Abstract
Background
The pstSCAB operon of Corynebacterium glutamicum, which encodes an ABC transport system for uptake of phosphate (Pi), is induced during the Pi starvation response. The two-component regulatory system PhoRS is involved in this response, but partial Pi starvation induction of pstSCAB in a ΔphoRS mutant indicated the involvement of additional regulator(s). Regulation of pstSCAB also involves the global transcriptional regulator GlxR.
Results
DNA affinity chromatography identified the regulator of acetate metabolism RamB as a protein binding to pstS promoter DNA in vitro. Gel mobility shift assays and mutational analysis of the pstS promoter region revealed that RamB binds to two sites localized at positions −74 to −88 and −9 to +2 with respect to the transcriptional start site of pstSCAB. Reporter gene studies supported the in vivo relevance of both binding sites for activation of pstSCAB by RamB. DNA microarray analysis revealed that expression of many Pi starvation genes reached higher levels during the Pi starvation response on minimal medium with glucose as sole carbon source than in Pi starved acetate-grown C. glutamicum cells.
Conclusions
In C. glutamicum, RamB is involved in expression control of pstSCAB operon. Thus, transcriptional regulation of pstSCAB is complex involving activation by the phosphate-responsive two-component regulatory system PhoSR and the regulators of carbon metabolism GlxR and RamB.
Keywords: Corynebacterium glutamicum, Phosphate starvation, pstS, RamB, Phosphorus metabolism, Carbon metabolism, Acetate metabolism, PhoR, GlxR
Background
Phosphorus is an essential component of all cells. In bacteria, phosphorus is typically assimilated as inorganic orthophosphate (Pi) via the reactions of the energy and carbon metabolism, thus, the phosphorus metabolism is closely intertwined with the energy and the central carbon metabolism. An optimal energy and carbon metabolism is possible only with sufficient phosphorus supply. As precursor metabolites for the biosynthesis of amino acids are derived from central carbon metabolism, the interplay of phosphorus and carbon metabolism is of particular interest in amino acid producing Corynebacterium glutamicum strains.
Pi is taken up into the cell by specific transport systems. When Pi becomes scarce, many bacteria induce the synthesis of proteins to use limiting concentrations of Pi more efficiently and to make alternative sources of phosphorus accessible. The regulation of the Pi starvation response of Escherichia coli [1] and Bacillus subtilis [2] has been studied in detail. In E. coli, the two component regulatory system PhoR-PhoB is responsible for the induction of the Pi starvation genes. Under Pi starvation conditions, the histidine kinase PhoR phosphorylates the response regulator PhoB and phosphorylated PhoB induces the transcription of at least 38 genes, the so-called PhoB regulon. Among these genes are the phoBR operon encoding two component regulatory system, the pstSCAB-phoU operon encoding an ABC transporter for high-affinity Pi uptake and an regulatory protein, and the ugpBAECQ operon encoding an sn-glycerol 3-phosphate ABC uptake system and glycerophosphoryl diester phosphodiesterase. The PhoB regulon in E. coli also comprises 21 genes important for uptake and degradation of phosphonates, e.g. the phnCDEFGHIJKLMNOP operon. In B. subtilis, the Pi starvation response is dependent on the two component system PhoP-PhoR for activation of Pho regulon, Spo0A for termination of the Pi starvation response and subsequent initiation of sporulation, ResDE for the full induction of the Pho regulon genes and the regulator AbrB. In addition, Pi starvation in B. subtilis leads to the induction of genes of the general stress response, mediated by σB and σM [3–6]. Under Pi starvation conditions, B. subtilis replaces teichoic acids in the cell-wall with the non-phosphate containing teichuronic acids due to repression of the teichoic acid biosynthesis operons tagAB and tagDEF and derepression of the teichuronic acid biosynthesis operon tuaABCDEFGH [7, 8].
C. glutamicum was isolated in 1957 as an L-glutamate excreting bacterium [9] and is used for the large scale biotechnological production of L-glutamate and L-lysine [10, 11]. This bacterium has been engineered for the production of other amino acids such as L-serine [12], L-isoleucine [13], L-valine [14, 15] or L-proline [16]. It has been also successfully engineered to produce derivatives or precursors of amino acids such as 1,4-diaminobutane [17, 18] 1,5-diaminopentane [19], 2-ketoisovalerate [20] and 2-ketoisocaproate [21, 22].
In C. glutamicum, phosphorus constitutes 1.5 % to 2.1 % of the cell dry weight [23]. Under Pi sufficient conditions, C. glutamicum accumulates cytoplasmic and granular polyphosphate [24–26]. Polyphosphate is synthesized by class II polyphosphate kinases [27]. For utilization, it is hydrolysed by exopolyphosphatases [28] and replaces ATP in the reactions of NAD kinase PpnK [29] and glucokinase PpgK [30]. Although intracellular polyphosphate was shown to serve as reservoir of phosphorus [27], expression of a number of genes involved in phosphorus metabolism is induced within 1 h after a shift from Pi sufficient to Pi limiting conditions [23, 31]. As determined by global gene expression analysis using whole-genome C. glutamicum DNA microarrays [31], the Pi starvation stimulon comprises among others pstSCAB encoding an ABC transporter for high affinity Pi uptake, ugpABCE encoding an sn-glycerol 3-phosphate ABC uptake system, ushA encoding a secreted enzyme with UDP sugar hydrolase and 5’nucleotidase activity [32], and the phoRS operon encoding for the two component system involved in the Pi starvation response of C. glutamicum [33]. Purified phosphorylated PhoR was shown to bind to the promoters of Pi starvation-inducible genes at sites containing a loosely conserved 8-bp direct repeat [34]. Transcriptome analyses of C. glutamicum WT and the deletion mutant ΔphoRS revealed that the known Pi starvation-inducible genes were not induced within 1 h after a shift from Pi excess to Pi limitation, with the exception of the pstSCAB operon, which was still partially induced in the deletion mutant [33]. This indicated that at least one additional regulator besides PhoR is involved in Pi-dependent regulation of the pstSCAB operon in C. glutamicum. GlxR, a global cAMP-dependent transcriptional regulator [35–37], was shown to bind to the pstS promoter −133 bps to −117 bps upstream of the transcriptional start site and activates the pstSCAB operon under phosphate limiting conditions in a carbon source dependent manner [38]. When glxR was overexpressed, growth was enhanced under phosphate limiting conditions on glucose as carbon source, but not on acetate [38]. Moreover, a metabolome analysis of C. glutamicum grown on acetate or glucose revealed a link between Pi limitation and accumulation of glycogen and maltose [39]. However, mutation of GlxR binding site in the pstS promoter sequence did not abolish the expression of the reporter gene. This indicated the existence of other factor(s) involved in regulation of pstS operon under Pi starvation conditions. The aim of this study was to characterize adaptation of C. glutamicum to Pi starvation in the absence of PhoS-PhoR and to identify additional regulator(s) of pstSCAB.
Results
Growth of C. glutamicum WT and ΔphoRS on different phosphorus sources and under Pi limiting conditions
To characterize the long-term response of C. glutamicum to Pi limitation and growth on alternative phosphorus sources, comparative growth experiments were performed with C. glutamicum WT and with the deletion mutant ΔphoRS, which lacks the two-component regulatory system PhoRS (Table 1) [33]. C. glutamicum WT and ΔphoRS were pre-cultured for 24 h in CGXII glucose medium without Pi in order to exhaust the intercellular phosphorus storages [25, 31] and inoculated into CGXII glucose medium with either a limiting Pi concentration of 0.065 mM or with 1 mM of the alternative phosphorus sources of adenosine 5’-monophosphate (5’AMP), L-α-glycerophosphate or UDP-glucose.
Table 1.
Strains and plasmids used in this study
| Strain or plasmid | Relevant characteristic | Reference |
|---|---|---|
| C. glutamicum | ||
| WT | wild type strain ATCC 13032 | [9] |
| ΔphoRS | deletion of the phoRS operon encoding the two component system PhoRS | [33] |
| ΔramB | Deletion of ramB encoding regulator of acetate metabolism B | [41] |
| E. coli | ||
| BL21(DE3) | ompT hsdS B(rB−mB−) gal dcm (DE3) | [64] |
| DH5α | F− thi-1 endA1 hsdr17(r−, m−) supE44 ΔblacU169 (ф80lacZΔM15) recA1 gyrA96 relA1 | [65] |
| Plasmids | ||
| pGEM-T | cloning vector | Promega, WI, USA |
| pET2 | promoter-probe vector | [54] |
| pET2-RF0 | pET2 with pstSCAB promoter fragment RF0 | This study |
| pET2-R0F0 | pET2 with pstSCAB promoterfragment R0F0 | This study |
| pET2-R1F0 | pET2 with pstSCAB promoter fragment R1F0 | This study |
| pET2-R2F0 | pET2 with pstSCAB promoter fragment R2F0 | This study |
| pET2-R3F0 | pET2 with pstSCAB promoter fragment R3F0 | This study |
| pET2-R0F1 | pET2 with pstSCAB promoter fragment R0F1 | This study |
| pET2-R0F2 | pET2 with pstSCAB promoter fragment R0F2 | This study |
| pET2-R0F3 | pET2 with pstSCAB promoter fragment R0F3 | This study |
| pET2-RcFc | pET2 with pstSCAB promoter fragment RcFc | This study |
| pET2-RcFm | pET2 with pstSCAB promoter fragment RcFm | This study |
| pET2-RmFc | pET2 with pstSCAB promoter fragment RmFc | This study |
| pET2-RmFm | pET2 with pstSCAB promoter fragment RmFm | This study |
| pET29-ramB-his | KanR; pET29-Histag derivative for over production of RamB with a C-terminal histidine tag | [41] |
With 0.065 mM Pi, which is below the Pi concentration of 0.1 mM that supported growth of C. glutamicum with a half-maximal growth rate [31], C. glutamicum WT showed a doubling time of 0.14 h−1 and formed 0.5 g DW l−1 biomass whereas the deletion mutant ΔphoRS showed a growth defect under Pi limiting conditions as expected from previous results (Table 2) [33].
