient cleavage sites were introduced into the SUMO protein allowing for the generation of a specific di-GG signature after tryptic digestion [66]. 84 out of 103 identified SUMO sites were located in direct, or an inverted SUMO consensus sequences and out of the 16 proteins that were identified, a new SUMOylation motif HCSM (Hydrophobic Cluster SUMOylation Motif) was identified.
Additionally, Tammsalu et al identified over 1000 SUMOylation sites within 539 human proteins involved in cell cycle, transcription and DNA repair. By using a His-tagged SUMO2 with a T90K mutation, they obtained a di-GG remnant after endoproteinase cleavage and the SUMO-enriched peptides were subsequently analyzed by mass spectrometry [67]. Very recently, Hendriks et al identified 4361 SUMOylation sites in 1606 proteins in human cells, both under normal growth conditions and in response to heat shock stress, as well as to SUMO protease and proteasome inhibition [96].
TAP-tagged and GST-tagged Nedd8 were respectively used to identify 75 and 496 neddylated proteins by Xirodimas et al and Jones et al 2008 [55, 118]. Various cullin and non-cullin neddylation substrates were discovered including proteins involved in RNA splicing, DNA replication and repair and proteasomal degradation [55, 118].
There are limited reports of large-scale identification of ISGylated proteins. By using high-throughput immunoblotting, for example, Malakhov et al discovered 76 ISG15 substrates involved in translation, glycolysis, stress response and cell motility [59]. Similarly, by using affinity selection and mass spectrometry Zhao et al identified 158 proteins ISGylated in response to interferon and functioning in diverse cellular processes [119].
In summary, the wide-assortment of multiple large-scale analyses has identified thousands of ubiquitination and UBL modification sites in numerous proteins. However, the large amount of data collected poses a huge challenge to validate the global proteomic studies. As discussed above, different strategies raise a variety of technical and analytical issues, such as contaminant protein recognition, pseudo modification, and false-positive assignments. Various algorithms may also have different levels of sensitivity and specificity that may bias the data obtained [105]. Therefore, individual examination and critical evaluation is essential to confirm global modification site profiling.
4. Post Replication DNA Repair (PRR) pathway
DNA damage tolerance pathways allow for temporal acceptance of the presence of DNA lesions in the genome when there is a risk of cell death. The most frequent and deleterious are DNA lesions that arise during the DNA replication. Very efficient and extremely faithful polymerases guarantee fast and accurate DNA duplication during S-phase. However, any distortion in the DNA structure can hinder this proficient process and cause the replication fork to stall. If stalled for too long, the replication fork can collapse, generating DNA double strand breaks that lead to genome instability. It is therefore critically important to resume replication even in the face of persistent DNA damage [120, 121]. Based on genetic studies, PRR utilizes two major mechanisms that allow blocked replication to resume. The first is indirect bypass using DNA damage avoidance mechanism and the second is DNA translesion synthesis (TLS). The mechanism of DNA damage avoidance is not clear, however it is thought to involve template switching where the undamaged sister chromatid is used as a temporary replication template or homologous recombination [122, 123]. In contrast, during TLS, the highly precise and efficient DNA replicases that are blocked by a DNA lesion are replaced by specialized, low-processivity TLS polymerases that are able to carry out DNA synthesis past the damaged site. These TLS DNA polymerases can either bypass the lesion unassisted, or with the help of another TLS polymerase in a two-step process [124].
4.1. PCNA modifications
It should be noted that in budding yeast, the decision as to which DNA damage tolerance pathway will be undertaken to rescue a stalled replication fork depends on the type of posttranslational modification to proliferating cell nuclear antigen (PCNA), which is a central player in the PRR pathway.
PCNA, a replication processivity factor, is a ring-shaped homotrimeric complex which encircles double stranded DNA and slides along the DNA [125]. The monomers, each comprising two structurally similar domains, are linked in head-to-tail mode. PCNA monomers interact with DNA through their DNA binding motifs (61–80 residues) located on an internal surface. On the outer surface, the N- and C-terminal halves of PCNA are linked by the interdomain-connecting loop (IDCL) positioned above a hydrophobic pocket that provides a docking site for the PIP (PCNA interacting peptide) motif of proteins that interact with PCNA [126]. PCNA interacts with multiple proteins involved in replication, cell cycle regulation and DNA repair and coordinates their access to replication forks (reviewed in [127, 128].
In response to replication fork stalling, PCNA undergoes monoubiquitination at K164 by Rad6 and Rad18 (E2-ubiquitin conjugation and E3-ubiquitin ligase enzyme, respectively) [129, 130]. Monoubiquitinated PCNA interacts with TLS polymerases via their ubiquitin binding domains (UBDs), thereby activating the TLS pathway. Rad6-Rad18 is the main source of PCNA monoubiquitination, though some residual, conditional ubiquitination can be observed in yeast and chicken cells lacking Rad6 or Rad18 [131–133]. There are also reports that human PCNA can be monoubiquitinated by CRL4Cdt2 or Rnf8 ubiquitin ligases [134, 135]. PCNA ubiquitination is reversible and modified PCNA can be deubiquitinated via USP1 or BPLF1 (only in human cells) [136–139].
PCNA that is monoubiquitinated at K164 can undergo further ubiquitination. In budding yeast, K63-linked polyubiquitin chains, catalyzed by the Ubc13-Mms2/Rad5 E2-E3 enzymes, promote template switching [130]. In human cells, there are two Rad5 homologues, HLTF and SHPRH [140–143] serving as the E3 ligases for K63-chain formation. Like yeast Rad5, they both interact with RAD6/RAD18 and MMS2/UBC13 complexes [140, 141], however, their role is not fully understood and their function in both damage avoidance and TLS sub-pathways of PRR have been suggested [144]. Furthermore, Krijger et al suggested the existence of yet another E3 ligase, as PCNA polyubiquitination was observed in HLTF/SHPRH double mutant mice [145]. An additional difference in PCNA polyubiquitination between lower and higher eukaryotes regards a requirement for MMS2, as the protein seems to be dispensable for this process in mammalian cells [146]. PCNA polyubiquitination, similar to PCNA monoubiquitination, is negatively regulated by USP1 [138, 141].
Human PCNA is a stable protein with an estimated half-life of over 20 hours. The stability can be attributed to proteins such as MUTH2, ERK8 and NRAGE, which protect PCNA from polyubiquitination by blocking proteasome degradation signaling through K48- or K11-linked chains [147–149].
Yeast PCNA can also be ubiquitinated at K107 in response to replication stress caused by the presence of unprocessed Okazaki fragments in ligase-deficient cells. K107 monoubiquitination signals checkpoint activation and both mono- and K29-linked polyubiquitination on K107 involves Mms2, Ubc4 and Rad5 [150]. Human PCNA is also ubiquitinated in ligase I depleted cells, but the modified residue has not yet been identified [150].
In addition to ubiquitination, PCNA can be modified by SUMO. Initially, this modification was identified in yeast, followed by a handful of other species including Xenopus and chicken DT40 cells [129, 131, 151]. Recently, a low level of SUMOylation of human PCNA has also been reported [152, 153]. SUMOylation is a reversible process and Ulp1 hydrolase removes SUMO from PCNA [154].
In budding yeast, PCNA SUMOylation occurs mostly during S-phase progression and, by influencing PCNA interactions with various partners, controls DNA replication and repair. PCNA SUMOylation promotes the binding of the Srs2 helicase, an inhibitor of recombination and thereby prevents unwanted recombination events at the replication fork [155]. Kim et al showed that the non-canonical Srs2 PIP box has relatively low affinity for unmodified PCNA but it enhances significantly upon PCNA SUMOylation [156]. Another interaction involving Elg1, the large subunit of an alternative clamp loading complex, implies the involvement of SUMOylation of PCNA in its unloading from DNA in yeast cells [157; 158]. Conversely, ATAD5, a human homologue of yeast Elg1, does not seem to have a preference for unloading SUMOlyated PCNA despite possessing a SUMO interacting motif [159, 160]. On the other hand, it has been shown to be involved in deubiquitinating PCNA by recruiting the USP1 complex to ubiquitinated PCNA [155]. PCNA SUMOylation in yeast cells has also been shown to inhibit Eco1-PCNA-dependent sister chromatid cohesion [161].
Yeast PCNA can be SUMOylated on K164 and to a lesser extent on K127, with the involvement of SUMO conjugating and ligating E2-E3 enzymes, Ubc9 and Siz1 (K164), or just Ubc9 itself (K127) [129]. Gali et al showed that hPCNA can be SUMOylated on K164 and K254 which prevent DSB formation and inappropriate recombination in response to replication fork arrest by DNA lesions [152]. In another study, the human analog of Srs2, PARI, was shown to promote the interaction with SUMOylated PCNA, correspondingly obstructing homologous recombination [153].
Monoubiquitination of PCNA on K164 strengthens the interaction with TLS polymerases, while yet another lysine modification, ISGylation, promotes release of polymerase η (polη) from PCNA [162]. Upon UV irradiation, either K164 or K168 is assumed to undergo ISG15 modification that induces PCNA de-ubiquitination and polη discharge followed by PCNA de-ISGylation and resumption of normal replication.
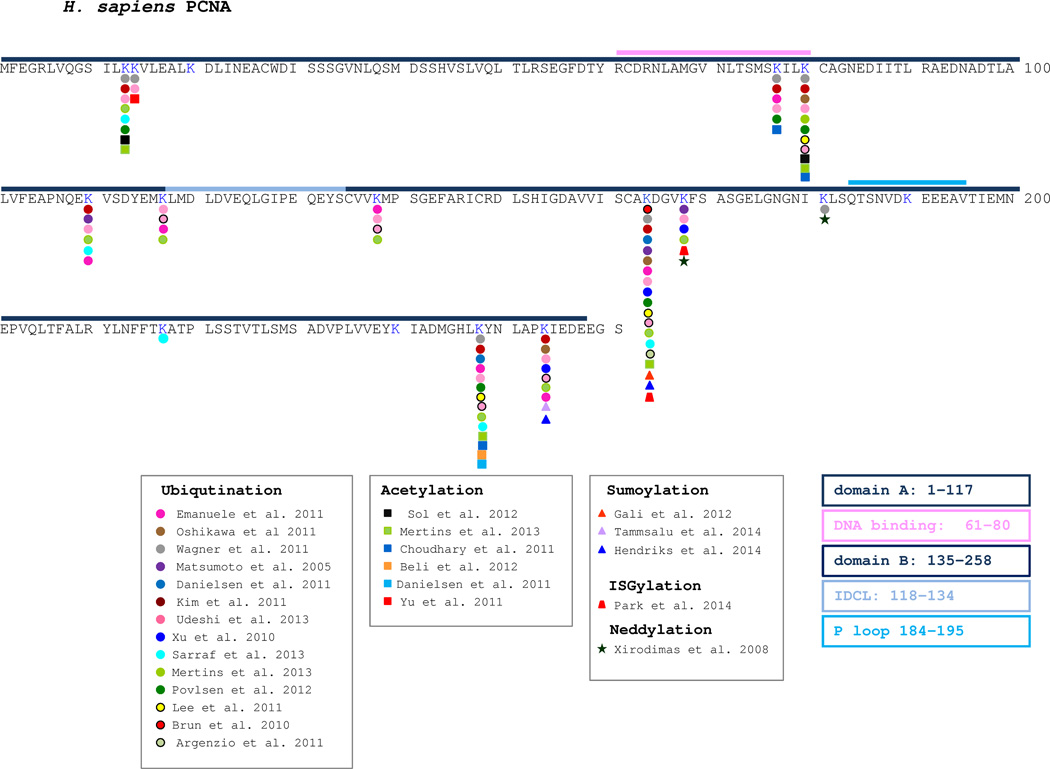
There are 16 lysine residues in human PCNA and 13 of them have been reported to be ubiquitinated, either in individual experiments (K164), or by numerous large-scale methods (Figure 1). Nine of the identified residues can also be UBL-modified, or acetylated, suggesting possible competition between different types of modification. Site-specific overlap between lysine ubiquitination and acetylation has been suggested for about 20% of identified protein ubiquitinations [61] and an interplay between ubiquitination and acetylation represents a common way to regulate protein stability. In most cases, lysine acetylation prevents ubiquitination and ubiquitin mediated proteolysis of modified protein, but there are some examples of acetylation-directed acceleration of protein degradation, by modulating protein-protein interactions (reviewed in [163]). Indeed, Yu et al showed that UV-induced PCNA acetylation at K14 caused dissociation of PCNA from a complex with MTH2, and as a consequence, shortens its half-life, as PCNA is more easily degraded by the proteasome[147].
