Abstract
Background
Prenatal mercury (Hg) exposure is associated with adverse child neurobehavioral outcomes. Because Hg can interfere with placental functioning and cross the placenta to target the fetal brain, prenatal Hg exposure can inhibit fetal growth and development directly and indirectly.
Objectives
We examined potential associations between prenatal Hg exposure assessed through infant toenail Hg, placental DNA methylation changes, and newborn neurobehavioral outcomes.
Methods
The methylation status of > 485,000 CpG loci was interrogated in 192 placental samples using Illumina’s Infinium HumanMethylation450 BeadArray. Hg concentrations were analyzed in toenail clippings from a subset of 41 infants; neurobehavior was assessed using the NICU Network Neurobehavioral Scales (NNNS) in an independent subset of 151 infants.
Results
We identified 339 loci with an average methylation difference > 0.125 between any two toenail Hg tertiles. Variation among these loci was subsequently found to be associated with a high-risk neurodevelopmental profile (omnibus p-value = 0.007) characterized by the NNNS. Ten loci had p < 0.01 for the association between methylation and the high-risk NNNS profile. Six of 10 loci reside in the EMID2 gene and were hypomethylated in the 16 high-risk profile infants’ placentas. Methylation at these loci was moderately correlated (correlation coefficients range, –0.33 to –0.45) with EMID2 expression.
Conclusions
EMID2 hypomethylation may represent a novel mechanism linking in utero Hg exposure and adverse infant neurobehavioral outcomes.
Citation
Maccani JZ, Koestler DC, Lester B, Houseman EA, Armstrong DA, Kelsey KT, Marsit CJ. 2015. Placental DNA methylation related to both infant toenail mercury and adverse neurobehavioral outcomes. Environ Health Perspect 123:723–729; http://dx.doi.org/10.1289/ehp.1408561
Introduction
Multiple studies have found associations between in utero, childhood, or early adulthood mercury (Hg) exposure and later neurologic and psychological impairment. One of the most cited is a study of Faroe Islands children exposed to Hg predominantly through a seafood-heavy diet, showing adverse neurobehavioral outcomes at 7 and 14 years of age (Grandjean et al. 1997). Early-life Hg exposure is associated with neurodevelopmental deficits (Counter and Buchanan 2004), including reduced newborn cerebellum size (Cace et al. 2011), adverse behavioral outcomes (Gao et al. 2007), central nervous system damage (Choi 1989), poor psychomotor development (Llop et al. 2012), cognitive developmental delays (Freire et al. 2010), and later-life effects (Rice 1996), including increased diabetes susceptibility (He et al. 2013).
The placenta is crucial in regulating fetal growth and development, including neurodevelopment (Lester and Padbury 2009; O’Keeffe and Kenny 2014). In utero environmental toxicant exposures may disrupt placental function, affecting growth factor and hormone production and detoxification activity (Maccani and Marsit 2009). Toxicants may interfere with placental function through epigenetic alterations, including changes in normal placental DNA methylation patterns (Burris et al. 2012; Suter et al. 2011; Wilhelm-Benartzi et al. 2012), which control the expression of genes involved in key placental cellular processes. Abnormal methylation alterations may have serious consequences for placental growth and functioning and, in turn, for developing infants’ health.
Hg crosses the placenta (Ilbäck et al. 1991; National Research Council 2000; Yang et al. 1997) and also accumulates within the placenta, where methylmercury (MeHg) concentrations can be double those of maternal blood (Ask et al. 2002) and disrupt placental functioning (Boadi et al. 1992). A common exposure source is fish consumption (Davidson et al. 2004), although occupational exposures and maternal dental amalgams with inorganic Hg (Davidson et al. 2004; Takahashi et al. 2001) can also increase placental Hg. A single amalgam restoration is associated with a 3- to 6-fold increase in placental Hg (Takahashi et al. 2001).
MeHg exposure has been associated with SEPP1 hypomethylation in adult blood (Goodrich et al. 2013). SEPP1 encodes a selenoprotein potentially involved in Hg toxicity protection (Goodrich et al. 2011), suggesting that methylation may be exposure-responsive. Although SEPP1 is expressed and active in the placenta (Kasik and Rice 1995), there have been no examinations of SEPP1 methylation or its relationship to Hg in the placenta. The placenta is active during development, and variation in placental methylation at various genes has been associated with fetal growth and development and neurobehavior (Banister et al. 2011; Bromer et al. 2012; Filiberto et al. 2011; Marsit et al. 2012a, 2012b; Wilhelm-Benartzi et al. 2012). Thus, Hg-associated placental alterations may mediate exposure-associated neurobehavioral outcomes, even at exposure levels commonly identified in the population. Previous studies (He et al. 2013; Hinners et al. 2012; Wickre et al. 2004; Xun et al. 2013) have assessed toenail Hg for integrated exposure estimates. We hypothesized that prenatal Hg exposure, assessed through infant toenail Hg, is associated with altered placental methylation patterns that are, in turn, associated with adverse infant neurobehavioral outcomes.
Methods
Study design. This sample included 192 infants with placental specimens from the Rhode Island Child Health Study (RICHS), a birth cohort of nonpathologic term pregnancies delivered at Women and Infants’ Hospital in Providence, Rhode Island. Participants underwent an informed consent process approved by the Institutional Review Boards of Women and Infants’ Hospital and Dartmouth College. Eligible infants were born at ≥ 37 weeks gestation, and small- and large-for-gestational-age (SGA and LGA) infants were oversampled. By definition, SGA infants weigh ≤ 10th percentile for their gestational age; 6 of 41 (14.6%) infants in the Hg subcohort and 36 of 151 (23.8%) infants in the NNNS (NICU Network Neurobehavioral Scales) subcohort had a birthweight percentile ≤ 10%. By definition, LGA infants weigh ≥ 90th percentile for their gestational age; 14 of 41 (34.1%) infants in the Hg subcohort and 45 of 151 (29.8%) infants in the NNNS subcohort had a birth weight percentile ≥ 90%. This analysis included 41 samples with Hg data and an independent subcohort of 151 samples with neurobehavioral assessments. Within 2 hr of birth, full-thickness sections were taken from the maternal side of the placenta and 2 cm from the umbilical cord-insertion site, free of maternal decidua. These sections were immediately placed in RNAlaterTM (AM7020; Applied Biosystems Inc.). Following ≥ 72 hr at 4°C, samples were blotted dry, snap-frozen in liquid nitrogen, homogenized via pulverization and stored at –80°C until analysis. Infants were examined with a newborn neurobehavioral assessment, the NNNS (Lester and Tronick 2004), after 24 hr of life, but before hospital discharge. Examinations were performed from 24 to 96 hr following birth.
Exposure assessment. First toenail clippings from all toes were requested from mothers as well as infants following discharge, and were available for 41 of 192 infants. Parents were asked to collect their own and their children’s toenail clippings and mail back toenail clippings in provided envelopes. Average time from birth to collection was 2.8 months (range, 0.3–7.1 months). Micrograms Hg per gram of toenail were analyzed (Rees et al. 2007) in the Dartmouth Trace Element Analysis laboratory. Within batches, samples below the limit of detection limit (LOD) were assigned a value half the lowest observed Hg value in that batch. Average LOD across batches was 0.382 μg/g; 26 samples were below LOD.
