Abstract
Hypocrea jecorina (anamorph Trichoderma reesei) is a saprophytic fungus that produces hydrolases, which are applied in different types of industries and used for the production of biofuel. A recombinant Hypocrea strain, which constantly expresses the main transcription activator of hydrolases (Xylanase regulator 1), was found to grow faster on xylan and its monomeric backbone molecule d-xylose. This strain also showed improved ability of clearing xylan medium on plates. Furthermore, this strain has a changed transcription profile concerning genes encoding for hydrolases and enzymes associated with degradation of (hemi)celluloses. We demonstrated that enzymes of this strain from a xylan cultivation favoured break down of hemicelluloses to the monomer d-xylose compared to the parental strain, while the enzymes of the latter one formed more xylobiose. Applying supernatants from cultivation on carboxymethylcellulose in enzymatic conversion of hemicelluloses, the enzymes of the recombinant strain were clearly producing more of both, d-xylose and xylobiose, compared to the parental strain. Altogether, these results point to a changed hydrolase expression profile, an enhanced capability to form the xylan-monomer d-xylose and the assumption that there is a disordered induction pattern if the Xylanase regulator 1 is de-regulated in Hypocrea.
Keywords: Trichoderma reesei, Hypocrea jecorina, Biopolymer degradation, Hydrolases, (Hemi)cellulases, Xyr1
1. Introduction
Hypocrea jecorina (anamorph Trichoderma reesei [1]) is a filamentous ascomycete, which abundantly occurs wherever biomass is available. This fungus is of noteworthy industrial importance, mainly because of its native extracellular enzymes that are employed in different types of industry. These hydrolases achieved a broad area of application covering pulp and paper industry [2], [3], [4], food and feed industry [5], [6], [7], textile industry [8], [9], [10] and most recently, biofuel production [11], [12], [13]. The set of hydrolytic enzymes produced by H. jecorina comprises amongst others two main cellobiohydrolases, CBHI and CBHII (EC 3.2.1.91) (e.g. [14]), endo-β-1,4-glucanases, EGLI to EGLV (EC 3.2.1.4) (e.g. [15]), two major specific endo-β-1,4-xylanases, XYNI and XYNII (EC 3.2.1.8) [16], and one β-xylosidase BXLI (EC 3.2.1.37) [17]. This set of hydrolases is synergistically working together to attain a complete degradation of biopolymeric substrates of which cellulose and xylan are predominant. In this particular decomposition process these enzymes cause hydrolysis to smaller, soluble oligo- and monosaccharides, which finally can act directly as low-molecular weight inducer substances (e.g. xylobiose, d-xylose) [18], [19].
For a certain industrial employment of these enzymes the composition of the cocktail is crucial: some applications demand for cellulases or hemicellulases exclusively, others need exact ratios of cellulases to hemicellulases. In the past, it was demonstrated that a simple deletion or insertion of multiple copies of various hydrolytic enzyme-encoding genes from/into Hypocrea genomes did not substantially alter the composition of produced enzymes [20], [21]. This insight provoked other strategies as the regulation of gene expression of (hemi)cellulases. Consequently, during the last years a major target of research in this field became the transcription factors involved in the regulation of hydrolase expression.
Some years ago, we identified the main transcriptional activator of hydrolase-encoding genes in H. jecorina: Xyr1 (Xylanase regulator 1) [22]. Xyr1 is a central regulatory protein responsible for the activation of the most important hydrolytic enzyme-encoding genes, including cbh1, cbh2, egl1, xyn1, xyn2, and bxl1. It also contributes to the regulation of d-xylose metabolism because it is essential for the activation of d-xylose reductase (Xyl1 [23]) expression [22]. This consequently implies that Xyr1 has also a notable influence on lactose metabolism, which is further supported by an indirect influence on transcription of bga1 (encoding β-galactosidase [24]) [25].
In 2008, different aspects of transcriptional regulation of xyr1 itself have been reported [26]. During this study, a H. jecorina strain, nx7, expressing the xyr1 gene under the H. atroviridis (T. atroviride) nag1 (N-acetyl glucoseaminidase-encoding) promoter was constructed to ensure a constant expression level of xyr1. This strain allows investigations on the effects of a constitutively expressed, de-regulated (e.g. independent from carbon catabolite repression caused by glucose) xyr1 expression [26]. In course of these analyses we found that the nx7 grows faster on media containing xylan as carbon source and that it clears this medium faster and completely compared to the parental strain [26].
