Abstract
Our previous analysis of 65 advanced dental caries lesions by traditional culture techniques indicated that lactobacilli were numerous in the advancing front of the progressive lesion. Production of organic acids by lactobacilli is considered to be important in causing decalcification of the dentinal matrix. The present study was undertaken to define more precisely the diversity of lactobacilli found in this environment and to quantify the major species and phylotypes relative to total load of lactobacilli by real-time PCR. Pooled DNA was amplified by PCR with Lactobacillus genus-specific primers for subsequent cloning, sequencing, and phylogenetic analysis. Based on 16S ribosomal DNA sequence comparisons, 18 different phylotypes of lactobacilli were detected, including strong representation of both novel and gastrointestinal phylotypes. Specific PCR primers were designed for nine prominent species, including Lactobacillus gasseri, L. ultunensis, L. salivarius, L. rhamnosus, L. casei, L. crispatus, L. delbrueckii, L. fermentum, and L. gallinarum. More than three different species were identified as being present in most of the dentine samples, confirming the widespread distribution and numerical importance of various Lactobacillus spp. in carious dentine. Quantification by real-time PCR revealed various proportions of the nine species colonizing carious dentine, with higher mean loads of L. gasseri and L. ultunensis than of the other prevalent species. The findings provide a basis for further characterization of the pathogenicity of Lactobacillus spp. in the context of extension of the carious lesion.
Dental caries continues to be a significant public health problem in many parts of the world. Although the bacteria responsible for caries initiation and early caries progression have been studied extensively, the microbiology of dentine caries has been reported to show considerable diversity and has not yet been fully characterized. Dissolution by acid of the surface enamel exposes the underlying avascular mineralized connective tissue matrix of dentine, which is prone to invasion. This occurs by migration of bacteria into the network of tubules occupied by processes of the pulpal odontoblasts. The early stage of invasion involves lactobacilli, Actinomyces spp., veillonellae, and mutans streptococci (for a review, see reference 19). This phase is followed by the invasion of a more diverse group of microorganisms including gram-negative anaerobes. There is evidence that interspecies cooperation enhances the migration of the mixed bacterial flora through the dentinal tubules (20, 27).
Lactobacilli have been reported to occur in high numbers in both superficial and deep caries (9), though they are not suspected of being involved in bacterial invasion of nonexposed dental pulp (12). Our previous analysis of lactobacilli by culture under microaerophilic conditions in 65 deep caries samples indicated that Lactobacillus acidophilus was numerically dominant, although Lactobacillus paracasei, Lactobacillus rhamnosus, and Lactobacillus fermentum were also present in many samples (22). In the present study, analysis of samples by quantitative molecular techniques indicated a greater abundance and unexpected diversity of lactobacilli, with representation by species that are not commonly found in the oral cavity.
MATERIALS AND METHODS
Bacterial strains.
Lactobacilli (Table 1) were obtained from the Institute of Dental Research collection and the Australian Starter Culture Research Centre (Werribee, Victoria, Australia) and cultured in MRS medium (Oxoid, Basingstoke, United Kingdom). Other bacteria were cultured as described previously (26).
TABLE 1.
Bacterial strains and the specificity of primers for the detection of bacteria by PCR
| Species | Strain | Specificity of primer pair for detection of bacteria
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| UniF, LactoR | LactoF, LactoR | LcaseF, LcaseR | LrhamF, LcaseR | LfermF, LactoR | LactoF, LgassR | LsaliF, LactoR | LcrisF, LcrisR | LgallF, LcrisR | LdelbF, LdelbR | LultF, LcrisR | ||
| Lactobacillus casei subsp. casei | ATCC 393 | + | + | − | − | − | − | − | − | − | − | − |
| Lactobacillus paracasei subsp. paracasei | ATCC 25302 | + | + | + | − | − | − | − | − | − | − | − |
| Lactobacillus rhamnosus | ATCC 7469 | + | + | − | + | − | − | − | − | − | − | − |
| Lactobacillus rhamnosus | CCUG 27772 | + | + | − | + | − | − | − | − | − | − | − |
| Lactobacillus fermentum | ATCC 14931 | + | + | − | − | + | − | − | − | − | − | − |
| Lactobacillus fermentum | ATCC 9338 | + | + | − | − | + | − | − | − | − | − | − |
| Lactobacillus gasseri | ATCC 33323 | + | + | − | − | − | + | − | − | − | − | − |
| Lactobacillus salivarius subsp. salivarius | ATCC 11741 | + | + | − | − | − | − | + | − | − | − | − |
| Lactobacillus salivarius subsp. salicinus | ATCC 11742 | + | + | − | − | − | − | + | − | − | − | − |
| Lactobacillus crispatus | ATCC 33820 | + | + | − | − | − | − | − | + | − | − | − |
| Lactobacillus gallinarum | ATCC 33199 | + | + | − | − | − | − | − | − | + | − | − |
| Lactobacillus delbrueckii subsp. bulgaricus | ATCC 11842 | + | + | − | − | − | − | − | − | − | + | − |
| Lactobacillus delbrueckii | ATCC 9649 | + | + | − | − | − | − | − | − | − | + | − |
| Lactobacillus acidophilus | ATCC 4356 | + | + | − | − | − | − | − | − | − | − | − |
| Lactobacillus brevis | ATCC 14869 | + | + | − | − | − | − | − | − | − | − | − |
| Lactobacillus reuteri | ATCC 23272 | + | + | − | − | − | − | − | − | − | − | − |
| Lactobacillus plantarum | ATCC 10241 | + | + | − | − | − | − | − | − | − | − | − |
| Lactobacillus plantarum | ATCC 14917 | + | + | − | − | − | − | − | − | − | − | − |
| Lactobacillus oris | NCFB 2160 | + | + | − | − | − | − | − | − | − | − | − |
| Lactobacillus oris | NCFB 2164 | + | + | − | − | − | − | − | − | − | − | − |
| Lactobacillus leichmanni | ATCC 4797 | + | + | − | − | − | − | − | − | − | − | − |
| Escherichia coli | XL-1 Blue | +/− | − | |||||||||
| Staphylococcus aureus | ATCC 12600 | + | − | |||||||||
| Pseudomonas aeruginosa | ATCC 19660 | − | − | |||||||||
| Haemophilus influenzae | MSHRIN-1 | − | − | |||||||||
| Streptococcus pyogenes | MSHRPY-1 | +/− | − | |||||||||
| Streptococcus pneumoniae | MSHRPN-1 | − | − | |||||||||
| Moraxella catarrhalis | MSHRCA-1 | − | − | |||||||||
| Porphyromonas gingivalis | ATCC 33277 | − | − | |||||||||
| Fusobacterium nucleatum | ATCC 25586 | +/− | − | |||||||||
| Peptostreptococcus micros | ATCC 33270 | +/− | − | |||||||||
| Serratia marcescens | ATCC 274 | − | − | |||||||||
Carious dentine samples.
