Abstract
Pseudorabies has been controlled efficiently in China for many years by vaccination. However, it suddenly broke out in many pig farms in 2012–2013 in southern China. In this study, a systematic investigation that included virus isolation, genetic and pathological studies, and immunogenicity analysis was carried out with the aim of understanding the pathogenetic and antigenic features of novel isolates of pseudorabies virus (PRV). Of 38 tissue samples collected from pigs with clinical signs of pseudorabies on 13 farms in 4 provinces in southern China in 2012–2013, 29 showed wild-type PRV infection by polymerase chain reaction. Sequence analysis of 5 isolates from the 4 provinces showed that they belonged to a relatively independent cluster that shared 2 insertions of a single amino acid in the gE gene and 1 insertion of 7 amino acids in the gC gene. In experiments, isolate ZJ01 caused death in 100% of pigs that were either 14 or 80 days old. The serum antibodies to the commercial PRV vaccines had significantly lower neutralizing activity against the ZJ01 isolate than against the vaccine strains. The antigenic relatedness between ZJ01 and the vaccine strains was 0.378 to 0.455. These findings indicated that a novel, highly virulent PRV strain with antigenic variance had spread widely in southern China.
Résumé
La pseudorage a été maitrisée de manière efficace en Chine pendant plusieurs années grâce à la vaccination. Toutefois, en 2012–2013 des poussées de cas sont apparues soudainement dans des fermes porcines dans le sud de la Chine. Dans la présente enquête, l’isolement viral, des études génétiques et pathologiques, et des analyses d’immunogénicité furent effectués avec l’objectif de comprendre les caractéristiques pathogénétiques et antigéniques des nouveaux isolats du virus de la pseudorage (VPR). À partir de 38 échantillons de tissu prélevés en 2012–2013 chez des porcs avec des signes cliniques de pseudorage provenant de 13 fermes dans quatre provinces du sud de la Chine, 29 se sont révélés positifs par réaction d’amplification en chaîne par la polymérase pour une infection par une souche sauvage du VPR. L’analyse de séquence de cinq isolats provenant des quatre provinces montrait qu’ils appartenaient à un regroupement relativement indépendant qui partageait deux insertions d’un acide aminé unique dans le gène gE et une insertion de sept acides aminés dans le gène gC. Lors d’expériences, l’isolat ZJ01 causait la mort chez 100 % des porcs qui étaient âgés de 14 ou 80 jours. Les anticorps sériques envers les vaccins VPR commerciaux avaient une activité neutralisante significativement moindre contre l’isolat ZJ01 que contre les souches vaccinales. La parenté antigénique entre ZJ01 et les souches vaccinales variait de 0,378 à 0,455. Ces trouvailles indiquent qu’une souche nouvelle et hautement virulente du VPR avec une variance antigénique s’est répandue largement dans le sud de la Chine.
(Traduit par Docteur Serge Messier)
Introduction
Pseudorabies virus (PRV), a member of the subfamily Alphaher-pesvirinae of the family Herpesviridae, causes fatal encephalitis in newborn pigs, respiratory disorders in growing and fattening pigs, and reproductive failure in sows (1–4). It establishes a lifelong latent infection in the peripheral nervous system, which increases the difficulty in eradicating pseudorabies (4,5). The most commonly used serologic assay in PRV diagnosis has been a blocking enzyme-linked immunosorbent assay (ELISA) for detecting the presence of antibody against glycoprotein gE in field strains of PRV but not PRV vaccine strains that lack gE, such as Bartha-K61 and Bucharest (6). By using the marker vaccines that lack the nonessential gE and an accompanying serologic test that differentiates infected and vaccinated animals, the United States and some European countries had eradicated pseudorabies from domestic pig populations (4,7).
In China, the first pseudorabies outbreak was reported in the 1950s, and antiviral vaccination had contributed effectively to pseudorabies control during the past several decades. The scale of pig farms developed quickly, and almost every pig herd was inoculated with the PRV live vaccines. Many farms had pigs that were seropositive for PRV antibody against glycoprotein gB but negative for antibody against gE for a long time. However, at the end of 2011, a severe PRV outbreak took place in many PRV-vaccinated pig farms and spread quickly among the pig herds in northern China, causing great economic losses to the swine industry. This outbreak was characterized by fatal encephalitis in newborn pigs and reproductive failure in sows (8–10). The onset of clinical signs in 2- to 3-day-old piglets was sudden, and death occurred on average after 5 h. In addition, grower–finisher pigs as well as newborn pigs were infected and died, which was unlike previous infection with classic PRV strains in China.