Fig. 1. Posttranslational modification of human PCNA.
Lysines that have been indicated to be ubiquitinated are shown with a colored circle. Lysines that have been indicated to be acetylated are shown with a colored square. Lysines that have been indicated to be SUMOylated are shown by a colored triangle. The lysine residue that is ISGylated is shown by a red trapezium. The lysines that are known to be neddylated are shown by a green star. References are given in the appropriate associated box.
In PCNA, six of the lysines located in the N- and C-terminal domains were found to be targets of both ubiquitination and acetylation. Naryzhny and Lee [164] showed in Chinese hamster ovary (CHO) cells that the PCNA acetylation status plays an important role in regulating its function. Indeed, they suggested the existence of three PCNA isoforms differing in their acetylation status and subcellular localization. Additionally, they implied that acetylated PCNA is involved in DNA replication, while its deacetylated form with replication termination, as acetylated PCNA has higher affinity for polδ and polβ, compared to the deacetylated form.
UBL modifications were identified at four lysine residues in the C-terminal half of the protein. The most modified lysine residue is K164, as all of the global ubiquitin profiling studies detected ubiquitination at this site. Additionally, K164 can also be a subject of SUMOylation, ISGylation and acetylation. The structural studies of ubiquitinated PCNA suggest that the ubiquitin moieties attached to K164 of PCNA monomers extend away from the ring and do not block the interactions of PIP box containing proteins with PCNA. It seems, however, that the type of modification can control the interacting partner by strengthening the interaction between the ubiquitin, or UBL, on PCNA and UBD, or UBL binding motif on the protein [165].
Two of the identified lysine residues, K77 and K80, are placed within the DNA binding domain, implying possible obstruction of DNA-PCNA binding upon modifications at any of these sites. Two other lysines, K110 and K117, in which ubiquitination was detected in multiple studies, lie within a segment of 101–120 amino acids found to play an essential role for PCNA nuclear location [164]. Even though both K110A and K117A mutants are efficiently imported to the nucleus, K110A does not co-localize with replication foci suggesting that ubiquitination of this residue might influence PCNA foci formation.
4.2. Modifications of Y-family polymerases
While it is apparent that many of the recently characterized DNA polymerases have the ability to facilitate TLS of certain DNA lesions, the best characterized from an historical perspective are the four Y-family DNA polymerases polη, polι, poκ, and Rev1 and the B-family DNA polymerase, polζ [167, 168]. Each of these polymerases presents a specific portfolio of DNA lesions they are able to bypass with differing efficiency and accuracy. The Y-family polymerases are multi-domain proteins. The catalytic domain generally occupies the N-terminus, while the C-terminus is engaged in protein-protein interactions and contains various protein-binding motifs. Polymerases η, ι and κ possess non-canonical PCNA-binding motifs (PIP box) and a Rev1 interacting region (RIR) [168]. Rev1 interacts with PCNA via a BRCT domain localized at its extreme N-terminus while the C-terminus of Rev1 interacts with polymerases η, ι and κ. All Y-family DNA polymerases have ubiquitin binding domains that bind non-covalently to ubiquitin, or ubiquitinated proteins. Polymerases ι and Rev1 possess two UBMs, while polymerases η and κ, have so called UBZs – UBDs that additionally bind a zinc atom (polη has one UBZ, whereas polκ has two UBZs) [168–170]. Besides possessing UBDs that facilitate the interaction with monoubiquitinated PCNA, human polη, polι, mouse polc and Rev1 as well as yeast and nematode polη have been shown to be subject to ubiquitination themselves [169, 171–177]. In general, most of the proteins that can non-covalently bind ubiquitin via different types of UBDs are themselves targets of monoubiquitination in a process called coupled monoubiquitination [178, 179]. In this process, ubiquitin attached to an E2 ubiquitin conjugating enzyme, or E3 ubiquitin ligase, is recruited to the UBD containing substrate which becomes ubiquitinated in an E3-dependent, or independent mode [180].
In addition to ubiquitination, Caenorabditis elegans polη and polκ have also been shown to be subject of SUMOylation [174, 181].
4.2.1. DNA polymerase η modifications
Polη is possibly the best-characterized Y-family DNA polymerase and is mainly known for efficient replication past cyclobutane pyrimidine dimers, which are the main DNA lesions induced after UV-irradiation. As a consequence, a dysfunction in human polη results in the variant form of Xeroderma Pigmentosum, which is characterized by sunlight sensitivity and a high incidence of skin cancer [181, 183].
It has previously been shown that human polη can be ubiquitinated in vivo in its nuclear localization signal (NLS) motif. K682 was identified as the main ubiquitination site, however when this residue is unavailable, three other close by lysines (K686, K694 or K709) can serve as a target [169, 171]. Pirh2, an E3 ligase, was discovered to interact with human polη and monoubiquitinate it at one of the four lysine residues at the C-terminus [184, 185]. Attaching a ubiquitin moiety to the C-terminus of polη prevents its interaction with PCNA and inhibits its ability to bypass UV-induced lesions and causes an increased sensitivity to UV radiation [171, 185]. Therefore, monoubiquitinated polη needs to be actively de-ubiquitinated prior to interacting with PCNA and its recruitment to a stalled replication fork [171]. Additionally, polη is a subject of polyubiquitination by another E3 ligase, Mdm2, that targets polη for proteasomal degradation and controls its stability in response to UV-induced DNA damage [186]. Wallace et al showed that human polη can also be polyubiquitinated by a RING E3 ligase, TRIP (TRAF-interacting protein (tumor necrosis factor receptor (TNFR)-associated factor) and TRIP promotes its localization in nuclear foci [187]. The TRIP homolog in Drosophila melanogaster, NOPO, enhances ubiquitination of polη during insect embryogenesis [187]. Most probably NOPO promotes non-proteolytic polyubiquitination, as its overexpression does not cause polη destabilisation and additionally, NOPO interacts with Bendless (Ben), the Drosophila homolog of Ubc13, suggesting the formation of K63-linked polyubiquitin chains [188].
Recently, the deubiquitinating enzyme, USP7, has been reported to regulate the stability of human polη in two ways. On one hand, USP7 can directly deubiquitinate polη which stabilises polη, and on the other hand, knockout of USP7 increases the steady-state level of polη by destabilising Mdm2 [189].
S.cerevisiae polη was also found to be ubiquitinated [173, 175, 176]. However, the particular ubiquitination sites were not identified. Nevertheless, it has been shown that similar to human polη, the ubiquitination depends on a functional UBZ domain [169, 175]. Ubiquitination of yeast polη is correlated with the cell cycle and increases during G1 and drops as cells enter S-phase, thereby allowing for the recruitment of polη to PCNA that is monoubiquitinated in response to a replication block [173]. There are contradicting reports about the stability of yeast polη [172, 173] because it appears that the half-life of the yeast enzyme largely depends on the epitope tag used to identify the recombinant enzyme [190, this issue].
Interestingly, in response to DNA damaging agents (MMS and UV), nematode polη becomes SUMOylated by GEI-17 SUMO E3 ligase at K85 and K260 and protects it from degradation mediated by CRL4-CDT-2-dependent ubiquitination [174, 181]. The SUMOylated lysine residues are conserved in human polη (K86 and K261 respectively), and very recently human polη has been reported to be SUMOylated at K163 (Patricia Kannouche and Emmanuelle Despras, Pers. Comm.).
In addition to ubiquitination and SUMOylation, human polη can also be phosphorylated by both ATR kinase and protein kinase C (PKC), in response to UV radiation. Potential phosphorylation sites were reported at S587, T617 and S601 by two independent groups [191, 192]. The phosphorylation of polη seems to be required for cell survival after UV radiation and provides a link between DNA damage-induced checkpoint control and translesion synthesis.
There are 47 lysine residues in human polη. In 7 out of 13 large-scale studies, polη was found to be ubiquitinated (Figure 2). In total, the ubiquitination of 14 lysine residues was reported, but 10 of them were identified in single analysis implying that they might represent non-specific modifications. All 7 global approaches that showed polη ubiquitination identified K682, thereby independently confirming the primary ubiquitination site originally found by Bienko et al [171]. Three neighboring lysine residues, K686, K694 and K709 were also identified as back-up sites of ubiquitination, again, as previously reported by Bienko et al [171].
Fig. 2. Posttranslational modification of human DNA polη.
Lysines that have been identified to be ubiquitinated are shown with a colored circle. The serine residues that are thought to be phosphorylated, are shown by a colored star. References are given in the appropriate associated box.
4.2.2. DNA polymerase ι modifications
DNA polι is a paralog of polη [193] and is thought to bypass a number of lesions in vivo especially when polη is absent (e.g. in XPV cells) and due to its reduced accuracy in synthesizing across photoproducts, polι-dependent TLS results in elevated mutagenesis [194–198]. Extensive biochemical studies performed in vitro with the highly purified enzyme suggest that polι is able to bypass a wide range of DNA lesions [199–202]. Interestingly, when copying an undamaged template, its accuracy varies 10,000-fold depending on the template base copied (reviewed in [203]). Its unusual preference of incorporating G opposite T (3 to 10-fold greater than the correct base, A) gives polι a distinctive signature [204, 205] and is a result of the specific structure of its active site [206, 207]. Another interesting feature of polι is that its N-terminus contains two partially overlapping catalytic domains; one with DNA polymerase activity and one with dRP lyase activity [208, 209]. The C-terminus contains motifs characteristic of other Y-family polymerases and includes PCNA-interacting (PIP), Rev1-interacting (RIR) and Ubiquitin-interacting (UBM1 and UBM2) motifs/domains.
To date, the only reported posttranslational modification in human polι is ubiquitination [169] and the cellular function of ubiquitinated polι is not fully understood. However, our previous studies implied that ubiquitination of either polη or polι is required for the two polymerases to physically interact [210].
There are 53 lysine residues in human polι. In 8 total large-scale studies, 24 lysine residues distributed along the entire length of the polymerase were found to be ubiquitinated (Figure 3). Ubiquitination of half of these residues was detected just once, suggesting that the modifications might have appeared either accidentally, or represent rather rare conditional cases. The remaining 12 modified lysines were found to be ubiquitinated in 2–4 independent approaches, often using different experimental strategies. We believe that multiple autonomous detection of the same residue thereby increases the probability that the ubiquitination of any particular lysine has a functional meaning and possibly a broader cellular effect. Nonetheless, no single lysine has appeared in all of the analyzed studies.
Fig. 3. Posttranslational modification of human DNA polι.
Lysines that have been indicated to be ubiquitinated are shown with a colored circle and the appropriate reference cited in the associated box.
Thirteen of the detected ubiquitination sites are located in the polymerase catalytic domain and the proposed dRP lyase domain, suggesting that ubiquitination of some of these lysines could possibly influence the enzymatic activities of polι. Ubiquitination of K309 was detected in four independent approaches and nearby K310 in two, giving these residues a higher possibility of bona fide modification. Ubiquitination targets were also reported in two adjacent lysines, K549 and K550, of the RIR motif suggesting control of the Rev1-polι interaction. The detection of ubiquitination of both lysines suggests that either one could serve as a modification target. Several other ubiquitination sites were detected at, or in close proximity to the UBM motifs, which could possibly affect the ability of polι to bind to ubiquitinated proteins (such as PCNA or polη).
4.2.3. DNA polymerase κ modifications
Polκ is able to bypass multiple types of DNA lesions including abasic sites and bulky adducts, but with rather low efficiency [211] and due to a constricted active site, cannot incorporate a base opposite a pyrimidine dimer [212]. When copying an undamaged template, polκ is quite accurate compared to other Y-family polymerases, but can also extend mispaired primer termini [213]. Furthermore, after UV-irradiation the activity of pok has been implicated in the gap-filling step of nucleotide excision repair [214].
Similar to other Y-family DNA polymerases, pok possesses a catalytic domain at its N-terminus while the C-terminal half of the protein contains domains involved in protein-protein interactions including an RIR, two UBZ domains and at the extreme C-terminus, a PIP box [215, 216]. The UBZ domains mediate the enhanced interaction with PCNA and some studies report that they are essential for UV-induced nuclear foci formation by polκ, but do not affect protein’s half-life [177]. On the other hand, Okada et al suggests that polκ can function independently of PCNA modification [216]. Recent studies of Wit et al concluded that dependent on the type of DNA damage, PCNA ubiquitination may, or may not, be required for polκ activation [217].
