Abstract
Genome sequences are now available for two macaque species used in infectious disease research and drug safety testing.
Macaques are the most widely used nonhuman primates in biomedical research. They serve as models for >70 human infectious diseases and have been instrumental in the development ~15 vaccines licensed in the United States1. In this issue, Yan et al. 2 report the genome sequence of two members of the genus Macaca: the cynomolgus/crab-eating macaque (Macaca fasicularis), and the Chinese rhesus macaque (Macaca mulatta lasiota). This work contributes to our understanding of the biology and pathology of two important animal models of human disease, sets the groundwork for the development of new medically important genomic tools and opens up new avenues for the study of comparative primate evolution.
The first primate to be sequenced was the human3 in 2001, followed by the chimpanzee4 in 2005, the Indian rhesus macaque5 (Macaca mulatta mulatta) in 2007 and the orangutan6 earlier this year. All these genomes were deciphered using a shotgun approach and Sanger sequencing. With the advent of short-read, next generation sequencing, sequencing of whole genomes has become feasible beyond the confines of large research consortia. Not surprisingly, genome sequencing projects for non-human primates have multiplied, with projects on gibbons, baboons, bonobos, gorillas, African green monkeys, squirrel monkeys, galagos (bushbabies), pigtailed macaques, aye-aye (lemur), sooty mangabeys and other species underway or planned (Fig. 1).
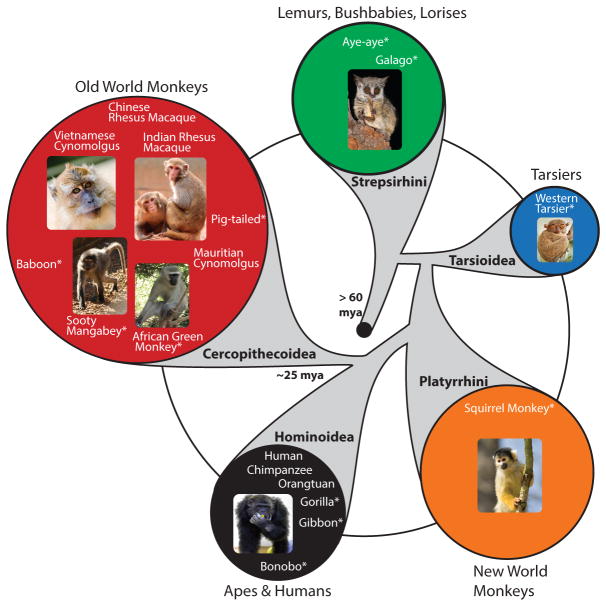
Figure 1. Sequenced primate genomes.
The genome sequences of human, chimpanzee, orangutan, Indian rhesus macaque, Chinese rhesus macaque and cynomolgus macaques have been reported. Several more primate species are currently undergoing sequencing (indicated by asterisk). Phylogeny of extant primate groups is indicated by a top-down view looking backwards in time. Estimated time of divergence from common ancestors is indicated at select branch points. The relative area of each circle indicates the number of species in each group. Phylogenetic tree courtesy of T. Preuss, modified from The Primate Visual System, Collins & Kaas (eds.)12. Photo credits: sooty mangabey, Mollie Bloomsmith/Yerkes National Primate Research Center; rhesus macaques, chimpanzee and African green monkey, Yerkes National Primate Research Center; Cynomolgus, Anna Yu/iStockphoto; squirrel monkey, EcoPic/iStockphoto; bushbaby, Nico Smit/iStockphoto.
The study by Yan et al.2 is remarkable in that reports the first primate genomes to be derived completely by de novo assembly of short-reads, without using existing primate genomes as templates. To achieve this, the authors constructed many sequencing libraries—19 for a female Chinese rhesus macaque and 18 for a Vietnamese female cynomolgus macaque monkey—containing fragments of varying lengths. Knowing the distance between paired-end reads from these fragments provided critical information for the de novo assembly process. Yan et al.2 also obtained high sequence coverage, reaching an average of 47× coverage for the rhesus and 54× for the cynomolgus macaque. In total, >140 billion base pairs were sequenced for each ~2.8 Gb genome. Of note, concurrent with the study by Yan et al., the genome sequences of a male Chinese rhesus macaque (5.5× coverage)7 and a female Mauritian cynomolgus macaque (6× coverage)8 were independently reported. The authors also used RNA-seq to sequence the transcriptome of several tissues; in combination with genomic sequences these data identified novel species-specific genes.
The initial analyses of Yan et al.2 have already revealed some intriguing findings. In AIDS research, infection of macaques with simian immunodeficiency virus (SIV), closely related to HIV, is used to study viral pathogenesis, therapy and prevention. Although the animal model of choice in this field has been the Indian rhesus macaque, sequenced in 2007 (ref. 5), restrictions on the export of these animals from India have increased demand for Chinese rhesus macaques and, to a lesser extent, for cynomolgus macaques, underscoring the need for reference genome sequences of these species.
