Significance
This research documents the spread of an opportunistic coral endosymbiont, Symbiodinium trenchii, from the Indo-Pacific into the Greater Caribbean, a region afflicted by human-related impacts including climate warming and environmental degradation. As a symbiont, it increases the resilience of photosynthetic corals to environmental perturbation but may diminish the animal’s capacity to calcify and build reefs. This work exposes a critical need to better understand the consequences of microbial introductions (even mutualistic species) on ecosystem stability and function and raises questions about the long-term impact of new, but maladapted, symbioses on the productivity of reef coral communities in the Atlantic Ocean.
Keywords: invasive species, climate change, reef corals, calcification, symbiosis
Abstract
Human-induced environmental changes have ushered in the rapid decline of coral reef ecosystems, particularly by disrupting the symbioses between reef-building corals and their photosymbionts. However, escalating stressful conditions enable some symbionts to thrive as opportunists. We present evidence that a stress-tolerant “zooxanthella” from the Indo-Pacific Ocean, Symbiodinium trenchii, has rapidly spread to coral communities across the Greater Caribbean. In marked contrast to populations from the Indo-Pacific, Atlantic populations of S. trenchii contained exceptionally low genetic diversity, including several widespread and genetically similar clones. Colonies with this symbiont tolerate temperatures 1–2 °C higher than other host–symbiont combinations; however, calcification by hosts harboring S. trenchii is reduced by nearly half, compared with those harboring natives, and suggests that these new symbioses are maladapted. Unforeseen opportunism and geographical expansion by invasive mutualistic microbes could profoundly influence the response of reef coral symbioses to major environmental perturbations but may ultimately compromise ecosystem stability and function.
Biological invasions and global climate change are drastically altering the diversity, distribution, and ecology of all biota on the planet (1–4), which negatively affects human well-being (5). Although the body of literature on invasive plants and animals is voluminous, invasions by nonpathogenic microbes and their impacts on ecosystem functions are poorly documented (6). Microbial invasions are difficult to detect, especially among free-living or mutualistic species, and their introduction and proliferation into new regions proceed without recognition. These invasions may alter the composition of a host community and affect the functional capacity of ecosystems (6, 7). Additional climate warming and ecosystem degradation will likely facilitate increasing numbers of successful invasions, which may further exacerbate negative ecological impacts (8).
Global change, especially ocean warming, continues to diminish the functional importance of reef corals by reducing their productivity, biodiversity, and capacity to precipitate calcium carbonate (9). The symbiotic dinoflagellates found in reef-building corals (Symbiodinium) are crucial to the maintenance and photosynthetic productivity of coral reef ecosystems, thus underscoring the importance of microbes and symbioses to ecosystem viability. Anomalous episodes of warm ocean temperatures destabilize these mutualistic partnerships, resulting in mass coral “bleaching” (i.e., symbiont expulsion) and mortality over broad geographic expanses (10). Predicting the response of present day host–symbiont combinations to increasing anthropogenic warming and environmental degradation is a matter of considerable debate and involves broad speculation on whether stress-tolerant symbionts might emerge to facilitate a rapid eco-physiological response to changing conditions (11–14). However, it is unclear how symbiont replacement, or displacement (referred to as “switching” or “shuffling”), will progress in diverse coral communities distributed across broad geographic areas over the decadal ecological timescales that are necessary to keep pace with the current rate of warming.
Research on the diversity and ecology of coral symbionts suggests that episodes of stressful warming may facilitate the spread of ecologically rare or opportunistic species (13). The severe mass bleaching and mortality of eastern Caribbean corals in 2005 corresponded with an increased prevalence and abundance of Symbiodinium trenchii (formerly D1a, or D1-4) (15), a species in Clade D. This group is distantly related to other lineages of Symbiodinium and is primarily known for occurring in animals at the margins of their environmental tolerance (15). Before, during, and after this bleaching event, S. trenchii spread to many coral species, especially among those ultimately showing the greatest signs of physiological trauma (i.e., bleaching). Colonies with high densities of this symbiont seemed to tolerate significant thermal stress. Indeed, photosynthetic rates of S. trenchii (in hospite) remain unaffected at temperatures that are typically stressful to other Symbiodinium spp. (16–18). When environmental conditions stabilized, however, S. trenchii was inexorably displaced (requiring months or years) by host-typical symbionts (13, 19). Therefore, the persistence of S. trenchii within a coral colony may depend on chronic, or acute, stressors.
We suspected, for several reasons, that S. trenchii populations in the northwestern tropical Atlantic (i.e., Greater Caribbean) were not native, motivating our population genetic investigation. Indeed, detailed phylogenetic analyses are unable to distinguish S. trenchii in the Greater Caribbean from those in the Indo-Pacific (15). Moreover, several other host-specialized Clade D Symbiodinium spp. often occur with S. trenchii in equatorial coral communities of the Indo-Pacific (20), but S. trenchii is the only species of Clade D found in the Atlantic. Finally, as mentioned above, S. trenchii behaves opportunistically and can successfully invade the colonies of many coral species experiencing physiological stress (analogous in some ways to “disturbed habitats”) (16, 21). Therefore, we tested the hypothesis that populations of Atlantic S. trenchii represent a recent invasion by examining interindividual genotypic diversity and divergence across the Indo-Pacific and Greater Caribbean using high-resolution microsatellite loci (22, 23).
Results
Population Genetic Diversity of S. trenchii in the Indo-West Pacific.
Most samples of reef corals with S. trenchii from the eastern Indian and western Pacific Oceans contained a single multilocus genotype (MLG), or strain, and different samples usually contained a distinct MLG (Fig. 1 A–C and Dataset S1). About 10–20% of samples contained mixtures of genotypes and, therefore, were not included in subsequent genetic analyses. Interindividual variation among genotypes was high, and allele sizes often differed at multiple (>4–6) loci (Fig. 2 and Tables S1 and S2). Although most MLGs were discovered only once (Figs. 1 B and C), those that were found on several occasions were obtained from host animals living in the same vicinity (1–50 m apart on the same reef). Host communities on reef systems separated by as little as 10 km contained distinct MLG assemblages of S. trenchii (Table 1 and Fig. 1 B and C). Moreover, populations from Indo-Pacific locations contained a high diversity of alleles, including many that were unique either to the eastern Indian or western Pacific Oceans (Table 1).
Fig. 1.
The diversity and prevalence of individual multilocus genotypes from populations of the endosymbiotic dinoflagellate, S. trenchii, obtained from reef corals in the Indian, Pacific, and Atlantic Oceans. (A) Available biogeographic data indicate that the “species” diversity among stress-tolerant Symbiodinium clade D is centered around Southeast Asia and probably Indonesia. (B) Most multilocus genotypes (MLGs = strains or clones) characterized from coral colonies in the Andaman Sea were unique (black shading) and seemed restricted in their distribution to a single reef (MLGs that were found more than once are designated by a color and Greek letter). (C) Similarly, populations characterized from coral communities dwelling in the rock island habits of Palau were genotypically diverse with no overlap in the distribution of MLGs between sites. (D) Samples of S. trenchii were collected from locations across the Greater Caribbean, where (E) genotypic diversity was low and populations were composed of several geographically widespread and genetically similar clones. The most prevalent of these genotypes, “α” (red), was found at seven of eight locations separated in some cases by thousands of kilometers.
Fig. 2.
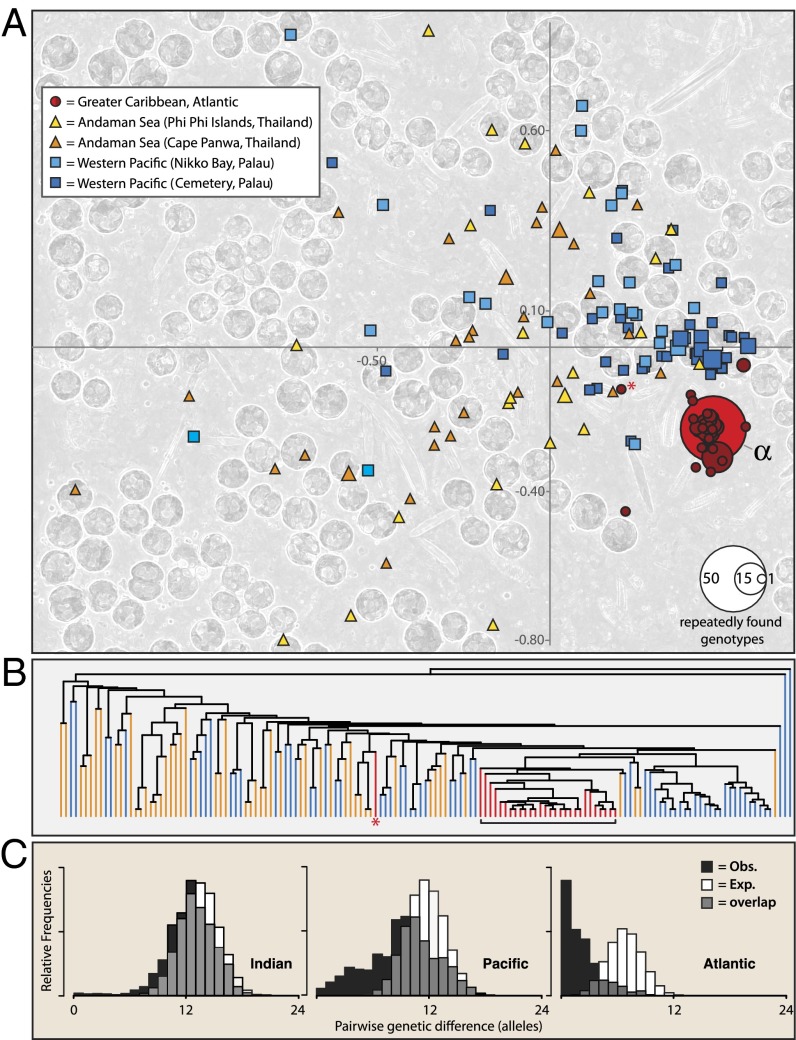
Low genetic diversity and high clonality of S. trenchii in the Greater Caribbean relative to the Indo-Pacific. Although many genotypes occurred, analyses of allelic differences across 12 loci show that most strains differed at only one or two loci in collections from the Atlantic. (A) Principal coordinate analysis of MLGs shows the extent of genetic differentiation found among genotypes (the two axes describe 34% and 17% of the total variation). Symbol shape corresponds to ocean basin whereas symbol size relates to the number of times that a particular MLG was recovered. The most represented and widely distributed clone was genotype α from the Greater Caribbean. Cells of S. trenchii found in each host colony often comprised a distinct clonal population, or MLG (background image). (B) UPGMA tree showing the distance-based relationship between unique MLGs of S. trenchii obtained from the Indian (orange), Pacific (blue), and Atlantic (red) Oceans. The red asterisk in A and B signifies an unusual Atlantic genotype found in the Gulf of Mexico (FGB). (C) Pairwise comparisons of genotypes found in each of three oceans further indicate that the frequency of related individuals in the Greater Caribbean (Atlantic) is significantly higher than expected in a panmictic population.