To date, several reports show evidence for posttranslational modifications to polκ in eukaryotic cells. Guo et al observed monoubiquitination of mouse polκ and confirmed that similar to other Y-family polymerases, monoubiquitination depends on UBDs [173]. Endogenously expressed murine polκ is a stable protein with a half-life estimated to be ∼ 5.4 h [177]. The turnover of exogenously expressed polκ is somewhat faster (3.7–4.2 h), but does not change in polκ UBZ mutants. The suggestion that human polκ might also be ubiquitinated comes from the publication of Wallace et al which reports an interaction between the C-terminus of polκ and the RING E3 ligase, TRIP [187]. On the other hand, the possibility of polκ E3-independent monoubiquitination has also been reported in vitro [179]. The biological function of polκ modification is not yet known. However, by influencing polκ interactions with other proteins, it may regulate its presence in replication factories.
Regarding other types polκ modification, Roerink et al suggested that GEI-17, which is known to SUMOylate and consequently protect nematode polη from proteasomal degradation, most likely also acts on polκ [181], so it is entirely possible that polκ may also be SUMOylated in vivo.
Until now, the information about which lysine residues can be modified in polκ comes from large-scale proteomics analysis. There are 84 lysine residues in human polκ. Just four of the over 13 independent global studies have reported ubiquitination of polκ at 36 lysines (Figure 4). It should be noted that most of these sites were detected in just one single analysis. Only K173, K461 and K541 were detected in two studies and K683 in three studies. Indeed, the majority of polκ expressed in vivo is not ubiquitinated [173], which probably explains the limited number of studies reporting ubiqutination of polκ. In a recent publication on proteome wide SUMOylation, Hendriks et al [96] reported SUMO2 modifications at six polκ lysine residues; half of which, K55, K224 and K814 overlap with known ubiquitination sites.
Fig. 4. Posttranslational modification of human DNA polκ.
Lysines that have been indicated to be ubiquitinated are shown with a colored circle. Lysines that have been indicated to be SUMOylated are shown by a colored triangle. The appropriate references are cited in the associated box.
4.2.4. Rev1 modifications
Rev1 is a unique enzyme among Y-family polymerases, as it is only able to incorporate dCMP nucleotides opposite undamaged or damaged template G and some DNA lesions, including an abasic site [218–221]. Besides its distinctive enzymatic activity, Rev1 also plays a non-catalytic role in TLS as a scaffolding protein that coordinates the other TLS polymerases. The extreme C-terminus of Rev1 in higher eukaryotes is devoted to the interaction with pols η, ι, κ and ζ [214, 222–227]. Additionally the two UBMs that are located close to the C-terminal domain permit the interaction with ubiquitin, ubiquitinated PCNA, or other ubiquitinated proteins [170, 228]. Interestingly, Rev1 lacks a well-conserved PIP box, that is characteristic of the other Y-family polymerases and the interaction with PCNA is, instead, through the N-terminal BRCT domain of Rev1 [229, 230].
In addition to non-covalent interacts with ubiquitin and ubiquitinated proteins, at least two studies have shown that mouse and human Rev1 can also be directly conjugated to ubiquitin [176, 231]. However, the mechanism and sites of the ubiquitination remain unknown. Kim et al reported that ubiquitinated human Rev1 can be recruited to nuclear foci by the Fanconi Anemia core complex, as it binds directly to the UBZ4 domain of FAAP20 protein [231]. Moreover, the level of S.cerevisiae Rev1 seems to be regulated in a cell cycle-dependent mechanism via ubiquitin-mediated proteasomal degradation, suggesting that it gets ubiquitinated before degradation [232].
There are 92 lysine residues in human Rev1. However, ubiquitination of Rev1 was detected in just one of the large-scale proteomic studies [64]. Out of 7 lysine modifications 5 localize to the N-terminus (K28, K41, K119, K134 and K140), which are located near, or within, the BRCT domain. The remaining ubiquitinated lysines were located in the middle of the protein (K678 and K770].
SUMO2 modification of human Rev1 at K99 has also been reported [96]. In another proteome-wide analysis, Drosophila melanogaster Rev1 was shown to be acetylated at K136 [233]. However, in human Rev1, this residue is replaced by arginine (R149). These results suggest that Rev1 modifications (most probably at the N-terminus), occurs either rarely, or under highly specific conditions not employed in the global proteomic studies.
5. Concluding remarks
Review of the published data reveals interesting differences in the extent of posttranslational modification of key TLS proteins. PCNA is the most highly modified protein and is subject to ubiquitination, acetylation, SUMOylation, ISGylation and neddylation. Human polη is a target of limited ubiquitination, primarily at one key residue, K682, and is also been reported to undergo phosphorylation and very recently SUMOylation. In contrast, nearly 50% of the lysines in polι can apparently be ubiquitinated. This is also in dramatic contrast to either the polκ or Rev1 proteins, which appear to undergo limited lysine (or any other) posttranslational modification. It will therefore be interesting to determine why polι is so highly ubiquitinated and the effects such ubiquitination have on the regulation and in vivo properties of the enigmatic polι enzyme.
Highlights.
Key TLS proteins are modified at multiple lysines.
PCNA is the most highly modified protein.
Polymerase η is a target of limited ubiquitination in its C-terminus.
At least half of polymerase ι lysines can be ubiquitinated in vivo.
Polymerase κ and Rev1 undergo limited posttranslational modifications.
Acknowledgements
Funding for this review was provided by the National Institutes of Health Intramural Research Program (to R.W.) and the Foundation for Polish Science HOMING PLUS/2013-7/10 Regulation of human DNA polymerase iota by ubiquitination (to J.M.). We would like to thank Ewa Sledziewska-Gojska, Alexandra Vaisman and Mary McLenigan for critical comments on the manuscript and members of the Mass Spectrometry Lab from IBB PAS for help with analyzing some of the mass spectrometry data.
Abbreviations
- TLS
translesion synthesis
- PCNA
proliferating cell nuclear antigen
- pol
polymerase
- PIP
PCNA-Interacting Peptide
- RIR
Rev1-interacting region
- UBM
Ubiquitin binding motif
- UBZ
ubiquitin binding zinc motif
- polι
DNA polymerase iota
- polη
DNA polymerase eta
- polκ
DNA polymerase kappa
- UBL
ubiquitin-like protein
- PTM
Posttranslational modification
- PRR
Post Replication DNA Repair
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Conflict of interest
The authors declare no conflict of interest
References
- 1.Khoury GA, Baliban RC, Floudas CA. Proteome-wide post-translational modification statistics: frequency analysis and curation of the swiss-prot database. Sci. Rep. 2011;1 doi: 10.1038/srep00090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Chen Y, Sprung R, Tang Y, Ball H, Sangras B, Kim SC, Falck JR, Peng J, Gu W, Zhao Y. Lysine propionylation and butyrylation are novel post-translational modifications in histones. Mol. Cell. Proteomics MCP. 2007;6:812–819. doi: 10.1074/mcp.M700021-MCP200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Zhang X, Wen H, Shi X. Lysine methylation: beyond histones. Acta Biochim. Biophys. Sin. 2012;44:14–27. doi: 10.1093/abbs/gmr100. [DOI] [PubMed] [Google Scholar]
- 4.Lee S. Post-translational modification of proteins in toxicological research: focus on lysine acylation. Toxicol. Res. 2013;29:81–86. doi: 10.5487/TR.2013.29.2.081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Friedberg EC, Friedberg EC. Eds DNA repair and mutagenesis. 2nd ed. Washington, D.C: ASM Press; 2006. [Google Scholar]
- 6.Pinder JB, Attwood KM, Dellaire G. Reading, writing, and repair: the role of ubiquitin and the ubiquitin-like proteins in DNA damage signaling and repair. Front. Genet. 2013;4 doi: 10.3389/fgene.2013.00045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Zhao Y, Brickner JR, Majid MC, Mosammaparast N. Crosstalk between ubiquitin and other post-translational modifications on chromatin during double-strand break repair. Trends Cell Biol. 2014;24:426–434. doi: 10.1016/j.tcb.2014.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Yang K, Weinacht CP, Zhuang Z. Regulatory Role of Ubiquitin in Eukaryotic DNA Translesion Synthesis. Biochemistry. 2013;52:3217–3228. doi: 10.1021/bi400194r. [DOI] [PubMed] [Google Scholar]
- 9.Ulrich HD. Two-way communications between ubiquitin-like modifiers and DNA. Nat. Struct. Mol. Biol. 2014;21:317–324. doi: 10.1038/nsmb.2805. [DOI] [PubMed] [Google Scholar]
- 10.Huang TT, D’Andrea AD. Regulation of DNA repair by ubiquitylation. Nat. Rev. Mol. Cell Biol. 2006;7:323–334. doi: 10.1038/nrm1908. [DOI] [PubMed] [Google Scholar]
- 11.Aguilar RC, Wendland B. Ubiquitin: not just for proteasomes anymore. Curr. Opin. Cell Biol. 2003;15:184–190. doi: 10.1016/s0955-0674(03)00010-3. [DOI] [PubMed] [Google Scholar]
- 12.Haglund K, Di Fiore PP, Dikic I. Distinct monoubiquitin signals in receptor endocytosis. Trends Biochem. Sci. 2003;28:598–603. doi: 10.1016/j.tibs.2003.09.005. [DOI] [PubMed] [Google Scholar]
- 13.Hicke L. Protein regulation by monoubiquitin. Nat. Rev. Mol. Cell Biol. 2001;2:195–201. doi: 10.1038/35056583. [DOI] [PubMed] [Google Scholar]
- 14.Sun L, Chen ZJ. The novel functions of ubiquitination in signaling. Curr. Opin. Cell Biol. 2004;16:119–126. doi: 10.1016/j.ceb.2004.02.005. [DOI] [PubMed] [Google Scholar]
- 15.Kessler BM. Ubiquitin — omics reveals novel networks and associations with human disease. Curr. Opin. Chem. Biol. 2013;17:59–65. doi: 10.1016/j.cbpa.2012.12.024. [DOI] [PubMed] [Google Scholar]
- 16.Hoeller D, Hecker C-M, Dikic I. Ubiquitin and ubiquitin-like proteins in cancer pathogenesis. Nat. Rev. Cancer. 2006;6:776–788. doi: 10.1038/nrc1994. [DOI] [PubMed] [Google Scholar]
- 17.Ciechanover A, Schwartz AL. The ubiquitin system: pathogenesis of human diseases and drug targeting. Biochim. Biophys. Acta. 2004;1695:3–17. doi: 10.1016/j.bbamcr.2004.09.018. [DOI] [PubMed] [Google Scholar]
- 18.Reinstein E, Ciechanover A. Narrative review: protein degradation and human diseases: the ubiquitin connection. Ann. Intern. Med. 2006;145:676–684. doi: 10.7326/0003-4819-145-9-200611070-00010. [DOI] [PubMed] [Google Scholar]
- 19.Petroski MD. The ubiquitin system, disease, and drug discovery. BMC Biochem. 2008;9(Suppl 1):S7. doi: 10.1186/1471-2091-9-S1-S7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Nalepa G, Rolfe M, Harper JW. Drug discovery in the ubiquitin-proteasome system. Nat. Rev. Drug Discov. 2006;5:596–613. doi: 10.1038/nrd2056. [DOI] [PubMed] [Google Scholar]