In this study, by clinical investigation of some pig farms in southern China in 2012–2013, along with virus isolation, genomic sequencing, pig-challenge experiments, and cross serum virus neutralization tests, we found that highly virulent, antigenic-variant novel strains had caused this unparalleled PRV outbreak that spread wildly in both northern and southern China.
Materials and methods
Clinical cases
Thirty-eight samples of brain, lung, liver, and lymph nodes were collected in 2012–2013 from 13 pig farms in Zhejiang, Guangdong, Guanxi, and Jiangxi provinces in southern China that had used vaccination with PRV live vaccines and in which pigs had clinical signs of pseudorabies, mainly high temperature (40°C to 41.5°C), abortion, and severe neurologic disorders in piglets; the morbidity rate was 30% to 80% and the mortality rate 20% to 60% among the farms. Infection with PRV was diagnosed by polymerase chain reaction (PCR) (5) with the primers shown in Table I.
Table I.
Oligonucleotide primers and probe used in this study
| Application; primers and probe | Primer sequence (5′–3′) |
|---|---|
| Diagnosis | |
| gE-CF | TGCGACGCCGTGGCGGTGACCA |
| gE-.R | ACGGTCAGGTGCGGCGACCACC |
| Amplify gE gene | |
| PRV-gE-F | ATGCGGCCCTTTCTGCTGCG |
| PRV-gE-R | TTAAGCGGGGCGGGACATCAACA |
| Amplify gC gene | |
| PRV-gC-F | GTACGTGACCGTCGCCATGTGT |
| PRV-gC-R | ACAAACAAACAACCGGACGCGAT |
| Real-time PCR | |
| PRV-gE-694F | CTTCCACTCGCAGCTCTTCTC |
| PRV-gE-765R | GTRAAGTTCTCGCGCGAGT |
| Probe | FAM-TTCGACCTGATGCCGC-TARAM |
F — forward; R — reverse; PRV — pseudorabies virus; PCR — polymerase chain reaction.
Virus isolation and identification
The brain tissue samples, from 5 pig farms in Zhejiang, Guangdong, Guanxi, and Jiangxi provinces, were homogenized with Dulbecco’s modified Eagle’s medium. The supernatant was inoculated into BHK-21 cells and incubated for 3 d for evidence of cytopathic effect. Collected viruses were plaque-purified for 3 cycles and their identity confirmed by PCR.
Sequencing analysis of gE and gC genes
Viral DNA was extracted from the isolates or tissue samples with the DNA Miniprep Kit (Omega, Norcross, Georgia, USA). Primers (Table I) were designed on the basis of PRV nucleotide sequences with GenBank accession numbers JF797218 and JF797219 (National Center for Biotechnology Information, Bethesda, Maryland, USA) to amplify the gE and gC genes of PRV isolates. The PCR products were cloned into pEASY-Blunt Zero plasmid and sequenced by Invitrogen (Shanghai, China). Phylogenetic trees were constructed according to the gE and gC gene sequences of PRV (1,2,5,11–14) in GenBank with MEGA software, version 6.0 (www.megasoftware.net) by the neighbor-joining method with 1000 bootstrap replicates (15). Evolutionary distances were computed by the pairwise distance method with the maximum composite likelihood model.
Animal-challenge experiments
The pigs used in the experiments were free of infection with PRV, Porcine circovirus type 2 (PCV2), Porcine reproductive and respiratory syndrome virus (PRRSV), Classical swine fever virus (CSFV), and Porcine parvovirus (PPV) and were housed in a separate animal facility. The experimental protocols were approved by the Institutional Animal Care and Ethics Committee of Nanjing Agricultural University (permit IACECNAU20130102) and met the standards of the International Guiding Principles for Biomedical Research Involving Animals (16). The pigs had been born to unvaccinated sows and were free of antibodies against PRV before the start of the experiment.
Ten 14-day-old piglets were randomly assigned to 2 groups of 5 piglets each and housed separately. Group 1 was challenged intranasally with the ZJ01 strain of PRV at a median tissue culture infective dose (TCID50) of 106.0 as previously described (17). Group 2 was inoculated with phosphate-buffered saline (PBS) and served as an uninfected control group. Fifteen 80-day-old piglets were randomly assigned to 3 groups of 5 piglets each and housed separately. Groups 1 and 2 were challenged intranasally with 107.0 and 106.0 TCID50 of ZJ01, respectively. Group 3 was inoculated with PBS.
After challenge, all the animals were monitored daily for 14 d for rectal temperature and clinical signs. Gross pathological changes were noted and organ tissues collected from the piglets after death or euthanasia at the end of the experimental period for real-time PCR and histopathological examination.