Table S1.
Description and summary data on microsatellite loci
| Locus | Primer sequence (5′–3′) | Repeat motif | Size range, bp | No. of alleles |
| D1Sym9 | F-CAGAAGCCCAATTATATGCGGCA (FAM) | (GTT)6 | 106–112 | 3 |
| R-AGGATGATGAGCATGCCGACG | ||||
| D1Sym11 | F-TGAAATCTCACTCAGAGTCGGAC (FAM) | (AC)13 | 149–161 | 7 |
| R-GCAGACAGTGATTTCAGTTCCGA | ||||
| D1Sym14 | F-TCTCAGTGGAAAGCATTGTGG (FAM) | (CT)11 AT (CT)4 | 167–191 | 7 |
| R-TCGTCTGAATCAGGATCTGACG | ||||
| D1Sym17 | F-TGTGAATGCTTCTTGGGGTG (HEX) | (CA)8 | 137–149 | 7 |
| R-TCATGCTTGTCCGTGAGCAG | ||||
| D1Sym34 | F-ACCTGAGACCTGAGTGTTGC (FAM) | (CAAA)9 CACA (CAAA)4 (GAAACAAA)2 (CAAA)13 | 350–433 | 29 |
| R-ATCATGGGCAGAGCTCCTGG | ||||
| D1Sym66 | F-CTCTGGGATGGACCCCGAA (PET) | (TCT)8 | 288–303 | 6 |
| R-GACAGTGTTTCACTGGTCGCA | ||||
| D1Sym67 | F-GAATCCAGATGGTGCCTGC (VIC) | (ATC)8 | 125–140 | 6 |
| R-CAAAGGTAGCCGATTGTCTC | ||||
| D1Sym82 | F-AAGACTTCCGCAAGCACAC (HEX) | (ACT)12 | 219–285 | 22 |
| R-CGCATCTACTGGTGGTCAGAC | ||||
| D1Sym87 | F-CCTATGACTCCAAGGGTGACG (FAM) | (GAAG)7 | 232–256 | 7 |
| R-AGACATACCTCGGTCTTGTC | ||||
| D1Sym88 | F-TTGTCAGACTGAATGCTCCA (NED) | (CTTT)3 G (TTTC)7 (TCTCTTTC)2 TCTTTTT (CT)3 (TCTT)2 (TTTC)2 T (CTT)4 | 232–250 | 6 |
| R-GTGTTCAAGCGACATCCCA | ||||
| D1Sym92 | F-GCGTTTGACACAAGGATCCCT (FAM) | (CCTA)6 (CCTG)3 | 124–136 | 4 |
| R-TTGGGATGCTCTTGGCGAC | ||||
| D1Sym93 | F-GCTCAAAGGAGCTCTAGGGGT (NED) | (AATC)6 | 141–173 | 8 |
| R-TGTCAAGGTAGAGAGCCTGGT |
Subscript numbers after the repeat motif indicate the number of repeats in the initial cloned sequence that was used to develop locus primers.
Table S2.
Allele frequencies by locus found in populations of S. trenchii from the Greater Caribbean (GC), Indian Ocean (IND), and Pacific Ocean (PAC)
| Locus | Allele size & sample no. (N) | GC | IND | PAC |
| Sym9 | N | 29 | 52 | 67 |
| 100 | 0.000 | 0.000 | 0.015 | |
| 106 | 0.517 | 0.577 | 0.463 | |
| 109 | 0.483 | 0.337 | 0.507 | |
| 112 | 0.000 | 0.087 | 0.015 | |
| Sym11 | N | 29 | 50 | 65 |
| 149 | 0.000 | 0.050 | 0.031 | |
| 151 | 0.500 | 0.340 | 0.408 | |
| 153 | 0.448 | 0.210 | 0.308 | |
| 155 | 0.052 | 0.130 | 0.138 | |
| 157 | 0.000 | 0.120 | 0.100 | |
| 159 | 0.000 | 0.120 | 0.008 | |
| 161 | 0.000 | 0.030 | 0.008 | |
| Sym14 | N | 28 | 51 | 53 |
| 173 | 0.000 | 0.020 | 0.000 | |
| 175 | 0.000 | 0.039 | 0.000 | |
| 177 | 0.518 | 0.451 | 0.491 | |
| 179 | 0.482 | 0.471 | 0.425 | |
| 181 | 0.000 | 0.010 | 0.085 | |
| 191 | 0.000 | 0.010 | 0.000 | |
| Sym17 | N | 28 | 48 | 66 |
| 137 | 0.000 | 0.021 | 0.000 | |
| 139 | 0.000 | 0.010 | 0.000 | |
| 141 | 0.482 | 0.458 | 0.500 | |
| 143 | 0.482 | 0.198 | 0.356 | |
| 145 | 0.018 | 0.271 | 0.114 | |
| 147 | 0.018 | 0.042 | 0.023 | |
| 149 | 0.000 | 0.000 | 0.008 | |
| Sym34 | N | 29 | 51 | 65 |
| 350 | 0.000 | 0.010 | 0.000 | |
| 353 | 0.000 | 0.010 | 0.008 | |
| 361 | 0.000 | 0.010 | 0.008 | |
| 363 | 0.000 | 0.010 | 0.000 | |
| 365 | 0.000 | 0.020 | 0.023 | |
| 367 | 0.000 | 0.059 | 0.008 | |
| 369 | 0.000 | 0.039 | 0.008 | |
| 371 | 0.000 | 0.078 | 0.000 | |
| 373 | 0.000 | 0.118 | 0.092 | |
| 375 | 0.000 | 0.049 | 0.008 | |
| 377 | 0.052 | 0.078 | 0.238 | |
| 379 | 0.034 | 0.020 | 0.038 | |
| 381 | 0.034 | 0.088 | 0.054 | |
| 383 | 0.121 | 0.049 | 0.038 | |
| 385 | 0.362 | 0.049 | 0.208 | |
| 387 | 0.362 | 0.039 | 0.038 | |
| 389 | 0.034 | 0.059 | 0.038 | |
| 391 | 0.000 | 0.020 | 0.031 | |
| 393 | 0.000 | 0.059 | 0.023 | |
| 395 | 0.000 | 0.029 | 0.008 | |
| 397 | 0.000 | 0.059 | 0.046 | |
| 399 | 0.000 | 0.010 | 0.008 | |
| 401 | 0.000 | 0.020 | 0.000 | |
| 403 | 0.000 | 0.010 | 0.023 | |
| 405 | 0.000 | 0.010 | 0.008 | |
| 407 | 0.000 | 0.000 | 0.015 | |
| 409 | 0.000 | 0.000 | 0.015 | |
| 429 | 0.000 | 0.000 | 0.008 | |
| 433 | 0.000 | 0.000 | 0.008 | |
| Sym92 | N | 29 | 52 | 66 |
| 124 | 0.017 | 0.260 | 0.129 | |
| 128 | 0.983 | 0.712 | 0.848 | |
| 132 | 0.000 | 0.019 | 0.015 | |
| 136 | 0.000 | 0.010 | 0.008 | |
| Sym66 | N | 29 | 49 | 65 |
| 288 | 0.000 | 0.031 | 0.015 | |
| 291 | 0.000 | 0.224 | 0.115 | |
| 294 | 1.000 | 0.633 | 0.592 | |
| 297 | 0.000 | 0.112 | 0.115 | |
| 300 | 0.000 | 0.000 | 0.146 | |
| 303 | 0.000 | 0.000 | 0.015 | |
| Sym93 | N | 29 | 50 | 64 |
| 141 | 0.000 | 0.000 | 0.008 | |
| 145 | 0.000 | 0.100 | 0.023 | |
| 149 | 0.362 | 0.400 | 0.508 | |
| 151 | 0.241 | 0.010 | 0.000 | |
| 153 | 0.000 | 0.120 | 0.070 | |
| 157 | 0.379 | 0.350 | 0.391 | |
| 161 | 0.017 | 0.010 | 0.000 | |
| 173 | 0.000 | 0.010 | 0.000 | |
| Sym67 | N | 29 | 51 | 67 |
| 125 | 0.000 | 0.000 | 0.015 | |
| 128 | 0.000 | 0.108 | 0.000 | |
| 131 | 0.983 | 0.412 | 0.485 | |
| 134 | 0.000 | 0.225 | 0.425 | |
| 137 | 0.017 | 0.176 | 0.045 | |
| 140 | 0.000 | 0.078 | 0.030 | |
| Sym82 | N | 29 | 47 | 62 |
| 219 | 0.000 | 0.000 | 0.008 | |
| 225 | 0.000 | 0.011 | 0.000 | |
| 228 | 0.000 | 0.011 | 0.000 | |
| 231 | 0.000 | 0.043 | 0.032 | |
| 234 | 0.431 | 0.074 | 0.177 | |
| 237 | 0.103 | 0.085 | 0.065 | |
| 240 | 0.052 | 0.170 | 0.097 | |
| 243 | 0.293 | 0.043 | 0.056 | |
| 246 | 0.069 | 0.032 | 0.081 | |
| 249 | 0.017 | 0.064 | 0.081 | |
| 252 | 0.000 | 0.085 | 0.113 | |
| 255 | 0.000 | 0.106 | 0.081 | |
| 258 | 0.000 | 0.074 | 0.024 | |
| 261 | 0.017 | 0.032 | 0.024 | |
| 264 | 0.000 | 0.043 | 0.056 | |
| 267 | 0.000 | 0.021 | 0.040 | |
| 270 | 0.017 | 0.032 | 0.032 | |
| 273 | 0.000 | 0.000 | 0.008 | |
| 276 | 0.000 | 0.000 | 0.008 | |
| 279 | 0.000 | 0.011 | 0.008 | |
| 282 | 0.000 | 0.032 | 0.000 | |
| 285 | 0.000 | 0.032 | 0.008 | |
| Sym87 | N | 29 | 51 | 64 |
| 232 | 0.000 | 0.000 | 0.016 | |
| 236 | 0.000 | 0.029 | 0.016 | |
| 240 | 0.500 | 0.471 | 0.438 | |
| 244 | 0.483 | 0.422 | 0.336 | |
| 248 | 0.017 | 0.059 | 0.195 | |
| 252 | 0.000 | 0.010 | 0.000 | |
| 256 | 0.000 | 0.010 | 0.000 | |
| Sym88 | N | 29 | 51 | 65 |
| 232 | 0.017 | 0.049 | 0.000 | |
| 234 | 0.000 | 0.343 | 0.054 | |
| 236 | 0.000 | 0.000 | 0.031 | |
| 240 | 0.966 | 0.588 | 0.877 | |
| 244 | 0.017 | 0.020 | 0.023 | |
| 250 | 0.000 | 0.000 | 0.015 |
Table 1.
Summary statistics by location, including number of colonies sampled, number of host taxa (genera) sampled, number of unique MLGs of S. trenchii, clonal richness (R), number of private alleles, information index (I), and observed (Ho) and expected (He) heterozygosity