- 21.Wong BR, Parlati F, Qu K, Demo S, Pray T, Huang J, Payan DG, Bennett MK. Drug discovery in the ubiquitin regulatory pathway. Drug Discov. Today. 2003;8:746–754. doi: 10.1016/s1359-6446(03)02780-6. [DOI] [PubMed] [Google Scholar]
- 22.Hershko A, Ciechanover A. The ubiquitin system. Annu. Rev. Biochem. 1998;67:425–479. doi: 10.1146/annurev.biochem.67.1.425. [DOI] [PubMed] [Google Scholar]
- 23.Ciechanover A, Ben-Saadon R. N-terminal ubiquitination: more protein substrates join in. Trends Cell Biol. 2004;14:103–106. doi: 10.1016/j.tcb.2004.01.004. [DOI] [PubMed] [Google Scholar]
- 24.Cadwell K, Coscoy L. Ubiquitination on nonlysine residues by a viral E3 ubiquitin ligase. Science. 2005;309:127–130. doi: 10.1126/science.1110340. [DOI] [PubMed] [Google Scholar]
- 25.Kravtsova-Ivantsiv Y, Sommer T, Ciechanover A. The lysine48-based polyubiquitin chain proteasomal signal: not a single child anymore. Angew. Chem. Int. Ed. 2013;52:192–198. doi: 10.1002/anie.201205656. [DOI] [PubMed] [Google Scholar]
- 26.Jentsch S. Ubiquitin-dependent protein degradation: a cellular perspective. Trends Cell Biol. 1992;2:98–103. doi: 10.1016/0962-8924(92)90013-d. [DOI] [PubMed] [Google Scholar]
- 27.Ciechanover A. The ubiquitin-proteasome pathway: on protein death and cell life. EMBO J. 1998;17:7151–7160. doi: 10.1093/emboj/17.24.7151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Glickman MH, Ciechanover A. The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol. Rev. 2002;82:373–428. doi: 10.1152/physrev.00027.2001. [DOI] [PubMed] [Google Scholar]
- 29.Kirisako T, Kamei K, Murata S, Kato M, Fukumoto H, Kanie M, Sano S, Tokunaga F, Tanaka K, Iwai K. A ubiquitin ligase complex assembles linear polyubiquitin chains. EMBO J. 2006;25:4877–4887. doi: 10.1038/sj.emboj.7601360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hofmann RM, Pickart CM. In vitro assembly and recognition of Lys-63 polyubiquitin chains. J. Biol. Chem. 2001;276:27936–27943. doi: 10.1074/jbc.M103378200. [DOI] [PubMed] [Google Scholar]
- 31.Welchman RL, Gordon C, Mayer RJ. Ubiquitin and ubiquitin-like proteins as multifunctional signals. Nat. Rev. Mol. Cell Biol. 2005;6:599–609. doi: 10.1038/nrm1700. [DOI] [PubMed] [Google Scholar]
- 32.Dimova NV, Hathaway NA, Lee B-H, Kirkpatrick DS, Berkowitz ML, Gygi SP, Finley D, King RW. APC/C-mediated multiple monoubiquitylation provides an alternative degradation signal for cyclin B1. Nat. Cell Biol. 2012;14:168–176. doi: 10.1038/ncb2425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Dammer EB, Na CH, Xu P, Seyfried NT, Duong DM, Cheng D, Gearing M, Rees H, Lah JJ, Levey AI, Rush J, Peng J. Polyubiquitin linkage profiles in three models of proteolytic stress suggest the etiology of Alzheimer disease. J. Biol. Chem. 2011;286:10457–10465. doi: 10.1074/jbc.M110.149633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Koegl M, Hoppe T, Schlenker S, Ulrich HD, Mayer TU, Jentsch S. A novel ubiquitination factor, E4, is involved in multiubiquitin chain assembly. Cell. 1999;96:635–644. doi: 10.1016/s0092-8674(00)80574-7. [DOI] [PubMed] [Google Scholar]
- 35.Richly H, Rape M, Braun S, Rumpf S, Hoege C, Jentsch S. A series of ubiquitin binding factors connects CDC48/p97 to substrate multiubiquitylation and proteasomal targeting. Cell. 2005;120:73–84. doi: 10.1016/j.cell.2004.11.013. [DOI] [PubMed] [Google Scholar]
- 36.Amerik AY, Hochstrasser M. Mechanism and function of deubiquitinating enzymes. Biochim. Biophys. Acta BBA - Mol. Cell Res. 2004;1695:189–207. doi: 10.1016/j.bbamcr.2004.10.003. [DOI] [PubMed] [Google Scholar]
- 37.Cajee U-F, Hull R, Ntwasa M. Modification by ubiquitin-like proteins: significance in apoptosis and autophagy pathways. Int. J. Mol. Sci. 2012;13:11804–11831. doi: 10.3390/ijms130911804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Herrmann J, Lerman LO, Lerman A. Ubiquitin and ubiquitin-like proteins in protein regulation. Circ. Res. 2007;100:1276–1291. doi: 10.1161/01.RES.0000264500.11888.f0. [DOI] [PubMed] [Google Scholar]
- 39.Meulmeester E, Melchior F. Cell biology: SUMO. Nature. 2008;452:709–711. doi: 10.1038/452709a. [DOI] [PubMed] [Google Scholar]
- 40.Hochstrasser M. Origin and function of ubiquitin-like proteins. Nature. 2009;458:422–429. doi: 10.1038/nature07958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kerscher O. SUMO junction—what’s your function? New insights through SUMO-interacting motifs. EMBO Rep. 2007;8:550–555. doi: 10.1038/sj.embor.7400980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Droescher M, Chaugule VK, Pichler A. SUMO Rules: Regulatory Concepts and Their Implication in Neurologic Functions. NeuroMolecular Med. 2013;15:639–660. doi: 10.1007/s12017-013-8258-6. [DOI] [PubMed] [Google Scholar]
- 43.Bettermann K, Benesch M, Weis S, Haybaeck J. SUMOylation in carcinogenesis. Cancer Lett. 2012;316:113–125. doi: 10.1016/j.canlet.2011.10.036. [DOI] [PubMed] [Google Scholar]
- 44.Krumova P, Weishaupt JH. Sumoylation in neurodegenerative diseases. Cell. Mol. Life Sci. 2013;70:2123–2138. doi: 10.1007/s00018-012-1158-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Rajan S, Torres J, Thompson MS, Philipson LH. SUMO downregulates GLP-1-stimulated cAMP generation and insulin secretion. AJP Endocrinol. Metab. 2012;302:E714–E723. doi: 10.1152/ajpendo.00486.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Haas AL, Ahrens P, Bright PM, Ankel H. Interferon induces a 15-kilodalton protein exhibiting marked homology to ubiquitin. J. Biol. Chem. 1987;262:11315–11323. [PubMed] [Google Scholar]
- 47.Zou W, Zhang D-E. The interferon-inducible ubiquitin-protein isopeptide ligase (E3) EFP also functions as an ISG15 E3 ligase. J. Biol. Chem. 2006;281:3989–3994. doi: 10.1074/jbc.M510787200. [DOI] [PubMed] [Google Scholar]
- 48.Dastur A, Beaudenon S, Kelley M, Krug RM, Huibregtse JM. Herc5 an interferon-induced HECT E3 enzyme, is required for conjugation of ISG15 in human cells. J. Biol. Chem. 2006;281:4334–4338. doi: 10.1074/jbc.M512830200. [DOI] [PubMed] [Google Scholar]
- 49.Jeon YJ, Yoo HM, Chung CH. ISG15 and immune diseases. Biochim. Biophys. Acta BBA - Mol. Basis Dis. 2010;1802:485–496. doi: 10.1016/j.bbadis.2010.02.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Morales DJ, Lenschow DJ. The antiviral activities of ISG15. J. Mol. Biol. 2013;425:4995–5008. doi: 10.1016/j.jmb.2013.09.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Desai SD. Elevated expression of ISG15 in tumor cells interferes with the ubiquitin/26S proteasome pathway. Cancer Res. 2006;66:921–928. doi: 10.1158/0008-5472.CAN-05-1123. [DOI] [PubMed] [Google Scholar]
- 52.Kamitani T, Kito K, Nguyen HP, Yeh ETH, Characterization of NEDD8. a developmentally down-regulated ubiquitin-like protein. J. Biol. Chem. 1997;272:28557–28562. doi: 10.1074/jbc.272.45.28557. [DOI] [PubMed] [Google Scholar]
- 53.Whitby FG, Xia G, Pickart CM, Hill CP. Crystal structure of the human ubiquitin-like protein NEDD8 and interactions with ubiquitin pathway enzymes. J. Biol. Chem. 1998;273:34983–34991. doi: 10.1074/jbc.273.52.34983. [DOI] [PubMed] [Google Scholar]
- 54.Pan Z-Q, Kentsis A, Dias DC, Yamoah K, Wu K. Nedd8 on cullin: building an expressway to protein destruction. Oncogene. 2004;23:1985–1997. doi: 10.1038/sj.onc.1207414. [DOI] [PubMed] [Google Scholar]
- 55.Xirodimas DP, Sundqvist A, Nakamura A, Shen L, Botting C, Hay RT. Ribosomal proteins are targets for the NEDD8 pathway. EMBO Rep. 2008;9:280–286. doi: 10.1038/embor.2008.10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Lin D-H, Sterling H, Wang Z, Babilonia E, Yang B, Dong K, Hebert SC, Giebisch G, Wang W-H. ROMK1 channel activity is regulated by monoubiquitination. Proc. Natl. Acad. Sci. U. S. A. 2005;102:4306–4311. doi: 10.1073/pnas.0409767102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Okumoto K, Noda H, Fujiki Y. Distinct modes of ubiquitination of peroxisome-targeting signal type 1 (PTS1)-receptor Pex5p regulate PTS1 protein import. J. Biol. Chem. 2014;289:14089–108. doi: 10.1074/jbc.M113.527937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Gentry MS, Worby CA, Dixon JE. Insights into Lafora disease: malin is an E3 ubiquitin ligase that ubiquitinates and promotes the degradation of laforin. Proc. Natl. Acad. Sci. U. S. A. 2005;102:8501–8506. doi: 10.1073/pnas.0503285102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Malakhov MP, Kim KI, Malakhova OA, Jacobs BS, Borden EC, Zhang D-E. High-throughput immunoblotting. Ubiquitiin-like protein ISG15 modifies key regulators of signal transduction. J. Biol. Chem. 2003;278:16608–16613. doi: 10.1074/jbc.M208435200. [DOI] [PubMed] [Google Scholar]
- 60.Wagner SA, Beli P, Weinert BT, Nielsen ML, Cox J, Mann M, Choudhary C. A, proteome-wide, quantitative survey of in vivo ubiquitylation sites reveals widespread regulatory roles. Mol. Cell. Proteomics. 2011;10 doi: 10.1074/mcp.M111.013284. M111.013284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Danielsen JMR, Sylvestersen KB, Bekker-Jensen S, Szklarczyk D, Poulsen JW, Horn H, Jensen LJ, Mailand N, Nielsen ML. Mass spectrometric analysis of lysine ubiquitylation reveals promiscuity at site level. Mol. Cell. Proteomics. 2011;10 doi: 10.1074/mcp.M110.003590. M110.003590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Kim W, Bennett EJ, Huttlin EL, Guo A, Li J, Possemato A, Sowa ME, Rad R, Rush J, Comb MJ, Harper JW, Gygi SP. Systematic and quantitative assessment of the ubiquitin-modified proteome. Mol. Cell. 2011;44:325–340. doi: 10.1016/j.molcel.2011.08.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Udeshi ND, Mani DR, Eisenhaure T, Mertins P, Jaffe JD, Clauser KR, Hacohen N, Carr SA. Methods for quantification of in vivo changes in protein ubiquitination following proteasome and deubiquitinase inhibition. Mol. Cell. Proteomics. 2012;11:148–159. doi: 10.1074/mcp.M111.016857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Udeshi ND, Mertins P, Svinkina T, Carr SA. Large-scale identification of ubiquitination sites by mass spectrometry. Nat. Protoc. 2013;8:1950–1960. doi: 10.1038/nprot.2013.120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Povlsen LK, Beli P, Wagner SA, Poulsen SL, Sylvestersen KB, Poulsen JW, Nielsen ML, Bekker-Jensen S, Mailand N, Choudhary C. Systems-wide analysis of ubiquitylation dynamics reveals a key role for PAF15 ubiquitylation in DNA-damage bypass. Nat. Cell Biol. 2012;14:1089–1098. doi: 10.1038/ncb2579. [DOI] [PubMed] [Google Scholar]
- 66.Matic I, Schimmel J, Hendriks IA, van Santen MA, van de Rijke F, van Dam H, Gnad F, Mann M, Vertegaal ACO. Site-specific identification of SUMO-2 targets in cells reveals an inverted SUMOylation motif and a hydrophobic cluster SUMOylation motif. Mol. Cell. 2010;39:641–652. doi: 10.1016/j.molcel.2010.07.026. [DOI] [PubMed] [Google Scholar]
