Abstract
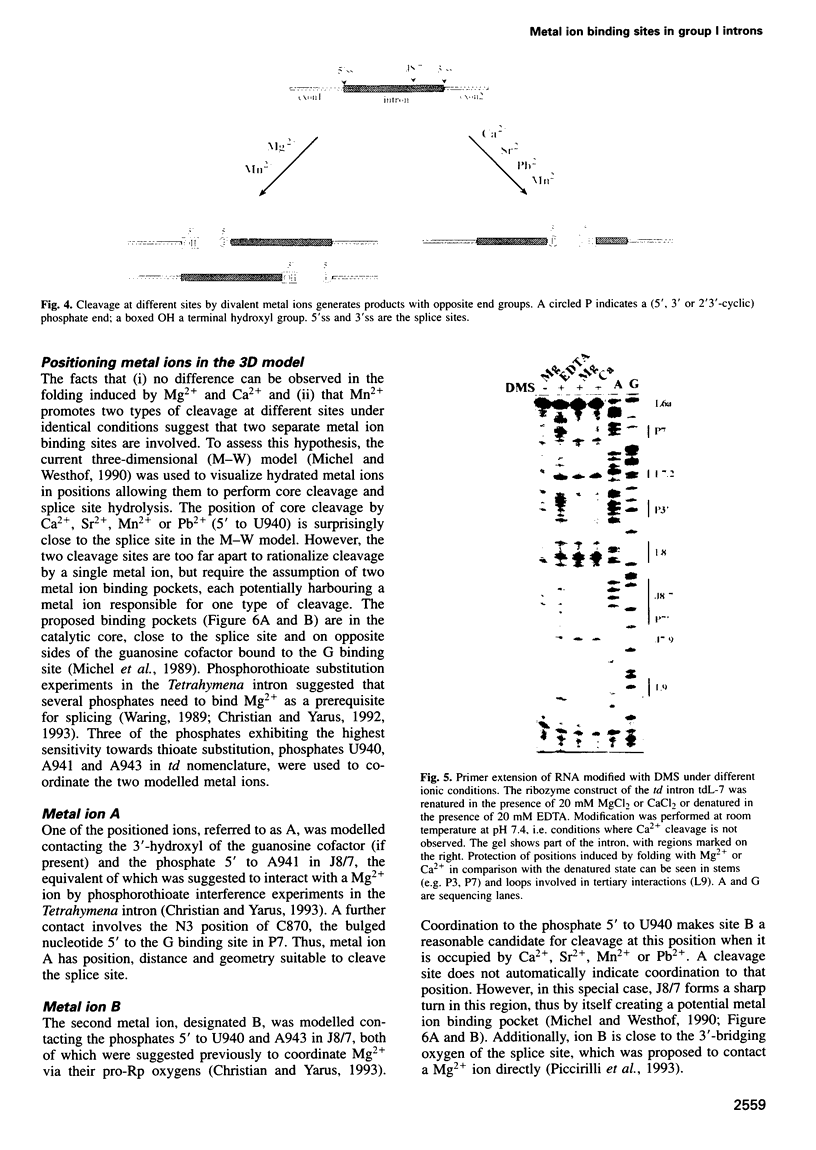
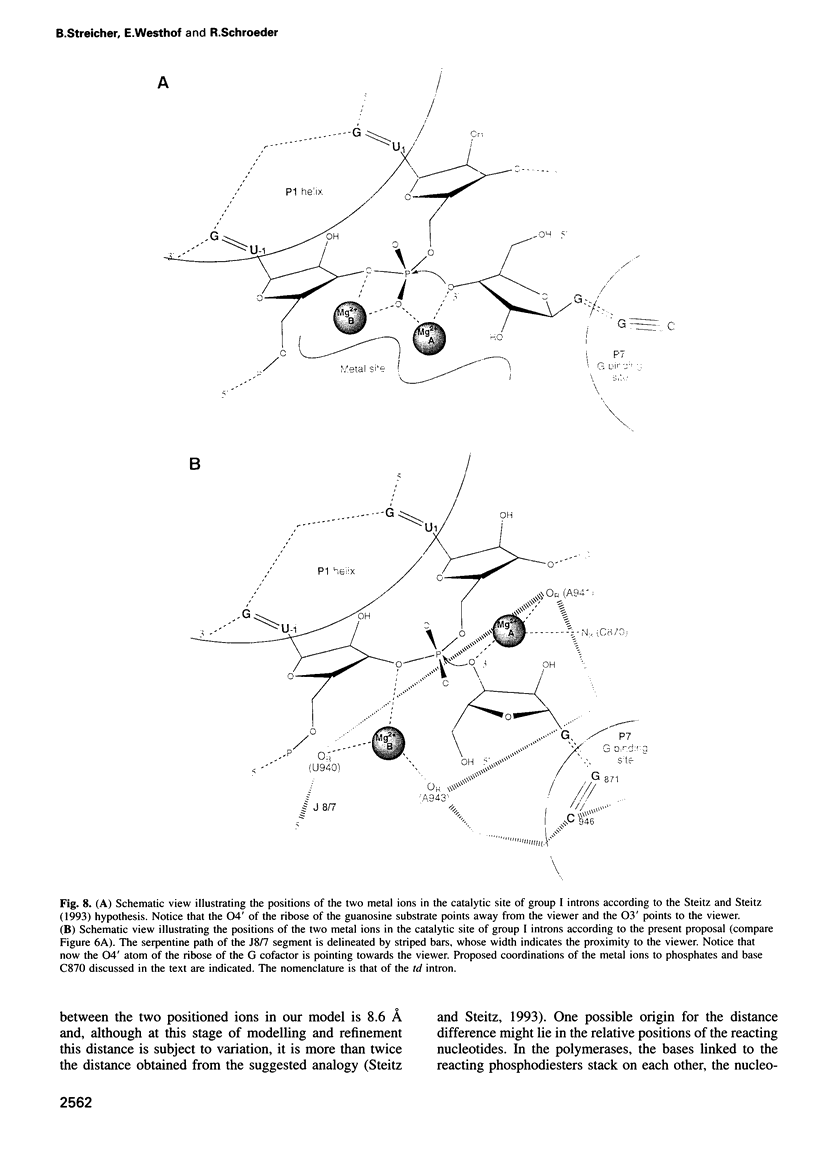
Several divalent metal ions (Ca2+, Sr2+ and Pb2+) do not promote splicing, but instead induce cleavage at a single site in the conserved group I intron core in the absence of the guanosine cofactor at elevated pH, generating products with 5'-OH and 3'-phosphate ends. The reaction is competed by Mg2+, which does not cleave at this position, but hydrolyses the splice sites producing 3'-OH and 5'-phosphate ends. Mn2+ promotes both core cleavage and splice site hydrolysis under identical conditions, suggesting that two different metal atoms are involved, each responsible for one type of cleavage, and with different chemical and geometric requirements. Based on the core cleavage position and on the previously proposed coordination sites for Mg2+, we propose a structural location for two metal ions surrounding the splice site in the Michel-Westhof three-dimensional model of the group I intron core. The proposed location was strengthened by a first mutational analysis which supported the suggested interaction between one of the metal ions and the bulged residue in P7.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Beese L. S., Steitz T. A. Structural basis for the 3'-5' exonuclease activity of Escherichia coli DNA polymerase I: a two metal ion mechanism. EMBO J. 1991 Jan;10(1):25–33. doi: 10.1002/j.1460-2075.1991.tb07917.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behlen L. S., Sampson J. R., DiRenzo A. B., Uhlenbeck O. C. Lead-catalyzed cleavage of yeast tRNAPhe mutants. Biochemistry. 1990 Mar 13;29(10):2515–2523. doi: 10.1021/bi00462a013. [DOI] [PubMed] [Google Scholar]
- Brown R. S., Dewan J. C., Klug A. Crystallographic and biochemical investigation of the lead(II)-catalyzed hydrolysis of yeast phenylalanine tRNA. Biochemistry. 1985 Aug 27;24(18):4785–4801. doi: 10.1021/bi00339a012. [DOI] [PubMed] [Google Scholar]
- Brown R. S., Hingerty B. E., Dewan J. C., Klug A. Pb(II)-catalysed cleavage of the sugar-phosphate backbone of yeast tRNAPhe--implications for lead toxicity and self-splicing RNA. Nature. 1983 Jun 9;303(5917):543–546. doi: 10.1038/303543a0. [DOI] [PubMed] [Google Scholar]
- Caprara M. G., Mohr G., Lambowitz A. M. A tyrosyl-tRNA synthetase protein induces tertiary folding of the group I intron catalytic core. J Mol Biol. 1996 Apr 5;257(3):512–531. doi: 10.1006/jmbi.1996.0182. [DOI] [PubMed] [Google Scholar]