The source of material for analysis was the collection of carious dentine from 65 extracted teeth described previously (protocol approved by the Human Ethics Committee of Central Sydney Area Health Service) (22, 26). Patients elected to have extractions for unrestored teeth which presented large coronal dentine caries that, by macroscopic examination, had not penetrated to the underlying vital pulp tissue and had adjacent periodontal probing depths of less than 4 mm. The carious zone of decalcified and partially decalcified dentine in proximity to the advancing front of the lesion in each tooth was excavated, weighed, and resuspended in reduced transport fluid (10 mg [wet weight] of dentine per ml) at 37°C inside an anaerobic chamber. The carious dentine fragments were initially dispersed in reduced transport fluid by vortexing for 20 s, followed by manual homogenization in a 2-ml glass homogenizer for 30 s prior to extraction of bacterial DNA (22).
Underlying pulpal tissue was examined for pathological change and categorized for the predominant presentation of essentially normal histology (category 1), hyaline soft tissue degeneration (category 2), extensive calcification (category 3), or infiltration of inflammatory cells (category 4) (22).
Isolation of DNA.
DNA was isolated from carious dentine as described previously (22) with the ATL buffer reagent (Qiagen, Clifton Hill, Victoria, Australia), which efficiently releases DNA from gram-negative bacteria and from organisms cultured under anaerobic conditions. DNA was extracted from reference Lactobacillus strains (Table 1) with the QIAamp DNA mini kit (Qiagen) according to the manufacturer's instructions.
PCR primers and conditions.
Primers specific for the genus Lactobacillus were designed from regions of identity within the 16S ribosomal DNA (rDNA) sequence from a wide diversity of Lactobacillus spp. (GenBank accession numbers in parentheses): Lactobacillus acetotolerans (M58801), Lactobacillus alimentarius (M58804), Lactobacillus amylolyticus (Y17361), Lactobacillus amylophilus (M58806), Lactobacillus animalis (M58807), Lactobacillus aviarius (M58808), Lactobacillus bifermentans (M58809), Lactobacillus brevis (M58810), Lactobacillus buchneri (M58811), Lactobacillus casei (AY196975), Lactobacillus collinoides (AB005893), Lactobacillus crispatus (AF257097), Lactobacillus delbrueckii (AJ414691), L. fermentum (AF302116), Lactobacillus fructivorans (M58818), Lactobacillus gallinarum (AJ417737), Lactobacillus gasseri (AF519171), Lactobacillus iners (Y16329), Lactobacillus jensenii (AF243176), Lactobacillus lactis (M58823), Lactobacillus lindneri (X95423), Lactobacillus manihotivorans (AF000162), Lactobacillus mucosae (AF126738), Lactobacillus nagelii (Y17500), Lactobacillus oris (X94229), Lactobacillus perolens (Y19168), Lactobacillus plantarum (AL935253), Lactobacillus pontis (X76329), Lactobacillus reuteri (L23507), L. rhamnosus (AF243146), Lactobacillus sakei (M58829), Lactobacillus salivarius (AF089108), Lactobacillus sharpeae (M58831), Lactobacillus vaginalis (AF243177), and Lactobacillus zeae (D86516).
Sequences were retrieved from GenBank and aligned with clustal w (35) together with sequences from the taxonomically related bacteria Bacillus subtilis (AB016721), Staphylococcus aureus (SA16SRRN), Listeria monocytogenes (S55472), Clostridium botulinum (CBA16S), Peptostreptococcus micros (PEP16SRR8), Streptococcus mutans (SM16SRNA), Enterococcus faecalis (AB012212), and Pediococcus acidilactici (X95976). The sequences of selected Lactobacillus-specific primers LactoF and LactoR are shown in Table 2. The specificity of the primer sequences was determined by BLAST (1) homology searches for short, nearly exact matches in GenBank. BLAST was accessed through the Australian National Genomic Information Service (ANGIS; http://www.angis.org.au). The specificity of the primers was confirmed by PCR on DNA templates from taxonomically related and unrelated bacteria in 25-μl reaction mixtures containing 1× HotStarTaq Master mix (Qiagen), 2 μl of template (≈40 ng of DNA), and 100 nM (each) primer. PCR was performed with the GeneAmp PCR System 9700 (Perkin Elmer, Wellesley, Mass.) with an initial denaturation step of 95°C for 15 min, followed by 40 cycles of 95°C for 15 s and 62°C for 1 min. A 10-μl aliquot of the PCR was subjected to electrophoresis on a 2% agarose gel containing ethidium bromide, and the DNA bands were visualized by UV illumination.
TABLE 2.
PCR primers
| Primer | Sequence (5′-3′) | Specificity | Locationa (5′-3′) | Region of 16S rRNAb | Annealing temp (°C) | Amplicon sizec (bp) |
|---|---|---|---|---|---|---|
| UniF | GAGAGTTTGATCCTGGCTCAG | Most bacteria | 7-27 | 5′-V1 | 60 | 391-406 |
| LactoF | TGGAAACAGRTGCTAATACCG | All lactobacilli | 157-167 | V2.1-V2.2 | 62 | 231-233 |
| LactoR | GTCCATTGTGGAAGATTCCC | All lactobacilli | 379-360 | V2.2-V3 | ||
| LcaseF | GCACCGAGATTCAACATGG | L. casei/L. paracasei | 86-104 | V1 | 60 | 117 |
| LrhamF | TGCTTGCATCTTGATTTAATTTTG | L. rhamnosus | 81-104 | V1 | 62 | 122 |
| LcaseR | GGTTCTTGGATYTATGCGGTATTAG | L. casei group | 196-169 | V2.2 | ||
| LfermR | GCACCTGATTGATTTTGGTCG | L. fermentum | 86-94 | V1 | 60 | 317 |
| LdelbF | GGRTGATTTGTTGGACGCTAG | L. delbrueckii | 92-102 | V1 | 60 | 138 |
| LdelbR | GCCGCCTTTCAAACTTGAATC | L. delbrueckii | 219-199 | V2.2 | ||
| LgassR | CAGTTACTACCTCTATCTTTCTTCACTAC | L. gasseri | 470-442 | V3 | 60 | 322 |
| LsaliF | CGAAACTTTCTTACACCGAATGC | L. salivarius | 67-91 | V1 | 60 | 332 |
| LcrisF | AGCGAGCGGAACTAACAGATTTAC | L. crispatus | 65-89 | V1 | 60 | 154 |
| LgallF | CGGTAATGACGCTGGGGAC | L. gallinarum | 78-97 | V1 | 64 | 128 |
| LultF | GAGCGGAACCAGCAGATCTG | L. ultunensis | 68-87 | V1 | 60 | 151 |
| LcrisR | AGCTGATCATGCGATCTGCTT | L. acidophilus groups A1-A4d | 205-185 | V2.2 | ||
| T7 primer | TAATACGACTCACTATAGGG | T7 promoter |
Locations relative to positions in the E. coli 16S rRNA gene.