- 67.Tammsalu T, Matic I, Jaffray EG, Ibrahim AFM, Tatham MH, Hay RT. Proteome-wide identification of SUMO2 modification sites. Sci. Signal. 2014;7:rs2. doi: 10.1126/scisignal.2005146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Chernorudskiy AL, Garcia A, Eremin EV, Shorina AS, Kondratieva EV, Gainullin MR. UbiProt: a database of ubiquitylated proteins. BMC Bioinformatics. 2007;8:126. doi: 10.1186/1471-2105-8-126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Lee W-C, Lee M, Jung JW, Kim KP, Kim D. SCUD: Saccharomyces Cerevisiae Ubiquitination Database. BMC Genomics. 2008;9:440. doi: 10.1186/1471-2164-9-440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Li H, Xing X, Ding G, Li Q, Wang C, Xie L, Zeng R, Li Y. SysPTM: A systematic resource for proteomic research on post-translational modifications. Mol. Cell. Proteomics. 2009;8:1839–1849. doi: 10.1074/mcp.M900030-MCP200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Liu Z, Wang Y, Gao T, Pan Z, Cheng H, Yang Q, Cheng Z, Guo A, Ren J, Xue Y. CPLM: a database of protein lysine modifications. Nucleic Acids Res. 2014;42:D531–D536. doi: 10.1093/nar/gkt1093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Catic A, Collins C, Church GM, Ploegh HL. Preferred in vivo ubiquitination sites. Bioinformatics. 2004;20:3302–3307. doi: 10.1093/bioinformatics/bth407. [DOI] [PubMed] [Google Scholar]
- 73.Tung C-W, Ho S-Y. Computational identification of ubiquitylation sites from protein sequences. BMC Bioinformatics. 2008;9:310. doi: 10.1186/1471-2105-9-310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Radivojac P, Vacic V, Haynes C, Cocklin RR, Mohan A, Heyen JW, Goebl MG, Iakoucheva LM. Identification, analysis, and prediction of protein ubiquitination sites. Proteins Struct. Funct. Bioinforma. 2010;78:365–380. doi: 10.1002/prot.22555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Cai Y, Huang T, Hu L, Shi X, Xie L, Li Y. Prediction of lysine ubiquitination with mRMR feature selection and analysis. Amino Acids. 2012;42:1387–1395. doi: 10.1007/s00726-011-0835-0. [DOI] [PubMed] [Google Scholar]
- 76.Chen Z, Chen Y-Z, Wang X-F, Wang C, Yan R-X, Zhang Z. Prediction of ubiquitination sites by using the composition of k-spaced amino acid pairs. PloS One. 2011;6:e22930. doi: 10.1371/journal.pone.0022930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Chen Z, Zhou Y, Song J, Zhang Z. hCKSAAP_UbSite: improved prediction of human ubiquitination sites by exploiting amino acid pattern and properties. Biochim. Biophys. Acta. 2013;1834:1461–1467. doi: 10.1016/j.bbapap.2013.04.006. [DOI] [PubMed] [Google Scholar]
- 78.Walsh I, Di Domenico T, Tosatto SCE. RUBI: rapid proteomic-scale prediction of lysine ubiquitination and factors influencing predictor performance. Amino Acids. 2014;46:853–862. doi: 10.1007/s00726-013-1645-3. [DOI] [PubMed] [Google Scholar]
- 79.Chen X, Qiu J-D, Shi S-P, Suo S-B, Huang S-Y, Liang R-P. Incorporating key position and amino acid residue features to identify general and species-specific Ubiquitin conjugation sites. Bioinforma. Oxf. Engl. 2013;29:1614–1622. doi: 10.1093/bioinformatics/btt196. [DOI] [PubMed] [Google Scholar]
- 80.Sampson DA, Wang M, Matunis MJ. The small ubiquitin-like modifier-1 (SUMO-1) consensus sequence mediates Ubc9 binding and is essential for SUMO-1 modification. J. Biol. Chem. 2001;276:21664–21669. doi: 10.1074/jbc.M100006200. [DOI] [PubMed] [Google Scholar]
- 81.Zhao Q, Xie Y, Zheng Y, Jiang S, Liu W, Mu W, Liu Z, Zhao Y, Xue Y, Ren J. GPS-SUMO: a tool for the prediction of sumoylation sites and SUMO-interaction motifs. Nucleic Acids Res. 2014;42:W325–W330. doi: 10.1093/nar/gku383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Xue Y, Zhou F, Zhu M, Ahmed K, Chen G, Yao X. GPS: a comprehensive www server for phosphorylation sites prediction. Nucleic Acids Res. 2005;33:W184–W187. doi: 10.1093/nar/gki393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Xue Y, Zhou F, Fu C, Xu Y, Yao X. SUMOsp: a web server for sumoylation site prediction. Nucleic Acids Res. 2006;34:W254–W257. doi: 10.1093/nar/gkl207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Zhou F-F, Xue Y, Chen G-L, Yao X. GPS: a novel group-based phosphorylation predicting and scoring method. Biochem. Biophys. Res. Commun. 2004;325:1443–1448. doi: 10.1016/j.bbrc.2004.11.001. [DOI] [PubMed] [Google Scholar]
- 85.Kim JH, Lee J, Oh B, Kimm K, Koh I. Prediction of phosphorylation sites using SVMs. Bioinformatics. 2004;20:3179–3184. doi: 10.1093/bioinformatics/bth382. [DOI] [PubMed] [Google Scholar]
- 86.Ren J, Gao X, Jin C, Zhu M, Wang X, Shaw A, Wen L, Yao X, Xue Y. Systematic study of protein sumoylation: Development of a site-specific predictor of SUMOsp 2.0. PROTEOMICS. 2009;9:3409–3412. doi: 10.1002/pmic.200800646. [DOI] [PubMed] [Google Scholar]
- 87.Teng S, Luo H, Wang L. Predicting protein sumoylation sites from sequence features. Amino Acids. 2012;43:447–455. doi: 10.1007/s00726-011-1100-2. [DOI] [PubMed] [Google Scholar]
- 88.Schwartz D. Prediction of lysine post-translational modifications using bioinformatic tools. Essays Biochem. 2012;52:165–177. doi: 10.1042/bse0520165. [DOI] [PubMed] [Google Scholar]
- 89.Emanuele MJ, Elia AEH, Xu Q, Thoma CR, Izhar L, Leng Y, Guo A, Chen Y-N, Rush J, Hsu PW-C, Yen H-CS, Elledge SJ. Global identification of modular cullin-RING ligase substrates. Cell. 2011;147:459–474. doi: 10.1016/j.cell.2011.09.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Lopitz-Otsoa F, Rodriguez-Suarez E, Aillet F, Casado-Vela J, Lang V, Matthiesen R, Elortza F, Rodriguez MS. Integrative analysis of the ubiquitin proteome isolated using Tandem Ubiquitin Binding Entities (TUBEs) J. Proteomics. 2012;75:2998–3014. doi: 10.1016/j.jprot.2011.12.001. [DOI] [PubMed] [Google Scholar]
- 91.Meierhofer D, Wang X, Huang L, Kaiser P. Quantitative analysis of global ubiquitination in HeLa cells by mass spectrometry. J. Proteome Res. 2008;7:4566–4576. doi: 10.1021/pr800468j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Mertins P, Qiao JW, Patel J, Udeshi ND, Clauser KR, Mani DR, Burgess MW, Gillette MA, Jaffe JD, Carr SA. Integrated proteomic analysis of post-translational modifications by serial enrichment. Nat. Methods. 2013;10:634–637. doi: 10.1038/nmeth.2518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Xu G, Deglincerti A, Paige JS, Jaffrey SR. Profiling lysine ubiquitination by selective enrichment of ubiquitin remnant-containing peptides. Methods Mol. Biol. Clifton NJ. 2014;1174:57–71. doi: 10.1007/978-1-4939-0944-5_4. [DOI] [PubMed] [Google Scholar]
- 94.Vertegaal ACO, Andersen JS, Ogg SC, Hay RT, Mann M, Lamond AI. Distinct and overlapping sets of SUMO-1 and SUMO-2 target proteins revealed by quantitative proteomics. Mol. Cell. Proteomics. 2006;12:2298–2310. doi: 10.1074/mcp.M600212-MCP200. [DOI] [PubMed] [Google Scholar]
- 95.Bursomanno S, Beli P, Khan AM, Minocherhomji S, Wagner SA, Bekker-Jensen S, Mailand N, Choudhary C, Hickson ID, Liu Y. Proteome-wide analysis of SUMO2 targets in response to pathological DNA replication stress in human cells. DNA Repair. 2015;25:84–96. doi: 10.1016/j.dnarep.2014.10.011. [DOI] [PubMed] [Google Scholar]
- 96.Hendriks IA, D’Souza RC, Yang B, Verlaan-de Vries M, Mann M, Vertegaal AC. Uncovering global SUMOylation signaling networks in a site-specific manner. Nat Struct Mol Biol. 2014;10:927–36. doi: 10.1038/nsmb.2890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Makarov A, Denisov E, Kholomeev A, Balschun W, Lange O, Strupat K, Horning S. Performance evaluation of a hybrid linear ion trap/orbitrap mass spectrometer. Anal. Chem. 2006;78:2113–2120. doi: 10.1021/ac0518811. [DOI] [PubMed] [Google Scholar]
- 98.Zubarev R, Mann M. On the proper use of mass accuracy in proteomics. Mol. Cell. Proteomics. 2007;6:377–381. doi: 10.1074/mcp.M600380-MCP200. [DOI] [PubMed] [Google Scholar]
- 99.Marshall AG, Hendrickson CL. High-resolution mass spectrometers. Annu. Rev. Anal. Chem. 2008;1:579–599. doi: 10.1146/annurev.anchem.1.031207.112945. [DOI] [PubMed] [Google Scholar]
- 100.Peng J, Elias JE, Thoreen CC, Licklider LJ, Gygi SP. Evaluation of multidimensional chromatography coupled with tandem mass spectrometry (LC/LC-MS/MS) for large-scale protein analysis: the yeast proteome. J. Proteome Res. 2003;2:43–50. doi: 10.1021/pr025556v. [DOI] [PubMed] [Google Scholar]
- 101.Franco M, Seyfried NT, Brand AH, Peng J, Mayor U. A novel strategy to isolate ubiquitin conjugates reveals wide role for ubiquitination during neural development. Mol. Cell. Proteomics. 2011;10 doi: 10.1074/mcp.M110.002188. M110.002188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Xu G, Paige JS, Jaffrey SR. Global analysis of lysine ubiquitination by ubiquitin remnant immunoaffinity profiling. Nat. Biotechnol. 2010;28:868–873. doi: 10.1038/nbt.1654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Matsumoto ML, Wickliffe KE, Dong KC, Yu C, Bosanac I, Bustos D, Phu L, Kirkpatrick DS, Hymowitz SG, Rape M, Kelley RF, Dixit VM. K11-linked polyubiquitination in cell cycle control revealed by a K11 linkage-specific antibody. Mol. Cell. 2010;39:477–484. doi: 10.1016/j.molcel.2010.07.001. [DOI] [PubMed] [Google Scholar]
- 104.Shi Y, Chan DW, Jung SY, Malovannaya A, Wang Y, Qin J. A data set of human endogenous protein ubiquitination sites. Mol. Cell. Proteomics. 2011;10 doi: 10.1074/mcp.M110.002089. M110.002089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Shi Y, Xu P, Qin J. Ubiquitinated proteome: ready for global? Mol. Cell. Proteomics. 2011;10 doi: 10.1074/mcp.R110.006882. R110.006882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Hjerpe R, Thomas Y, Kurz T. NEDD8 overexpression results in neddylation of ubiquitin substrates by the ubiquitin pathway. J. Mol. Biol. 2012;421:27–29. doi: 10.1016/j.jmb.2012.05.013. [DOI] [PubMed] [Google Scholar]
- 107.Matsumoto ML, Dong KC, Yu C, Phu L, Gao X, Hannoush RN, Hymowitz SG, Kirkpatrick DS, Dixit VM, Kelley RF. Engineering and structural characterization of a linear polyubiquitin-specific antibody. J. Mol. Biol. 2012;418:134–144. doi: 10.1016/j.jmb.2011.12.053. [DOI] [PubMed] [Google Scholar]
- 108.Lee KA, Hammerle LP, Andrews PS, Stokes MP, Mustelin T, Silva JC, Black RA, Doedens JR. Ubiquitin ligase substrate identification through quantitative proteomics at both the protein and peptide levels. J. Biol. Chem. 2011;286:41530–41538. doi: 10.1074/jbc.M111.248856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Sarraf SA, Raman M, Guarani-Pereira V, Sowa ME, Huttlin EL, Gygi SP, Harper JW. Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization. Nature. 2013;496:372–376. doi: 10.1038/nature12043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Stes E, Laga M, Walton A, Samyn N, Timmerman E, De Smet I, Goormachtig S, Gevaert K. A COFRADIC Protocol To Study Protein Ubiquitination. J. Proteome Res. 2014;13:3107–3113. doi: 10.1021/pr4012443. [DOI] [PubMed] [Google Scholar]
