Summary
Spindle assembly requires the coordinated action of multiple cellular structures to nucleate and organize microtubules in a precise spatiotemporal manner. Among them the contributions of centrosomes, chromosomes and microtubules have been well studied, yet the involvement of membrane-bound organelles remains largely elusive. Here we provide mechanistic evidence for a membrane-based, Golgi-derived microtubule assembly pathway in mitosis. Upon mitotic entry, the Golgi matrix protein GM130 interacts with importin α via a classical nuclear localization signal that recruits importin α to the Golgi membranes. Sequestration of importin α by GM130 liberates the spindle assembly factor TPX2, which activates Aurora-A kinase and stimulates local microtubule nucleation. Upon filament assembly, nascent microtubules are further captured by GM130, thus linking Golgi membranes to the spindle. Our results reveal an active role for the Golgi in regulating spindle formation to ensure faithful organelle inheritance.
Graphical Abstract
Introduction
Successful cell division requires concerted efforts to assemble a microtubule-based spindle that partitions genomic information and intracellular contents. In animal cells, spindle assembly is predominately orchestrated by centrosomes that nucleate and organize microtubules into a bipolar spindle. However, spindles also form when centrosomes are physically ablated from cultured cells (Khodjakov et al., 2000) or genetically depleted in developing embryos (Basto et al., 2006; Bazzi and Anderson, 2014), indicating that non-centrosomal components contribute to microtubule generation and spindle morphogenesis. For example, microtubules are induced at the chromosomes in an action that requires the small GTPase Ran (Carazo-Salas et al., 1999; Kalab et al., 1999; Ohba et al., 1999; Wilde and Zheng, 1999). GTP-loaded Ran is generated in the vicinity of chromatin where its guanine nucleotide exchange factor RCC1 is localized. RanGTP binding dissolves the complex of cargo and importins (Wiese et al., 2001). As a consequence, a set of nuclear localization signal (NLS)-bearing cargos, collectively termed spindle assembly factors (SAFs), are released to promote microtubule nucleation, stabilization and spindle organization (Kalab and Heald, 2008). In addition, microtubules also nucleate and amplify themselves throughout the spindle (Goshima et al., 2008; Petry et al., 2013), further suggesting multiple microtubule origins in dividing cells.
Although different cellular structures have been described to facilitate spindle assembly, the contribution of membrane organelles is poorly understood (Zheng, 2010; Meunier and Vernos, 2012). Among them, the Golgi apparatus has been shown to function as a microtubule-organizing center that builds up an acentrosomal microtubule array in both proliferating (Chabin-Brion et al., 2001; Efimov et al., 2007) and differentiated cells (Ori-McKenney et al., 2012; Oddoux et al., 2013). In support, several components of the microtubule-nucleating complex such as γ-tubulin have been localized to Golgi membranes (Chabin-Brion et al., 2001; Rivero et al., 2009). During cell division, the single mammalian Golgi undergoes extensive vesiculation in preparation for its partitioning (Wei and Seemann, 2009a). Despite the disruption of its characteristic ribbon structure, a subset of Golgi membranes cluster around the spindle poles and reside in close proximity to microtubules (Jokitalo et al., 2001; Shima et al., 1998), suggesting an intimate link between nascent spindle microtubules and mitotic Golgi membranes. Furthermore, as the spindle delivers Golgi ribbon-linking factors into progeny (Wei and Seemann, 2009b), it remains elusive how the Golgi establishes the connection to the spindle upon disassembly of the interphase microtubule network.
Mitotic Golgi disassembly is primarily driven by the master mitotic kinase Cdk1 that directly phosphorylates the Golgi matrix protein GM130 at S25 (Lowe et al., 1998). Phosphorylation of GM130 inhibits the binding of the vesicle docking protein p115, which prevents tethering and fusion of vesicles with Golgi cisternae (Levine et al., 1996; Nakamura et al., 1997; Lowe et al., 1998, 2000). As vesicles continue to bud, the Golgi is converted into clusters of vesicles and tubules. Such disassembly as well as the displacement of p115 from mitotic GM130 not only physically exposes GM130 that is mostly hidden at the central part of cisternae (Diao et al., 2003), it may also biochemically render GM130 accessible to mitotic regulators, thus licensing its participation in spindle assembly and/or other mitotic events. In agreement, depletion of GM130 by morpholino in mouse oocytes compromises meiotic spindle formation and asymmetric cell division (Zhang et al., 2011), although the mechanism awaits elucidation. Likewise, down-regulation of GM130 by RNAi in mammalian culture cells causes abnormal spindles, which was attributed as a secondary carryover effect to its interphase function in centrosome biogenesis (Kodani and Sütterlin, 2008).
To unravel the complex interplay between the Golgi and microtubules in mitosis, we set out to identify the mechanisms that underpin GM130 involvement in spindle assembly. Here, we show that the N-terminal domain of GM130 contains a previously unknown, high-affinity classical NLS (cNLS) and is essential for proper spindle assembly and cell division. Via this motif GM130 binds and sequesters importin α to the Golgi membranes, which locally activates the spindle assembly factor TPX2 that stimulates microtubule nucleation and its downstream effector Aurora-A. GM130 then further captures newly assembled microtubules and links mitotic Golgi membranes to the spindle. In this way, the Golgi actively regulates spindle assembly in a precise spatiotemporal manner to ensure its correct inheritance.
Results
Acute perturbation of GM130 in mitosis impairs spindle assembly and cell division
To unambiguously dissect the mitotic function of GM130 from its interphase roles in secretion (Seemann et al., 2000), centrosome biogenesis (Kodani and Sütterlin, 2008) and signaling (Preisinger et al., 2004; Baschieri et al., 2014), we used microinjection to acutely perturb GM130 in mitosis without affecting its interphase functions and analyzed the immediate effects. GM130 is a peripheral membrane protein tightly bound to the cytoplasmic face of the Golgi. Structurally, it contains extensive coiled-coil regions in the middle, a Golgi-targeting domain at the C-terminus, and a positively charged p115-interacting motif at the N-terminus (residues 1-74, N74) (Nakamura et al., 1997). To evaluate the physiological significance of the mitotically regulated N-terminal domain, we microinjected affinity-purified antibodies against N74 or control IgG into normal rat kidney (NRK) cells in pro/prometaphase, and then assessed division by the co-injected fluorescent marker (Figures 1A and 1B). While control-injected cells divided normally, anti-N74 antibodies caused multiple defects including mitotic arrest, multipolar division, multiple nuclei and micronuclei (Figure 1A). Only 38 ± 5% of the anti-N74-injected cells underwent bipolar division compared to the IgG control (Figures 1B and S1). Similar mitotic defects were also observed upon microinjection of N74-GST protein followed by phase-contrast time-lapse microscopy (Figures 1C and 1D). N74-GST, but not T7-GST or GST-SV40 NLS controls, induced mitotic defects ranging from multipolar cell division to prolonged mitotic arrest that, in some cases, led to mitotic catastrophe (Figure 1C). Only 44 ± 5% of the N74-GST-injected cells divided in a bipolar fashion compared to the T7-GST or GST-SV40 NLS controls (Figure 1D).
Figure 1.
