ABSTRACT
HIV-1 envelope glycoprotein (Env) spikes are prime vaccine candidates, at least in principle, but suffer from instability, molecular heterogeneity and a low copy number on virions. We anticipated that chemical cross-linking of HIV-1 would allow purification and molecular characterization of trimeric Env spikes, as well as high copy number immunization. Broadly neutralizing antibodies bound tightly to all major quaternary epitopes on cross-linked spikes. Covalent cross-linking of the trimer also stabilized broadly neutralizing epitopes, although surprisingly some individual epitopes were still somewhat sensitive to heat or reducing agent. Immunodepletion using non-neutralizing antibodies to gp120 and gp41 was an effective method for removing non-native-like Env. Cross-linked spikes, purified via an engineered C-terminal tag, were shown by negative stain EM to have well-ordered, trilobed structure. An immunization was performed comparing a boost with Env spikes on virions to spikes cross-linked and captured onto nanoparticles, each following a gp160 DNA prime. Although differences in neutralization did not reach statistical significance, cross-linked Env spikes elicited a more diverse and sporadically neutralizing antibody response against Tier 1b and 2 isolates when displayed on nanoparticles, despite attenuated binding titers to gp120 and V3 crown peptides. Our study demonstrates display of cross-linked trimeric Env spikes on nanoparticles, while showing a level of control over antigenicity, purity and density of virion-associated Env, which may have relevance for Env based vaccine strategies for HIV-1.
IMPORTANCE The envelope spike (Env) is the target of HIV-1 neutralizing antibodies, which a successful vaccine will need to elicit. However, native Env on virions is innately labile, as well as heterogeneously and sparsely displayed. We therefore stabilized Env spikes using a chemical cross-linker and removed non-native Env by immunodepletion with non-neutralizing antibodies. Fixed native spikes were recognized by all classes of known broadly neutralizing antibodies but not by non-neutralizing antibodies and displayed on nanoparticles in high copy number. An immunization experiment in rabbits revealed that cross-linking Env reduced its overall immunogenicity; however, high-copy display on nanoparticles enabled boosting of antibodies that sporadically neutralized some relatively resistant HIV-1 isolates, albeit at a low titer. This study describes the purification of stable and antigenically correct Env spikes from virions that can be used as immunogens.
INTRODUCTION
A vaccine for HIV-1 will most likely need to elicit broadly neutralizing antibodies (bNAbs) in order to protect against the vast diversity of circulating primary isolates (1–3). Neutralizing antibodies bind to the envelope glycoprotein (Env) spike and block the virus from productively infecting target cells prior to, or in some cases following, its engagement with cellular receptors (i.e., CD4 and chemokine coreceptor) (4, 5). Innate features of Env appear to limit elicitation of bNAbs, including the presence of variable residues and heavy glycosylation immediately adjacent to conserved epitopes, and a low copy number of ∼10 to 14 spikes per virion (2, 6–9). Thus, native Env often elicits rather weak or isolate specific neutralizing antibody in small animal models (8).
Mature Env spikes, trimers of gp120-gp41 heterodimers, are formed once immature gp160 has been glycosylated in the host cell and processed by furin proteases into gp120 (surface) and gp41 (transmembrane) subunits (4). Immature gp160, gp41 stumps, monomeric gp120, and nontrimeric Env debris often contaminate virions and other preparations of Env and can elicit non-neutralizing antibodies that fail to recognize mature spikes (10–13). HIV-1 spikes can also spontaneously inactivate and physically dissociate over time, which can be further accelerated in the presence of ligands or antibodies to various regions of Env (14–17). When used as an immunogen such decay of Env presumably contributes to ineffective, “off-target” antibody responses.
Attempts have been made to stabilize Env in soluble form using genetic mutations, trimerization domains, and disruption of the furin cleavage site (18–23). Most notably, BG505 SOSIP, is a soluble (gp140) trimer that contains an engineered disulfide bond between gp120 and gp41 and a mutation (I559P) to disfavor aberrant forms of gp41 (18). This engineered trimer is recognized by most bNAbs but not by non-neutralizing antibodies, while cryo-electron microscopy (cryo-EM) and X-ray crystallography have provided near-atomic-level models with strong topological similarity to virion-associated spikes (24–26). However, mimicry with these molecules is imperfect, since, for example, some non-neutralizing antibodies can bind to BG505 SOSIP and not native spikes (18). Domains of gp41 were also truncated from SOSIP gp140s (e.g., membrane-proximal external region [MPER], transmembrane [TM], and cytoplasmic tail [CT]) that alters display and eliminates bNAb epitopes (27–29). Thus, methods are desired to stabilize more complete Env molecules, especially since, to date, no immunogen has reportedly elicited bNAb titers of sufficient breadth and potency to form the basis of a vaccine for HIV-1 (22, 27).
Cross-linkers have been used to probe conformational states of Env (30–34) and typically diminish the binding of non-neutralizing antibodies, as well as V3 antibodies with lesser effects on binding by many bNAbs (31, 32). The use of glutaraldehyde or bis(sulfosuccinimidyl) suberate (BS3) to cross-link soluble gp140s and/or complexes between gp120 or gp140 and CD4 has reportedly helped elicit cross-isolate neutralizing antibodies (35, 36), although in the latter case antibodies to host CD4 complicated analyses (37). To date, no immunization study has been described that uses purified native spikes, cross-linked or otherwise.
Formaldehyde-fixed virions have been used to immunize mice and reportedly elicited weak neutralizing antibodies against heterologous isolates (38, 39). In these studies, however, Env was subjected to harsh heat treatment, molecular heterogeneity in Env was not studied, and the low copy number of Env was not addressed, necessitating high doses of HIV-1 virions that also elicit antibodies to cellular proteins that incorporate onto virions (8, 39, 40). The presence of misfolded Env on virions can facilitate recognition of HIV-1 by non-neutralizing antibodies, and the decay of native Env can alter or decrease recognition by bNAbs (10, 15).
Here, we sought to use cross-linker to stabilize native spikes of HIV-1 while simultaneously mitigating problems of immunizing with insufficient copies of Env per virion, as well as the presence of misfolded Env and other proteins found in virion preparations. The relatively homogeneously trimeric Env of HIV-1 JR-FL was treated with a chemically defined cross-linker, BS3, and purified from virion debris via an affinity tag in mild detergent. We further immunodepleted non-native forms of Env using non-neutralizing antibodies. These steps resulted in highly stable trimers, free of virion debris, that were recognized by bNAbs and not by typical V3 crown antibodies and non-neutralizing antibodies. We find that cross-linker can alter bNAb epitopes in a concentration-dependent manner. Moreover, cross-linker shows a limited capacity to protect quaternary antibody epitopes from disruption by heat and reduction of disulfide bonds. Notably, when arrayed on proteoliposome nanoparticles (PLNs), cross-linked spikes elicited antibodies that sporadically neutralized relatively resistant tier 1b and tier 2 isolates of HIV-1, albeit at a relatively low titer. Meanwhile, the neutralization of sensitive tier 1a strains, associated with V3 crown antibodies, was clearly diminished. This study provides information relevant to the rational development of nanoparticle immunogens displaying chemically stabilized Env spikes of HIV-1.
MATERIALS AND METHODS
Reagents. (i) Plasmids.
Molecular clone pLAI-JR-FL contains the env ectodomain (amino acids [aa] 24 to 692) of JR-FL in the pLAI.2 backbone plasmid (15). pcDNA-JR-FL is an Env complementation vector that matches env from pLAI-JR-FL (10). To make epitope tagged JR-FL plasmid, FLAG tag (DYKDDDDK) was introduced by QuikChange mutagenesis (Agilent) at the C-terminal end of Env (gp160). A tripeptide linker “NSG” was included N-terminal of the tag. Env-complementation plasmids pSVIII-JR-FL, JR-CSF, SF162, ADA, HxB2, and pCAGGS-SIVmac239 have been described previously (41, 42). The clade B Env reference panel and pSG3ΔEnv were obtained from the National Institutes of Health AIDS Research and Reference Reagent Program (NIH ARRRP), contributed by D. Montefiori and F. Gao (43) and by J. Kappes and X. Wu (44), respectively. Plasmid pMV-5001 and related variants that encode codon-optimized env from the JR-CSF isolate, as well as env variants with domain swaps between JR-CSF and JR-FL, were generously provided by R. Whalen (Altravax).
(ii) Cells.
TZM-bl cells (44) were obtained from the NIH ARRRP (contributed by J. Kappes and X. Wu), 293T cells were purchased from the American Type Culture Collection, and MT2-CCR5ΔCT cells were obtained from D. Mosier (TSRI). Cell lines were maintained in Dulbecco modified Eagle medium (TZM-bl and 293T cells) or RPMI medium (MT2 cells) containing 10% fetal calf serum, 20 mM l-glutamine, 100 U of penicillin/ml, and 100 μg of streptomycin/ml.
(iii) Antibodies.
Anti-HIV-1 monoclonal antibodies (human IgG unless otherwise stated) were obtained from the following sources: 2G12 (45), 4E10 (46), 2F5 (47), and 447-52D (48) from H. Katinger (Polymun); PGT151 (49), PGV04 (50), b12 (51), and b6 (52) from D. Burton (TSRI); VRC01 (53) from J. Mascola (VRC, NIH); PGT121, PGT126, and PGT128 (54) and PG9 and PG16 (55) from Steven Fling (International AIDS Vaccine Initiative Neutralizing Antibody Consortium); 35O22 (56) and 10E8 (57) from Mark Connors (NIH); 7B2 (58), C11 (59), 19b (60), and 17b (61) from J. Robinson (Tulane); murine D49 and D50 (62) from P. Earl (NIH); F425-B4e8 (63) from L. Cavacini (Harvard Medical School); and CD4-IgG2 (64) from W. Olson (Progenics). Z13e1 (65) was produced in-house. Mouse anti-FLAG M2 antibody (Sigma) was obtained commercially.
(iv) Proteins.
Trimut core, Trimut 368/370, gp140-foldon (FT) trimers (66), and JR-FL gp140 SOSIP trimers (18) were kindly donated by R. Wyatt (TSRI). Monomeric JR-FL gp120 was purchased from Progenics, and JR-FL gp41 MBP-fusion protein M41xt (aa 535 to 681) (65) was produced in Escherichia coli in-house. CD4bs scaffold RSC3 (53) was obtained from the NIH ARRRP (Z-Y. Yang, P. Kwong, G. Nabel, and J. Mascola). N332 supersite scaffold 1GUT was generously contributed by J. Zhu (TSRI).
Virus production.
HIV-1 virions were produced by transient transfection of 293T cells using the transfection reagent polyethyleneimine (PEI [67]) and harvested after 72 h as previously described (15). For cross-linking and trimer purification, culture supernatants were cleared of cell debris by centrifugation at 2,000 × g for 15 min; supernatants were then centrifuged at 60,000 × g for 1 h, and pellets were resuspended in phosphate-buffered saline (PBS) at 100 times the original concentration. Virus replication in MT2-CCR5ΔCT cells was carried out as previously described (23).
Cross-linking of virus.
Cross-linkers bis(sulfosuccinimidyl)suberate (BS3; Pierce), and 3,3′-dithiobis(sulfosuccinimidylpropionate) (DTSSP; Pierce) were dissolved in PBS immediately before use. Unless otherwise noted, cross-linkers were added to a final concentration of 1 mM. Cross-linkers and virus were incubated for 20 min at room temperature, and the reaction was quenched by the addition of 20 mM Tris-HCl (pH 7.5) for 15 min. Virus was then pelleted to remove unreacted cross-linker by spinning at 20,000 × g for 45 min in a microcentrifuge and resuspended in fresh PBS.
BN-PAGE.
Blue native-polyacrylamide gel electrophoresis (BN-PAGE) and Western blotting was performed using the NativePAGE Gel System (Invitrogen) as previously described (10). Coomassie blue staining was performed using Simply Blue Safe Stain (Invitrogen) according to the manufacturer's instructions. In some cases, virus was heated for 1 h or incubated at 37°C for up to 96 h prior to running on the gel. In gel mobility shift experiments, antibodies and virus were incubated for 20 min on ice prior to sample preparation. Antibodies were added at a concentration of 10 μg/ml (for bNAbs) or 40 μg/ml (for non-neutralizing antibodies).
SDS-PAGE.
Virus or protein samples were incubated in Laemmli buffer (Bio-Rad) containing 50 mM dithiothreitol (DTT) for 5 min at 100°C prior to loading on 4 to 15% Tris-HCl gels (Bio-Rad). Gels were run at room temperature using running buffer (25 mM Tris, 192 mM glycine, 0.1% SDS) at 120V for 1 h and then transferred to polyvinylidene difluoride membrane and Western blotted as previously described (10).
ELISA. (i) Capture ELISA.