Table 2.
Growth of C. glutamicum WT and ΔphoRS on different phosphorus sources
| Phosphorus source | Strain | Biomass formed [g/l] | μ [h−1] | Duration of lag phase [h] | UDP-glucose hydrolase activity in supernatants [nmol min−1 ml−1] a |
|---|---|---|---|---|---|
| Low Pi, 0.065 mMb | WT | 2 | 0.14 | 0 | 27 |
| ΔphoRS | 1 | 0.07 | 6 | 39 | |
| Glycerol-3-phosphate, 1 mM | WT | 11 | 0.16 | 0 | 9 |
| ΔphoRS | 9 | 0.11 | 9 | 13 | |
| 5’AMP, 1 mM | WT | 9 | 0.08 | 11 | 6 |
| ΔphoRS | 7 | 0.08 | 34 | 12 | |
| UDP-glucose, 1 mM | WT | 9 | 0.06 | 39 | 6 |
| ΔphoRS | 8 | 0.09 | 63 | 3 |
aUDP-glucose hydrolase activity was measured after 180 h of cultivation. No UDP-glucose hydrolase activity was detectable (<1 nmol min−1 ml−1) in supernatants of cells grown under Pi sufficient conditions (13 mM)
bThis concentration is below the Pi concentration of 0.1 mM which supports the half-maximal growth rate in C. glutamicum [31]
C. glutamicum ΔphoRS could utilize the alternative phosphorus sources L-α-glycerophosphate, 5’AMP and UDP-glucose, however, it showed longer lag phases, lower growth rates and lower biomass yields than C. glutamicum WT (Table 2). As growth of C. glutamicum on 5'-AMP and UDP-glucose requires the Pi starvation inducible gene ushA, which encodes a secreted enzyme with UDP-glucose hydrolase and 5'-nucleotidase activity [32], UDP-glucose hydrolase activity of supernatants of these cultures were measured. While UDP-glucose hydrolase activity could not be detected under Pi sufficient conditions (data not shown), supernatants of C. glutamicum WT and ΔphoRS grown with L-α-glycerophosphate, 5’AMP and UDP-Glucose as sole phosphorus sources showed UDP-glucose hydrolase activity (Table 2). Taken together, PhoRS is not essential for growth with these organophosphates and other regulators apparently allow C. glutamicum to induce ushA and possibly other genes necessary for the Pi starvation response in the absence of PhoRS.
Deletion analysis of the pstS promoter
To identify cis-regulatory sequences of the pstS promoter for the PhoR-dependent and PhoR-independent control, a deletion analysis of the pstS promoter region was performed using different oligonucleotides (Table 3). The pstS promoter fragment (RF0) and the promoter fragments either lacking the 5' region (R0F0, R1F0, and R2F0) or the 3' region (R0F1, R0F2, and R0F3) were fused to the promoter-less chloramphenicol acetyl transferase (CAT) gene (Fig. 1). The resulting plasmids pET2-RF0, pET2-R0F0, pET2-R1F0, pET2-R2F0, pET2-R0F1, pET2-R0F2 and pET2-R0F3 were transferred into C. glutamicum WT and ΔphoRS. Expression of these fusions was assayed before and 90 min after a shift from Pi rich to Pi lacking medium. The fusion with fragment R3F0 was not expressed as it lacked the previously determined transcriptional start site and the −10 and −35 binding regions of the RNA polymerase (Fig. 2a, b) [33]. All other fusions were expressed and showed Pi starvation-inducible expression both in C. glutamicum WT and ΔphoRS (Fig. 2a, b).
Table 3.
Oligonucleotides used in this study
| Oligonucleotide Sequence (5’ → 3’) | |
|---|---|
| pstsRforward | CCCCTCGAGTAAAAAAGAGACTTGCTAAAAACCT (XhoI) |
| pstsR0forward | CCCCTCGAGTAAGAATCGGTGATTTTCGTTCC (XhoI) |
| pstsR1forward | CCCCTCGAGAGAGTCTCCAAATGTTACGAGTGAA (XhoI) |
| pstsR2forward | CCCCTCGAGCCTGAGTTAGTCATTTCAAGGTCTTA (XhoI) |
| pstsR3forward | CCCCTCGAGGCCCGCCTACAGGATCTGCTCA (XhoI) |
| pstsF0reverse | CGTCTAGATGCGGACTGCTGGGAAGATG (XbaI) |
| pstsF1reverse | CGTCTAGACCTCAATGGATGCAGCATCGGAAG (XbaI) |
| pstsF2reverse | CGTCTAGATCAGACTCATTGGAGTCGGAGCAA (XbaI) |
| pstsF3reverse | CGTCTAGAGTTCACGGGGAAGCCTTTCCGG (XbaI) |
| pstsF4reverse | CGTCTAGATAAGACCTTGAAATGACTAACTCAGG (XbaI) |
| pstsFc_reverse | CGGTTTCCCTCCGGATTGCTCACGACTTAAAAACCTA |
| pstsFm_reverse | CGGTTTCCCTCCGGATTGCGCGCGGAGTAAAAACCTA |
| pstsRc_forward | CCCGATGTGGGTAGTGGCAGAATTTGCCGAACGAT |
| pstsRm_forward | CCCGATGTGGGTAGTGGCAGAAGAGGCCGAACGAT |
| pstsF0biotin | Biotin-TGCGGACTGCTGGGAAGATGCAC |
*In some cases oligonucleotides were designed to introduce recognition sites for restriction endonucleases (recognition sites in italics)
Fig. 1.
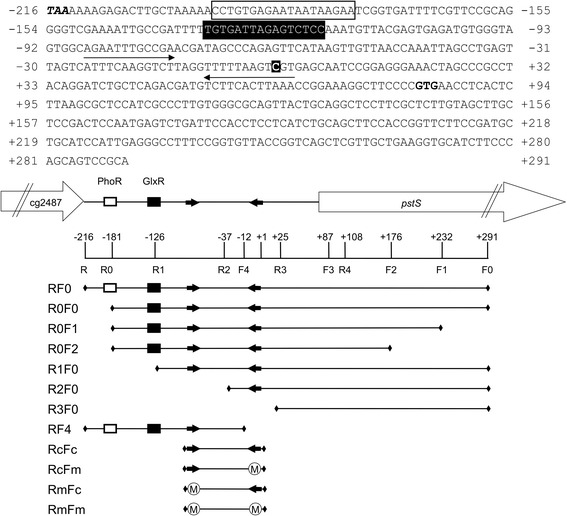
Overview of the pstS promoter region and the fragments used in this study. Several DNA fragments were used to analyze RamB binding to the pstS promoter in the gel mobility shift assays and the reporter gene assay. The PhoR binding site (open box), GlxR binding site (black box) and two putative RamB binding sites (black arrows) are indicated in the sequence and diagrams. The stop codon of cg2487 (TAA with bold italic), the transcriptional start site of pstS (C in a black box), and the pstS start codon (GTG in bold) are indicated in the sequence. The number in the diagram indicates the respective position of nucleotide from the transcription start site (+1) of pstS and the coverage of each fragment is indicated. A mutation introduced into a RamB binding site is indicated as circled M in the diagram
Fig. 2.
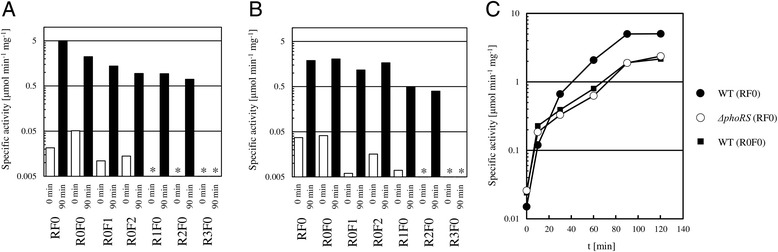
Expression of reporter gene with various promoter fragments in C. glutamicum WT and ΔphoRS. Expression levels of the fusions in C. glutamicum WT (a) and in C. glutamicum ΔphoRS (b). Expression levels of the CAT gene fusions were measured before (open bar) and 90 min (filled bar) after the shift from Pi sufficient to Pi limiting conditions. RF0 to R3F0 indicates the fragment used in the experiment. Expressions are given as specific activity of chloramphenicol acetyltransferase. (*, the specific activity < 0.005) (c) Expression levels of the fusions in a time dependent manner. Expression of fusions was measured after a medium shift to medium lacking Pi. C. glutamicum WT (filled) or ΔphoRS (open) carrying the promoter fragment RF0 (circle) or R0F0 (square) was used
Expression of the reporter gene fused to the full-length pstS promoter in C. glutamicum WT (pET2-RF0) was about threefold higher than in C. glutamicum ΔphoRS (pET2-RF0), while expression of the other fusions did not differ much between WT and ΔphoRS (Fig. 2a, b). This indicated that fragment R0F0 lacked a cis regulatory sequence required for activation by PhoRS under Pi starvation conditions and it is consistent with the finding of a PhoRS binding site in this region [34]. Also the fusions in pET2-R1F0 and pET2-R2F0, which lack the previously determined GlxR binding site, were expressed in C. glutamicum WT as well as in ΔphoRS upon Pi starvation.