DNA extraction and modification. DNA was extracted, quantified, and bisulfite modified via QIAmp DNA Mini Kit (51304; Qiagen), ND-1000 spectrophotometer (NanoDrop) and EZ DNA Methylation Kit (D5008; Zymo Research).
Methylation profiling. Placental methylation was assessed at the University of Minnesota Genomics Center via Illumina Infinium HumanMethylation450 BeadArray (Illumina). Samples were randomized across batches stratified by birth weight group and sex. β-values—the ratio of fluorescent signals from methylated (M) and unmethylated (U) alleles—were used as the measure of methylation status at each locus, where β = Max(M,0)/[Max(M,0) + Max(U,0) + 100]. β-values ranged from 0 (no methylation) to 1 (complete methylation). Array quality assurance was assessed; poor-performing loci, X- and Y-linked loci, and SNP (single nucleotide polymorphism)–associated loci were removed (Banister et al. 2011), yielding 384,474 loci for 192 infants.
Statistical analysis. Figure 1 presents our analysis strategy. Before analysis, we assured random sample distribution across batches by Hg tertile and neurobehavioral profile; there were no associations between Hg exposure tertile and the chip or plate on which the placenta DNA sample was arrayed (p > 0.05). Methylation data were adjusted for plate effects via ComBat (Johnson et al. 2007), which performs effectively compared with competing adjustment methodologies. Effectiveness of this adjustment was assessed using principal components analysis and assuring no association between plate or chip and the top three principal components (all p > 0.05). In 41 infants with Hg data, the omnibus association between Hg tertile and methylation over 384,474 loci was tested via permutation test (Westfall and Young 1993), involving 384,474 linear regression models, one per locus, each permuted 1,000 times and controlled for maternal age (in years), birth weight percentile (continuous), delivery method (vaginal or cesarean section), and infant sex. Minimum p-value (over individual regression models for 384,474 loci) was used as a test statistic. Its null distribution was obtained by 1,000 draws from the permutation distribution obtained by permuting infant toenail Hg with respect to methylation and putative confounders. To avoid assuming linear response, to allow capture of relationships at the highest exposures, and to limit bias due to detection limits, we used tertiles in all analyses (Kuan et al. 2010). Individual, locus-specific p-values for Hg tertile were computed via standard F-test for H0:β1 = β2 = 0, where coefficients β1 and β2 correspond to nonreferent tertiles. Δβ-values were calculated as the difference in mean β-values between any tertile pairs. To balance sensitivity (i.e., the need to identify a comprehensive list of loci) and specificity (i.e., the need to limit false discovery), we limited the analysis of methylation and the high-risk neurobehavioral outcome to loci with Δβ > 0.125 for at least one pair of Hg tertiles.
Figure 1.
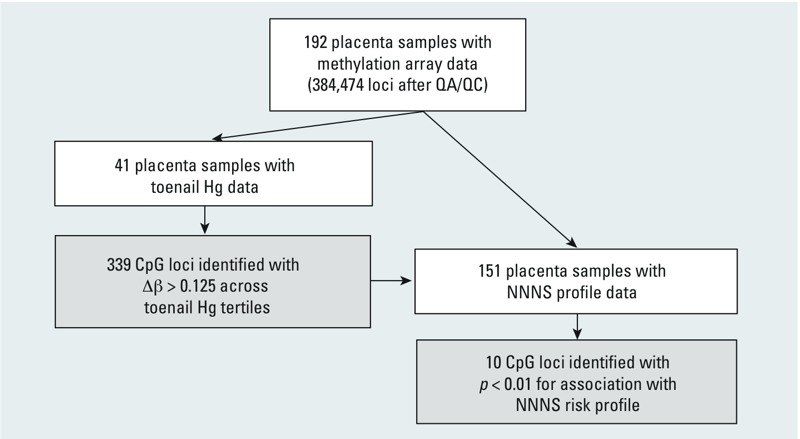
Analysis strategy: 192 placental samples were arrayed on a HumanMethylation450 BeadArray. Following quality assurance, sex-linked and SNP-associated loci were removed. Forty-one samples with Hg data were analyzed for Hg-associated placental methylation differences; 339 loci with methylation differences between Hg tertiles > 0.125 were analyzed for associations with high-risk NNNS profile in an independent set of 151 samples with NNNS data.
Similar to latent profile analyses described for NNNS scores (Liu et al. 2010), mutually exclusive neurobehavioral profiles based on 13 NNNS scores were defined using recursively partitioned mixture modeling (Lesseur et al. 2013). From this analysis, seven profiles were identified, with one profile demonstrating similar attributes to that described as high-risk by Liu et al. (2010). We defined these infants as “high-risk” compared with all other infants in further analyses. Infants in this high-risk group demonstrated poorer quality of movement, poorer self-regulation, increased signs of physiologic stress and abstinence, greater excitability and the need for additional techniques for handling the infant to change state. For details on each of the summary scores between the high-risk group and other infants, see Supplemental Material, Table S1. For loci with greatest methylation differences by Hg level, we estimated the null multivariate distribution of test statistics via permutation distribution (controlled for potential confounders above) to investigate associations with the high-risk infant neurobehavioral profile in an independent sample (n = 151) of infants from the same study population, for whom NNNS data were available. Socioeconomic status (measured by maternal education) was examined as a possible confounder; no significant associations were identified between Hg or NNNS profile and maternal education (p = 0.37 and p = 0.70, respectively; chi-square tests), so these were not included in final models for parsimony.
Heatmaps were created in R (R Core Team 2014), using a Euclidean distance measure, to cluster placenta samples and loci based on methylation of 339 Hg-associated loci.
Gene expression. Total RNA was extracted via RNeasy Mini Kit (Qiagen), quantified via Nanodrop 2000 (ThermoFisher Scientific), aliquoted, and stored at –80C. Expression analysis was performed via CFX Connect Real-Time PCR Detection System (BioRad). First-strand reactions were performed in triplicate with BioRad iScript cDNA Synthesis Kit and qPCR (quantitative polymerase chain reaction) reaction with BioRad iQ SYBER Green Supermix. The sample with lowest expression served as a reference sample for delta-delta-Ct normalization. EMID2 and SDHA expression were measured using primers: EMID2: forward 5´-TTTCAGCCTTGGACTTAGCGA, reverse 5´-GCCAAAATCCTGT CCTTGTCA, SDHA: forward 5´-TGCTCAGTATCCAGTAGTGGA, reverse 5´-TTCTCTTACCTGTGCTGCAA.
Results
Table 1 describes the two study groups (infants with toenail Hg data, n = 41; and infants with NNNS data, n = 151). All infants were born at ≥ 37 weeks gestation, as required for the parent study. There is oversampling for SGA and LGA infants. Children in the two study groups were generally similar with regard to maternal age, infant sex, birth weight, or gestation time. No mothers of Hg-subcohort infants reported smoking. Among NNNS-subcohort infants, there were higher percentages of non-Caucasian mothers and cesarean section births than in the Hg-subcohort infants. Low (referent) Hg tertile ranged from 0.005 μg/g to 0.031 μg/g; medium, 0.032 μg/g to 0.076 μg/g; high, 0.077 μg/g to 0.425 μg/g. These values fall largely within a toenail Hg reference range of 0.07–0.38 μg/g derived from 130 healthy volunteers in a French study (Goullé et al. 2009). Within the Hg subcohort, there were more male infants within the medium tertile, and birth weights were higher amongst this tertile; thus, these variables were included in all models.