| Location | No. of samples | No. of host taxa | Unique MLGs | Clonal richness (R) | Private alleles | Information index (I) | Observed (Ho) | Expected (He) |
| Greater Caribbean | 112 | 8 | 29 | 0.25 | 0 | 0.70 (0.15) | 0.65 (0.13) | 0.40 (0.08) |
| Indian Ocean (Thailand) | 57 | 22 | 52 | 0.91 | 16 | 1.42 (0.21) | 0.76 (0.06) | 0.66 (0.05) |
| Pacific (Palau) | 76 | 29 | 66 | 0.87 | 16 | 1.24 (0.20) | 0.78 (0.08) | 0.59 (0.06) |
Numbers in parenthesis represent the SE for a particular statistic.
Population Genetic Diversity of S. trenchii in the Greater Caribbean.
Atlantic populations of S. trenchii were highly clonal and consisted of only a few MLGs (= clonal lineages) relative to the number of samples analyzed (Table 1). Many clones were widely distributed throughout the Greater Caribbean basin, found in multiple locations separated by hundreds of kilometers (Figs. 1 D and E). The most notable of these common clones was the one genotype comprising 42% of all samples (Figs. 1E and 2A). The distribution of this single clone (genotype α) (Fig. 1E) spanned thousands of kilometers and was found in every Greater Caribbean location, except the Gulf of Mexico (Fig. 1 D and E). There were several other common and widely distributed clones (Fig. 1E), but all were genetically similar to genotype α (Fig. 2A) and differed from each other by just 1 or 2 loci, out of 24 (Fig. 2 A and B). The principal coordinate analysis plot (with axis 1 and axis 2 explaining 34% and 17% of variation, respectively) relates the low diversity in the Greater Caribbean, where all but a few individuals clustered tightly together, to the more genetically divergent genotypes from the Indo-West Pacific scattered across the plot (Fig. 2A). The low diversity in the Greater Caribbean, accompanied by noticeably fewer rare alleles (Table 1, Supporting Information, and Fig. S1), indicates a reduced effective population size either from a recent population bottleneck or from a founding event (24, 25). Finally, of additional significance, no alleles were restricted to the Greater Caribbean, meaning that all alleles found in this region were also found in populations from the Indo-Pacific (Table 1).
Fig. S1.
A comparison of allele frequencies for S. trenchii populations from the Indian, Pacific, and Atlantic Oceans. Unlike populations from the Indo-Pacific, there were many alleles that occurred at high frequency [i.e., the result of a decrease in rare alleles or “phase shift” (25)] in the Greater Caribbean, indicative of a recent population bottleneck or from a founding event.
The unweighted pair group method with arithmetic mean (UPGMA) tree clustered all MLGs from the Greater Caribbean using short branch lengths (with the exception of one MLG from the Flower Garden Banks in the Gulf of Mexico) (Fig. 2B). The Atlantic cluster was embedded within the range of genetic differentiation found in the Indo-Pacific MLGs and not a distinct lineage outside of the Indo-Pacific grouping (Fig. 2B). The occurrence of a divergent genotype in the Flower Garden Banks southwest of the entrance to the Mississippi River suggests that independent introductions of S. trenchii might have occurred (signified by asterisks in Fig. 2 A and B). In contrast, most of the individuals from the Indo-Pacific fall at the end of long branches, reflecting large differences in allelic composition among MLGs from this region.
Disparity in Clonal Structure Among Populations of S. trenchii.
For each population, we generated an expected distribution of pairwise differences under an assumed model of sexual recombination based on the number and frequency of observed alleles found (code available in Supporting Information). Although clonality was evident in each, populations in the Andaman Sea, Indian Ocean exhibited the broadest diversity of genotypes and minimal clonality, which was consistent with our expected distribution (Fig. 2C). Such patterns of genotypic diversity and disparity (number of pairwise differences between MLGs) are typical for populations of other Symbiodinium species (Fig. S2). The bimodal distribution in pairwise differences found for populations of S. trenchii from the Pacific Ocean (Palau) shows that some proportion of the population comprises similar genotypes and repeated clones (Fig. 2C). This distribution pattern was influenced most by samples from the southern location in Palau (Cemetery), which contained a higher proportion of clones that were found more than once (Figs. 1C and 2A). The population examined from the Greater Caribbean exhibited the highest degree of clonality and the lowest mean in pairwise differences. The distribution of pairwise differences for this population had the greatest departure from the expected (Fig. 2C).
Fig. S2.
The distribution of pairwise genetic difference (PWD) among individual multilocus genotypes (MLGs) of typical Symbiodinium populations. Frequency of pairwise genetic difference typical of MLGs found for native Symbiodinium spp. [types A3 (A), B7 (B), and C7 (C)] from the Greater Caribbean (GC), and S. “glynni,“ a member of clade D and sibling species to S. trenchii, from the Eastern Pacific (D). The frequency distributions depict pairwise genetic differences between MLGs among individuals for observed data (black bars) and those expected for a panmictic (randomly mating) population (white bars). Gray bars indicate the overlap between expected and observed. All observed pairwise genetic difference were significantly different (α < 0.05) than the expected pairwise genetic difference, determined by comparing the variance in pairwise genetic difference of observed and expected values in a permutation test, indicating that all lineages do undergo significant asexual reproduction. However, the low allelic diversity and significantly lower average pairwise genetic difference than expected average pairwise genetic difference (from our permutation tests) exhibited by S. trenchii in the Atlantic Ocean seems unique in this basin to this lineage, and not simply a characteristic of Symbiodinium populations from the Caribbean in general (Fig. 2C).
Photosynthesis and Instantaneous Calcification Rates in Corals with Different Symbiodinium spp.
