Abstract
Background
Premature leaf senescence induced by external stress conditions, e.g. drought stress, is a main factor for yield losses in barley. Research in drought stress tolerance has become more important as due to climate change the number of drought periods will increase and tolerance to drought stress has become a goal of high interest in barley breeding. Therefore, the aim is to identify quantitative trait loci (QTL) involved in drought stress induced leaf senescence and drought stress tolerance in early developmental stages of barley (Hordeum vulgare L.) by applying genome wide association studies (GWAS) on a set of 156 winter barley genotypes.
Results
After a four weeks stress period (BBCH 33) leaf colour as an indicator of leaf senescence, electron transport rate at photosystem II, content of free proline, content of soluble sugars, osmolality and the aboveground biomass indicative for drought stress response were determined in the control and stress variant in greenhouse pot experiments. Significant phenotypic variation was observed for all traits analysed. Heritabilities ranged between 0.27 for osmolality and 0.61 for leaf colour in stress treatment and significant effects of genotype, treatment and genotype x treatment were estimated for most traits analysed. Based on these phenotypic data and 3,212 polymorphic single nucleotide polymorphisms (SNP) with a minor allele frequency >5 % derived from the Illumina 9 k iSelect SNP Chip, 181 QTL were detected for all traits analysed. Major QTLs for drought stress and leaf senescence were located on chromosome 5H and 2H. BlastX search for associated marker sequences revealed that respective SNPs are in some cases located in proteins related to drought stress or leaf senescence, e.g. nucleotide pyrophosphatase (AVP1) or serine/ threonin protein kinase (SAPK9).
Conclusions
GWAS resulted in the identification of many QTLs involved in drought stress and leaf senescence of which two major QTLs for drought stress and leaf senescence were located on chromosome 5H and 2H. Results may be the basis to incorporate breeding for tolerance to drought stress or leaf senescence in barley breeding via marker based selection procedures.
Electronic supplementary material
The online version of this article (doi:10.1186/s12870-015-0524-3) contains supplementary material, which is available to authorized users.
Keywords: Barley, Leaf senescence, Drought stress, GWAS, QTL
Background
Barley (Hordeum vulgare L.) is one of the first cereals domesticated in the Fertile Crescent [1] and today it is the fourth most important crop species concerning acreage next to wheat, maize, and rice [2]. Worldwide, barley is mainly used for animal feed and malting and only a very small amount is used for direct human consumption and bakery. Average yield of barley on the worldwide level is 2.9 t/ha but in some European countries, e.g. Germany average yield is up to 6.5 t/ha [2]. Barley yield in many parts of the world is reduced by biotic stress but also by abiotic stress e.g. heat, salt, deficits in nitrogen nutrition and drought [3–6]. Especially, in the juvenile stages from sowing to tillering, drought can severely influence barley development already reducing the potential yield [7]. Research on drought stress tolerance has become more important worldwide as due to climate change the number of drought periods will increase in the future [8, 9]. Up to now, most studies conducted in barley focused on effects of terminal drought stress whereas drought in juvenile stages is less well documented [10].
Drought tolerance is a complex quantitative trait, that is controlled by various mechanisms [11, 12]. Abscisic acid (ABA) is a key phytohormone involved in adaption to environmental stresses and regulation of plant development. It promotes the closure of stomata under drought stress conditions initiated by a loss of turgor [13]. Furthermore, it increases the hydraulic conductivity of water, promotes chlorophyll breakdown and leads to leaf senescence [14]. Another relevant protein is ubiquitin which regulates the degradation of proteins [15, 16]. Moreover, late embryo abundant (LEA) proteins and heat shock proteins that are involved in the protection of functional proteins are induced in response to various abiotic stresses [17–19]. For example protein kinases and protein phosphatases which activate or deactivate proteins by phosphorylation and dephosphorilation [20], as well as the LEA protein dehydrin which is described to have different functions in different stresses [21] is often present along with drought stress [22]. A lot of parameters indicative for drought stress influenced by these and additional genes were analysed in different crops [23, 24]. For example, biomass production [25], yield [26], photosynthesis rate [27], as well as the content of free proline [28], total content of soluble sugars [29], or osmolality [30] are parameters which are affected by drought stress in barley.
Another factor relevant for yield improvement is leaf senescence [31], which is a natural degradation process at the final stage of the development of organs and plants. This process is divided into three steps and starts with reprogramming of gene expression to turn on senescence activating genes. Before programmed cell death in the terminal phase occurs, the second step in which nutrients and metabolites are transported from source (e.g. roots, leaves) to sink (e.g. fruits, seed) is important for yield and quality of the seeds harvested [32]. During this reorganization phase, degradation of chlorophyll and a decrease in photosynthesis is observed [33, 34]. Because of degradation of chlorophyll, yellowing of the leaves is a symptom of leaf senescence [35], which in many studies is rated visually [36, 37], but can be more precisely determined by Soil Plant Analysis Development (SPAD) readings which estimate leaf greenness [38]. Degradation of chlorophyll is regulated by chlorophyllase, pheophorbide α oxygenase and red chlorophyll catabolite reductase among others [34], but so far for regulation of leaf senescence only a few genes are known [39, 40]. Leaf senescence is a process which is influenced by a lot of external stress conditions e.g. drought stress [24, 41]. Stress often results in premature induction of leaf senescence and therefore leads to an inefficient recycling of resources and a massive yield loss [42–44]. In contrast, plants showing delayed leaf senescence under stress, represented by a “stay green effect”, minimize yield loss [45].
Genome wide association studies (GWAS) are a powerful tool to subdivide such complex pathways as drought stress and leaf senescence by the detection of quantitative trait loci (QTL) out of the regression analysis of genotypic and phenotypic data [19, 46–48]. Up to now, some QTLs involved in drought stress response were published in barley [49–52], whereas for leaf senescence only few QTLs are known [42, 53]. Besides this, QTLs which are involved in the response to other abiotic stresses, e.g. salt stress were identified in barley [6, 54]. Molecular markers, such as single nucleotide polymorphisms (SNP) flanking QTLs having a significant influence on the respective trait can be used for efficient marker assisted selection and smart breeding procedures [55].
The aim of the present study is therefore the identification of QTLs for drought stress and drought stress induced leaf senescence in early developmental stages of barley suited to be used in future barley breeding programs using GWAS followed by the identification of the function of these QTLs associated to respective traits.
Material & methods
Plant material and experimental setup
A set of 156 winter barley genotypes (Additional file 1) consisting of 113 German winter barley cultivars [49 two-rowed and 64 six-rowed, [56] and 43 accessions of the Spanish barley core collection (SBCC) [57] were used to investigate drought stress induced leaf senescence in juvenile barley plants. Drought stress was applied in greenhouses of the Julius Kühn-Institut in Groß Lüsewitz, Germany according to Honsdorf et al. [51]. Trials were conducted in a split plot design with three replications per genotype and variant (control, drought stress). Ten seeds of each accession were sown per plastic pot (16x16x16 cm) containing 1,500 g of a mixed clay soil ED73 (H. Nitsch & Sohn GmbH & Co. KG, Dorsten Germany). After germination, seedlings were reduced to seven plants per pot. Plants were grown under semi-controlled long day conditions in a temperature range from 20 to 22 °C at day (16 h) and 17 to 19 °C at night (8 h). If natural radiation was below 20 klx, additional light was applied from 6 a.m. to 10 p.m. Drought stress started at the primary leaf stage (BBCH 10) seven days after sowing (das). At this time watering of the stress variant was stopped till the soil reached 20 % of the maximal soil water capacity, and then this level was kept by weighting each pot and re-watering. Control plants were continuously watered to 70 % of the maximal soil water capacity. Water capacity was calculated of the saturated soil weight and drought weight according to DIN ISO 11465 1996–12 [58]. At the end of a four weeks stress period (BBCH 33) physiological traits were determined and above ground biomass was harvested (experimental setup A). Experimental setup A was repeated in three years. A modified experimental setup was conducted to optimize lightening conditions of the primary leaves for the measurement of leaf colour (SPAD) and electron transport rate at photosystem II (ETR) (experimental setup B). General settings were the same as in setup A, but only four plants were grown in smaller pots (12x12x12 cm) to allow wider spacing of pots and all leaves except the primary leaves were tied up to reduce shading. Experimental setup B was repeated in two years.
Physiological parameters determined
Six physiological traits, i.e. leaf colour (SPAD), electron transport rate at photosystem II (ETR), content of free proline (CFP), content of soluble sugars (CSS), osmolality (OA) and the aboveground biomass yield (BY) were determined in the control and stress treatment. Measurement and sampling respectively were conducted on primary leaves.
Chlorophyll content which was used as the main indicator for drought stress induced leaf senescence was measured 33–34 das by Minolta SPAD readings (Konica Minolta Chlorophyll Meter SPAD-502 Plus, Osaka Japan), which gives a value for leaf colour. Three primary leaves of three plants for each pot were measured at five positions per leaf. These SPAD readings turned out to be correlated to the chlorophyll content analysed photometrically [59, 60]. Because of this relation, the chlorophyll content can be indirectly measured by SPAD [61].
At 34–35 das chlorophyll fluorescence was measured in all genotypes using light adapted plants with the OS1P-Chlorophyll Fluorometer (OPTI-SCIENCE, Hudson USA) in the middle of three primary leaves per pot at one position per leaf. The relative electron transport rate at photosystem II (PSII) (ETR = Y(II) * PAR * 0, 84 * 0, 5) was calculated including the photosynthetically active radiation (PAR), as well as the quantum photosynthetic yield of PSII (Y(II)) and constants representing light which is absorbed by the leaf (0.84) and light which is equally absorbed by PSI and PSII (0.5) [62].
At 36 das five primary leaves per pot were harvested and cut in pieces of 1 cm length for the analysis of CFP, CSS and OA. These samples were frozen in liquid nitrogen immediately and samples for CFP and CSS measurement were freeze dried. For CFP measurement the ninhydrin method [63] was applied, and for CSS measurement the anthron method [64] was used. Both traits were measured photometrical using a spectrophotometer. The concentration of these ingredients was determined with a standard curve calculated on a dry weight basis. To assess OA, frozen leaf samples were grinded in a swing mill (30/s for 3 min), filled up with 200 μl water and centrifuged at 15,000 rpm for 15 min to get cell sap for the measurement of osmolality with a freezing-point osmometer (Osmomat O-30 Gonotec, Berlin Germany). Osmolality was corrected for the water content of fresh and dry weight. Above ground biomass was harvested 36 das, too. Leaf material was dried in a compartment dryer at 105 °C and weighted.