Variable regions according to Neefs et al. (28).
Theoretical amplicon sizes based on the reference sequence for that species; for L. gasseri, L. salivarius, and L. fermentum, either LactoF or LactoR was the complementary primer.
L. acidophilus groups according to Johnson et al. (15).
Real-time quantification of total Lactobacillus load in carious dentine.
Quantitative PCRs were performed in a reaction volume of 25 μl containing 1× SYBR Green PCR Master Mix (Applied Biosystems, Foster City, Calif.), 100 nM each of the LactoF and LactoR primers, and 2 μl of DNA extracted from the carious dentine samples. The amount of DNA in the 65 carious dentine samples was determined in triplicate, and the mean values were calculated. Amplification and detection of DNA were performed with the ABI-Prism 7700 sequence detection system (Applied Biosystems) with optical grade 96-well PCR plates and optical caps. The reaction conditions were 50°C for 2 min and 95°C for 10 min, followed by 40 cycles of 95°C for 15 s and 62°C for 1 min. Data analysis was conducted with Sequence Detection Software version 1.6.3, supplied by Applied Biosystems.
Purified genomic DNA in the range 10 fg to 1 ng of Lactobacillus delbrueckii subsp. bulgaricus (ATCC 11842) was used as the standard for determining the amount of Lactobacillus DNA by real-time PCR. This was equivalent to approximately 4.0 to 4.0 × 105 copies of the genome (genome size of 2.3 Mb). DNA concentrations were determined with the PicoGreen double-stranded DNA quantitation kit (Molecular Probes, Eugene, Oreg.) and Luminescence spectrometer model LS 50B (Perkin Elmer).
Enumeration of Lactobacillus species.
Species- or phylotype-specific primers were designed from either the V1 or V2 variable region (28) from the sequence alignment of the above-mentioned Lactobacillus sequences together with representatives of the major phylotypes identified from the Lactobacillus diversity profile. In most cases, a specific complementary primer could not be designed, and either the LactoF or LactoR primer was used (Table 2). For L. gasseri, the reverse primer was designed from the V3 region. Specificity was checked by BLAST analysis of the sequence databases and confirmed by PCR (Table 1). PCR primers could not be designed to differentiate Lactobacillus ultunensis and the oral Lactobacillus clone represented by L5 (Fig. 1), as their 16S rDNA amplicon sequences differed on average by only 1.5%. A common forward primer, LultF, was therefore designed to be specific for both of these species or phylotypes.
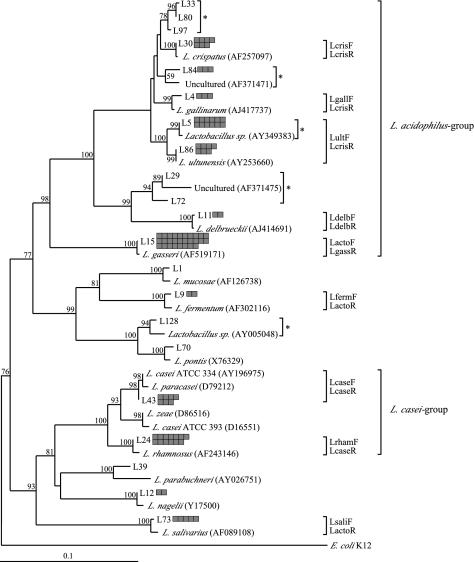
FIG. 1.
Neighbor-joining tree of the Lactobacillus species and phylotypes identified from 58 carious dentine samples, based on 100 sequences of the Lactobacillus-specific 16S rRNA amplicon (≈400 bp) and closely related amplicon regions from Lactobacillus 16S rRNA gene sequences in GenBank (accession numbers in parentheses). Sequences were aligned with clustalw, and the distance matrix was calculated with the Jukes-Cantor algorithm. Shaded boxes adjacent to the representative sequence correspond to the number of clones of the different Lactobacillus spp. or phylotypes identified. Lactobacillus spp. or phylotypes quantified by real-time PCR in the carious dentine samples are marked by the species-specific primer pairs used. Novel phylotypes (*) were defined to differ by >2% in sequence comparisons of their 16S rDNA amplicons from their closest related species. Bootstrap values (>50) near the nodes are represented as percentages of 100 replicates. The phylogenetic tree was rooted with the appropriate amplicon region of the 16S rRNA gene sequence of E. coli K-12 strain MG1655. The scale bar represents the genetic distance.
PCR primers were designed to be optimal for real-time PCR and for the amplification of sequences within the size range detectable with this system. Real-time PCR analysis was conducted with the SYBR Green PCR Master Mix (Applied Biosystems) with the ABI-Prism 7700 sequence detection system (Applied Biosystems). For the quantification of L. casei (including Lactobacillus paracasei), L. crispatus, L. delbrueckii, L. fermentum, L. gallinarum, L. gasseri, L. rhamnosus, and L. salivarius, purified genomic DNA from strains Lactobacillus paracasei subsp. paracasei (ATCC 25302), L. crispatus (ATCC 33820), L. delbrueckii subsp. bulgaricus (ATCC 11842), L. fermentum (ATCC 14931), L. gallinarum (ATCC 33199), L. gasseri (ATCC 33323), L. rhamnosus (ATCC 7469), and Lactobacillus salivarius subsp. salivarius (ATCC 11741), respectively, were used as standards in 10-fold dilution series in the range from 10 fg to 1 ng DNA. For the quantification of L. ultunensis and its related phylotype, purified plasmid DNA containing the appropriate L. ultunensis 16S rDNA insert was used as the standard in a 10-fold dilution series in the range from 100 ag to 10 pg of DNA. The standard DNA concentrations were determined with the PicoGreen double-stranded DNA quantitation kit (Molecular Probes) as described above.
Calculation of bacterial cell numbers.
Conversion of the amount of Lactobacillus DNA in the carious dentine samples determined by real-time PCR to theoretical genome equivalents required the assumption that the genome size and 16S rRNA gene copy number for all lactobacilli was similar. From the review by Klaenhammer et al. (16) of current and completed Lactobacillus genomic sequencing projects, the average genome size for lactobacilli commonly found in the oral cavity of humans is estimated to be 2.2 Mb, so that each cell contains approximately 2.4 fg of DNA.
Identification of Lactobacillus 16S rDNA amplicons.
DNAs from 58 of the 65 carious dentine samples were diluted in sterile H2O to contain 20 pg of Lactobacillus DNA μl−1 and pooled. Seven samples containing <50 pg of Lactobacillus DNA (mg [wet weight] of dentine)−1 were excluded.
PCR of the pooled carious dentine samples was performed with primers UniF and LactoR (Table 2) as described above except that 30 cycles of amplification and 50-μl reaction volumes were used. Aliquots (10 μl) from four independent PCRs were verified by electrophoresis with 2% agarose gels containing ethidium bromide, followed by visualization under UV illumination to confirm the generation of ≈400-bp amplicons. Amplified DNA was pooled, purified with the UltraClean PCR Clean-up kit (Mo Bio Laboratories, Carlsbad, Calif.), and ligated into linearized pGEM-T Easy vectors (Promega, Sydney, New South Wales, Australia), according to the manufacturer's protocol.