Acute Perturbation of GM130 in Mitosis Impairs Spindle Assembly and Cell Division
(A) Microinjection of antibodies against the N-terminal domain of GM130 (residues 1-74, N74) interferes with bipolar division. NRK cells in pro/prometaphase were injected with 9 mg/ml affinity-purified anti-N74 or control IgG together with fluorescent dextran as an injection marker. Cell division was then assessed by fluorescence microscopy. (B) Anti-N74 injection decreased bipolar division to 38% compared to control IgG. 373 (ctl IgG) and 311 (anti-N74) cells from 4 experiments were analyzed, p<0.0001. (C) Microinjection of N74 protein interferes with bipolar division. Recombinant T7-GST, N74-GST or GST-SV40 NLS was injected as in (A). Mitotic progression was then monitored by phase-contrast time-lapse microscopy. T7 and SV40-injected cells progressed through mitosis and divided normally. N74-injected cells frequently stalled in mitosis and died after a prolonged arrest. (D) N74 injection decreased bipolar division to 44%, while T7 and SV40 had little effect. 83 (T7), 81 (N74), and 138 (SV40) cells from 3-4 experiments were analyzed, p<0.005. (E) Microinjection of N74 induces spindle assembly defects. Injected cells were stained for DNA and α-tubulin (MT). N74-injected cells showed various defects in chromosome alignment and spindle assembly. Notably, category II phenocopied RanQ69L-injection that induced ectopic asters. (F) Percentage of mitotic phenotypes in injected cells. 75 (T7), 201 (N74), and 190 (SV40) cells from 3-6 experiments were analyzed. Bars, 25 μm. Error bars represent SEM. See also Figure S1.
To determine whether the division defects induced by GM130 imbalance were ascribed to deficits in spindle assembly, we examined the chromosome configuration and spindle morphology by immunofluorescence microscopy (Figures 1E and 1F). 87% of T7-GST and 81% of GST-SV40 NLS-injected cells completed mitosis and entered the next interphase, as determined by reformed nuclei and radial microtubule arrays. By contrast, N74-GST-injections caused profound defects in chromosome alignment and spindle assembly, which were classified into five categories. Most cells were arrested with partially congressed chromosomes and a bipolar-like spindle (categories I, 28% and II, 10%). Strikingly, we detected ectopic asters (category II, arrowhead), which phenocopied spindle defects induced by microinjection of active RanQ69L (Kalab et al., 2006). 11% of the injected cells assembled multipolar spindles with more robust pole-like structures (category III). Some cells divided in a bipolar fashion (categories IV, 19% and V, 32%), although frequently with chromosomal bridges (category IV, arrow). Taken together, interfering with GM130 in mitosis impairs spindle formation and cell division.
The N-terminal domain of GM130 harbors a classical NLS that binds importin α
To gain insights into the molecular mechanisms underlying these mitotic defects, we inspected the sequence of the injected N74 protein. The N-terminal domain of GM130 is highly conserved across mammals, suggesting it preserves a critical activity in higher animals (Figure S2). Notably, N74 contains two K/R clusters that flank the Cdk1 phosphorylation site at S25 (Figure 2A, K/R highlighted in blue and S in red). The K/R-rich motifs are reminiscent of the cNLS that has a pivotal role in Ran-mediated signaling. Since mitotic injection of N74 caused ectopic asters, a distinct spindle defect also induced by RanQ69L protein injection (Kalab et al., 2006), we explored whether GM130 carries a cNLS. First, we evaluated its ability to directly bind importin α, the adaptor that recognizes the cNLS in cargo and bridges it to importin β (Figure 2B). T7-GST did not pull down importin α (imp α ΔIBB, NLS-recognition domain), and neither did the non-classical M9 NLS that binds transportin without an adaptor (Pollard et al., 1996). By contrast, N74 pulled down a comparable amount of importin α as the well-characterized cNLS of the SV40 large T-antigen. The direct binding was abolished when the two K/R clusters in N74 were mutated to alanine (AA, NLS mutant), but was not affected when S25 was changed to aspartic acid (S25D), a previously characterized Cdk1 phospho-mimicking mutation that prevents p115 binding (Lowe et al., 1998) (Figure 2C).
Figure 2.
GM130 Harbors a bona fide Classical NLS
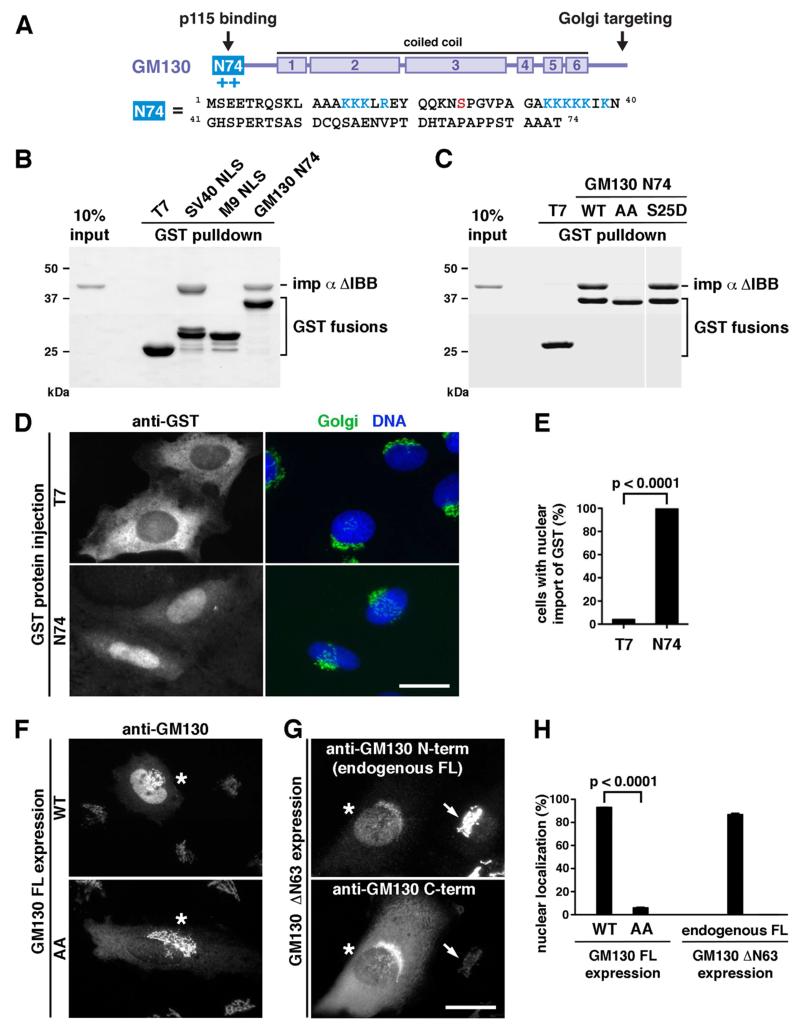
(A) Domain structure. GM130 is comprised of six coiled-coil regions in the middle, a Golgi-targeting domain at the C-terminus and a p115-interacting motif at the N-terminus (N74). N74 contains two K/R-rich clusters (blue) and a Cdk1 phosphorylation site S25 (red). (B) GM130 N74 directly interacts with importin α in vitro. Upon incubation with the NLS-interacting domain of importin α (ΔIBB), GST fusion proteins were pulled down and analyzed by SDS-PAGE and Coomassie blue staining. N74 and the classical SV40 NLS bound importin α, but not T7 or the non-classical M9 NLS. (C) Pulldowns were performed as in (B) with the N74 NLS mutant AA (K/R clusters changed to alanine) or the Cdk1 phospho-mimicking mutant S25D. Importin α binding required the K/R residues and was not affected by S25 phospho-mimicking mutation. The blank space indicates removal of irrelevant lanes. (D) N74 is recognized and translocated into the nucleus by the endogenous nuclear import machinery in cells. NRK cells were microinjected into the cytoplasm with T7-GST or N74-GST and then stained for DNA, GST, and the Golgi (with an antibody against the C-terminal domain of GM130). (E) N74-GST was imported into the nucleus in 99.7% of the injected cells. 379 (T7) and 358 (N74) cells from 3 experiments were analyzed, p<0.0001. (F) Over-expressed full-length GM130 targets to the Golgi and enters the nucleus. 93% of GM130 FL WT- and 6% of GM130 FL AA-expressing cells (marked by asterisks) showed nuclear localization of GM130. (G) Endogenous GM130 translocates into the nucleus once mobilized. Upon expression of a Golgi-targeting, N-terminal truncation mutant of GM130 (ΔN63), endogenous GM130 was displaced from the Golgi and entered the nucleus in 87% of ΔN63-expressing cells, as detected with an antibody against the N-terminal domain. Expressing and non-expressing cells are marked by asterisks and arrows, respectively. (H) Quantitation of (F) and (G). 170 (FL WT), 167 (FL AA), and 358 (ΔN63) cells from 3-4 experiments were analyzed, p<0.0001. Bars, 25 μm. Error bars represent SEM. See also Figure S2.