For all traits an outlier test was calculated to exclude extreme deviations [65]. To get information on the stability of all analysed traits in response to drought stress compared to the control, the drought susceptibility index (DSI) was calculated [66] for each trait and across treatments according to the formula:
DSI is a relative value estimated for each genotype and trait. According to the formula genotypes revealing a DSI close to one are highly susceptible to drought and those close to zero or showing a negative value are tolerant.
Statistical analysis of phenotypic data
Statistical analyses were performed with SAS 9.3 [67]. Least square means (LSMeans) were calculated with GLM procedure for the replications of each genotype in the respective years and for both control and drought conditions. Descriptive statistics was calculated out of LSMeans by PROC UNIVARIATE. Analysis of variance (ANOVA) was calculated using PROC MIXED with genotypes (G), drought stress treatment (T) and GxT as fixed effects. Replication (R), year (Y) and row type are chosen as random factors. The heritability (h2) was calculated with SAS in two steps. First, the variance components for the genotypes (VG), variance associated with the genotype by year interaction (VGY) and VE which is the error variance were calculated with PROC VARCOMP. Next, h2 was calculated with the following formula: VG/(VG + VGY/Y + VE/RY) for both well watered and drought stress conditions. Furthermore, the coefficient of correlation (PROC CORR) by Pearson was calculated with SAS based on LSMeans.
Genotyping and genome wide association study (GWAS)
For genotyping the whole set of genotypes was analysed with the barley Illumina 9 k iSelect SNP-chip [68]. Population structure was calculated with STRUCTURE 2.3.4 [69] based on 51 simple sequence repeat (SSR) markers covering the whole genome. The STRUCTURE programme was run 20 times for pre-defined k (the number of population groups) from 1 to 5 each. To get the number of calculated subpopulations (k) with highest likelihood the procedure of Evanno et al. [70] was applied. An independent run with 500,000 iterations of a Monte Carlo Markov Chain with a length of the burn in period of 500,000 was conducted for the k with the highest likelihood to obtain the q-matrix. Kinship was calculated with SPAGeDi 1.3d [71] based on 51 SSRs and allele size correlation coefficient [72] with 5,000 permutations.
Out of 3,886 genetically mapped SNP markers (398 at 1H, 690 at 2H, 583 at 3H, 342 at 4H, 781 at 5H, 546 at 6H and 546 at 7H) [68], 3,212 polymorphic markers with minor allele frequencies higher than 5 % were taken into account. Based on these data and respective phenotypic data (LSMeans) GWAS was conducted applying a mixed linear model (MLM) using TASSEL 3.0 [73]. All results with p values <0.001 were considered as significant marker trait associations. Linkage disequilibrium (LD) was calculated on mapped polymorphic SNPs with R [74] by an estimate of the average decay [75] over all barley chromosomes.
Sequences of significantly associated SNP-markers (p <0.001) were downloaded from the James Hutton Institute [http://bioinf.hutton.ac.uk/iselect/app] and respective sequences were compared against the plant proteome in the UniProtKB/Swiss-Prot protein database by BlastX (Basic Local Alignment Search Tool, p <10−5 or query cover of minimum 80 % in NCBI [https://www.ncbi.nlm.nih.gov] accessed Oct 2014) to get information on the proteins coded by these sequences [76]. Using UniProt [77] the involvement of respective proteins in drought stress and leaf senescence processes was analysed. In a last step a genetic map with all significantly associated SNPs in genes coding for proteins known to be involved in drought stress tolerance was generated using MapChart 2.2 [78].
Results
Phenotyping
The experiments revealed variability for genotype and treatment in all analysed traits as shown in Table 1. For the traits biomass yield (BY), leaf colour (SPAD) and the electron transport rate (ETR) the mean values for the stress treatment were lower than in the well watered variant. An exception are some genotypes of the SBCC (SBCC 3, 12, 14, 76, 80, 138 and 140) showing no decrease in SPAD and ETR or even an increase, represented by negative values across treatments (DSI). In contrast to the above mentioned traits, osmolality (OA), content of free proline (CFP) and total content of soluble sugars (CSS) increased under drought stress.
Table 1.
Descriptive statistics, heritability (h2) and number of significant (p <0.001) quantitative trait loci (QTL)
| Traita | Description | Treat.b | Unit | Minc | Maxc | Meanc | SDc | CVc | LSDc | h2 | No. QTL (SNPs) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| BY | Total above ground biomass yield | Control | g | 4.47 | 18.18 | 10.70 | 2.68 | 25.08 | 4.12 | 0.80 | 72 (179) |
| Stress | g | 2.73 | 9.98 | 5.57 | 1.83 | 32.80 | 1.45 | 0.58 | 19 (32) | ||
| DSI | unit free | 0.65 | 1.28 | 0.98 | 50 (87) | ||||||
| SPAD | Leaf colour | Control | unit free | 4.70 | 48.80 | 35.76 | 8.33 | 23.30 | 23.39 | 0.64 | 8 (12) |
| Stress | unit free | 7.83 | 46.77 | 34.16 | 6.56 | 19.20 | 21.41 | 0.61 | 3 (6) | ||
| DSI | unit free | −7.70 | 9.41 | 0.93 | 2 (2) | ||||||
| ETR | Relative electron transport rate at PSII | Control | Y(II) x PAR x 0,84 x 0,5 | 6.12 | 32.26 | 15.73 | 4.63 | 29.46 | 19.47 | 0.08 | |
| Stress | Y(II) x PAR x 0,84 x 0,5 | 2.00 | 30.70 | 13.41 | 4.54 | 33.85 | 16.38 | 0.50 | 2 (2) | ||
| DSI | unit free | −4.09 | 4.82 | 0.90 | |||||||
| OA | Osmolality | Control | osmol kg−1 | 0.23 | 0.78 | 0.46 | 0.05 | 11.57 | 0.19 | 0.00 | |
| Stress | osmol kg−1 | 0.40 | 1.16 | 0.67 | 0.08 | 11.48 | 0.23 | 0.27 | 22 (29) | ||
| DSI | unit free | 0.44 | 2.00 | 1.02 | 0.26 | 1 (1) | |||||
| CFP | Content of free proline | Control | μmol g−1 | 0.00 | 22.36 | 3.66 | 3.11 | 84.89 | 7.33 | 0.13 | |
| Stress | μmol g−1 | 1.02 | 78.57 | 23.94 | 16.86 | 70.43 | 44.30 | 0.29 | |||
| DSI | unit free | 0.00 | 3.46 | 1.15 | 1 (2) | ||||||
| CSS | Total content of soluble sugars | Control | μmol g−1 | 86.07 | 568.07 | 231.40 | 85.51 | 36.95 | 240.72 | 0.13 | |
| Stress | μmol g−1 | 164.61 | 981.40 | 419.64 | 164.03 | 39.09 | 411.52 | 0.30 | 1 (1) | ||
| DSI | unit free | 0.08 | 2.95 | 1.07 |
aBY: biomass yield, SPAD: leaf colour, ETR: electron transport rate at PSII, OA: osmolality, CFP: content of free proline, CSS: total content of soluble sugars
bControl and drought stress treatment, as well as DSI: drought susceptibility index across treatments by Fischer & Maurer (1978)
cMinimum, maximum, mean, standard deviation in % (SD), coefficient of variation (CV) (standard deviation divided by mean) and least significant difference (LSD)
The coefficient of variation (CV) was comparable for control and drought stress treatment (Table 1) for all six traits. Heritabilities (h2) estimated ranged between 0 for OA to 0.80 for BY in the control treatment and 0.27 for OA and 0.61 for SPAD in the stress treatment. Generally, h2 was higher for the stress treatment except for BY and SPAD. Analysis of variance (ANOVA) revealed significant (p <0.001) genotype and treatment effects for all investigated traits and genotype x treatment interactions for BY, CFP and CSS (Table 2).
Table 2.
Analysis of variance (ANOVA) of analysed traits showing F and p values
| Traita | Effectb | F value | P value |
|---|---|---|---|
| BY | Genotype | 7.61 | <.0001 |
| Treatment | 10878.9 | <.0001 | |
| G x T | 4.16 | <.0001 | |
| SPAD | Genotype | 8.81 | <.0001 |
| Treatment | 30.74 | <.0001 | |
| G x T | 0.83 | 0.9348 | |
| ETR | Genotype | 2.08 | <.0001 |
| Treatment | 43.69 | <.0001 | |
| G x T | 0.97 | 0.6007 | |
| OA | Genotype | 1.45 | 0.0004 |
| Treatment | 3737.86 | <.0001 | |
| G x T | 1.16 | 0.0962 | |
| CFP | Genotype | 2.59 | <.0001 |
| Treatment | 2544.91 | <.0001 | |
| G x T | 2.89 | <.0001 | |
| CSS | Genotype | 2.85 | <.0001 |
| Treatment | 1984.1 | <.0001 | |
| G x T | 2.42 | <.0001 |
aBY: biomass yield, SPAD: leaf colour, ETR: electron transport rate at PSII, OA: osmolality, CFP: content of free proline, CSS: total content of soluble sugars
bGenotype, Treatment and GxT: genotype x treatment interaction effect
To get information on the influence of the physiological parameters estimated on biomass yield as the indicator for drought stress and SPAD as the indicator for drought stress induced leaf senescence, correlations to these traits were calculated (Table 3). For control and stress treatment BY is significantly correlated to SPAD with r = 0.39 and r = 0.36, respectively. A significant correlation was also determined for CSS to SPAD with r = 0.42 and for CFP with r = 0.42 in the drought stress treatment whereas for the control treatment significantly negative correlations were found. Low but nevertheless significant correlations to SPAD were also detected for ETR and OA in the control treatment. Similar correlations were detected for BY. High and significant correlations were found between BY and CSS (r = 0.36) and CFP (r = 0.31) for the drought stress treatment. Under control conditions the SBCC being a sub-population of its own, influences the correlation by producing less BY which results in reduced shading of the primary leaves and a negative correlation especially to CFP and CSS. By correlating only the German cultivars, these effects are excluded and no correlations (r = −0.16 for CFP and r = −0.03 for CSS) were observed. ETR and OA were not significantly correlated to BY.
Table 3.