Microtiter wells were coated with Galanthus nivalis lectin (GNL) at 5 μg/ml in PBS overnight at 4°C. Plates were then washed (all washes were performed using PBS containing 0.05% Tween [PBST]) and blocked with 4% nonfat dry milk (NFDM) in PBS for 1 h at 37°C. Next, the plates were washed, and virus solubilized in 1% Empigen was added. After incubation for 1 h at 37°C, the plates were washed, and primary antibody in PBST containing 0.4% NFDM was added for 1 h at 37°C. The plates were washed again, goat anti-human Fcγ-horseradish peroxidase (HRP) secondary antibody (Jackson) was added, and the plates were incubated for 45 min at room temperature. After another wash, TMB substrate (Pierce) was added for 5 min, and the absorbance was read at 450 nm.
(ii) Direct ELISA.
Direct ELISAs were performed as described above, except that antigens were coated directly onto microtiter wells, sera from immunized animals were used as primary antibodies, and the secondary antibody was anti-rabbit IgG-HRP (Jackson).
(iii) Competition ELISA.
Monomeric JR-FL gp120 or JR-FL gp41-MBP fusion protein M41xt were directly coated onto microtiter wells. After blocking, a constant concentration of biotinylated monoclonal antibody (selected to give 50% maximum signal) was prepared, and a titration of immune serum was added at the same time, followed by incubation for 1 h at 37°C. Streptavidin-HRP (Jackson) was used to detect antibody binding.
(iv) ELISA to quantify Env.
To quantify Env for immunization, GNL capture ELISAs were performed, as described above, in which a dilution series of Empigen-lysed virus was added to GNL-coated wells. Env was detected using an antibody cocktail consisting of 2G12 and b12 (2 μg/ml each). The concentration of viral Env was interpolated against a sequence-matched JR-FL gp120 (Progenics) standard curve by Prism software (GraphPad).
(v) p24 ELISA.
Microtiter wells were coated overnight at 4°C using 50 μl of a 5-μg/ml concentration of sheep anti-p24 (Aalto) and then blocked for 1 h at 37°C with 4% NFDM in PBS. Virions were lysed by adding 1% Empigen in PBS, and 50 μl was transferred to anti-p24-coated wells. After 2 h of incubation at 37°C, p24 was probed with sheep anti-p24-AP (Aalto) and detected using an AMPAK amplification kit (Argene) according to the manufacturer's directions.
Purification of viral Env spikes.
We devised a two-step protocol for purifying BS3-treated HIV-1 spikes involving immunodepletion of misfolded Env molecules using non-neutralizing antibodies, followed by purification via a C-terminal affinity tag. For step 1, immunodepletion using non-neutralizing antibodies was carried out. IgGs b6 and D49 were bound individually to protein G-Sepharose beads (GE Healthcare) by coincubation for 1 h in PBS, followed by the addition of 20 mM dimethylpimelimidate (DMP) in 0.2 M sodium borate (pH 9.0) for 30 min at room temperature. The coupling reaction was quenched with 0.2 M ethanolamine (pH 8.0) for 2 h at room temperature. BS3-cross-linked virions were solubilized in PBS containing 0.1% n-dodecyl-β-d-maltoside (DDM). Detergent-treated samples were sequentially incubated with b6- or D49-coated beads for 1 h at room temperature in PBS, and the remaining supernatant was saved. For step 2, FLAG-tagged trimeric Env spikes were purified. Cross-linked virion-associated Env in (immunodepleted) supernatant was added at a 5:1 (vol/vol) ratio to anti-FLAG M2 IgG-coupled agarose beads (Sigma) for 3 h at room temperature. Beads were washed four times for 5 min in PBS containing 0.1% DDM (PBSD). To elute bound trimers, beads were incubated with FLAG-peptide at 250 μg/ml in PBSD for 1 h at room temperature. The eluted supernatant was saved, the elution step was repeated twice more, and the supernatants were combined.
Incorporation of Env spikes into proteoliposomes.
For PLNs, mouse anti-FLAG M2 IgG (Sigma) was covalently coupled to magnetic Amino-Adembeads (Ademtech; 200 nm) according to the manufacturer's protocol. Antibody-coupled magnetic beads were incubated with detergent-treated cross-linked viral supernatants for 3 h at room temperature and then washed three times with PBSD. Separately, Cholesterol/sphingomyelin/DOPE/POPC/DOPS/biotinylated DOPE were mixed in a 45:27:19:9:9:1 ratio meant to mimic the composition of the viral membrane (68) and dried for 2 h in a fume hood. Lipids were resuspended in PBS by sonicating for 5 min on a 4°C water bath, and DDM was added to a final concentration of 1%. The lipids were then added to magnetic beads containing cross-linked trimers and incubated for 15 min at room temperature. Detergent was removed by dialysis against PBS overnight at 4°C. Beads were recovered, washed three times in PBS, and stored in PBS at 4°C.
Immunization of rabbits.
Four groups of three New Zealand White rabbits (Aldevron, Fargo, ND) each received two DNA primes consisting of pMV-5001 (Altravax), a plasmid encoding codon-optimized env from the HIV-1 strain JR-CSF. DNA (400 μg) was given by intramuscular electroporation using an Inovio twin injector device on days 0 and 28. At day 98, rabbits in each group were given the following boosts by intramuscular injection: group A received JR-FL virus-like particle (VLP) containing 25-μg Env equivalents, group B received BS3-cross-linked JR-FL VLP containing 25-μg Env equivalents, group C received BS3-cross-linked and purified JR-FL Env trimers in PBS containing 0.1% DDM (50 μg of Env), and group D received BS3-cross-linked and purified JR-FL Env trimers incorporated in PLNs (50 μg of Env). Groups A, B, and D received 100 μg of CpG ODN 2007 as an adjuvant, while group C received both 100 μg of CpG ODN 2007 and alum (Pierce), mixed at a 1:1.4 alum/boosting protein ratio. Ear bleeds were performed 14 days postprime and postboost.
Electron microscopy.
Preparation of negative stain electron microscopy grids, data collection, and image processing were performed as stated elsewhere (69). Briefly, BS3-cross-linked Env trimers were diluted to ∼0.01 mg/ml. adsorbed onto carbon coated 400-mesh Cu grids, and then stained with 2% uranyl formate. Images were collected via Leginon (70) using a 4k×4k Tietz Temcam F-416 at ×52,000 magnification on a Tecnai T12 electron microscope, resulting in a pixel size of 2.05 Å/pixel on the specimen plane. Particles were picked and stacked in Appion (71). Reference-free two-dimensional (2D) class averages were generated using the Sparx software package (72).
Neutralization assays.
Neutralization was determined in a single-cycle neutralization assay using pseudotyped virus and TZM-bl cells as target cells. Serum, heat inactivated for 1 h at 56°C, was added to virus in 100 μl of 293T cell culture media. Virus-serum mixtures were incubated for 1 h at 37°C prior to addition to target cells. After a 72 h of incubation at 37°C, the cells were lysed, Bright-Glo luciferase reagent (Promega) was added, and the luminescence in relative light units was measured using an Orion luminometer (Berthold).
RESULTS
Virion spikes stabilized by cross-linking.
To fix HIV-1 Env in its native state, we chose to use the JR-FL molecular clone produced by transfection of 293T cells, as we previously found these virions to produce relatively homogeneous and well-cleaved Env trimers (10). After screening several common cross-linkers, we settled on BS3 due to its defined chemical properties and clarity of the cross-linked trimer band on BN-PAGE (Fig. 1A and data not shown). When tested for its resistance to destabilizing conditions, BS3-cross-linked Env remained trimeric after 1 h at 57°C, after 96 h in DDM detergent, and even after boiling for 5 min in SDS-PAGE buffer, while non-cross-linked trimers completely dissociated under these conditions (Fig. 1B to D). BS3 was used at 1 mM for this and all subsequent experiments, unless noted otherwise, because this concentration prevented any trimer dissociation upon heat treatment (data not shown).
FIG 1.

Cross-linked Env trimers are highly stable. (A) Chemical structures of BS3 and DTSSP. (B) Virions were treated with 1 mM BS3 or left untreated and then incubated at either 37°C or 57°C for 1 h. Heat-treated virions were solubilized in DDM, run on BN-PAGE, and probed by Western blotting using a gp41 (4E10, 2F5, and Z13e1) antibody cocktail. (C) Untreated and BS3-cross-linked virions were boiled in Laemmli buffer with DTT for 5 min, run on SDS-PAGE, and visualized by Western blotting with an anti-gp120 (b12, 2G12, and F425-B4e8) antibody cocktail. (D) Untreated and BS3-cross-linked virions were incubated in 1% DDM detergent at 37°C for up to 4 days and then analyzed by BN-PAGE and Western blotting as described above.
Antigenicity of BS3-cross-linked trimeric spikes.
To assess how cross-linking affects relevant bNAb epitopes, BS3-cross-linked virions were detergent solubilized, and then Env was captured onto immobilized GNL and probed using a panel of antibodies to gp120 and gp41 (Fig. 2). This panel of bNAbs included PG9 and PG16 (gp120 V2/V3), as well as PGT151 and 35O22 (gp120-gp41 interface), that recognize quaternary epitopes preferentially exposed on trimeric Env (49, 55, 56). Also included were PGT121, PGT126, and PGT128, which target the N332 glycan at the base of V3 (N332 supersite); F425-B4e8 and 447-52D (V3 crown); VRC01, PGV04, b12, and CD4-IgG2 (CD4 binding site; CD4bs); 2G12 (gp120 glycans); and 10E8, 4E10, 2F5, and Z13e1 (gp41 MPER).
FIG 2.

Relative binding of bNAbs and non-neutralizing antibodies to BS3-cross-linked Env spikes. HIV-1 virions were either treated with 1 mM BS3 or left untreated, the virions were detergent solubilized, and Env was captured on GNL. Antibody binding was determined by ELISA and the fold change in binding (50% effective concentration [EC50]) to cross-linked Env versus un-cross-linked Env was determined (EC50 untreated/EC50 cross-linked). bNAbs used have an IC50 of <50 μg/ml against JR-FL, and non-neutralizing antibodies have an IC50 of >50 μg/ml. PG9/PG16 binding was determined using JRFL mutant E168R. PG9, PG16, PGT151, and 35O22 did not reach an EC50 against un-cross-linked Env. The results shown are averages of at least three independent experiments.
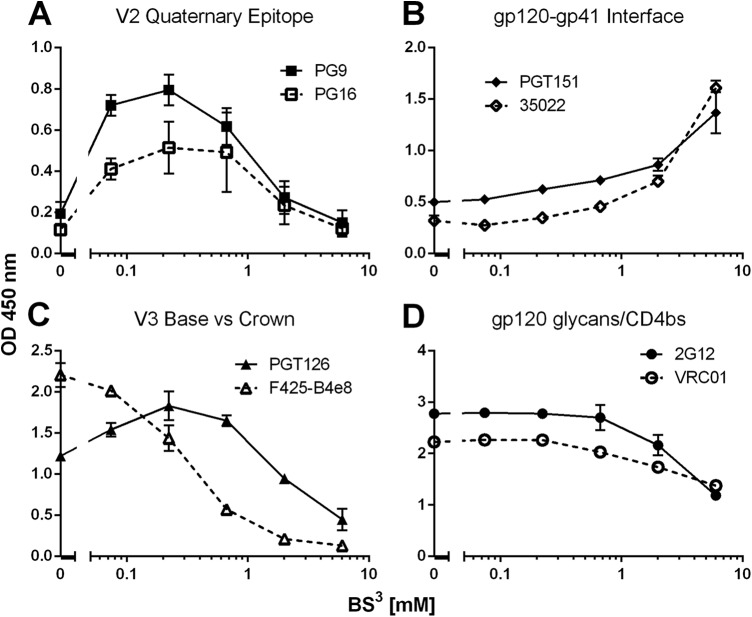
Trimer-preferring bNAbs PG9/PG16, PGT151, and 35O22 all recognized BS3-cross-linked Env but not un-cross-linked Env (Fig. 2). We note that the Env used, JR-FL, contained the mutation E168R, since a Lys or Arg at position 168 is required for PG9/PG16 recognition and Arg is far less likely than Lys to be modified by BS3 (55, 73). At concentrations above 0.2 mM BS3, PG9/PG16 binding decreased, whereas PGT151 and 35O22 binding continued to increase up to 6 mM BS3 (Fig. 3A and B). Remarkably, binding of bNAbs to the N332 supersite was improved 2- to 3-fold by treatment with BS3, whereas binding by V3 crown antibodies was diminished 15-fold (Fig. 2). A dose-response analysis revealed a maximum 6-fold enhancement of PGT126 binding to Env (EC50) at 0.2 mM BS3, which began to decrease at higher concentrations, whereas F425-B4e8 binding only decreased with BS3 concentration in a dose-dependent manner (Fig. 3C). BS3-cross-linking did not alter binding by 2G12 and MPER antibodies (Fig. 2). However, BS3 diminished binding of CD4bs ligands b12 and CD4-IgG2 by 4- and 15-fold, respectively, but had negligible effect on VRC01 and PGV04 (Fig. 2 and 3D). In summary, BS3 cross-linking of virion-displayed spikes occludes the V3 crown, slightly restricts some CD4bs epitopes, has no effect on the gp41 MPER and largely preserves quaternary epitopes on V2, the gp120-gp41 interface, and the N332 glycan supersite.