Pi starvation induction of the pstSCAB operon is stronger and faster than that of other Pi starvation inducible genes of C. glutamicum [31] and its induction is partially retained in the absence of PhoRS [33]. Therefore, the time dependent expression from pET2-RF0 and pET2-R0F0 was analyzed in C. glutamicum WT and ΔphoRS under Pi starvation. After a shift from Pi-sufficient to Pi-limiting conditions, expression of the pstS promoter fusion in pET2-RF0 was induced in C. glutamicum WT and ΔphoRS before 60 min (Fig. 2c). However, Pi starvation induction of the pstS promoter in the phoRS mutant followed slower kinetics and reached a two to three fold lower level than in C. glutamicum WT. On the other hand, induction was very similar between the full-length pstS promoter (pET2-RF0) in the phoRS mutant and the pstS promoter lacking PhoR binding site (pET2-R0F0) in the wild type. Thus, expression control of the pstS promoter by PhoRS in vivo required the cognate PhoR binding site, which is present in the full-length promoter fragment (RF0), but absent from the 35 nucleotides shorter fragment (R0F0). Furthermore, the fragment R0F0 apparently contains all cis regulatory sequences required for Pi starvation induction independent of PhoRS. Moreover, the fusions lacking the PhoR and the GlxR binding sites (pET2-R1F0, pET2-R2F0) were still induced under Pi starvation conditions. Thus, besides PhoRS, which is required for maximal Pi starvation induction of pstSCAB, and GlxR, (an) aditional unknown regulator(s) are involved in control of pstSCAB expression during adaptation of C. glutamicum to Pi limitation.
Identification of RamB as a protein binding to the pstS promoter
In order to identify (a) regulatory protein(s) binding to the pstS promoter region, we coupled the biotinylated pstS promoter fragment R0F0 to Dynabeads® streptavidin for DNA affinity purification experiments. DNA affinity chromatography was performed with crude extracts from C. glutamicum WT (data not shown) and deletion mutant ΔphoRS in CGXII minimal medium with 4 % (w/v) glucose (Fig. 3a). In these experiments, a number of proteins bound to the promoter DNA fragment. By tryptic finger print analysis using MALDI-TOF mass spectrometry, some of these proteins could be identified. Among proteins binding the promoter DNA in a sequence-independent manner (e.g. subunits of RNA polymerase or topoisomerase) the transcriptional regulator RamB was identified (Fig. 3a). The regulator of acetate metabolism RamB is known to repress transcription of the pta-ack operon, the aceA and aceB genes encoding enzymes for acetate activation and of the glyoxylate cycle [40, 41]. Therefore, the DNA affinity chromatography experiments were repeated using crude extracts of C. glutamicum WT cultivated on acetate minimal medium under Pi starvation conditions. As a result, GlxR and RamB were found to bind to the full-length pstS promoter DNA (data not shown). Binding of RamB to the pstS promoter DNA suggested its involvement in direct control of the pstSCAB operon.
Fig. 3.
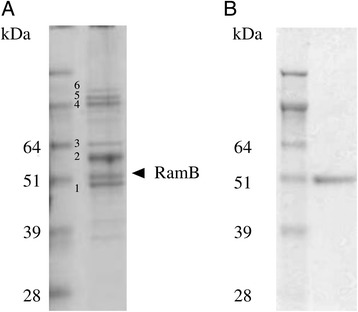
SDS-PAGE images of DNA affinity chromatography experiment and purified RamB protein. (a) Proteins eluted from a DNA affinity chromatography experiment using the pstS promoter. For the DNA affinity chromatography experiment, the pstS promoter fragment R0F0 was used as a probe and incubated with cell extracts of C. glutamicum ΔphoRS grown under Pi sufficient conditions in minimal medium with 4 % (w/v) glucose (Right lane). 1: DNA-polymerase I, 2: Acetyl/propionyl-CoA carboxylase subunit, 3: Acetyl/propionyl-CoA carboxylase subunit, 4: DNA gyrase, 5: DNA-directed RNA polymerase β-subunit, 6: DNA-directed RNA polymerase β’-subunit. Left lane: protein standard Seeblue II prestained Standard (Invitrogen, Karlsruhe) (b) Purified His-tagged RamB. His-tagged RamB was over produced in E. coli, purified and separated on a 10 % (w/v) SDS-polyacrylamide gel. Gel was stained with Coomassie Blue. Left lane: protein standard Seeblue II prestained Standard (Invitrogen, Karlsruhe), Right lane: purified His-tagged RamB obtained after imidazol elution from a nickel-chelate affinity column
Purified RamB binds to two binding motifs in the pstS promoter in vitro
RamB binding sites (AA/GAACTTTGCAAA) are present upstream of many genes encoding enzymes of the central carbon metabolism that belong to the acetate stimulon [41]. However, a RamB binding site within the pstS promoter region has not yet been reported. Inspection of the pstS promoter DNA suggested the occurrence of two partially conserved RamB binding sites, motif A and motif B: AGAA-TTTGCCGA (−74 to −88) and the reverse complement of ACGACTT-AAAAA (+2 to −9).
In order to test whether RamB directly binds to the pstS promoter DNA, band shift assays with purified RamB were performed. RamB containing a C-terminal His-Tag was overproduced in E. coli BL21 (DE3) and purified to apparent homogeneity by affinity chromatography (Fig. 3b). Gel shift assays showed that RamB bound with a high affinity to the full-length pstS promoter, but not to the negative control fragment cg0527 (Fig. 4a). Gel shift assays with the different fragments of the pstS promoter lacking the 5' region (RF0, R0F1, R1F0, R2F0, R3F0) showed binding of RamB to respective DNA fragments except for the fragment R3F0, which lacked both of the predicted RamB binding sites (Fig. 4a). RamB bound weaker to the fragment R2F0, which contains one of the predicted binding site (motif A), than to other fragments which contain both of the predicted binding sites (RF0, R0F0, R1F0). Similarly, the affinity of RamB to fragment RF4, which contains only one of the predicted RamB binding site (motif B), was weaker than that to the full-length pstS promoter fragment (RF0) (Fig. 4b). These results suggested the presence of two RamB binding sites in the full-length pstS promoter fragment.
Fig. 4.
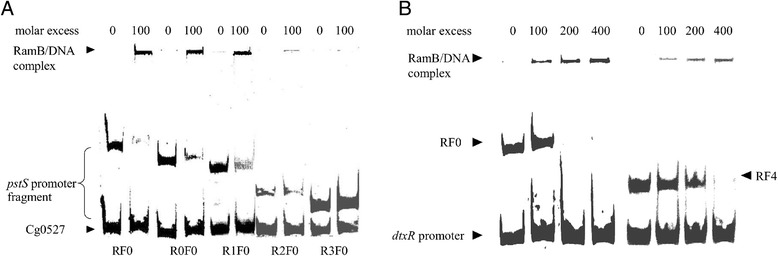
Binding of RamB to various pstS promoter fragments. (a) Gel shift assay with RamB and the fragment of the pstS promoter lacking 5' region. RamB protein (0, 100 fold molar excess) was incubated with the full-length pstS promoter (RF0, 507 bp, 15 nM) or the different fragments of the pstS promoter lacking 5' region (R0F0, R1F0, R2F0, R3F0, final concentrations 61 nM – 15 nM) and applied for native polyacrylamide gel electrophoresis. A 185 bp promoter fragment of cg0527 served as a negative control. (b) Gel shift assay with RamB and the fragment of the pstS promoter lacking 3' region. RamB protein (0, 100, 200, 400-fold molar excess) was incubated with the pstS promoter (RF0, 507 bp, 15 nM) or fragment of the pstS promoter lacking 3' region (RF4, 230 bp, 33 nM) and applied for native polyacrylamide gel electrophoresis. A 122 bp promoter fragment of dtxR served as a negative control
A mutational analysis was performed to determine whether both of the partially conserved RamB binding motifs are required for interaction of RamB with the pstS promoter. Mutations of RamB binding motif A (AGAAGAGGCCGA instead of AGAATTTGCCGA in fragment RmFc), RamB binding motif B (GCGGGAGTAAAAA instead of TCAGACTTAAAAA in fragment RcFm), or of both RamB binding motifs (in fragment RmFm) were introduced into the pstS promoter fragment RcFc, which contained both putative binding sites within a 124 bp region (Fig. 1). RamB did not bind to the fragment RmFm containing both mutated binding sites. RamB interacted stronger with non-mutated fragment RcFc than with the fragments RcFm and RmFc, each only containing one intact binding site (Fig. 5). Thus, both binding sites contribute to binding of RamB to the pstS promoter in vitro.
Fig. 5.