We then tested whether N74 can be recognized by the endogenous nuclear import machinery in cells. We microinjected T7-GST or N74-GST into the cytoplasm of NRK cells and scored its nuclear import. In line with our in vitro data, N74-GST but not T7-GST translocated into the nucleus (Figures 2D and 2E). Similarly, ectopically overexpressed full-length GM130 was imported into the nucleus in addition to being targeted to the Golgi. Such import requires a cNLS and is mediated by importin α, as the GM130 NLS mutant AA failed to translocate into the nucleus (Figures 2F and 2H). Moreover, upon overexpression of a Golgi-targeting, N-terminal truncation mutant that lacks the cNLS (GM130 ΔN63), endogenous GM130 was partially displaced from the Golgi and entered the nucleus (Figures 2G and 2H). This result demonstrates that endogenous GM130 interacts with importin α in the same manner as its N74 counterpart and gets imported into the nucleus once it is released from the Golgi membranes. In conclusion, the N-terminal domain of GM130 harbors a functional cNLS that binds importin α in vitro and in cells.
Importin α is recruited to Golgi membranes in mitosis by GM130
By interacting with GM130, importin α should be targeted to Golgi membranes. To test this, we fractionated a post-nuclear supernatant (PNS) from exponentially growing HeLa cells into cytosol (C) and membranes (M) (Figure 3A). Consistent with a previous report where importin α was implicated in nuclear envelope assembly in Xenopus extracts (Hachet et al., 2004), we found a portion of importin α, but not importin β, on membranes. We then compared the membrane association between interphase and mitosis (Figure 3B). GM130 was mitotically phosphorylated as indicated by its slower mobility on SDS-PAGE (Nakamura et al., 1997) with no significant change in abundance. Similarly, the levels of other organelle markers including giantin (Golgi), calnexin (ER), PDI (ER) and TIMM50 (mitochondria), as well as membrane-associated γ-tubulin remained unaltered. By contrast, dynein was displaced from mitotic membranes as a result of Cdk1 phosphorylation (Niclas et al., 1996). Most notably, membrane-bound importin α increased dramatically during mitosis (4-5 fold), implying a specialized role for this pool.
Figure 3.
Importin α Is Recruited to Mitotic Golgi Membranes by GM130
(A) Importin α is partially membrane-bound. A post-nuclear supernatant (PNS) from unsynchronized HeLa cells was fractionated into cytosol (C) and membranes (M). Equal-volume fractions were analyzed by Western blotting. While importin β was restricted to the cytosol, a portion of importin α bound to membranes. (B) Membrane-bound importin α is elevated in mitosis. Equal protein amounts of membrane fractions from interphase and mitotic HeLa cells were analyzed by Western blotting. In contrast to the unaltered levels of membrane markers, including GM130 and giantin (Golgi), calnexin and PDI (ER), TIMM50 (mitochondria) and membrane-bound γ-tubulin, membrane association of importin α increased in mitosis. (C) Importin α co-sediments with mitotic Golgi membranes. A post-chromosomal supernatant (PCS) from mitotic HeLa cells was centrifuged through a glycerol gradient. Membranes from each fraction were re-isolated and analyzed by Western blotting. Importin α co-migrated with the Golgi markers GM130 and GRASP65 in the top fractions, while ER (calnexin) and mitochondria (TIMM50) markers were found in the bottom. (D) Pre-incubation of the PCS with N74 WT-GST or anti-N74, but not T7-GST, N74 AA-GST or pre-immune serum, selectively removed the pool of Golgi-bound importin α in the top fractions. Input: 2-4% PCS. n=3, **p<0.01. (E) Importin α forms a complex with GM130 specifically in mitosis. Endogenous GM130 from interphase and mitotic HeLa lysates was immunoprecipitated and analyzed by Western blotting. Endogenous importin α co-precipitated with GM130 from mitotic but not interphase lysates. Error bars represent SEM. See also Figure S3.
We next analyzed whether importin α is localized to Golgi membranes in mitosis. A post-chromosomal supernatant (PCS) from mitotic HeLa cells was centrifuged through a glycerol velocity gradient to separate smaller mitotic Golgi clusters from larger membranes (Figure 3C) (Jesch and Linstedt, 1998). While ER and mitochondria were found near the bottom of the gradient, importin α co-fractionated with the Golgi markers GM130 and GRASP65 in the upper half of the gradient, which lends strong support for mitotic recruitment of endogenous importin α to the Golgi. Incubation of the PCS prior to gradient centrifugation with N74 WT-GST, but not T7-GST or N74 AA-GST, selectively reduced Golgi-bound importin α in the top fractions (Figure 3D), indicating that importin α-Golgi association relies on the NLS. This is in contrast to importin α that binds nuclear envelope membranes in an NLS-independent manner (Hachet et al., 2004). To further corroborate that GM130 targets importin α to the mitotic Golgi, we incubated the PCS prior to gradient centrifugation with the anti-N74 antibodies that inhibited mitotic progression in cells (Figure 1A). Indeed, anti-N74 but not pre-immune serum dissociated importin α from the membranes in the top fractions (Figure 3D). Consistently, anti-N74 also displaced importin α from the membranes of the pooled fractions 2 and 3 following gradient centrifugation (Figure S3A). Moreover, importin α co-immunoprecipitated with endogenous GM130 but not golgin-84 or control IgG from mitotic but not interphase lysates (Figures 3E and S3B). Altogether, our results show that GM130 recruits importin α to Golgi membranes during mitosis via its NLS-containing N74 domain.
GM130 competes importin α off the spindle assembly factor TPX2 via its high affinity NLS
In mitosis importin α prevents microtubule formation by binding and inhibiting NLS-bearing SAFs including TPX2, a microtubule nucleator essential for spindle assembly (Gruss et al., 2001; Schatz et al., 2003). Mitotic binding of GM130 to importin α may therefore hinder its ability to silence TPX2, which provides an explanation for N74-triggered aster formation in mitotic cells (Figure 1E, II). Mechanistically, GM130 can initiate microtubule assembly around the Golgi by competing importin α off TPX2. To evaluate this, we first identified the NLS of mouse TPX2 by sequence alignment with the previously mapped NLS-containing domain of frog TPX2 (Giesecke and Stewart, 2010; Schatz et al., 2003) (Figure 4A) and then determined if it binds importin α ΔIBB (Figure 4B). Indeed, importin α was pulled down by the GST-tagged mouse TPX2 NLS (residues 302-322), but not by its NLS mutant with two key residues changed to alanine (K315A K316A, mut, marked in bold in Figure 4A). We then measured the binding affinity of importin α with TPX2 NLS, N74 WT, and N74 S25D using isothermal titration calorimetry (ITC) (Figure 4C). All pairs bound with equimolar stoichiometry. N74 WT and N74 S25D bound importin α equally well but with approximately 9-fold higher affinity than TPX2.
Figure 4.