FIG 3.
Effect of BS3 treatment on antibody recognition of virion-associated Env as a function of cross-linker concentration. Virions were treated using a 3-fold dilution series of BS3. Env was detergent solubilized and probed by bNAbs PG9 and PG16 (A), PGT151 and 35O22 (B), PGT126 and F425-B4e8 (C), and 2G12 and VRC01 (D) in a lectin-capture ELISA. The optical density (OD) at 450 nm is reported using PG9, PG16, PGT151, and 35O22 at 10 μg/ml and PGT126, F425-B4e8, 2G12, and VRC01 at 0.08 μg/ml.
In contrast to the relatively modest effect on bNAb epitopes, BS3 cross-linking strongly reduced binding of six different non-neutralizing antibodies by up to 3 orders of magnitude regardless of epitope; CD4bs antibody b6 was the exception in which binding was reduced only 4-fold (Fig. 2). Since these virions display predominantly native trimers (10) (Fig. 1B), we infer that BS3 reduces non-neutralizing antibody binding largely by preventing dissociation of Env subunits into non-native species within the time of the assay, which is also consistent with the observed enhancement in binding to the N332 supersite and quaternary bNAb epitopes.
To assess how the observed effects of cross-linking on bNAb binding relate to antibody occupancy of trimeric spikes, we used a BN-PAGE gel mobility shift assay in which binding by bNAb causes Env spikes to run slower on the gel (11). IgG fragments antigen-binding (Fabs) of b12, 2G12, PGT151, 10E8, PGT126, PG9, 4E10, and Z13e1 all retarded the electrophoretic mobility of cross-linked spikes, whereas non-neutralizing antibodies b6 and D49, and V3 crown binder F425-B4e8 had no effect (Fig. 4 and data not shown). However, Fab b12 bound cross-linked spikes with an occupancy of about one Fab per trimer less than untreated trimers under the same conditions (Fig. 4B). Taken as a whole, we conclude that BS3 cross-linking does not broadly alter occupancy of antibodies on JR-FL spikes for the majority of bNAb epitopes.
FIG 4.

Occupancy by bNAbs on cross-linked Env spikes determined using BN-PAGE gel mobility shift. (A) JR-FL virions, either BS3 cross-linked or untreated, were incubated with various antibodies, and Env was resolved using BN-PAGE. Fab fragments were used in this experiment to avoid the cross-linking of spikes that occurs with whole IgGs. Gel mobility shift with PG9 was determined using Env containing the mutation E168R that permits binding by PG9 to JR-FL. (B) Gel mobility shifts induced by antibodies were quantified using imaging software, including some antibodies not shown in panel A. The stoichiometry was calculated by measuring the distance of the gel mobility shift and comparing Fabs of known stoichiometry against un-cross-linked JR-FL (PG9 = one Fab/trimer, PGT151 = two Fabs/trimer, and PGT126 = three Fabs/trimer). Note that domain-swapped 2G12 is an F(ab′)2 and induces a gel mobility shift that is approximately twice what would be expected. Statistically significant changes were determined using an unpaired two-tailed Student t test.
Although the stability of BS3-treated spikes is high at the oligomeric level (Fig. 1), for vaccine development the integrity of individual epitopes is more relevant, so we investigated this using BN-PAGE and ELISA. Among the bNAbs tested, including PG9 and PGT151 to quaternary epitopes, all shifted cross-linked trimers after treatment at 57°C for 1 h, which is a condition that causes un-cross-linked JR-FL spikes to fully dissociate (Fig. 5A). Cross-linking also rescued binding to spikes by PGT151, 35O22, PG16, VRC01, b12, PGT128, PGT121, and 2G12 in an ELISA after heat treatment (Fig. 5B). Notably, under the conditions of this assay bNAb epitopes on cross-linked viral Env were generally more stable than on sequence-matched JR-FL SOSIP gp140. However, incubation at higher temperatures (i.e., ≥70°C) largely eliminated bNAb binding to Env in all samples, with the exception of more linear epitopes (i.e., MPER and V3 crown). In sum, cross-linking significantly enhances thermostability of bNAb epitopes, but an extreme temperature disrupts the antigenicity of Env despite preservation of the trimeric state.
FIG 5.
Conformational epitopes on cross-linked Env spikes are relatively thermostable. (A) BS3-cross-linked Env was incubated at 57°C for 1 h and examined using BN-PAGE gel mobility shift. (B) Untreated JR-FL virions, BS3-cross-linked JR-FL virions, or soluble JR-FL SOSIP were incubated at a range of temperatures for 1 h. The Env was solubilized in 1% Empigen detergent (detergent was also added to JR-FL SOSIP for consistency), and bNAb binding was determined using a lectin-capture ELISA. All antibodies were tested at 1 μg/ml, and the percent binding at each temperature is shown relative to a control sample that was incubated at 4°C for 1 h. No binding was detected to un-cross-linked viral Env with PG16, PGT151, and 35O22, and no binding was detected to JR-FL SOSIP by 10E8, since the MPER has been removed from this construct.
Reversible cross-linker chemistry reveals stability features of Env quaternary epitopes.
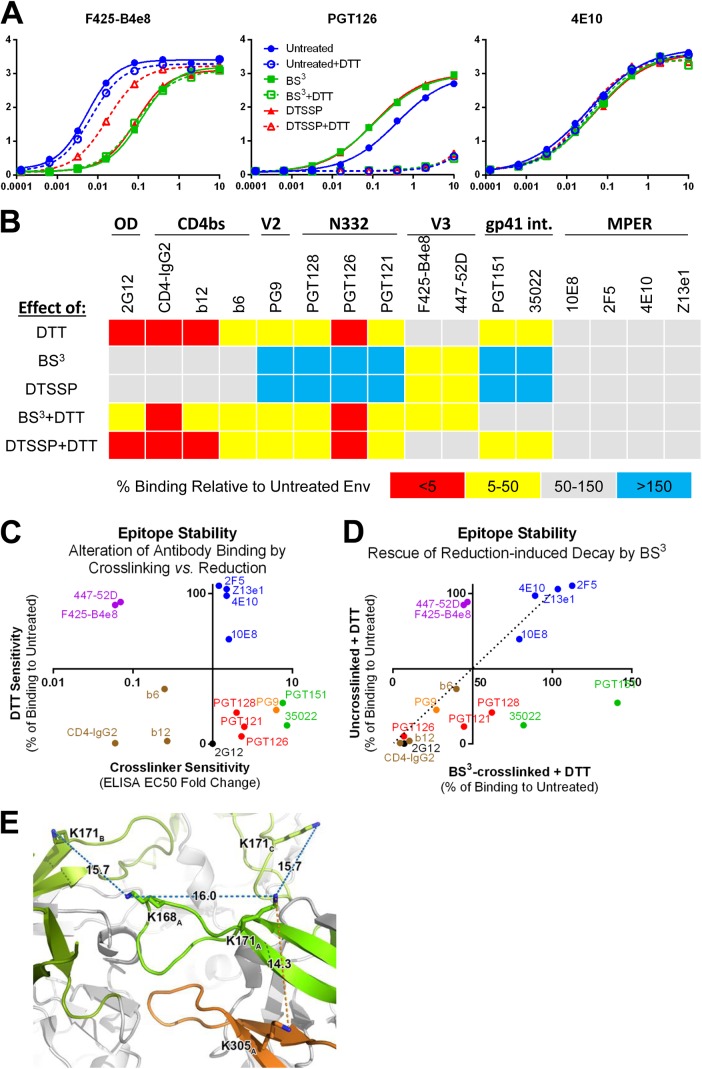
To determine whether effects of BS3 on binding by certain antibodies (Fig. 2) were due to direct chemical modification of epitopes or indirect (allosteric) fixation of Env quaternary structure, we made use of a disulfide containing cleavable homolog of BS3, DTSSP (Fig. 1A). We first verified that both cross-linkers caused a comparable diminution in binding to Env by V3 crown binders, F425-B4e8 and 447-52D (Fig. 6A and B). Next, we found that upon treatment with DTT, F425-B4e8 and 447-52D binding was substantially restored to spikes treated with DTSSP but not those treated with BS3. These results are consistent with a model in which neither DTSSP nor BS3 destroy V3 crown epitopes by direct chemical modification but rather lock Env conformation such that it becomes inaccessible to antibody. This likely occurs via V1-V1 interprotomer cross-links at Lys171 or Lys168 and/or intraprotomer linkage involving Lys171 in V1 and Lys305 at the V3 base (Fig. 6E). In contrast, DTT treatment had no effect on binding by MPER antibodies. Surprisingly, DTT virtually abrogated binding to the CD4bs, N332 supersite, PG9 epitope, and 2G12 epitope irrespective of cross-linker treatment, but disruption of epitopes in the gp120-gp41 interface (PGT151 and 35O22) caused by DTT treatment was alleviated by BS3 treatment (Fig. 6B and D). Notably, conformational epitopes that were stabilized by cross-linking were also most sensitive to reduction by DTT (Fig. 6C). We conclude that BS3 helps occlude V3 and non-neutralizing epitopes while stabilizing the gp120-gp41 interface primarily through conformational restriction; however, BS3 shows little capacity to protect bNAb epitopes from susceptibility to Env reduction-induced disruption.
FIG 6.
The V3 crown is revealed upon cleavage of the cross-link, but bNAb epitopes on spikes are sensitive to reduction regardless of cross-linking. (A) JR-FL virions were cross-linked using BS3 or its disulfide-containing analog DTSSP. DTT breaks DTSSP linkages but does not affect BS3. Detergent-solubilized Env from virions was captured on GNL and probed in an ELISA. Full binding curves of a few representative antibodies are shown. (B) Virion-associated Env was treated as in panel A against a broader panel of bNAbs at a single concentration (1 μg/ml). The percent change in OD at 450 nm is plotted for each condition relative to untreated Env. (C) The effect of cross-linking on binding (fold change in EC50; Fig. 2) is plotted against the effect of DTT reduction (% of wild-type binding) for each antibody. Color-coding accords with class of antibody as follows: 2G12, black; CD4bs, brown; V3 crown, purple; MPER, blue; N332 supersite, red; PG9, orange; and gp120-gp41 interface, green. (D) Effect of DTT on binding to un-cross-linked Env versus BS3-cross-linked Env (% of wild-type binding) for each antibody is plotted. The dashed line denotes where antibodies would be plotted if cross-linking does not affect their binding to DTT-treated Env. Binding by antibodies plotted to the right of the dashed line is preserved when Env is cross-linked before DTT treatment, whereas binding by antibodies to the left of the line is inhibited by cross-linking. (E) Possible cross-links that could be formed at the trimer apex are shown on the crystal structure of BG505 SOSIP.664 (PDB ID 4TVP) (26). Only lysine residues that are shared between BG505 and JR-FL are shown. V1/V2 from the three gp120 protomers is shown in shades of green, and V3 of protomer A is shown in orange. The distance between the side chain primary amines of two lysine residues is labeled by the dotted line (blue, V1/V2-V1/V2; orange, V1-V3) and measured in angstroms. The subscript in the residue number indicates which gp120 protomer the residue belongs to. A combination of an intraprotomer cross-link between V3 and V1/V2 and an interprotomer cross-link between two V1/V2 regions could block accessibility to the V3 crown epitope. The measured distances are not absolute, since in solution the lysines can assume different rotameric states and the V1/V2/V3 loops can be conformationally flexible. BS3 is 11.4 Å long, and the amide bond formed upon cross-linking adds ∼1.5 Å to each end.
Cross-linker effects on bNAb epitopes on native trimer is distinct from that of soluble gp120 and uncleaved Env trimers.
We compared the effects of BS3 cross-linking on the exposure of bNAb epitopes on different states of Env in an ELISA involving cleaved viral spikes, uncleaved gp140 foldon trimers (gp140-FT) and monomeric gp120 (all JR-FL). In contrast to viral spikes in which binding by V3 crown antibodies decreased ∼10-fold due to BS3, V3 antibody binding to gp120 and gp140-FT was diminished only 2- to 3-fold (Fig. 7). N332 supersite bNAb binding to native spikes and soluble Env molecules increased 2- to 4-fold and decreased >10-fold, respectively. These results show that V3 and proximal N332 epitopes are fixed by BS3 on native spikes in a closed, prefusion conformation that is fundamentally different from that of soluble gp120 and uncleaved soluble gp140 molecules.
FIG 7.
BS3-cross-linking stabilizes a conformation of V3 and N332 supersite on native spikes that is distinct from that on soluble gp120 and uncleaved gp140. Effects of BS3 treatment on monomeric gp120, gp140-foldon (FT) trimers, and solubilized spikes (all JR-FL) as probed by antibodies are presented as the fold change in ELISA binding (EC50 cross-linked/EC50 untreated). nb, no binding. Statistically significant changes were determined using a two-tailed t test (*, P < 0.05; **, P < 0.01; ***, P < 0.001).
Purification of Env spikes away from Env debris using non-neutralizing antibody immunodepletion.