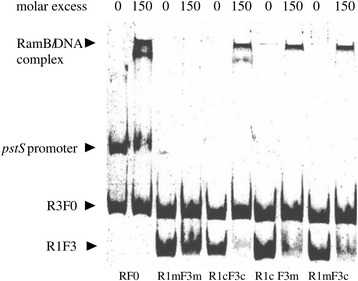
Binding of RamB to various pstS promoter fragments. RamB protein (0, 150 fold molar excess) was incubated with the full-length pstS promoter fragment (RF0, 5 nM, 507-bp) or the partial length pstS promoter fragments (RmFm, RcFm, RmFc, RcFc, 61 nM, 124 bp) and applied for native polyacrylamide gel electrophoresis. A 267 bp fragment of R3F0, which lacked both of RamB binding sites, served as a negative control
Role of RamB sites for regulation of the pstS promoter in vivo
In order to determine the role of RamB for Pi starvation induction of the pstS promoter in vivo, expression of pstS promoter fusion to the promoter-less CAT reporter gene was analyzed in C. glutamicum WT on different carbon sources after a shift from Pi-sufficient to Pi starvation conditions. These medium shift experiments were performed with minimal medium containing either 4 % (w/v) glucose or 2 % (w/v) potassium acetate as sole carbon source. Expression of the pstS promoter fusion R0F0 after a shift from Pi-sufficient to Pi starvation conditions was higher on glucose than on acetate (2.10 compared to 0.22 μmol min−1 mg−1, Table 4). Pi starvation induced expression of the fusion with the shorter RcFc promoter fragment, which lacks the PhoR and GlxR binding sites, and induction was six fold higher onglucose than on acetate (0.61 as compared to 0.10 μmol min−1 mg−1, Table 4). When mutations were introduced in only one of RamB binding sites (fragments RmFc and RcFm), expression was reduced both on glucose and acetate. The RmFm fusion carrying mutations in both RamB binding sites showed almost no activity after medium shift on both carbon source (Table 4).
Table 4.
Expression of various pstS promoter fragment cat fusions in C. glutamicum WT
| Promoter fragment in transcriptional fusion | Carbon source | sp. act. of chloramphenicol acetyltransferase [μmol min−1 mg−1]a | |
|---|---|---|---|
| 0 min | 90 minb | ||
| R0F0 | Glucose | 0.02 | 2.10 |
| Acetate | 0.01 | 0.22 | |
| RcFc | Glucose | <0.005 | 0.61 |
| Acetate | <0.005 | 0.10 | |
| RcFm | Glucose | <0.005 | 0.12 |
| Acetate | <0.005 | 0.01 | |
| RmFc | Glucose | <0.005 | 0.10 |
| Acetate | <0.005 | 0.01 | |
| RmFm | Glucose | <0.005 | 0.02 |
| Acetate | <0.005 | <0.005 | |
aAt least three determinations of two independent cultivations were performed. Average values are given with experimental imprecision < 20 %
bThe specific activity of chloramphenicol acetyltransferase was measured 0 and 90 min after a shift from Pi sufficient to Pi limiting conditions
In addition, expression of the pstS promoter fusion R0F0 was assayed in the deletion mutant ΔramB growing in 4 % (w/v) glucose as a carbon source before and after Pi starvation induction. Before Pi starvation, expression of the pstS promoter fusion was low, both in WT and in ΔramB (0.02 and 0.02 μmol min−1 mg−1, respectively), while Pi starvation induction was higher in WT as compared to ΔramB (2.10 and 1.23 μmol min−1 mg−1, respectively, data not shown). Taken together, RamB as well as both RamB binding sites are important for Pi starvation induction of the pstS promoter in C. glutamicum in vivo.
Comparison of Pi starvation inducible gene expression on glucose and acetate minimal medium
As Pi starvation induction of the pstS promoter differed with respect to the carbon source, DNA microarray analysis was performed to compare the gene expression profile on minimal medium containing either glucose or acetate during the Pi starvation response. C. glutamicum cells growing exponentially on glucose or acetate minimal medium with 13 mM Pi were shifted to minimal medium containing either glucose or acetate but lacking Pi. RNA was prepared 90 min after the medium shift. As expected for acetate dependent regulation in C. glutamicum [41], the DNA microarray analysis revealed two to 100 fold higher mRNA levels for genes belonging to the acetate stimulon on acetate than on glucose: pta encoding phosphotransacetylase, aceA and aceB encoding isocitrate lyase and malate synthase, pck encoding gluconeogenetic PEP carboxykinase, acn encoding aconiatase and gltA encoding citrate synthase (Table 5). Expression of ramB was about four fold higher on glucose than on acetate due to autoregulation by RamB and control by RamA [40]. Expression of genes of the pstSCAB operon was higher on glucose than on acetate in response to Pi starvation, which is consistent with the pstS promoter fusion experiments in this study (Table 5). In addition, expression of other genes belonging to the Pi starvation stimulon reached higher levels on glucose than on acetate: ushA encoding 5’-nucleotidase, psiB encoding a putative alkaline phosphatase, phoH1 encoding a putative ATPase, cg1224 encoding a PhnB-like protein, pctC of the pctABCD operon encoding an ABC transport system and ugpA and ugpE of the ugpEABC operon encoding an glycerol-3-phosphate uptake system (Table 5). Unlike other genes of the Pi starvation stimulon, expression of phoS and phoR encoding phosphate sensor kinase and its response regulator was lower on glucose than on acetate during Pi starvation (Table 5).
Table 5.
Genes differentially expressed in either glucose or acetate minimal medium cultures of C. glutamicum WT after a shift from Pi-sufficient to Pi-limiting conditions
| Gene identifier | Annotationa | Relative mRNA levelb glucose/acetate |
|---|---|---|
| cg2560 | aceA, isocitrate lyase | 0.01 |
| cg2559 | aceB, malate synthase | 0.05 |
| cg3169 | pck, phosphoenolpyruvate carboxykinase | 0.22 |
| cg3048 | pta, phosphoacetyltransferase | 0.27 |
| cg2887 | phoR, phosphate response regulator | 0.33 |
| cg1737 | acn, aconitase | 0.34 |
| cg0949 | gltA, citrate synthase | 0.46 |
| cg2406 | ctaE, cytochrome aa 3 oxidase, subunit | 0.47 |
| cg2888 | phoS, phosphate sensor kinase | 0.47 |
| cg2843 | pstB, Pi ABC transporter, ATPase | 2.0 |
| cg1569 | ugpE, glycerol 3-phosphate ABC transporter, permease | 2.1 |
| cg1224 | phnB1, PhnB-like protein | 2.3 |
| cg0397 | ushA, UDP sugar hydrolase/5’-nucleotidase | 2.4 |
| cg0444 | ramB, regulator of acetate metabolism B | 3.5 |
| cg1647 | psiB, putative alkaline phosphatase | 4.2 |
| cg3393 | phoC, putative secreted phosphoesterase | 4.3 |
| cg0085 | phoH1, ATPase | 5.2 |
| cg1650 | pctC, ABC transporter, permease | 5.2 |
| cg2868 | nucH, putative nuclease | 5.4 |
| cg0812 | accD1, acetyl-CoA carboxylase subunit | 11.3 |
| cg1568 | ugpA, glycerol 3-phosphate ABC transporter, permease | 31.2 |
aGene identifiers and annotations are given according to BX927147
bThe mRNA levels were derived from two independent cultivations
Discussion
Here we have shown that RamB is involved in expression control of the pstSCAB operon during the Pi starvation response of C. glutamicum. The two component regulatory system PhoR-PhoS is neither essential for Pi starvation induction of pstSCAB nor for growth on media with the organophosphates glycerol-3-phosphate, 5’-AMP and UDP-glucose as sole phosphorus source. However, PhoR-PhoS ensures rapid and maximal Pi starvation induction of pstSCAB. The regulator of acetate metabolism RamB was shown to bind to two binding sites in the pstS promoter fragment in vitro and both of two binding sites were shown to influence the activity of the pstS promoter fragment in vivo by reporter gene assay. Pi starvation induction of the pstS promoter fragment reached 10 fold higher levels on glucose minimal medium than on acetate minimal medium. Microarray experiments showed that Pi starvation induction of ramB and the Pi starvation stimulon including pstSCAB reached higher RNA levels with glucose as carbon source than with acetate as carbon source. These findings support and extend a regulatory link between phosphorus and carbon metabolism in C. glutamicum [38, 39].
The regulator of acetate metabolism RamB represses transcription of the pta-ack operon and the aceA and aceB genes, which encode enzymes for acetate activation and for the glyoxylate cycle [41]. Deletion and mutation analysis of the promoter regions of these genes allowed identifying conserved 13-bp motifs as RamB binding sites [41]. A bioinformatics analysis of the genome sequence revealed that variants of the cis-regulatory motif for RamB binding were identified upstream of aceA, aceB, pta-ack and also occur in the promoter regions of 28 other genes, 11 of which were differentially expressed in acetate- and glucose-grown C. glutamicum cells. These genes code for enzymes of e.g. glucose uptake, glycolysis, glucoeneogenesis, anaplerosis and the tricarboxylic acid cycle [41]. While this bioinformatic analysis searched for variants of the RamB binding site (AA/GAACTTTGCAAA or its complement) with maximal mismatches of two nucleotides [41], the newly identified RamB binding sites in the pstSCAB promoter were not recognized previously as they contain 3 (AGAA-TTTGCCGA) and 5 mismatches (complement of ACGACTT-AAAAA)), respectively. Mutational analysis of the RamB binding sites in the pstS promoter fragment showed that RamB binds to both of the newly identified RamB binding sites in vitro and that both binding sites are relevant for regulation of the pstS promoter under Pi limiting condition in vivo. Thus, RamB appears to activate pstSCAB expression under Pi limiting conditions. While RamB mostly represses its target genes, RamB was shown to activate aceE encoding the E1p subunit of the pyruvate dehydrogenase complex [42].