- Cech T. R., Damberger S. H., Gutell R. R. Representation of the secondary and tertiary structure of group I introns. Nat Struct Biol. 1994 May;1(5):273–280. doi: 10.1038/nsb0594-273. [DOI] [PubMed] [Google Scholar]
- Cech T. R. Self-splicing of group I introns. Annu Rev Biochem. 1990;59:543–568. doi: 10.1146/annurev.bi.59.070190.002551. [DOI] [PubMed] [Google Scholar]
- Christian E. L., Yarus M. Analysis of the role of phosphate oxygens in the group I intron from Tetrahymena. J Mol Biol. 1992 Dec 5;228(3):743–758. doi: 10.1016/0022-2836(92)90861-d. [DOI] [PubMed] [Google Scholar]
- Christian E. L., Yarus M. Metal coordination sites that contribute to structure and catalysis in the group I intron from Tetrahymena. Biochemistry. 1993 May 4;32(17):4475–4480. doi: 10.1021/bi00068a001. [DOI] [PubMed] [Google Scholar]
- Ciesiołka J., Wrzesinski J., Górnicki P., Podkowiński J., Krzyzosiak W. J. Analysis of magnesium, europium and lead binding sites in methionine initiator and elongator tRNAs by specific metal-ion-induced cleavages. Eur J Biochem. 1989 Dec 8;186(1-2):71–77. doi: 10.1111/j.1432-1033.1989.tb15179.x. [DOI] [PubMed] [Google Scholar]
- Dahm S. C., Derrick W. B., Uhlenbeck O. C. Evidence for the role of solvated metal hydroxide in the hammerhead cleavage mechanism. Biochemistry. 1993 Dec 7;32(48):13040–13045. doi: 10.1021/bi00211a013. [DOI] [PubMed] [Google Scholar]
- Freemont P. S., Friedman J. M., Beese L. S., Sanderson M. R., Steitz T. A. Cocrystal structure of an editing complex of Klenow fragment with DNA. Proc Natl Acad Sci U S A. 1988 Dec;85(23):8924–8928. doi: 10.1073/pnas.85.23.8924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grosshans C. A., Cech T. R. Metal ion requirements for sequence-specific endoribonuclease activity of the Tetrahymena ribozyme. Biochemistry. 1989 Aug 22;28(17):6888–6894. doi: 10.1021/bi00443a017. [DOI] [PubMed] [Google Scholar]
- Heuer T. S., Chandry P. S., Belfort M., Celander D. W., Cech T. R. Folding of group I introns from bacteriophage T4 involves internalization of the catalytic core. Proc Natl Acad Sci U S A. 1991 Dec 15;88(24):11105–11109. doi: 10.1073/pnas.88.24.11105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hingerty B., Brown R. S., Jack A. Further refinement of the structure of yeast tRNAPhe. J Mol Biol. 1978 Sep 25;124(3):523–534. doi: 10.1016/0022-2836(78)90185-7. [DOI] [PubMed] [Google Scholar]