Table 1.
Study population demographics.
| Variable | Subset 1: Infants with toenail Hg data (n = 41) | Subset 2: Infants with NNNS data (n = 151) | |||||
|---|---|---|---|---|---|---|---|
| Low Hg tertile (n = 14) | Medium Hg tertile (n = 13) | High Hg tertile (n = 14) | p-Value | Non-high-risk profile (n = 135) | High-risk profile (n = 16) | p-Value | |
| Infant sex | |||||||
| Female [n (%)] | 8 (57.1) | 3 (23.1) | 10 (71.4) | 0.037 | 65 (48.1) | 12 (75.0) | 0.08 |
| Male [n (%)] | 6 (42.9) | 10 (76.9) | 4 (28.6) | 70 (51.9) | 4 (25.0) | ||
| Maternal age (years) | |||||||
| Mean ± SD | 31.9 ± 3.1 | 32.8 ± 4.4 | 31.4 ± 3.3 | 0.76 | 28.4 ± 6.0 | 26.8 ± 6.1 | 0.33 |
| Median (range) | 32.5 (26–39) | 33 (23–39) | 30 (26–38) | 29 (18–40) | 26.5 (18–38) | ||
| Tobacco use during pregnancya | |||||||
| Yes [n (%)] | 0 (0) | 0 (0) | 0 (0) | NA | 7 (5.2) | 2 (12.5) | 0.55 |
| No [n (%)] | 14 (100) | 13 (100) | 14 (100) | 126 (93.3) | 14 (87.5) | ||
| Birth weight (g) | |||||||
| Mean ± SD | 3647.5 ± 628.2 | 3978.4 ±473.3 | 3175.7 ± 524.4 | 0.046 | 3462.9 ± 737.7 | 3443.8 ± 779.6 | 0.93 |
| Median (range) | 3,740 (2,280–4,465) | 4,185 (3,035–4,530) | 3,230 (2,160–4,090) | 3,415 (1,705–5,465) | 3,385 (2,370–4,570) | ||
| Gestational age (weeks) | |||||||
| Mean ± SD | 39.8 ± 1.0 | 39.5 ± 1.0 | 39.5 ± 1.3 | 0.53 | 39.2 ± 1.1 | 39.7 ± 1.3 | 0.44 |
| Median (Range) | 40 (37.4–41.3) | 39.7 (37.3–41.1) | 39.8 (37.1–41.1) | 39.3 (37–41.9) | 39.5 (37–41.3) | ||
| Maternal ethnicity | |||||||
| Caucasian [n (%)] | 13 (92.9) | 13 (100) | 11 (78.6) | 0.53 | 99 (73.3) | 10 (62.5) | 0.62 |
| Non-Caucasian [n (%)] | 1 (7.1) | 0 (0) | 3 (21.4) | 36 (26.7) | 6 (37.5) | ||
| Cesarean section delivery | |||||||
| Yes [n (%)] | 8 (57.1) | 9 (69.2) | 10 (71.4) | 0.69 | 71 (52.6) | 7 (43.8) | 0.69 |
| No [n (%)] | 6 (42.9) | 4 (30.8) | 4 (28.6) | 64 (47.4) | 9 (56.3) | ||
| Recreational drug use during pregnancy | |||||||
| Yes [n (%)] | 0 (0) | 0 (0) | 1 (7.1) | 0.37 | 3 (2.2) | 1 (6.3) | 0.9 |
| No [n (%)] | 14 (100) | 13 (100) | 13 (92.9) | 132 (97.8) | 15 (93.8) | ||
| Maternal educationb | |||||||
| High school graduate/equivalent or less [n (%)] | 2 (14.3) | 0 (0) | 1 (7.1) | 0.37 | 49 (36.3) | 5 (31.3) | 0.79 |
| Junior college graduate/equivalent or greater [n (%)] | 12 (85.7) | 13 (100) | 12 (85.7) | 86 (63.7) | 11 (68.8) | ||
| NA, not applicable.aOne sample with Hg data missing smoking data. bOne sample with Hg data missing education data. | |||||||
Mean methylation β-values were calculated for each locus by Hg tertile. Placental methylation epigenome-wide was associated with Hg (omnibus p = 0.017). At 339 loci, methylation differed by > 0.125 between tertiles (Figure 2; see also Supplemental Material, Table S2); generally, samples clustered by Hg and sex, but not by maternal ethnicity, maternal age, or birthweight group. Mean β-values increased monotonically with increasing Hg tertiles for 79 loci, 34 loci had a monotonic decrease with increasing tertiles, and 226 loci had a non-monotonic relationship across tertiles.
Figure 2.
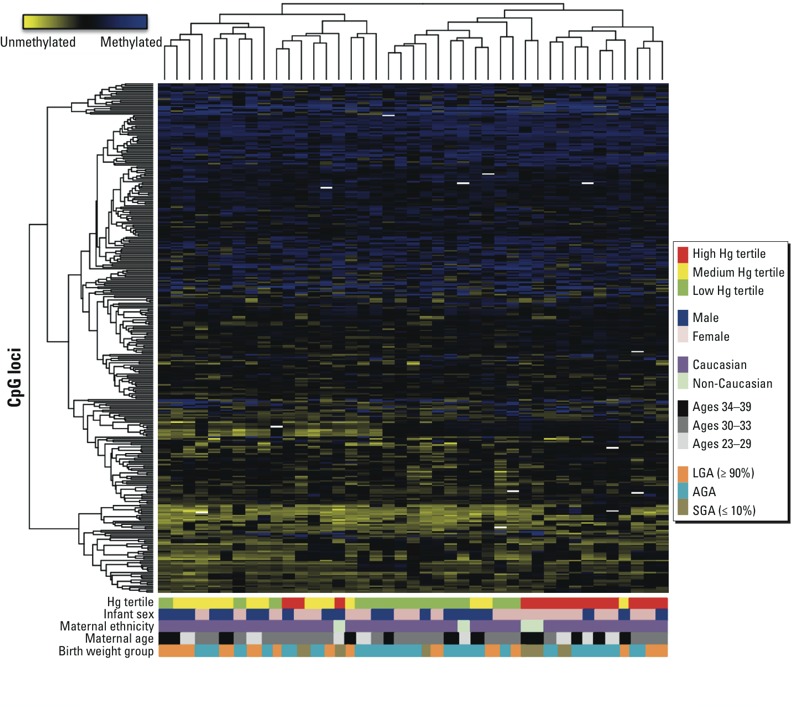
Heat map demonstrating Hg tertile differences > 0.125. Placental samples in columns; 339 loci in rows. Methylation β-values indicated by key at top left. Below figure, color bars indicate Hg tertiles (green, low tertile; yellow, medium; red, high), infant sex (blue, males; pink, females), maternal ethnicity (purple, Caucasian; green, non-Caucasian), maternal age tertiles (light gray, 23–29 years; dark gray, 30–33 years; black, 34–39 years), birth weight group [orange, LGA (≥ 90%); teal, appropriate-for-gestational-age (AGA); olive, SGA (≤ 10%)].