Nontrimeric and/or immature species of Env that bear non-neutralizing epitopes account for ∼5 to 20% of Env associated with JR-FL virions (10). To enhance purity, non-neutralizing antibodies D49 and b6, against gp41 and gp120, respectively, were used to sequentially deplete any reactive (non-native) Env molecules from detergent-solubilized virions (Fig. 8A). BN-PAGE analysis revealed that this immunodepletion step removed heterogeneous Env contaminants, leaving behind only trimeric spikes, whereas SDS-PAGE showed that uncleaved gp160 was also removed from virus lysates leaving behind cleaved Env (Fig. 8B and C). Moreover, ELISA showed that the binding of six different non-neutralizing antibodies was reduced to near background levels by immunodepletion (Fig. 8D). We conclude that immunodepletion of solubilized Env molecules using non-neutralizing antibodies can effectively remove contaminating species of Env from cross-linked virion preparations and eliminate binding of non-neutralizing antibodies against epitopes that do not overlap the depleting antibodies.
FIG 8.
Immunodepletion using non-neutralizing antibodies removes immature gp160 and other Env debris from cross-linked spike preparations. (A) A diagram demonstrating how non-native Env is removed by immunodepletion using non-neutralizing b6 and D49 antibodies. (B) Virions were cross-linked using cleavable BS3 analog, DTSSP. Detergent-solubilized non-native Env was depleted using b6, D49, or both sequentially. The DTSSP cross-link was reduced using DTT, and Env was resolved by SDS-PAGE and blotted with anti-gp120 antibodies. (C) JR-FL virions were BS3 cross-linked, solubilized in detergent, and then incubated sequentially with b6- and D49-coupled Sepharose beads. Immunodepleted and untreated samples were run side by side in a BN-PAGE gel mobility shift assay using Fabs b12 and b6. The Western blots shown were blotted with anti-gp41 antibodies. The arrow denotes the gp120-gp41 trimer. (D) Samples were analyzed for binding to a panel of neutralizing and non-neutralizing antibodies before or after being immunodepleted as described above using a GNL-capture ELISA. Shown is the average OD at 450 nm for each antibody at 10 μg/ml from at least three independent experiments. Binding by the negative-control antibody DEN3 has been subtracted.
C-terminal epitope tagging of Env spikes.
To facilitate enrichment, immobilization, and presentation of Env spikes at higher densities than that found on VLPs, such as might be advantageous for immunizations (74), a FLAG epitope tag was genetically fused to the C terminus of full-length gp160. FLAG-tagged JR-FL virions showed similar infectivity and Env expression levels as wild-type virions and replicated in MT2-CCR5 cells (Fig. 9A). Importantly, BN-PAGE showed that anti-FLAG tag antibodies recognized cognate Env trimers (Fig. 9B). BS3-cross-linked FLAG-tagged spikes were purified using the immunodepletion-affinity-purification scheme, as described above, with ∼80% yield, whereas p24 levels were undetectable (Fig. 9C). In addition, no contaminating proteins were detected using BN-PAGE and Coomassie blue staining (Fig. 9D).
FIG 9.
Cross-linked virion spikes can be purified using a FLAG epitope tag. (A) A FLAG tag was genetically fused to the C terminus of full-length gp160 JR-FL Env. FLAG-tagged virions were assayed for infectivity in a single-cycle infectivity assay using TZM-bl cells (solid bar), gp120 incorporation (normalized for p24 input and measured by GNL-capture ELISA; checkered bar), and replication in MT2-CCR5ΔCT cells (p24 equivalents at day 7 relative to wild type; hatched bar). (B) Gp160 FLAG-tagged virions were analyzed using BN-PAGE mobility shift assay using Fabs b12 and b6, as well as anti-FLAG IgG. The Western blot was probed using an anti-gp41 antibody cocktail. (C) JR-FL FLAG-tagged gp160 samples were probed in an ELISA using antibody to gp120 (left panel) and p24 (right panel) before and after immunoaffinity purification. (D) Affinity-purified BS3-cross-linked JR-FL trimers were run on BN-PAGE beside recombinant JR-FL gp120 (5 μg of each protein). The purity of the proteins were analyzed using Coomassie blue staining.
BS3-cross-linked Env spikes share architectural features with native Env trimers.
Purified BS3-cross-linked Env spikes were analyzed using negative-stain electron microscopy (EM) and reference-free 2D class averages were generated (Fig. 10A and B). The global architecture observed with the purified cross-linked spikes is similar to published EM structures of HIV-1 Env spikes (Fig. 10C) and shows the typical trilobed apex and a narrow stem region (25, 69, 75, 76). The globular structure at the base of the spike presumably corresponds to the CT of gp41 and detergent micelle, the latter of which is also observed with MPER-containing soluble Env trimers (69, 77, 78).
FIG 10.
Reference-free 2D class averaged EM reconstructions of BS3-cross-linked JR-FL spikes extracted from viral membrane. (A) EM image of BS3-cross-linked JR-FL Env spikes. (B) Select reference-free 2D class averages of cross-linked JR-FL Env trimers. In the enlarged image, gp120 and gp41 ectodomain regions are shaded blue, while the TM and CT domains encapsulated in a DDM micelle are shaded yellow. The white scale bar in the top left indicates 10 nm. (C) The X-ray crystal structure of the trimer ectodomain (gp120, cyan; gp41, blue; PDB ID 4TVP) (26) is shown for comparison. The predicted location of TM and CT are shown as a yellow circle.
Immunization using cross-linked spikes on VLPs and PLNs.
The production yields of virion-derived spikes were 2 to 3 orders of magnitude lower than that of typical soluble gp140s (79). To immunize with cross-linked spikes we therefore opted for a DNA prime-protein boost strategy that would be dose sparing (Fig. 11A). A plasmid encoding codon-optimized env from HIV-1 JR-CSF (92% sequence homology with JR-FL) was used for the DNA prime, since this plasmid DNA was found to be immunogenic in pilot studies (data not shown). Rabbits were immunized with two DNA primes and divided into four groups of three rabbits each. Groups A and B were boosted with un-cross-linked and BS3-cross-linked JR-FL VLPs, respectively. Groups C and D were boosted with purified cross-linked trimers in detergent (group C) or captured onto PLNs (see below; group D).
FIG 11.
Neutralization properties of sera from rabbits immunized using Env in a DNA prime-cross-linked spike boost format. (A) Schematic illustrating the major events in the immunization study. (B) Cross-linked spikes were captured onto 200-nm magnetic particles to generate PLNs. PLNs were analyzed by GNL-capture ELISA. Biotinylated DOPE was added at 1% of total lipid content for determination of lipid incorporation onto spike-bearing PLNs. A 5-fold dilution series of PLNs was added to GNL-coated microtiter wells and then probed using a constant concentration of anti-gp120 antibody cocktail (b12, 2G12, and F425-B4e8; left) or streptavidin (right). (C) Sera from day 0 (prebleed), day 42 (post-DNA prime), or day 98 (post-protein boost) were tested for neutralization activity against a panel of five well-characterized clade B isolates: JR-FL, JR-CSF, and ADA (tier 2); SF162 (tier 1a), and HxB2 (tier 1b). SIVmac239 and VSV-g were included as negative controls to test for nonspecific inhibition. Serum titers (IC50) are color-coded according to potency, as indicated. (D) Day 98 sera from groups A and D were tested further for neutralization activity against a reference panel of 12 clade B isolates. Serum titer IC50s are color-coded as in panel C. (E) Sera in groups A and D were compared for breadth (number of tested isolates neutralized) and potency of neutralization against SF162, HxB2, and JRCSF. The differences were not statistically significant (breadth, P = 0.19; SF162 potency, P = 0.15; HxB2 potency, P = 0.35; JRCSF potency, P = 0.36).
We prepared PLNs to present cross-linked Env as a particulate boosting agent and to prevent cross-linked spikes from aggregating via exposed hydrophobic MPER and TM domains (27), as well as to enhance valency of Env display. Cross-linked spikes were captured onto nanobeads conjugated to anti-FLAG antibody (80), and lipids were then added at a ratio that mimics that of the native viral membrane (68). The PLNs used for immunization were produced using 200-nm magnetic beads; these virus-like dimensions allow the PLNs access to all areas of the lymph node and can stimulate B cell responses more efficiently than micron-sized particles (81). ELISA analysis confirmed incorporation of Env spikes and lipids into PLNs (Fig. 11B).
PLNs but not VLPs elicit distinct Tier 2 isolate neutralizing antibodies.
Not unexpectedly, sera from group A rabbits (VLP boost) neutralized only the sensitive Tier 1a strain, SF162 (50% inhibitory concentrations [IC50s] = 1,000 to 3,900 dilutions) (Fig. 11C). Group B sera (BS3-cross-linked VLP boost) and group C sera (BS3-cross-linked purified trimer boost) neutralized SF162 on average 2 orders of magnitude less potently (IC50 = 8 to 38 dilutions), indicating that cross-linking Env decreased immunogenicity of Tier 1a-restricted epitope(s). Notably, group D sera (cross-linked trimer PLN boost) exhibited a different neutralization pattern that shifted from Tier 1a neutralization to Tier 1b/Tier 2 neutralization. In addition to some SF162 neutralizing activity, serum 7547 showed potent neutralization against the Tier 2 isolate JR-CSF. Serum 7549 potently neutralized the Tier 1b strain HxB2, which we and others observed relatively infrequently compared to SF162 neutralization (82; data not shown). Finally, serum 7548 displayed negligible neutralization potential. Collectively, these results show that BS3-cross-linking JR-FL Env spikes redirects antibody responses from those that neutralize only a Tier 1a sensitive strain to those that can neutralize, at least sporadically, more resistant isolates.
Sera from groups A and D were further compared against a larger panel of 12 clade B isolates using the TZM-bl assay. Cross-linked trimer PLN boosted sera (group D) 7549 and 7547 showed a modest but consistent neutralization (IC50 = 5 to 11 dilution) against 8/12 and 2/12 isolates, respectively, whereas none of the VLP boosted sera (group A) showed any activity against the panel (Fig. 11D). Hence, boosting with cross-linked trimer PLNs appeared to elicit more broadly active, but quite weak, neutralizing antibodies to various clade B isolates than VLPs, although the differences were not statistically significant (Fig. 11E).
Neutralization of SF162 is frequently mediated by antibodies to the V3 crown that are elicited readily by monomeric gp120 but do not generally neutralize Tier 2 isolates. Indeed, addition of a soluble V3 peptide blocked a majority of neutralization of SF162 by group A sera (60 to 83%), whereas neutralization of SF162 and JR-CSF by group D sera was less affected (39 to 54% and ∼20%, respectively; Table 1). Thus, cross-linking Env likely reduces immunogenicity of the V3 crown, thereby diminishing neutralization of strains sensitive to V3 antibodies.
TABLE 1.
Competition of HIV-1 neutralizing activity from sera using V3 peptide (305KRIHIGPGRAFYTTK321)
| Group | Serum ID | % Competition of neutralization |
||
|---|---|---|---|---|
| SF162 | HxB2 | JR-CSF | ||
| A | 7538 | 60 | NAa | NA |
| 7539 | 83 | NA | NA | |
| 7540 | 60 | NA | 16 | |
| D | 7547 | 54 | NA | 17 |
| 7548 | NA | –2 | NA | |
| 7549 | 39 | –29 | 22 | |
NA, not applicable because the serum did not neutralize the corresponding virus.
To determine the antibody specificities in serum 7547 responsible for potent neutralization of JR-CSF, we used a panel of JR-CSF or JR-FL virions in which major domains of gp120 have been substituted (83). Substitutions of C3 or V5 JR-FL into JR-CSF abrogated neutralization, and reciprocal introduction of JR-CSF C3 and V5 together (but not individually) into JR-FL raised neutralization levels to that observed against JR-CSF (Fig. 12A and B). Other domain swaps had no effect. In addition, the glycan knockout mutation N301Q, which frequently enhances neutralization (83), and replacement of central sequences in V4 with Gly or Ser significantly increased JR-CSF neutralization, whereas substitutions in V5 abrogated neutralization (Fig. 12C and E). We conclude that neutralization of JR-CSF by serum 7547 is largely V5 and C3 dependent and that V4 appears to interfere with antibody recognition of this epitope in some way. With the other PLN-boosted sera, 7548 and 7549, N301Q also enhanced their weak neutralization of JR-CSF ∼10-fold; however, in contrast to 7547, a V4 Gly-Ser mutation reduced neutralization (Fig. 12D).
FIG 12.
Neutralizing antibodies in rabbit 7547 target C3 and V5 on JR-CSF gp120. Day 98 serum from rabbit 7547 was tested for neutralization against a panel of gp120 “domain swap” pseudotyped viruses that contain substitutions of major domains from JR-FL gp120 into a JR-CSF backbone (A), major domains from JR-CSF gp120 into a JR-FL backbone (B), and V4 or V5 with substitutions to Gly/Ser residues or the mutation N301Q (C). (D) Sera 7548 and 7549 were also tested for neutralization of JR-CSF mutants V4-GGS33 and N301Q. (E) Sequence changes introduced into V4 and V5 in the mutant Envs used in panels C and D.