Transformation was done with electrocompetent Escherichia coli XL-1 Blue cells and plated onto Luria-Bertani agar plates supplemented with 25 μg of ampicillin ml−1, 30 μg of 5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside (X-Gal) ml−1 and 20 μg of isopropyl-β-d-thiogalactopyranoside (IPTG) ml−1 and incubated overnight at 37°C. White transformants were picked randomly, transferred to Luria-Bertani medium in either 15-ml tubes or 2.2-ml deep-well plates, and grown with shaking for 18 h at 37°C. Plasmid DNA was extracted either individually with a Wizard Plus SV miniprep kit (Promega) or in 96-well blocks with the Perfectprep Plasmid 96 Vac Direct Bind system (Eppendorf, North Ryde, New South Wales, Australia) with a Perfect Vac manifold (Eppendorf). Purified plasmids containing the cloned 16S rDNA amplicons were sequenced by cycle sequencing at the Westmead DNA Sequencing Facility at Westmead Hospital, Wentworthville, New South Wales, Australia. All clones were sequenced with the T7 promoter sequence primer in order to provide full coverage of the ≈400-bp insert.
Sequence analysis of lactobacillus 16S rDNA amplicons.
Sequences were compared with the 16S rRNA gene sequences in the Ribosomal Database Project (6) and in GenBank to identify related sequences in the databases. Sequences which were found to have ≥99% identity were grouped, and representative sequences were aligned with the 16S rDNA sequences of closely related Lactobacillus spp. in GenBank with CLUSTALW (35). The distance matrix was calculated with DNADIST with the Jukes-Cantor model (15), and the phylogenetic tree was constructed by the neighbor-joining method of Saitou and Nei (31) with NEIGHBOR. The corresponding region of the 16S rDNA sequence of E. coli was used to root the phylogenetic tree. Phylogenetic data were subjected to bootstrap analysis of 100 replicates with SEQBOOT and consense, accessed through ANGIS.
Possible chimeric structures among the sequences that were identified by the program chimera_check, accessed through the Ribosomal Database Project (6), or by visual detection of anomalous clustering in the phylogenetic tree, were excluded from the analysis.
Statistical analyses.
Correlations between Lactobacillus sp. and between lactobacilli and pulp tissue responses were determined by a nonparametric Spearman test. Differences were analyzed by nonparametric analysis with the Freidman test, followed by Dunn's post test for comparison of multiple paired samples.
RESULTS
Specificity of LactoF and LactoR primers.
The specificity of the Lactobacillus genus-specific primers was evaluated by PCR with DNA purified from the strains listed in Table 1. For all 21 Lactobacillus strains tested, a ≈230-bp PCR product was obtained. When tested against non-Lactobacillus strains, no amplicons were generated at an annealing temperature of 62°C, confirming the specificity of the primers. LactoF and LactoR are almost identical in sequence to the primary Lactobacillus primers designed by McOrist et al. (25) for the detection of lactobacilli in human fecal samples and therefore are also predicted to exclude DNA from bacteria commonly isolated from the gastrointestinal tract of humans.
Quantification of total Lactobacillus DNA by real-time PCR.
Lactobacillus DNA was found in all 65 carious dentine samples, and quantification by real-time PCR found that it ranged from 25 pg to 359 ng (mg [wet weight] of dentine)−1 (Table 3). When converted to theoretical cell numbers, these values corresponded to levels of lactobacilli in the carious lesions ranging from 1.0 × 104 to 1.4 × 108 cells (mg [wet weight] of dentine)−1 (Table 3). Compared with the number of Lactobacillus CFU for each dentine sample (22), the values obtained with real-time PCR were, on average, 34-fold higher.
TABLE 3.
Lactobacilli detected in carious dentine by real-time PCR
| Species | No. of positive samples (%) | DNA (pg [mg(wet wt) of dentine]−1)
|
Cell equivalentsa (mg [wet wt] of dentine)−1
|
||||
|---|---|---|---|---|---|---|---|
| Range | Median | Mean ± SEM | Range | Median | Mean ± SEM | ||
| Total lactobacilli | 65 (100) | 25-3.6 × 105 | 1.5 × 104 | 4.7 × 104 ± 7.0 × 104 | 1.0 × 104-1.4 × 108 | 5.9 × 106 | 1.9 × 107 ± 2.8 × 107 |
| L. rhamnosus | 35 (54) | 2.0 × 102-3.4 × 104 | 4.1 × 103 | 7.6 × 103 ± 8.8 × 103 | 7.6 × 104-1.3 × 107 | 1.5 × 106 | 2.9 × 106 ± 3.4 × 106 |
| L. casei/L. paracasei | 26 (40) | 4.0-2.2 × 104 | 6.4 × 102 | 2.7 × 103 ± 5.1 × 103 | 1.4 × 103-7.6 × 106 | 2.2 × 105 | 9.6 × 105 ± 17.6 × 105 |
| L. fermentum | 14 (22) | 1.5 × 102-4.7 × 104 | 9.3 × 102 | 4.7 × 102 ± 12.4 × 103 | 6.3 × 104-2.0 × 107 | 3.9 × 105 | 1.9 × 106 ± 5.1 × 106 |
| L. delbrueckii | 4 (6) | 1.0-8.8 × 104 | 5.9 × 10 | 2.1 × 104 ± 3.7 × 104 | 3.8 × 102-3.5 × 107 | 2.2 × 104 | 8.4 × 106 ± 14.8 × 106 |
| L. gasseri | 35 (54) | 6.1 × 10-1.6 × 105 | 7.4 × 103 | 1.7 × 104 ± 3.0 × 104 | 3.1 × 104-8.0 × 107 | 3.8 × 106 | 8.7 × 106 ± 15.2 × 106 |
| L. salivarius | 39 (60) | 1.3 × 10-5.3 × 104 | 2.1 × 103 | 9.2 × 103 ± 12.5 × 103 | 5.5 × 103-2.2 × 107 | 8.7 × 105 | 3.8 × 106 ± 5.2 × 106 |
| L. crispatus | 29 (45) | 1.4 × 10-3.9 × 104 | 2.1 × 103 | 5.9 × 103 ± 9.0 × 103 | 5.9 × 103-1.6 × 107 | 8.5 × 105 | 2.4 × 106 ± 3.7 × 106 |
| L. gallinarum | 6 (9) | 1.4 × 102-2.6 × 104 | 3.3 × 103 | 8.4 × 103 ± 10.8 × 103 | 5.7 × 104-1.1 × 107 | 1.4 × 106 | 3.5 × 106 ± 4.5 × 106 |
| L. ultunensis/Lactobacillus sp. oral clone | 33 (51) | 5.3 × 10-1.8 × 105 | 3.2 × 103 | 2.0 × 104 ± 3.5 × 104 | 2.2 × 104-7.4 × 107 | 1.3 × 106 | 8.1 × 106 ± 14.4 × 106 |
The genome sizes assumed for calculating cell numbers were as follows: L. casei, 2.6 Mb; L. rhamnosus, 2.4 Mb; L. fermentum, L. salivarius, L. crispatus, L. gallinarum and L. ultunensis, 2.2 Mb; L. gasseri, 1.8 Mb; and L. delbrueckii, 2.3 Mb.