Real-time PCR
The load of PRV in the tissue samples from the infected pigs was determined by real-time PCR for the gE gene as previously described (18) and was expressed as log10 copies per gram.
Cross serum neutralization assay
Twenty 30-day-old piglets free of specific pathogens were randomly assigned to 4 groups of 5 piglets each and housed separately. Groups 1 to 3 were inoculated intranasally with live PRV vaccine containing the strains Bartha-K61 (Pfizer Animal Health, Lincoln, Nebraska, USA), Bucharest (Pfizer Animal Health), and HB-98 (Wuhan Keqian Animal Biological Products, Wuhan, China), respectively, according to the manufacturers’ instructions. Group 4 was inoculated intranasally with 102.0 TCID50 of ZJ01. Blood samples were collected from each pig on day 28 after inoculation.
All tested serum was studied with PRV commercial ELISA kits (IDEXX Laboratories, Westbrook, Maine, USA). The titers of serum neutralization antibody to the different PRV strains were calculated by the Reed–Muench method (19,20,24). The antigenic relatedness (R) between the ZJ01 isolate and the vaccine strain Bartha-K61 was calculated as a geometric mean of 2 ratios between the heterologous and homologous neutralizing antibody titers, as previously described (20–22), such that
where: r1 = heterologous titer 1/homologous titer 1; r2 = heterologous titer 2/homologous titer 2; R > 80% means that the 2 viruses have similar antigenicity and belong to the same subtype, and 80 ≤ R ≤ 25 means that the 2 viruses have antigenic variance and belong to different subtypes.
Statistical analysis
Comparisons of viral loads, neutralizing antibodies titers, and antigenic relatedness were carried out with use of the SYSTAT 9 software package (SPSS, Chicago, Illinois, USA). Differences were considered statistically significant at P < 0.05.
Results
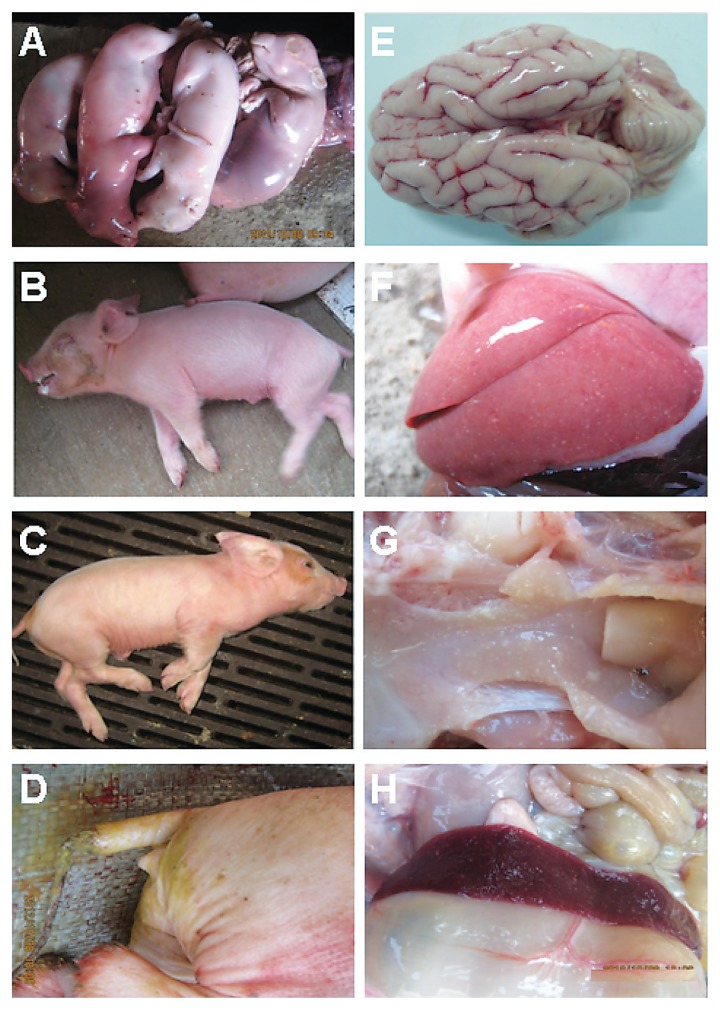
The gross lesions of the ill pigs at the 13 investigated pig farms in southern China included severe hemorrhages in the brain and multiple small focal areas of necrosis in some organs, such as lung, tonsil, and spleen (Figure 1).
Figure 1.
Clinical signs and gross lesions observed in the pigs with pseudorabies. A — Abortion. B and C — Severe neurologic disorders. D — Diarrhea. E — Cerebral edema and hemorrhage. F, G, and H — Lung, tonsil, and spleen with multiple small focal areas of necrosis.