To determine whether neutralizing antibodies were elicited against the CD4bs, sera were absorbed using the probe TriMut, or control probe TriMut368/370, prior to addition to virus (66). TriMut is based on the gp120 core and contains three mutations that prevent binding to CD4, but not to most CD4bs antibodies, whereas TriMut368/370 contains two mutations that also prevent binding by most CD4bs antibodies (66). Most of the neutralization by PLN-boosted sera 7549 and 7548 against HxB2 and SF162 was depleted by TriMut, similar to control antibody VRC01 (Fig. 13). However, neutralization mediated by 7549 against JRCSF, TRO.11, and SC422661 was not affected by TriMut (Fig. 13A). Neutralization by PLN-boosted serum 7547 and VLP-boosted serum 7540 was also unaffected by TriMut. Thus, TriMut cross-reactive CD4bs antibodies were elicited in two of the PLN-boosted sera, but they appear to only neutralize sensitive Tier 1 isolates and do not appear to constitute the antibody specificities mediating broad but weak Tier 2 neutralization in serum 7549.
FIG 13.
CD4 binding site antibodies in PLN-boosted sera 7549 and 7548 neutralize tier 1 and not tier 2 isolates. (A) Sera were incubated with TriMut core or TriMut368/370 at 10 μg/ml, or with an equivalent volume of medium, for 1 h at 37°C. Sera were then tested for neutralization of HxB2, JR-CSF, SF162, TRO.11, and SC422661. (B) Control CD4bs antibody VRC01 was tested in the assay described above. Similar results with VRC01 were observed with all tested isolates.
Antibody binding specificities boosted by HIV-1 spikes.
The immune sera were also assayed by ELISA against a panel of antigens to look for binding patterns associated with cross-linking the Env immunogen. All VLP-boosted (group A) sera reacted very strongly with monomeric gp120, as expected, whereas gp120 binding titers with PLN-boosted (group D) sera were lower and more variable (Fig. 14A). Serum binding titers to recombinant gp41 and un-cross-linked solubilized Env mirrored that of gp120, but when tested against purified cross-linked spikes the group A and D sera showed equal binding titers. Group B and C antibody titers were not boosted against any of the tested antigens, indicating a lack of immunogenicity. Off-target binding to BS3 cross-links rather than to Env, measured by ELISA binding to BS3-cross-linked bovine serum albumin, was only observed with sera from group B, perhaps because BS3-cross-linked VLPs may present a relatively high number and diversity of BS3 epitopes in the immunogen.
FIG 14.
Binding properties of sera from immunized rabbits. (A) Binding of immune serum against a panel of antigens, including recombinant gp120 and gp41, detergent-dissociated JR-FL viral Env, purified cross-linked spikes, and BS3-cross-linked bovine serum albumin (BSA) as determined by ELISA. All Env molecules were from JR-FL. The data points correspond to individual sera from day 42 (○) and day 98 (◼), and cognate means are shown as dashed and solid lines, respectively. Statistically significant changes were determined using a two-tailed Student t test (*, P < 0.05; **, P < 0.01; ***, P < 0.001). (B) Group A and D sera were tested for binding in ELISA to the scaffold RSC3 that was designed to specifically bind neutralizing CD4bs antibodies. (C) Group A and D sera were tested for the ability to block binding by neutralizing and non-neutralizing antibodies to monomeric JR-FL gp120 or gp41. The IC50 of each serum against each antibody is shown. (D) Serum binding to gp120 V3 and gp41 DSL peptides was determined by using ELISA. Statistical significance was determined as in panel A. (E) Relationship between binding of day 98 sera (EC50) to V3 peptide versus neutralization (IC50) of SF162 virus.
To identify antibodies in the sera to particular epitopes, several assays were used. First, sera were tested for CD4bs antibodies using the scaffold RSC3 that specifically binds CD4bs bNAbs (53). Several sera bound to RSC3, most notably 7549 and 7540 (Fig. 14B), although binding was weak relative to VRC01 and neutralization was adsorbed using TriMut only with serum 7549 (Fig. 13A). In addition, sera were tested in a competition ELISA to determine whether they could block representative bNAbs from binding to recombinant JR-FL gp120 or gp41. All of the sera blocked b12 binding, suggesting the presence of CD4bs antibodies; however, competition was weaker against the more potent bNAb VRC01 in which only serum 7549 achieved 50% inhibition (Fig. 14C). In the competition ELISA, sera that potently neutralized SF162 (i.e., VLP boosted sera and serum 7547) competed with PGT128 for binding to gp120 (Fig. 14C); however, none of the sera reacted with a scaffold, glycan V3-ST09, designed to display the N332 supersite (84). Hence, the PGT128 competition we observed most likely relates to the ability of V3 crown antibodies to sterically block the N332 supersite on monomeric gp120 (54).
Antibodies against immunodominant V3 and gp41 disulfide loop (DSL) epitopes were detected using a peptide ELISA. Binding titers to V3 were higher in sera that potently neutralized SF162 and those same sera also potently blocked F425-B4e8 binding to gp120 (Fig. 14C and D). Notably, binding titers to V3 peptide showed a statistically significant correlation with SF162 neutralization (Fig. 14E). Moreover, reactivity to gp41 DSL peptides was notably high with VLP boosted (group A) sera that also blocked anti-DSL IgG D49 from binding to gp41; however, sera from animals boosted with D49-depleted cross-linked spikes showed little binding to DSL and did not block D49 binding. (Fig. 14C and D). Thus, non-neutralizing antibody immunodepletion and PLN display of cross-linked Env spikes (group D) not only reduced immunodominant antibody responses to gp120 (V3) and gp41 (DSL) but also directed responses to novel, apparently less immunogenic epitopes.
DISCUSSION
Virion-displayed HIV-1 spikes are important vaccine candidates as, by definition, they embody Env in its native form, but they also suffer from inherent lability, heterogeneity and low copy number on virion particles. Here, we used cross-linker to fix native spikes, which largely preserves bNAb epitopes, as well as occludes the V3 crown and non-neutralizing epitopes. We show that mature spikes can be purified and separated from non-native forms of Env using non-neutralizing antibody immunodepletion and can be captured onto nanoparticles at high valency for immunization. Although the overarching goal of consistently eliciting potent and broad neutralizing responses remains to be reached, cross-linked spikes on PLNs did elicit intriguing neutralizing antibody responses employing strategies that seem eligible for improvement.
BS3-stabilized spikes display most bNAb epitopes, including the N332 supersite, gp41 MPER, PG9/PG16 epitopes on V2, and 35O22 and PGT151 epitopes at the gp120-gp41 interface. The latter two quaternary epitopes, as well as the N332 supersite, were shown in binding assays to be enhanced relative to un-cross-linked Env. The reactivity of certain CD4bs bNAbs was slightly reduced by cross-linking spikes, perhaps because the cross-links prevent small conformational changes induced by these antibodies (76). Nevertheless, we observed antibodies to the CD4bs in antisera boosted by Env spikes, including some that neutralized Tier 1 isolates, suggesting that the CD4bs on cross-linked spikes remained immunogenic.
The V3 crown is a site of neutralization on sensitive isolates of HIV-1 and was not only occluded by cross-linking, but its immunogenicity was also reduced. This provides a clear example of a modification to an immunogen that attenuates an immunodominant antibody response. V3 has been shown to become partially blocked by glutaraldehyde treatment of virion-displayed Env but not soluble gp140 trimers (32). We observed a decrease in V3 antibody binding following cross-linking that was much greater with virion-displayed Env than with soluble gp120 or gp140-FT, showing that native Env affords a unique context for V3. The recovery of F425-B4e8 binding to trimers cross-linked using the cleavable cross-linker, DTSSP, further provides direct biochemical evidence that the V3 crown is occluded on native Env from antibody via quaternary packing. Since V1V2 can shield access to V3 on Env trimers of primary isolates (24, 85, 86), and V1 covers V3 in the recent crystal structure of BG505 SOSIP (26), cross-linking may “staple” V1V2 to V3 in a quaternary context that is absent on soluble monomers and uncleaved soluble trimers. PGT121, PGT126, and PGT128 recognize epitopes involving glycans at or near N332, as well as protein elements at the base of V3, using different angles of approach (54). BS3 seems to stabilize the entire N332 epitope cluster in a uniquely accessible conformation on native Env. In contrast, binding of N332 antibodies to monomeric gp120 and gp140-FT trimers was decreased by BS3 cross-linking, indicating that the N332 epitope cluster is presented differently on these molecules than in the native conformation, potentially making them less desirable for immunizations.
Formalin-fixed virions have been used previously to immunize mice, but spikes were in low copy number and were not purified away from misfolded Env or host proteins (38, 39). Soluble gp120 and gp140 have been cross-linked in complex with CD4 for immunogens that mimic a receptor-activated form of Env (35). More recently, glutaraldehyde-treated, uncleaved gp140 was shown to elicit improved neutralization of Tier 1 isolates, most likely due to increased levels of weakly neutralizing CD4bs antibodies, but did not improve neutralization of tier 2 isolates (36). To our knowledge, the present study is the first that uses cross-linked cleaved Env spikes purified from HIV-1 virions as an immunogen.
Importantly, we show that immunodepletion using non-neutralizing antibodies efficiently eliminates immature gp160 and other misfolded Env debris from virion preparations. Non-neutralizing antibodies that bind to immunodominant “cluster” epitopes on gp41 (e.g., DSL and six helix bundle), as well as to structurally ill-defined epitopes on the inner domain of gp120, V1V2, V3, and others, are commonly elicited both during acute natural infection and by gp120 and gp140 immunizations (87–90). Immunizations using untreated VLPs in the present study yielded antibodies of this type that only neutralized the sensitive strain SF162. Cross-linking and immunodepletion complemented one another in eliminating non-native forms of Env: cross-linking prevents decay of trimeric Env, whereas immunodepletion using different non-neutralizing antibodies enriches for Env that is trimeric, cleaved, and more native-like.
Cross-linking, however, did reduce overall binding titers to Env (i.e., immunogenicity) elicited in VLP-boosted animals. One explanation is that cross-linking Env negatively impacted processing into peptides for major histocompatibility complex (MHC) display and thus reduced the level of T cell help that developing B cells need (91). This might be overcome by adding T cell epitopes (92), adjuvants, limiting cross-linker density, and increasing antigen dose. In addition, the concentration of BS3 used here (1 mM) was chosen because it prevented any trimer dissociation after heat treatment, but lower concentrations might be used that would increase antigen processing. Alternatively, limiting exposure of immunodominant epitopes (e.g., V3 and DSL) might have negatively impacted the total antibody response. For example, site-directed hyperglycosylation of V3 on soluble gp140 trimers was shown to reduce V3 specific antibody responses but did not elicit a stronger response to other regions of Env (93). A third possibility is that cross-linking restricts conformational flexibility of the spike such that it triggers fewer germ line/naive B cell receptors (BCRs). This effect might be overcome with modifications to Env, such as removing glycans that restrict access to certain epitopes (94), or by screening for mutations that might increase reactivity to germ line antibodies despite a loss of conformational flexibility.
Despite eliciting relatively modest titers of antibodies specific to Env, cross-linked spike PLNs elicited a detectably broader and more diverse neutralization response than that of VLPs. Due to the small size of the study, the differences were not statistically significant, but since we did not observe any neutralization by prebleed sera or neutralization of negative virus controls, the qualitative differences elicited by cross-linked trimer PLNs seem likely to be real. Although PLN-immunized rabbits received 2-fold more total Env than VLP-immunized rabbits, the anti-Env ELISA titers were no higher and cross-linked spikes alone at the same dose did not appear to be immunogenic. Particles often enhance immunogenicity relative to soluble proteins (95). Thus, PLN display of Env spikes in higher density than VLPs and HIV-1 may have enhanced engagement of more diverse BCRs through avidity effects and/or improved local stimulation of immune cells (e.g., T cells). The diminished levels of antibody elicited to the V3 crown by cross-linked spikes may have unmasked a B cell response to conserved epitopes nearby. Whatever the reason for the increased response with group D sera, it seems a reasonable path forward for native trimer immunization to employ some form of particle display. In the present study, we chose to prime with DNA and then boost once with cross-linked Env protein because the yield of Env trimers purified from virions is exceedingly low, under the theory that trimers stabilized in the correct conformation would only boost BCRs that recognize the native Env spike and not other BCRs elicited by the DNA prime. Giving animals multiple boosts of cross-linked trimers might further expand the weak neutralization observed in some individuals here. On the other hand, priming with DNA that produces un-cross-linked Env may actually increase triggering of CD4+ T cells relative to cross-linked trimer only immunization, since cross-linking may limit antigen processing and display by MHC class II. Improvements to immunizations with cross-linked spikes will likely need to be determined empirically.