The identity of the resident Symbiodinium spp. in colonies of the dominant reef builder, Orbicella faveolata, did not seem to significantly affect rates of photosynthesis over a broad range of irradiances (Fig. 3 A and B, Fig. S3, and Table S3). Maximum photosynthesis (Pmax) values, inferred from rates of oxygen evolution, for S. trenchii (2.8 ± 0.29 SE μmol O2 cm−2⋅h−1) fall well within the range and is not significantly different in a pairwise t test from values measured for the three other Symbiodinium spp. associated with O. faveolata (Fig. 3B; P values listed on Table S3). This similarity suggests that photosynthetic rates for S. trenchii in hospite are similar to the Symbiodinium spp. normally found with O. faveolata and that, therefore, it can generate similar amounts of fixed carbon.
Fig. 3.

Comparison of photosynthetic rates and instantaneous calcification among different coral-Symbiodinium combinations. (A) In the Greater Caribbean, S. trenchii associates with reef building colonies of Orbicella spp. living in warm and/or turbid environments, or in colonies recovered from episodes of severe bleaching (photo by Dustin Kemp). (Inset) Magnified Symbiodinium that measure about 8–12 µm in cell diameter. (B) In hospite S. trenchii photosynthesizes (Pmax) at rates statistically indistinguishable from host–typical symbionts (undescribed species A3, B17, and C7) (Table S3) but (C) can significantly reduce rates of coral calcification measured over a broad range of temperatures. Calcification rates (under an irradiance of ∼400 μmol quanta m-2⋅s−1) in colonies with S. trenchii were statistically different (up to 50% less) than colonies with A3 and B17 at all temperatures (Table S4 and Fig. S4). All error bars were drawn as ± SE.
Fig. S3.
Example of ITS2-DGGE fingerprints diagnostic of Symbiodinium spp. living in small colonies of O. faveolata. Colonies designated by an asterisk were used for photosynthesis and calcification measurements.
Table S3.
P values for assessment of significance in all pairwise comparisons in mean Pmax involving different host–symbiont combinations
| A3 | B17 | C7 | |
| A3 | — | — | — |
| B17 | 0.61 | — | — |
| C7 | 0.17 | 0.82 | — |
| S. trenchii | 0.61 | 0.91 | 0.82 |
Fig. S4.
The statistical assessment of pairwise differences in instantaneous calcification rates based on symbiont identity. Colonies with S. trenchii calcified significantly slower than colonies with either Symbiodinium A3, or B17, when integrated in the linear mixed model over the five temperature treatments (24, 26, 28, 30, and 32 °C). Boxes depict the estimated differences between symbiont types (based on differences in the coefficient of the linear model as assessed by least square means), with error bars depicting 95% confidence intervals around this estimate. Boxes were colored by significance (P values) for each pairwise comparison.
However, significant differences in instantaneous calcification rates (up to 50%), measured at temperatures ranging between 24 °C and 32 °C, were observed in colonies dominated by S. trenchii (Fig. 3C and Table S4). The dichotomy between photosynthesis and calcification by S. trenchii symbioses suggests that, whereas S. trenchii may assimilate carbon at rates similar to other symbionts, it probably does not contribute the same benefits (e.g., translocated organic carbon) to its host. A wide variance was observed in calcification rates measured from colony to colony, which may be attributed to genotypic differences between host individuals and symbiont clones. In pairwise comparisons, instantaneous calcification rates in hosts with Symbiodinium A3 and B17 were significantly higher than colonies with S. trenchii (Supporting Information and Table S4). However, no significance difference was observed between colonies with C7 and S. trenchii across the range of temperatures investigated (Fig. 3C and Table S4). Calcification rates in corals with C7 were also not significantly different from those with A3 and B17. Symbiodinium C7 is a low-light adapted species, and colonies with this symbiont may perform better under lower irradiances relative to these other partner combinations. Pooled comparisons, whereby the rates of the native symbioses (A3, B17, and C7) were compared with S. trenchii alone, show that colonies with S. trenchii calcify at a significantly lower rate than do colonies with native symbiont species (P = 0.0015).
Table S4.
P values for assessment of significance in all pairwise comparisons in mean instantaneous calcification rates
| A3 | B17 | C7 | |
| A3 | — | — | — |
| B17 | 0.282 | — | — |
| C7 | 0.234 | 0.039 | — |
| S. trenchii | 0.006 | 0.001 | 0.068 |
Values in bold font indicate rates that were significantly different.
Discussion
Invasion of an Alien Endosymbiont.
The concordance of these population genetic data with phylogenetic, biogeographic, and ecological evidence compels us to conclude that S. trenchii was introduced into the Greater Caribbean by human-related activities (possibly multiple times and as recently as several decades ago). S. trenchii is part of an adaptive radiation of clade D centered in the Indo-West Pacific proposed to have occurred during the Pleistocene, which produced many genetically and ecologically distinct (i.e., host-specialized) species (20). Consistent with this epoch timeline, the high frequency of rare alleles and high genotypic diversity in populations of S. trenchii from the Indo-Pacific indicate a temporal existence lasting hundreds of thousands to millions of years (Table 1 and Fig. 2 A and B) (24, 25). Other members of this group, including Symbiodinium “glynni” (nomen nudum = type D1), also exhibit high allelic and genotypic diversity in the Indo-Pacific (20, 22) (Fig. S2D).
The widespread clonality exhibited by S. trenchii across the entire Western Atlantic is highly unusual in light of population genetic evidence from other Symbiodinium spp. (Fig. S2), and not merely a characteristic of Symbiodinium populations from the Greater Caribbean region. Diversity (allele and MLGs) in these S. trenchii populations was a fraction of what was recovered from minimal host sampling over small areas of Indo-Pacific reefs (less than 50 m2) (Fig. 1). It could be argued that the low genetic diversity in the Atlantic may have arisen from the recent expansion of a small, resident population that was subjected to a severe bottleneck in response to regional environmental degradation (26, 27). However, populations recovering from acute natural selection and reduction in population size should retain relatively high amounts of neutral allelic diversity (28). Additionally, the emergence from one small native population does not account for the lack of alleles unique to the Atlantic, which should exist in a population isolated for millions of years. The most parsimonious explanation is that S. trenchii was introduced to the Greater Caribbean in recent decades from the Indo-Pacific and has rapidly proliferated over the entire region.
S. trenchii possesses many of the physiological and ecological characteristics commonly attributed to successful biological invaders (6). It is common as a host–generalist in a broad diversity of stony coral taxa living in warm, turbid, low-pH Indo-Pacific habitats (20). Furthermore, populations of this Symbiodinium exhibit high genetic connectivity over thousands of kilometers (15). S. trenchii, therefore, is predisposed for ecological opportunism, especially during times of severe stress, or in coral colonies persisting in marginal environments.
Environmental instability brought on by global climate change favors the successful invasion of species with physiological or life history traits that incline them to respond opportunistically (1, 8). The coral reef ecosystems of the Greater Caribbean are among the most impacted in the world from warming climate, elevated pCO2, overfishing, and pollution (27). The rate of climate warming and large-scale environmental degradation in recent decades may have conditioned coral communities to opportunistic invasion by S. trenchii. Several species of symbiotic cnidarians have been introduced to the Greater Caribbean from the Indo-Pacific, yet there is no evidence that the Symbiodinium spp. they carried have become established in native hosts (29, 30). Similarly, the brown sea anemone, Exaiptasia (= Aiptasia), has spread successfully via human transport to locations around the Indo-Pacific (31). However, its symbiont, Symbiodinium minutum, while continually released to the new environments where this animal now persists, does not seem to be compatible with Indo-Pacific cnidarians (32).
Marine pathogens and introduced species have taken only a few years to disperse across the Greater Caribbean (33, 34), supporting the feasibility that a successfully introduced Symbiodinium could spread rapidly. The earliest introductions may have occurred at southern Caribbean ports (Panama, Barbados, and Curaçao) where populations of S. trenchii possess greater genotypic diversity, relative to other Greater Caribbean locations (Fig. 1E). The first documentation of this species in the Atlantic Ocean Basin was from Panama in the mid-to-late 1990s (21, 35). Microorganisms, including dinoflagellates, are transported globally through the ballast water of cargo ships (36), and shipping traffic through the Panama Canal alone may be the source of this introduction. Recent Caribbean-wide surveys have detected S. trenchii at background concentrations in numerous coral genera (37–39), and, as environmental conditions worsen, its prevalence and persistence may increase, especially in corals that are the most sensitive to stress (13, 16).
Under stressful conditions S. trenchii can displace native symbiont populations and dominate a host colony for periods of time lasting a few years, or more (13, 19). It remains unknown what biotic or abiotic factors ultimately drive this competitive outcome. When native symbiont populations decline in hospite during coral bleaching, increases in inorganic nitrogen from partial tissue necrosis may promote the proliferation of low abundance background populations of S. trenchii (40). Therefore, it may be a better competitor when inorganic nitrogen concentrations are high (41). Conversely, its resilience to thermal stress may allow it to assimilate available nitrogen when other dominant symbiont species have physiologically lost this capacity, leading to increased proliferation of this species while other symbionts decline.
Ecosystem Implications.
The long-term effect of a nonnative symbiont on native coral communities may have significant consequences (42). The introduction of S. trenchii to a distinct Atlantic fauna with no coevolutionary history generates associations that are probably suboptimal (i.e., maladapted). Although colonies with this symbiont may cope with increasingly unstable environmental conditions, significant physiological trade-offs may persist that diminish functional performance (43–47). Indeed, our measurements showed that calcification rates were affected in colonies of Orbicella harboring S. trenchii in comparison with native Symbiodinium spp. (when subjected to the environmental conditions of our experiment).