- 111.Golebiowski F, Matic I, Tatham MH, Cole C, Yin Y, Nakamura A, Cox J, Barton GJ, Mann M, Hay RT. System-wide changes to SUMO modifications in response to heat shock. Sci Signal. 2009;72:ra24. doi: 10.1126/scisignal.2000282. [DOI] [PubMed] [Google Scholar]
- 112.Bruderer R, Tatham MH, Plechanovova A, Matic I, Garg AK, Hay RT. Purification and identification of endogenous polySUMO conjugates. EMBO Rep. 2011;12:142–8. doi: 10.1038/embor.2010.206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Becker J, Barysch SV, Karaca S, Dittner C, Hsiao HH, Berriel Diaz M, Herzig S, Urlaub H. F. Melchior Detecting endogenous SUMO targets in mammalian cells and tissues. Nat Struct Mol Biol. 2013;20:525–31. doi: 10.1038/nsmb.2526. [DOI] [PubMed] [Google Scholar]
- 114.Zhao Y, Kwon SW, Anselmo A, Kaur K, White MA. Broad spectrum identification of cellular small ubiquitin-related modifier (SUMO) substrate proteins. J. Biol. Chem. 2004;279:20999–21002. doi: 10.1074/jbc.M401541200. [DOI] [PubMed] [Google Scholar]
- 115.Filosa G, Barabino SML, Bachi A. Proteomics strategies to identify SUMO targets and acceptor sites: a survey of RNA-binding proteins SUMOylation. Neuromolecular Med. 2013;15:661–676. doi: 10.1007/s12017-013-8256-8. [DOI] [PubMed] [Google Scholar]
- 116.Wilson VG, Heaton PR. Ubiquitin proteolytic system: focus on SUMO. Expert Rev. Proteomics. 2008;5:121–135. doi: 10.1586/14789450.5.1.121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Chicooree N, Griffiths JR, Connolly Y, Tan C-T, Malliri A, Eyers CE, Smith DL. A novel approach to the analysis of SUMOylation with the independent use of trypsin and elastase digestion followed by database searching utilising consecutive residue addition to lysine. Rapid Commun. Mass Spectrom. 2013;27:127–134. doi: 10.1002/rcm.6425. [DOI] [PubMed] [Google Scholar]
- 118.Jones J, Wu K, Yang Y, Guerrero C, Nillegoda N, Pan Z-Q, Huang L. A targeted proteomic analysis of the ubiquitin-like modifier nedd8 and associated proteins. J. Proteome Res. 2008;7:1274–1287. doi: 10.1021/pr700749v. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Zhao C, Denison C, Huibregtse JM, Gygi S, Krug RM. Human ISG15 conjugation targets both IFN-induced and constitutively expressed proteins functioning in diverse cellular pathways. Proc. Natl. Acad. Sci. 2005;102:10200–10205. doi: 10.1073/pnas.0504754102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Lopes M, Foiani M, Sogo JM. Multiple mechanisms control chromosome integrity after replication fork uncoupling and restart at irreparable UV lesions. Mol. Cell. 2006;21:15–27. doi: 10.1016/j.molcel.2005.11.015. [DOI] [PubMed] [Google Scholar]
- 121.Branzei D. Ubiquitin family modifications and template switching. FEBS Lett. 2011;585:2810–2817. doi: 10.1016/j.febslet.2011.04.053. [DOI] [PubMed] [Google Scholar]
- 122.Xiao W, Chow BL, Broomfield S, Hanna M. The Saccharomyces cerevisiae RAD6 Group Is Composed of an Error-Prone and Two Error-Free Postreplication Repair Pathways. Genetics. 2000;155:1633–1641. doi: 10.1093/genetics/155.4.1633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Broomfield S, Hryciw T, Xiao W. DNA postreplication repair and mutagenesis in Saccharomyces cerevisiae . Mutat Res. 2001;486:167–184. doi: 10.1016/s0921-8777(01)00091-x. [DOI] [PubMed] [Google Scholar]
- 124.Woodgate R. Evolution of the two-step model for UV-mutagenesis. Mutat. Res. 2001;485:83–92. doi: 10.1016/s0921-8777(00)00076-8. [DOI] [PubMed] [Google Scholar]
- 125.Krishna TS, Kong XP, Gary S, Burgers PM, Kuriyan J. Crystal structure of the eukaryotic DNA polymerase processivity factor PCNA. Cell. 1994;79:1233–1243. doi: 10.1016/0092-8674(94)90014-0. [DOI] [PubMed] [Google Scholar]
- 126.Kirchmaier AL. Ub-family modifications at the replication fork: Regulating PCNA-interacting components. FEBS Lett. 2011;585:2920–2928. doi: 10.1016/j.febslet.2011.08.008. [DOI] [PubMed] [Google Scholar]
- 127.Warbrick E. The puzzle of PCNA’s many partners. BioEssays News Rev. Mol. Cell. Dev. Biol. 2011;22:997–1006. doi: 10.1002/1521-1878(200011)22:11<997::AID-BIES6>3.0.CO;2-#. [DOI] [PubMed] [Google Scholar]
- 128.Moldovan G-L, Pfander B, Jentsch S. PCNA the maestro of the replication fork. Cell. 2007;129:665–679. doi: 10.1016/j.cell.2007.05.003. [DOI] [PubMed] [Google Scholar]
- 129.Hoege C, Pfander B, Moldovan G-L, Pyrowolakis G, Jentsch S. RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature. 2002;419:135–141. doi: 10.1038/nature00991. [DOI] [PubMed] [Google Scholar]
- 130.Parker JL, Ulrich HD. Mechanistic analysis of PCNA poly-ubiquitylation by the ubiquitin protein ligases Rad18 and Rad5. EMBO J. 2009;28:3657–3666. doi: 10.1038/emboj.2009.303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Arakawa H, Moldovan G-L, Saribasak H, Saribasak NN, Jentsch S, Buerstedde J-M. A role for PCNA ubiquitination in immunoglobulin hypermutation. PLoS Biol. 2006;4:e366. doi: 10.1371/journal.pbio.0040366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Simpson LJ, Ross A-L, Szüts D, Alviani CA, Oestergaard VH, Patel KJ, Sale JE. RAD18-independent ubiquitination of proliferating-cell nuclear antigen in the avian cell line DT40. EMBO Rep. 2006;7:927–932. doi: 10.1038/sj.embor.7400777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Kats ES, Enserink JM, Martinez S, Kolodner RD. The Saccharomyces cerevisiae Rad6 Postreplication repair and Siz1/Srs2 homologous recombination-inhibiting pathways process DNA damage that arises in asf1 mutants. Mol. Cell. Biol. 2009;29:5226–5237. doi: 10.1128/MCB.00894-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Terai K, Abbas T, Jazaeri AA, Dutta A. CRL4(Cdt2) E3 ubiquitin ligase monoubiquitinates PCNA to promote translesion DNA synthesis. Mol. Cell. 2010;37:143–149. doi: 10.1016/j.molcel.2009.12.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Zhang S, Chea J, Meng X, Zhou Y, Lee EYC, Lee MYWT. PCNA is ubiquitinated by RNF8. Cell Cycle Georget. Tex. 2008;7:3399–3404. doi: 10.4161/cc.7.21.6949. [DOI] [PubMed] [Google Scholar]
- 136.Huang TT, Nijman SMB, Mirchandani KD, Galardy PJ, Cohn MA, Haas W, Gygi SP, Ploegh HL, Bernards R, D’Andrea AD. Regulation of monoubiquitinated PCNA by DUB autocleavage. Nat. Cell Biol. 2006;8:339–347. doi: 10.1038/ncb1378. [DOI] [PubMed] [Google Scholar]
- 137.Niimi A, Brown S, Sabbioneda S, Kannouche PL, Scott A, Yasui A, Green CM, Lehmann AR. Regulation of proliferating cell nuclear antigen ubiquitination in mammalian cells. Proc. Natl. Acad. Sci. U. S. A. 2008;105:16125–16130. doi: 10.1073/pnas.0802727105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Brun J, Chiu RK, Wouters BG, Gray DA. Regulation of PCNA polyubiquitination in human cells. BMC Res. Notes. 2010;3 doi: 10.1186/1756-0500-3-85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Whitehurst CB, Vaziri C, Shackelford J, Pagano JS. Epstein-Barr virus BPLF1 deubiquitinates PCNA and attenuates polymerase η recruitment to DNA damage sites. J. Virol. 2012;86:8097–8106. doi: 10.1128/JVI.00588-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Motegi A, Sood R, Moinova H, Markowitz SD, Liu PP, Myung K. Human SHPRH suppresses genomic instability through proliferating cell nuclear antigen polyubiquitination. J. Cell Biol. 2006;175:703–708. doi: 10.1083/jcb.200606145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Motegi A, Liaw H-J, Lee K-Y, Roest HP, Maas A, Wu X, Moinova H, Markowitz SD, Ding H, Hoeijmakers JHJ, Myung K. Polyubiquitination of proliferating cell nuclear antigen by HLTF and SHPRH prevents genomic instability from stalled replication forks. Proc. Natl. Acad. Sci. U. S. A. 2008;105:12411–12416. doi: 10.1073/pnas.0805685105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Unk I, Hajdú I, Fátyol K, Szakál B, Blastyák A, Bermudez VJ, Hurwitz J, Prakash L, Prakash S, Haracska L. Human SHPRH is a ubiquitin ligase for Mms2-Ubc13-dependent polyubiquitylation of proliferating cell nuclear antigen. Proc Natl Acad Sci U S A. 2006;103:18107–12. doi: 10.1073/pnas.0608595103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Unk I, Hajdú I, Fátyol K, Hurwitz J, Yoon J, Prakash L, Prakash S, Haracska L. Human HLTF functions as a ubiquitin ligase for proliferating cell nuclear antigen polyubiquitination. PNAS. 2008;105:3768–3773. doi: 10.1073/pnas.0800563105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Lin J-R, Zeman MK, Chen J-Y, Yee M-C, Cimprich KA. SHPRH and HLTF act in a damage-specific manner to coordinate different forms of postreplication repair and prevent mutagenesis. Mol Cell. 2011;48:237–249. doi: 10.1016/j.molcel.2011.02.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Krijger PH, Lee KY, Wit N, van den Berk PC, Wu X, Roest HP, Maas A, Ding JH, Hoeijmakers JH, Myung K, Jacobs H. HLTF and SHPRH are not essential for PCNA polyubiquitination, survival and somatic hypermutation: existence of an alternative E3 ligase. DNA Repair. 2011;10:438–44. doi: 10.1016/j.dnarep.2010.12.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Brun J, Chiu R, Lockhart K, Xiao W, Wouters BG, Gray DA. hMMS2 serves a redundant role in human PCNA polyubiquitination. BMC Mol Biol. 2008;9:9–24. doi: 10.1186/1471-2199-9-24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Yu Y, Cai J-P, Tu B, Wu L, Zhao Y, Liu X, Li L, McNutt MA, Feng J, He Q, Yang Y, Wang H, Sekiguchi M, Zhu W-G. Proliferating cell nuclear antigen is protected from degradation by forming a complex with MutT Homolog2. J. Biol. Chem. 2009;284:19310–19320. doi: 10.1074/jbc.M109.015289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Groehler AL, Lannigan DA. A chromatin-bound kinase, ERK8, protects genomic integrity by inhibiting HDM2-mediated degradation of the DNA clamp PCNA. J. Cell Biol. 2010;190:575–586. doi: 10.1083/jcb.201002124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Yang Q, Ou C, Liu M, Xiao W, Wen C, Sun F. NRAGE promotes cell proliferation by stabilizing PCNA in a ubiquitin-proteasome pathway in esophageal carcinomas. Carcinogenesis. 2014;35:1643–1651. doi: 10.1093/carcin/bgu084. [DOI] [PubMed] [Google Scholar]
- 150.Das-Bradoo S, Nguyen HD, Wood JL, Ricke RM, Haworth JC, Bielinsky A-K. Defects in DNA ligase I trigger PCNA ubiquitylation at Lys 107. Nat. Cell Biol. 2010;12:74–79. doi: 10.1038/ncb2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Leach CA, Michael WM. Ubiquitin/SUMO modification of PCNA promotes replication fork progression in Xenopus laevis egg extracts. J. Cell Biol. 2005;171:947–954. doi: 10.1083/jcb.200508100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Gali H, Juhasz S, Morocz M, Hajdu I, Fatyol K, Szukacsov V, Burkovics P, Haracska L. Role of SUMO modification of human PCNA at stalled replication fork. Nucleic Acids Res. 2012;40:6049–6059. doi: 10.1093/nar/gks256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Moldovan G-L, Dejsuphong D, Petalcorin MIR, Hofmann K, Takeda S, Boulton SJ, D’Andrea AD. Inhibition of homologous recombination by the PCNA-interacting protein PARI. Mol. Cell. 2012;45:75–86. doi: 10.1016/j.molcel.2011.11.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Panse VG, Küster B, Gerstberger T, Hurt E. Unconventional tethering of Ulp1 to the transport channel of the nuclear pore complex by karyopherins. Nat. Cell Biol. 2003;5:21–27. doi: 10.1038/ncb893. [DOI] [PubMed] [Google Scholar]