One serum elicited by the PLN boost (7547) displayed relatively potent neutralization against tier 2 isolate JR-CSF. Domain swap neutralization experiments showed the antibody response was to an epitope involving V5 and C3 of gp120 rather than the more typical V1V2V3 epitopes. Potent neutralization of JR-CSF involving V4 and V5 has been observed previously (83), suggesting that these regions of Env may be frequent targets of autologous neutralization. Encouragingly, single residue changes were sufficient to knock out these epitopes in the present study, which may be exploited to redirect immunization responses onto more conserved regions.
Group D serum 7549 neutralized HxB2 relatively potently and displayed some breadth against the clade B panel. This activity only appeared after the PLN boost. The neutralizing activity of 7549 against HxB2 and SF162 was largely depleted using the CD4bs probe TriMut, showing that neutralization of these tier 1 isolates was mediated by CD4bs antibodies. However, neutralization of Tier 2 isolates JR-CSF, TRO.11, and SC422661 was not altered by TriMut depletion. Interestingly, neutralization of JR-CSF by serum 7549 was mostly abrogated by mutations to V4, so the broad but weak neutralization may be mediated by antibodies to an epitope near V4. Weak neutralization of JR-CSF by serum 7548 was also diminished by V4 mutation, so antibodies of this specificity may have been elicited at low levels in two of three PLN-boosted sera.
Surprisingly, boosting with JR-FL Env did not elicit antibodies able to neutralize autologous virus. This may be a case of “original antigenic sin” (96), whereby B cells triggered by JR-CSF in the DNA prime were boosted by low affinity engagement of BCRs amplified by multivalent presentation of JR-FL spikes on the PLNs (avidity effect). It is also possible that JR-CSF Env produced in the DNA prime persisted at low levels (97) and JR-FL spikes on PLNs boosted antibody titers via enhanced T cell help, or, alternatively, cross-linked spikes may present novel epitopes in vivo to immune cells through an unappreciated mechanism. Finally, BS3 produces random Lys-Lys cross-links in Env so it is possible that the B cell response triggered by a given cross-linked trimer may not recognize a significant proportion of the total cross-linked trimer population, in which case cognate binding may be underestimated. These issues may be addressed by use of site-specific cross-linkers (98–101) or genetic stabilization (23). Identification of single B cells (and possibly T cells) against improved trimeric immunogens will allow a higher resolution for addressing these issues.
Cross-linking native Env, immunodepleting misfolded Env contaminants, and purifying cross-linked spikes for nanoparticle-based immunization expands the methods available for native Env immunization and appears to redirect and slightly diversify HIV-1 neutralizing antibody responses in comparison to VLPs. Future directions will be aimed at determining whether more consistent and potent neutralizing antibody responses can be elicited by using (i) specific cross-linking or genetic stabilization strategies, (ii) Envs capable of binding germ line precursors of neutralizing antibodies, (iii) higher Env doses, and (iv) a more ordered display of trimers.
ACKNOWLEDGMENTS
We are grateful to Phil Dawson and Syna Gift for helpful discussions, as well as Arthur Kim and Katie Bauer for technical assistance. We thank Robert Whalen for the generous donation of env plasmids. We acknowledge Richard Wyatt for the TriMut and gp140-foldon constructs and Dennis Burton and Mark Connors for providing antibodies.
Funding was provided by the National Institutes of Health/National Institute of Allergy and Infectious Diseases (R01 AI093287 and R01 AI098602), the International AIDS Vaccine Initiative (IAVI) Neutralizing Antibody Center and the Collaboration for AIDS Vaccine Discovery (CAVD), The Bill and Melinda Gates Foundation, Scripps CHAVI-ID (UM1 AI100663), and a California HIV/AIDS Research Program Dissertation Fellowship.
REFERENCES
- 1.Mascola JR, Montefiori DC. 2010. The role of antibodies in HIV vaccines. Annu Rev Immunol 28:413–444. doi: 10.1146/annurev-immunol-030409-101256. [DOI] [PubMed] [Google Scholar]
- 2.Walker LM, Burton DR. 2010. Rational antibody-based HIV-1 vaccine design: current approaches and future directions. Curr Opin Immunol 22:358–366. doi: 10.1016/j.coi.2010.02.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Stephenson KE, Barouch DH. 2013. A global approach to HIV-1 vaccine development. Immunol Rev 254:295–304. doi: 10.1111/imr.12073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wyatt R, Sodroski J. 1998. The HIV-1 envelope glycoproteins: fusogens, antigens, and immunogens. Science 280:1884–1888. doi: 10.1126/science.280.5371.1884. [DOI] [PubMed] [Google Scholar]
- 5.Zwick MB, Burton DR. 2007. HIV-1 neutralization: mechanisms and relevance to vaccine design. Curr HIV Res 5:608–624. doi: 10.2174/157016207782418443. [DOI] [PubMed] [Google Scholar]
- 6.Zhu P, Liu J, Bess J Jr, Chertova E, Lifson JD, Grise H, Ofek GA, Taylor KA, Roux KH. 2006. Distribution and three-dimensional structure of AIDS virus envelope spikes. Nature 441:847–852. doi: 10.1038/nature04817. [DOI] [PubMed] [Google Scholar]
- 7.Ota T, Doyle-Cooper C, Cooper AB, Huber M, Falkowska E, Doores KJ, Hangartner L, Le K, Sok D, Jardine J, Lifson J, Wu X, Mascola JR, Poignard P, Binley JM, Chakrabarti BK, Schief WR, Wyatt RT, Burton DR, Nemazee D. 2012. Anti-HIV B Cell lines as candidate vaccine biosensors. J Immunol 189:4816–4824. doi: 10.4049/jimmunol.1202165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Crooks ET, Moore PL, Franti M, Cayanan CS, Zhu P, Jiang P, de Vries RP, Wiley C, Zharkikh I, Schulke N, Roux KH, Montefiori DC, Burton DR, Binley JM. 2007. A comparative immunogenicity study of HIV-1 virus-like particles bearing various forms of envelope proteins, particles bearing no envelope, and soluble monomeric gp120. Virology 366:245–262. doi: 10.1016/j.virol.2007.04.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.West AP Jr, Scharf L, Scheid JF, Klein F, Bjorkman PJ, Nussenzweig MC. 2014. Structural insights on the role of antibodies in HIV-1 vaccine and therapy. Cell 156:633–648. doi: 10.1016/j.cell.2014.01.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Leaman DP, Kinkead H, Zwick MB. 2010. In-solution virus capture assay helps deconstruct heterogeneous antibody recognition of human immunodeficiency virus type 1. J Virol 84:3382–3395. doi: 10.1128/JVI.02363-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Moore PL, Crooks ET, Porter L, Zhu P, Cayanan CS, Grise H, Corcoran P, Zwick MB, Franti M, Morris L, Roux KH, Burton DR, Binley JM. 2006. Nature of nonfunctional envelope proteins on the surface of human immunodeficiency virus type 1. J Virol 80:2515–2528. doi: 10.1128/JVI.80.5.2515-2528.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Poignard P, Moulard M, Golez E, Vivona V, Franti M, Venturini S, Wang M, Parren PW, Burton DR. 2003. Heterogeneity of envelope molecules expressed on primary human immunodeficiency virus type 1 particles as probed by the binding of neutralizing and nonneutralizing antibodies. J Virol 77:353–365. doi: 10.1128/JVI.77.1.353-365.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Liao HX, Chen X, Munshaw S, Zhang R, Marshall DJ, Vandergrift N, Whitesides JF, Lu X, Yu JS, Hwang KK, Gao F, Markowitz M, Heath SL, Bar KJ, Goepfert PA, Montefiori DC, Shaw GC, Alam SM, Margolis DM, Denny TN, Boyd SD, Marshal E, Egholm M, Simen BB, Hanczaruk B, Fire AZ, Voss G, Kelsoe G, Tomaras GD, Moody MA, Kepler TB, Haynes BF. 2011. Initial antibodies binding to HIV-1 gp41 in acutely infected subjects are polyreactive and highly mutated. J Exp Med 208:2237–2249. doi: 10.1084/jem.20110363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ruprecht CR, Krarup A, Reynell L, Mann AM, Brandenberg OF, Berlinger L, Abela IA, Regoes RR, Gunthard HF, Rusert P, Trkola A. 2011. MPER-specific antibodies induce gp120 shedding and irreversibly neutralize HIV-1. J Exp Med 208:439–454. doi: 10.1084/jem.20101907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Agrawal N, Leaman DP, Rowcliffe E, Kinkead H, Nohria R, Akagi J, Bauer K, Du SX, Whalen RG, Burton DR, Zwick MB. 2011. Functional stability of unliganded envelope glycoprotein spikes among isolates of human immunodeficiency virus type 1 (HIV-1). PLoS One 6:e21339. doi: 10.1371/journal.pone.0021339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Moore JP, McKeating JA, Weiss RA, Sattentau QJ. 1990. Dissociation of gp120 from HIV-1 virions induced by soluble CD4. Science 250:1139–1142. doi: 10.1126/science.2251501. [DOI] [PubMed] [Google Scholar]
- 17.Kim AS, Leaman DP, Zwick MB. 2014. Antibody to gp41 MPER alters functional properties of HIV-1 Env without complete neutralization. PLoS Pathog 10:e1004271. doi: 10.1371/journal.ppat.1004271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Sanders RW, Vesanen M, Schuelke N, Master A, Schiffner L, Kalyanaraman R, Paluch M, Berkhout B, Maddon PJ, Olson WC, Lu M, Moore JP. 2002. Stabilization of the soluble, cleaved, trimeric form of the envelope glycoprotein complex of human immunodeficiency virus type 1. J Virol 76:8875–8889. doi: 10.1128/JVI.76.17.8875-8889.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Chakrabarti BK, Kong WP, Wu BY, Yang ZY, Friborg J, Ling X, King SR, Montefiori DC, Nabel GJ. 2002. Modifications of the human immunodeficiency virus envelope glycoprotein enhance immunogenicity for genetic immunization. J Virol 76:5357–5368. doi: 10.1128/JVI.76.11.5357-5368.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Yang X, Farzan M, Wyatt R, Sodroski J. 2000. Characterization of stable, soluble trimers containing complete ectodomains of human immunodeficiency virus type 1 envelope glycoproteins. J Virol 74:5716–5725. doi: 10.1128/JVI.74.12.5716-5725.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zhang CW, Chishti Y, Hussey RE, Reinherz EL. 2001. Expression, purification, and characterization of recombinant HIV gp140. The gp41 ectodomain of HIV or simian immunodeficiency virus is sufficient to maintain the retroviral envelope glycoprotein as a trimer. J Biol Chem 276:39577–39585. [DOI] [PubMed] [Google Scholar]
- 22.Forsell MN, Schief WR, Wyatt RT. 2009. Immunogenicity of HIV-1 envelope glycoprotein oligomers. Curr Opin HIV AIDS 4:380–387. doi: 10.1097/COH.0b013e32832edc19. [DOI] [PubMed] [Google Scholar]
- 23.Leaman DP, Zwick MB. 2013. Increased functional stability and homogeneity of viral envelope spikes through directed evolution. PLoS Pathog 9:e1003184. doi: 10.1371/journal.ppat.1003184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Julien JP, Cupo A, Sok D, Stanfield RL, Lyumkis D, Deller MC, Klasse PJ, Burton DR, Sanders RW, Moore JP, Ward AB, Wilson IA. 2013. Crystal structure of a soluble cleaved HIV-1 envelope trimer. Science 342:1477–1483. doi: 10.1126/science.1245625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lyumkis D, Julien JP, de Val N, Cupo A, Potter CS, Klasse PJ, Burton DR, Sanders RW, Moore JP, Carragher B, Wilson IA, Ward AB. 2013. Cryo-EM structure of a fully glycosylated soluble cleaved HIV-1 envelope trimer. Science 342:1484–1490. doi: 10.1126/science.1245627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Pancera M, Zhou T, Druz A, Georgiev IS, Soto C, Gorman J, Huang J, Acharya P, Chuang GY, Ofek G, Stewart-Jones GB, Stuckey J, Bailer RT, Joyce MG, Louder MK, Tumba N, Yang Y, Zhang B, Cohen MS, Haynes BF, Mascola JR, Morris L, Munro JB, Blanchard SC, Mothes W, Connors M, Kwong PD. 2014. Structure and immune recognition of trimeric pre-fusion HIV-1 Env. Nature 514:455–461. doi: 10.1038/nature13808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Depetris RS, Julien JP, Khayat R, Lee JH, Pejchal R, Katpally U, Cocco N, Kachare M, Massi E, David KB, Cupo A, Marozsan AJ, Olson WC, Ward AB, Wilson IA, Sanders RW, Moore JP. 2012. Partial enzymatic deglycosylation preserves the structure of cleaved recombinant HIV-1 envelope glycoprotein trimers. J Biol Chem 287:24239–24254. doi: 10.1074/jbc.M112.371898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Harris A, Borgnia MJ, Shi D, Bartesaghi A, He H, Pejchal R, Kang YK, Depetris R, Marozsan AJ, Sanders RW, Klasse PJ, Milne JL, Wilson IA, Olson WC, Moore JP, Subramaniam S. 2011. Trimeric HIV-1 glycoprotein gp140 immunogens and native HIV-1 envelope glycoproteins display the same closed and open quaternary molecular architectures. Proc Natl Acad Sci U S A 108:11440–11445. doi: 10.1073/pnas.1101414108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Dey AK, David KB, Lu M, Moore JP. 2009. Biochemical and biophysical comparison of cleaved and uncleaved soluble, trimeric HIV-1 envelope glycoproteins. Virology 385:275–281. doi: 10.1016/j.virol.2008.12.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Finnegan CM, Berg W, Lewis GK, DeVico AL. 2001. Antigenic properties of the human immunodeficiency virus envelope during cell-cell fusion. J Virol 75:11096–11105. doi: 10.1128/JVI.75.22.11096-11105.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sattentau QJ. 1995. Conservation of HIV-1 gp120 neutralizing epitopes after formalin inactivation. AIDS 9:1383–1385. doi: 10.1097/00002030-199512000-00017. [DOI] [PubMed] [Google Scholar]