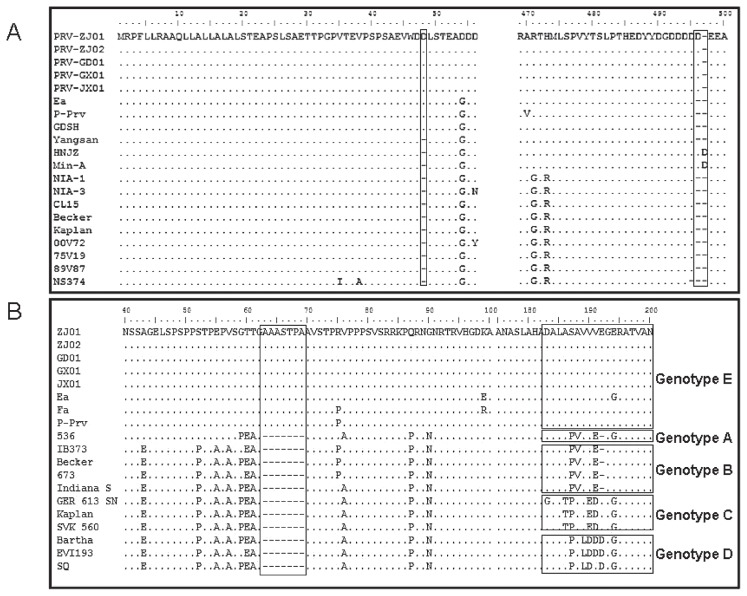
Of the 38 samples of brain, lung, liver, and lymph nodes from clinically ill pigs on the farms, 29 were found to be positive for the PRV gE gene by PCR. Five plaque-purified PRV isolates, ZJ01, ZJ02, GD01, GX01, and JX01, were obtained from 5 of the farms, which were in 4 provinces. The gE and gC genes of PRV were amplified and sequenced from the 5 new isolates. As shown in Figure 2, these 5 isolates all shared an insertion of 1 amino acid (D) at position 48 in the gE gene and an insertion of 7 amino acids (AAASTPA) between positions 63 and 69 in the gC gene as compared with the classic PRV strains Becker and Kaplan; only strains Ea, and P-Prv shared these insertions. Another insertion of 1 amino acid (D), at position 496 in the gE gene, was shared by the 5 new isolates but not by strains Ea, P-Prv, and SA215. The phylogenetic trees (Figure 3) showed that these 5 isolates were in an independent cluster based on the gE and gC genes and were closely related to the traditional PRV isolates in China and other Asian countries.
Figure 2.
Amino acid sequence alignment of the gE and gC genes of pseudorabies virus (PRV). A — New PRV strains ZJ01, ZJ02, GD01, GX01, and JX01 have an insertion of 1 amino acid at positions 48 and 496 in gE. B — An insertion of 7 amino acids between positions 63 and 69 in gC and several mutations between positions 180 and 200 are shared by the 5 new strains and some strains isolated previously in China but not others.
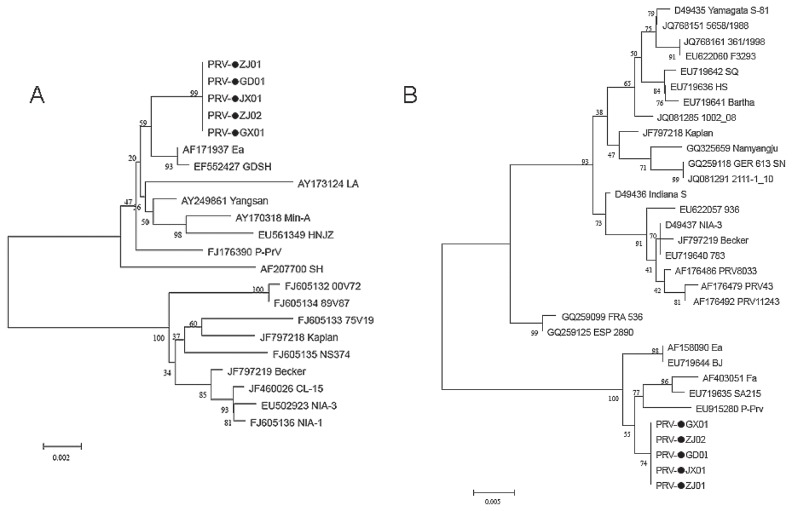
Figure 3.
Phylogenetic trees of PRV based on the gE (A) and gC (B) genes. The 5 new strains are all in an independent cluster that is closely related to the traditional PRV isolates in China and other Asian countries.