In this study, we report in detail on the ability of the nx7 strain to grow on different biopolymers and to form clearing zones on xylan plates. We demonstrate that the constitutive expression of the regulator protein Xyr1 leads to a different transcription pattern of genes encoding for hydrolases and enzymes associated with decomposition of (hemi)celluloses. Consequently, we investigated the degree of brake down of biopolymers to di- and monosaccharides by enzymatic catalysis using supernatants of H. jecorina nx7 in comparison to supernatants of its parental strain.
2. Materials and methods
2.1. Fungal strains
H. jecorina (T. reesei) QM9414 (ATCC 26921) was used as parental strain throughout this study and nx7 strain as a recombinant strain constitutively expressing xyr1 [26]. Both strains were maintained on malt agar.
2.2. Determination of fungal growth and clearing properties
Strains were cultivated at 30 °C on agar plates containing Mandels–Andreotti (MA) medium [27] without peptone, applying 1% (w/v) d-xylose (Sigma, St. Louis, MO), birch wood xylan (Sigma), crystalline cellulose (Merck Schuchardt OHG, München, Germany) or carboxymethylcellulose (CMC) (Calbiochem, part of EMD Chemicals, affiliate of Merck, Darmstadt, Germany) as the sole carbon source. After 2 days agar pieces of 0.5 cm diameter were cut out and placed on another agar plate containing the same carbon source. Then growth diameters and clearing zones (if xylan was used) were measured after 16.5, 24, 42, 48, 69.5, and 88.5 h. Values given are means of three biological replicates, standard deviations are below 5%.
2.3. Cultivation in a bioreactor
Cultivation in a bench top fermenter (Bioengineering AG, Wald, Switzerland) was carried out using 1 litre medium adjusted to pH 4.5 comprising 20 g xylan (Lenzing, AG, Lenzing, Austria) or CMC, 2.8 g (NH4)2SO4, 1 g MgSO4·7H2O, 4 g KH2PO4, 0.5 g NaCl, 0.5 g Tween 80, 0.1 g peptone, 5 mg FeSO4·7H2O, 1.7 mg MnSO4·H2O, 1.4 mg ZnSO4·7H2O, and 2 mg CaCl2·2H2O dissolved/resuspended in distilled water. Some drops glanapon (Becker, Wien, Austria) were added to the medium to avoid excessive foam formation. 108 conidia per litre (final concentration) were used as inoculums. Cultivation was performed at 30 °C, pH 4.5, 0.3 vvm aeration rate and 500 rpm agitation rate until autolysis started. Each sample drawing was followed by a microscopic analysis for infection control. Culture supernatant and mycelia were separated by filtration through a Miracloth filter (Calbiochem). Mycelia were stored in liquid nitrogen for subsequent mRNA extraction.
2.4. RNA-extraction, reverse transcription, qPCR
Harvested mycelia were homogenized in 1 ml peqGOLD TriFast DNA/RNA/protein purification system (PEQLAB Biotechnologie, Erlangen, Germany) using a FastPrep FP120 BIO101 ThermoSavant cell disrupter (Qbiogene, Carlsbad, US). DNA and RNA were simultaneously isolated in a two-step-process according to the manufacturer's instructions.
Synthesis of cDNA from mRNA was carried out applying RevertAid™ H Minus First Strand cDNA synthesis Kit (Fermentas, St. Leon-Rot, Germany) according to the manufacturer's instructions.
All quantitative PCRs (qPCRs) were performed in an iCycler iQ, Real-Time Detection System (Bio-Rad, Herkules, US). The software of the iCycler (iCycler iQ, Optical System Software, Version 3.0a, Bio-Rad) was used to compile PCR protocols and to define plate set-ups. All PCRs were carried out in triplicate in 25 μl reaction mixtures including 1× iQ SYBR Green Supermix (Bio-Rad), 0.1 μM forward primer, 0.1 μM reverse primer and as template cDNA (100-fold diluted). Primers pairs are given in Table 1. Each run included a blank (sterile bi-distilled water instead of sample) and a no-amplification control (0.01% SDS added to the reaction mixture). The following PCR protocols were followed: 3 min initial denaturation at 95 °C, followed by 45 cycles of 15 s at 95 °C, 15 s at 59 °C (for the target genes) or 60 °C (for act) and 15 s at 72 °C. The Optical System Software set the threshold level automatically to noise-to-signal ratio conditions. Results of transcription analyses are given as relative transcript ratios with reference to the actin gene (act).
Table 1.