GM130 Competes Importin α off TPX2 through its High Affinity NLS
(A) Mapping of the NLS in mouse TPX2 (NP_082385.3) by protein sequence alignment with frog TPX2 (AAH68637). NLS-containing peptides (residues 270-289 in frog; 302-322 in mouse) are grey-shaded. The key NLS residues (K284, R285 in frog; K315, K316 in mouse) are highlighted in bold. (B) TPX2 binding to importin α requires a functional NLS. T7-GST, GST-TPX2 NLS wildtype (wt), and GST-TPX2 NLS mutant (K315A K316A, mut) were incubated with importin α ΔIBB, pulled down and analyzed by SDS-PAGE and Coomassie blue staining. Importin α only bound the wt NLS of TPX2 but not the NLS mutant. (C) N74 WT and the phospho-mimicking S25D mutant bind importin α with a nine-fold higher affinity than TPX2. The dissociation constants (Kd, represented as mean ± SEM) were measured by ITC. n≥3. (D) GST-pulldown competition assay. N74 WT but not AA competes importin α off the preformed importin α-TPX2 complex. (E) N74 does not directly bind TPX2-NLS. T7-GST and GST-TPX2 NLS were incubated with N74 WT-His or importin α ΔIBB, pulled down and analyzed by SDS-PAGE and Coomassie blue staining. GST-TPX2 NLS bound importin α ΔIBB but not N74-His.
We next examined whether N74 directly competes importin α off the preformed importin α-TPX2 complex (Figure 4D). Indeed, increasing N74 concentrations significantly reduced the amount of importin α associated with TPX2. The dissociation requires the importin α binding site in N74, because no competition was observed with the N74 NLS mutant (N74 AA-His). To rule out the possibility that N74 might directly bind TPX2 and destabilizes the preformed importin α-TPX2 complex, we tested for their direct interaction and found that GST-TPX2 NLS only interacts with importin α but not N74-His (Figure 4E). Collectively, these data demonstrate that N74 directly competes importin α off TPX2 via its high affinity NLS.
GM130 promotes Aurora-A activation and mitotic aster formation in a TPX2-dependent manner
By competing importin α off TPX2, N74 should promote microtubule nucleation in mitotic extracts, as it does in mitotic cells (Figure 1E). We thus tested whether N74 induces aster formation in mitotic HeLa extracts (Figures 5A, 5B and 5D) (Chang et al., 2009). In sharp contrast to N74 AA and SV40 NLS controls, both beads-bound and soluble N74 WT stimulated microtubule generation and aster assembly. To determine whether full-length GM130 also has similar activity in its native membrane environment, we replaced recombinant N74 with mitotic Golgi membranes generated by phosphorylation of rat liver Golgi with mitotic cytosol (Nakamura et al., 1997). The mitotic Golgi membranes showed strong microtubule nucleation and aster formation activity (Figures 5C, 5E and S4). This activity was abolished if the Cdk1 inhibitor Purvalanol A was added to the mitotic cytosol during pre-treatment of the Golgi, suggesting that Cdk1-mediated phosphorylation primes the Golgi for the subsequent reaction. On the other hand, addition of the anti-N74 antibodies that inhibited mitotic progression in cells (Figure 1A) blocked Golgi-stimulated aster assembly, thereby corroborating the requirement for the N74 domain. Asters also failed to form with N74 AA, or N74 WT co-incubated with importin α ΔIBB or full-length importins α/ß, indicative of a strict requirement for the NLS motif (Figure 5D). In addition, removal of mitotic extracts abolished aster assembly, suggesting that the extracts contain the factor(s) indispensable for the process (Figure 5D). We thus immunodepleted TPX2 from the extracts to evaluate its direct involvement (Figures 5F and 5G). TPX2-depleted extracts failed to assemble asters in the presence of N74 WT, suggesting that N74 acts upstream of TPX2 to induce mitotic microtubules.
Figure 5.
GM130 Induces Aurora-A Activation and Triggers Aster Formation by Liberating TPX2
(A and B) N74 WT, but not N74 AA or SV40 NLS, stimulates aster formation in mitotic extracts. GST fusion proteins immobilized on beads (A) or as soluble proteins (B) were incubated with mitotic HeLa extracts and fluorescent tubulin. Microtubule asters and autofluorescent beads were detected in the same channel by fluorescence microscopy. The inset in (B) shows a magnified representative aster. (C and E) Aster formation is triggered by mitotic Golgi membranes and blocked by antibodies against N74. The activity was also abolished if Cdk1 was inhibited during pre-treatment of the Golgi. (D) N74-stimulated aster assembly was blocked by addition of importin α ΔIBB or full-length importins α/ß. n=3, ****p<0.0001. (F) N74-stimulated aster formation requires TPX2. N74 failed to induce asters in TPX2-immunodepleted (ID) mitotic extracts. n=3, ****p<0.0001. (G) Immunodepletion of TPX2. Mitotic HeLa extracts were incubated for three consecutive rounds with protein A-conjugated beads pre-coupled to control IgG or anti-TPX2. Extracts and beads were analyzed by Western blotting with antibodies against α-tubulin and TPX2. After the third round, TPX2 was depleted from the mitotic extracts. (H) N74 activates Aurora-A, a downstream target of TPX2. Upon incubation with mitotic cytosol and microtubules, N74-WT but not AA induced Aurora-A T288 phosphorylation (left panel). Addition of importins α/β (middle panels) or antibodies against N74 (Ab, right panel) blocked Aurora-A activation. Bars, 20 μm. Error bars represent SEM. See also Figure S4.
Given that N74 liberates pre-occupied TPX2 from importin α in vitro and stimulates microtubule generation in mitotic extracts, we further investigated whether the competition triggers TPX2 downstream signaling. Upon release from importin α, TPX2 activates Aurora-A kinase in a microtubule-dependent manner, which itself is also an essential regulator for spindle assembly (Tsai et al., 2003; Sardon et al., 2008). TPX2 locks Aurora-A in the active state by inducing its conformational change that renders the key residue T288 inaccessible to the phosphatase (Bayliss et al., 2003). We therefore tested whether GM130 induces Aurora-A T288 phosphorylation by liberating TPX2 from importin α inhibition in mitotic cytosol (Figure 5H). Upon incubation with mitotic HeLa cytosol and microtubules, N74 WT, but not T7 or N74 AA, induced Aurora-A T288 phosphorylation (Figure 5H, left panel). The activation requires N74 and its NLS motif, as phosphorylation was impaired by adding importins α/ß or anti-N74 antibodies (Figure 5H, middle and right panels).
GM130 associates with the mitotic spindle by direct binding to microtubules
Microtubule nucleation and assembly around mitotic Golgi membranes is most relevant to its own inheritance since the spindle delivers essential Golgi factors into the daughter cells (Wei and Seemann, 2009b). To establish this connection upon mitotic entry requires coupling of nascent spindle microtubules to Golgi membranes. To our surprise, we found that GM130 per se possesses such activity. In agreement with the proteome of HeLa spindles (Sauer et al., 2005), we detected endogenous GM130 together with the spindle proteins γ-tubulin, α-tubulin and Plk1 on isolated spindles, while membrane markers of the ER (calnexin, PDI), mitochondria (TIMM50, Cox IV) and the cytosolic protein GAPDH were depleted (Figures 6A and 6B). Furthermore, because tubulin was identified by mass spectrometry as a mitotic interactor of N74 in our proteomic analysis (data not shown), we tested its direct binding to microtubules using a microtubule-copelleting assay (Figure 6C). N74 WT, the phospho-mimicking N74 S25D mutant and the yeast EB1 homolog Mal3 co-pelleted with microtubules, whereas BSA and T7-GST did not bind microtubules and remained in the supernatant. We then visualized the effect of N74 on microtubules using fluorescently-labeled filaments (Figure 6D). Individual microtubules became bundled, as shown by a significant increase in length, width and fluorescence intensity across the filaments. Interestingly, despite inhibiting microtubule nucleation and aster formation (Figure 5D), pre-binding of N74 with importin α ΔIBB did not prevent microtubule bundling (Figure 6D), suggesting that residues other than the NLS are involved.
Figure 6.