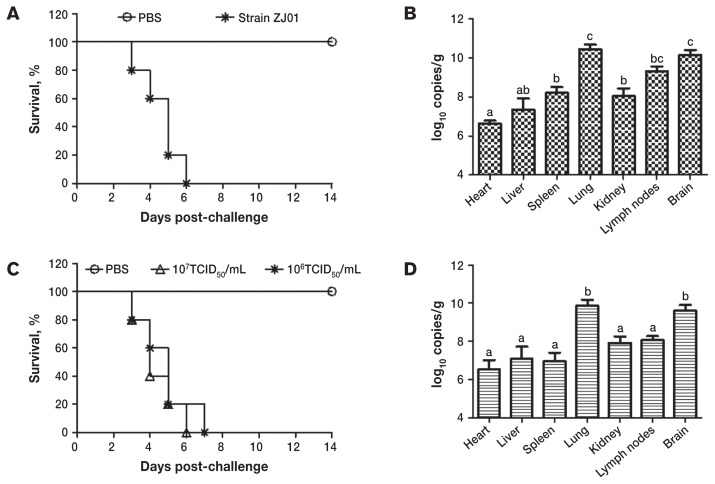
After challenge with PRV strain ZJ01 the 14-day-old and 80-day-old piglets showed the same clinical signs, which included high temperature (40°C to 41.5°C), cough, and severe neurologic disorders. All died within 7 d after challenge (Figures 4A and 4C). Gross lesions were found in multiple organs, including brain, lung, and liver (Figures 5A to 5C); the histologic changes are shown in Figures 5D to 5F. The viral DNA loads in brain and lung tissue from the infected piglets were higher than those in other organ tissues (Figures 4B and 4D).
Figure 4.
Survival and viral DNA load, as determined by real-time polymerase chain reaction, after challenge with PRV strain ZJ01 in 14-day-old pigs (A and B) and 80-day-old pigs (C and D); for the latter, 2 different inoculation doses were used. In each experiment a control group was inoculated with phosphate-buffered saline (PBS). TCID50— median tissue culture infective dose. Different letters (a to c) indicate a significant difference (P < 0.05) between the tissues.
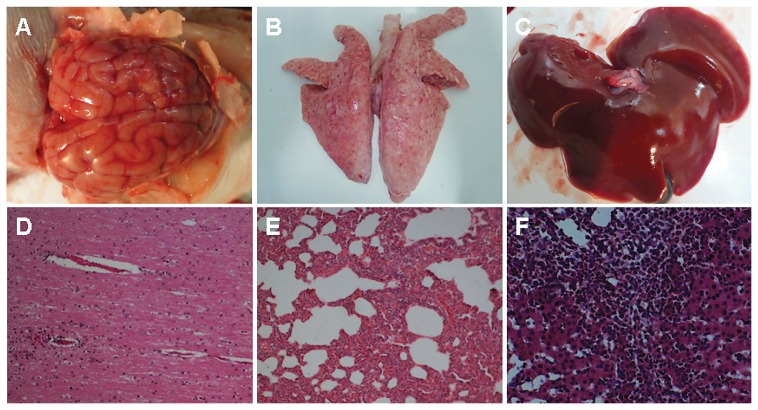
Figure 5.
Gross lesions and histopathological features of tissues stained with hematoxylin and eosin from piglets challenged with ZJ01. A — Cerebral edema and hemorrhage. B — Lung with many blood spots. C — Liver with multiple small focal areas of necrosis. D — Nonsuppurative meningoencephalitis; original magnification ×100. E — Lung tissue with severe hemorrhage and congestion; original magnification ×100. F — Liver tissue with hemorrhage, hepatocyte necrosis, and inflammatory cell infiltration; original magnification ×200.
As shown in Table II, the anti-PRV serum antibodies induced by the vaccine strains provided poor neutralizing activity against strain ZJ01. However, the neutralizing titers of anti-ZJ01 serum against ZJ01 were similar to those against the vaccine strains. The antigenic relatedness between ZJ01 and the vaccine strains was 0.378 to 0.455, which indicates that the ZJ01 isolate changed in antigen compared with the vaccine strains.
Table II.