Coefficient of correlation (PEARSON) for control and drought stress treatment
| Treatment | SPAD | ETR | OA | CFP | CSS | |
|---|---|---|---|---|---|---|
| BY | Control | 0.395*** | 0.091 | −0.127 | −0.328*** | −0.220** |
| Stress | 0.361*** | −0.087 | −0.124 | 0.307*** | 0.367*** | |
| SPAD | Control | 0.160* | −0.185* | −0.239** | −0.192* | |
| Stress | −0.105 | 0.034 | 0.425*** | 0.418*** |
aBY: biomass yield, SPAD: leaf colour, ETR: electron transport rate at PSII, OA: osmolality, CFP: content of free proline, CSS: total content of soluble sugars
r is significant with *p <0.05, **p <0.01 and ***p <0.001
Genotyping
The set of genotypes was analysed with the 9 k iSelect SNP-chip available for barley. In summary 6,807 SNPs turned out to be polymorphic. Out of these, 3,212 SNPs are mapped on the seven barley chromosomes [68], showing a minor allele frequency (MAF) >5 %. This set of SNPs was used for the calculation of the linkage disequilibrium decay (LD), which turned out be on average 2.52 cM for this set of genotypes. The number of subpopulations was estimated at k = 4 (Fig. 1).
Fig. 1.
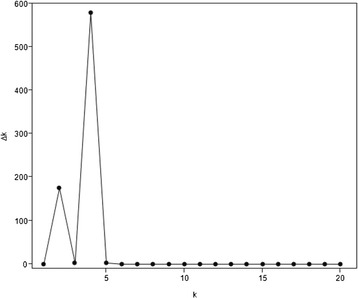
Optimal k of the population structure. The number of subpopulations within the set of barley genotypes was estimated at k = 4 by calculation described in Evanno et al. [70]
Genome wide association study (GWAS)
Results of GWAS are shown in detail in Additional file 2 and summarized in Tables 4 and 5. 191 SNPs significantly (p <0.001) associated to traits estimated in the control variant, 70 significantly associated SNPs in the stress treatment and 92 significantly associated SNPs across treatments (DSI), were detected using the MLM analysis in TASSEL. Significant associations were found on all barley chromosomes. Most of the significant marker trait associations were located on barley chromosome 2H and 5H. A large number of SNPs on chromosome 5H around 45 cM turned out to be associated to SPAD and BY in the stress treatment (Fig. 2). Most significant associations for these traits were detected on chromosome 5H. The highest number of marker trait associations was detected for BY. No associations were observed for CFP in control and stress treatment, whereas across treatments one significant association was found. For ETR two significant associations and for CSS one significant association in the stress treatment explaining 5.5 % (ETR) and 1.6 % (CSS) of the phenotypic variance respectively were detected. The strongest association was observed on chromosome 1H for BY across treatments with a (−log p) value of 7.57 explaining 7.1 % of the phenotypic variance. For OA 29 significant associations were detected in the stress treatment located over all barley chromosomes, whereas in the control treatment no associations and across treatments only one association was found on chromosome 4H.
Table 4.
Significant markers traits associations detected under drought stress conditions at a significance of p <0.001
| Traita | Number of genomic regions associated with the traits on the seven linkage groups (barley chromosomes)b,c | |||||||
|---|---|---|---|---|---|---|---|---|
| 1H | 2H | 3H | 4H | 5H | 6H | 7H | Total QTL | |
| BY | 81.7 cM (3 SNP) | 2 cM (3 SNP) | 76.2 cM (1 SNP) | 99.1 cM (1 SNP) | 46.7 cM (8 SNP) | 48.3 cM (1 SNP) | 19 (32 SNPs) | |
| 92.2 cM (1 SNP) | 5.5 cM (1 SNP) | 135.5 cM (1 SNP) | 59.7 cM (1 SNP) | 70.2 cM (1 SNP) | ||||
| 12.1 cM (1 SNP) | 80.3 cM (1 SNP) | 133.9 cM (1 SNP) | ||||||
| 90.2 cM (3 SNP) | 110.1 cM (1 SNP) | |||||||
| 139.1 cM (1 SNP) | ||||||||
| 152.4 cM (1 SNP) | ||||||||
| 167.7 cM (1 SNP) | ||||||||
| SPAD | 49.2 cM (1 SNP) | 44.2 cM (4 SNP) | 128.3 cM (1 SNP) | 3 (6 SNPs) | ||||
| ETR | 59.4 cM (1 SNP) | 2.1 cM (1 SNP) | 2 (2 SNPs) | |||||
| OA | 116.8 cM (1 SNP) | 51.8 cM (1 SNP) | 2.4 cM (1 SNP) | 52.3 cM (1 SNP) | 46.5 cM (1 SNP) | 10.3 cM (1 SNP) | 106.5 cM (1 SNP) | 22 (29 SNPs) |
| 60.8 cM (2 SNP) | 36.8 cM (2 SNP) | 110.2 cM (1 SNP) | 55.7 cM (1 SNP) | 47.5 cM (1 SNP) | ||||
| 81.5 cM (4 SNP) | 51.8 cM (1 SNP) | 95 cM (1 SNP) | 51 cM (2 SNP) | |||||
| 135.8 cM (1 SNP) | 61.9 cM (1 SNP) | 137.9 cM (1 SNP) | ||||||
| 146.5 cM (1 SNP) | 89.4 cM (1 SNP) | |||||||
| 100.7 cM (2 SNP) | ||||||||
| CSS | 95.8 cM (1 SNP) | 1 (1 SNP) | ||||||
| Total QTL | 4 (6 SNPs) | 10 (18 SNPs) | 8 (10 SNPs) | 3 (3 SNPs) | 12 (22 SNPs) | 4 (5 SNPs) | 6 (6 SNPs) | 47 (70 SNPs) |
aBY: biomass yield, SPAD: leaf colour, ETR: electron transport rate at PSII, OA: osmolality, CSS: total content of soluble sugars
bOne genomic region up to 2.6 cM (LD); the chromosomal position in cM was taken from the respective SNP with the highest R2
cChromosome positions are based on Comadran et al. (2012)
Table 5.
Significant blasted proteins related to drought stress or leaf senescence
| Protein (Top BlastX hit with p <10−5 or query cover >80 %) | Protein abbr. | Accession | Functiona | Markerb | Chr.b | Pos. in cMb | Traitc | Treat.d |
|---|---|---|---|---|---|---|---|---|
| Protease Do-like | DEGP2 | [Swiss-Prot:O82261.2] | ls | BOPA1_8166-525 | 1H | 47.5 | BY | C, DSI |
| Cullin-1 | CUL1 | [Swiss-Prot:Q94AH6.1] | ls | SCRI_RS_85918 | 1H | 47.7 | BY | C |
| Serine/threonine-protein phosphatase PP1 isozyme 3 | TOPP1 | [Swiss-Prot:P48483.1] | ds | SCRI_RS_17924 | 1H | 47.7 | BY | C, DSI |
| Electron transfer flavoprotein-ubiquinone oxidoreductase | ETFQO | [Swiss-Prot:O22854.1] | ls | SCRI_RS_132604 | 1H | 48.4 | BY | C, DSI |
| ATP-dependent zinc metalloprotease | FTSH3 | [Swiss-Prot:Q84WU8.1] | ls | BOPA1_2881-935 | 1H | 81.7 | BY | S |
| ABC transporter G family member 43 | ABCG43 | [Swiss-Prot:Q7PC81.1] | ds | BOPA2_12_31319 | 1H | 92.4 | SPAD | C, DSI |
| Probable pectinesterase 49 | PME49 | [Swiss-Prot:Q9LY18.1] | ds | SCRI_RS_235724 | 1H | 95.8 | CSS | S |
| Sucrose synthase 4 | SUS4 | [Swiss-Prot:Q9M111.1] | ls | SCRI_RS_239231 | 2H | 49.2 | SPAD | S |
| Metal-nicotianamine transporter YSL | YSL2 | [Swiss-Prot:Q6H3Z6.2] | ds | SCRI_RS_221886 | 2H | 80.9 | BY | C |
| Glutamate dehydrogenase 2 | GDH2 | [Swiss-Prot:Q38946.1] | ls | BOPA1_3469-1152 | 2H | 81.5 | BY | C |
| Probable glutamate carboxypeptidase 2 | AMP1 | [Swiss-Prot:Q9M1S8.3] | ds | SCRI_RS_156090 | 2H | 81.5 | BY, OA | C, S |
| Probable phospholipid hydroperoxide glutathione peroxidase | GPX1 | [Swiss-Prot:O23968.1] | ds | BOPA1_1635-691 | 2H | 89.8 | BY | S |
| Ethylene receptor 1 | ETR1 | [Swiss-Prot:Q9SSY6.1] | ls | SCRI_RS_185665 | 2H | 114.9 | BY | C, DSI |
| Cullin-3A 3B | CUL3A CUL3B | [Swiss-Prot:Q9ZVH4.1] [Swiss-Prot:Q9C9L0.1] | ls | BOPA1_3608-2133 | 2H | 129.7 | BY | C |
| Senescence-induced receptor-like serine/threonine-protein kinase | SIRK | [Swiss-Prot:O64483.1] | ls | SCRI_RS_8420 | 2H | 139.9 | BY | C |
| Putative F-box/LRR-repeat protein 21 | FBL21 | [Swiss-Prot:Q9M0U8.1] | ds | SCRI_RS_115423 | 3H | 36.3 | OA | S |
| 1-aminocyclopropane-1-carboxylate oxidase | ACO1 | [Swiss-Prot:Q9ZQZ1.1] | ds | SCRI_RS_167825 | 3H | 100.3 | BY, OA | C, S, DSI |
| Dehydrin | DHN 3 | [Swiss-Prot:P12948.1] | ds | BOPA1_ABC13753-1-2-167 | 3H | 105.3 | BY | C |
| ABC transporter D family member 1 | ABCA1 | [Swiss-Prot:Q94FB9.1] | ds | SCRI_RS_142818 | 3H | 148.2 | BY | C |
| Abscisic acid receptor | PYL5 | [Swiss-Prot:Q9FLB1.1] | ds | SCRI_RS_157396 | 4H | 52.3 | OA | S |
| Ethylene-responsive transcription factor | ERF011 | [Swiss-Prot:Q9SNE1.1] | ls | SCRI_RS_9164 | 4H | 113.7 | BY | C, DSI |
| Nucleotide pyrophosphatase/phosphodiesterase | AVP1 | [Swiss-Prot:Q687E1.2] | ds | BOPA1_9766-787 | 5H | 44 | BY | S |
| Abscisic acid-inducible protein kinase | TRIUR3 | [Swiss-Prot:Q02066.1] | ds | SCRI_RS_102075 | 5H | 44 | SPAD | S |
| Serine/threonine-protein kinase | ATM | [Swiss-Prot:Q75H77.1] | ds | BOPA1_ABC08327-1-1-353 | 5H | 44 | SPAD | S |
| Serine/threonine-protein kinase | SAPK9 | [Swiss-Prot:Q75V57.1] | ds | SCRI_RS_102075 | 5H | 44 | SPAD | S |
| Anthocyanin regulatory R-S protein | R-S | [Swiss-Prot:P13027.1] | ds | BOPA1_12045-83 | 5H | 46.7 | BY | S |
| Fasciclin-like arabinogalactan protein 2 | FLA2 | [Swiss-Prot:Q9SU13.1] | ds | BOPA1_5004-375 | 5H | 83.5 | BY | C |
| Serine/threonine-protein kinase | ATM | [Swiss-Prot:Q9M3G7.1] | ds | BOPA1_6315-914 | 5H | 94.7 | SPAD | C |
| Dehydration-responsive element-binding protein 1A | DREB1A | [Swiss-Prot:Q64MA1.1] | ds | BOPA2_12_30852 | 5H | 95 | BY, OA | C, S |
| Probable zinc metalloprotease EGY1 | EGY1 | [Swiss-Prot:Q852K0.3] | ls | SCRI_RS_208686 | 5H | 137.4 | BY | C, DSI |
| Cation/H(+) antiporter 2 | CHX | [Swiss-Prot:Q9SAK8.1] | ds | SCRI_RS_160297 | 7H | 2.1 | ETR | S |
| Ethylene-responsive transcription factor | ERF062 | [Swiss-Prot:Q9SVQ0.1] | ls | SCRI_RS_150783 | 7H | 48.3 | BY | S |
aFunction of the proteins related to drought stress (ds) or leaf senescence (ls), given by UniProt
bMarkers and chromosome positions are based on Comadran et al. (2012)
cBY: biomass yield, CSS: total content of soluble sugars, ETR: electron transport rate at PSII, CFP: content of free proline, OA: osmolality, SPAD: leaf colour
dC: control treatment, S: stress treatment, DSI: drought susceptibility index across treatments by Fischer & Maurer (1978)
Fig. 2.