Coral reef accretion requires positive carbonate budgets in which constructive processes, mainly by coral calcification, are larger than the destructive processes of physical, chemical, or biological erosion (48). Some heat-tolerant symbionts have depressed rates of CO2 fixation (44) and may limit the relative amount of photosynthates translocated to the host, which are necessary for calcification (49). Our Pmax and calcification data indicate that S. trenchii may fix amounts of carbon similar to native symbionts (Fig. 3B) but retain a larger portion of these photosynthetic products for its own use. Reduced nutrient translocation in symbioses with S. trenchii would ultimately deprive the host of energy and therefore limit maximum rates of calcification. If extrapolated across large numbers of coral colonies, severe reductions in calcification may be sufficient to influence a coral’s ability to compete successfully for space and ultimately affect reef growth (43, 50). However, additional investigations are needed to determine whether the presence of S. trenchii also imposes limitations in calcification when occurring in other species of Caribbean coral.
The establishment of S. trenchii in the Greater Caribbean highlights a critical need to better understand the consequences of microbial introductions on ecosystem stability and function (6). The responses of coral–dinoflagellate symbioses to climate change are often discussed from the standpoint of the host controlling its partnerships with thermally tolerant symbionts. The behavior of S. trenchii counters such host-centric perspectives by indicating that ecological opportunism and the range expansion by only a few, or one, symbiont species can drive the dynamic response of reef coral symbioses to climate change. These de novo mutualisms open another dimension of research into the functional ecology and evolutionary genetics of coral–dinoflagellate symbioses (51). The ecological opportunism of S. trenchii, for better or worse, will likely affect the regional response of Atlantic coral communities to continued ocean warming.
Materials and Methods
Sample Collections.
Samples of S. trenchii (initially called type D1a, or D1-4) (15) analyzed in this study were originally obtained during large regional investigations of Symbiodinium diversity in the Atlantic (e.g., ref. 52) (Dataset S1), the eastern Indian Ocean, and the western Pacific Ocean (e.g., ref. 20). Small specimens obtained from Cnidarians (mostly scleractianians) were collected by self-contained underwater breathing apparatus (SCUBA) from reefs at eight locations around the Greater Caribbean and included Barbados, Belize, Curaçao, Florida Keys, Flower Garden Banks, Mexico, Panama and St. Croix. S. trenchii was also obtained from two locations in the eastern Indian Ocean (Andaman Sea, Thailand) and two locations in the western Pacific Ocean (Palau) (Fig. 1 B and C). Small (1–2 cm2) fragments with tissue were preserved in either a high salt, 20% DMSO buffer (53) or 95% ethanol and stored at −20 °C until DNA extraction. Collection totals and range of dates for each location are as follows: Greater Caribbean (GC; n = 112, recovered from the screening of over 1,500 samples) from 1999 to 2013, Andaman Sea in the Indian Ocean (IND; n = 57) in 2007, and Palau in the west Pacific (PAC; n = 76) in 2009 (Dataset S1).
Genetic and Physiological Analyses.
Additional methods, along with supporting tables, figures, and access to source codes, are available in Supporting Information.
SI Materials and Methods
Molecular–Genetic Identification.
Nucleic acid extractions were conducted using a modified Promega Wizard genomic DNA extraction protocol (54). The dominant resident symbiont was initially identified by denaturing gradient gel electrophoresis (DGGE) fingerprinting of the partial 5.8S and internal transcribed spacer region 2 (ITS2) (55, 56). All samples used in this study for multilocus genotyping were verified by this protocol to contain high abundances of S. trenchii (20–100% of the total symbiont population). For colonies used in our physiological measurements, only those with homogenous populations of a particular symbiont were selected (Fig. S3).
Microsatellite Allele Calling.
Twelve microsatellite loci (Table S1) developed for clade D Symbiodinium were used to determine the genetic diversity of S. trenchii populations (57, 58). Many of these loci were used previously for other species of clade D Symbiodinium (e.g., Symbiodinium boreum and Symbiodinium eurythalpos) (15) and S. “glynni” (nomen nudum) (22); however, S. trenchii has experienced a partial or whole genome duplication making these loci diallelic (15, 57, 58). Although S. trenchii exists predominantly in the haploid stage, like other dinoflagellates, we treated these loci as diploid in our analyses, unless otherwise noted. Microsatellite fragments were amplified and analyzed according to published methods, and multilocus genotypes (MLGs) were constructed according to previously described methods (22, 59).
Population Genetic Data from Other Species of Symbiodinium.
To investigate the allelic diversity and pairwise genetic differences among MLGs for other Symbiodinium and contrast them with those of S. trenchii in the Greater Caribbean, three previously published datasets representing symbionts within clades A, B, and C were analyzed. Populations of these symbionts were collected from the Greater Caribbean; Symbiodinium “fitti” (nomen nudum = A3) in Acropora palmata from across this coral’s entire Atlantic distribution (n = 652) (23), Symbiodinium B7 in corals of the genus Madracis spp. (n = 35), also from locations around the Greater Caribbean (60), and Symbiodinium C7 in Orbicella spp. (n = 47) from Belize, western Caribbean (61). These three symbiont lineages (formal species descriptions are pending) have experienced similar environmental conditions in their recent evolutionary past, and their population data provide a method to investigate the demographics and genetic variation among populations of Symbiodinium species within this region. A forth Symbiodinium sp. was also included, S. “glynni” (nomen nudum = type D1) in Pocillopora type I from the Gulf of California (n = 109) ( 22) and represents another species of clade D Symbiodinium and a sibling lineage to S. trenchii, and thus provides a comparison of diversity and divergence among MLGs between species within the same clade based on the same microsatellite loci.
Microsatellite Data Analysis.
Populations of organisms that commonly reproduce asexually tend to have a proportion of individuals with identical MLGs (i.e., a clonal cell lineage equivalent to an individual), and, therefore, statistical calculations based on allele frequencies can be negatively biased (discussed in ref. 59). For this reason, all summary statistics and analyses (unless stated otherwise) were conducted with duplicated MLGs in each sampling location removed from datasets. The probability of identity (PI) was calculated to determine the power to resolve genetically distinct individuals, which estimates the probability that two unrelated individuals drawn at random will by chance have the same MLG (62). PIs were calculated for each locus, and then the overall PI for discerning individual clones was determined from the product of all loci. Because these values are affected by population substructure (63), PI was calculated for each regional location. Along with PI, additional summary statistics were calculated for each regional location using GenAlEx (version 6.4) (62) and included observed and expected heterozygosity (Ho and He), the information index (I), and the number of private alleles.
Clonal richness (R) is the frequency of unique MLGs and was calculated to give an indication of the contribution of asexual reproduction within each region. Pairwise genetic difference (PWD) was calculated in the statistical program R (R Core Team) (64) by determining the number of nonidentical alleles between genotypes, and the observed distribution of PWD was compared with that expected in a panmictic population. The expected distribution was calculated by permuting the alleles of the observed genotypes to yield datasets of unlinked multilocus genotypes of equal diversity and size as the observed dataset. One thousand permuted MLG datasets were created, and the PWDs of the permuted genotypes, along with the summary statistics of variance in pairwise genetic difference and the index of association (Ia) (65), were calculated for each of the permuted datasets. The same summary statistics were calculated from the observed data to determine whether there was significant departure from panmixia (R code written for this task is available at doi:10.5061/dryad.d4152). PWD was further used to construct a principal coordinate analysis (PCoA), using GenAlEx, to graphically depict the relatedness between individuals within and between regional populations of S. trenchii. Lastly, a dendrogram of genetic divergence among unique MLGs of S. trenchii was generated using an unweighted pair group method with arithmetic mean (UPGMA) in the Neighbor routine from the PHYLIP package (66) and based on pairwise differences in alleles at each locus.
Photosynthesis and Calcification Measurements.
To investigate the potential implications of S. trenchii in the Greater Caribbean, the physiology of Orbicella faveolata colonies with either S. trenchii, A3, B17, or C7 were examined by measuring rates of photosynthesis and calcification. Fragments (∼11 cm2 SD ± 3.3) from four to six healthy colonies of O. faveolata per symbiont were collected at depths ranging between 4 m and 7 m from La Bocana and La Pared reef sites off Puerto Morelos, Mexico in October 2010. To compensate for possible variations in the photo-physiological state of individual samples derived from differences in the relative position in the colony and depth, fragments were maintained in outdoor flowing seawater tables for 2 mo before been analyzed at constant 28 ± 0.5 °C under neutral density shade cloth providing a maximal midday irradiance of ∼400 μmol quanta m-2⋅s−1. Acclimation in the photophysiology of corals generally requires 1–4 wk (67). Temperature settings were close to the yearly average for the reef whereas the 30% attenuation of the neutral density filters provides irradiance levels similar to the collection sites. Photosynthesis to irradiance curves (P-E) were performed to estimate photosynthetic functioning of each symbiont and according to published methods (68). Briefly, coral fragments were incubated in airtight water-jacked acrylic chambers equipped with Clark-type O2 electrodes (Hansatech) and magnetic stir bars to provide water circulation. The temperature within the chambers was maintained at 28 °C (± 0.5), and the irradiance was supplied using 6-W pure white LED bulbs. Coral fragments were sequentially exposed to 0, 6, 20, 40, 51, 96, 604, and 1,392 μmol quanta m-2⋅s−1 to obtain the rate of oxygen evolution (μmol O2 h−1) at each irradiance. P-E curve parameters were obtained by a nonlinear fitting to a hyperbolic tangent function. Three to five replicate fragments representing each host–symbiont combination were used to obtain an average rate of maximal photosynthesis (Pmax) for each of the four host–symbiont combinations. Pmax was normalized to fragment surface area obtained by imaging each coral specimen and applying the trace function using ImageJ (69).