We performed supervised clustering of samples with NNNS profiles using 339 Hg-associated loci (see Supplemental Material, Figure S1), but observed no obvious clustering pattern of high-risk neurobehavioral profile. Thus, we examined individual association of loci with high-risk profile using a series of linear models; comparison of the distribution of p-values obtained from these models to a null distribution determined by permutation suggested that some degree of variability in risk for high-risk neurobehavioral profile membership could be attributed to methylation variation at these loci (omnibus p = 0.007). See Supplemental Material, Table S2, for profiles of the results of individual models. Ten loci (Table 2) residing in CPLX1, TTC23, and EMID2 were associated with high-risk profile (p < 0.01). Six of 10 reside in a CpG island within EMID2, the only gene with multiple loci associated with high-risk profile within those loci at p < 0.01.
Table 2.
Ten loci associated with infant toenail Hg tertile (omnibus p = 0.017 and Δβ > 0.125 between any two Hg tertiles) and high-risk NNNS profile (p < 0.01).
| Illumina CpG designation | Genomic position | Relation to CpG island | Gene symbol | p-Value for high-risk NNNS profile |
|---|---|---|---|---|
| cg13267931 | Chr 7: 101006308 | Island | EMID2 | 8.25 × 10–6 |
| cg14175932 | Chr 14: 23018807a | 2.84 × 10–5 | ||
| cg27179533 | Chr 7: 101006052 | Island | EMID2 | 5.46 × 10–5 |
| cg14874750 | Chr 7: 101006063 | Island | EMID2 | 6.06 × 10–5 |
| cg23424003 | Chr 7: 101006035 | Island | EMID2 | 7.30 × 10–5 |
| cg27528510 | Chr 7: 101006058 | Island | EMID2 | 9.00 × 10–5 |
| cg14048874 | Chr 7: 101006573 | Island | EMID2 | 0.0023 |
| cg17128947 | Chr 4: 779480 | Island | CPLX1 | 0.0054 |
| cg25385940 | Chr 15: 99789637 | N Shoreb | TTC23 | 0.0059 |
| cg10470368 | Chr 11: 64146517a | 0.0075 | ||
| Chr, chromosome. aAccording to Illumina array annotation, these loci are not located within an annotated CpG region and are not associated with any gene. bThe north shore of a CpG island is defined as the region just upstream (5’) of the CpG island region. | ||||
Four of six loci are within 200 bases of EMID2’s transcription start site: cg13267931 is in the 5´ untranslated region upstream of the first exon, and cg14048874 in the gene body. Figure 3 illustrates their methylation by Hg tertile. In general, those infants in the highest tertile of exposure demonstrated the highest extent of methylation at each of the CpGs present on the array.
Figure 3.
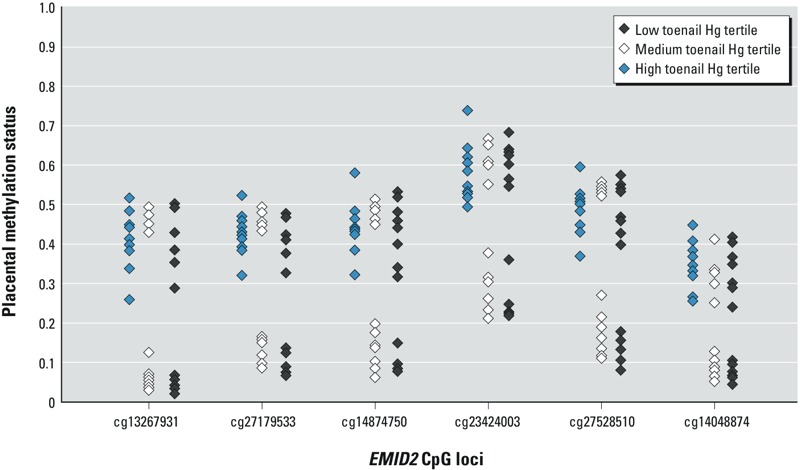
Plot of six Hg- and high-risk profile–associated EMID2 loci in 41 samples with Hg data by tertile. y-Axis represents EMID2 methylation β-value. Loci in order of appearance (+ strand, 5’ to 3’).
We then examined the average extent of methylation across all of the EMID2 loci in the NNNS subset, comparing those infants in the high-risk and non–high-risk groups. As shown in Figure 4, those in the high-risk group demonstrated hypomethylation of this gene. qRT-PCR in a subset of samples revealed moderate correlations between placental methylation at these loci and EMID2 gene expression, with correlation coefficients for individual CpG loci and expression ranging from –0.33 to –0.45 (see Supplemental Material, Figure S2).
Figure 4.
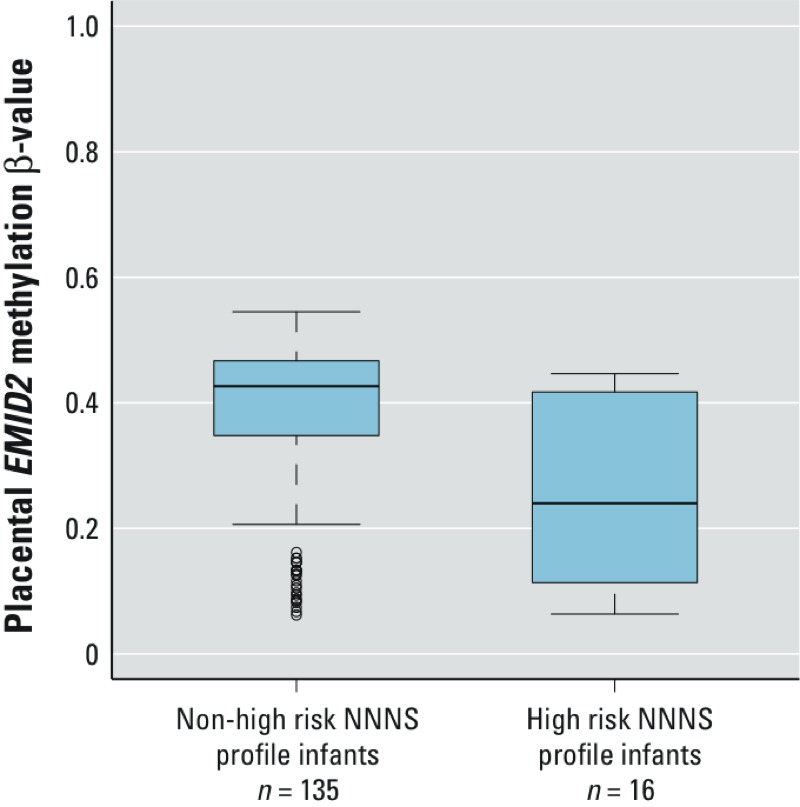
Average β-value across six EMID2 loci associated with Hg and high-risk NNNS profile in an independent subset of 151 infants. Boxes extend from 25th to 75th percentile, horizontal bars represent medians, and whiskers extend 1.5 times the length of the interquartile range above and below the 75th and 25th percentiles, respectively. Outliers are represented as points.