- Holbrook S. R., Sussman J. L., Warrant R. W., Church G. M., Kim S. H. RNA-ligant interactions. (I) Magnesium binding sites in yeast tRNAPhe. Nucleic Acids Res. 1977 Aug;4(8):2811–2820. doi: 10.1093/nar/4.8.2811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inoue T., Sullivan F. X., Cech T. R. New reactions of the ribosomal RNA precursor of Tetrahymena and the mechanism of self-splicing. J Mol Biol. 1986 May 5;189(1):143–165. doi: 10.1016/0022-2836(86)90387-6. [DOI] [PubMed] [Google Scholar]
- Jaeger L., Westhof E., Michel F. Monitoring of the cooperative unfolding of the sunY group I intron of bacteriophage T4. The active form of the sunY ribozyme is stabilized by multiple interactions with 3' terminal intron components. J Mol Biol. 1993 Nov 20;234(2):331–346. doi: 10.1006/jmbi.1993.1590. [DOI] [PubMed] [Google Scholar]
- Lin C. W., Hanna M., Szostak J. W. Evidence that the guanosine substrate of the Tetrahymena ribozyme is bound in the anti conformation and that N7 contributes to binding. Biochemistry. 1994 Mar 8;33(9):2703–2707. doi: 10.1021/bi00175a045. [DOI] [PubMed] [Google Scholar]
- Massire C., Gaspin C., Westhof E. DRAWNA: a program for drawing schematic views of nucleic acids. J Mol Graph. 1994 Sep;12(3):201-6, 196. doi: 10.1016/0263-7855(94)80088-x. [DOI] [PubMed] [Google Scholar]
- Michel F., Hanna M., Green R., Bartel D. P., Szostak J. W. The guanosine binding site of the Tetrahymena ribozyme. Nature. 1989 Nov 23;342(6248):391–395. doi: 10.1038/342391a0. [DOI] [PubMed] [Google Scholar]
- Michel F., Westhof E. Modelling of the three-dimensional architecture of group I catalytic introns based on comparative sequence analysis. J Mol Biol. 1990 Dec 5;216(3):585–610. doi: 10.1016/0022-2836(90)90386-Z. [DOI] [PubMed] [Google Scholar]
- Piccirilli J. A., Vyle J. S., Caruthers M. H., Cech T. R. Metal ion catalysis in the Tetrahymena ribozyme reaction. Nature. 1993 Jan 7;361(6407):85–88. doi: 10.1038/361085a0. [DOI] [PubMed] [Google Scholar]
- Pley H. W., Flaherty K. M., McKay D. B. Three-dimensional structure of a hammerhead ribozyme. Nature. 1994 Nov 3;372(6501):68–74. doi: 10.1038/372068a0. [DOI] [PubMed] [Google Scholar]