To estimate instantaneous rates of calcification, fragments were incubated in 200-mL acrylic chambers containing filtered reef water (0.45 μm), each fitted with a stir bar to provide adequate water circulation. Samples were incubated for 1 h at constant temperature (± 0.5 °C) and irradiance (400 μmol quanta m-2⋅s−1) using the same lighting described above. At the end of 1 h, incubation seawater was collected and measured for total alkalinity (± 2.5 μ Eq kg−1) using an Ocean Optics USB4000 spectrophotometer following published procedures (70). Instantaneous rates of calcification (G) were calculated following the alkalinity anomaly principle on the basis that, for each mole of CaCO3 precipitated, total alkalinity is reduced by two molar equivalents (71). Calcification was measured over a range of ecologically relevant temperatures (24, 26, 28, 30, and 32 °C) for all host–symbiont combinations. Four to six O. faveolata fragments (from different parent colonies) representing each partner combination (A3, n = 6 colonies; B17, n = 4; C7, n = 6; S. trenchii, n = 5) were used to measure calcification at each temperature treatment. The same fragments were used across temperature treatments so that, after incubation at one temperature, the fragments were place in tanks at the next highest temperature and allowed to acclimate >2 d between incubations. Instantaneous rates of calcification (μmol CaCO3 h−1) were normalized to coral surface area as described above.
Physiological Data Analyses.
Statistical analyses were performed in the program R V 3.1.1 (R Core Team) using the packages lme4 (72) and lmerTest (73). To test the effect of symbiont identity on coral calcification across all four temperatures, we performed a nested linear mixed effects model, modeling temperature and symbiont identity as fixed effects and coral colony as a random effect nested in symbiont identity. We then calculated the significance (α = 0.05) of pairwise differences of the levels of the fixed effects in lmerTest. Pairwise t tests (data not transformed) were conducted in R to determine significant differences in symbiont photosynthetic rates.
SI Results
ITS2-DGGE Fingerprinting and Sequencing.
DGGE-ITS2 fingerprinting allowed for the rapid detection of S. trenchii in host specimens where this symbiont species was abundant. Its profile is characterized by two codominant ITS2 sequences (sequence “1” and sequence “a,” or “4”; thus, this symbiont was originally called type D1a and later D1-4) (15) that are numerically dominant in the ribosomal array and appear as two bands (plus two heteroduplexes) on a denaturing gel (13, 55, 74). Samples with this fingerprint alone or in mixture with another species of Symbiodinium were used for multilocus genotyping. The significant genetic divergence among major Symbiodinium clades means that microsatellites developed for clade D do not work on Symbiodinium from other clades and thus samples with mixed symbiont assemblages can be analyzed without complications.
Colony fragments of O. faveolata collected for physiological measurements were analyzed before measurements of photosynthesis and calcification. Fragments exhibiting high homogeneity in their symbiont population were selected for physiological analyses (Fig. S1).
Microsatellite Allele Diversity.
The number of alleles per locus ranged from 4 to 29 (Table S1), with the allele frequencies for each location given in Table S2. The frequencies of putative null alleles were rare among loci ranging from 0 for loci D1Sym9 to a value of 0.109 for D1Sym14. Although all of the alleles in the Greater Caribbean were shared with the Indo-Pacific locations, certain alleles were found only in populations from the Indian Ocean or the West Pacific (Table 1). In Palau there were 16 alleles across nine loci (D1Sym9, -17, -34, -66, -67, -82, -87, -88, and -93) that were unique to S. trenchii populations from that region whereas the Andaman Sea populations possessed 16 other alleles distributed across seven loci that were distinct to that geographic location (D1Sym14, -17, -34, -67, --82, -87, and-93). Allele frequency distributions for the Indo-Pacific populations show a dominance in the proportion of rare alleles (0.1 or less) (Fig. S3). In contrast, the proportion of rare alleles was substantially lower in the Greater Caribbean relative to the Indo-Pacific, with a small number of alleles found at higher frequencies (Fig. S3). A total of 147 different genotypes were scored among the 245 MLGs obtained from the entire sample set (Indo-Pacific and Atlantic). Twenty-nine of these MLGs were from the Greater Caribbean, 52 from the Pacific Ocean (Palau), and 66 from the Indian Ocean (Andaman Sea; Table 1).
Observed heterozygosity ranged from 0.65 to 0.78 and was lowest in the Greater Caribbean (Table 1). Observed heterozygosity was higher than expected in every region. Similar to heterozygosity, the information index (I) was higher in the Indo-Pacific (1.24–1.42) than in the Caribbean (0.70). These high values are expected under the assumption that S. trenchii’s genome was recently duplicated and that its “diploid” condition frequently results in two different alleles per locus (15). The probability that two samples with the same MLGs may not have originated from the same clone lineage (PI) was exceedingly low for most locations and ranged from 6.1 × 10−4 (within populations from the Atlantic) to 2.7 × 10−7 (within populations from the Andaman Sea). All of these values were at or well below the 0.0001 criteria recommended by ref. 63 and are therefore adequate for population-level investigations.
Genotypic Diversity, Prevalence, and Distribution.
Clone richness (R), or genotypic richness, was high at both Indo-Pacific locations, ranging from 0.87 to 0.91 (Table 1). The most abundant clone, or MLG, found in the Indo-Pacific was observed in just four samples (Andaman Sea), and no clones were shared between sample locations (Figs. 1B and 1C). By contrast, R was low in populations from the Atlantic Ocean (0.25) where distinct clones were found multiple times within and between sampling locations as far as 3,000 km apart (Table 1 and Fig. 1E). In the most extreme case, genotype “α” was found in 47 of the 112 samples from around the Greater Caribbean and at every location except the Flower Garden Banks (Figs. 1E and 2A). A list of all S. trenchii MLGs is provided in Table S1, organized by region, location, and host taxon.
The mean pairwise genetic difference between individuals was high from locations in the Andaman Sea (Indian Ocean) and Pacific Ocean (15.16 and 12.03 differences, respectively) but was much lower in the Atlantic Ocean (4.32) (Fig. 2C). The Andaman Sea and Palau showed a bell-shaped distribution around the mean pairwise genetic difference for each population, and their observed distribution closely matched that expected in sexually recombining populations. In contrast, pairwise genetic difference from the Atlantic Ocean was skewed toward zero and was substantially different from the distribution expected from that region’s allele frequency data (and higher Ho than He). The skewed distribution of the Atlantic Ocean is attributed to the high similarity between individuals within this region, many of which (∼96%) differ by only four alleles, or fewer, and involving one or two loci.
Genotypic Variation Among Other Pacific and Atlantic Symbiodinium.
The results from pairwise genetic difference analyses of Symbiodinium representatives from clades A, B, and C, and a sibling lineage in clade D, S. “glynni,” (whose genomes are not duplicated) were similar to that observed for S. trenchii populations in the Indo-Pacific, where the observed distribution closely overlaps that expected in a panmictic population (Fig. S2D). These results indicate that other lineages of Symbiodinium are genetically diverse and that therefore the low genetic variation and high clonality observed for S. trenchii in the Greater Caribbean are not typical of the region. The average pairwise genetic differences calculated for these other Symbiodinium were as follows: S. “fitti” (6.43, examining the 13 microsatellites) (23), Symbiodinium B7 (6.25, examining the 9 microsatellite loci) (58), Symbiodinium C7 (1.73, examining 10 microsatellite loci) (59), and S. “glynni” (5.50, examining 11 microsatellite loci) (22). All datasets showed a significant departure from panmixia, probably arising from regional genetic structure, asexual reproduction, or both processes. Because each locus is haploid for these species (i.e., one allele per locus), the mean pairwise genetic difference is correspondingly lower than calculated for an equivalent number of loci in S. trenchii populations.
Source Code Availability.
Source codes and data files for the analysis of observed pairwise genetic difference, expected pairwise genetic difference under panmixia, and permutation test for departure from expected pairwise genetic difference among MLGs from datasets for S. trenchii populations in the Indian, Pacific, and Atlantic Oceans, as well as for S. “glynni” (type D1), type B7, type A3, and C7 were deposited in the DYRAD database.
Supplementary Material
Acknowledgments
We thank the numerous colleagues who helped with collections of specimens used in the analysis of S. trenchii genetics, including Mark Warner and Dustin Kemp, who provided samples from Curaçao; Kemp provided additional samples from Belize, and Jorge Pinzón collected samples from Panama. We also appreciated the constructive suggestions and criticisms provided by anonymous reviewers, which helped improve the manuscript. This work was funded in part by Pennsylvania State University, Florida International University, and National Science Foundation Grants OCE-0928764 and IOS-1258058, -1258065, and -1258063, the Canon Foundation (Japan), and Programa de Apoyos a Proyectos de Investigación e Innovación Tecnológica Grant IN206413 (to R.I.-P.), and a Professional Association of Diving Instructors (PADI) grant (to D.T.P.).