GM130 Associates with the Spindle and Directly Binds to Microtubules
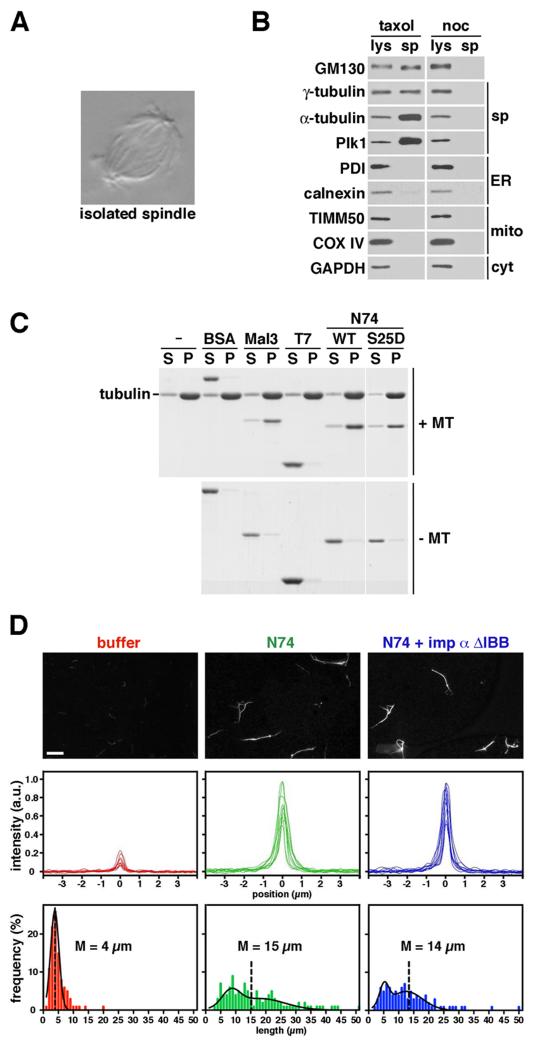
(A and B) GM130 co-purifies with spindles. Spindles isolated from mitotic HeLa cells were visualized by DIC microscopy (A) or analyzed by Western blotting (B). Whole cell lysates (lys) and isolated spindles (sp) were probed for the indicated marker proteins. Nocodazole (noc) treatment served as a negative control. (C) N74 binds directly to microtubules. Preformed microtubules were incubated with the indicated proteins and then pelleted by centrifugation. Equal volumes of the supernatant (S) and pellet (P) fractions were analyzed by SDS-PAGE and Coomassie blue staining. N74 WT, N74 S25D, and the microtubule binding protein Mal3/EB1 co-pelleted with microtubules, but not T7-GST or BSA. (D) N74 promotes microtubule bundling. Fluorescently-labeled microtubules treated with buffer, N74, or importin α ΔIBB-prebound N74 were visualized by fluorescence microscopy (top). Intensity line-scans from 10 cross-sections of microtubule structures were superimposed (middle) and length distribution was plotted (bottom, 204 structures on average per condition). M represents mean. Bar, 10 μm.
Discussion
In this study, we demonstrate that the Golgi resident protein GM130 activates the spindle assembly factor TPX2 to nucleate microtubules around the Golgi (Figures 5C and S4) and further captures them to couple mitotic membranes to the spindle (Figure 6). This function requires the N-terminal domain (N74) of GM130 that, when injected into mitotic cells or blocked by inhibitory antibodies, acutely impairs spindle assembly and bipolar division (Figure 1). We identified a cNLS in the N74 domain that binds the cargo adaptor importin α in vitro and in cells (Figure 2). This NLS motif is responsible for recruiting importin α to the mitotic Golgi membranes (Figure 3). By competing importin α, GM130 lifts TPX2 from inhibition and thus induces mitotic asters in cell extracts and intact cells (Figures 1E, 4 and 5). This activity is achieved by the high-affinity NLS of N74, which has a 9-fold lower dissociation constant than TPX2 and the majority of the cNLS (Figure 4C) (Cutress et al., 2008). In accordance, we found that microinjection of the cNLS of SV40 did not interfere with mitotic progression and spindle assembly in live mammalian cells (Figure 1), which differs from Xenopus extracts where an excess of SV40-NLS peptide is sufficient to induce aster formation in vitro (Gruss et al., 2001). As both experiments used comparable NLS concentrations, the discrepancy may be attributed to a tighter control present in vivo or to a specific regulation in mammalian cells.
In addition to the high affinity of its NLS, the local cellular concentration of GM130 could enhance its ability to liberate TPX2. The extensive coiled-coil domains of GM130 form oligomers that are immobilized onto the two-dimensional surface of the Golgi membranes (Ishida et al., 2015). In contrast to other NLS-containing cargos that diffuse throughout the cytoplasm, membrane confinement of GM130 greatly increases its effective local concentration (200-400-fold enrichment over PNS (Taylor et al., 1997)), making it a strong competitor for NLS-bearing SAFs. By binding and sequestering importin α, GM130 locally disrupts the import complex and thus liberates active SAFs. As a low level of SAF activation is sufficient to drive microtubule polymerization (Kalab et al., 2006), GM130 may thus induce microtubules around Golgi membranes. This is analogous to RanGTP, which destabilizes the import complex by binding to importin β and thus drives microtubule assembly around chromosomes. Given that both N74 and mitotic Golgi membranes are sufficient to trigger aster assembly in cell extracts, it would be interesting to determine whether in mitotic cells GM130 acts in parallel or synergistically with RanGTP to induce microtubules in the proximity of the Golgi.
Sequestration of importin α by highly concentrated and localized NLS-containing membrane proteins (acting as an “indirect activator” or “de-repressor”) may serve as a general mechanism for compartmentalized activation of SAFs in mitosis. As shown in our study, importin α was partially bound to membranes (Figure 3A), which was likely contributed by a small population of mitotic cells, as its membrane association was dramatically elevated during mitosis (Figure 3B). In addition to GM130 on the Golgi, other membrane-bound proteins may also employ a similar “sequestration-activation” mechanism, as hinted by a subset of importin α associated with light membranes devoid of the Golgi (Figure 3C, fraction 1). The NLS of these competitive activators may be temporarily masked by associated proteins or silenced via post-translational modifications, while upon specific signals become exposed or activated. In line with this, at a later stage of developing embryos importin α becomes sequestered to membranes, leading to the activation of the kinesin Kif2a that depolymerizes microtubules and thus down-scales the spindle size (Wilbur and Heald, 2013).
The specificity of GM130-triggered TPX2 activation might be partially attributable to an unusual binding mode of TPX2 to importin α (Giesecke and Stewart, 2010). While most cNLS cargos predominantly interact with the major NLS-binding pocket of importin α, TPX2 atypically binds to the minor NLS-binding groove. This differential binding preference is thought to prevent unspecific TPX2 activation by abundant cNLS cargos that are released from the nucleus upon nuclear envelope breakdown in early mitosis. Since our results show that GM130 N74 competes importin α off TPX2, it is possible that GM130 may bind to the same minor NLS-binding pocket of importin α as TPX2, or occupy both NLS-binding sites as GM130 contains two K/R clusters. Furthermore, the precise spatiotemporal control adds another layer of regulation. In interphase, the two molecules are spatially separated as GM130 is immobilized on the Golgi whereas TPX2 is sequestered in the nucleus (Heidebrecht et al., 1997). In early mitosis, Cdk1 dissociates p115 from GM130 and thus triggers mitotic Golgi disassembly (Levine et al., 1996; Nakamura et al., 1997; Shima et al., 1997; Lowe et al., 1998, 2000). Meanwhile, the elevated Cdk1 activity breaks down the nucleus, which releases TPX2 into the cytoplasm where it becomes accessible for activation by GM130.