Discussion
Placental methylation patterns were associated with infant toenail Hg and a potential high-risk infant neurobehavioral profile in our study population. Many of the 339 loci with greatest differences by Hg (see Supplemental Material, Table S2) reside in neurodevelopment-, neurogenesis- and behavior-related genes based on mutant or knockout gene studies in animal models, gene expression and knockdown studies, as well as whole genome and/or in silico studies (Barreto-Valer et al. 2013; Glynn et al. 2007; Heinrich et al. 2012; Ju et al. 2014; Kivimäe et al. 2011; Konno et al. 2012; Kremerskothen et al. 2002; Larsson et al. 2008; Morimura et al. 2006; Porro et al. 2010; Shimizu et al. 2010; Silver et al. 2012). Some have been associated with neurodevelopmental disorders: schizophrenia (DIXDC1, ARVCF, MAGI2, ZIC2) (Bradshaw and Porteous 2012; Chen et al. 2005; Sim et al. 2012), ADHD (attention deficit/hyperactivity disorder) (TCERG1L) (Neale et al. 2010), movement disorders (NOL3, TP53INP2) (Bennetts et al. 2007; Russell et al. 2012), Huntington’s disease (H2AFY2, AGPAT1) (Cong et al. 2012; Hu et al. 2011), Parkinson’s disease (LMX1B) (Tian et al. 2012), and autism (PLXNA4, WNT2) (Kalkman 2012; Suda et al. 2011). Others have been associated with diabetes (ZBED3) (Ohshige et al. 2011), asthma (EMID2) (Pasaje et al. 2011), and cancer (FBXO3, HOOK2, MT2A, EIF3E, RPH3AL, PTRF, MT1M, STK32A) (Cha et al. 2011; Krzeslak et al. 2013; Liu et al. 2012; Mao et al. 2012; Shimada et al. 2005).
Because of previously reported links between Hg and neurodevelopmental deficits and numerous Hg-variable genes involved in neurodevelopment, we examined these loci for associations with a high-risk newborn neurobehavioral profile defined by the NNNS, which are associated with later-life behavioral outcomes (Lester and Tronick 2004; Liu et al. 2010). In this analysis, 16 infants were observed to have a high-risk NNNS profile. We used a latent profile methodology to account for correlations between these scales and reduce data dimensionality. Liu et al. (2010) reported associations of such profiles with later-childhood outcomes: acute medical and behavior problems, school readiness, and IQ through 4.5 years of age. Of 339 loci, 10 (Table 2) were associated with a high-risk profile (p < 0.01) similar to that of Liu et al. (2010); 6 of 10 resided in the EMID2 promoter.
Although EMID2’s placental function is unknown, its genetic variation has been associated with aspirin-sensitive asthma (Pasaje et al. 2011), and with hearing and vision side effects of the antidepressant citalopram (Adkins et al. 2012). EMID2 (or COL26A1) contains an emilin and two collagen domains primarily expressed in testes and ovary (Sato et al. 2002). EMID2 is linked to a sonic hedgehog (SHH) enhancer adoption mutation, where an EMID2 enhancer drives ectopic SHH expression (Lettice et al. 2011), although the loci identified are not located within that enhancer element. Future investigation is warranted to define EMID2’s placental role and how its modulation can impact neurodevelopment. It may be of interest to explore its role in SHH, which plays roles in neural tube patterning and cell survival (Ho and Scott 2002; McCarthy and Argraves 2003).
Interestingly, this potential risk neurobehavioral profile was associated with EMID2 hypomethylation in low- and medium-Hg tertiles, with greatest hypomethylation in the mid-range group. This suggests a nonmonotonic and potentially complicated relationship between exposure, methylation, and outcome. We were limited in our ability to address these relationships in the same individuals. In addition, as we were making use of infant toenail samples, a large proportion were below the limits of detection for the assay, so extrapolation to a dose response may not be possible. Therefore, we urge caution in this interpretation, particularly until these results can be expanded and validated in a larger population.
Evidence from an autopsy study of adults has suggested strong correlations between levels of total Hg in toenails and MeHg levels in blood or occipital cortex (Björkman et al. 2007), suggesting that toenails are relevant biomarkers. Because of slow growth of toenails, toenail Hg likely reflects exposures in the past 3–5 months (Goullé et al. 2009). Thus, infant toenail Hg likely reflects prenatal exposures occurring over most of pregnancy. We note that toenail Hg observed in this cohort falls within toenail Hg reference ranges (Goullé et al. 2009), suggesting we are likely examining common, low-level variation in exposure and associations with methylation, which potentially could contribute to later developmental deficiencies. An important future direction will be investigating potential postnatal epigenetic × environment interactions in high-risk profile children in addition to confirming these findings in additional cohorts.
Limitations to this study include undetermined Hg exposure sources, infant toenail Hg as a proxy for prenatal exposure, use of term placentas, a relatively small sample size (including n = 16 high-risk NNNS profile infants), independent sample sets for Hg and neurobehavior analyses, which limits our ability to examine direct relationships between them, and a large proportion of samples falling below the limit of detection. Future analyses may benefit from examining, in larger data sets with Hg and NNNS data, whether high-risk profile infants were also exposed to more Hg. Since EMID2 methylation has not been associated with Hg or neurodevelopment, and its placental function is unknown, it is unclear whether hypo- or hypermethylation with high-risk profile is expected, and future mechanistic and epidemiologic studies should investigate this.
Conclusions
This study provides evidence for a potential role for placental epigenetic alterations as a mechanism linking Hg exposure and adverse infant neurodevelopment, and specifically a role for EMID2. This suggests possible associations between prenatal Hg exposure, placental methylation changes, and the developmental origins of mental/behavioral and physical health and disease.
Supplemental Material
Footnotes
This research was funded by grants R01MH094609 from the National Institute of Mental Health/National Institutes of Health (NIH); R01ES022223, P01 ES022832, and T32ES007272 from the National Institute of Environmental Health Sciences/NIH; and RD83544201 from the U.S. Environmental Protection Agency. J.Z.J.M. received support from grant T32HL076134-04 from the National Heart, Lung, and Blood Institute for a postdoctoral fellowship at Brown University and the Miriam Hospital. J.Z.J.M. is now at the Penn State College of Medicine Tobacco Center of Regulatory Science (TCORS) and is funded by grant P50-DA-036107-01 from the NIH.
The authors declare they have no actual or potential competing financial interests.
