Abstract
The RNA polymerase III (Pol III)-specific transcription factor Bdp1 is crucial to Pol III recruitment and promoter opening in transcription initiation, yet structural information is sparse. To examine its protein-binding targets within the preinitiation complex at the residue level, photoreactive amino acids were introduced into Saccharomyces cerevisiae Bdp1. Mutations within the highly conserved SANT domain cross-linked to the transcription factor IIB (TFIIB)-related transcription factor Brf1, consistent with the findings of previous studies. In addition, we identified an essential N-terminal region that cross-linked with the Pol III catalytic subunit C128 as well as Brf1. Closer examination revealed that this region interacted with the C128 N-terminal region, the N-terminal half of Brf1, and the C-terminal domain of the C37 subunit, together positioning this region within the active site cleft of the preinitiation complex. With our functional data, our analyses identified an essential region of Bdp1 that is positioned within the active site cleft of Pol III and necessary for transcription initiation.
INTRODUCTION
RNA polymerase III (Pol III) transcribes tRNAs and certain noncoding RNAs, such as 5S rRNA and U6 spliceosomal RNA (1). Pol III is a 17-subunit complex with a 12-subunit core structure homologous to Pol II and five other specific subunits (2). In the Pol III core structure, the two largest subunits, C160 and C128, form the active site cleft, and the other smaller subunits are localized in the periphery. The five Pol III-specific subunits form two separate subcomplexes, C53/C37 and C82/C34/C31, which are homologous to transcription factor IIF (TFIIF) and TFIIE, respectively, in the Pol II system. To initiate transcription, three transcription factors, TFIIIA, TFIIIB, and TFIIIC, cooperate to recruit Pol III to different types of gene promoters and form the preinitiation complex (PIC) (3, 4). For PIC formation at transfer DNA (tDNA) genes, TFIIIC recognizes the gene-internal box A and B elements, while TFIIIB localizes further upstream. TFIIIA is required only for the transcription of 5S rRNA, where it recognizes the gene-internal control element to further recruit TFIIIC and TFIIIB.
TFIIIB is composed of three subunits: the TATA box-binding protein (TBP), TFIIB-related factor 1 (Brf1), and Bdp1. TBP and Brf1 bind tightly, allowing copurification, whereas Bdp1 mostly dissociates during purification (5). The copurified TBP/Brf1 fraction is termed B′, and the Bdp1 fraction is termed B″ (B double prime). TBP is required for all three nuclear transcription systems, and Brf1 belongs to the class of TFIIB-related proteins that also include Rrn7 (in Saccharomyces cerevisiae) and TAF1B (in human) in the Pol I system (6, 7). The N-terminal half of Brf1 contains zinc ribbon and cyclin fold repeat domains related to TFIIB, while the structural features in the C-terminal half of Brf1 are Pol III specific. To date, no transcription factor homologous to Bdp1 has been found in the Pol I and Pol II systems.
Biochemical analysis suggests that the TFIIIB complex is assembled on the promoter in a stepwise fashion, with initial loading of TBP and Brf1 at the promoter and subsequent addition of Bdp1. On the basis of the results of DNA cross-linking and DNase I and hydroxyl radical protection analyses, both Brf1 and Bdp1 approximately localize on the promoter DNA from bases −40 to −2 upstream of the transcription start site (referred to as the +1 base), while TBP binds the DNA from bases −20 to −31 (8–15). The two termini of Brf1 have different functions, with the C-terminal half (amino acids [aa] 165 to 596) nucleating the TBP-Brf1-Bdp1 complex upstream of the transcription start site and the N-terminal half positioning within the Pol III active site cleft to assist with promoter opening (16–18).