Results of enzyme-linked immunosorbent assay (ELISA) of neutralizing serum antibodies against 3 PRV vaccine strains as well as the ZJ01 isolate in cross-reaction assays and antigenic relatedness (R) of the strains
| Neutralization titer (mean ± standard error) | |||||
|---|---|---|---|---|---|
|
|
|||||
| Serum antibody | gB-ELISAa | gE-ELISAa | Bartha-K61 | ZJ01 | R |
| Anti-Bartha-K61 | + | − | 1:54.5 ± 17.7 | 1:6.5 ± 3.8 | 0.432 |
| Anti-Bucharest | + | − | 1:53.4 ± 11.1 | 1:7.7 ± 5.4 | 0.378 |
| Anti-HB-98 | + | − | 1:66.6 ± 14.1 | 1:13.9 ± 8.3 | 0.455 |
| Anti-ZJ01 | + | + | 1:69.1 ± 14.7 | 1:69.8 ± 11.4 | |
S/N < 0.60 = positive; S/N > 0.60 and ≤ 0.70 = suspect; S/N > 0.70 = negative.
Discussion
Although pseudorabies results in fetal death and/or abortion in pregnant sows and is often fatal in young piglets, it causes only poor weight gain in grower–finisher pigs unless there is a secondary infection (1,2,4). During the epidemic period of 2011–2013 vaccinated grower–finisher pigs had an average mortality rate of 3% to 6% in comparison with the previous rate of 1% to 2% with the PRV strains that had been traditional in China (e.g., Ea, LA, FA and SC) (10,23,24). After clinical investigation, pathological examination, and detection of wild-type PRV in piglets from 4 provinces, we proved that PRV had spread in southern China again. Susceptibility to PRV infection was mainly dependent on the virulence of the virus strain, the dose of the virus, the age of the pigs, and the immune condition of the animals. In this study, the animal experiments showed that PRV strain ZJ01 induced severe disease not only in 14-day-old piglets but also in 80-day-old pigs, with a 100% death rate and substantial multiorgan lesions. Maybe the high mortality rate was due to the high dose used in the experiment, because even less virulent strains can cause symptoms at a dose of 107.0 TCID50 (17). But another experiment in 80-day-old pigs also showed that ZJ01 caused a 100% death rate, whereas the classic virulent PRV strain LA caused no deaths with the same challenge dose of 106.0 TCID50 (data not shown). Thus, ZJ01 is highly virulent. The average mortality rate of 3% to 6% in grower–finisher pigs during the epidemic period is likely due to the fact that grower–finisher pigs have been given PRV vaccines under field conditions. Many pathogens (e.g., PCV2, PRRSV, CSFV, and PPV) may coinfect with PRV (25,26), but all were ruled out in our diagnostic experiment with PCR or by real-time PCR (data not shown). Moreover, the experimental pigs were free of PRV, PCV2, PRRSV, CSFV, and PPV infection. Of course, we did not know whether other pathogens existed in the clinical samples and the experimental animals. The pathogenicity of the new isolate should be studied in depth and compared with that of the traditional PRV strains.
Live vaccines have been widely used to control pseudorabies, and many countries had eradicated the disease (1,4,6). In China the disease had been controlled in some regions for several decades: cases were rarely reported, and only 3% to 5% of serum samples were positive for PR gE antibody during 2005–2010 according to our testing documents. However, the disease broke out suddenly and spread wildly in many pig farms in which the commercial PRV vaccines had recently been used. This suggests that the current vaccines provide poor protection against novel strains. Cross serum neutralization tests could be used for virus typing (20–22). In this study the test results showed that the serum antibodies to the Bartha-K61, Bucharest, and HB-98 vaccine strains had significantly lower neutralizing activity against the PRV ZJ01 strain compared with the vaccine strains. However, the neutralizing titers of the anti-ZJ01 serum against ZJ01 were similar to those against the vaccine strains. The antigenic relatedness between ZJ01 and the PRV vaccine strains of 0.378 to 0.455 suggests that ZJ01 could be defined as a new serum subtype of PRV. In another experiment in pigs, all commercial PRV vaccines could not completely protect against ZJ01 challenge (data not shown), which is consistent with a recent report that Bartha-K61 vaccine provided poor protection against infection with the novel PRV isolate HeN1 (10). Although PRV is usually not differentiated serologically, this seems to be an exception. The cross serum neutralization test should be done with other strains from China to determine whether samples from China are really different or whether the difference occurs only with ZJ01.