- Pyle A. M. Ribozymes: a distinct class of metalloenzymes. Science. 1993 Aug 6;261(5122):709–714. doi: 10.1126/science.7688142. [DOI] [PubMed] [Google Scholar]
- Quigley G. J., Teeter M. M., Rich A. Structural analysis of spermine and magnesium ion binding to yeast phenylalanine transfer RNA. Proc Natl Acad Sci U S A. 1978 Jan;75(1):64–68. doi: 10.1073/pnas.75.1.64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubin J. R., Sundaralingam M. Lead ion binding and RNA chain hydrolysis in phenylalanine tRNA. J Biomol Struct Dyn. 1983 Dec;1(3):639–646. doi: 10.1080/07391102.1983.10507471. [DOI] [PubMed] [Google Scholar]
- Schroeder R., von Ahsen U., Belfort M. Effects of mutations of the bulged nucleotide in the conserved P7 pairing element of the phage T4 td intron on ribozyme function. Biochemistry. 1991 Apr 2;30(13):3295–3303. doi: 10.1021/bi00227a018. [DOI] [PubMed] [Google Scholar]
- Scott W. G., Finch J. T., Klug A. The crystal structure of an all-RNA hammerhead ribozyme: a proposed mechanism for RNA catalytic cleavage. Cell. 1995 Jun 30;81(7):991–1002. doi: 10.1016/s0092-8674(05)80004-2. [DOI] [PubMed] [Google Scholar]
- Steitz T. A., Smerdon S. J., Jäger J., Joyce C. M. A unified polymerase mechanism for nonhomologous DNA and RNA polymerases. Science. 1994 Dec 23;266(5193):2022–2025. doi: 10.1126/science.7528445. [DOI] [PubMed] [Google Scholar]
- Steitz T. A., Steitz J. A. A general two-metal-ion mechanism for catalytic RNA. Proc Natl Acad Sci U S A. 1993 Jul 15;90(14):6498–6502. doi: 10.1073/pnas.90.14.6498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Streicher B., von Ahsen U., Schroeder R. Lead cleavage sites in the core structure of group I intron-RNA. Nucleic Acids Res. 1993 Jan 25;21(2):311–317. doi: 10.1093/nar/21.2.311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waring R. B. Identification of phosphate groups important to self-splicing of the Tetrahymena rRNA intron as determined by phosphorothioate substitution. Nucleic Acids Res. 1989 Dec 25;17(24):10281–10293. doi: 10.1093/nar/17.24.10281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Westhof E., Sundaralingam M. Restrained refinement of the monoclinic form of yeast phenylalanine transfer RNA. Temperature factors and dynamics, coordinated waters, and base-pair propeller twist angles. Biochemistry. 1986 Aug 26;25(17):4868–4878. doi: 10.1021/bi00365a022. [DOI] [PubMed] [Google Scholar]