In the yeast Saccharomyces cerevisiae, Bdp1 is a 594-amino-acid polypeptide with an apparent molecular mass of 90 kDa by gel electrophoresis (14, 19). The C-terminal region contains a conserved SANT (SWI3, ADA2, N-CoR, and TFIIIB B″; aa 415 to 472) domain that generally functions as a DNA-binding module (Fig. 1A) (20). A nuclear magnetic resonance (NMR) study has revealed a triple α-helical fold of the SANT domain, and further binding analysis suggests an interaction with the C terminus of Brf1 (21). Sequence regions flanking the SANT domain are also conserved; however, structural information is still lacking (4, 22). According to previous genetic analysis of yeast Bdp1, the SANT domain, together with its flanking sequences, forms the essential region I, and two additional essential sequence blocks were also identified, including region II (RII; aa 269 to 312) and region III (aa 158 to 252) (Fig. 1A) (23). To date, structural information and protein interactions for regions II and III are still lacking.
FIG 1.
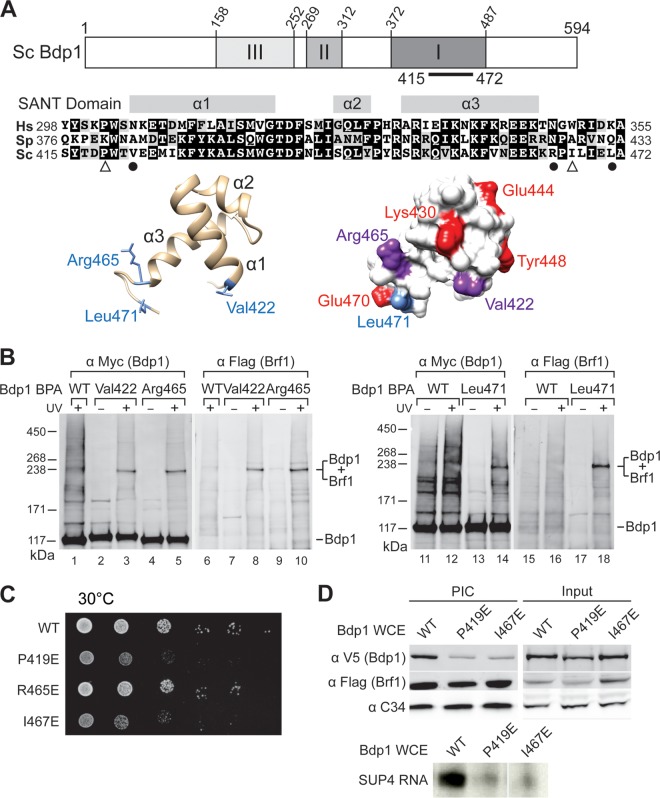
The Bdp1 SANT domain cross-links to Brf1 and functions in PIC formation. (A) (Top) Essential regions I, II, and III in Saccharomyces cerevisiae (Sc) Bdp1 are shaded in the schematic. The black bar indicates the SANT domain. (Middle) Sequence alignment of the SANT domain from Bdp1 homologues. Hs, Homo sapiens (UniProt [http://www.uniprot.org/] accession number A6H8Y1; Sp, Schizosaccharomyces pombe, accession number O94481; Sc, Saccharomyces cerevisiae, UniProt accession number P46678. The three α helices in the SANT domain are indicated. Identical and similar residues are shaded in black and gray, respectively. Symbols below the sequence alignment: dots, residues with BPA cross-linking to Brf1; triangles, point mutations resulting in slow cell growth and defective transcription activity. (Bottom) Structural model of the SANT domain in a backbone trace (left) and the molecular surface (right). The model was generated by the Modeler program with the sequence with PDB accession number 2CU7 as the template. The blue side chains in the backbone trace model indicate residues involved in cross-linking with Brf1. In the molecular surface model, red residues are proposed to directly interact with Brf1, on the basis of NMR analysis. The residues interacting with Brf1, derived from both NMR analysis and BPA cross-linking, are in purple. Leu471 is proposed to interact only with Brf1 on the basis of BPA cross-linking. (B) Western analysis of photo-cross-linking. The BPA-substituted residues in the SANT domain are indicated above the lanes. Anti-Myc antibody revealed Myc epitope-tagged Bdp1 and fusion protein (lanes 1 to 5 and lanes 11 to 14). Anti-Flag antibody revealed the Flag epitope-tagged Brf1 in the fusion protein (lanes 6 to 10 and lanes 15 to 18). UV + and UV −, UV irradiation and no UV irradiation, respectively; WT, wild-type Bdp1 without BPA. (C) Cell growth analysis of Bdp1 SANT domain mutants by serial dilution spot assay. Representative slow cell growth at 30°C is shown. (D) Bdp1 SANT domain mutations affect PIC formation. (Top) Western blot analysis of proteins isolated from PIC formation. As indicated, Bdp1 mutant or WT whole-cell extracts were used in the IMT assay to isolate PICs on DNA containing the SUP4 gene. Individual proteins were probed with the antibodies indicated on the left. (Bottom) Autoradiogram of the synthesized SUP4 preliminary tRNA from PICs of the IMT assay.