Primers used throughout this study.
| Name | Sequence (5′–3′) | Employment |
|---|---|---|
| actf | TGAGAGCGGTGGTATCCACG | act qPCR |
| actr | GGTACCACCAGACATGACAATGTTG | act qPCR |
| axe1fw | CTGCGGAGGAGGCGATC | axe1 qPCR |
| axe1r | GGACGGGCGTCGAAACC | axe1 qPCR |
| bga1f | CGTTTGATCCTTTCGGCGGCT | bga1 qPCR |
| bga1r | CCAAAGGTCATGTATATGTTGAAGATGGTC | bga1 qPCR |
| bxl1f | GCCAACTTCGCCACCAAGG | bxl1 qPCR |
| bxl1r | CGGCAATCTGGTGGATCAATGTG | bxl1 qPCR |
| chb1f | GATGATGACTACGCCAACATGCTG | cbh1 qPCR |
| cbh1r | ACGGCACCGGGTGTGG | cbh1 qPCR |
| cbh2f | CTATGCCGGACAGTTTGTGGTG | cbh2 qPCR |
| cbh2r | GTCAGGCTCAATAACCAGGAGG | cbh2 qPCR |
| egl1f | CTGCAACGAGATGGATATCCTGG | egl1 qPCR |
| egl1r | GTAGTAGCTTTTGTAGCCGCTGC | egl1 qPCR |
| glr1fw | GGATACAGCGGCAACCTC | glr1 qPCR |
| glr11r | GTATGCTCAAACCTGGCGGC | glr1 qPCR |
| swo1fw | CTTGCGTCAACCGGCGG | swo1 qPCR |
| swo1r | CGGCAATGCTGGGGTAAGG | swo1 qPCR |
| xorf | CTGTGACTATGGCAACGAAAAGGAG | xyl1 qPCR |
| xorr | CACAGCTTGGACACGATGAAGAG | xyl1 qPCR |
| xyn1f | CAGCTATTCGCCTTCCAACAC | xyn1 qPCR |
| xyn1r | CCAAAGTTGATGGGAGCAGAA | xyn1 qPCR |
| xyn2f | GGTCCAACTCGGGCAACTTT | xyn2 qPCR |
| xyn2r | CCGAGAAGTTGATGACCTTGTTC | xyn2 qPCR |
2.5. Enzyme assay
The 1 mL-assay for the enzymatic conversion of biopolymers was performed in duplicates as follows. 1% (w/v) beech wood xylan (Lenzing), birch wood xylan (Sigma), crystalline cellulose (Merck Schuchardt OHG), carboxymethylcellulose (Calbiochem) or Miscanthus giganteus in 50 mM Na-acetate buffer (pH 4.7) was incubated with 100 μL of the supernatants of the cultivations in a bioreactor using xylan or CMC as a substrate for 0, 20, 40, 60, 120, and 180 min at 40 °C. Afterwards the reaction was stopped by adding 300 μL 2% (w/v) Trizma base (Sigma). Solid residues have been removed by centrifugation at 14,000 × g and 4 °C for 10 min prior to HPLC analysis. The M. giganteus was pre-treated by acidic hydrolysis as described elsewhere for chitin [28]. But instead of washing with water, the pH was adjusted with NaOH until neutral and the M. giganteus suspension, containing NaCl, was dialyzed against water. Supernatants have been adjusted to the same protein concentrations before applied in the assay. Protein concentration was measured using the Bradford assay (Bio-Rad).
2.6. HPLC-analysis
Analyses were performed using a Thermo Finnigan Surveyor HPLC instrument (Thermo Fisher Scientific, MA, US). All 10 μL samples were injected onto a Rezex RHM-Monosaccharide column (H+, 8%, 150 mm × 7.8 mm; Phenomenex, CA, US). Water was used as the mobile phase and isocratic elution was followed at 85 °C, applying a flow rate of 0.6 mL/min for 20 min. The concentration was determined using d-xylose, xylobiose and glucose as a standard.