GlxR also links regulation of carbon and phosphorus metabolism in C. glutamicum. GlxR is known to regulate more than 100 genes and is one of the global hubs within the C. glutamicum gene-regulatory network [35]. GlxR was shown to bind to the pstS promoter in a cAMP-dependent manner in vitro [38] and the interaction of GlxR with pstS promoter DNA was higher on glucose than on acetate as carbon source in C. glutamicum [38, 43]. In this study, expression of the reporter gene fusion with the full length pstS promoter (RF0) was higher under Pi starvation conditions than expression of the fusion lacking the PhoR binding site (R0F0) and even higher than expression of the fusion lacking both the PhoR and GlxR binding sites (R1F0) (Fig. 2). Thus, the three transcriptional regulators PhoR, GlxR and RamB synergistically activate expression of the pstS operon under Pi starvation conditions.
GlxR, RamA and RamB also regulate transcription of their genes, e.g. GlxR activates ramA and represses ramB [35], RamA activates ramB [40] and GlxR, RamA and RamB show negative autoregulation [44–46]. Moreover, a number of target genes of RamB and RamA are also regulated by GlxR, e,g, adhA and ald encoding alcohol dehydrogenase and acetaldehyde dehydrogenase [41] as well as gltA encoding citrate synthase [44] are repressed by both GlxR and RamB, but activated by RamA, rpf2 encoding resuscitation promoting factor 2 is activated by RamA and GlxR, but repressed by RamB [45]. Negative autoregulation of RamB, carbon source-dependent activation of ramB by RamA [40] and cAMP-dependent activation of ramB by GlxR fine-tune regulation of carbon metabolism and also serve to integrate regulation of carbon and phosphorus metabolism in C. glutamicum.
Regulation of pstSCAB in C. glutamicum is complex, involves at least three transcriptional regulators: PhoR [33], GlxR [38] and RamB (this study) and differs from regulation of the pstS promoter in M. tuberculosis, E. coli and B. subtilis. Notably, in the related actinomycete Mycobacterium tuberculosis transcription of the pst operon is not induced upon Pi starvation. Since M. tuberculosis can replicate in the phagosomes of macrophages, an acidic and Pi poor environment, constitutive expression of pst may be a consequence of this intracellular life style [47]. In E. coli, the pstS promoter is regulated by integration host factor (IHF) and PhoB [48, 49], whereas this promoter is regulated in B. subtilis by PhoP [50].
Conclusions
In C. glutamicum, RamB is involved in expression control of the pstSCAB operon and two binding sites are relevant for activation by RamB in vitro. These finding support the notion that phosphorus and carbon metabolism in C. glutamicum are regulated in dependence of each other. Transcriptional regulation of pstSCAB is complex involving activation by the phosphate-responsive two-component regulatory system PhoSR and the regulators of carbon metabolism GlxR and RamB.
Methods
Bacterial strains, media, and growth conditions
Bacterial strains and plasmids used in this work are listed in Table 1. E. coli DH5α (Invitrogen) was used as host during the construction of recombinant plasmids and grown aerobically at 37 °C on a rotary shaker (120 rpm) in Luria-Bertani (LB) medium [51]. E. coli BL21 (DE3) was used for overproduction of RamB protein and grown aerobically at 37 °C on a rotary shaker (120 rpm) in LB medium. When appropriate, ampicillin was added at a concentration of 100 μg/ml. C. glutamicum wild-type strain ATCC 13032 (WT) and the ΔphoRS deletion mutant [33] were grown aerobically at 30 °C on a rotary shaker (120 rpm) in 500 ml baffled shake flasks with 60 ml BHI complex medium or CGXII minimal medium [52]. C. glutamicum cells were inoculated from 5 ml LB medium overnight culture to an optical density at 600 nm (OD600) of 0.6 in 60 ml CGXII-medium with 0.03 g/l protocatechuic acid as iron chelator and 40 g/l glucose or 20 g/l sodium acetate as carbon and energy source. For medium shift experiments, cells were harvested 14–18 h after inoculation by centrifugation at 4 °C, washed with CGXII without Pi and carbon sources, and inoculated in 60 ml CGXII medium with sufficient Pi (13 mM) to an optical density at 600 nm (OD600) of 0.6. These main cultures were cultivated until OD600 of 4 – 5 h. The cells were harvested and either stored at −20 °C for further analysis or washed with CGXII without Pi and carbon source, and resuspended in an equal volume of fresh CGXII medium that contained either a limiting Pi concentration (0.065 mM) or no Pi. After incubation at 30 °C for 10, 30, 60, 90 and 120 min in the Pi low or Pi free medium, cells were harvested and stored at −20 °C for further analysis. For comparative growth experiments on different phosphorus sources, C. glutamicum cells growing exponentially on CGXII medium with sufficient Pi (13 mM) were inoculated in 60 ml Pi-free CGXII medium to an OD600 of 0.6 and cultured for 24 h at 30 °C to deplete intracellular polyphosphate storage. Afterwards, these cells were harvested, washed with CGXII without Pi and carbon source, and inoculated to an OD600 of 0.6 in CGXII medium containing either 0.065 mM Pi, 1 mM adenosine 5’-monophosphate (5’AMP), 1 mM L-α-glycerophosphate or 1 mM UDP-Glucose as sole phosphorus source.
Preparation of supernatants and assay to determine UDP-glucose hydrolase activity
Cell cultures were centrifuged for 10 min at 5,000 g and 4 °C. Supernatants were passed through a 0.2 μm sterile filter and concentrated about 50 fold by ultrafiltration using Amicon Ultra MW 10.000 membranes (Millipore, Bedford, USA). UDP-sugar hydrolase activity was determined at 37 °C in a coupled spectrophotometric assay essentially as described before [53]. Briefly, reactions of the mixture containing 35 mM Tris–HCl, pH 8.0, 35 mM MgCl2, 3.1 μM glucose-1,6-bisphosphate, 0.7 mM NADP+, rabbit muscle phosphoglucomutase (1 U/ml) and Leuconostoc mesenteroides glucose-6-phosphate dehydrogenase (2.5 U/ml) were started by the addition of 1.4 mM UDP-glucose to the final volume of 1 ml. Glucose-1-phosphate formed by the reaction of UDP-sugar hydrolase was converted to glucose-6-phosphate and subsequently to 6-phosphogluconate by coupling of phosphoglucomutase and glucose-6-dehydrogenase, and the concomitant formation of NADPH (ε 340 nm = 6.3 mM−1 cm−1) was measured at 340 nm.
Construction of transcriptional fusions and chloramphenicol acetyltransferase (CAT) assays
Different parts of the upstream region of the pstSCAB operon were amplified using the primers respectively named pstsR, pstsR0, pstsR1, pstsR2, pstsR3, pstsF0, pstsF1, pstsF2, pstsF3, pstsRc, pstsRm, pstsFc and pstsFm (Table 3) and cloned into the corynebacterial promoter-probe vector pET2 [54]. The vector pGEM-T (Table 1) was used for subcloning. The correct sequence of the cloned promoter fragments was verified by sequencing (AGOWA, Berlin, Germany). The constructed promoter-probe vectors were introduced into C. glutamicum WT as well as into the ΔphoRS mutant by electroporation using the following conditions: 25 μF, 600 Ω and 2.5 kV/cm (Bio-Rad Gene Pulser Xcell, Bio-Rad Laboratories, Hercules, Canada). After electroporation, 1 ml BHI/sorbitol medium was added immediately to the sample [55]. The cell suspension was exposed to 46 °C for 6 min and incubated at 30 °C for 90 min for regeneration. The CAT assays were performed as described previously [56].