Diversity of lactobacilli in carious dentine.
In contrast to the traditional culture methods used to define species, 16S rDNA amplicon sequences can determine the similarity between phylotypes or species when the sequences of the 16S rDNA amplicons differ by ≤1%. Thus, the phylogenetic inferences among the 100 Lactobacillus 16S rDNA amplicon sequences obtained from carious dentine could be compared with the equivalent regions of reference 16S rDNA sequences of known species and phylotypes (Fig. 1). However, since the analysis was based on a ≈400-bp sequence comprising the V1 and V2 regions of the 16S rRNA gene, the phylogenetic tree only provides an indication of the diversity and relationships of the Lactobacillus spp. found in carious dentine samples. The phylogenetic distances between the species or phylotypes may not be truly representative of the genetic distances between the species if the entire 16S rRNA gene sequences had been used for the analysis (13).
Considerable diversity was displayed among the 100 Lactobacillus 16S rDNA amplicon sequences. Based on the definition that 16S rDNA sequences of phylotypes differ from one another by ≥2% of nucleotide sites within the amplified region, 18 different phylotypes were identified. Among these, 12 phylotypes showed ≥99% identity with the 16S rDNA amplicon sequences of known lactobacilli. The remaining phylotypes were either novel species or showed relatedness to uncharacterized lactobacilli from various environments.
Among the 100 Lactobacillus 16S rDNA amplicon sequences, L. gasseri was the most frequently identified species, represented by 28 randomly selected clones. This was followed by L. rhamnosus (13 clones) and a possible novel oral Lactobacillus sp., L5, represented by 12 clones. Other species that were also identified at higher frequency were L. crispatus, L. casei, L. ultunensis (each represented by seven clones), and L. salivarius (five clones).
Novel Lactobacillus phylotypes in carious dentine.
Several sequences were found on multiple occasions among the sequenced clones that were either novel species or phylotypes (Fig. 1) or showed high identity to sequences previously identified from sites other than the human oral cavity. The third most frequently isolated 16S rDNA amplicon, that of the novel Lactobacillus phylotype represented by L5 (12 clones), showed a single nucleotide difference over a 393-bp sequence from an uncultured Lactobacillus sp. identified in the human oral cavity (GenBank accession no. AY349383) (B. J. Paster, unpublished data). Closely related is the 391-bp 16S rDNA amplicon from Lactobacillus phylotypes represented by L86 (7 clones), which was identical to the 16S rDNA sequence from L. ultunensis, which was recently isolated from human stomach mucosa (GenBank accession no. AY253660) (S. Roos, unpublished data).
Another novel phylotype in the L. acidophilus group included three clones represented by L84 (Fig. 1), which was most closely related to an uncultured bacterium from the pig gastrointestinal tract (18) and probably represented a novel species. Within 395 bp, these sequences differed by only 2% on average but were 3% different from the most closely related 16S rDNA, that of L. gallinarum.
Additionally, within the L. acidophilus group were clones L33, L80, and L97 (Fig. 1), whose 16S rDNA amplicon sequences differed on average by 0.7%. These three clones may also represent a novel species, since their 16S rDNA amplicon sequences differed by 2% from that of L. crispatus, the closest related known species, a level of difference sufficient to classify a species within the L. acidophilus group.
Distribution of major species of lactobacilli in carious dentine samples.
A qualitative screen of the 65 carious dentine samples by PCR analysis with species-specific primer sets (Table 2) showed that in most dentine samples, at least three different species or phylotypes were present. The qualitative screen revealed that members of the L. casei group, which includes L. casei, L. paracasei, and L. rhamnosus, were the most prevalent species, being present in 68% of the samples. L. rhamnosus and L. casei/L. paracasei were found in 54 and 40% of samples, respectively. In decreasing order, other prevalent species or phylotypes identified in the dentine were L. salivarius (60%), L. gasseri (54%), L. ultunensis and related phylotype (52%), and L. crispatus (45%). Less prevalent were L. fermentum (22%), a heterofermentative species frequently associated with the initiation of dental caries (4, 21, 33), L. gallinarum (9%), and L. delbrueckii (6%).
Enumeration of Lactobacillus species and phylotypes.
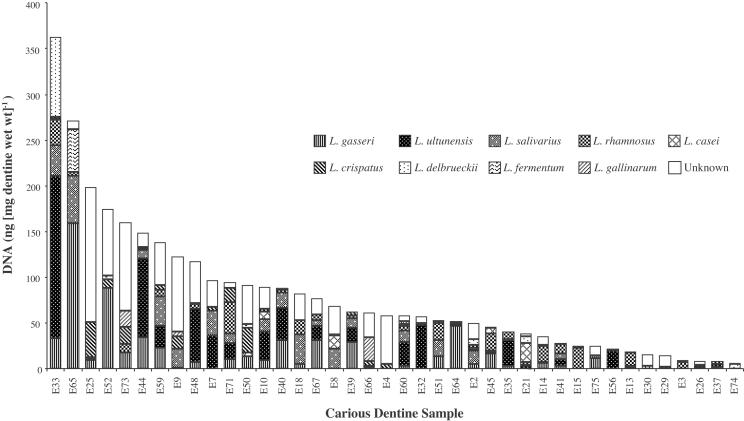
Among the more prevalent species identified by population analysis of pooled DNA (Fig. 1), real-time PCR analysis showed that L. gasseri and L. ultunensis (and its related phylotype) were present in higher numbers than the other species, with mean loads of 8.7 × 106 and 8.1 × 106 cells (mg [wet weight] of dentine)−1, respectively (Table 3). L. gasseri and L. ultunensis also comprised the highest loads of any lactobacilli in any sample, with levels as high as 8.0 × 107 and 7.4 × 107 cells (mg [wet weight] of dentine)−1, respectively (Table 3), suggesting an association between these species and advanced dental caries. In several carious dentine samples, these two prevalent species were also estimated to constitute the majority of the Lactobacillus spp. present. L. ultunensis (and its related phylotype) were found to constitute 84 and 68% of the total Lactobacillus load in samples E32 and E35, respectively (Fig. 2). In sample E56, L. ultunensis constituted approximately 90% of the total Lactobacillus load. Similarly, L. gasseri was found to constitute 91 and 58% of the total Lactobacillus load in samples E64 and E65, respectively (Fig. 2).
FIG. 2.