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
Data deposition: Source codes and data files for the analysis of observed pairwise genetic difference (PWD), expected pairwise genetic difference under panmixia, and permutation test for departure from expected pairwise genetic difference among MLGs from datasets for S. trenchii populations in the Indian, Pacific, and Atlantic Oceans, as well as for S. “glynni” (type D1), type B7, type A3, and C7 reported in this paper have been deposited in the Dryad database, www.datadryad.org (doi: 10.5061/dryad.d4152).
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1502283112/-/DCSupplemental.
References
- 1.Dukes JS, Mooney HA. Does global change increase the success of biological invaders? Trends Ecol Evol. 1999;14(4):135–139. doi: 10.1016/s0169-5347(98)01554-7. [DOI] [PubMed] [Google Scholar]
- 2.Mack RN, et al. Biotic invasions: Causes, epidemiology, global consequences, and control. Ecol Appl. 2000;10:689–710. [Google Scholar]
- 3.Walther GR, et al. Ecological responses to recent climate change. Nature. 2002;416(6879):389–395. doi: 10.1038/416389a. [DOI] [PubMed] [Google Scholar]
- 4.Parmesan C. Ecological and evolutionary responses to recent climate change. Annu Rev Ecol Evol Syst. 2006;37:637–669. [Google Scholar]
- 5.Pejchar L, Mooney HA. Invasive species, ecosystem services and human well-being. Trends Ecol Evol. 2009;24(9):497–504. doi: 10.1016/j.tree.2009.03.016. [DOI] [PubMed] [Google Scholar]
- 6.Litchman E. Invisible invaders: Non-pathogenic invasive microbes in aquatic and terrestrial ecosystems. Ecol Lett. 2010;13(12):1560–1572. doi: 10.1111/j.1461-0248.2010.01544.x. [DOI] [PubMed] [Google Scholar]
- 7.Vellinga EC, Wolfe BE, Pringle A. Global patterns of ectomycorrhizal introductions. New Phytol. 2009;181(4):960–973. doi: 10.1111/j.1469-8137.2008.02728.x. [DOI] [PubMed] [Google Scholar]
- 8.Walther GR, et al. Alien species in a warmer world: Risks and opportunities. Trends Ecol Evol. 2009;24(12):686–693. doi: 10.1016/j.tree.2009.06.008. [DOI] [PubMed] [Google Scholar]
- 9.Hoegh-Guldberg O, et al. Coral reefs under rapid climate change and ocean acidification. Science. 2007;318(5857):1737–1742. doi: 10.1126/science.1152509. [DOI] [PubMed] [Google Scholar]
- 10.Fitt WK, Brown BE, Warner ME, Dunne RP. Coral bleaching: Interpretation of thermal tolerance limits and thermal thresholds in tropical corals. Coral Reefs. 2001;20:51–65. [Google Scholar]
- 11.Baker AC, Starger CJ, McClanahan TR, Glynn PW. Coral reefs: Corals’ adaptive response to climate change. Nature. 2004;430(7001):741. doi: 10.1038/430741a. [DOI] [PubMed] [Google Scholar]
- 12.Berkelmans R, van Oppen MJ. The role of zooxanthellae in the thermal tolerance of corals: A ‘nugget of hope’ for coral reefs in an era of climate change. Proc Biol Sci. 2006;273(1599):2305–2312. doi: 10.1098/rspb.2006.3567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.LaJeunesse TC, Smith RT, Finney J, Oxenford H. Outbreak and persistence of opportunistic symbiotic dinoflagellates during the 2005 Caribbean mass coral ‘bleaching’ event. Proc Biol Sci. 2009;276(1676):4139–4148. doi: 10.1098/rspb.2009.1405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jones AM, Berkelmans R, van Oppen MJ, Mieog JC, Sinclair W. A community change in the algal endosymbionts of a scleractinian coral following a natural bleaching event: Field evidence of acclimatization. Proc Biol Sci. 2008;275(1641):1359–1365. doi: 10.1098/rspb.2008.0069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.LaJeunesse TC, et al. Ecologically differentiated stress-tolerant endosymbionts in the dinoflagellate genus Symbiodinium (Dinophyceae) Clade D are different species. Phycologia. 2014;53:305–319. [Google Scholar]
- 16.Grottoli AG, et al. The cumulative impact of annual coral bleaching can turn some coral species winners into losers. Glob Change Biol. 2014;20(12):3823–3833. doi: 10.1111/gcb.12658. [DOI] [PubMed] [Google Scholar]
- 17.Kemp DW, Hernandez-Pech X, Iglesias-Prieto R, Fitt WK, Schmidt GW. Community dynamics and physiology of Symbiodinium spp. before, during, and after a coral bleaching event. Limnol Oceanogr. 2014;59:788–797. [Google Scholar]
- 18.Silverstein RN, Cunning R, Baker AC. Change in algal symbiont communities after bleaching, not prior heat exposure, increases heat tolerance of reef corals. Glob Change Biol. 2014;21(1):236–249. doi: 10.1111/gcb.12706. [DOI] [PubMed] [Google Scholar]
- 19.Thornhill DJ, LaJeunesse TC, Kemp DW, Fitt WK, Schmidt GW. Multi-year, seasonal genotypic surveys of coral-algal symbioses reveal prevalent stability or post-bleaching reversion. Mar Biol. 2006;148:711–722. [Google Scholar]
- 20.LaJeunesse TC, et al. Long‐standing environmental conditions, geographic isolation and host–symbiont specificity influence the relative ecological dominance and genetic diversification of coral endosymbionts in the genus Symbiodinium. J Biogeogr. 2010;37:785–800. [Google Scholar]
- 21.Toller WW, Rowan R, Knowlton N. Repopulation of Zooxanthellae in the Caribbean corals Montastraea annularis and M. faveolata following experimental and disease-associated bleaching. Biol Bull. 2001;201(3):360–373. doi: 10.2307/1543614. [DOI] [PubMed] [Google Scholar]
- 22.Pettay DT, Wham DC, Pinzón JH, LaJeunesse TC. Genotypic diversity and spatial-temporal distribution of Symbiodinium clones in an abundant reef coral. Mol Ecol. 2011;20(24):5197–5212. doi: 10.1111/j.1365-294X.2011.05357.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Baums IB, Devlin-Durante MK, LaJeunesse TC. New insights into the dynamics between reef corals and their associated dinoflagellate endosymbionts from population genetic studies. Mol Ecol. 2014;23(17):4203–4215. doi: 10.1111/mec.12788. [DOI] [PubMed] [Google Scholar]
- 24.Nei M, Chakraborty R, Fuerst PA. Infinite allele model with varying mutation rate. Proc Natl Acad Sci USA. 1976;73(11):4164–4168. doi: 10.1073/pnas.73.11.4164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Luikart G, Allendorf FW, Cornuet JM, Sherwin WB. Distortion of allele frequency distributions provides a test for recent population bottlenecks. J Hered. 1998;89(3):238–247. doi: 10.1093/jhered/89.3.238. [DOI] [PubMed] [Google Scholar]
- 26.Gardner TA, Côté IM, Gill JA, Grant A, Watkinson AR. Long-term region-wide declines in Caribbean corals. Science. 2003;301(5635):958–960. doi: 10.1126/science.1086050. [DOI] [PubMed] [Google Scholar]
- 27.Mora C. A clear human footprint in the coral reefs of the Caribbean. Proc Biol Sci. 2008;275(1636):767–773. doi: 10.1098/rspb.2007.1472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wootton JC, et al. Genetic diversity and chloroquine selective sweeps in Plasmodium falciparum. Nature. 2002;418(6895):320–323. doi: 10.1038/nature00813. [DOI] [PubMed] [Google Scholar]
- 29.LaJeunesse TC, Lee S, Bush S, Bruno JF. Persistence of non-Caribbean algal symbionts in Indo-Pacific mushroom corals released to Jamaica 35 years ago. Coral Reefs. 2005;24:157–159. [Google Scholar]
- 30.LaJeunesse TC, Loh W, Trench RK. Do introduced endosymbiotic dinoflagellates ‘take’ to new hosts? Biol Inv. 2009;11:995–1003. [Google Scholar]
- 31.Thornhill DJ, Xiang Y, Pettay DT, Zhong M, Santos SR. Population genetic data of a model symbiotic cnidarian system reveal remarkable symbiotic specificity and vectored introductions across ocean basins. Mol Ecol. 2013;22(17):4499–4515. doi: 10.1111/mec.12416. [DOI] [PubMed] [Google Scholar]
- 32.LaJeunesse TC, et al. Host-symbiont recombination versus natural selection in the response of coral-dinoflagellate symbioses to environmental disturbance. Proc Biol Sci. 2010;277(1696):2925–2934. doi: 10.1098/rspb.2010.0385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lessios HA, Robertson DR, Cubit JD. Spread of diadema mass mortality through the Caribbean. Science. 1984;226(4672):335–337. doi: 10.1126/science.226.4672.335. [DOI] [PubMed] [Google Scholar]
- 34.Hines A, Acero P, Ortí G, Wilbur AE, Freshwater DW. Reconstructing the lionfish invasion: Insights into Greater Caribbean biogeography. J Biogeogr. 2011;38:1281–1293. [Google Scholar]