DNA affinity chromatography
The purification of DNA-binding proteins was performed essentially as described previously [57]. Briefly, pstS promoter fragments were generated by PCR using genomic DNA from C. glutamicum and the primer pair pstsR0/pstsF0bio. Primer pstsF0bio was tagged with biotin via a TEG linker (Operon, Cologne, Germany). Unincorporated oligonucleotides were removed by the Qiaquick PCR purification kit (Qiagen, Hilden, Germany). About 100 pmol of biotin-labeled PCR product was coupled to 5 mg of Dynabeads streptavidin (Dynal, Oslo, Norway) and free DNA was removed by magnetic separation. The coupled Dynabeads were stored at 4 °C. Cultures (900 ml) of C. glutamicum were grown on CGXII minimal medium, harvested at an optical density at 600 nm (OD600) of about 4, washed with 1 volume of TN buffer (50 mM NaCl, 50 mM Tris–HCl, pH 7.6) and suspended in 6 ml of TGED buffer (50 mM Tris–HCl (pH 7.6), 1 mM dithiothreitol, 10 mM MgCl2, 1 mM EDTA, 10 % (v/v) glycerol, 10 μM phenylmethylsulfonyl fluoride). The resuspended cell pellet was passed six times through a French pressure cell (SLM Amino, Spectronic Instruments, Rochester, NY) at 207 MPa. Cellular debris was removed by centrifugation at 8,000 g and 4 °C for 10 min and at 15,000 g and 4 °C for 60 min. Directly before incubation with the C. glutamicum crude extracts and the coupled Dynabeads, the beads were equilibrated with 300 μl of binding buffer (20 mM Tris–HCl pH 7,5, 1 mM EDTA, 10 % (v/v) glycerol, 0.01 % (v/v) Triton X-100, 100 mM NaCl and 1 mM dithiothreitol) for 2 min. The crude extract (about 6 ml) and 500 μg genomic DNA from C. glutamicum were incubated with the coupled Dynabeads for 1 h at room temperature with enough shaking to prevent sedimentation of the paramagnetic beads (150 rpm). Subsequently, the reaction was transferred into microcentrifuge tubes, washed once with 1 ml of TGED buffer, twice with 1 ml of TGED buffer including 400 μg of chromosomal DNA from C. glutamicum and finally with 1 ml of TGED buffer. Proteins bound to the immobilized DNA were eluted by washing the beads twice with 350 μl of elution buffer (TGED buffer containing 2 M NaCl). The eluates were pooled, concentrated and desalted with Microcon 3 microconcentrators (Millipore, Bedford, USA) and analysed by denaturing PAGE [51]. Gels were stained subsequently using a colloidal Coomassie blue staining kit (Novex, Frankfurt/Main, Germany).
MALDI-TOF mass spectrometry
For peptide mass fingerprinting, the protein band of interest was cut out from gels and subjected to in-gel digestion with trypsin essentially as described previously [58]. Briefly, gel pieces were washed twice with 750 μl of 0.1 M ammonium bicarbonate in 30 % (v/v) acetonitrile for 10 min. The destained and shrunken gel pieces were vacuum-dried for 20 min in a conventional vacuum centrifuge and subsequently rehydrated with 6 μl of 3 mM Tris–HCl (pH 8.8) containing trypsin (10 ng/μl). After 20 min, 6 μl of 3 mM Tris–HCl (pH 8.8) without trypsin was added. Digestion was allowed to proceed overnight at room temperature. Peptides were then extracted by sequential addition of 6 μl of water and 10 μl of 0.1 % (v/v) trifluoroacetic acid in 30 % (v/v) acetonitrile. A total of 0.5 μl of the resulting peptide solution was mixed on a stainless steel sample plate with 0.5 μl of a saturated μ-cyano-4-hydroxy-trans cinnamic acid solution in 50 % (v/v) acetonitrile – 0.1 % (v/v) trifluoroacetic acid. Close external calibration using calibration mixtures 1 and 2 of a Sequazyme peptide mass standard kit (Applied Biosystems, Weiterstadt, Germany) was performed. Samples were analyzed manually in positive-reflector mode with 20 kV of accelerating voltage and 63 % grid voltage; the delay time was set at 125 ns. Data acquisition and analysis were performed using Voyager Control Panel software (version 5.0) and Voyager Data Explorer software (version 3.5) (Applied Biosystems). The generated mass lists and MS-Fit were used to search the National Center for Biotechnology Information (NCBI) database [59].
Overproduction and purification of RamB
The RamB fusion protein was prepared essentially as described previously [41, 60]. Briefly, E. coli Bl21 (DE3) carrying the plasmid pET29-ramB-his was grown at 30 °C in 500 ml LB with 50 μg/ml kanamycin to an OD of 0.5 before adding 1 mM isopropyl ß-D-thiogalactoside. Four hours after induction, cells were harvested by centrifugation and stored at – 20 °C. For cell extract preparation, thawed cells were resuspended in 10 ml of TNGI5 buffer (20 mM Tris/HCl, pH 7.9, 300 mM NaCl, 5 % (v/v) glycerol, 5 mM imidazol) containing 1 mM diisopropylfluorophosphate and 1 mM phenylmethylsulfonyl fluoride. The cell suspension was passed six times through a French pressure cell (SLM Amino, Spectronic Instruments, Rochester, NY) at 207 MPa. Cell debris and intact cells were removed by centrifugation for 10 min at 5,000 g amd 4 °C, and the cell-free extract was subjected to centrifugation again for 1 h at 15,000 g and 4 °C. After centrifugation, the supernatant was purified by nickel affinity chromatography using Ni-NTA agarose (Novagen, San Diego, USA). The column was washed with TNGI20 and TNGI50 buffer (which contained 20 mM or 50 mM imidazol). The RamB protein was eluted with TNGI200 buffer (which contained 200 mM imidazol). Fractions containing RamB were pooled, and the elution buffer was exchanged against BS buffer (100 mM Tris/HCl, 20 % (v/v) glycerol, 100 mM KCl, 20 mM MgCl2, 1 mM EDTA, pH 7.5). From 250 ml of culture, ~ 4 mg of RamB was purified to apparent homogeneity (Fig. 3b).
Gel mobility shift assays
Gel shift assays with RamB were prepared as described previously [60]. Briefly, overexpressed and purified RamB was mixed with the putative target promoter pstS (RF0) or promoter fragments (R0F0, R1F0, R2F0, R3F0, RF4, FcRc, FmRc, FcRm and FmRm) (124 bps – 507 bps, final concentrations 61 nM – 15 nM) (Figs. 4, 5) in a total volume of 20 μl. The binding buffer contained 100 mM Tris/HCl, 20 % (v/v) glycerol, 100 mM KCl, 20 mM MgCl2, 1 mM EDTA, pH 7.5. Approximately 40 nM of a nontarget promoter fragment (Pcg0527, PdtxR or R3F0) (Figs. 4, 5) were added as a negative control. After incubation for 30 min at room temperature, the samples were separated on a 10 % native polyacrylamide gel at room temperature and 170 V using 1x TBE (89 mM Tris base, 89 mM boric acid, 2 mM EDTA) as electrophoresis buffer. The gels were subsequently stained with Sybr Green I (Sigma, Rödermark, Germany) and photographed.
DNA microarray analysis
Total RNA was isolated from exponentially growing cells by using the RNAeasy system (QIAGEN, Hilden, Germany) with on-column DNase I treatment prepared as described [61]. Quantity and quality of purified RNA was analyzed by UV-spectrometry and stored at −20 °C until use. DNA microarrays are based on PCR products of C. glutamicum genes [62]. Synthesis of fluorescently labelled cDNA from total RNA, microarray hybridization, washing and gene expression analysis were carried out as described previously [61–63]. The data are available as Gene Expression Omnibus GSE67012 data set at http://www.ncbi.nlm.nih.gov/geo/.
Acknowledgements
We thank Karin Niermann (Münster) for technical assistance, Hermann Sahm (Jülich) for support during the initial phase of the project and Michael Bott (Jülich) for access to and help with the mass spectrometry facility. We thank Michael Bott, Martina Kocan (Jülich) and Sarah Schaaf (Jülich) for fruitful discussions. We thank Annette Cramer and Bernhard J. Eikmanns (Ulm) for providing deletion strain ΔramB, plasmid pET29-ramB-his and for advice on purification of RamB.
Footnotes
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
USH and VFW planned and designed the experiments. USH performed the analysis and analysed data. HT analysed data. USH and HT drafted the manuscript. VFW coordinated the study, analysed data and finalized the manuscript. All authors read and approved the manuscript.
Contributor Information
Ulrike Sorger-Herrmann, Email: ulrike.sorger-herrmann@sandoz.com.
Hironori Taniguchi, Email: thiro@cebitec.uni-bielefeld.de.