- 155.Papouli E, Chen S, Davies AA, Huttner D, Krejci L, Sung P, Ulrich HD. Crosstalk between SUMO and ubiquitin on PCNA is mediated by recruitment of the helicase Srs2p. Mol. Cell. 2005;19:123–133. doi: 10.1016/j.molcel.2005.06.001. [DOI] [PubMed] [Google Scholar]
- 156.Kim SO, Yoon H, Park SO, Lee M, Shin J-S, Ryu K-S, Lee J-O, Seo Y-S, Jung HS, Choi B-S. Srs2 possesses a non-canonical PIP box in front of its SBM for precise recognition of SUMOylated PCNA. J. Mol. Cell Biol. 2012;4:258–261. doi: 10.1093/jmcb/mjs026. [DOI] [PubMed] [Google Scholar]
- 157.Kubota T, Nishimura K, Kanemaki MT, Donaldson AD. The Elg1 replication factor C- like complex functions in PCNA unloading during DNA replication. Mol. Cell. 2013;50:273–280. doi: 10.1016/j.molcel.2013.02.012. [DOI] [PubMed] [Google Scholar]
- 158.Parnas O, Zipin-Roitman A, Pfander B, Liefshitz B, Mazor Y, Ben-Aroya S, Jentsch S, Kupiec M. Elg1, an alternative subunit of the RFC clamp loader, preferentially interacts with SUMOylated. PCNA. EMBO J. 2010;29:2611–22. doi: 10.1038/emboj.2010.128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Lee KY, Yang K, Cohn MA, Sikdar N, D’Andrea AD, Myung K. Human ELG1 regulates the level of ubiquitinated proliferating cell nuclear antigen (PCNA) through Its interactions with PCNA and USP1. J Biol Chem. 2010;285:10362–9. doi: 10.1074/jbc.M109.092544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Lee KY, Fu H, Aladjem MI, Myung K. ATAD5 regulates the lifespan of DNA replication factories by modulating PCNA level on the chromatin. J Cell Biol. 2013;200:31–44. doi: 10.1083/jcb.201206084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Skibbens RV. Establishment of sister chromatid cohesion. Curr. Biol. 2009;19:R1126–R1132. doi: 10.1016/j.cub.2009.10.067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Park JM, Yang SW, Yu KR, Ka SH, Lee SW, Seol JH, Jeon YJ, Chung CH. Modification of PCNA by ISG15 plays a crucial role in termination of error-prone translesion DNA synthesis. Mol. Cell. 2014;54:626–638. doi: 10.1016/j.molcel.2014.03.031. [DOI] [PubMed] [Google Scholar]
- 163.Caron C, Boyault C, Khochbin S. Regulatory cross-talk between lysine acetylation and ubiquitination: role in the control of protein stability. BioEssays News Rev. Mol. Cell. Dev. Biol 27. 2005;4:408–415. doi: 10.1002/bies.20210. [DOI] [PubMed] [Google Scholar]
- 164.Naryzhny SN, Lee H. The post-translational modifications of proliferating cell nuclear antigen: acetylation not phosphorylation, plays an important role in the regulation of its function. J. Biol. Chem. 2004;279:20194–20199. doi: 10.1074/jbc.M312850200. [DOI] [PubMed] [Google Scholar]
- 165.Zhang Z, Zhang S, Lin SHS, Wang X, L Wu, Lee EYC, Lee MYWT. Structure of monoubiquitinated PCNA: implications for DNA polymerase switching and Okazaki fragment maturation. Cell Cycle Georget. Tex. 2012;11:2128–2136. doi: 10.4161/cc.20595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Kim BJ, Lee H. Lys-110 is essential for targeting PCNA to replication repair foci, and the K110A mutant activates apoptosis. Biol. Cell. 2008;100:675–686. doi: 10.1042/BC20070158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Goodman MF, Woodgate R. Translesion DNA Polymerases. Cold Spring Harb. Perspect. Biol. 2013;5:a010363–a010363. doi: 10.1101/cshperspect.a010363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Waters SL, Minesinger BK, Wiltrout ME, D’Souza S, Woodruff RV, Walker GC. Eukaryotic translesion polymerases and their roles and regulation in DNA damage Tolerance. Microbiol. Mol. Biol. Rev. 2009;73:134–154. doi: 10.1128/MMBR.00034-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Bienko M, Green CM, Crosetto N, Rudolf F, Zapart G, Coull B, Kannouche P, Wider G, Peter M, Lehmann AR, Hofmann K, Dikic I. Ubiquitin-binding domains in Y-family polymerases regulate translesion synthesis. Science. 2005;310:1821–1824. doi: 10.1126/science.1120615. [DOI] [PubMed] [Google Scholar]
- 170.Bomar MG, D’Souza S, Bienko M, Dikic I, Walker GC, Zhou P. Unconventional ubiquitin recognition by the ubiquitin-binding motif within the Y Family DNA polymerases ι and Rev1. Mol. Cell. 2010;37:408–417. doi: 10.1016/j.molcel.2009.12.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.Bienko M, Green CM, Sabbioneda S, Crosetto N, Matic I, Hibbert RG, Begovic T, Niimi A, Mann M, Lehmann AR, Dikic I. Regulation of translesion synthesis DNA polymerase η by monoubiquitination. Mol. Cell. 2010;37:396–407. doi: 10.1016/j.molcel.2009.12.039. [DOI] [PubMed] [Google Scholar]
- 172.Skoneczna A, McIntyre J, Skoneczny M, Policinska Z, Sledziewska-Gojska E. Polymerase eta is a short-lived, proteasomally degraded protein that is temporarily stabilized following UV irradiation in Saccharomyces cerevisiae . J. Mol. Biol. 2007;366:1074–1086. doi: 10.1016/j.jmb.2006.11.093. [DOI] [PubMed] [Google Scholar]
- 173.Pabla R, Rozario D, Siede W. Regulation of Saccharomyces cerevisiae DNA polymerase η transcript and protein. Radiat. Environ. Biophys. 2008;47:157–168. doi: 10.1007/s00411-007-0132-1. [DOI] [PubMed] [Google Scholar]
- 174.Kim S-H, Michael WM. Regulated proteolysis of DNA polymerase eta during the DNA damage response in C. elegans . Mol. Cell. 2008;32:757–766. doi: 10.1016/j.molcel.2008.11.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.L Parker J, Bielen AB, Dikic I, Ulrich HD. Contributions of ubiquitin- and PCNA-binding domains to the activity of Polymerase η in Saccharomyces cerevisiae . Nucleic Acids Res. 2007;35:881–889. doi: 10.1093/nar/gkl1102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Guo C, Tang T-S, Bienko M, L Parker J, Bielen AB, Sonoda E, Takeda S, Ulrich HD, Dikic I, Friedberg EC. Ubiquitin-binding motifs in REV1 protein are required for its role in the tolerance of DNA damage. Mol. Cell. Biol. 2006;26:8892–8900. doi: 10.1128/MCB.01118-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Guo C, Tang TS, Bienko M, Dikic I, Friedberg EC. Requirements for the interaction of mouse Polκ with ubiquitin and its biological significance. J Biol Chem. 2008;283:4658–64. doi: 10.1074/jbc.M709275200. [DOI] [PubMed] [Google Scholar]
- 178.Hoeller D, Crosetto N, Blagoev B, Raiborg C, Tikkanen R, Wagner S, Kowanetz K, Breitling R, Mann M, Stenmark H, Dikic I. Regulation of ubiquitin-binding proteins by monoubiquitination. Nat. Cell Biol. 2006;8:163–169. doi: 10.1038/ncb1354. [DOI] [PubMed] [Google Scholar]
- 179.Hoeller D, Hecker C-M, Wagner S, Rogov V, Dötsch V, Dikic I. E3-independent monoubiquitination of ubiquitin-binding proteins. Mol. Cell. 2007;26:891–898. doi: 10.1016/j.molcel.2007.05.014. [DOI] [PubMed] [Google Scholar]
- 180.Haglund K, Stenmark H. Working out coupled monoubiquitination. Nat. Cell Biol. 2006;8:1218–1219. doi: 10.1038/ncb1106-1218. [DOI] [PubMed] [Google Scholar]
- 181.Roerink SF, Koole W, Stapel LC, Romeijn RJ, Tijsterman M. A Broad requirement for TLS polymerases η and, κ interacting sumoylation nuclear pore proteins, in lesion bypass during C. elegans embryogenesis. PLoS Genet. 2012;8:e1002800. doi: 10.1371/journal.pgen.1002800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182.Masutani C, Kusumoto R, Yamada A, Dohmae N, Yokoi M, Yuasa M, Araki M, Iwai S, Takio K, Hanaoka F. The XPV (xeroderma pigmentosum variant) gene encodes human DNA polymerase η. Nature. 1999;399:700–704. doi: 10.1038/21447. [DOI] [PubMed] [Google Scholar]
- 183.Johnson RE, Washington MT, Prakash S, Prakash L. Fidelity of human DNA polymerase η. J. Biol. Chem. 2000;275:7447–7450. doi: 10.1074/jbc.275.11.7447. [DOI] [PubMed] [Google Scholar]
- 184.Jung Y-S, Liu G, Chen X. Pirh2 E3 ubiquitin ligase targets DNA polymerase eta for 20S proteasomal degradation. Mol. Cell. Biol. 2010;30:1041–1048. doi: 10.1128/MCB.01198-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Jung Y-S, Hakem A, Hakem R, Chen X. Pirh2 E3 ubiquitin ligase monoubiquitinates DNA polymerase eta to suppress translesion DNA synthesis. Mol. Cell. Biol. 2011;31:3997–4006. doi: 10.1128/MCB.05808-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186.Jung Y-S, Qian Y, Chen X. DNA polymerase eta is targeted by Mdm2 for polyubiquitination and proteasomal degradation in response to ultraviolet irradiation. DNA Repair. 2012;11:177–184. doi: 10.1016/j.dnarep.2011.10.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 187.Wallace HA, Merkle JA, Yu MC, Berg TG, Lee E, Bosco G, Lee AL. TRIP/NOPO E3 ubiquitin ligase promotes ubiquitylation of DNA polymerase η. Dev. Camb. Engl. 2014;141:1332–1341. doi: 10.1242/dev.101196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 188.Merkle JA, L Rickmyre J, Garg A, Loggins EB, Jodoin JN, Lee E, Wu LP, Lee AL. no poles encodes a predicted E3 ubiquitin ligase required for early embryonic development of Drosophila. Dev. Camb. Engl. 2009;136:449–459. doi: 10.1242/dev.027599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Qian J, Pentz K, Zhu Q, Wang Q, He J, Srivastava AK, Wani AA. USP7 modulates UV-induced PCNA monoubiquitination by regulating DNA polymerase eta stability. Oncogene. 2014 doi: 10.1038/onc.2014.394. [Epub ahead of print] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190.Plachta M, Halas A, McIntyre J, Sledziewska-Gojska E. The steady-state level and stability of TLS polymerase eta are cell cycle dependent in the yeast S. cerevisiae. doi: 10.1016/j.dnarep.2015.02.015. [this issue] [DOI] [PubMed] [Google Scholar]
- 191.Chen YW, Cleaver JE, Hatahet Z, Honkanen RE, Chang J-Y, Yen Y, M.Chou K. Human DNA polymerase η activity and translocation is regulated by phosphorylation. Proc. Natl. Acad. Sci. 2008;105:16578–16583. doi: 10.1073/pnas.0808589105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192.Gohler T, Sabbioneda S, Green CM, Lehmann AR. ATR-mediated phosphorylation of DNA polymerase η is needed for efficient recovery from UV damage. J. Cell Biol. 2011;192:219–227. doi: 10.1083/jcb.201008076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.McDonald JP, Rapić-Otrin V, Epstein JA, Broughton BC, Wang X, Lehmann AR, Wolgemuth DJ, Woodgate R. Novel human and mouse homologs of Saccharomyces cerevisiae DNA polymerase η. Genomics. 1999;60:20–30. doi: 10.1006/geno.1999.5906. [DOI] [PubMed] [Google Scholar]