3. Results and discussion
3.1. De-regulated xyr1 expression leads to faster growth and better degradation of xylan
Xyr1, the main transcriptional activator of hydrolase-encoding genes in H. jecorina [22], was expressed under the nag1 promoter of H. atroviridis to study the effects of a de-regulated (e.g. glucose repression independent) xyr1 expression [26]. It was previously reported, that an according strain, nx7, was able to clear a xylan-containing medium considerably earlier than the parental strain QM9414 [26]. Therefore, we decided to determine growth rates and clearing rates (if appropriate) on different biopolymers. We found that growth rates of nx7 on birch wood xylan and on its monosaccharide d-xylose was elevated compared to QM9414 (Fig. 1A). Accordingly, the clearing zone was formed faster by nx7 (Fig. 1A). In contrast, on carboxymethylcellulose (CMC) the nx7 growth rate was lower than the one from the parental strain. The mycelium of both strains on crystalline cellulose was so thin that it was not possible to measure the growth diameters (data not shown). QM9414 and nx7 also differed in their phenotype on all those carbon sources (Fig. 1B). On CMC, nx7 produced a few, white spores, whereas those of the parental strain were greenish, on d-xylose, nx7 sporulated strongly circular and white, but the parental strain produced green spores and a flashy, yellow metabolite, and on xylan, nx7 sporulation was white to brownish while that from QM9414 was white to greenish and again circular in both strains (Fig. 1B).
Fig. 1.
Growth and clearing properties of H. jecorina QM9414 and nx7. (A) Growth rates of both strains have been determined by cultivation on agar plates containing the indicated carbon source or biopolymer at 30 °C. Additionally, the rate of clearing of the xylan-containing medium was determined by measuring the clearing zone. Values are means of three biological replicates; standard deviation was below 5%. (B) Pictures of H. jecorina QM9414 and nx7 strains on the plates used for determination of growth rates.
3.2. De-regulation of xyr1 expression leads to a modified hydrolase expression pattern in H. jecorina
The observation that nx7 is able to clear a xylan-containing medium importantly faster than the parental strain and that the growth rate is increased as well, prompted us to look at the transcription of genes encoding for enzymes associated with degradation of (hemi)cellulosic biopolymers. Therefore, both strains were cultivated in a bioreactor using xylan as the sole carbon source. When enough mycelium was built, we started to draw samples, which was done after 30, 40 and 48 h of cultivation. After extraction of mRNA, it was reverse transcribed into cDNA, which was used for subsequent quantitative PCR (qPCR) in order to analyse gene expression. The target genes we have chosen are three important cellulase-encoding genes (cbh1, cbh2, and egl1), three genes encoding for the major xylanases (xyn1, xyn2, and bxl1), two genes encoding for side-chain cleaving enzymes: axe1 (encoding for the acetylxylanesterase 1) and glr1 (encoding for the glucuronidase 1), two inducer-providing enzyme-encoding genes (bga1 and xyl1), and an accession-providing enzyme-encoding gene, swo1 (encoding for swollenin [29]). Because the transcript ratios of both strains for all three sample-drawing time points were very similar, we have chosen exemplarily the results from the analysis of the 40 h sample (Table 2). The first column gives a comparison of the transcription of the analysed genes in QM9414. The second column compares transcription levels of a certain gene in nx7 to those in QM9414. Altogether, expression of six genes is up-regulated, that of three is uninfluenced and expression of two genes is down-regulated (Table 2). Interestingly, the expression of the xylanase-encoding genes xyn1 and xyn2 is reduced in nx7, while bxl1 is elevated. It also should be noted that two cellulase-encoding genes (cbh1, egl1) are elevated in the xyr1 de-regulated strain (nx7) even if the cultivation was performed on xylan. Both issues will be addressed later. Notably, we could identify no direct relation between the function of the enzymes for which the genes encode and the extent of changing their transcription (e.g. two cellulases are up-regulated, one is unaffected). Nor we found a clear relation between the circumstance if Xyr1 regulates genes and the extent of their transcription modification (e.g. cbh2 is known to be regulated via Xyr1 [22], but has not changed its transcription level). The latter observation may be explained by the fact that although Xyr1 is an indispensable activator, the constant expression level of it alone is not always enough to change the transcription of a certain gene. Certainly, one should consider that other regulatory mechanisms and even other transcription factors play a role. Summarizing, it can be stated that a quite simple intervention as the de-regulation of the main transcription activator Xyr1 leads to a modified gene expression profile in H. jecorina.
Table 2.
Transcription analysis of genes encoding for enzymes involved in the degradation of (hemi)cellulose in H. jecorina QM9414 and nx7 after 40 h of cultivation on xylan.
| Gene | QM9414a | nx7 compared to QM9414b |
|---|---|---|
| axe1 | ++ | ∼ |
| bga1 | + | ∼ |
| bxl1 | ++ | + |
| cbh1 | +++ | + |
| cbh2 | + | ∼ |
| egl1 | + | + |
| glr1 | +++ | + |
| swo1 | +++ | + |
| xyl1 | ++ | + |
| xyn1 | + | − |
| xyn2 | ++++ | − |
Transcription of given genes compared to xyn1; +, expressed; ++, stronger expressed; +++ importantly stronger expressed.