References
- Adkins DE, Clark SL, Åberg K, Hettema JM, Bukszár J, McClay JL, et al. 2012Genome-wide pharmacogenomic study of citalopram-induced side effects in STAR*D. Transl Psychiatry 2e129; 10.1038/tp.2012.57 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ask K, Åkesson A, Berglund M, Vahter M. Inorganic mercury and methylmercury in placentas of Swedish women. Environ Health Perspect. 2002;110:523–526. doi: 10.1289/ehp.02110523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Banister CE, Koestler DC, Maccani MA, Padbury JF, Houseman EA, Marsit CJ. Infant growth restriction is associated with distinct patterns of DNA methylation in human placentas. Epigenetics. 2011;6(7):920–927. doi: 10.4161/epi.6.7.16079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barreto-Valer K, Lopez-Bellido R, Rodriguez RE. Cocaine modulates the expression of transcription factors related to the dopaminergic system in zebrafish. Neuroscience. 2013;231:258–271. doi: 10.1016/j.neuroscience.2012.11.052. [DOI] [PubMed] [Google Scholar]
- Bennetts JS, Rendtorff ND, Simpson F, Tranebjaerg L, Wicking C. The coding region of TP53INP2, a gene expressed in the developing nervous system, is not altered in a family with autosomal recessive non-progressive infantile ataxia on chromosome 20q11-q13. Dev Dyn. 2007;236(3):843–852. doi: 10.1002/dvdy.21064. [DOI] [PubMed] [Google Scholar]
- Björkman L, Lundekvam BF, Laegreid T, Bertelsen BI, Morild I, Lilleng P, et al. Mercury in human brain, blood, muscle and toenails in relation to exposure: an autopsy study. Environ Health. 2007;11:6–30. doi: 10.1186/1476-069X-6-30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boadi WY, Urbach J, Brandes JM, Yannai S. In vitro exposure to mercury and cadmium alters term human placental membrane fluidity. Toxicol Appl Pharmacol. 1992;116(1):17–23. doi: 10.1016/0041-008x(92)90139-j. [DOI] [PubMed] [Google Scholar]
- Bradshaw NJ, Porteous DJ. DISC1-binding proteins in neural development, signalling and schizophrenia. Neuropharmacology. 2012;62(3):1230–1241. doi: 10.1016/j.neuropharm.2010.12.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bromer C, Marsit CJ, Armstrong DA, Padbury JF, Lester B. Genetic and epigenetic variation of the glucocorticoid receptor (NR3C1) in placenta and infant neurobehavior. Dev Psychobiol. 2012;55(7):673–683. doi: 10.1002/dev.21061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burris HH, Rifas-Shiman SL, Baccarelli A, Tarantini L, Boeke CE, Kleinman K, et al. Associations of LINE-1 DNA methylation with preterm birth in a prospective cohort study. J Dev Orig Health Dis. 2012;3(3):173–181. doi: 10.1017/s2040174412000104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cace IB, Milardovic A, Prpic I, Krajina R, Petrovic O, Vukelic P, et al. Relationship between the prenatal exposure to low-level of mercury and the size of a newborn’s cerebellum. Med Hypotheses. 2011;76(4):514–516. doi: 10.1016/j.mehy.2010.12.005. [DOI] [PubMed] [Google Scholar]
- Cha JD, Kim HJ, Cha IH. Genetic alterations in oral squamous cell carcinoma progression detected by combining array-based comparative genomic hybridization and multiplex ligation-dependent probe amplification. Oral Surg Oral Med Oral Pathol Oral Radiol Endod. 2011;111(5):594–607. doi: 10.1016/j.tripleo.2010.11.020. [DOI] [PubMed] [Google Scholar]
- Chen HY, Yeh JI, Hong CJ, Chen CH. Mutation analysis of ARVCF gene on chromosome 22q11 as a candidate for a schizophrenia gene. Schizophr Res. 2005;72(2–3):275–277. doi: 10.1016/j.schres.2004.03.004. [DOI] [PubMed] [Google Scholar]
- Choi BH. The effects of methylmercury on the developing brain. Prog Neurobiol. 1989;32(6):447–470. doi: 10.1016/0301-0082(89)90018-x. [DOI] [PubMed] [Google Scholar]
- Cong WN, Cai H, Wang R, Daimon CM, Maudsley S, Raber K, et al. 2012Altered hypothalamic protein expression in a rat model of Huntington’s disease. PLoS One 710e47240; 10.1371/journal.pone.0047240 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Counter SA, Buchanan LH. Mercury exposure in children: a review. Toxicol Appl Pharmacol. 2004;198(2):209–230. doi: 10.1016/j.taap.2003.11.032. [DOI] [PubMed] [Google Scholar]
- Davidson PW, Myers GJ, Weiss B. Mercury exposure and child development outcomes. Pediatrics. 2004;113(4) suppl:1023–1029. [PubMed] [Google Scholar]
- Filiberto AC, Maccani MA, Koestler D, Wilhelm-Benartzi C, Avissar-Whiting M, Banister CE, et al. Birthweight is associated with DNA promoter methylation of the glucocorticoid receptor in human placenta. Epigenetics. 2011;6(5):566–572. doi: 10.4161/epi.6.5.15236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Freire C, Ramos R, Lopez-Espinosa MJ, Diez S, Vioque J, Ballester F, et al. Hair mercury levels, fish consumption, and cognitive development in preschool children from Granada, Spain. Environ Res. 2010;110(1):96–104. doi: 10.1016/j.envres.2009.10.005. [DOI] [PubMed] [Google Scholar]
- Gao Y, Yan CH, Tian Y, Wang Y, Xie HF, Zhou X, et al. Prenatal exposure to mercury and neurobehavioral development of neonates in Zhoushan City, China. Environ Res. 2007;105(3):390–399. doi: 10.1016/j.envres.2007.05.015. [DOI] [PubMed] [Google Scholar]
- Glynn D, Sizemore RJ, Morton AJ. Early motor development is abnormal in complexin 1 knockout mice. Neurobiol Dis. 2007;25(3):483–495. doi: 10.1016/j.nbd.2006.10.011. [DOI] [PubMed] [Google Scholar]
- Goodrich JM, Basu N, Franzblau A, Dolinoy DC. Mercury biomarkers and DNA methylation among Michigan dental professionals. Environ Mol Mutagen. 2013;54(3):195–203. doi: 10.1002/em.21763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodrich JM, Wang Y, Gillespie B, Werner R, Franzblau A, Basu N. Glutathione enzyme and selenoprotein polymorphisms associate with mercury biomarker levels in Michigan dental professionals. Toxicol Appl Pharmacol. 2011;257(2):301–308. doi: 10.1016/j.taap.2011.09.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goullé JP, Saussereau E, Mahieu L, Bouige D, Groenwont S, Guerbet M, et al. Application of inductively coupled plasma mass spectrometry multielement analysis in fingernail and toenail as a biomarker of metal exposure. J Anal Toxicol. 2009;33:92–98. doi: 10.1093/jat/33.2.92. [DOI] [PubMed] [Google Scholar]
- Grandjean P, Weihe P, White RF, Debes F, Araki S, Yokoyama K, et al. Cognitive deficit in 7-year-old children with prenatal exposure to methylmercury. Neurotoxicol Teratol. 1997;19(6):417–428. doi: 10.1016/s0892-0362(97)00097-4. [DOI] [PubMed] [Google Scholar]