- 32.Yuan W, Bazick J, Sodroski J. 2006. Characterization of the multiple conformational states of free monomeric and trimeric human immunodeficiency virus envelope glycoproteins after fixation by cross-linker. J Virol 80:6725–6737. doi: 10.1128/JVI.00118-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Yuan W, Li X, Kasterka M, Gorny MK, Zolla-Pazner S, Sodroski J. 2009. Oligomer-specific conformations of the human immunodeficiency virus (HIV-1) gp41 envelope glycoprotein ectodomain recognized by human monoclonal antibodies. AIDS Res Hum Retroviruses 25:319–328. doi: 10.1089/aid.2008.0213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Haim H, Salas I, Sodroski J. 2013. Proteolytic processing of the human immunodeficiency virus envelope glycoprotein precursor decreases conformational flexibility. J Virol 87:1884–1889. doi: 10.1128/JVI.02765-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Fouts T, Godfrey K, Bobb K, Montefiori D, Hanson CV, Kalyanaraman VS, DeVico A, Pal R. 2002. Cross-linked HIV-1 envelope-CD4 receptor complexes elicit broadly cross-reactive neutralizing antibodies in rhesus macaques. Proc Natl Acad Sci U S A 99:11842–11847. doi: 10.1073/pnas.182412199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Schiffner T, Kong L, Duncan CJ, Back JW, Benschop JJ, Shen X, Huang PS, Stewart-Jones GB, DeStefano J, Seaman MS, Tomaras GD, Montefiori DC, Schief WR, Sattentau QJ. 2013. Immune focusing and enhanced neutralization induced by HIV-1 gp140 chemical cross-linking. J Virol 87:10163–10172. doi: 10.1128/JVI.01161-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lewis GK, Fouts TR, Ibrahim S, Taylor BM, Salkar R, Guan Y, Kamin-Lewis R, Devico AL. 2011. Identification and characterization of an immunogenic hybrid epitope formed by both HIV gp120 and human CD4 proteins. J Virol 85:13097–13104. doi: 10.1128/JVI.05072-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Poon B, Hsu JF, Gudeman V, Chen IS, Grovit-Ferbas K. 2005. Formaldehyde-treated, heat-inactivated virions with increased human immunodeficiency virus type 1 env can be used to induce high-titer neutralizing antibody responses. J Virol 79:10210–10217. doi: 10.1128/JVI.79.16.10210-10217.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Poon B, Safrit JT, McClure H, Kitchen C, Hsu JF, Gudeman V, Petropoulos C, Wrin T, Chen IS, Grovit-Ferbas K. 2005. Induction of humoral immune responses following vaccination with envelope-containing, formaldehyde-treated, thermally inactivated human immunodeficiency virus type 1. J Virol 79:4927–4935. doi: 10.1128/JVI.79.8.4927-4935.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Esser MT, Graham DR, Coren LV, Trubey CM, Bess JW Jr, Arthur LO, Ott DE, Lifson JD. 2001. Differential incorporation of CD45, CD80 (B7-1), CD86 (B7-2), and major histocompatibility complex class I and II molecules into human immunodeficiency virus type 1 virions and microvesicles: implications for viral pathogenesis and immune regulation. J Virol 75:6173–6182. doi: 10.1128/JVI.75.13.6173-6182.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sullivan N, Sun Y, Binley J, Lee J, Barbas CF III, Parren PW, Burton DR, Sodroski J. 1998. Determinants of human immunodeficiency virus type 1 envelope glycoprotein activation by soluble CD4 and monoclonal antibodies. J Virol 72:6332–6338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Zwick MB, Kelleher R, Jensen R, Labrijn AF, Wang M, Quinnan GV Jr, Parren PW, Burton DR. 2003. A novel human antibody against human immunodeficiency virus type 1 gp120 is V1, V2, and V3 loop dependent and helps delimit the epitope of the broadly neutralizing antibody immunoglobulin G1 b12. J Virol 77:6965–6978. doi: 10.1128/JVI.77.12.6965-6978.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Li M, Gao F, Mascola JR, Stamatatos L, Polonis VR, Koutsoukos M, Voss G, Goepfert P, Gilbert P, Greene KM, Bilska M, Kothe DL, Salazar-Gonzalez JF, Wei X, Decker JM, Hahn BH, Montefiori DC. 2005. Human immunodeficiency virus type 1 env clones from acute and early subtype B infections for standardized assessments of vaccine-elicited neutralizing antibodies. J Virol 79:10108–10125. doi: 10.1128/JVI.79.16.10108-10125.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wei X, Decker JM, Liu H, Zhang Z, Arani RB, Kilby JM, Saag MS, Wu X, Shaw GM, Kappes JC. 2002. Emergence of resistant human immunodeficiency virus type 1 in patients receiving fusion inhibitor (T-20) monotherapy. Antimicrob Agents Chemother 46:1896–1905. doi: 10.1128/AAC.46.6.1896-1905.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Trkola A, Purtscher M, Muster T, Ballaun C, Buchacher A, Sullivan N, Srinivasan K, Sodroski J, Moore JP, Katinger H. 1996. Human monoclonal antibody 2G12 defines a distinctive neutralization epitope on the gp120 glycoprotein of human immunodeficiency virus type 1. J Virol 70:1100–1108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Buchacher A, Predl R, Strutzenberger K, Steinfellner W, Trkola A, Purtscher M, Gruber G, Tauer C, Steindl F, Jungbauer A, et al. 1994. Generation of human monoclonal antibodies against HIV-1 proteins; electrofusion and Epstein-Barr virus transformation for peripheral blood lymphocyte immortalization. AIDS Res Hum Retroviruses 10:359–369. doi: 10.1089/aid.1994.10.359. [DOI] [PubMed] [Google Scholar]
- 47.Muster T, Steindl F, Purtscher M, Trkola A, Klima A, Himmler G, Ruker F, Katinger H. 1993. A conserved neutralizing epitope on gp41 of human immunodeficiency virus type 1. J Virol 67:6642–6647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Gorny MK, Conley AJ, Karwowska S, Buchbinder A, Xu JY, Emini EA, Koenig S, Zolla-Pazner S. 1992. Neutralization of diverse human immunodeficiency virus type 1 variants by an anti-V3 human monoclonal antibody. J Virol 66:7538–7542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Falkowska E, Le KM, Ramos A, Doores KJ, Lee JH, Blattner C, Ramirez A, Derking R, van Gils MJ, Liang CH, McBride R, von Bredow B, Shivatare SS, Wu CY, Chan-Hui PY, Liu Y, Feizi T, Zwick MB, Koff WC, Seaman MS, Swiderek K, Moore JP, Evans D, Paulson JC, Wong CH, Ward AB, Wilson IA, Sanders RW, Poignard P, Burton DR. 2014. Broadly neutralizing HIV antibodies define a glycan-dependent epitope on the prefusion conformation of gp41 on cleaved envelope trimers. Immunity 40:657–668. doi: 10.1016/j.immuni.2014.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Falkowska E, Ramos A, Feng Y, Zhou T, Moquin S, Walker LM, Wu X, Seaman MS, Wrin T, Kwong PD, Wyatt RT, Mascola JR, Poignard P, Burton DR. 2012. PGV04, an HIV-1 gp120 CD4 binding site antibody, is broad and potent in neutralization but does not induce conformational changes characteristic of CD4. J Virol 86:4394–4403. doi: 10.1128/JVI.06973-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Burton DR, Pyati J, Koduri R, Sharp SJ, Thornton GB, Parren PW, Sawyer LS, Hendry RM, Dunlop N, Nara PL, et al. 1994. Efficient neutralization of primary isolates of HIV-1 by a recombinant human monoclonal antibody. Science 266:1024–1027. doi: 10.1126/science.7973652. [DOI] [PubMed] [Google Scholar]
- 52.Roben P, Moore JP, Thali M, Sodroski J, Barbas CF III, Burton DR. 1994. Recognition properties of a panel of human recombinant Fab fragments to the CD4 binding site of gp120 that show differing abilities to neutralize human immunodeficiency virus type 1. J Virol 68:4821–4828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Wu X, Yang ZY, Li Y, Hogerkorp CM, Schief WR, Seaman MS, Zhou T, Schmidt SD, Wu L, Xu L, Longo NS, McKee K, O'Dell S, Louder MK, Wycuff DL, Feng Y, Nason M, Doria-Rose N, Connors M, Kwong PD, Roederer M, Wyatt RT, Nabel GJ, Mascola JR. 2010. Rational design of envelope identifies broadly neutralizing human monoclonal antibodies to HIV-1. Science 329:856–861. doi: 10.1126/science.1187659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Walker LM, Huber M, Doores KJ, Falkowska E, Pejchal R, Julien JP, Wang SK, Ramos A, Chan-Hui PY, Moyle M, Mitcham JL, Hammond PW, Olsen OA, Phung P, Fling S, Wong CH, Phogat S, Wrin T, Simek MD, Koff WC, Wilson IA, Burton DR, Poignard P. 2011. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 477:466–470. doi: 10.1038/nature10373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Walker LM, Phogat SK, Chan-Hui PY, Wagner D, Phung P, Goss JL, Wrin T, Simek MD, Fling S, Mitcham JL, Lehrman JK, Priddy FH, Olsen OA, Frey SM, Hammond PW, Kaminsky S, Zamb T, Moyle M, Koff WC, Poignard P, Burton DR. 2009. Broad and potent neutralizing antibodies from an African donor reveal a new HIV-1 vaccine target. Science 326:285–289. doi: 10.1126/science.1178746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Huang J, Kang BH, Pancera M, Lee JH, Tong T, Feng Y, Georgiev IS, Chuang GY, Druz A, Doria-Rose NA, Laub L, Sliepen K, van Gils MJ, de la Pena AT, Derking R, Klasse PJ, Migueles SA, Bailer RT, Alam M, Pugach P, Haynes BF, Wyatt RT, Sanders RW, Binley JM, Ward AB, Mascola JR, Kwong PD, Connors M. 2014. Broad and potent HIV-1 neutralization by a human antibody that binds the gp41-gp120 interface. Nature 515:138–142. doi: 10.1038/nature13601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Huang J, Ofek G, Laub L, Louder MK, Doria-Rose NA, Longo NS, Imamichi H, Bailer RT, Chakrabarti B, Sharma SK, Alam SM, Wang T, Yang Y, Zhang B, Migueles SA, Wyatt R, Haynes BF, Kwong PD, Mascola JR, Connors M. 2012. Broad and potent neutralization of HIV-1 by a gp41-specific human antibody. Nature 491:406–412. doi: 10.1038/nature11544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Pincus SH, Fang H, Wilkinson RA, Marcotte TK, Robinson JE, Olson WC. 2003. In vivo efficacy of anti-glycoprotein 41, but not anti-glycoprotein 120, immunotoxins in a mouse model of HIV infection. J Immunol 170:2236–2241. doi: 10.4049/jimmunol.170.4.2236. [DOI] [PubMed] [Google Scholar]
- 59.Moore JP, Willey RL, Lewis GK, Robinson J, Sodroski J. 1994. Immunological evidence for interactions between the first, second, and fifth conserved domains of the gp120 surface glycoprotein of human immunodeficiency virus type 1. J Virol 68:6836–6847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Scott CF Jr, Silver S, Profy AT, Putney SD, Langlois A, Weinhold K, Robinson JE. 1990. Human monoclonal antibody that recognizes the V3 region of human immunodeficiency virus gp120 and neutralizes the human T-lymphotropic virus type IIIMN strain. Proc Natl Acad Sci U S A 87:8597–8601. doi: 10.1073/pnas.87.21.8597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Thali M, Moore JP, Furman C, Charles M, Ho DD, Robinson J, Sodroski J. 1993. Characterization of conserved human immunodeficiency virus type 1 gp120 neutralization epitopes exposed upon gp120-CD4 binding. J Virol 67:3978–3988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Earl PL, Broder CC, Doms RW, Moss B. 1997. Epitope map of human immunodeficiency virus type 1 gp41 derived from 47 monoclonal antibodies produced by immunization with oligomeric envelope protein. J Virol 71:2674–2684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Cavacini L, Duval M, Song L, Sangster R, Xiang SH, Sodroski J, Posner M. 2003. Conformational changes in env oligomer induced by an antibody dependent on the V3 loop base. AIDS 17:685–689. doi: 10.1097/00002030-200303280-00006. [DOI] [PubMed] [Google Scholar]