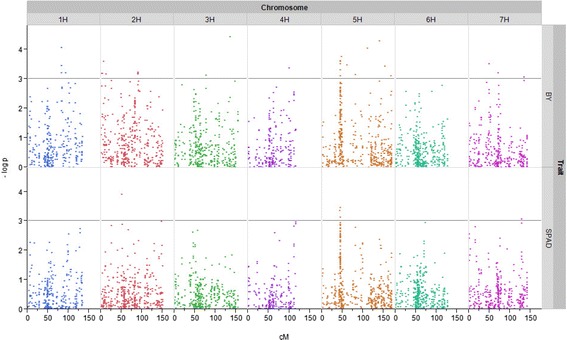
Manhattan plots. Showing –log p of association between Illumina SNPs and the analysed traits biomass yield (BY) and leaf colour (SPAD) against the position of the markers on all seven linkage groups (barley chromosomes) for stress treatment. Threshold indicates a significance level of p <0.001
For SPAD as an indicator of leaf senescence, twelve significant (p <0.001) marker trait associations in the control treatment, six under stress conditions and two across treatments were detected. For the drought stress indicator biomass yield 179 significant marker trait associations were found in the control treatment, 32 in the stress treatment and 87 across treatments. Significant marker trait associations for BY were evenly distributed over all chromosomes. Out of these, eight were identical in the stress treatment, control treatment and across treatments and a high number of 65 marker trait associations were identical in control treatment and across treatments.
In total the 191 significant associations estimated in the control treatment account for 80 genomic regions (LD = 2.52 cM), the 70 significant associations determined in the stress treatment represent 47 genomic regions and the 92 significant associations across treatments (DSI) account for 54 genomic regions. By comparing the localisation of marker trait associations detected, it turned out that significant associations were found for different traits at same positions, e.g. on chromosome 2H at 50 cM for SPAD and OA in the stress treatment and at 120 cM for BY and SPAD in the control treatment. Furthermore, on chromosome 5H at 45 cM significant marker trait associations for BY, SPAD and OA were detected in the stress treatment, and at 95 cM significant associations for BY and SPAD in the control treatment.
Summarizing, overlapping of QTL was found across treatments and for different traits, especially for BY and SPAD, which are also significantly correlated (Table 3). One interesting QTL was observed on chromosome 5H at 45 cM where a significant association to BY and SPAD in the drought stress treatment was found, which is also within the LD of a significant association to OA (Table 4). Therefore, at this position a putative major QTL for drought stress and leaf senescence may be located.
From the 353 significantly associated SNPs detected in the control treatment, stress treatment and across treatments (DSI), 127 proteins were identified by an NCBI Blast of the marker sequences. Out of these 19 proteins turned out to be related to drought stress, 10 proteins related to leaf senescence and 98 proteins turned out to be not related to drought stress or leaf senescence. Out of the 29 proteins for drought stress and leaf senescence (Table 5), 16 revealed associations under drought stress conditions. These were in a next step assigned to the barley chromosomes by the known genetic localization of respective SNPs (Fig. 3). Most of these were located at barley chromosome 2H and 5H, none were mapped on chromosome 6H.
Fig. 3.
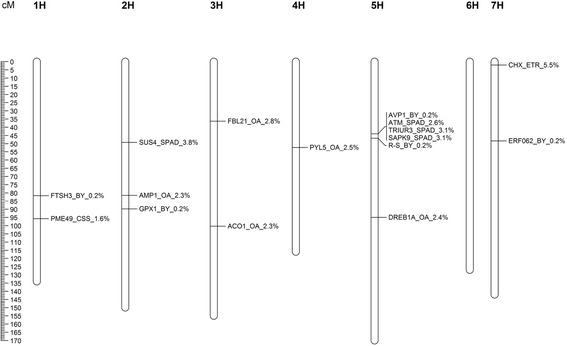
Genetic map. Shows QTL including the significant associated SNP marker positions for significant blasted proteins (BlastX) linked to drought stress or leaf senescence, related traits for drought stress treatment and percentage of phenotypic variance (explained R2 in %) of the SNPs for all linkage groups (barley chromosomes)
Discussion
Using the experimental design described above a clear cut influence of drought stress on biomass yield and physiological parameters was observed (Table 1). This effect of reduced biomass under drought conditions in juvenile stages was also reported in Honsdorf et al. [51] with an even higher reduction due to drought in early developmental stages of barley by Jamieson et al. [25]. In experiments on terminal drought stress application in barley this effect was not so pronounced [25, 79, 80] giving hint that barley is most susceptible to drought stress in early developmental stages. Furthermore, a reduction of the chlorophyll content under drought stress conditions in barley has been observed [80, 81], but there are also reports on adverse effects, e.g. for rapeseed or potato [82, 83], which may be due to a reduced leaf growth under drought stress conditions resulting in a reduced cell expansion leading to a relatively higher chlorophyll density in the leaves. In the present study measurement was done on primary leaves, which were fully expanded at the initiation of drought stress, so that this effect was excluded. For biomass yield a significant correlation to the leaf senescence parameter chlorophyll content was observed (Table 3). This correlation may be based on a true genetic relationship between these parameters, as it is also reported in drought stress field studies on wheat [59, 84] and in glasshouse experiments [85].
The electron transport rate at PS II as a parameter for the chlorophyll fluorescence decreased under drought stress indicating the degradation of chlorophyll during drought stress induced leaf senescence (Table 1), as already shown by Fang et al. [86], Li et al. [27], Netto et al. [87] and Silva et al. [88].
As expected, osmolality increased under drought stress thereby protecting cells against a turgor loss [30]. This is also reported in other drought stress studies on barley [29], but is more often detected in barley under salt stress conditions [89, 90]. For OA no correlation was found to BY, as also shown in a drought stress study on spring barley [91].
The amino acid proline has been described as an osmo-protectant [92] and is accumulated along with several abiotic stresses, such as drought stress, as seen in the present study (Table 1). This effect was also found in other drought stress studies, for example on barley in pot experiments [28], in greenhouse [93] and on Arabidopsis thaliana in climate chamber experiments [94]. The role of proline accumulation is still controversially discussed as it is described to function as a radical scavenger, antioxidant and is involved in the regulation of apoptosis and in seed development [95, 96, 97]. High correlations were detected for CFP to SPAD and BY giving hint that this trait is involved in drought stress tolerance and leaf senescence (Table 3). The correlation of the proline content to SPAD was also found in winter survival studies of barley [98] and in studies on salt stress tolerance in Trigonella foenum-graecum [99]. Up to now no correlations of CFP to BY under drought stress have been described for barley, but positive correlations to yield in wheat were observed under drought stress [100].
Soluble sugars are acting also as osmo-protectants and consequently like in our study an increase was detected in several drought stress experiments on barley [29], wheat [101], potato [102] and also pea [103]. Furthermore, studies showed that an increase of soluble sugars occurs along with leaf senescence [104] and that CSS was correlated to leaf senescence and biomass production [105, 106]. Interactions between sugar and ABA signalling may be responsible for the induction of senescence during drought stress [107].
Quite high values for the heritability of respective traits estimated under drought stress conditions (Table 1) give hint that such an experimental design is suited together with a set of diverse genotypes and the respective number of SNP-markers to detect QTLs using a genome wide association approach (GWAS). Like in other studies [49, 108, 109], the highest number of associations was detected for the traits with the highest heritability. In the present study these were SPAD and BY. Most associations were found on barley chromosomes 2H and 5H on which QTLs were located at 50 cM and at 45 cM, respectively. Also in other GWAS studies of barley significant QTLs for SPAD and BY were located on these chromosomes. Close to the QTL for SPAD located on chromosome 2H (50 cM) a QTL for SPAD under drought stress was also mapped by Li et al. [50]. Moreover, on chromosome 2H at 115 cM a QTL for SPAD was identified in a pot experiment with post-flowering drought stress [53]. QTLs for SPAD were also located on chromosome 2H at 102.7 cM and on chromosome 5H at 165.2 cM in Mediterranean dry land experiments (110), but no significant marker trait associations were detected at these positions in our experiments. The same holds true for a QTL for SPAD on chromosome 5H at 139 cM [49]. Varshney et al. [49] also detected a QTL for biomass yield on chromosome 5H at 95 cM and 156 cM, which is near to associations, which were found in our study on chromosome 5H at 152 cM and 167 cM. In addition, in the present study a QTL for SPAD and BY under drought stress treatment was detected on chromosome 5H at 45 cM, which has not been described before. Furthermore, a lot of significant marker trait associations were observed for osmolality under stress treatment distributed over all barley chromosomes. This was also reported for barley based on growth chamber drought experiments [29].