Upon activating the microtubule nucleator TPX2, GM130 exerts its second activity toward assembled microtubules. We demonstrate that GM130 is a genuine microtubule-associated protein (MAP) that directly binds and bundles microtubules (Figures 6C and 6D). With this activity, GM130 may stabilize the initial contact with nascent mitotic microtubules that can be further organized by other MAPs and/or motors during spindle assembly. Consistently, we found GM130 on isolated spindles (Figure 6B) and Golgi-triggered microtubule asters on mitotic Golgi membranes (Figure S4), supporting the in vivo observation that mitotic Golgi clusters reside in close vicinity to spindle microtubules (Seemann et al., 2002). The association of membrane-bound organelles with the spindle is not restricted to the Golgi, as Rab11-containing recycling endosomes also associate with the spindle where they contribute to microtubule anchoring, spindle organization and orientation (Hehnly and Doxsey, 2014).
Altogether, we present a pathway for Golgi-initiated microtubule assembly in mitosis and the relevance to its inheritance (Figure 7). Upon mitotic entry Cdk1 directly phosphorylates S25 of GM130, which dissociates its interphase binding partner p115 (Levine et al., 1996; Nakamura et al., 1997; Lowe et al., 1998, 2000). The N-terminal domain of GM130 locally recruits importin α via its cNLS and liberates TPX2, which drives microtubule nucleation around mitotic Golgi clusters. Upon filament assembly, microtubules are then captured and stabilized by GM130, thus coupling Golgi membranes to the spindle. Here GM130 not only serves as a SAF activator but also as a microtubule receptor on the Golgi. These activities are tightly coordinated with correct timing and defined location in mitosis, and must be under precise control as either too little or too much of GM130 impairs division (Figure 1) (Kodani and Sütterlin, 2008; Zhang et al., 2011). In this way, the Golgi shapes the spindle to facilitate its partitioning, illustrating the dynamic interplay between two cellular machines during cell division.
Figure 7.
Model of GM130-Mediated, Golgi-Based Microtubule Assembly in Mitosis
❶Cdk1 phosphorylates GM130 upon mitotic entry; ❷ p115 dissociates from GM130; ❸ GM130 sequesters importin α to mitotic Golgi membranes; ❹ TPX2 is released to stimulate microtubule nucleation and growth in the proximity; ❺ GM130 captures and stabilizes nascent microtubules, thereby anchoring mitotic Golgi clusters to the spindle.
Experimental Procedures
Microinjection
For effects of anti-N74 on cell division, NRK cells in pro/prometaphase were injected with 1.5 or 9 mg/ml of anti-N74 or control IgG together with 2 mg/ml FITC-dextran. After 1 h at 37°C, images of live fluorescent cells were taken. For effects of N74 on cell division, NRK cells in pro/prometaphase were injected with 9 mg/ml of GST proteins and followed by phase-contrast imaging at 10-15 min intervals for 16 h. For effects of N74 on spindle assembly, NRK cells in pro/prometaphase were injected with 9 mg/ml of GST proteins together with 1 mg/ml of purified IgG. Injected cells were identified by staining the co-injected IgG. For nuclear import of N74, 0.5 mg/ml of GST proteins were injected into the cytoplasm of NRK cells. Injected cells were labeled with GST antibodies. For nuclear import of full-length GM130, 0.2 mg/ml of expression plasmids were injected into the nuclei of NRK cells, incubated for 4 h at 37°C and analyzed by immunofluorescence.
Cell Fractionation
The post-nuclear supernatant (PNS) of exponentially growing HeLa cells was fractionated by ultracentrifugation for 30 min at 100,000 g at 4°C into cytosol (C) and membranes (M). Equal-volume fractions of PNS, C and M were analyzed by Western blotting. To compare interphase and mitotic membranes, HeLa cells mitotically arrested with nocodazole for 16-18 h were collected by shake-off and the remaining cells were pooled as interphase cells. Equal protein amounts of total membranes were analyzed by Western blotting. Mitotic Golgi membranes were further separated by gradient centrifugation as described (Jesch and Linstedt, 1998). Membranes from each fraction were recovered and analyzed by Western blotting. For competition, the post-chromosomal supernatant (PCS) was pre-incubated with proteins or antibodies for 1.5-2 h at 4°C prior to gradient centrifugation. HeLa spindles were isolated as described (Sauer et al., 2005). In the control, taxol was replaced with nocodazole.
Isothermal Titration Calorimetry
Binding affinities of GST-TPX2 NLS, N74 WT-GST and N74 S25D-GST to untagged importin α ΔIBB were measured by ITC (MicroCal Omega VP-ITC) at 20°C and analyzed by MicroCal Origin 7.0. All proteins were dialyzed against ITC buffer (20 mM HEPES pH 7.4, 100 mM NaCl, 10% glycerol, 2 mM β-mercaptoethanol). Protein concentrations were determined spectrophotometrically with the following extinction coefficients (M−1cm−1): GST-TPX2 NLS 44600; GM130 N74-GST 44600; importin α ΔIBB 47815.
Aurora-A Activation and Aster Formation
For the Aurora-A activation assay, mitotic HeLa cytosol (Lowe et al., 1998) was incubated with GST proteins for 30 min at room temperature and then pre-assembled microtubules were added for additional 30 min. Microtubule-bound proteins were recovered by centrifugation for 30 min at 100,000 g at 25°C and analyzed by Western blotting.
The aster formation assay using mitotic HeLa extracts was carried out as described (Chang et al., 2009). To visualize asters, mitotic extract supplemented with 5 μM HiLyte488-labeled tubulin was incubated with 2 μM free proteins, protein-coated beads, or Golgi membranes for 20 min at room temperature and then imaged immediately. The aster number per field was obtained from 16 non-overlapping, randomly selected fields per condition and averaged from at least three independent reactions. For Golgi-stimulated aster formation, mitotic Golgi membranes were generated by incubating rat liver Golgi with mitotic HeLa cytosol and an ATP-regenerating system (Tang et al., 2010). The animal procedures were approved by the Institutional Animal Care and Use Committee of UT Southwestern. The mitotic Golgi was recollected in HK buffer with or without anti-N74 antibodies. For the Cdk1 inhibitor control, Purvalanol A was added to mitotic cytosol and the ATP-regenerating system was omitted. For detection, mitotic Golgi membranes were stained with 1 μg/ml FM 4-64 for 15 min on ice, washed and resuspended in HK buffer.
For immunodepletion of TPX2, 10 μl of mitotic extract was incubated with 10 μl protein A-Dynabeads pre-coupled to 1 μg of anti-TPX2 or control IgG for 30 min at 4°C. Immunodepletion was performed three rounds. For preparation of the protein-coated beads, 10 μl of protein A-Dynabeads were incubated with 2.5 μg anti-GST antibody for 1 h at 4°C. 0.5 μl of the anti-GST beads were further incubated with 2 μM of the indicated GST proteins for additional 1 h at 4°C.
Microtubule Binding and Bundling
Pre-assembled microtubules were incubated with the indicated proteins at room temperature for 30 min and pelleted through cushion buffer containing taxol for 40 min at 100,000 g at 25°C. The supernatants and the pellets were analyzed by SDS-PAGE and Coomassie blue staining. For microtubule bundling, HiLyte488-labeled filaments were incubated with the indicated protein for 30 min at room temperature, fixed 5 min with 1% glutaraldehyde and spotted onto glass slides. Intensity line-scans were performed using ImageJ to evaluate bundling effects.
A comprehensive description of materials and methods can be found in Extended Experimental Procedures.
Supplementary Material
Highlights.