Among the many proteins related to the virulence of PRV (2), glycoprotein gE is important for neurovirulence and plays a role in transneuronal spread, although it is not essential for virus replication. Previous studies revealed that a genome deletion that removed gE and glycoprotein I affected the virulence of the vaccine strain Bartha and many virulent strains in vivo (2,3). Our results show that the newly circulating isolates in China have 2 insertions of 1 amino acid (D), in positions 48 and 496, in the gE gene. In addition, these 5 new strains have many strain-specific amino acid mutations compared with other PRV strains, including Becker and Kaplan (2). But we do not know whether the insertions and mutations of these pandemic strains in China are related to virulence and antigenic features. Moreover, the gC gene of PRV is known to vary between strains and is the most efficacious target for virus-neutralizing antibodies (3). In this study, the 5 PRV isolates and other strains previously isolated in China had insertions of 7 amino acids and many amino acid mutations in the gC gene. Of these, the mutation of E43A also occurred in the Bartha, Becker, and Kaplan strains (2). Furthermore, the N-terminal partition of gC was widely used for genotyping. Recent reports have revealed that PRV could be divided into 5 genotypes (A to E) on the basis of partial gC gene sequence (11–14,27), the strains of genotypes A to C being isolated mainly from European countries and North America, and the strains of genotypes D to E being isolated mainly from other countries, such as China and Brazil (14,27). In our study the new isolates could be classified into genotype E (Figure 2) according to the previous description (14,27). However, the molecular mechanisms underlying the high virulence and antigenic variance of these isolates are unknown. It is necessary to explore the full genome sequence of the novel viruses.
In this study a novel, highly virulent PRV with antigenic variance has been isolated and identified, and we have confirmed that highly virulent PRV spread widely in southern China. This poses serious challenges for the control and possible eradication of PRV in China.
Acknowledgments
This work was supported by the China Agricultural Research System Foundation (CARS-36), the Special Fund for Agro-scientific Research in the Public Interest (201203039), and a grant from Jiangsu province (BE2012368, PAPD).
References
- 1.Klupp BG, Hengartner CJ, Mettenleiter TC, Enquist LW. Complete, annotated sequence of the pseudorabies virus genome. J Virol. 2003;78:424–440. doi: 10.1128/JVI.78.1.424-440.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Szpara ML, Tafuri YR, Parsons L, et al. A wide extent of inter-strain diversity in virulent and vaccine strains of alphaherpes-viruses. PLoS Pathog. 2011;7:e1002282. doi: 10.1371/journal.ppat.1002282. Epub 2011 Oct 13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Brittle EE, Reynolds AE, Enquist LW. Two modes of pseudorabies virus neuroinvasion and lethality in mice. J Virol. 2004;78:12951–12963. doi: 10.1128/JVI.78.23.12951-12963.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bouma A. Determination of the effectiveness of pseudorabies marker vaccines in experiments and field trials. Biologicals. 2005;33:241–245. doi: 10.1016/j.biologicals.2005.08.011. [DOI] [PubMed] [Google Scholar]
- 5.Hahn EC, Fadl-Alla B, Lichtensteiger CA. Variation of Aujeszky’s disease viruses in wild swine in USA. Vet Microbiol. 2010;143:45–51. doi: 10.1016/j.vetmic.2010.02.013. Epub 2010 Feb 11. [DOI] [PubMed] [Google Scholar]
- 6.Thawley DG, Morrison RB. Programs for the elimination of pseudorabies virus from large herds of swine. J Am Vet Med Assoc. 1988;193:184–190. [PubMed] [Google Scholar]
- 7.Maresch C, Lange E, Teifke JP, et al. Oral immunization of wild boar and domestic pigs with attenuated live vaccine protects against pseudorabies virus infection. Vet Microbiol. 2012;161:20–25. doi: 10.1016/j.vetmic.2012.07.002. [DOI] [PubMed] [Google Scholar]
- 8.Wu R, Bai C, Sun J, Chang S, Zhang X. Emergence of virulent pseudorabies virus infection in northern China. J Vet Sci. 2013;14:363–365. doi: 10.4142/jvs.2013.14.3.363. Epub 2013 Jun 30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Yu X, Zhou Z, Hu D, et al. Pathogenic pseudorabies virus, China, 2012. Emerg Infect Dis. 2014;20:102–104. doi: 10.3201/eid2001.130531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.An TQ, Peng JM, Tian ZJ, et al. Pseudorabies virus variant in Bartha-K61-vaccinated pigs, China, 2012. Emerg Infect Dis. 2013;19:1749–1755. doi: 10.3201/eid1911.130177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ishikawa K, Tsutsui M, Taquchi K, Saitoh A, Muramats M. Sequence variation of the gC gene among pseudorabies virus strains. Vet Microbiol. 1996;49:267–272. doi: 10.1016/0378-1135(95)00182-4. [DOI] [PubMed] [Google Scholar]
- 12.Goldberg TL, Weigel RM, Hahn EC, Scherba G. Comparative utility of restriction fragment length polymorphism analysis and gene sequencing to the molecular epidemiological investigation of a viral outbreak. Epidemiol Infect. 2001;126:415–424. doi: 10.1017/s0950268801005489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Müller TF, Kluup BG, Freuling C, et al. Characterization of pseudorabies virus of wild boar origin from Europe. Epidemiol Infect. 2010;138:1590–1600. doi: 10.1017/S0950268810000361. [DOI] [PubMed] [Google Scholar]
- 14.Fonseca AA, Jr, Camargos MF, de Oliveira AM, et al. Molecular epidemiology of Brazilian pseudorabies virus isolates. Vet Microbiol. 2010;141:238–245. doi: 10.1016/j.vetmic.2009.09.018. [DOI] [PubMed] [Google Scholar]
- 15.Tamura K, Dudley J, Nei M, Kumar S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol. 2007;24:1596–1599. doi: 10.1093/molbev/msm092. [DOI] [PubMed] [Google Scholar]
- 16.Council for International Organizations of Medical Sciences and the International Council for Laboratory Animal Science. International Guiding Principles for Biomedical Research Involving Animals. 2012. [Last accessed March 16, 2015]. Available from: http://www.cioms.ch/images/stories/CIOMS/IGP2012.pdf.