Proteins involved in drought stress and leaf senescence were detected by a blast of SNP marker sequences and it turned out that they are distributed over all barley chromosomes, except 6H with a focus on chromosomes 2H and 5H (Table 5). Most interesting proteins detected in the drought stress treatment (Fig. 3) are discussed in detail.
On chromosome 1H an ATP-dependent zinc metalloprotease (FTSH3) which is a regulator of heat shock proteins turned out to be associated to BY under drought stress. This protein is involved in the thylakoid formation and in the removal of damaged D1 in the photosystem II, preventing cell death under high-intensity light conditions. In interaction with heat shock proteins it reduces chlorophyll a/b ratios in heat tolerance regulation in Arabidopsis thaliana [111]. Heat stress often occurs simultaneously with drought and also leads to leaf senescence. Besides this, on chromosome 1H the pectin methylesterase 49 (PME49) was found by sequence alignment to be associated to CSS. This protein acts in the modification of cell walls via demethylesterification of pectin and turned out to be up-regulated by drought stress in rice [112]. It influences the mechanical stability of cell walls and thereby also of leaves.
On chromosome 2H a sucrose synthase 4 (SUS4) turned out to be associated with SPAD. This is a sucrose-cleaving enzyme that provides UDP-glucose and fructose for various metabolic pathways and is involved in nucleic acid break down during leaf senescence as revealed by expression analysis e.g. in cucumber and rapeseed [113]. The SNP marker with the homolog sequences to this protein was associated to the leaf senescence parameter SPAD at 49.2 cM. So there may be a direct relationship between the SPAD values and the activation of this enzyme, especially because this SNP marker explains 3.8 % of the phenotypic variance. Furthermore, a probable glutamate carboxypeptidase (AMP) revealed an association to OA which plays an important role in shoot apical meristem development and phytohormone homeostasis. By microarray analysis it turned out that AMP mediates ABA production and is involved in abiotic stress response such as drought stress in Arabidopsis thaliana [114]. Moreover, a marker with a sequence homologue to a phospholipid hydroperoxide glutathione peroxidase (GPX1) associated to BY was found which protects cells and enzymes from oxidative damage. Photometrical analyses of protein quantity and activity showed that the expression of GPX1 and GPX3 is reduced under drought stress and restored after recovery in winter wheat [115].
On chromosome 3H an F-box protein was detected (FBL21) associated to OA. These proteins are ubiquitin related and negatively regulate ABA mediated drought stress response in Arabidopsis thaliana [116]. Furthermore, an association of OA to 1-aminocyclopropane-1-carboxylate oxidase (ACO1) was detected which limits leaf growth by inhibiting the ethylene biosynthesis and so leads to drought tolerance. This was figured out in barley by expression analyses of protein related genes [117]. Surprisingly, in the control treatment associated to BY, a well known drought stress related protein, i.e. dehydrin (DHN) [118] was found which was also in another study located on chromosome 3H [119]. This protein belongs to the family of late embryogenesis abundant (LEA) proteins and is reported to be up-regulated in the protection mechanisms activated by plants in response to drought stress in wheat [120].
On chromosome 4H an ABA receptor (PYL5) was located and associated to OA that activates ABA signalling and ABA-mediated responses such as stomatal closure and germination inhibition. Immuno-detection experiments of protein extracts revealed that ABA signalling is involved in several stresses for example drought stress in Arabidopsis thaliana [121].
On chromosome 5H nucleotide pyrophosphatase/phosphodiesterase (AVP1) was found associated to BY which facilitates auxin transport by modulating apoplastic pH and regulating auxin-mediated developmental processes. Increased expression of protein related genes in transgenic barley confers tolerance to NaCl and to drought by increasing ion retention [122]. Furthermore, three protein kinases associated to SPAD were located on this chromosome regulating protein activity by phosphorylation. First a serine/threonine-protein kinase (ATM) which leads to stress induced programmed cell death, shown in Arabidopsis thaliana by expression profiles of protein related genes [123], second a serine/threonine-protein kinase (SAPK9) which is activated by hyperosmotic stress in rice [124] and third the abscisic acid-inducible protein kinase (TRIUR3) observed in wheat, which is also involved in dehydration stress response [125]. Moreover, an anthocyanin regulatory protein (R-S) was detected associated to BY. Anthocyanin is often accumulated in abiotic stress response, among others in drought stress with a photoprotective function as shown in Arabidopsis thaliana [94, 126]. All of these proteins showed homologies to sequences of SNPs at chromosome 5H around 45 cM and were associated significantly to BY or SPAD, representing an interesting candidate QTL for drought stress and leaf senescence. In addition, a dehydration-responsive element-binding protein (DREB1A) was found associated to OA, which delays water stress symptoms and promotes expression of drought tolerance genes in transgenic wheat [127].
On chromosome 7H another ethylene responsive protein was found to be associated to BY under drought stress. The transcription factor ERF062 is involved in the regulation of gene expression by stress factors (transcriptional repressors) and progression of leaf senescence in Arabidopsis thaliana [128]. Besides, on chromosome 7H the cation/H(+) antiporter 2 (CHX) was detected associated to ETR, which is important for pH gradients in the cell. This protein plays a vital role in maintaining both cellular and intercellular ionic balances under stresses such as drought stress as observed in Arabidopsis thaliana [129].
In summary the blast of the associated SNPs to protein data bases revealed many proteins which are known to be involved in drought stress response or leaf senescence, respectively giving hint that the GWAS approach is well suited for the genetic dissection of these traits in barley. Out of the QTL detected, the ones on chromosome 2H at 50 cM and chromosome 5H at 45 cM are of prime importance and may be involved in breeding barley for drought tolerance in the future due to the quite high amount of phenotypic variance explained.
Conclusions
By GWAS marker trait associations for above ground biomass and physiological traits involved in drought stress tolerance and leaf senescence in early developmental stages of barley were detected. Major QTL for BY and SPAD under drought stress were located at chromosome 2H at 50 cM and chromosome 5H at 45 cM, giving hint that in these regions putative major QTLs for drought stress and leaf senescence are located. With respect to the QTL on chromosome 2H, QTLs for drought stress and leaf senescence were located at comparable positions in other GWAS studies while the one on chromosome 5H was detected for the first time. By BlastX of respective SNP carrying sequences, 29 proteins were identified being involved in drought stress or leaf senescence, respectively. Respective QTLs may be the starting point for marker based selection in barley for drought stress tolerance in the juvenile stage.
Acknowledgements
The authors thank Heike de Lahr for excellent technical assistance, Dr. Ernesto Igartua ICCS, Spain for providing seeds of the SBCC, the Interdisciplinary Center for Crop Plant Research (IZN) of the Martin-Luther-University of Halle-Wittenberg for funding this project and Prof. Dr. Klaus Pillen for close collaboration.
Abbreviations
- e.g.
for example
- i.e.
id est
- GWAS
Genome wide association study
- QTL
Quantitative trait locus
- SNP
Single nucleotide polymorphism
- ABA
Abscisic acid
- LEA
Late embryogenesis abundant protein
- SPAD
Soil Plant Analysis Development; measurement of chlorophyll content by colour
- SBCC
Spanish Barley Core Collection
- das
Days after sowing
- ETR
Electron transport rate
- CFP
Content of free proline
- CSS
Total content of soluble sugars
- OA
Osmolality
- BY
Biomass yield
- PSII
Photosystem two
- PAR
Photosynthetic active radiation
- DSI
Drought susceptibility index; value across treatments
- LSMeans
Last Square Means
- h2
Heritability
- CV
Coefficient of variation
- SSR
Simple sequence repeat
- MAF
Minor allele frequency
- k
number of subpopulations
- MLM
Mixed linear model
- LD
Linkage disequilibrium
- Blast
Basic Local Alignment Search Tool
Additional files
Overview of the 156 analysed genotypes. aSBCC: Spanish Barley Core Collection.
Significant associations (p <0.001) of the genome wide association study and blasted proteins out of the marker sequences. aBY: biomass yield, CSS: total content of soluble sugars, ETR: electron transport rate at PSII, CFP: content of free proline, OA: osmolality, SPAD: leaf colour. bC: control treatment, S: stress treatment, DSI: drought susceptibility index across treatments by Fischer & Maurer (1978). cMarkers and chromosome positions are based on Comadran et al. (2012).
Footnotes
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
GW conducted all experiments, including statistical and bioinformatics analyses, as well as the sequence alignment and mainly wrote the manuscript. ME participated in the GWAS and LD calculation. CB, KH and FO designed the research, supervised the experimental design, data analysis and participated in writing the manuscript. All authors approved the final manuscript.
Contributor Information
Gwendolin G Wehner, Email: gwendolin.wehner@jki.bund.de.
Christiane C Balko, Email: christiane.balko@jki.bund.de.
Matthias M Enders, Email: matthias.enders@jki.bund.de.
Klaus K Humbeck, Email: klaus.humbeck@pflanzenphys.uni-halle.de.
Frank F Ordon, Email: frank.ordon@jki.bund.de.
References
- 1.Harlan JR, Zohary D. Distribution of wild wheats and barley. Science. 1966;153(3740):1074–80. doi: 10.1126/science.153.3740.1074. [DOI] [PubMed] [Google Scholar]
- 2.FAOSTAT. http://faostat3.fao.org/faostat-gateway/go/to/home/E (2014). Accessed 07 Jan 2015.