- He K, Xun P, Liu K, Morris S, Reis J, Guallar E. Mercury exposure in young adulthood and incidence of diabetes later in life: the CARDIA Trace Element Study. Diabetes Care. 2013;36(6):1584–1589. doi: 10.2337/dc12-1842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heinrich J, Proepper C, Schmidt T, Linta L, Liebau S, Boeckers TM. The postsynaptic density protein Abelson interactor protein 1 interacts with the motor protein Kinesin family member 26B in hippocampal neurons. Neuroscience. 2012;221:86–95. doi: 10.1016/j.neuroscience.2012.06.055. [DOI] [PubMed] [Google Scholar]
- Hinners T, Tsuchiya A, Stern AH, Burbacher TM, Faustman EM, Mariën K.2012Chronologically matched toenail-Hg to hair-Hg ratio: temporal analysis within the Japanese community (U.S.). Environ Health 1181; 10.1186/1476-069X-11-81 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ho KS, Scott MP. Sonic hedgehog in the nervous system: functions, modifications and mechanisms. Curr Opin Neurobiol. 2002;12(1):57–63. doi: 10.1016/s0959-4388(02)00290-8. [DOI] [PubMed] [Google Scholar]
- Hu Y, Chopra V, Chopra R, Locascio JJ, Liao Z, Ding H, et al. Transcriptional modulator H2A histone family, member Y (H2AFY) marks Huntington disease activity in man and mouse. Proc Natl Acad Sci USA. 2011;108(41):17141–17146. doi: 10.1073/pnas.1104409108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ilbäck NG, Sundberg J, Oskarsson A. Methyl mercury exposure via placenta and milk impairs natural killer (NK) cell function in newborn rats. Toxicol Lett. 1991;58(2):149–158. doi: 10.1016/0378-4274(91)90169-7. [DOI] [PubMed] [Google Scholar]
- Johnson WE, Li C, Rabinovic A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics. 2007;8(1):118–127. doi: 10.1093/biostatistics/kxj037. [DOI] [PubMed] [Google Scholar]
- Ju XD, Guo Y, Wang NN, Huang Y, Lai MM, Zhai YH, et al. Both Myosin-10 isoforms are required for radial neuronal migration in the developing cerebral cortex. Cereb Cortex. 2014;24(5):1259–1268. doi: 10.1093/cercor/bhs407. [DOI] [PubMed] [Google Scholar]
- Kalkman HO.2012A review of the evidence for the canonical Wnt pathway in autism spectrum disorders. Mol Autism 3110; 10.1186/2040-2392-3-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kasik JW, Rice EJ. Selenoprotein P expression in liver, uterus and placenta late in pregnancy. Placenta. 1995;16(1):67–74. doi: 10.1016/0143-4004(95)90082-9. [DOI] [PubMed] [Google Scholar]
- Kivimäe S, Martin PM, Kapfhamer D, Ruan Y, Heberlein U, Rubenstein JL, et al. 2011Abnormal behavior in mice mutant for the Disc1 binding partner, Dixdc1. Transl Psychiatry 1e43; 10.1038/tp.2011.41 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konno D, Iwashita M, Satoh Y, Momiyama A, Abe T, Kiyonari H, et al. 2012The mammalian DM domain transcription factor Dmrta2 is required for early embryonic development of the cerebral cortex. PLoS One 710e46577; 10.1371/journal.pone.0046577 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kremerskothen J, Teber I, Wendholt D, Liedtke T, Böckers TM, Barnekow A. Brain-specific splicing of α-actinin 1 (ACTN1) mRNA. Biochem Biophys Res Commun. 2002;295(3):678–681. doi: 10.1016/s0006-291x(02)00734-9. [DOI] [PubMed] [Google Scholar]
- Krzeslak A, Forma E, Chwatko G, Jozwiak P, Szymczyk A, Wilkosz J, et al. Effect of metallothionein 2A gene polymorphism on allele-specific gene expression and metal content in prostate cancer. Toxicol Appl Pharmacol. 2013;268(3):278–285. doi: 10.1016/j.taap.2013.02.013. [DOI] [PubMed] [Google Scholar]
- Kuan PF, Wang S, Zhou X, Chu H. A statistical framework for Illumina DNA methylation arrays. Bioinformatics. 2010;26(22):2849–2855. doi: 10.1093/bioinformatics/btq553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsson J, Forsberg M, Brännvall K, Zhang XQ, Enarsson M, Hedborg F, et al. Nuclear receptor binding protein 2 is induced during neural progenitor differentiation and affects cell survival. Mol Cell Neurosci. 2008;39(1):32–39. doi: 10.1016/j.mcn.2008.05.013. [DOI] [PubMed] [Google Scholar]
- Lesseur C, Armstrong DA, Paquette AG, Koestler DC, Padbury JF, Marsit CJ. Tissue-specific Leptin promoter DNA methylation is associated with maternal and infant perinatal factors. Mol Cell Endocrinol. 2013;381(1–2):160–167. doi: 10.1016/j.mce.2013.07.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lester BM, Padbury JF. Third pathophysiology of prenatal cocaine exposure. Dev Neurosci. 2009;31(1–2):23–35. doi: 10.1159/000207491. [DOI] [PubMed] [Google Scholar]
- Lester BM, Tronick EZ. History and description of the Neonatal Intensive Care Unit Network Neurobehavioral Scale. Pediatrics. 2004;113(3 pt 2):634–640. [PubMed] [Google Scholar]
- Lettice LA, Daniels S, Sweeney E, Venkataraman S, Devenney PS, Gautier P, et al. Enhancer-adoption as a mechanism of human developmental disease. Hum Mutat. 2011;32(12):1492–1499. doi: 10.1002/humu.21615. [DOI] [PubMed] [Google Scholar]
- Liu J, Bann C, Lester B, Tronick E, Das A, Lagasse L, et al. Neonatal neurobehavior predicts medical and behavioral outcome. Pediatrics. 2010;125(1):e90–e98. doi: 10.1542/peds.2009-0204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu YP, Shan BE, Wang XL, Ma L. Correlation between MTA2 overexpression and tumour progression in esophageal squamous cell carcinoma. Exp Ther Med. 2012;3(4):745–749. doi: 10.3892/etm.2012.475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Llop S, Guxens M, Murcia M, Lertxundi A, Ramon R, Riaño I, et al. Prenatal exposure to mercury and infant neurodevelopment in a multicenter cohort in Spain: study of potential modifiers. Am J Epidemiol. 2012;175(5):451–465. doi: 10.1093/aje/kwr328. [DOI] [PubMed] [Google Scholar]
- Maccani MA, Marsit CJ. Epigenetics in the placenta. Am J Reprod Immunol. 2009;62(2):78–89. doi: 10.1111/j.1600-0897.2009.00716.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mao J, Yu H, Wang C, Sun L, Jiang W, Zhang P, et al. Metallothionein MT1M is a tumor suppressor of human hepatocellular carcinomas. Carcinogenesis. 2012;33(12):2568–2577. doi: 10.1093/carcin/bgs287. [DOI] [PubMed] [Google Scholar]
- Marsit CJ, Lambertini L, Maccani MA, Koestler DC, Houseman EA, Padbury JF, et al. Placenta-imprinted gene expression association of infant neurobehavior. J Pediatr. 2012a;160(5):854–860.e2. doi: 10.1016/j.jpeds.2011.10.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marsit CJ, Maccani MA, Padbury JF, Lester BM.2012bPlacental 11-beta hydroxysteroid dehydrogenase methylation is associated with newborn growth and a measure of neurobehavioral outcome. PLoS One 73e33794; 10.1371/journal.pone.0033794 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCarthy RA, Argraves WS. Megalin and the neurodevelopmental biology of sonic hedgehog and retinol. J Cell Sci. 2003;116(Pt 6):955–960. doi: 10.1242/jcs.00313. [DOI] [PubMed] [Google Scholar]