Graphic representation of the DNA loads of the nine predominant Lactobacillus species or phylotypes in carious dentine determined by real-time PCR. Forty samples with the highest total Lactobacillus loads are shown. The difference in the total Lactobacillus load with the cumulative total of the nine species in each sample (referred to as unknown) was attributed to the presence of other, less-prevalent Lactobacillus spp. that were not quantified by real-time PCR. Patient samples are indicated on the x axis.
The mean loads of the other prevalent species varied from 9.5 × 105 cells (mg [wet weight] of dentine)−1 for L. casei to 3.8 × 106 cells (mg [wet weight] of dentine)−1 for L. salivarius (Table 3). Although the less prevalent species L. fermentum, L. gallinarum, and L. delbrueckii were present in fewer samples, when they were found, they constituted comparable loads.
Associations between Lactobacillus load and specific Lactobacillus species and the histopathology of pulpitis.
The pulps from the 65 carious teeth were divided into four histopathological categories on the basis of the dominant pathology described previously (22). In each of the four histopathological categories, multivariate analyses were performed on the mean values of DNA loads estimated by real-time PCR for the total lactobacilli and for the different species with the Freidman test. For the total load of lactobacilli, no statistically significant relationships with histological category of pulp response were observed. Similarly, no significant relationships between the different Lactobacillus spp. and histological response were detected.
Possible relationships between pairs of Lactobacillus spp. in each carious dentine sample were determined with the Spearman correlation (Table 4). For most of the associations, the correlation coefficients (r) were low and insignificant (P > 0.05). However, analysis revealed multiple positive associations between L. gasseri and L. salivarius with the other Lactobacillus spp. examined. Several of these associations were found to be significant, e.g., L. gasseri and L. salivarius (r = 0.5434, P < 0.0001) and L. gasseri and L. rhamnosus (r = 0.5313, P < 0.0001), indicating that the presence of one of these species in the carious site was associated with the colonization and proliferation of the other species.
TABLE 4.
Correlations between Lactobacillus species within the carious dentine
| Lactobacillus sp. | Microbial associationa
|
|||
|---|---|---|---|---|
|
L. gasseri
|
L. salivarius
|
|||
| r | P | r | P | |
| L. casei | 0.1638 | NS | 0.2075 | NS |
| L. rhamnosus | 0.5313 | <0.0001 | 0.375 | 0.0021 |
| L. crispatus | 0.4078 | 0.0007 | 0.3003 | 0.0151 |
| L. ultunensis | 0.3027 | 0.0142 | 0.3687 | 0.0025 |
| L. gasseri | 0.5434 | <0.0001 | ||
| L. salivarius | 0.5434 | <0.0001 | ||
| L. fermentum | 0.4305 | 0.0003 | 0.323 | 0.0087 |
| L. delbrueckii | 0.08768 | NS | 0.1425 | NS |
| L. gallinarum | 0.1774 | NS | 0.05456 | NS |
r, correlation coefficients were calculated with the Spearman test. NS, not significant (P > 0.05).
When pairs of Lactobacillus spp. within the four histopathological categories were analyzed separately, several significant relationships were revealed (Table 5), indicating that positive associations between certain Lactobacillus spp. are correlated with changes in disease progression. Interestingly, in all histopathological states, a positive relationship between L. gasseri and another species was observed.
TABLE 5.
Correlations between Lactobacillus species and histopathological categoriesa
| Histological category | Microbial association | r | P |
|---|---|---|---|
| Minimal inflammatory change | L. gasseri, L. salivarius | 0.6729 | <0.0001 |
| L. salivarius, L. rhamnosus | 0.5124 | 0.0053 | |
| L. gasseri, L. rhamnosus | 0.5083 | 0.0057 | |
| Soft tissue degenerative change | L. rhamnosus, L. gasseri | 0.6553 | 0.0023 |
| L. rhamnosus, L. ultunensis | 0.6375 | 0.0033 | |
| Hard tissue degenerative change | L. fermentum, L. salivarius | 0.8748 | 0.0123 |
| L. fermentum, L. rhamnosus | 0.8669 | 0.0123 | |
| L. gasseri, L. crispatus | 0.7952 | 0.0341 | |
| Inflammatory degenerative change | L. gasseri, L. crispatus | 0.7075 | 0.0149 |
| L. gasseri, L. ultunensis | 0.6392 | 0.0342 |
Correlation coefficients were calculated with the Spearman test.
Presence of other lactobacilli in carious dentine.
Comparisons of the total Lactobacillus DNA with the cumulative contribution from individual species showed that there was a significant component of Lactobacillus load that remained unidentified in several samples (Fig. 2). Although the differences in the amount of Lactobacillus DNA could be due to differences in the genome sizes and/or the copy number of 16S rRNA genes relative to that in L. delbrueckii subsp. bulgaricus, which was used as the standard, there was a substantial proportion of undetected species in several of the lesions, such as samples E25 and E73, where 74 and 61%, respectively, of the total Lactobacillus load remained unaccounted for (Fig. 2). Random cloning and sequencing of 16S rDNA amplicons in the samples that contained high proportions of unknown lactobacilli revealed the presence of species of lactobacilli other than the predominant species determined from pooling the carious dentine samples (data not shown). In the case of sample E25, for example, in which 74% of the total Lactobacillus load was unidentified, 67% of the 16S rDNA amplicon sequences showed identity with the phylotype cluster represented by Lactobacillus clones L33, L80, and L97 in the L. acidophilus group (Fig. 1). Other species present in these carious dentine samples but not identified among the sequences of the 100 16S rDNA amplicons were Lactobacillus panis, Lactobacillus colehominis, and Lactobacillus nagelii (unpublished data). It appears that these species and the other previously identified species that were not quantified by real-time PCR most likely constitute the unidentified lactobacilli in these carious dentine samples.
DISCUSSION
Compared with lactobacillus counts determined by culture-based methods (22), the values reported in this study determined by real-time PCR were considerably higher, as found previously with the use of this enumeration technique (22, 26). The molecular approaches used in this study also led to the identification of 18 different phylotypes of lactobacilli. Of these, only 12 showed relatedness to known Lactobacillus spp., with only about half of these being similar to species previously isolated from the human oral cavity (2-4, 9, 33). The remaining phylotypes were novel or related to uncultured bacteria. Many of these novel phylotypes were found in significant numbers, indicating their possible association with the progression of the disease. For example, 30% of the clones belonged to the L. acidophilus group and were either new species or phylotypes or similar to species not associated with the human oral cavity, such as clones L84 and L29/L72, which were related to uncultured bacteria identified in the gastrointestinal tract of pigs (18). The failure to detect these lactobacilli in carious dentine by culture-based techniques could simply reflect the difficulties encountered in distinguishing between species of this group with standard biochemical tests (8). Alternatively, members of this group may have additional nutrient requirements that impede their cultivation by conventional methods.