In this study, Yan et al.2 sequenced the gene encoding for the retroviral restriction factor TRIM5α in a population of 33 unrelated cynomolgus and 28 Chinese rhesus macaques. They found that none of the Chinese rhesus macaques show the ‘TrimCyp’ allele that is common in Indian animals and encodes for a TRIM5a-Cyclophillin fusion protein that is associated with reduced replication of certain SIV strains9. In addition, they observed that the a 6-bp deletion in the SPRY domain of this molecule (previously described as TF339–340ΔΔ, ref. 10) that increases the permissivity to SIV replication is nearly ubiquitous among cynomolgus macaques, but present at similar frequencies in Chinese and Indian rhesus macaques (50% and 36%, respectively). Given the influence of TRIM5α alleles on SIV transmission and pathogenesis in vivo, these findings enhance the use of these macaque species in studies of AIDS pathogenesis and prevention.
Cynomolgus macaques are used mostly in pharmacology as the reference non-human primate model for drug safety testing. In the Indian and Chinese rhesus macaques, and in the cynomolgus macaque, Yan et al.2 identified orthologs of biomedically relevant human genes, including proteins with druggable domains. These data should facilitate the interpretation of preclinical drug testing results in these animals.
Identifying intra- and interspecies polymorphisms is certainly useful, but more sophisticated evolutionary analyses are possible, especially with the two highly related species sequenced by Yan et al.2. By identifying portions of the genome that were transferred from one species to another, called regions of introgression, the authors estimated that ~30% of the cynomolgus macaque genome originated from the Chinese rhesus macaque. This observation supports models in which interspecies breeding within overlapping habitat ranges in Indochina caused an ancestral introgression into the cynomolgus macaque genome.
The availability of sequence data from three macaque species also allowed interesting comparisons between macaques and humans. Yan et al.2 scanned the genomes for regions with little variation between the three macaque genomes but normal levels of divergence between macaques and humans. These regions could be evidence of selective pressure that maintains sequences unique to the macaque genus. The authors identified 217 such regions undergoing ‘selective sweeps’. Notably, many of them contained only a single gene, and in some no genes were found. These results are consistent with concurrent work showing high sequence conservation between humans and macaques in protein-coding sequences but substantially greater divergence in untranslated flanking regions8. Thus, a picture is emerging in which variation in intergenic regions may be more important than the genes themselves in determining the differences between humans and monkeys.
The results of Yan et al.2 provide a tantalizing glimpse of the value of sequencing the full genomes of additional species of nonhuman primates. Moreover, their transcriptome sequencing data suggest that characterizing transcripts in individual cell or tissue types may help elucidate the mechanisms responsible for species-specific pathogenic processes acting at the level of gene regulation.
Among the nonhuman primate species whose genome sequence will be highly relevant to research are the African monkey species that are natural hosts for SIV, such as the sooty mangabeys, African green monkeys, and mandrills. In striking contrast to pathogenic HIV infection of humans and SIV infection of macaques, natural SIV infections are typically nonpathogenic despite robust virus replication. Moreover, they are characterized by low levels of immune activation and a specific pattern of infected cells11. Genome sequences for these species may lead to better understanding of the molecular and cellular mechanisms underlying the development of AIDS in HIV-infected individuals. For instance, the technique of scanning for selective sweeps could be applied to identify regions undergoing positive selection in African species that are natural hosts to SIV compared with non-natural Asian hosts. The results of Yan et al.2 suggest that this approach is effective even among closely related nonhuman primate subspecies.
Primate genomics promise to yield evolutionary insight, new research tools and improved disease models. The work of Yan et al.2 and other studies published this year6–8 suggest that an era of explosive growth in the field is underway.
Footnotes
S.E.B. & Z.P.J. are Co-Directors of the Yerkes Non-Human Primate Genomics Core.
References
- 1.Shedlock DJ, Silvestri G, Weiner DB. Nat Rev Immunol. 2009;9:717–728. doi: 10.1038/nri2636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Yan G, et al. Nat Biotechnol. 2011;29:XX–XX. [Google Scholar]
- 3.Lander ES, et al. Nature. 2001;409:860–921. doi: 10.1038/35057062. [DOI] [PubMed] [Google Scholar]
- 4.Mikkelsen TS, et al. Nature. 2005;437:69–87. [Google Scholar]
- 5.Gibbs RA, et al. Science. 2007;316:222–234. doi: 10.1126/science.1139247. [DOI] [PubMed] [Google Scholar]
- 6.Locke DP, et al. Nature. 2011;469:529–533. doi: 10.1038/nature09687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Fang X, et al. Genome Biol. 2011;12:R63. doi: 10.1186/gb-2011-12-7-r63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ebeling M, et al. Genome Res. 2011;10:1746–56. doi: 10.1101/gr.123117.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kirmaier A, et al. PLoS Biol. 2010;8:e1000462. doi: 10.1371/journal.pbio.1000462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lim SY, et al. PLoS Genet. 2010;6:e1000997. doi: 10.1371/journal.pgen.1000997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Paiardini M, et al. Nat Med. 2011;17:830–836. doi: 10.1038/nm.2395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Preuss TM. In: The Primate Visual System. Kaas JH, Collins CE, editors. CRC Press; 2004. pp. 231–259. [Google Scholar]