Transcript ratio of nx7 compared to QM9414 for a certain gene; −, less expressed; ∼, equally expressed; +, stronger expressed.
3.3. H. jecorina nx7 enzymes produced in xylan fermentation favour break down of hemi-celluloses to the monomer d-xylose
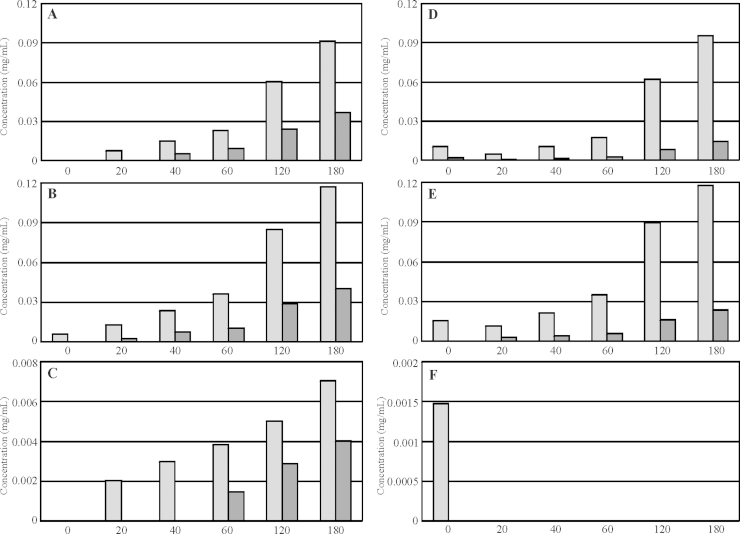
As mentioned above, transcription analysis of nx7 and the parental strain grown on xylan revealed higher expression of bxl1, but lower expression of xyn1 and xyn2 in nx7 compared to QM9414 (Table 2). We were interested if this observation could be confirmed by the enzymatic activity of the fungus. Thus, supernatants from the fermentations of both strains on xylan have been used for enzymatic conversion of hemicellulosic biopolymers. After incubation of the hemicelluloses with the supernatants at 40 °C for 0, 20, 40, 60, 120, 180 min HPLC analyses were done. We found that the decomposition to the monomer d-xylose (Fig. 2A–C) was clearly elevated in the nx7 strain compared to the parental strain, regardless if birch wood xylan (Fig. 2A), beech wood xylan (Fig. 2B) or M. giganteus (Fig. 2C) was used as substrate. Notably, the disaccharide xylobiose was formed in higher amounts applying the QM9414 supernatant, if the substrate was birch wood xylan (Fig. 2D) or beech wood xylan (Fig. 2E), and if M. giganteus was the substrate it was only detectable in small amounts in QM9414 (Fig. 2F). This perfectly fits the transcription analysis results (Table 2), which indicated that bxl1, responsible for enzymatic catalysis of xylobiose to d-xylose, is higher expressed in nx7, whereas xyn1 and xyn2, cleaving the xylan-backbone into oligo- and disaccharides is less expressed in nx7.
Fig. 2.
Analysis of the conversion of biopolymers to mono- and disaccharides via enzymatic catalysis using supernatants from H. jecorina fermentations on birch wood xylan. The biopolymers birch wood xylan (A and D), beech wood xylan (B and E), and M. giganteus (C and F) were incubated with supernatants from H. jecorina nx7 (light grey) and QM9414 (dark grey) for 0, 20, 40, 60, 120, 180 min at 40 °C. Afterwards HPLC-analysis was performed using d-xylose (A–C) and xylobiose (D–F) as standards. Values are means of four replicates (two biological replicates used in duplicates in the enzyme assay); standard deviation was below 5%.