- 35.Toller WW, Rowan R, Knowlton N. Zooxanthellae of the Montastraea annularis species complex: Patterns of distribution of four taxa of Symbiodinium on different reefs and across depths. Biol Bull. 2001;201(3):348–359. doi: 10.2307/1543613. [DOI] [PubMed] [Google Scholar]
- 36.Hallegraeff GM. Transport of toxic dinoflagellates via ships' ballast water: Bioeconomic risk assessment and efficacy of possible ballast water management strategies. Mar Ecol Prog Ser. 1998;168:297–309. [Google Scholar]
- 37.Smith RT. 2008. Specificity, stability and comparative physiology of coral-Symbiodinium mutualisms: Evaluating the potential for acclimation and/or adaptation in reef corals. PhD dissertation (Florida International University, Miami, FL)
- 38.Correa AM, McDonald MD, Baker AC. Development of clade-specific Symbiodinium primers for quantitative PCR (qPCR) and their application to detecting clade D symbionts in Caribbean corals. Mar Biol. 2009;156:2403–2411. [Google Scholar]
- 39.Kennedy EV, Foster NL, Mumby PJ, Stevens JR. Widespread prevalence of cryptic Symbiodinium D in the key Caribbean reef builder, Orbicella annularis. Coral Reefs. 2015;34:519–531. [Google Scholar]
- 40.Fitt WK, Spero HJ, Halas J, White MW, Porter JW. Recovery of the coral Montastrea annularis in the Florida Keys after the 1987 Caribbean “bleaching event”. Coral Reefs. 1993;12:57–64. [Google Scholar]
- 41.Baker DM, Andras JP, Jordán-Garza AG, Fogel ML. Nitrate competition in a coral symbiosis varies with temperature among Symbiodinium clades. ISME J. 2013;7(6):1248–1251. doi: 10.1038/ismej.2013.12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ortiz JC, González-Rivero M, Mumby PJ. Can a thermally tolerant symbiont improve the future of Caribbean coral reefs? Glob Change Biol. 2013;19(1):273–281. doi: 10.1111/gcb.12027. [DOI] [PubMed] [Google Scholar]
- 43.Little AF, van Oppen MJ, Willis BL. Flexibility in algal endosymbioses shapes growth in reef corals. Science. 2004;304(5676):1492–1494. doi: 10.1126/science.1095733. [DOI] [PubMed] [Google Scholar]
- 44.Cantin NE, van Oppen MJ, Willis BL, Mieog JC, Negri AP. Juvenile corals can acquire more carbon from high-performance algal symbionts. Coral Reefs. 2009;28:405–414. [Google Scholar]
- 45.Abrego D, Ulstrup KE, Willis BL, van Oppen MJ. Species-specific interactions between algal endosymbionts and coral hosts define their bleaching response to heat and light stress. Proc Biol Sci. 2008;275(1648):2273–2282. doi: 10.1098/rspb.2008.0180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Mieog JC, et al. The roles and interactions of symbiont, host and environment in defining coral fitness. PLoS ONE. 2009;4(7):e6364. doi: 10.1371/journal.pone.0006364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Jones A, Berkelmans R. Potential costs of acclimatization to a warmer climate: Growth of a reef coral with heat tolerant vs. sensitive symbiont types. PLoS ONE. 2010;5(5):e10437. doi: 10.1371/journal.pone.0010437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Adey WH. Coral reef morphogenesis: A multidimensional model. Science. 1978;202(4370):831–837. doi: 10.1126/science.202.4370.831. [DOI] [PubMed] [Google Scholar]
- 49.Colombo-Pallotta MF, Rodríguez-Román A, Iglesias-Prieto R. Calcification in bleached and unbleached Montastraea faveolata: Evaluating the role of oxygen and glycerol. Coral Reefs. 2010;29:899–907. [Google Scholar]
- 50.Kennedy EV, et al. Avoiding coral reef functional collapse requires local and global action. Curr Biol. 2013;23(10):912–918. doi: 10.1016/j.cub.2013.04.020. [DOI] [PubMed] [Google Scholar]
- 51.Lee CE. Evolutionary genetics of invasive species. Trends Ecol Evol. 2002;17:386–391. [Google Scholar]
- 52.Finney JC, et al. The relative significance of host-habitat, depth, and geography on the ecology, endemism, and speciation of coral endosymbionts in the genus Symbiodinium. Microb Ecol. 2010;60(1):250–263. doi: 10.1007/s00248-010-9681-y. [DOI] [PubMed] [Google Scholar]
- 53.Seutin G, White BN, Boag PT. Preservation of avian blood and tissue samples for DNA analyses. Can J Zool. 1991;69:82–92. [Google Scholar]
- 54.LaJeunesse TC, et al. Low symbiont diversity in southern Great Barrier Reef corals, relative to those of the Caribbean. Limnol Oceanogr. 2003;48:2046–2054. [Google Scholar]
- 55.LaJeunesse TC. Diversity and community structure of symbiotic dinoflagellates from Caribbean coral reefs. Mar Biol. 2002;141:387–400. [Google Scholar]
- 56.Lajeunesse TC, Trench RK. Biogeography of two species of Symbiodinium (Freudenthal) inhabiting the intertidal sea anemone Anthopleura elegantissima (Brandt) Biol Bull. 2000;199(2):126–134. doi: 10.2307/1542872. [DOI] [PubMed] [Google Scholar]
- 57.Pettay DT, Lajeunesse TC. Microsatellite loci for assessing genetic diversity, dispersal and clonality of coral symbionts in ‘stress-tolerant’ clade D Symbiodinium. Mol Ecol Resour. 2009;9(3):1022–1025. doi: 10.1111/j.1755-0998.2009.02561.x. [DOI] [PubMed] [Google Scholar]
- 58.Wham DC, Pettay DT, LaJeunesse TC. Microsatellite loci for the host-generalist “zooxanthella” Symbiodinium trenchi and other Clade D Symbiodinium. Conserv Genet Resour. 2011;3:541–544. [Google Scholar]
- 59.Pettay DT, LaJeunesse TC. Long-range dispersal and high-latitude environments influence the population structure of a “stress-tolerant” dinoflagellate endosymbiont. PLoS ONE. 2013;8(11):e79208. doi: 10.1371/journal.pone.0079208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Pettay DT, LaJeunesse TC. Microsatellites from clade B Symbiodinium spp. specialized for Caribbean corals in the genus Madracis. Mol Ecol Notes. 2007;7:1271–1274. [Google Scholar]
- 61.Thornhill DJ, Lewis AM, Wham DC, LaJeunesse TC. Host-specialist lineages dominate the adaptive radiation of reef coral endosymbionts. Evolution. 2014;68(2):352–367. doi: 10.1111/evo.12270. [DOI] [PubMed] [Google Scholar]
- 62.Peakall R, Smouse PE. GENALEX 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes. 2006;6:288–295. doi: 10.1093/bioinformatics/bts460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Waits LP, Luikart G, Taberlet P. Estimating the probability of identity among genotypes in natural populations: Cautions and guidelines. Mol Ecol. 2001;10(1):249–256. doi: 10.1046/j.1365-294x.2001.01185.x. [DOI] [PubMed] [Google Scholar]
- 64. R Core Team (2014) R: A language and environment for statistical computing (R Foundation for Statistical Computing, Vienna)
- 65.Anthony KRN, Hoegh-Guldberg O. Kinetics of photoacclimation in corals. Oecologia. 2003;134(1):23–31. doi: 10.1007/s00442-002-1095-1. [DOI] [PubMed] [Google Scholar]
- 66.Smith JM, Smith NH, O’Rourke M, Spratt BG. How clonal are bacteria? Proc Natl Acad Sci USA. 1993;90(10):4384–4388. doi: 10.1073/pnas.90.10.4384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Felsenstein J. 2004. PHYLIP: Phylogeny Inference Package (Department of Genome Sciences, University of Washington, Seattle, WA), Version 3.6.
- 68.Rodríguez-Román A, et al. Photosynthesis and light utilization in the Caribbean coral Montastraea faveolata recovering from a bleaching event. Limnol Oceanogr. 2006;51:2701–2710. [Google Scholar]
- 69.Abramoff MD, Magalhaes PJ, Ram SJ. Image processing with ImageJ. Biophoton Int. 2004;11:36–42. [Google Scholar]
- 70.Yao W, Byrne RH. Simplified seawater alkalinity analysis: Use of linear array spectrometers. Deep Sea Res Part I Oceanogr Res Pap. 1998;45:1383–1392. [Google Scholar]
- 71.Smith SV, Kinsey DW. 1978. Calcification and organic carbon metabolism as indicated by carbon dioxide. Coral Reefs: Research Methods (Monographs on Oceanographic Methodology), eds Stoddart DR, Johannes RE (UNESCO, Paris), Number 5, pp 469–484.
- 72.Bates D, Maechler M, Bolker BM, Walker S. 2014 Fitting linear mixed-effects models using Ime4. arxiv.org/abs/1406.5823.
- 73.Kuznetsova A, Brockhoff PB, Christensen RHB. 2014 lmerTest: Tests in Linear Mixed Effects Models. cran.r-project.org.
- 74.Thornhill DJ, Lajeunesse TC, Santos SR. Measuring rDNA diversity in eukaryotic microbial systems: How intragenomic variation, pseudogenes, and PCR artifacts confound biodiversity estimates. Mol Ecol. 2007;16(24):5326–5340. doi: 10.1111/j.1365-294X.2007.03576.x. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.