- 194.Wang Y, Woodgate R, McManus TP, Mead S, McCormick JJ, Maher VM. Evidence that in xeroderma pigmentosum variant cells which lack DNA polymerase η, DNA polymerase ι causes the very high frequency and unique spectrum of UV-induced mutations. Cancer Res. 2007;67:3018–3026. doi: 10.1158/0008-5472.CAN-06-3073. [DOI] [PubMed] [Google Scholar]
- 195.Ohkumo T, Kondo Y, Yokoi M, Tsukamoto T, Yamada A, Sugimoto T, Kanao R, Higashi Y, Kondoh H, Tatematsu M, Masutani C, Hanaoka F. UV-B radiation induces epithelial tumors in mice lacking DNA polymerase η and mesenchymal tumors in mice deficient for DNA polymerase ι. Mol. Cell. Biol. 2006;26:7696–7706. doi: 10.1128/MCB.01076-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.Wang M, Devereux TR, Vikis HG, McCulloch SD, Holliday W, Anna C, Wang Y, Bebenek K, Kunkel TA, Guan K, You M, Pol ι is a candidate for the mouse pulmonary adenoma resistance 2 locus. a major modifier of chemically induced lung neoplasia. Cancer Res. 2004;64:1924–1931. doi: 10.1158/0008-5472.can-03-3080. [DOI] [PubMed] [Google Scholar]
- 197.Ziv O, Geacintov N, Nakajima S, Yasui A, Livneh Z. DNA polymerase ζ cooperates with polymerases κ and ι in translesion DNA synthesis across pyrimidine photodimers in cells from XPV patients. Proc. Natl. Acad. Sci. U. S. A. 2009;106:11552–11557. doi: 10.1073/pnas.0812548106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 198.Jansen JG, Temviriyanukul P, Wit N, Delbos F, Reynaud C-A, Jacobs H, de Wind N. Redundancy of mammalian Y family DNA polymerases in cellular responses to genomic DNA lesions induced by ultraviolet light. Nucleic Acids Res. 2014;42:11071–11082. doi: 10.1093/nar/gku779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 199.Frank EG, Woodgate R. Increased catalytic activity and altered fidelity of human DNA polymerase ι in the presence of manganese. J. Biol. Chem. 2007;282:24689–24696. doi: 10.1074/jbc.M702159200. [DOI] [PubMed] [Google Scholar]
- 200.Tissier A, Frank EG, McDonald JP, Iwai S, Hanaoka F, Woodgate R. Misinsertion and bypass of thymine-thymine dimers by human DNA polymerase ι. EMBO J. 2000;19:5259–5266. doi: 10.1093/emboj/19.19.5259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 201.Pence MG, Blans P, Zink CN, Hollis T, Fishbein JC, Perrino FW. Lesion bypass of N2-ethylguanine by human DNA polymerase ι. J. Biol. Chem. 2009;284:1732–1740. doi: 10.1074/jbc.M807296200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 202.Washington MT, Minko IG, Johnson RE, Wolfle WT, Harris TM, Lloyd RS, Prakash S, Prakash L. Efficient and error-free replication past a minor-groove DNA adduct by the sequential action of human DNA polymerases ι and κ. Mol. Cell. Biol. 2004;24:5687–5693. doi: 10.1128/MCB.24.13.5687-5693.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 203.Washington MT, Carlson KD, Freudenthal BD, Pryor JM. Variations on a theme: eukaryotic Y-family DNA polymerases. Biochim. Biophys. Acta. 2010;1804:1113–1123. doi: 10.1016/j.bbapap.2009.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204.Tissier A, Frank EG, McDonald JP, Vaisman A, Fernàndez de Henestrosa AR, Boudsocq F, McLenigan MP, Woodgate R. Biochemical characterization of human DNA polymerase ι provides clues to its biological function. Biochem. Soc. Trans. 2001;29:183–187. doi: 10.1042/0300-5127:0290183. [DOI] [PubMed] [Google Scholar]
- 205.Zhang Y, Yuan F, Wu X, Wang Z. Preferential incorporation of G opposite template T by the low-fidelity human DNA polymerase ι. Mol. Cell. Biol. 2000;20:7099–7108. doi: 10.1128/mcb.20.19.7099-7108.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206.Kirouac KN, Ling H. Unique active site promotes error-free replication opposite an 8- oxo-guanine lesion by human DNA polymerase iota. Proc. Natl. Acad. Sci. U. S. A. 2011;108:3210–3215. doi: 10.1073/pnas.1013909108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207.Nair DT, Johnson RE, Prakash L, Prakash S, Aggarwal AK. Human DNA polymerase ι incorporates dCTP opposite template G via a G.C + Hoogsteen base pair. Struct. 2005;13:1569–1577. doi: 10.1016/j.str.2005.08.010. [DOI] [PubMed] [Google Scholar]
- 208.Bebenek K, Tissier A, Frank EG, McDonald JP, Prasad R, Wilson SH, Woodgate R, Kunkel TA. 5’-Deoxyribose phosphate lyase activity of human DNA polymerase ι in vitro. Science. 2001;291:2156–2159. doi: 10.1126/science.1058386. [DOI] [PubMed] [Google Scholar]
- 209.Prasad R, Bebenek K, Hou E, Shock DD, Beard WA, Woodgate R, Kunkel TA, Wilson SH. Localization of the deoxyribose phosphate lyase active site in human DNA polymerase ι by controlled proteolysis. J. Biol. Chem. 2003;278:29649–29654. doi: 10.1074/jbc.M305399200. [DOI] [PubMed] [Google Scholar]
- 210.McIntyre J, Vidal AE, McLenigan MP, Bomar MG, Curti E, McDonald JP, Plosky BS, Ohashi E, Woodgate R. Ubiquitin mediates the physical and functional interaction between human DNA polymerases η and ι. Nucleic Acids Res. 2013;41:1649–1660. doi: 10.1093/nar/gks1277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.Ohashi E, Ogi T, Kusumoto R, Iwai S, Masutani C, Hanaoka F, Ohmori H. Error-prone bypass of certain DNA lesions by the human DNA polymerase κ. Genes Dev. 2000;14:1589–1594. [PMC free article] [PubMed] [Google Scholar]
- 212.Lone S, Townson SA, Uljon SN, Johnson RE, Brahma A, Nair DT, Prakash S, Prakash L, Aggarwal AK. Human DNA polymerase κ encircles DNA: implications for mismatch extension and lesion bypass. Mol. Cell. 2007;25:601–614. doi: 10.1016/j.molcel.2007.01.018. [DOI] [PubMed] [Google Scholar]
- 213.Washington MT, Johnson RE, Prakash L, Prakash S. Human DINB1-encoded DNA polymerase κ is a promiscuous extender of mispaired primer termini. Proc. Natl. Acad. Sci. U. S. A. 2002;99:1910–1914. doi: 10.1073/pnas.032594399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 214.Ogi T, Lehmann AR. The Y-family DNA polymerase κ (pol κ) functions in mammalian nucleotide-excision repair. Nat. Cell Biol. 2006;8:640–642. doi: 10.1038/ncb1417. [DOI] [PubMed] [Google Scholar]
- 215.Sale JE, Lehmann AR, Woodgate R. Y-family DNA polymerases and their role in tolerance of cellular DNA damage. Nat Rev Mol Cell Biol. 2012;13:141–52. doi: 10.1038/nrm3289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 216.Okada T, Sonoda E, Yamashita YM, Koyoshi S, Tateishi S, Yamaizumi M, Takata M, Ogawa O, Takeda S. Involvement of vertebrate polκ in Rad18-independent postreplication repair of UV damage. J. Biol. Chem. 2002;277:48690–48695. doi: 10.1074/jbc.M207957200. [DOI] [PubMed] [Google Scholar]
- 217.Wit N, Buoninfante OA, van den Berk PC, Jansen JG, Hogenbirk MA, de Wind N, Jacobs H. Roles of PCNA ubiquitination and TLS polymerases κ and η in the bypass of methyl methanesulfonate-induced DNA damage. Nucleic Acids Res. 2015;43:282–94. doi: 10.1093/nar/gku1301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 218.Nelson JR, Lawrence CW, Hinkle DC. Deoxycytidyl transferase activity of yeast REV1 protein. Nature. 1996;382:729–731. doi: 10.1038/382729a0. [DOI] [PubMed] [Google Scholar]
- 219.Nelson JR, Gibbs PE, Nowicka AM, Hinkle DC, Lawrence CW. Evidence for a second function for Saccharomyces cerevisiae Rev1p. Mol. Microbiol. 2000;37:549–554. doi: 10.1046/j.1365-2958.2000.01997.x. [DOI] [PubMed] [Google Scholar]
- 220.Lin W, Xin H, Zhang Y, Wu X, Yuan F, Wang Z. The human REV1 gene codes for a DNA template-dependent dCMP transferase. Nucleic Acids Res. 1999;27:4468–4475. doi: 10.1093/nar/27.22.4468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 221.Zhang Y. Response of human REV1 to different DNA damage: preferential dCMP insertion opposite the lesion. Nucleic Acids Res. 2002;30:1630–1638. doi: 10.1093/nar/30.7.1630. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 222.Ross A-L. Vertebrate DNA damage tolerance requires the C-terminus but not BRCT or transferase domains of REV1. Nucleic Acids Res. 2005;33:1280–1289. doi: 10.1093/nar/gki279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 223.Ohashi E, Murakumo Y, Kanjo N, Akagi J-I, Masutani C, Hanaoka F, Ohmori H. Interaction of hREV1 with three human Y-family DNA polymerases. Genes Cells. 2004;9:523–531. doi: 10.1111/j.1356-9597.2004.00747.x. [DOI] [PubMed] [Google Scholar]
- 224.Ohashi E, Hanafusa T, Kamei K, Song I, Tomida J, Hashimoto H, Vaziri C, Ohmori H. Identification of a novel REV1-interacting motif necessary for DNA polymerase κ function. Genes Cells. 2009;14:101–111. doi: 10.1111/j.1365-2443.2008.01255.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 225.Guo C, L Fischhaber P, Luk-Paszyc MJ, Masuda Y, Zhou J, Kamiya K, Kisker C, Friedberg EC. Mouse Rev1 protein interacts with multiple DNA polymerases involved in translesion DNA synthesis. EMBO J. 2003;22:6621–6630. doi: 10.1093/emboj/cdg626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 226.Tissier A, Kannouche P, Reck M-P, Lehmann AR, Fuchs RPP, Cordonnier A. Co-localization in replication foci interaction of human Y-family members, DNA polymerase pol η and REV1 protein. DNA Repair. 2004;3:1503–1514. doi: 10.1016/j.dnarep.2004.06.015. [DOI] [PubMed] [Google Scholar]
- 227.Pozhidaeva A, Pustovalova Y, D’Souza S, Bezsonova I, Walker GC, Korzhnev DM. NMR structure and dynamics of the C-terminal domain from human Rev1 and its complex with Rev1 interacting region of DNA polymerase η. Biochemistry. 2012;51:5506–5520. doi: 10.1021/bi300566z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 228.Wood A, Garg P, Burgers PMJ. A ubiquitin-binding motif in the translesion DNA polymerase Rev1 mediates its essential functional interaction with ubiquitinated proliferating cell nuclear antigen in response to DNA damage. J. Biol. Chem. 2007;282:20256–20263. doi: 10.1074/jbc.M702366200. [DOI] [PubMed] [Google Scholar]
- 229.Pustovalova Y, Maciejewski MW, Korzhnev DM. NMR mapping of PCNA interaction with translesion synthesis DNA polymerase Rev1 mediated by Rev1-BRCT domain. J. Mol. Biol. 2013;425:3091–3105. doi: 10.1016/j.jmb.2013.05.029. [DOI] [PubMed] [Google Scholar]
- 230.Guo C, Sonoda E, Tang T, Parker J, Bielen A, Takeda S, Ulrich H, Friedberg E. REV1 Protein Interacts with PCNA: Significance of the REV1 BRCT Domain in vitro and in vivo. Mol. Cell. 2006;23:265–271. doi: 10.1016/j.molcel.2006.05.038. [DOI] [PubMed] [Google Scholar]
- 231.Kim H, Yang K, Dejsuphong D, D’Andrea AD. Regulation of Rev1 by the Fanconi anemia core complex. Nat. Struct. Mol. Biol. 2012;19:164–170. doi: 10.1038/nsmb.2222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 232.Wiltrout ME, Walker GC. Proteasomal regulation of the mutagenic translesion DNA polymerase, Saccharomyces cerevisiae Rev1. DNA Repair. 2011;10:169–75. doi: 10.1016/j.dnarep.2010.10.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 233.Weinert BT, Wagner SA, Horn H, Henriksen P, Liu WR, Olsen JV, Jensen JL, Choudhary C. Proteome-wide mapping of the Drosophila acetylome demonstrates a high degree of conservation of lysine acetylation. Sci. Signal. 2011;4 doi: 10.1126/scisignal.2001902. ra48, 2011. [DOI] [PubMed] [Google Scholar]