3.4. H. jecorina nx7 enzymes, produced in CMC fermentation, increase the overall degradation of hemicelluloses
The finding that genes encoding for cellulases were stronger expressed in the nx7 strain even if grown on xylan (Table 2), pointed at a de-regulation of induction mechanisms and suggested to examine vice versa the ability of enzymes in supernatants of both strains, grown on CMC, to catalyze degradation of hemicelluloses. Thus, the enzyme assay was performed according to the one using supernatants from cultivation on xylan (vide supra). This time, we found that both d-xylose formation (Fig. 3A–C) and xylobiose formation (Fig. 3D–F) was higher in nx7 whatever hemicellulosic biopolymer was applied: birch wood xylan (Fig. 3A and D), beech wood xylan (Fig. 3B and E) or M. giganteus (Fig. 3C and F). This allows the assumption that the de-regulation of xyr1 also leads to a shifted induction pattern. Although if xylanases and cellulases are always expressed in a coordinate manner on (hemi)celluloses [30], the released oligo- or disaccharides (e.g. xylobiose) favour the induction of certain genes responsible for their formation (e.g. xyn1, and xyn2) (e.g. [31], [32], [33]). In case of the nx7 strain, these inducer specificities seem to be more diffuse, which may result from constitutive expression of xyr1 that mediates all induction signals (received from d-xylose, xylobiose, etc.) [34], [35].
Fig. 3.
Analysis of the conversion of biopolymers to mono- and disaccharides via enzymatic catalysis using supernatants from H. jecorina fermentations on CMC. The biopolymers birch wood xylan (A and D), beech wood xylan (B and E), and M. giganteus (C and F) were incubated with supernatants from H. jecorina nx7 (light grey) and QM9414 (dark grey) for 0, 20, 40, 60, 120, 180 min at 40 °C. Afterwards HPLC-analysis was performed using d-xylose (A–C) and xylobiose (D–F) as standards. Values are means of four replicates (two biological replicates used in duplicates in the enzyme assay); standard deviation was below 5%.
The single xylobiose peak at time point zero in Fig. 3F obviously results from a low amount of xylobiose present in the fermentation supernatant of nx7 and this peak most probably disappears at later time points because the xylobiose from the supernatant is then completely degraded to d-xylose as well as the in the assay decomposed M. giganteus.
3.5. De-regulation of xyr1 expression leads to increased degradation of crystalline cellulose even with enzymes produced in xylan fermentation
As already emphasized, the gene expression analysis of both strains after cultivation on xylan surprisingly showed elevated transcription of cellulase-encoding genes in the xyr1 de-regulated strain (nx7) (Table 2). Therefore, we decided to compare the decomposition rate of crystalline cellulose after enzymatic catalysis using supernatants from fermentation on xylan and CMC of both strains. The enzyme assay was performed according to the previous ones (vide supra), and HPLC analysis was performed using the monomer of cellulose, glucose, as standard. The formation of glucose was equally or even elevated using the supernatant of nx7 grown on xylan (Fig. 4A) compared to the supernatant of nx7 grown on CMC (Fig. 4B). Moreover, the formation of glucose was equally or elevated (for 180 min of incubation time) using enzymes from the xylan fermentation of nx7 compared to those of QM9414 (Fig. 4A). Of course, also the parental strain QM9414 was able to degrade crystalline cellulose to glucose even if supernatant from cultivation on xylan was applied (Fig. 4A), most likely due to the coordinate expression of xylanases and cellulases on (hemi)cellulosic substrates [30]. Nevertheless, glucose formation using supernatant of QM9414 grown on CMC was slightly higher than if that from xylan cultivation was applied (compare Fig. 4B and A). These findings again point at a possible shift in induction patterns if expression of xyr1 is de-regulated.
Fig. 4.
Analysis of the conversion of crystalline cellulose to its monosaccharide via enzymatic catalysis using supernatants from different H. jecorina fermentations. The biopolymer was incubated with supernatants from H. jecorina nx7 (light grey) and QM9414 (dark grey) fermentations on birch wood xylan (A) and CMC (B) for 0, 20, 40, 60, 120, 180 min at 40 °C. Afterwards HPLC-analysis was performed using glucose as standard. Values are means of four replicates (two biological replicates used in duplicates in the enzyme assay); standard deviation was below 5%.
Accordingly, we performed an enzyme assay using CMC as the biopolymer to be converted, catalyzed by supernatants from cultivation of both strains on xylan. Surprisingly, this does never lead to an improved degradation of CMC to glucose applying the supernatant of nx7 compared to the supernatant of the parental strain (Fig. 5A). So, this clearly differs concerning the 180 min incubation time from the previous result using crystalline cellulose as substrate (for better comparison again pictured in Fig. 5B). In contrast to the amorphous CMC, crystalline cellulose rather needs an enzyme improving the accessibility of substrates like swollenin. Therefore, the increased expression of swollenin in nx7 (Table 2), might be a possible explanation for the finding that nx7 supernatant decomposes crystalline cellulose better, but CMC worse than the parental strain. Of course, this is rather a speculation demanding for further proof.
Fig. 5.