Although the PCR amplification and cloning technique used in this study may bias the reflection of microbial diversity (38), the fact that individual dentine samples were diluted to contribute equal quantities of Lactobacillus DNA to a pooled sample prior to phylogenetic analysis meant that the numerical frequency of the identified species in the clone library reflected their overall numerical predominance within the pooled carious dentine samples. Thus, L. gasseri, L. rhamnosus, and a novel Lactobacillus phylotype (represented by clone L5) were the most numerically dominant species within the 65 carious dentine samples. With classic culture-based methods, L. casei, L. rhamnosus, and L. acidophilus are the most frequently isolated species (4, 9, 23, 32, 33).
Of particular note was that neither L. brevis nor L. plantarum was identified among the 100 16S rDNA amplicons that were analyzed even though these two species were previously isolated in significant numbers (2, 5, 9, 11, 33). Other species cultured from carious dentine include Lactobacillus coryniformis, L. sakei, L. lactis, and L. fermentum (9), whereas those identified at low frequency in the current study showed phylogenetic relatedness to lactobacilli not typically found in the human oral cavity. These included phylotypes isolated from the gastrointestinal tract of animals (30), sourdough (37), maize silage (17), and semifermented wine (7). An exception was clone L128, which was related to the oral clone CX036, obtained from human subgingival plaque (29). Whether the presence of numerous Lactobacillus spp. in pooled carious dentine reflects the diverse dietary habits of the patients (10, 34) remains to be determined. Some of these species were poorly represented in the pooled sample, suggesting their presence in a small number of individual carious dentine samples.
Among the more prevalent lactobacilli identified by population analysis, at least three different species or phylotypes were found in the majority of the 65 carious dentine samples (Fig. 2). This is consistent with previous reports from culture-based studies (3). In accordance with these previous findings (4), members of the L. casei group were the most prevalent, followed by L. salivarius, L. gasseri, L. ultunensis (and its related phylotype), and L. crispatus. However, quantification of the prevalent species by real-time PCR showed that L. gasseri and L. ultunensis (and its related phylotype) were present in considerably higher numbers than the other species, consistent with the findings of their numerical predominance in the pooled carious dentine sample (Fig. 1 and 2). These two species also constituted the majority of lactobacilli present in several dentine samples, suggesting that they may possess a selective advantage for colonization and proliferation in decaying dentine. This notion is consistent with the observation of higher proportions of lactobacilli in severely demineralized dentine (36) and their affinity for collagen type I (24). Other prevalent species were found to constitute variable proportions in the dentine sample, and their role in caries progression cannot be overlooked (Fig. 2).
In conclusion, the findings of the present study indicate a complex and diverse presentation of lactobacilli in the advancing front of dentinal carious lesions. Phylogenetic analysis based on regions of the 16S rRNA genes enabled both an appreciation of this diversity and a high degree of precision in classifying representative lactobacilli. Although the combined approach of population analysis and real-time PCR quantification indicated a complete profile of the genus for many lesions, it was also evident that additional and as yet undefined lactobacilli were present in some lesions (Fig. 2). On the basis of known habitats, it is postulated that environmental influences, particularly dietary influences, have a determining role in colonization profiles within the lesions. It is also evident that both synergistic and antagonistic interactions would determine the final profile of Lactobacillus spp. and phylotypes within these lesions.
REFERENCES
- 1.Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. [DOI] [PubMed] [Google Scholar]
- 2.Ando, N., and E. Hoshino. 1990. Predominant obligate anaerobes invading the deep layers of root canal dentin. Int. Endodont. J. 23:20-27. [DOI] [PubMed] [Google Scholar]
- 3.Bjorndal, L., and T. Larsen. 2000. Changes in the cultivable flora in deep carious lesions following a stepwise excavation procedure. Caries Res. 34:502-508. [DOI] [PubMed] [Google Scholar]
- 4.Botha, S. J., S. C. Boy, F. S. Botha, and R. Senekal. 1998. Lactobacillus species associated with active caries lesions. J. Dent. Assoc. S. Afr. 53:3-6. [PubMed] [Google Scholar]
- 5.Brown, L. R., R. J. Billings, and A. G. Kaster. 1986. Quantitative comparisons of potentially cariogenic microorganisms cultured from noncarious and carious root and coronal tooth surfaces. Infect. Immun. 51:765-770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Cole, J. R., B. Chai, T. L. Marsh, R. J. Farris, Q. Wang, S. A. Kulam, S. Chandra, D. M. McGarrell, T. M. Schmidt, G. M. Garrity, and J. M. Tiedje. 2003. The Ribosomal Database Project (RDP-II): previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomy. Nucleic Acids Res. 31:442-443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Edwards, C., M. Collins, P. Lawson, and A. Rodriguez. 2000. Lactobacillus nagelii sp. nov., an organism isolated from a partially fermented wine. Int. J. Syst. E vol. Microbiol. 50:699-702. [DOI] [PubMed] [Google Scholar]
- 8.Fujisawa, T., Y. Benno, T. Yaeshima, and T. Mitsuoka. 1992. Taxonomic study of the Lactobacillus acidophilus group, with recognition of Lactobacillus gallinarum sp. nov. and Lactobacillus johnsonii sp. nov. and synonymy of Lactobacillus acidophilus group A3 (Johnson et al. 1980) with the type strain of Lactobacillus amylovorus (Nakamura 1981). Int. J. Syst. Bacteriol. 42:487-491. [DOI] [PubMed] [Google Scholar]
- 9.Hahn, C. L., W. A. Falkler, Jr., and G. E. Minah. 1991. Microbiological studies of carious dentine from human teeth with irreversible pulpitis. Arch. Oral Biol. 36:147-153. [DOI] [PubMed] [Google Scholar]
- 10.Helderman, W. H. V., M. I. N. Matee, J. S. vanderHoeven, and F. H. M. Mikx. 1996. Cariogenicity depends more on diet than the prevailing mutans streptococcal species. J. Dent. Res. 75:535-545. [DOI] [PubMed] [Google Scholar]
- 11.Hoshino, E. 1985. Predominant obligate anaerobes in human carious dentin. J. Dent. Res. 64:1195-1198. [DOI] [PubMed] [Google Scholar]
- 12.Hoshino, E., N. Ando, M. Sato, and K. Kota. 1992. Bacterial invasion of non-exposed dental pulp. Int. Endodont. J. 25:2-5. [DOI] [PubMed] [Google Scholar]
- 13.Hugenholtz, P., B. M. Goebel, and N. R. Pace. 1998. Impact of culture-independent studies on the emerging phylogenetic view of bacterial diversity. J. Bacteriol. 180:4765-4774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Johnson, J. L., C. F. Phelps, C. S. Cummins, J. London, and F. Gasser. 1980. Taxonomy of the Lactobacillus acidophilus group. Int. J. Syst. Bacteriol. 30:53-68. [Google Scholar]
- 15.Jukes, T. H., and C. R. Cantor. 1969. Evolution of protein molecules, p. 21-132. In H. N. Munro (ed.), Mammalian protein metabolism. Academic Press, New York, N.Y.