- 17.Müller TF, Teuffert J, Zellmer R, Conraths FJ. Experimental infection of European wild boars and domestic pigs with pseudorabies viruses with differing virulence. Am J Vet Res. 2001;62:252–258. doi: 10.2460/ajvr.2001.62.252. [DOI] [PubMed] [Google Scholar]
- 18.Ma W, Lager KM, Richt JA, Stoffregen WC, Zhou F, Yoon KJ. Development of real-time polymerase chain reaction assays for rapid detection and differentiation of wild-type pseudorabies and gene-deleted vaccine viruses. J Vet Diagn Invest. 2008;20:440–447. doi: 10.1177/104063870802000405. [DOI] [PubMed] [Google Scholar]
- 19.Ju C, Fan H, Tan Y, et al. Immunogenicity of a recombinant pseudorabies virus expressing ORF1–ORF2 fusion protein of porcine circovirus type 2. Vet Microbiol. 2005;109:179–190. doi: 10.1016/j.vetmic.2005.06.001. [DOI] [PubMed] [Google Scholar]
- 20.Lu CP. Veterinary Microbiology. 3rd ed. Beijing, China: China Agriculture Press; 2001. Serological techniques in immunology; pp. 164–165. [Google Scholar]
- 21.Lee MS, Chen JS. Predicting antigenic variants of influenza A/ H3N2 viruses. Emerg Infect Dis. 2004;10:1385–1390. doi: 10.3201/eid1008.040107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yassine HM, Lee CW, Suarez DL, Saif YM. Genetic and antigenic relatedness of H3 subtype influenza A viruses isolated from avian and mammalian species. Vaccine. 2008;26:966–977. doi: 10.1016/j.vaccine.2007.11.094. Epub 2007 Dec 26. [DOI] [PubMed] [Google Scholar]
- 23.Zhu L, Yi Y, Xu Z, Cheng L, Tang S, Guo W. Growth, physico-chemical properties, and morphogenesis of Chinese wild-type PRV Fa and its gene-deleted mutant strain PRV SA215. Virol J. 2011;8:272. doi: 10.1186/1743-422X-8-272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lin Y, Qigai H, Xiaolan Y, Weicheng B, Huanchun C. The co-administrating of recombinant porcine IL-2 could enhance protective immune responses to PRV inactivated vaccine in pigs. Vaccine. 2005;23:4436–4441. doi: 10.1016/j.vaccine.2005.03.034. [DOI] [PubMed] [Google Scholar]
- 25.Pérez LJ, Perera CL, Frías MT, Núñez JI, Ganges L, de Arce HD. A multiple SYBR Green I-based real-time PCR system for the simultaneous detection of porcine circovirus type 2, porcine parvovirus, pseudorabies virus and Torque teno sus virus 1 and 2 in pigs. J Virol Methods. 2012;179:233–241. doi: 10.1016/j.jviromet.2011.11.009. Epub 2011 Nov 18. [DOI] [PubMed] [Google Scholar]
- 26.Xu XG, Chen GD, Huang Y, et al. Development of multiplex PCR for simultaneous detection of six swine DNA and RNA viruses. J Virol Methods. 2012;183:69–74. doi: 10.1016/j.jviromet.2012.03.034. [DOI] [PubMed] [Google Scholar]
- 27.Fonseca AA, Jr, Camargos MF, Sales ML, Heinemann MB, Leite RC, Reis JK. Pseudorabies virus can be classified into five genotypes using partial sequences of UL44. Braz J Microbiol. 2012;43:1632–1640. doi: 10.1590/S1517-838220120004000048. [DOI] [PMC free article] [PubMed] [Google Scholar]