- 3.Thompson G, Woodward F. Some influences of CO2 enrichment, nitrogen nutrition and competition on grain yield and quality in spring wheat and barley. J Exp Bot. 1994;45(7):937–42. [Google Scholar]
- 4.Jamieson P, Martin R, Francis G. Drought influences on grain yield of barley, wheat, and maize. New Zeal J Crop Hort. 1995;23(1):55–66. [Google Scholar]
- 5.Rollins J, Habte E, Templer S, Colby T, Schmidt J, von Korff M. Leaf proteome alterations in the context of physiological and morphological responses to drought and heat stress in barley (Hordeum vulgare L.) J Exp Bot. 2013;64(11):3201–12. doi: 10.1093/jxb/ert158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Long NV, Dolstra O, Malosetti M, Kilian B, Graner A, Visser RG, et al. Association mapping of salt tolerance in barley (Hordeum vulgare L.) TAG Theoretical and applied genetics Theoretische und angewandte Genetik. 2013;126(9):2335–51. doi: 10.1007/s00122-013-2139-0. [DOI] [PubMed] [Google Scholar]
- 7.El Hafid R, Smith DH, Karrou M, Samir K. Physiological responses of spring durum wheat cultivars to early-season drought in a Mediterranean environment. Ann Bot-London. 1998;81(2):363–70. [Google Scholar]
- 8.Coumou D, Robinson A. Historic and future increase in the global land area affected by monthly heat extremes. Environ Res Let. 2013;8(3):034018. [Google Scholar]
- 9.Ahuja I, de Vos RC, Bones AM, Hall RD. Plant molecular stress responses face climate change. Trends Plant Sci. 2010;15(12):664–74. doi: 10.1016/j.tplants.2010.08.002. [DOI] [PubMed] [Google Scholar]
- 10.Tyagi K, Park MR, Lee HJ, Lee CA, Rehman S, Steffenson B, et al. Fertile crescent region as source of drought tolerance at early stage of plant growth of wild barley (Hordeum vulgare L. ssp. spontaneum) Pakistan J Bot. 2011;43(1):475–86. [Google Scholar]
- 11.Beck EH, Fettig S, Knake C, Hartig K, Bhattarai T. Specific and unspecific responses of plants to cold and drought stress. J Biosci. 2007;32(3):501–10. doi: 10.1007/s12038-007-0049-5. [DOI] [PubMed] [Google Scholar]
- 12.Anjum SA, Xie X-y, Wang L-c, Saleem MF, Man C, Lei W. Morphological, physiological and biochemical responses of plants to drought stress. African J Agri Res. 2011;6(9):2026–32. [Google Scholar]
- 13.Shinozaki K, Yamaguchi-Shinozaki K. Gene networks involved in drought stress response and tolerance. J Exp Bot. 2007;58(2):221–7. doi: 10.1093/jxb/erl164. [DOI] [PubMed] [Google Scholar]
- 14.Lim PO, Kim HJ, Nam HG. Leaf senescence. Ann Rev Plant Biol. 2007;58:115–36. doi: 10.1146/annurev.arplant.57.032905.105316. [DOI] [PubMed] [Google Scholar]
- 15.Lee HK, Cho SK, Son O, Xu Z, Hwang I, Kim WT. Drought stress-induced Rma1H1, a RING membrane-anchor E3 ubiquitin ligase homolog, regulates aquaporin levels via ubiquitination in transgenic Arabidopsis plants. Plant Cell. 2009;21(2):622–41. doi: 10.1105/tpc.108.061994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Qin F, Sakuma Y, Tran L-SP, Maruyama K, Kidokoro S, Fujita Y, et al. Arabidopsis DREB2A-interacting proteins function as RING E3 ligases and negatively regulate plant drought stress–responsive gene expression. Plant Cell. 2008;20(6):1693–707. doi: 10.1105/tpc.107.057380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Xu D, Duan X, Wang B, Hong B, Ho T-HD WR. Expression of a late embryogenesis abundant protein gene, HVA1, from barley confers tolerance to water deficit and salt stress in transgenic rice. Plant Physiol. 1996;110(1):249–57. doi: 10.1104/pp.110.1.249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wang W, Vinocur B, Altman A. Plant responses to drought, salinity and extreme temperatures: towards genetic engineering for stress tolerance. Planta. 2003;218(1):1–14. doi: 10.1007/s00425-003-1105-5. [DOI] [PubMed] [Google Scholar]
- 19.Cattivelli L, Rizza F, Badeck F-W, Mazzucotelli E, Mastrangelo AM, Francia E, et al. Drought tolerance improvement in crop plants: An integrated view from breeding to genomics. Field Crop Res. 2008;105(1–2):1–14. [Google Scholar]
- 20.Zhu J-K. Salt and drought stress signal transduction in plants. Ann Rev Plant Biol. 2002;53:247–73. doi: 10.1146/annurev.arplant.53.091401.143329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tunnacliffe A, Wise MJ. The continuing conundrum of the LEA proteins. Naturwissenschaften. 2007;94(10):791–812. doi: 10.1007/s00114-007-0254-y. [DOI] [PubMed] [Google Scholar]
- 22.Campbell SA, Close TJ. Dehydrins: genes, proteins, and associations with phenotypic traits. New Phytol. 1997;137(1):61–74. [Google Scholar]
- 23.Barnabas B, Jager K, Feher A. The effect of drought and heat stress on reproductive processes in cereals. Plant, Cell Environ. 2008;31(1):11–38. doi: 10.1111/j.1365-3040.2007.01727.x. [DOI] [PubMed] [Google Scholar]
- 24.Farooq M, Wahid A, Kobayashi N, Fujita D, Basra SMA. Plant drought stress: effects, mechanisms and management. Agron Sust Dev. 2009;29(1):185–212. [Google Scholar]
- 25.Jamieson PD, Martin RJ, Francis GS, Wilson DR. Drought effects on biomass production and radiation-use efficiency in barley. Field Crop Res. 1995;43:77–86. [Google Scholar]
- 26.Gonzales A, Martín I, Ayerbe L. Barley yield in water-stress conditions. The influence of precocity, osmotic adjustment and stomatal conductance. Field Crop Res. 1999;62:23–34. [Google Scholar]
- 27.Li R-h, Guo P-g, Michael B, Stefania G, Salvatore C. Evaluation of Chlorophyll Content and Fluorescence Parameters as Indicators of Drought Tolerance in Barley. Agr Sci China. 2006;5(10):751–7. [Google Scholar]
- 28.Sayed MA, Schumann H, Pillen K, Naz AA, Leon J. AB-QTL analysis reveals new alleles associated to proline accumulation and leaf wilting under drought stress conditions in barley (Hordeum vulgare L.) BMC Genet. 2012;13(61):1471–2156. doi: 10.1186/1471-2156-13-61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Teulat B, Borries C, This D. New QTLs identified for plant water status, water-soluble carbohydrate and osmotic adjustment in a barley population grown in a growth-chamber under two water regimes. TAG Theoretical and applied genetics Theoretische und angewandte Genetik. 2001;103(1):161–70. [Google Scholar]
- 30.Blum A. Osmotic adjustment and growth of barley genotypes under drought stress. Crop Sci. 1989;29:230–3. [Google Scholar]
- 31.Verma V, Foulkes MJ, Worland AJ, Sylvester-Bradley R, Caligari PDS, Snape JW. Mapping quantitative trait loci for flag leaf senescence as a yield determinant in winter wheat under optimal and drought-stressed environments. Euphytica. 2004;135:255–63. [Google Scholar]
- 32.Munne-Bosch S, Alegre L. Die and let live: leaf senescence contributes to plant survival under drought stress. Funct Plant Biol. 2004;31:203–16. doi: 10.1071/FP03236. [DOI] [PubMed] [Google Scholar]
- 33.Miersch I, Heise J, Zelmer I, Humbeck K. Differential degradation of the photosynthetic apparatus during leaf senescence in barley (Hordeum vulgare L.) Plant Biol. 2000;2:618–23. [Google Scholar]
- 34.Hörtensteiner S. Chlorophyll drgradation during senescence. Ann Rev Plant Biol. 2006;57:55–77. doi: 10.1146/annurev.arplant.57.032905.105212. [DOI] [PubMed] [Google Scholar]
- 35.Breeze E, Harrison E, McHattie S, Hughes L, Hickman R, Hill C, et al. High-resolution temporal profiling of transcripts during Arabidopsis leaf senescence reveals a distinct chronology of processes and regulation. Plant Cell. 2011;23(3):873–94. doi: 10.1105/tpc.111.083345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Rae A, Ferris R, Tallis M, Taylor G. Elucidating genomic regions determining enhanced leaf growth and delayed senescence in elevated CO2. Plant, Cell Environ. 2006;29(9):1730–41. doi: 10.1111/j.1365-3040.2006.01545.x. [DOI] [PubMed] [Google Scholar]
- 37.Robson PR, Farrar K, Gay AP, Jensen EF, Clifton-Brown JC, Donnison IS. Variation in canopy duration in the perennial biofuel crop Miscanthus reveals complex associations with yield. Journal of experimental botany. 2013;64(8):1–11. [DOI] [PMC free article] [PubMed]
- 38.Rodriguez IR, Miller GL. Using a Chlorophyll Meter to Determine the Chlorophyll Concentration, Nitrogen Concentration, and Visual Quality of St. Augustinegrass. Hort Sci. 2000;34(4):751–4. [Google Scholar]
- 39.Buchanan-Wollaston V, Earl HJ, Harrison E, Mathas E, Navabpour S, Page T, et al. The molecular analysis of leaf senescence – a genomics approach. Plant Biotechnol J. 2003;1:3–22. doi: 10.1046/j.1467-7652.2003.00004.x. [DOI] [PubMed] [Google Scholar]
- 40.Fischer AM. The Complex Regulation of Senescence. Crit Rev Plant Sci. 2012;31(2):124–47. [Google Scholar]
- 41.Buchanan-Wollaston V, Page T, Harrison E, Breeze E, Lim PO, Nam HG, et al. Comparative transcriptome analysis reveals significant differences in gene expression and signalling pathways between developmental and dark/starvation-induced senescence in Arabidopsis. The Plant journal : for cell and molecular biology. 2005;42(4):567–85. doi: 10.1111/j.1365-313X.2005.02399.x. [DOI] [PubMed] [Google Scholar]
- 42.Emebiri LC. QTL dissection of the loss of green colour during post-anthesis grain maturation in two-rowed barley. TAG Theoretical and applied genetics Theoretische und angewandte Genetik. 2013;126(7):1873–84. doi: 10.1007/s00122-013-2102-0. [DOI] [PubMed] [Google Scholar]
- 43.Talukder A, McDonald GK, Gill GS. Effect of short-term heat stress prior to flowering and early grain set on the grain yield of wheat. Field Crop Res. 2014;160:54–63. [Google Scholar]