- 16.Klaenhammer, T., E. Altermann, F. Arigoni, A. Bolotin, F. Breidt, J. Broadbent, R. Cano, S. Chaillou, J. Deutscher, M. Gasson, M. van de Guchte, J. Guzzo, A. Hartke, T. Hawkins, P. Hols, R. Hutkins, M. Kleerebezem, J. Kok, O. Kuipers, M. Lubbers, E. Maguin, L. McKay, D. Mills, A. Nauta, R. Overbeek, H. Pel, D. Pridmore, M. Saier, D. van Sinderen, A. Sorokin, J. Steele, D. O'Sullivan, W. de Vos, B. Weimer, M. Zagorec, and R. Siezen. 2002. Discovering lactic acid bacteria by genomics. Antonie van Leeuwenhoek 82:29-58. [DOI] [PubMed] [Google Scholar]
- 17.Krooneman, J., F. Faber, A. C. Alderkamp, S. Oude Elferink, F. Driehuis, I. Cleenwerck, J. Swings, J. C. Gottschal, and M. Vancanneyt. 2002. Lactobacillus diolivorans sp. nov., a 1,2-propanediol-degrading bacterium isolated from aerobically stable maize silage. Int. J. Syst. E vol. Microbiol. 52:639-646. [DOI] [PubMed] [Google Scholar]
- 18.Leser, T. D., J. Z. Amenuvor, T. K. Jensen, R. H. Lindecrona, M. Boye, and K. Moller. 2002. Culture-independent analysis of gut bacteria: the pig gastrointestinal tract microbiota revisited. Appl. Environ. Microbiol. 68:673-690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Love, R. M., and H. F. Jenkinson. 2002. Invasion of dentinal tubules by oral bacteria. Crit. Rev. Oral Biol. Med. 13:171-183. [DOI] [PubMed] [Google Scholar]
- 20.Love, R. M., M. D. McMillan, Y. Park, and H. F. Jenkinson. 2000. Coinvasion of dentinal tubules by Porphyromonas gingivalis and Streptococcus gordonii depends upon binding specificity of streptococcal antigen I/II adhesin. Infect. Immun. 68:1359-1365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Marchant, S., S. R. Brailsford, A. C. Twomey, G. J. Roberts, and D. Beighton. 2001. The predominant microflora of nursing caries lesions. Caries Res. 35:397-406. [DOI] [PubMed] [Google Scholar]
- 22.Martin, F. E., M. A. Nadkarni, N. A. Jacques, and N. Hunter. 2002. Quantitative microbiological study of human carious dentine by culture and real-time PCR: association of anaerobes with histopathological changes in chronic pulpitis. J. Clin. Microbiol. 40:1698-1704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Massey, W. L., D. M. Romberg, N. Hunter, and W. R. Hume. 1993. The association of carious dentin microflora with tissue changes in human pulpitis. Oral Microbiol. Immunol. 8:30-35. [DOI] [PubMed] [Google Scholar]
- 24.McGrady, J. A., W. G. Butcher, D. Beighton, and L. M. Switalski. 1995. Specific and charge interactions mediate collagen recognition by oral lactobacilli. J. Dent. Res. 74:649-657. [DOI] [PubMed] [Google Scholar]
- 25.McOrist, A. L., M. Jackson, and A. R. Bird. 2002. A comparison of five methods for extraction of bacterial DNA from human faecal samples. J. Microbiol. Methods 50:131-139. [DOI] [PubMed] [Google Scholar]
- 26.Nadkarni, M. A., F. E. Martin, N. A. Jacques, and N. Hunter. 2002. Determination of bacterial load by real-time PCR using a broad-range (universal) probe and primers set. Microbiology 148:257-266. [DOI] [PubMed] [Google Scholar]
- 27.Nagaoka, S., Y. Miyazaki, H. J. Liu, Y. Iwamoto, M. Kitano, and M. Kawagoe. 1995. Bacterial invasion into dentinal tubules of human vital and nonvital teeth. J. Endodont. 21:70-73. [DOI] [PubMed] [Google Scholar]
- 28.Neefs, J. M., Y. Van de Peer, P. De Rijk, S. Chapelle, and R. De Wachter. 1993. Compilation of small ribosomal subunit RNA structures. Nucleic Acids Res. 21:3025-3049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Paster, B. J., S. K. Boches, J. L. Galvin, R. E. Ericson, C. N. Lau, V. A. Levanos, A. Sahasrabudhe, and F. E. Dewhirst. 2001. Bacterial diversity in human subgingival plaque. J. Bacteriol. 183:3770-3783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Roos, S., F. Karner, L. Axelsson, and H. Jonsson. 2000. Lactobacillus mucosae sp nov, a new species with in vitro mucus-binding activity isolated from pig intestine. Int. J. Syst. E vol. Microbiol. 50:251-258. [DOI] [PubMed] [Google Scholar]
- 31.Saitou, N., and M. Nei. 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. E vol. 4:406-425. [DOI] [PubMed] [Google Scholar]
- 32.Shovlin, F. E., and R. E. Gillis. 1968. Biochemical and antigenic studies of lactobacilli isolated from deep dentinal caries. I. Biochemical aspects. J. Dent. Res. 48:356-360. [DOI] [PubMed] [Google Scholar]
- 33.Smith, S. I., A. J. Aweh, A. O. Coker, K. O. Savage, D. A. Abosede, and K. S. Oyedeji. 2001. Lactobacilli in human dental caries and saliva. Microbios 105:77-85. [PubMed] [Google Scholar]
- 34.Staat, R. H., T. H. Gawronski, D. E. Cressey, R. S. Harris, and L. E. Folke. 1975. Effects of dietary sucrose levels on the quantity and microbial composition of human dental plaque. J. Dent. Res. 54:872-880. [DOI] [PubMed] [Google Scholar]
- 35.Thompson, J. D., D. G. Higgins, and T. J. Gibson. 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22:4673-4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.van Strijp, A. J., T. J. van Steenbergen, and J. M. ten Cate. 1997. Bacterial colonization of mineralized and completely demineralized dentine in situ. Caries Res. 31:349-355. [DOI] [PubMed] [Google Scholar]
- 37.Vogel, R. F., G. Bocker, P. Stolz, M. Ehrmann, D. Fanta, W. Ludwig, B. Pot, K. Kersters, K. H. Schleifer, and W. P. Hammes. 1994. Identification of lactobacilli from sourdough and description of Lactobacillus pontis sp. nov. Int. J. Syst. Bacteriol. 44:223-229. [DOI] [PubMed] [Google Scholar]
- 38.von Wintzingerode, F., U. B. Gobel, and E. Stackebrandt. 1997. Determination of microbial diversity in environmental samples: pitfalls of PCR-based rRNA analysis. FEMS Microbiol. Rev. 21:213-229. [DOI] [PubMed] [Google Scholar]