- 64.Allaway GP, Davis-Bruno KL, Beaudry GA, Garcia EB, Wong EL, Ryder AM, Hasel KW, Gauduin MC, Koup RA, McDougal JS, et al. 1995. Expression and characterization of CD4-IgG2, a novel heterotetramer that neutralizes primary HIV type 1 isolates. AIDS Res Hum Retroviruses 11:533–539. doi: 10.1089/aid.1995.11.533. [DOI] [PubMed] [Google Scholar]
- 65.Nelson JD, Brunel FM, Jensen R, Crooks ET, Cardoso RM, Wang M, Hessell A, Wilson IA, Binley JM, Dawson PE, Burton DR, Zwick MB. 2007. An affinity-enhanced neutralizing antibody against the membrane-proximal external region of human immunodeficiency virus type 1 gp41 recognizes an epitope between those of 2F5 and 4E10. J Virol 81:4033–4043. doi: 10.1128/JVI.02588-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Feng Y, McKee K, Tran K, O'Dell S, Schmidt SD, Phogat A, Forsell MN, Karlsson Hedestam GB, Mascola JR, Wyatt RT. 2012. Biochemically defined HIV-1 envelope glycoprotein variant immunogens display differential binding and neutralizing specificities to the CD4-binding site. J Biol Chem 287:5673–5686. doi: 10.1074/jbc.M111.317776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Reed SE, Staley EM, Mayginnes JP, Pintel DJ, Tullis GE. 2006. Transfection of mammalian cells using linear polyethylenimine is a simple and effective means of producing recombinant adeno-associated virus vectors. J Virol Methods 138:85–98. doi: 10.1016/j.jviromet.2006.07.024. [DOI] [PubMed] [Google Scholar]
- 68.Brugger B, Glass B, Haberkant P, Leibrecht I, Wieland FT, Krausslich HG. 2006. The HIV lipidome: a raft with an unusual composition. Proc Natl Acad Sci U S A 103:2641–2646. doi: 10.1073/pnas.0511136103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Blattner C, Lee JH, Sliepen K, Derking R, Falkowska E, de la Pena AT, Cupo A, Julien JP, van Gils M, Lee PS, Peng W, Paulson JC, Poignard P, Burton DR, Moore JP, Sanders RW, Wilson IA, Ward AB. 2014. Structural delineation of a quaternary, cleavage-dependent epitope at the gp41-gp120 interface on intact HIV-1 Env trimers. Immunity 40:669–680. doi: 10.1016/j.immuni.2014.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Carragher B, Kisseberth N, Kriegman D, Milligan RA, Potter CS, Pulokas J, Reilein A. 2000. Leginon: an automated system for acquisition of images from vitreous ice specimens. J Struct Biol 132:33–45. doi: 10.1006/jsbi.2000.4314. [DOI] [PubMed] [Google Scholar]
- 71.Voss NR, Yoshioka CK, Radermacher M, Potter CS, Carragher B. 2009. DoG Picker and TiltPicker: software tools to facilitate particle selection in single particle electron microscopy. J Struct Biol 166:205–213. doi: 10.1016/j.jsb.2009.01.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Penczek P, Radermacher M, Frank J. 1992. Three-dimensional reconstruction of single particles embedded in ice. Ultramicroscopy 40:33–53. doi: 10.1016/0304-3991(92)90233-A. [DOI] [PubMed] [Google Scholar]
- 73.Doores KJ, Burton DR. 2010. Variable loop glycan dependency of the broad and potent HIV-1-neutralizing antibodies PG9 and PG16. J Virol 84:10510–10521. doi: 10.1128/JVI.00552-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Klein JS, Bjorkman PJ. 2010. Few and far between: how HIV may be evading antibody avidity. PLoS Pathog 6:e1000908. doi: 10.1371/journal.ppat.1000908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Bartesaghi A, Merk A, Borgnia MJ, Milne JL, Subramaniam S. 2013. Prefusion structure of trimeric HIV-1 envelope glycoprotein determined by cryo-electron microscopy. Nat Struct Mol Biol 20:1352–1357. doi: 10.1038/nsmb.2711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Tran EE, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JL, Subramaniam S. 2012. Structural mechanism of trimeric HIV-1 envelope glycoprotein activation. PLoS Pathog 8:e1002797. doi: 10.1371/journal.ppat.1002797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Ringe RP, Sanders RW, Yasmeen A, Kim HJ, Lee JH, Cupo A, Korzun J, Derking R, van Montfort T, Julien JP, Wilson IA, Klasse PJ, Ward AB, Moore JP. 2013. Cleavage strongly influences whether soluble HIV-1 envelope glycoprotein trimers adopt a native-like conformation. Proc Natl Acad Sci U S A 110:18256–18261. doi: 10.1073/pnas.1314351110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Khayat R, Lee JH, Julien JP, Cupo A, Klasse PJ, Sanders RW, Moore JP, Wilson IA, Ward AB. 2013. Structural characterization of cleaved, soluble HIV-1 envelope glycoprotein trimers. J Virol 87:9865–9872. doi: 10.1128/JVI.01222-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Sellhorn G, Kraft Z, Caldwell Z, Ellingson K, Mineart C, Seaman MS, Montefiori DC, Lagerquist E, Stamatatos L. 2012. Engineering, expression, purification, and characterization of stable clade A/B recombinant soluble heterotrimeric gp140 proteins. J Virol 86:128–142. doi: 10.1128/JVI.06363-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Grundner C, Mirzabekov T, Sodroski J, Wyatt R. 2002. Solid-phase proteoliposomes containing human immunodeficiency virus envelope glycoproteins. J Virol 76:3511–3521. doi: 10.1128/JVI.76.7.3511-3521.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Bachmann MF, Jennings GT. 2010. Vaccine delivery: a matter of size, geometry, kinetics and molecular patterns. Nat Rev Immunol 10:787–796. doi: 10.1038/nri2868. [DOI] [PubMed] [Google Scholar]
- 82.Chakrabarti BK, Feng Y, Sharma SK, McKee K, Karlsson Hedestam GB, Labranche CC, Montefiori DC, Mascola JR, Wyatt RT. 2013. Robust neutralizing antibodies elicited by HIV-1 JRFL envelope glycoprotein trimers in nonhuman primates. J Virol 87:13239–13251. doi: 10.1128/JVI.01247-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Narayan KM, Agrawal N, Du SX, Muranaka JE, Bauer K, Leaman DP, Phung P, Limoli K, Chen H, Boenig RI, Wrin T, Zwick MB, Whalen RG. 2013. Prime-boost immunization of rabbits with HIV-1 gp120 elicits potent neutralization activity against a primary viral isolate. PLoS One 8:e52732. doi: 10.1371/journal.pone.0052732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Zhou T, Zhu J, Yang Y, Gorman J, Ofek G, Srivatsan S, Druz A, Lees CR, Lu G, Soto C, Stuckey J, Burton DR, Koff WC, Connors M, Kwong PD. 2014. Transplanting supersites of HIV-1 vulnerability. PLoS One 9:e99881. doi: 10.1371/journal.pone.0099881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Rusert P, Krarup A, Magnus C, Brandenberg OF, Weber J, Ehlert AK, Regoes RR, Gunthard HF, Trkola A. 2011. Interaction of the gp120 V1V2 loop with a neighboring gp120 unit shields the HIV envelope trimer against cross-neutralizing antibodies. J Exp Med 208:1419–1433. doi: 10.1084/jem.20110196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Liu L, Cimbro R, Lusso P, Berger EA. 2011. Intraprotomer masking of third variable loop (V3) epitopes by the first and second variable loops (V1V2) within the native HIV-1 envelope glycoprotein trimer. Proc Natl Acad Sci U S A 108:20148–20153. doi: 10.1073/pnas.1104840108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Phogat S, Wyatt R. 2007. Rational modifications of HIV-1 envelope glycoproteins for immunogen design. Curr Pharm Des 13:213–227. doi: 10.2174/138161207779313632. [DOI] [PubMed] [Google Scholar]
- 88.Wei X, Decker JM, Wang S, Hui H, Kappes JC, Wu X, Salazar-Gonzalez JF, Salazar MG, Kilby JM, Saag MS, Komarova NL, Nowak MA, Hahn BH, Kwong PD, Shaw GM. 2003. Antibody neutralization and escape by HIV-1. Nature 422:307–312. doi: 10.1038/nature01470. [DOI] [PubMed] [Google Scholar]
- 89.Tomaras GD, Yates NL, Liu P, Qin L, Fouda GG, Chavez LL, Decamp AC, Parks RJ, Ashley VC, Lucas JT, Cohen M, Eron J, Hicks CB, Liao HX, Self SG, Landucci G, Forthal DN, Weinhold KJ, Keele BF, Hahn BH, Greenberg ML, Morris L, Karim SS, Blattner WA, Montefiori DC, Shaw GM, Perelson AS, Haynes BF. 2008. Initial B-cell responses to transmitted human immunodeficiency virus type 1: virion-binding immunoglobulin M (IgM) and IgG antibodies followed by plasma anti-gp41 antibodies with ineffective control of initial viremia. J Virol 82:12449–12463. doi: 10.1128/JVI.01708-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.McMichael AJ, Borrow P, Tomaras GD, Goonetilleke N, Haynes BF. 2010. The immune response during acute HIV-1 infection: clues for vaccine development. Nat Rev Immunol 10:11–23. doi: 10.1038/nri2674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Victora GD, Schwickert TA, Fooksman DR, Kamphorst AO, Meyer-Hermann M, Dustin ML, Nussenzweig MC. 2010. Germinal center dynamics revealed by multiphoton microscopy with a photoactivatable fluorescent reporter. Cell 143:592–605. doi: 10.1016/j.cell.2010.10.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.del Guercio MF, Alexander J, Kubo RT, Arrhenius T, Maewal A, Appella E, Hoffman SL, Jones T, Valmori D, Sakaguchi K, Grey HM, Sette A. 1997. Potent immunogenic short linear peptide constructs composed of B cell epitopes and Pan DR T helper epitopes (PADRE) for antibody responses in vivo. Vaccine 15:441–448. doi: 10.1016/S0264-410X(97)00186-2. [DOI] [PubMed] [Google Scholar]
- 93.Forsell MN, Soldemo M, Dosenovic P, Wyatt RT, Karlsson MC, Karlsson Hedestam GB. 2013. Independent expansion of epitope-specific plasma cell responses upon HIV-1 envelope glycoprotein immunization. J Immunol 191:44–51. doi: 10.4049/jimmunol.1203087. [DOI] [PubMed] [Google Scholar]
- 94.McGuire AT, Hoot S, Dreyer AM, Lippy A, Stuart A, Cohen KW, Jardine J, Menis S, Scheid JF, West AP, Schief WR, Stamatatos L. 2013. Engineering HIV envelope protein to activate germline B cell receptors of broadly neutralizing anti-CD4 binding site antibodies. J Exp Med 210:655–663. doi: 10.1084/jem.20122824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Kushnir N, Streatfield SJ, Yusibov V. 2012. Virus-like particles as a highly efficient vaccine platform: diversity of targets and production systems and advances in clinical development. Vaccine 31:58–83. doi: 10.1016/j.vaccine.2012.10.083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Francis T., Jr 1960. On the doctrine of original antigenic sin. Proc Am Philos Soc 104:572–578. [Google Scholar]
- 97.Sheets RL, Stein J, Manetz TS, Duffy C, Nason M, Andrews C, Kong WP, Nabel GJ, Gomez PL. 2006. Biodistribution of DNA plasmid vaccines against HIV-1, Ebola, severe acute respiratory syndrome, or West Nile virus is similar, without integration, despite differing plasmid backbones or gene inserts. Toxicol Sci 91:610–619. doi: 10.1093/toxsci/kfj169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Ngo TT, Yam CF, Lenhoff HM, Ivy J. 1981. p-Azidophenylglyoxal: a heterobifunctional photoactivable cross-linking reagent selective for arginyl residues. J Biol Chem 256:11313–11318. [PubMed] [Google Scholar]
- 99.Ban H, Gavrilyuk J, Barbas CF III. 2010. Tyrosine bioconjugation through aqueous ene-type reactions: a click-like reaction for tyrosine. J Am Chem Soc 132:1523–1525. doi: 10.1021/ja909062q. [DOI] [PubMed] [Google Scholar]
- 100.Zeng Y, Ramya TN, Dirksen A, Dawson PE, Paulson JC. 2009. High-efficiency labeling of sialylated glycoproteins on living cells. Nat Methods 6:207–209. doi: 10.1038/nmeth.1305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Chin JW, Martin AB, King DS, Wang L, Schultz PG. 2002. Addition of a photo-cross-linking amino acid to the genetic code of Escherichia coli. Proc Natl Acad Sci U S A 99:11020–11024. doi: 10.1073/pnas.172226299. [DOI] [PMC free article] [PubMed] [Google Scholar]