- 44.Inoue T, Inanaga S, Sugimoto Y, An P, Eneji A. Effect of drought on ear and flag leaf photosynthesis of two wheat cultivars differing in drought resistance. Photosynthetica. 2004;42(2):559–65. [Google Scholar]
- 45.Borrell AK, Hammer GL, Henzell RG. Does maintaining green leaf area in sorghum improve yield under drought? II. Dry matter production and yield. Crop Sci. 2000;40(4):1037–48. [Google Scholar]
- 46.Ashraf M. Inducing drought tolerance in plants: recent advances. Biotechnol Adv. 2010;28(1):169–83. doi: 10.1016/j.biotechadv.2009.11.005. [DOI] [PubMed] [Google Scholar]
- 47.Zhu C, Gore M, Buckler ES, Yu J. Status and Prospects of Association Mapping in Plants. Plant Genome J. 2008;1(1):5. [Google Scholar]
- 48.Miles CM, Wayne M. Quantitative Trait Locus (QTL) Analysis. Nature Education. 2008;1(1):1–6. [Google Scholar]
- 49.Varshney RK, Paulo MJ, Grando S, van Eeuwijk FA, Keizer LCP, Guo P, et al. Genome wide association analyses for drought tolerance related traits in barley (Hordeum vulgare L.) Field Crop Res. 2012;126:171–80. [Google Scholar]
- 50.Li W-T, Liu C, Liu Y-X, Pu Z-E, Dai S-F, Wang J-R, et al. Meta-analysis of QTL associated with tolerance to abiotic stresses in barley. Euphytica. 2013;189(1):31–49. [Google Scholar]
- 51.Honsdorf N, March TJ, Hecht A, Eglinton J, Pillen K. Evaluation of juvenile drought stress tolerance and genotyping by sequencing with wild barley introgression lines. Mol Breeding. 2014;34(3):1475–95. [Google Scholar]
- 52.Teulat B, This D, Khairallah M, Borries C, Ragot C, Sourdille P, et al. Several QTLs involved in osmotic-adjustment trait variation in barley (Hordeum vulgare L.) TAG Theoretical and applied genetics Theoretische und angewandte Genetik. 1998;96(5):688–98. [Google Scholar]
- 53.Guo P, Baum M, Varshney RK, Graner A, Grando S, Ceccarelli S. QTLs for chlorophyll and chlorophyll fluorescence parameters in barley under post-flowering drought. Euphytica. 2008;163(2):203–14. [Google Scholar]
- 54.Nevo E, Chen G. Drought and salt tolerances in wild relatives for wheat and barley improvement. Plant, Cell Environ. 2010;33(4):670–85. doi: 10.1111/j.1365-3040.2009.02107.x. [DOI] [PubMed] [Google Scholar]
- 55.McCouch S. Diversifying selection in plant breeding. PLoS Biol. 2004;2(10):e347. doi: 10.1371/journal.pbio.0020347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Rode J, Ahlemeyer J, Friedt W, Ordon F. Identification of marker-trait associations in the German winter barley breeding gene pool (Hordeum vulgare L.) Mol Breeding. 2012;30(2):831–43. [Google Scholar]
- 57.Igartua E, Gracia MP, Lasa JM, Medina B, Molina-Cano JL, Montoya JL, et al. The Spanish barley core collection. Genet Resour Crop Ev. 1998;45:475–81. doi: 10.1111/j.1601-5223.2001.00217.x. [DOI] [PubMed] [Google Scholar]
- 58.Paech K, Simonis W. Pflanzenphysiologische Praktika Band I Übungen zur Stoffwechselphysiologie der Pflanzen. Berlin Göttingen Heidelberg: Springer Verlag; 1952. [Google Scholar]
- 59.Zaharieva M, Gaulin E, Havaux M, Acevedo E, Monneveux P. Drought and Heat Responses in the Wild Wheat Relative Roth. Crop Sci. 2001;41(4):1321–9. [Google Scholar]
- 60.Xu W, Rosenow DT, Nguyen HT. Stay green trait in grain sorghum: relationship between visual rating and leaf chlorophyll concentration. Plant Breed. 2000;119:365–7. [Google Scholar]
- 61.Cartelat A, Cerovic ZG, Goulas Y, Meyer S, Lelarge C, Prioul JL, et al. Optically assessed contents of leaf polyphenolics and chlorophyll as indicators of nitrogen deficiency in wheat (Triticum aestivum L.) Field Crop Res. 2005;91(1):35–49. [Google Scholar]
- 62.Krall JP, Edwards GE. RelaticHiship between photosystem II activity and CO2 fixation in leaves. Pysiol Plantarum. 1992;86:180–7. [Google Scholar]
- 63.Bates LS, Waldren RP, Teare ID. Rapid determination of free proline for water-stress studies. Plant Soil. 1973;39:205–7. [Google Scholar]
- 64.Yemm EW, Willis AJ. The estimation of carbohydrates in plant extractes by anthrone. Biochem J. 1954;57:508–14. doi: 10.1042/bj0570508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Grubbs FE. Procedures for detecting outlying observations in samples. Technometrics. 1969;11(1):1–21. [Google Scholar]
- 66.Fischer RA, Maurer R. Drought Resistance in Spring Wheat Cultivars. I Grain Yield Responses. Austral J Agr Res. 1978;29:897–912. [Google Scholar]
- 67.SAS. Enterprise Guide 4.3 of SAS System for Microsoft Windows. In Administering SAS Enterprise Guide 4.3. Cary NC, editor. USA: SAS Institute Inc.; 2008.
- 68.Comadran J, Kilian B, Russell J, Ramsay L, Stein N, Ganal M, et al. Natural variation in a homolog of Antirrhinum CENTRORADIALIS contributed to spring growth habit and environmental adaptation in cultivated barley. Nature Genet. 2012;44(12):1388–92. doi: 10.1038/ng.2447. [DOI] [PubMed] [Google Scholar]
- 69.Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics. 2000;155(2):945–59. doi: 10.1093/genetics/155.2.945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Evanno G, Regnaut S, Goudet J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol. 2005;14(8):2611–20. doi: 10.1111/j.1365-294X.2005.02553.x. [DOI] [PubMed] [Google Scholar]
- 71.Hardy OJ, Vekemans X. SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes. 2002;2(4):618–20. [Google Scholar]
- 72.Streiff R, Labbe T, Bacilieri R, Steinkellner H, Glössl J, Kremer A. Within‐population genetic structure in Quercus robur L. and Quercus petraea (Matt.) Liebl. assessed with isozymes and microsatellites. Mol Ecol. 1998;7(3):317–28. [Google Scholar]
- 73.Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics. 2007;23(19):2633–5. doi: 10.1093/bioinformatics/btm308. [DOI] [PubMed] [Google Scholar]
- 74.RCore . R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2012. [Google Scholar]
- 75.Hill WG, Weir BS. Variances and Covariances Disequilibria of Squared Linkage in Finite Populations. Theor Popul Biol. 1988;33:54–78. doi: 10.1016/0040-5809(88)90004-4. [DOI] [PubMed] [Google Scholar]
- 76.Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403–10. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- 77.UniProt The universal protein resource (UniProt) Nucleic Acids Res. 2008;36(suppl 1):D190–D5. doi: 10.1093/nar/gkm895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Voorrips RE. MapChart: Software for the Graphical Presentation of Linkage Maps and QTLs. J Heredity. 2002;93(1):77–8. [DOI] [PubMed]
- 79.Teulat B, Monneveux P, Wery J, Borries C, Souyris I, Charrier A, et al. Relationships between relative water content and growth parameters under water stress in barley: a QTL study. New Phytol. 1997;137:99–107. [Google Scholar]
- 80.Zhao J, Sun H, Dai H, Zhang G, Wu F. Difference in response to drought stress among Tibet wild barley genotypes. Euphytica. 2010;172(3):395–403. [Google Scholar]
- 81.Mamnouie E, Fotouhi Ghazvini R, Esfahani M, Nakhoda B. The effects of water deficit on crop yield and the physiological characteristics of barley (Hordeum vulgare L.) varieties. J Agr Sci Technol. 2010;8:211–9. [Google Scholar]
- 82.Ramírez DA, Yactayo W, Gutiérrez R, Mares V, De Mendiburu F, Posadas A, et al. Chlorophyll concentration in leaves is an indicator of potato tuber yield in water-shortage conditions. Sci Hortic -Amsterdam. 2014;168:202–9. [Google Scholar]
- 83.Khan MA, Ashraf M, Mujtaba S, Shirazi M, Khan M, Shereen A, et al. Evaluation of high yielding canola type Brassica genotypes/mutants for drought tolerance using physiological indices as screening tool. Pakistan J Bot. 2010;42(6):3807–16. [Google Scholar]
- 84.Babar M, Reynolds M, Van Ginkel M, Klatt A, Raun W, Stone M. Spectral reflectance to estimate genetic variation for in-season biomass, leaf chlorophyll, and canopy temperature in wheat. Crop Sci. 2006;46(3):1046–57. [Google Scholar]
- 85.Fotovat R, Valizadeh M, Toorchi M. Association between water-use efficiency components and total chlorophyll content (SPAD) in wheat (Triticum aestivum L.) under well-watered and drought stress conditions. J Food Agr Env. 2007;5(3/4):225. [Google Scholar]
- 86.Fang Z, Bouwkamp JC, Solomos T. Chlorophyllase activities and chlorophyll degradation during leaf senescence in non-yellowingmutant and wild type of Phaseolus vulgaris L. J Exp Bot. 1998;49(320):503–10. [Google Scholar]
- 87.Netto AT, Campostrini E, Oliveira JG, Bressan-Smith RE. Photosynthetic pigments, nitrogen, chlorophyll a fluorescence and SPAD-502 readings in coffee leaves. Sci Hortic -Amsterdam. 2005;104(2):199–209. [Google Scholar]
- 88.Silva MA, Jifon JL, Da Silva JAG, Sharma V. Use of physiological parameters as fast tools to screen for drought tolerance in sugarcane. Braz J Plant Physiol. 2007;3(19):193–201. [Google Scholar]
- 89.Veselov D, Sharipova G, Veselov S, Kudoyarova G. The effects of NaCl treatment on water relations, growth, and ABA content in barley cultivars differing in drought tolerance. J Plant Growth Regul. 2008;27(4):380–6. [Google Scholar]
- 90.Fricke W, Peters WS. The biophysics of leaf growth in salt-stressed barley. A study at the cell level. Plant Physiol. 2002;129(1):374–88. doi: 10.1104/pp.001164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Szira F, Bálint AF, Börner A, Galiba G. Evaluation of Drought-Related Traits and Screening Methods at Different Developmental Stages in Spring Barley. J Agr Crop Sci. 2008;194(5):334–42. [Google Scholar]
- 92.Delauney AJ, Verma DPS. Proline biosynthesis and osmoregulation in plants. The Plant journal : for cell and molecular biology. 1993;4(2):215–23. [Google Scholar]