- Morimura N, Inoue T, Katayama K, Aruga J. Comparative analysis of structure, expression and PSD95-binding capacity of Lrfn, a novel family of neuronal transmembrane proteins. Gene. 2006;380(2):72–83. doi: 10.1016/j.gene.2006.05.014. [DOI] [PubMed] [Google Scholar]
- National Research Council. Washington, DC: National Academy Press; 2000. Toxicological Effects of Methylmercury. [Google Scholar]
- Neale BM, Medland S, Ripke S, Anney RJ, Asherson P, Buitelaar J, et al. Case-control genome-wide association study of attention-deficit/hyperactivity disorder. J Am Acad Child Adolesc Psychiatry. 2010;49(9):906–920. doi: 10.1016/j.jaac.2010.06.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohshige T, Iwata M, Omori S, Tanaka Y, Hirose H, Kaku K, et al. 2011Association of new loci identified in European genome-wide association studies with susceptibility to type 2 diabetes in the Japanese. PLoS One 610e26911; 10.1371/journal.pone.0026911 [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Keeffe GW, Kenny LC. Predicting infant neurodevelopmental outcomes using the placenta? Trends Mol Med. 2014;20(6):303–305. doi: 10.1016/j.molmed.2014.04.005. [DOI] [PubMed] [Google Scholar]
- Pasaje CF, Kim JH, Park BL, Cheong HS, Kim MK, Choi IS, et al. A possible association of EMID2 polymorphisms with aspirin hypersensitivity in asthma. Immunogenetics. 2011;63(1):13–21. doi: 10.1007/s00251-010-0490-8. [DOI] [PubMed] [Google Scholar]
- Porro F, Rosato-Siri M, Leone E, Costessi L, Iaconcig A, Tongiorgi E, et al. β-Adducin (Add2) KO mice show synaptic plasticity, motor coordination and behavioral deficits accompanied by changes in the expression and phosphorylation levels of the α- and γ-adducin subunits. Genes Brain Behav. 2010;9(1):84–96. doi: 10.1111/j.1601-183X.2009.00537.x. [DOI] [PubMed] [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing. Vienna, Austria:R Foundation for Statistical Computing. 2014. Available: http://R-project.org [accessed 1 March 2014]
- Rees JR, Sturup S, Chen C, Folt C, Karagas MR. Toenail mercury and dietary fish consumption. J Expo Sci Environ Epidemiol. 2007;17(1):25–30. doi: 10.1038/sj.jes.7500516. [DOI] [PubMed] [Google Scholar]
- Rice DC. Evidence for delayed neurotoxicity produced by methylmercury. Neurotoxicology. 1996;17(3–4):583–596. [PubMed] [Google Scholar]
- Russell JF, Steckley JL, Coppola G, Hahn AF, Howard MA, Kornberg Z, et al. Familial cortical myoclonus with a mutation in NOL3. Ann Neurol. 2012;72(2):175–183. doi: 10.1002/ana.23666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sato K, Yomogida K, Wada T, Yorihuzi T, Nishimune Y, Hosokawa N, et al. Type XXVI collagen, a new member of the collagen family, is specifically expressed in the testis and ovary. J Biol Chem. 2002;277(40):37678–37684. doi: 10.1074/jbc.M205347200. [DOI] [PubMed] [Google Scholar]
- Shimada H, Nakashima K, Ochiai T, Nabeya Y, Takiguchi M, Nomura F, et al. Serological identification of tumor antigens of esophageal squamous cell carcinoma. Int J Oncol. 2005;26(1):77–86. [PubMed] [Google Scholar]
- Shimizu T, Nakazawa M, Kani S, Bae YK, Shimizu T, Kageyama R, et al. Zinc finger genes Fezf1 and Fezf2 control neuronal differentiation by repressing Hes5 expression in the forebrain. Development. 2010;137(11):1875–1885. doi: 10.1242/dev.047167. [DOI] [PubMed] [Google Scholar]
- Silver M, Janousova E, Hua X, Thompson PM, Montana G, Alzheimer’s Disease Neuroimaging Initiative. Identification of gene pathways implicated in Alzheimer’s disease using longitudinal imaging phenotypes with sparse regression. Neuroimage. 2012;63(3):1681–1694. doi: 10.1016/j.neuroimage.2012.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sim K, Chan WY, Woon PS, Low HQ, Lim L, Yang GL, et al. ARVCF genetic influences on neurocognitive and neuroanatomical intermediate phenotypes in Chinese patients with schizophrenia. J Clin Psychiatry. 2012;73(3):320–326. doi: 10.4088/JCP.10m06491. [DOI] [PubMed] [Google Scholar]
- Suda S, Iwata K, Shimmura C, Kameno Y, Anitha A, Thanseem I, et al. 2011Decreased expression of axon-guidance receptors in the anterior cingulate cortex in autism. Mol Autism 2114; 10.1186/2040-2392-2-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suter M, Ma J, Harris A, Patterson L, Brown KA, Shope C, et al. Maternal tobacco use modestly alters correlated epigenome-wide placental DNA methylation and gene expression. Epigenetics. 2011;6(11):1284–1294. doi: 10.4161/epi.6.11.17819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takahashi Y, Tsuruta S, Hasegawa J, Kameyama Y, Yoshida M. Release of mercury from dental amalgam fillings in pregnant rats and distribution of mercury in maternal and fetal tissues. Toxicology. 2001;163(2–3):115–126. doi: 10.1016/s0300-483x(01)00390-0. [DOI] [PubMed] [Google Scholar]
- Tian LP, Zhang S, Zhang YJ, Ding JQ, Chen SD. Lmx1b can promote the differentiation of embryonic stem cells to dopaminergic neurons associated with Parkinson’s disease. Biotechnol Lett. 2012;34(7):1167–1174. doi: 10.1007/s10529-012-0888-5. [DOI] [PubMed] [Google Scholar]
- Westfall PH, Young SS. New York: John Wiley & Sons Inc; 1993. Resampling-Based Multiple Testing: Examples and Methods for p-Value Adjustment. [Google Scholar]
- Wickre JB, Folt CL, Sturup S, Karagas MR. Environmental exposure and fingernail analysis of arsenic and mercury in children and adults in a Nicaraguan gold mining community. Arch Environ Health. 2004;59(8):400–409. doi: 10.3200/AEOH.59.8.400-409. [DOI] [PubMed] [Google Scholar]
- Wilhelm-Benartzi CS, Houseman EA, Maccani MA, Poage GM, Koestler DC, Langevin SM, et al. 2012In utero exposures, infant growth, and DNA methylation of repetitive elements and developmentally related genes in human placenta. Environ Health Perspect 120296–302.; 10.1289/ehp.1103927 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xun P, Liu K, Morris JS, Jordan JM, He K. Distributions and determinants of mercury concentrations in toenails among American young adults: the CARDIA Trace Element Study. Environ Sci Pollut Res Int. 2013;20(3):1423–1430. doi: 10.1007/s11356-012-1126-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang J, Jiang Z, Wang Y, Qureshi IA, Wu XD. Maternal-fetal transfer of metallic mercury via the placenta and milk. Ann Clin Lab Sci. 1997;27(2):135–141. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