Perturbation of GM130 in mitosis impairs spindle assembly and cell division
GM130 contains a classical NLS that recruits importin α to the Golgi
By sequestering importin α, GM130 activates TPX2 to stimulate microtubule nucleation
GM130 directly binds microtubules and associates with the mitotic spindle
Acknowledgments
We thank Rebecca Heald and Kara Helmke for insightful discussion and support during project development, Matthew Beard for GM130 constructs, Yuh Min Chook for importin constructs, Luke Rice for Mal3 protein and Beatriz Fontoura for suggestions. This work is supported by the NIH grant GM096070 to J.S.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Author Contributions
J.H.W. and J.S. conceived, designed and performed the experiments. R.M.W. performed ITC. Z.C.Z. contributed reagents. J.H.W. and J.S. wrote the manuscript.
References
- Baschieri F, Confalonieri S, Bertalot G, Di Fiore PP, Dietmaier W, Leist M, Crespo P, Macara IG, Farhan H. Spatial control of Cdc42 signalling by a GM130-RasGRF complex regulates polarity and tumorigenesis. Nat. Commun. 2014;5:4839. doi: 10.1038/ncomms5839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Basto R, Lau J, Vinogradova T, Gardiol A, Woods CG, Khodjakov A, Raff JW. Flies without centrioles. Cell. 2006;125:1375–1386. doi: 10.1016/j.cell.2006.05.025. [DOI] [PubMed] [Google Scholar]
- Bayliss R, Sardon T, Vernos I, Conti E. Structural basis of Aurora-A activation by TPX2 at the mitotic spindle. Mol. Cell. 2003;12:851–862. doi: 10.1016/s1097-2765(03)00392-7. [DOI] [PubMed] [Google Scholar]
- Bazzi H, Anderson KV. Acentriolar mitosis activates a p53-dependent apoptosis pathway in the mouse embryo. Proc. Natl. Acad. Sci. U. S. A. 2014;111:E1491–E1500. doi: 10.1073/pnas.1400568111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carazo-Salas RE, Guarguaglini G, Gruss OJ, Segref A, Karsenti E, Mattaj IW. Generation of GTP-bound Ran by RCC1 is required for chromatin-induced mitotic spindle formation. Nature. 1999;400:178–181. doi: 10.1038/22133. [DOI] [PubMed] [Google Scholar]
- Chabin-Brion K, Marceiller J, Perez F, Settegrana C, Drechou A, Durand G, Poüs C. The Golgi complex is a microtubule-organizing organelle. Mol. Biol. Cell. 2001;12:2047–2060. doi: 10.1091/mbc.12.7.2047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang P, Coughlin M, Mitchison TJ. Interaction between Poly(ADP-ribose) and NuMA contributes to mitotic spindle pole assembly. Mol. Biol. Cell. 2009;20:4575–4585. doi: 10.1091/mbc.E09-06-0477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cutress ML, Whitaker HC, Mills IG, Stewart M, Neal DE. Structural basis for the nuclear import of the human androgen receptor. J. Cell Sci. 2008;121:957–968. doi: 10.1242/jcs.022103. [DOI] [PubMed] [Google Scholar]
- Diao A, Rahman D, Pappin DJC, Lucocq J, Lowe M. The coiled-coil membrane protein golgin-84 is a novel rab effector required for Golgi ribbon formation. J. Cell Biol. 2003;160:201–212. doi: 10.1083/jcb.200207045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dumont J, Petri S, Pellegrin F, Terret M-E, Bohnsack MT, Rassinier P, Georget V, Kalab P, Gruss OJ, Verlhac M-H. A centriole- and RanGTP-independent spindle assembly pathway in meiosis I of vertebrate oocytes. J. Cell Biol. 2007;176:295–305. doi: 10.1083/jcb.200605199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Efimov A, Kharitonov A, Efimova N, Loncarek J, Miller PM, Andreyeva N, Gleeson P, Galjart N, Maia ARR, McLeod IX, et al. Asymmetric CLASP-dependent nucleation of noncentrosomal microtubules at the trans-Golgi network. Dev. Cell. 2007;12:917–930. doi: 10.1016/j.devcel.2007.04.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giesecke A, Stewart M. Novel binding of the mitotic regulator TPX2 (target protein for Xenopus kinesin-like protein 2) to importin-alpha. J. Biol. Chem. 2010;285:17628–17635. doi: 10.1074/jbc.M110.102343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goshima G, Mayer M, Zhang N, Stuurman N, Vale RD. Augmin: a protein complex required for centrosome-independent microtubule generation within the spindle. J. Cell Biol. 2008;181:421–429. doi: 10.1083/jcb.200711053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gruss OJ, Carazo-Salas RE, Schatz CA, Guarguaglini G, Kast J, Wilm M, Le Bot N, Vernos I, Karsenti E, Mattaj IW. Ran induces spindle assembly by reversing the inhibitory effect of importin alpha on TPX2 activity. Cell. 2001;104:83–93. doi: 10.1016/s0092-8674(01)00193-3. [DOI] [PubMed] [Google Scholar]
- Hachet V, Köcher T, Wilm M, Mattaj IW. Importin alpha associates with membranes and participates in nuclear envelope assembly in vitro. EMBO J. 2004;23:1526–1535. doi: 10.1038/sj.emboj.7600154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hehnly H, Doxsey S. Rab11 endosomes contribute to mitotic spindle organization and orientation. Dev. Cell. 2014;28:497–507. doi: 10.1016/j.devcel.2014.01.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heidebrecht HJ, Buck F, Steinmann J, Sprenger R, Wacker HH, Parwaresch R. p100: a novel proliferation-associated nuclear protein specifically restricted to cell cycle phases S, G2, and M. Blood. 1997;90:226–233. [PubMed] [Google Scholar]
- Ishida R, Yamamoto A, Nakayama K, Sohda M, Misumi Y, Yasunaga T, Nakamura N. GM130 is a parallel tetramer with a flexible rod-like structure and N-terminally open (Y-shaped) and closed (I-shaped) conformations. FEBS J. 2015 doi: 10.1111/febs.13271. [DOI] [PubMed] [Google Scholar]
- Jesch SA, Linstedt AD. The Golgi and endoplasmic reticulum remain independent during mitosis in HeLa cells. Mol. Biol. Cell. 1998;9:623–635. doi: 10.1091/mbc.9.3.623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jokitalo E, Cabrera-Poch N, Warren G, Shima DT. Golgi clusters and vesicles mediate mitotic inheritance independently of the endoplasmic reticulum. J. Cell Biol. 2001;154:317–330. doi: 10.1083/jcb.200104073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalab P, Heald R. The RanGTP gradient - a GPS for the mitotic spindle. J. Cell Sci. 2008;121:1577–1586. doi: 10.1242/jcs.005959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalab P, Pu RT, Dasso M. The Ran GTPase regulates mitotic spindle assembly. Curr. Biol. 1999;9:481–484. doi: 10.1016/s0960-9822(99)80213-9. [DOI] [PubMed] [Google Scholar]
- Kalab P, Pralle A, Isacoff EY, Heald R, Weis K. Analysis of a RanGTP-regulated gradient in mitotic somatic cells. Nature. 2006;440:697–701. doi: 10.1038/nature04589. [DOI] [PubMed] [Google Scholar]
- Khodjakov A, Cole RW, Oakley BR, Rieder CL. Centrosome-independent mitotic spindle formation in vertebrates. Curr. Biol. 2000;10:59–67. doi: 10.1016/s0960-9822(99)00276-6. [DOI] [PubMed] [Google Scholar]
- Kodani A, Sütterlin C. The Golgi protein GM130 regulates centrosome morphology and function. Mol. Biol. Cell. 2008;19:745–753. doi: 10.1091/mbc.E07-08-0847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levine TP, Rabouille C, Kieckbusch RH, Warren G. Binding of the vesicle docking protein p115 to Golgi membranes is inhibited under mitotic conditions. J. Biol. Chem. 1996;271:17304–17311. doi: 10.1074/jbc.271.29.17304. [DOI] [PubMed] [Google Scholar]