- 93.Bandurska H. Does proline accumulated in leaves of water deficit stressed barley plants confine cell membrane injury? I. Free proline accumulation and membrane injury index in drought and osmotically stressed plants. Acta Physiol Plant. 2000;22(4):409–15. [Google Scholar]
- 94.Sperdouli I, Moustakas M. Interaction of proline, sugars, and anthocyanins during photosynthetic acclimation of Arabidopsis thaliana to drought stress. J Plant Physiol. 2012;169(6):577–85. doi: 10.1016/j.jplph.2011.12.015. [DOI] [PubMed] [Google Scholar]
- 95.Kuznetsov VV, Shevyakova N. Proline under stress: biological role, metabolism, and regulation. Russ J Plant Physl. 1999;46(2):274–87. [Google Scholar]
- 96.Kishor P, Kavi B, Sreenivasulu N. Is proline accumulation per se correlated with stress tolerance or is proline homeostasis a more critical issue? Plant, Cell Environ. 2014;37(2):300–11. doi: 10.1111/pce.12157. [DOI] [PubMed] [Google Scholar]
- 97.Kocsy G, Laurie R, Szalai G, Szilágyi V, Simon‐Sarkadi L, Galiba G, et al. Genetic manipulation of proline levels affects antioxidants in soybean subjected to simultaneous drought and heat stresses. Physiol Plant. 2005;124(2):227–35. [Google Scholar]
- 98.Sofalian O, Behi M. Assessment of winter survival in barley (Hordeum vulgare L.) genotypes using molecular markers and some physiological traits. J Agr Fac Gaziosmanpasa University. 2013;30(2):45–54. [Google Scholar]
- 99.Al-Saady N, Khan A, Rajesh L, Esechie H. Effect of salt stress on germination, proline metabolism and chlorophyll content of fenugreek (Trigonella foenum gracium L.) Plant science : an international journal of experimental plant biology. 2012;7:176–85. [Google Scholar]
- 100.Keyvan S. The effects of drought stress on yield, relative water content, proline, soluble carbohydrates and chlorophyll of bread wheat cultivars. J Anim Plant Sci. 2010;8:1051–60. [Google Scholar]
- 101.Bajji M, Lutts S, Kinet J-M. Water deficit effects on solute contribution to osmotic adjustment as a function of leaf ageing in three durum wheat (Triticum durum Desf.) cultivars performing differently in arid conditions. Plant science : an international journal of experimental plant biology. 2001;160(4):669–81. doi: 10.1016/s0168-9452(00)00443-x. [DOI] [PubMed] [Google Scholar]
- 102.Basu PS, Sharma A, Sukumaran NP. Changes in net photosynthetic rate and chlorophyll fluorescence in potato leaves induced by water stress. Photosynthetica. 1998;35(1):13–9. [Google Scholar]
- 103.Sanchez FJ, Manzanares M, de Andres EF LTJ, Ayerbe L. Tugor maintenance, osmotic adjustment and soluble sugar and proline accumulation in 49 pea cultivars in response to water stress. Field Crop Res. 1998;59:225–35. [Google Scholar]
- 104.Wingler A, Purdy S, MacLean JA, Pourtau N. The role of sugars in integrating environmental signals during the regulation of leaf senescence. J Exp Bot. 2006;57(2):391–9. doi: 10.1093/jxb/eri279. [DOI] [PubMed] [Google Scholar]
- 105.Rajcan I, Dwyer LM, Tollenaar M. Note on relationship between leaf soluble carbohydrate and chlorophyll concentrations in maize during leaf senescence. Field Crop Res. 1999;63(1):13–7. [Google Scholar]
- 106.Zebarjadi A, Kakaei M, Mostafaie A. Genetic variability of some traits in Rapeseed (Brassica napus L.) under drought stress and non-stress conditions. Biharean Biologist. 2011;5(2):127–31. [Google Scholar]
- 107.Wingler A, Roitsch T. Metabolic regulation of leaf senescence: interactions of sugar signalling with biotic and abiotic stress responses. Plant Biol. 2008;10(Suppl 1):50–62. doi: 10.1111/j.1438-8677.2008.00086.x. [DOI] [PubMed] [Google Scholar]
- 108.Wang M, Jiang N, Jia T, Leach L, Cockram J, Waugh R, et al. Genome-wide association mapping of agronomic and morphologic traits in highly structured populations of barley cultivars. Theor Appl Genet. 2012;124(2):233–46. doi: 10.1007/s00122-011-1697-2. [DOI] [PubMed] [Google Scholar]
- 109.Cockram J, White J, Zuluaga DL, Smith D, Comadran J, Macaulay M, et al. Genome-wide association mapping to candidate polymorphism resolution in the unsequenced barley genome. Proc Natl Acad Sci U S A. 2010;107(50):21611–6. doi: 10.1073/pnas.1010179107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.von Korff M, Grando S, Del Greco A, This D, Baum M, Ceccarelli S. Quantitative trait loci associated with adaptation to Mediterranean dryland conditions in barley. TAG Theoretical and applied genetics Theoretische und angewandte Genetik. 2008;117(5):653–69. doi: 10.1007/s00122-008-0787-2. [DOI] [PubMed] [Google Scholar]
- 111.Wagner R, Aigner H, Funk C. FtsH proteases located in the plant chloroplast. Physiol Plant. 2012;145(1):203–14. doi: 10.1111/j.1399-3054.2011.01548.x. [DOI] [PubMed] [Google Scholar]
- 112.Liu J-X, Bennett J. Reversible and irreversible drought-induced changes in the anther proteome of rice (Oryza sativa L.) genotypes IR64 and Moroberekan. Mol Plant. 2011;4(1):59–69. doi: 10.1093/mp/ssq039. [DOI] [PubMed] [Google Scholar]
- 113.Buchanan-Wollaston V. The molecular biology of leaf senescence. J Exp Bot. 1997;48(307):181–99. [Google Scholar]
- 114.Shi Y, Wang Z, Meng P, Tian S, Zhang X, Yang S. The glutamate carboxypeptidase AMP1 mediates abscisic acid and abiotic stress responses in Arabidopsis. New Phytol. 2013;199(1):135–50. doi: 10.1111/nph.12275. [DOI] [PubMed] [Google Scholar]
- 115.Simova-Stoilova L, Demirevska K, Petrova T, Tsenov N, Feller U. Antioxidative protection in wheat varieties under severe recoverable drought at seedling stage. Plant Soil Environ. 2008;54(12):529–36. [Google Scholar]
- 116.Lyzenga WJ, Stone SL. Abiotic stress tolerance mediated by protein ubiquitination. Journal of experimental botany. 2012;63(2):1–18. [DOI] [PubMed]
- 117.Sobeih WY, Dodd IC, Bacon MA, Grierson D, Davies WJ. Long-distance signals regulating stomatal conductance and leaf growth in tomato (Lycopersicon esculentum) plants subjected to partial root-zone drying. J Exp Bot. 2004;55(407):2353–63. doi: 10.1093/jxb/erh204. [DOI] [PubMed] [Google Scholar]
- 118.Rizhsky L, Liang H, Mittler R. The combined effect of drought stress and heat shock on gene expression in tobacco. Plant Physiol. 2002;130(3):1143–51. doi: 10.1104/pp.006858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Choi D-W, Zhu B, Close T. The barley (Hordeum vulgare L.) dehydrin multigene family: sequences, allele types, chromosome assignments, and expression characteristics of 11 Dhn genes of cv Dicktoo. TAG Theoretical and applied genetics Theoretische und angewandte Genetik. 1999;98(8):1234–47. [Google Scholar]
- 120.Rampino P, Pataleo S, Gerardi C, Mita G, Perrotta C. Drought stress response in wheat: physiological and molecular analysis of resistant and sensitive genotypes. Plant, Cell Environ. 2006;29(12):2143–52. doi: 10.1111/j.1365-3040.2006.01588.x. [DOI] [PubMed] [Google Scholar]
- 121.Santiago J, Rodrigues A, Saez A, Rubio S, Antoni R, Dupeux F, et al. Modulation of drought resistance by the abscisic acid receptor PYL5 through inhibition of clade A PP2Cs. The Plant journal : for cell and molecular biology. 2009;60(4):575–88. doi: 10.1111/j.1365-313X.2009.03981.x. [DOI] [PubMed] [Google Scholar]
- 122.Schilling RK, Marschner P, Shavrukov Y, Berger B, Tester M, Roy SJ, et al. Expression of the Arabidopsis vacuolar H + −pyrophosphatase gene (AVP1) improves the shoot biomass of transgenic barley and increases grain yield in a saline field. Plant Biotechnol J. 2013;12(3):378–86. doi: 10.1111/pbi.12145. [DOI] [PubMed] [Google Scholar]
- 123.Garcia V, Bruchet H, Camescasse D, Granier F, Bouchez D, Tissier A. AtATM is essential for meiosis and the somatic response to DNA damage in plants. Plant Cell. 2003;15(1):119–32. doi: 10.1105/tpc.006577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Kobayashi Y, Yamamoto S, Minami H, Kagaya Y, Hattori T. Differential activation of the rice sucrose nonfermenting1–related protein kinase2 family by hyperosmotic stress and abscisic acid. Plant Cell. 2004;16(5):1163–77. doi: 10.1105/tpc.019943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Anderberg RJ, Walker-Simmons M. Isolation of a wheat cDNA clone for an abscisic acid-inducible transcript with homology to protein kinases. Proc Natl Acad Sci U S A. 1992;89(21):10183–7. doi: 10.1073/pnas.89.21.10183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Chalker‐Scott L. Environmental significance of anthocyanins in plant stress responses. Photochem Photobiol. 1999;70(1):1–9. [Google Scholar]
- 127.Pellegrineschi A, Reynolds M, Pacheco M, Brito RM, Almeraya R, Yamaguchi-Shinozaki K, et al. Stress-induced expression in wheat of the Arabidopsis thaliana DREB1A gene delays water stress symptoms under greenhouse conditions. Genome/National Research Council Canada = Genome/Conseil national de recherches Canada. 2004;47(3):493–500. doi: 10.1139/g03-140. [DOI] [PubMed] [Google Scholar]
- 128.Koyama T, Nii H, Mitsuda N, Ohta M, Kitajima S, Ohme-Takagi M, et al. A regulatory cascade involving class II ETHYLENE RESPONSE FACTOR transcriptional repressors operates in the progression of leaf senescence. Plant Physiol. 2013;162(2):991–1005. doi: 10.1104/pp.113.218115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Song W-Y, Zhang Z-B, Shao H-B, Guo X-L, Cao H-X, Zhao H-B, et al. Relationship between calcium decoding elements and plant abiotic-stress resistance. Int J Biol Sci. 2008;4(2):116. doi: 10.7150/ijbs.4.116. [DOI] [PMC free article] [PubMed] [Google Scholar]