Analysis of the conversion of cellulosic biopolymers to its monosaccharide via enzymatic catalysis using supernatants from H. jecorina fermentations on birch wood xylan. The biopolymers CMC (A) and crystalline cellulose (B) were incubated with supernatants from H. jecorina nx7 (light grey) and QM9414 (dark grey) fermentations for 0, 20, 40, 60, 120, 180 min at 40 °C. Afterwards HPLC-analysis was performed using glucose as standard. Values are means of four replicates (two biological replicates used in duplicates in the enzyme assay); standard deviation was below 5%.
4. Conclusions
The intervention in gene regulation by means of changing the expression of a transcription factor, like Xyr1, led to a modified gene expression profile, different inducer specificity, and maybe an improved accessibility of substrates in H. jecorina. This is a quite simple action and has a distinct impact, which recommends itself as an important tool for designing strains that produce requested compositions of hydrolases. A useful extension of this strategy would be the precise engineering of the regulatory proteins (e.g. their permanent activation).
Acknowledgements
This study was supported by a grant from the Austrian Science Fund FWF (P20192-B03) and by a grant from the Vienna University of Technology (“DemoTech”, Innovative Project), which are gratefully acknowledged. M.E. Pucher is recipient of a DOC-fFORTE fellowship from the Austrian Academy of Sciences at the Institute of Chemical Engineering in the Vienna University of Technology. Moreover, we wish to thank COST Action CM0903.
References
- 1.Kuhls K., Lieckfeldt E., Samuels G.J., Kovacs W., Meyer W., Petrini O., Gams W., Borner T., Kubicek C.P. Proc. Natl. Acad. Sci. U.S.A. 1996;93:7755. doi: 10.1073/pnas.93.15.7755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Buchert J., Oksanen T., Pere J., Siika-aho M., Suurnäkki A., Viikari L. In: Harman G.E., Kubicek C.P., editors. vol. 2. Taylor & Francis Ltd.; London, UK: 1998. p. p343. (Trichoderma & Gliocladium). [Google Scholar]
- 3.Noé P C.J., Mora F., Comtat J. J. Wood Chem. Technol. 1986;6:167. [Google Scholar]
- 4.Welt T., Dinus R.J. Prog. Paper Recycl. 1995;4:36. [Google Scholar]
- 5.Lanzarini G., Pifferi P.G. In: Biotechnology Applications in Beverage Production. Cantarelli C., Lanzarini G., editors. Elsevier Science; London: 1989. p. 189. [Google Scholar]
- 6.Walsh G.A., Power R.F., Headon D.R. Trends Biotechnol. 1993;11:424. doi: 10.1016/0167-7799(93)90006-U. [DOI] [PubMed] [Google Scholar]
- 7.Galante Y.M., Monteverdi R., Inama S., Caldini C., De Conti A., Lavelli V., Bonomi F. Ital. Biochem. Soc. Trans. 1993;4:34. [Google Scholar]
- 8.Pedersen G.P., Screws G.A., Cereoni D.A. Can. Text. J. 1992:31. [Google Scholar]
- 9.Kumar A., Lepola M., Purtell C. Text. Chem. Color. 1994;26:25. [Google Scholar]
- 10.Koo H., Ueda M., Wakida T., Yoshimura Y., Igarashi T. Text. Res. J. 1994;64:70. [Google Scholar]
- 11.Hahn-Hägerdal B., Galbe M., Gorwa-Grauslund M.F., Liden G., Zacchi G. Trends Biotechnol. 2006;24:549. doi: 10.1016/j.tibtech.2006.10.004. [DOI] [PubMed] [Google Scholar]
- 12.Himmel M.E., Ding S.Y., Johnson D.K., Adney W.S., Nimlos M.R., Brady J.W., Foust T.D. Science (New York, NY) 2007;315:804. doi: 10.1126/science.1137016. [DOI] [PubMed] [Google Scholar]
- 13.Ragauskas A.J., Williams C.K., Davison B.H., Britovsek G., Cairney J., Eckert C.A., Frederick W.J., Jr., Hallett J.P., Leak D.J., Liotta C.L., Mielenz J.R., Murphy R., Templer R., Tschaplinski T. Science (New York, NY) 2006;311:484. doi: 10.1126/science.1114736. [DOI] [PubMed] [Google Scholar]