- Lowe M, Rabouille C, Nakamura N, Watson R, Jackman M, Jämsä E, Rahman D, Pappin DJ, Warren G. Cdc2 kinase directly phosphorylates the cis-Golgi matrix protein GM130 and is required for Golgi fragmentation in mitosis. Cell. 1998;94:783–793. doi: 10.1016/s0092-8674(00)81737-7. [DOI] [PubMed] [Google Scholar]
- Lowe M, Gonatas NK, Warren G. The mitotic phosphorylation cycle of the cis-Golgi matrix protein GM130. J. Cell Biol. 2000;149:341–356. doi: 10.1083/jcb.149.2.341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meunier S, Vernos I. Microtubule assembly during mitosis - from distinct origins to distinct functions? J. Cell Sci. 2012;125:2805–2814. doi: 10.1242/jcs.092429. [DOI] [PubMed] [Google Scholar]
- Nakamura N, Lowe M, Levine TP, Rabouille C, Warren G. The vesicle docking protein p115 binds GM130, a cis-Golgi matrix protein, in a mitotically regulated manner. Cell. 1997;89:445–455. doi: 10.1016/s0092-8674(00)80225-1. [DOI] [PubMed] [Google Scholar]
- Niclas J, Allan VJ, Vale RD. Cell cycle regulation of dynein association with membranes modulates microtubule-based organelle transport. J. Cell Biol. 1996;133:585–593. doi: 10.1083/jcb.133.3.585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oddoux S, Zaal KJ, Tate V, Kenea A, Nandkeolyar SA, Reid E, Liu W, Ralston E. Microtubules that form the stationary lattice of muscle fibers are dynamic and nucleated at Golgi elements. J. Cell Biol. 2013;203:205–213. doi: 10.1083/jcb.201304063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohba T, Nakamura M, Nishitani H, Nishimoto T. Self-organization of microtubule asters induced in Xenopus egg extracts by GTP-bound Ran. Science. 1999;284:1356–1358. doi: 10.1126/science.284.5418.1356. [DOI] [PubMed] [Google Scholar]
- Ori-McKenney KM, Jan LY, Jan Y-N. Golgi Outposts Shape Dendrite Morphology by Functioning as Sites of Acentrosomal Microtubule Nucleation in Neurons. Neuron. 2012;76:921–930. doi: 10.1016/j.neuron.2012.10.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petry S, Groen AC, Ishihara K, Mitchison TJ, Vale RD. Branching Microtubule Nucleation in Xenopus Egg Extracts Mediated by Augmin and TPX2. Cell. 2013;152:768–777. doi: 10.1016/j.cell.2012.12.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pollard VW, Michael WM, Nakielny S, Siomi MC, Wang F, Dreyfuss G. A novel receptor-mediated nuclear protein import pathway. Cell. 1996;86:985–994. doi: 10.1016/s0092-8674(00)80173-7. [DOI] [PubMed] [Google Scholar]
- Preisinger C, Short B, De Corte V, Bruyneel E, Haas A, Kopajtich R, Gettemans J, Barr FA. YSK1 is activated by the Golgi matrix protein GM130 and plays a role in cell migration through its substrate 14-3-3zeta. J. Cell Biol. 2004;164:1009–1020. doi: 10.1083/jcb.200310061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rivero S, Cardenas J, Bornens M, Rios RM. Microtubule nucleation at the cis-side of the Golgi apparatus requires AKAP450 and GM130. EMBO J. 2009;28:1016–1028. doi: 10.1038/emboj.2009.47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sardon T, Peset I, Petrova B, Vernos I. Dissecting the role of Aurora A during spindle assembly. EMBO J. 2008;27:2567–2579. doi: 10.1038/emboj.2008.173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sauer G, Körner R, Hanisch A, Ries A, Nigg EA, Silljé HHW. Proteome analysis of the human mitotic spindle. Mol. Cell. Proteomics. 2005;4:35–43. doi: 10.1074/mcp.M400158-MCP200. [DOI] [PubMed] [Google Scholar]
- Schatz CA, Santarella R, Hoenger A, Karsenti E, Mattaj IW, Gruss OJ, Carazo-Salas RE. Importin alpha-regulated nucleation of microtubules by TPX2. EMBO J. 2003;22:2060–2070. doi: 10.1093/emboj/cdg195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seemann J, Jokitalo EJ, Warren G. The role of the tethering proteins p115 and GM130 in transport through the Golgi apparatus in vivo. Mol. Biol. Cell. 2000;11:635–645. doi: 10.1091/mbc.11.2.635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seemann J, Pypaert M, Taguchi T, Malsam J, Warren G. Partitioning of the matrix fraction of the Golgi apparatus during mitosis in animal cells. Science. 2002;295:848–851. doi: 10.1126/science.1068064. [DOI] [PubMed] [Google Scholar]
- Shima DT, Haldar K, Pepperkok R, Watson R, Warren G. Partitioning of the Golgi apparatus during mitosis in living HeLa cells. J. Cell Biol. 1997;137:1211–1228. doi: 10.1083/jcb.137.6.1211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shima DT, Cabrera-Poch N, Pepperkok R, Warren G. An ordered inheritance strategy for the Golgi apparatus: visualization of mitotic disassembly reveals a role for the mitotic spindle. J. Cell Biol. 1998;141:955–966. doi: 10.1083/jcb.141.4.955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang D, Xiang Y, Wang Y. Reconstitution of the cell cycle-regulated Golgi disassembly and reassembly in a cell-free system. Nat. Protoc. 2010;5:758–772. doi: 10.1038/nprot.2010.38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor RS, Jones SM, Dahl RH, Nordeen MH, Howell KE. Characterization of the Golgi complex cleared of proteins in transit and examination of calcium uptake activities. Mol. Biol. Cell. 1997;8:1911–1931. doi: 10.1091/mbc.8.10.1911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsai M-Y, Wiese C, Cao K, Martin O, Donovan P, Ruderman J, Prigent C, Zheng Y. A Ran signalling pathway mediated by the mitotic kinase Aurora A in spindle assembly. Nat. Cell Biol. 2003;5:242–248. doi: 10.1038/ncb936. [DOI] [PubMed] [Google Scholar]
- Wei J-H, Seemann J. Mitotic division of the mammalian Golgi apparatus. Semin. Cell Dev. Biol. 2009a;20:810–816. doi: 10.1016/j.semcdb.2009.03.010. [DOI] [PubMed] [Google Scholar]
- Wei J-H, Seemann J. The mitotic spindle mediates inheritance of the Golgi ribbon structure. J. Cell Biol. 2009b;184:391–397. doi: 10.1083/jcb.200809090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wiese C, Wilde A, Moore MS, Adam SA, Merdes A, Zheng Y. Role of importin-beta in coupling Ran to downstream targets in microtubule assembly. Science. 2001;291:653–656. doi: 10.1126/science.1057661. [DOI] [PubMed] [Google Scholar]
- Wilbur JD, Heald R. Mitotic spindle scaling during Xenopus development by kif2a and importin α. eLife. 2013;2:e00290. doi: 10.7554/eLife.00290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilde A, Zheng Y. Stimulation of microtubule aster formation and spindle assembly by the small GTPase Ran. Science. 1999;284:1359–1362. doi: 10.1126/science.284.5418.1359. [DOI] [PubMed] [Google Scholar]
- Zhang C-H, Wang Z-B, Quan S, Huang X, Tong J-S, Ma J-Y, Guo L, Wei Y-C, Ouyang Y-C, Hou Y, et al. GM130, a cis-Golgi protein, regulates meiotic spindle assembly and asymmetric division in mouse oocyte. Cell Cycle. 2011;10:1861–1870. doi: 10.4161/cc.10.11.15797. [DOI] [PubMed] [Google Scholar]
- Zheng Y. A membranous spindle matrix orchestrates cell division. Nat. Rev. Mol. Cell Biol. 2010;11:529–535. doi: 10.1038/nrm2919. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.