Volker F. Wendisch, Phone: +49 521 106 5611, Email: volker.wendisch@uni-bielefeld.de
References
- 1.Neidhardt FC. Escherichia coli and Salmonella: Cellular and Molecular Biology. Washington, DC, USA: ASM Press; 1996. [Google Scholar]
- 2.Sonenshein AL, Hoch JA, Losick R. Bacillus subtilis and Its Closest Relatives. Washington, DC, USA: ASM Press; 2002. [Google Scholar]
- 3.Hecker M, Völker U. Non-specific, general and multiple stress resistance of growth-restricted Bacillus subtilis cells by the expression of the σB regulon. Mol Microbiol. 1998;29:1129–1136. doi: 10.1046/j.1365-2958.1998.00977.x. [DOI] [PubMed] [Google Scholar]
- 4.Prágai Z, Allenby NEE, O’Connor N, Dubrac S, Rapoport G, Msadek T, et al. Transcriptional regulation of the phoPR Operon in Bacillus subtilis. J Bacteriol. 2004;186:1182–1190. doi: 10.1128/JB.186.4.1182-1190.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Prágai Z, Harwood CR. Regulatory interactions between the Pho and σB-dependent general stress regulons of Bacillus subtilis. Microbiology. 2002;148:1593–1602. doi: 10.1099/00221287-148-5-1593. [DOI] [PubMed] [Google Scholar]
- 6.Price CW, Fawcett P, Cérémonie H, Su N, Murphy CK, Youngman P. Genome-wide analysis of the general stress response in Bacillus subtilis. Mol Microbiol. 2001;41:757–774. doi: 10.1046/j.1365-2958.2001.02534.x. [DOI] [PubMed] [Google Scholar]
- 7.Antelmann H, Scharf C, Hecker M. Phosphate starvation-inducible proteins of Bacillus subtilis: proteomics and transcriptional analysis. J Bacteriol. 2000;182:4478–4490. doi: 10.1128/JB.182.16.4478-4490.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Minnig K, Lazarevic V, Soldo B, Mauël C. Analysis of teichoic acid biosynthesis regulation reveals that the extracytoplasmic function sigma factor σM is induced by phosphate depletion in Bacillus subtilis W23. Microbiology. 2005;151:3041–3049. doi: 10.1099/mic.0.28021-0. [DOI] [PubMed] [Google Scholar]
- 9.Abe S, Takayama K-I, Kinoshita S. Taxonomical studies on glutamic acid-producing bacteria. J Gen Appl Microbiol. 1967;13:279–301. doi: 10.2323/jgam.13.279. [DOI] [Google Scholar]
- 10.Hermann T. Industrial production of amino acids by coryneform bacteria. J Biotechnol. 2003;104:155–172. doi: 10.1016/S0168-1656(03)00149-4. [DOI] [PubMed] [Google Scholar]
- 11.Leuchtenberger W, Huthmacher K, Drauz K. Biotechnological production of amino acids and derivatives: current status and prospects. Appl Microbiol Biotechnol. 2005;69:1–8. doi: 10.1007/s00253-005-0155-y. [DOI] [PubMed] [Google Scholar]
- 12.Peters-Wendisch P, Stolz M, Etterich H, Kennerknecht N, Sahm H, Eggeling L. Metabolic engineering of Corynebacterium glutamicum for L-serine production. Appl Environ Microbiol. 2005;71:7139–7144. doi: 10.1128/AEM.71.11.7139-7144.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Morbach S, Sahm H, Eggeling L. L-Isoleucine production with Corynebacterium glutamicum: further flux increase and limitation of export. Appl Environ Microbiol. 1996;62:4345–4351. doi: 10.1128/aem.62.12.4345-4351.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Blombach B, Schreiner ME, Holátko J, Bartek T, Oldiges M, Eikmanns BJ. L-valine production with pyruvate dehydrogenase complex-deficient Corynebacterium glutamicum. Appl Environ Microbiol. 2007;73:2079–2084. doi: 10.1128/AEM.02826-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Radmacher E, Vaitsikova A, Burger U, Krumbach K, Sahm H, Eggeling L. Linking central metabolism with increased pathway flux: L-valine accumulation by Corynebacterium glutamicum. Appl Environ Microbiol. 2002;68:2246–2250. doi: 10.1128/AEM.68.5.2246-2250.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Jensen JVK, Wendisch VF. Ornithine cyclodeaminase-based proline production by Corynebacterium glutamicum. Microb Cell Factories. 2013;12:63. doi: 10.1186/1475-2859-12-63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Schneider J, Eberhardt D, Wendisch VF. Improving putrescine production by Corynebacterium glutamicum by fine-tuning ornithine transcarbamoylase activity using a plasmid addiction system. Appl Microbiol Biotechnol. 2012;95:169–178. doi: 10.1007/s00253-012-3956-9. [DOI] [PubMed] [Google Scholar]
- 18.Schneider J, Wendisch VF. Putrescine production by engineered Corynebacterium glutamicum. Appl Microbiol Biotechnol. 2010;88:859–868. doi: 10.1007/s00253-010-2778-x. [DOI] [PubMed] [Google Scholar]
- 19.Mimitsuka T, Sawai H, Hatsu M, Yamada K. Metabolic engineering of Corynebacterium glutamicum for cadaverine fermentation. Biosci, Biotechnol, Biochem. 2007;71:2130–2135. doi: 10.1271/bbb.60699. [DOI] [PubMed] [Google Scholar]
- 20.Krause FS, Blombach B, Eikmanns BJ. Metabolic engineering of Corynebacterium glutamicum for 2-ketoisovalerate production. Appl Environ Microbiol. 2010;76:8053–8061. doi: 10.1128/AEM.01710-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Bückle-Vallant V, Krause FS, Messerschmidt S, Eikmanns BJ. Metabolic engineering of Corynebacterium glutamicum for 2-ketoisocaproate production. Appl Microbiol Biotechnol. 2014;98:297–311. doi: 10.1007/s00253-013-5310-2. [DOI] [PubMed] [Google Scholar]
- 22.Vogt M, Haas S, Polen T, van Ooyen J, Bott M. Production of 2-ketoisocaproate with Corynebacterium glutamicum strains devoid of plasmids and heterologous genes. Biotechnol: Microb; 2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wendisch VF, Bott M. Handbook of Corynebacterium glutamicum. Boca Raton, USA: CRC Press; 2005. Phosphorus Metabolism; pp. 377–396. [Google Scholar]
- 24.Klauth P, Pallerla SR, Vidaurre D, Ralfs C, Wendisch VF, Schoberth SM. Determination of soluble and granular inorganic polyphosphate in Corynebacterium glutamicum. Appl Microbiol Biotechnol. 2006;72:1099–1106. doi: 10.1007/s00253-006-0562-8. [DOI] [PubMed] [Google Scholar]
- 25.Lambert C, Weuster-Botz D, Weichenhain R, Kreutz EW, de Graaf AA, Schoberth SM. Monitoring of inorganic polyphosphate dynamics in Corynebacterium glutamicum using a novel oxygen sparger for real time 31P in vivo NMR. Acta Biotechnol. 2002;22:245–260. doi: 10.1002/1521-3846(200207)22:3/4<245::AID-ABIO245>3.0.CO;2-E. [DOI] [Google Scholar]
- 26.Pallerla SR, Knebel S, Polen T, Klauth P, Hollender J, Wendisch VF, et al. Formation of volutin granules in Corynebacterium glutamicum. FEMS Microbiol Lett. 2005;243:133–140. doi: 10.1016/j.femsle.2004.11.047. [DOI] [PubMed] [Google Scholar]
- 27.Lindner SN, Vidaurre D, Willbold S, Schoberth SM, Wendisch VF. NCgl2620 encodes a class II polyphosphate kinase in Corynebacterium glutamicum. Appl Environ Microbiol. 2007;73:5026–5033. doi: 10.1128/AEM.00600-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lindner SN, Knebel S, Wesseling H, Schoberth SM, Wendisch VF. Exopolyphosphatases PPX1 and PPX2 from Corynebacterium glutamicum. Appl Environ Microbiol. 2009;75:3161–3170. doi: 10.1128/AEM.02705-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lindner SN, Niederholtmeyer H, Schmitz K, Schoberth SM, Wendisch VF. Polyphosphate/ATP-dependent NAD kinase of Corynebacterium glutamicum: biochemical properties and impact of ppnK overexpression on lysine production. Appl Microbiol Biotechnol. 2010;87:583–593. doi: 10.1007/s00253-010-2481-y. [DOI] [PubMed] [Google Scholar]
- 30.Lindner SN, Knebel S, Pallerla SR, Schoberth SM, Wendisch VF. Cg2091 encodes a polyphosphate/ATP-dependent glucokinase of Corynebacterium glutamicum. Appl Microbiol Biotechnol. 2010;87:703–713. doi: 10.1007/s00253-010-2568-5. [DOI] [PubMed] [Google Scholar]
- 31.Ishige T, Krause M, Bott M, Wendisch VF, Sahm H. The phosphate starvation stimulon of Corynebacterium glutamicum determined by DNA microarray analyses. J Bacteriol. 2003;185:4519–4529. doi: 10.1128/JB.185.15.4519-4529.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Rittmann D, Sorger-Herrmann U, Wendisch VF. Phosphate starvation-inducible gene ushA encodes a 5’ nucleotidase required for growth of Corynebacterium glutamicum on media with nucleotides as the phosphorus source. Appl Environ Microbiol. 2005;71:4339–4344. doi: 10.1128/AEM.71.8.4339-4344.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kočan M, Schaffer S, Ishige T, Sorger-Herrmann U, Wendisch VF, Bott M. Two-component systems of Corynebacterium glutamicum: deletion analysis and involvement of the PhoS-PhoR system in the phosphate starvation response. J Bacteriol. 2006;188:724–732. doi: 10.1128/JB.188.2.724-732.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Schaaf S, Bott M. Target genes and DNA-binding sites of the response regulator PhoR from Corynebacterium glutamicum. J Bacteriol. 2007;189:5002–5011. doi: 10.1128/JB.00121-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Jungwirth B, Sala C, Kohl TA, Uplekar S, Baumbach J, Cole ST, et al. High-resolution detection of DNA binding sites of the global transcriptional regulator GlxR in Corynebacterium glutamicum. Microbiology. 2013;159:12–22. doi: 10.1099/mic.0.062059-0. [DOI] [PubMed] [Google Scholar]