- 14.Teeri T., Salovouri I., Knowles J. Biotechnology. 1983;1:696. [Google Scholar]
- 15.Penttilä M., Lehtovaara P., Nevalainen H., Bhikhabhai R., Knowles J. Gene. 1986;45:253. doi: 10.1016/0378-1119(86)90023-5. [DOI] [PubMed] [Google Scholar]
- 16.Törrönen A., Harkki A., Rouvinen J. EMBO J. 1994;13:2493. doi: 10.1002/j.1460-2075.1994.tb06536.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Herrmann M.C., Vrsanska M., Jurickova M., Hirsch J., Biely P., Kubicek C.P. Biochem. J. 1997;321(Pt 2):375. doi: 10.1042/bj3210375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Mach R.L., Seiboth B., Myasnikov A., Gonzalez R., Strauss J., Harkki A.M., Kubicek C.P. Mol. Microbiol. 1995;16:687. doi: 10.1111/j.1365-2958.1995.tb02430.x. [DOI] [PubMed] [Google Scholar]
- 19.Zeilinger S., Mach R.L., Schindler M., Herzog P., Kubicek C.P. J. Biol. Chem. 1996;271:25624. doi: 10.1074/jbc.271.41.25624. [DOI] [PubMed] [Google Scholar]
- 20.Kubicek C.P., Messner R., Gruber F., Mach R.L., Kubicek-Pranz E.M. Enzyme Microb. Technol. 1993;15:90. doi: 10.1016/0141-0229(93)90030-6. [DOI] [PubMed] [Google Scholar]
- 21.Kubicek-Pranz E.M., Gruber F., Kubicek C.P. J. Biotechnol. 1991;20:83. [Google Scholar]
- 22.Stricker A.R., Grosstessner-Hain K., Würleitner E., Mach R.L. Eukaryot. Cell. 2006;5:2128. doi: 10.1128/EC.00211-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Seiboth B., Gamauf C., Pail M., Hartl L., Kubicek C.P. Mol. Microbiol. 2007;66:890. doi: 10.1111/j.1365-2958.2007.05953.x. [DOI] [PubMed] [Google Scholar]
- 24.Seiboth B., Hartl L., Salovuori N., Lanthaler K., Robson G.D., Vehmaanpera J., Penttilä M.E., Kubicek C.P. Appl. Environ. Microbiol. 2005;71:851. doi: 10.1128/AEM.71.2.851-857.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Stricker A.R., Steiger M.G., Mach R.L. FEBS Lett. 2007;581:3915. doi: 10.1016/j.febslet.2007.07.025. [DOI] [PubMed] [Google Scholar]
- 26.Mach-Aigner A.R., Pucher M.E., Steiger M.G., Bauer G.E., Preis S.J., Mach R.L. Appl. Environ. Microbiol. 2008;74:6554. doi: 10.1128/AEM.01143-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Mandels M. Biochem. Soc. Trans. 1985;13:414. doi: 10.1042/bst0130414. [DOI] [PubMed] [Google Scholar]
- 28.Roberts W., Selitrennikoff C. J. Gen. Microbiol. 1988;134:169. [Google Scholar]
- 29.Saloheimo M., Paloheimo M., Hakola S., Pere J., Swanson B., Nyyssonen E., Bhatia A., Ward M., Penttilä M. Eur. J. Biochem. 2002;269:4202. doi: 10.1046/j.1432-1033.2002.03095.x. [DOI] [PubMed] [Google Scholar]
- 30.Hrmova M., Biely P., Vrsanska M. Arch. Microbiol. 1986;144:307. [Google Scholar]
- 31.Margolles-Clark E., Ilmén M., Penttilä M. J. Biotechnol. 1997;57:167. [Google Scholar]
- 32.Rauscher R., Würleitner E., Wacenovsky C., Aro N., Stricker A.R., Zeilinger S., Kubicek C.P., Penttilä M., Mach R.L. Eukaryot. Cell. 2006;5:447. doi: 10.1128/EC.5.3.447-456.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Würleitner E., Pera L., Wacenovsky C., Cziferszky A., Zeilinger S., Kubicek C.P., Mach R.L. Eukaryot. Cell. 2003;2:150. doi: 10.1128/EC.2.1.150-158.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Stricker A.R., Mach R.L., de Graaff L.H. Appl. Microbiol. Biotechnol. 2008;78:211. doi: 10.1007/s00253-007-1322-0. [DOI] [PubMed] [Google Scholar]
- 35.Stricker A.R., Mach R.L. In: Current Advances in Molecular Mycology. Gherbawy Y., Mach R.L., Rai M., editors. Nova Science Publishers, Inc.; NY: 2008. [Google Scholar]