- 36.Townsend PD, Jungwirth B, Pojer F, Bußmann M, Money VA, Cole ST, et al. The Crystal Structures of Apo and cAMP-Bound GlxR from Corynebacterium glutamicum Reveal Structural and Dynamic Changes upon cAMP Binding in CRP/FNR Family Transcription Factors. PLoS One. 2014;9 doi: 10.1371/journal.pone.0113265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Toyoda K, Teramoto H, Inui M, Yukawa H. Genome-wide identification of in vivo binding sites of GlxR, a cyclic AMP receptor protein-type regulator in Corynebacterium glutamicum. J Bacteriol. 2011;193:4123–4133. doi: 10.1128/JB.00384-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Panhorst M, Sorger-Herrmann U, Wendisch VF. The pstSCAB operon for phosphate uptake is regulated by the global regulator GlxR in Corynebacterium glutamicum. J Biotechnol. 2011;154:149–155. doi: 10.1016/j.jbiotec.2010.07.015. [DOI] [PubMed] [Google Scholar]
- 39.Woo HM, Noack S, Seibold GM, Willbold S, Eikmanns BJ, Bott M. Link between phosphate starvation and glycogen metabolism in Corynebacterium glutamicum, revealed by metabolomics. Appl Environ Microbiol. 2010;76:6910–6919. doi: 10.1128/AEM.01375-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Cramer A, Auchter M, Frunzke J, Bott M, Eikmanns BJ. RamB, the transcriptional regulator of acetate metabolism in Corynebacterium glutamicum, is subject to regulation by RamA and RamB. J Bacteriol. 2007;189:1145–1149. doi: 10.1128/JB.01061-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Gerstmeir R, Cramer A, Dangel P, Schaffer S, Eikmanns BJ. RamB, a novel transcriptional regulator of genes involved in acetate metabolism of Corynebacterium glutamicum. J Bacteriol. 2004;186:2798–2809. doi: 10.1128/JB.186.9.2798-2809.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Blombach B, Cramer A, Eikmanns BJ, Schreiner M. RamB is an activator of the pyruvate dehydrogenase complex subunit E1p gene in Corynebacterium glutamicum. J Mol Microbiol Biotechnol. 2009;16:236–239. doi: 10.1159/000108782. [DOI] [PubMed] [Google Scholar]
- 43.Kim H-J, Kim T-H, Kim Y, Lee H-S. Identification and characterization of glxR, a gene involved in regulation of glyoxylate bypass in Corynebacterium glutamicum. J Bacteriol. 2004;186:3453–3460. doi: 10.1128/JB.186.11.3453-3460.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Van Ooyen J, Emer D, Bussmann M, Bott M, Eikmanns BJ, Eggeling L. Citrate synthase in Corynebacterium glutamicum is encoded by two gltA transcripts which are controlled by RamA, RamB, and GlxR. J Biotechnol. 2011;154:140–148. doi: 10.1016/j.jbiotec.2010.07.004. [DOI] [PubMed] [Google Scholar]
- 45.Jungwirth B, Emer D, Brune I, Hansmeier N, Pühler A, Eikmanns BJ, et al. Triple transcriptional control of the resuscitation promoting factor 2 (rpf2) gene of Corynebacterium glutamicum by the regulators of acetate metabolism RamA and RamB and the cAMP-dependent regulator GlxR. FEMS Microbiol Lett. 2008;281:190–197. doi: 10.1111/j.1574-6968.2008.01098.x. [DOI] [PubMed] [Google Scholar]
- 46.Cramer A, Gerstmeir R, Schaffer S, Bott M, Eikmanns BJ. Identification of RamA, a Novel LuxR-Type Transcriptional Regulator of Genes Involved in Acetate Metabolism of Corynebacterium glutamicum. J Bacteriol. 2006;188:2554–2567. doi: 10.1128/JB.188.7.2554-2567.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Rengarajan J, Bloom BR, Rubin EJ. Genome-wide requirements for Mycobacterium tuberculosis adaptation and survival in macrophages. Proc Natl Acad Sci U S A. 2005;102:8327–8332. doi: 10.1073/pnas.0503272102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Kimura S, Makino K, Shinagawa H, Amemura M, Nakata A. Regulation of the phosphate regulon of Escherichia coli: characterization of the promoter of the pstS gene. Mol Gen Genet MGG. 1989;215:374–380. doi: 10.1007/BF00427032. [DOI] [PubMed] [Google Scholar]
- 49.Spira B, Yagil E. The integration host factor (IHF) affects the expression of the phosphate-binding protein and of alkaline phosphatase in Escherichia coli. Curr Microbiol. 1999;38:80–85. doi: 10.1007/s002849900407. [DOI] [PubMed] [Google Scholar]
- 50.Liu W, Qi Y, Hulett FM. Sites internal to the coding regions of phoA and pstS bind PhoP and are required for full promoter activity. Mol Microbiol. 1998;28:119–130. doi: 10.1046/j.1365-2958.1998.00779.x. [DOI] [PubMed] [Google Scholar]
- 51.Sambrook J. Molecular Cloning: A Laboratory Manual, Third Edition. N.Y: Cold Spring Harbor Laboratory Press; 2001. [Google Scholar]
- 52.Keilhauer C, Eggeling L, Sahm H. Isoleucine synthesis in Corynebacterium glutamicum: molecular analysis of the ilvB-ilvN-ilvC operon. J Bacteriol. 1993;175:5595–5603. doi: 10.1128/jb.175.17.5595-5603.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Edwards CJ, Innes DJ, Burns DM, Beacham IR. UDP-sugar hydrolase isozymes in Salmonella enterica and Escherichia coli: silent alleles of ushA in related strains of group I Salmonella isolates, and of ushB in wild-type and K12 strains of E. coli, indicate recent and early silencing events, respectively. FEMS Microbiol Lett. 1993;114:293–298. doi: 10.1111/j.1574-6968.1993.tb06588.x. [DOI] [PubMed] [Google Scholar]
- 54.Vasicová P, Abrhámová Z, Nesvera J, Pátek M, Sahm H, Eikmanns B. Integrative and autonomously replicating vectors for analysis of promoters in Corynebacterium glutamicum. Biotechnol Tech. 1998;12:743–746. doi: 10.1023/A:1008827609914. [DOI] [Google Scholar]
- 55.Van der Rest ME, Lange C, Molenaar D. A heat shock following electroporation induces highly efficient transformation of Corynebacterium glutamicum with xenogeneic plasmid DNA. Appl Microbiol Biotechnol. 1999;52:541–545. doi: 10.1007/s002530051557. [DOI] [PubMed] [Google Scholar]
- 56.Engels V, Wendisch VF. The DeoR-type regulator SugR represses expression of ptsG in Corynebacterium glutamicum. J Bacteriol. 2007;189:2955–2966. doi: 10.1128/JB.01596-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Gabrielsen OS, Hornes E, Korsnes L, Ruet A, Oyen TB. Magnetic DNA affinity purification of yeast transcription factor tau–a new purification principle for the ultrarapid isolation of near homogeneous factor. Nucleic Acids Res. 1989;17:6253–6267. doi: 10.1093/nar/17.15.6253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Schaffer S, Weil B, Nguyen VD, Dongmann G, Günther K, Nickolaus M, et al. A high-resolution reference map for cytoplasmic and membrane-associated proteins of Corynebacterium glutamicum. Electrophoresis. 2001;22:4404–4422. doi: 10.1002/1522-2683(200112)22:20<4404::AID-ELPS4404>3.0.CO;2-2. [DOI] [PubMed] [Google Scholar]
- 59.Clauser KR, Baker P, Burlingame AL. Role of accurate mass measurement (±10 ppm) in protein identification strategies employing MS or MS/MS and database searching. Anal Chem. 1999;71:2871–2882. doi: 10.1021/ac9810516. [DOI] [PubMed] [Google Scholar]
- 60.Wennerhold J, Krug A, Bott M. The AraC-type regulator RipA represses aconitase and other iron proteins from Corynebacterium under iron limitation and is itself repressed by DtxR. J Biol Chem. 2005;280:40500–40508. doi: 10.1074/jbc.M508693200. [DOI] [PubMed] [Google Scholar]
- 61.Netzer R, Krause M, Rittmann D, Peters-Wendisch PG, Eggeling L, Wendisch VF, et al. Roles of pyruvate kinase and malic enzyme in Corynebacterium glutamicum for growth on carbon sources requiring gluconeogenesis. Arch Microbiol. 2004;182:354–363. doi: 10.1007/s00203-004-0710-4. [DOI] [PubMed] [Google Scholar]
- 62.Wendisch VF. Genome-wide expression analysis in Corynebacterium glutamicum using DNA microarrays. J Biotechnol. 2003;104:273–285. doi: 10.1016/S0168-1656(03)00147-0. [DOI] [PubMed] [Google Scholar]
- 63.Polen T, Schluesener D, Poetsch A, Bott M, Wendisch VF. Characterization of citrate utilization in Corynebacterium glutamicum by transcriptome and proteome analysis. FEMS Microbiol Lett. 2007;273:109–119. doi: 10.1111/j.1574-6968.2007.00793.x. [DOI] [PubMed] [Google Scholar]
- 64.Studier FW, Rosenberg AH, Dunn JJ, Dubendorff JW. Use of T7 RNA polymerase to direct expression of cloned genes. Methods Enzymol. 1990;185:60–89. doi: 10.1016/0076-6879(90)85008-c. [DOI] [PubMed] [Google Scholar]
- 65.Hanahan D. Studies on transformation of Escherichia coli with plasmids. J Mol Biol. 1983;166:557–580. doi: 10.1016/S0022-2836(83)80284-8. [DOI] [PubMed] [Google Scholar]