Here, we dissected the residue-level interactions of Bdp1 within the Pol III PIC. We incorporated the photo-cross-linker p-benzoyl-l-phenylalanine (BPA) as a nonnatural amino acid into Bdp1 to site-specifically identify its protein interactions within the Pol III PIC. Consistent with previously proposed interactions, BPA cross-linking revealed that the SANT domain of Bdp1 binds Brf1. Additionally, BPA replacement in region II of Bdp1 resulted in cross-links with the C128 subunit of Pol III. Further protein mapping suggests that Bdp1 region II interacts with the N-terminal region of C128, suggesting that it is positioned within the Pol III active site cleft. This localization of Bdp1 region II is also supported by its interactions with the N-terminal half of Brf1 and the C-terminal domain of the C37 subunit of Pol III, both of which are known to reside in the Pol III active site cleft (17, 24). Our findings thus confer a novel functional role to Bdp1 region II within the Pol III active center.
MATERIALS AND METHODS
Plasmids and yeast strains.
The coding sequence for Saccharomyces cerevisiae Bdp1 was cloned into the yeast high-copy-number vector pRS425 (2μ) with a LEU2 selection marker (25). Subsequent subcloning with DNA insertions flanking the Bdp1-coding sequence resulted in the plasmid pHL1 (Adh1-Bdp1c13Myc-pRS425), which contains the 414-bp ADH1 promoter and the 13-Myc epitope tag at the C terminus of Bdp1. This plasmid was used to generate by site-directed mutagenesis individual Bdp1 mutant plasmids harboring single TAG (amber) nonsense codon substitutions at specific amino acid positions. All TAG-containing Bdp1 mutant plasmids are generally referred to as Bdp1-amber plasmids.
A yeast plasmid shuffle strategy was utilized to generate all Bdp1 mutant strains. We generated a strain, based on yeast strain BY4705 (26), containing a chromosomal deletion of Bdp1 that is compensated for by the Bdp1 gene in a URA3-marked low-copy-number (cen ars) plasmid, and the new strain was named HLy1 [MATα ade2::his3G his3Δ200 leu2Δ met15Δ lys2Δ trp1Δ63 ura3Δ (bdp1::KanMX4) Bdp1-pRS316 (URA3+)]. To generate individual strains into which the nonnatural amino acid BPA could be incorporate into Bdp1, we initially cotransformed a Bdp1-amber plasmid and the pLH157 plasmid into HLy1. Plasmid pLH157 contains the tRNA–aminoacyl-tRNA synthetase pair for nonsense suppression of the TAG codon to incorporate the BPA cross-linker at a selected amino acid position (27, 28). Following plasmid shuffling on medium containing 5-fluoroorotic acid (5-FOA) and the nonnatural amino acid BPA, viable yeast strains retaining the Bdp1-amber and pLH157 plasmids were selected and are referred to as Bdp1-amber strains.
Yeast strains for Bdp1 mutational analysis (strains with internal deletions and single/multiple point mutations) were similarly generated by plasmid shuffling. However, all Bdp1 mutant plasmids were generated on the basis of the LEU2-marked low-copy-number (cen ars) plasmid pHL2 (Bdp1-V5/pRS315), which contains its endogenous promoter and a C-terminal V5 epitope tag in Bdp1. For cell growth analysis, the wild-type and mutant strains were grown in yeast extract-peptone-dextrose (YPD) medium until the optical density at 600 nm (OD600) was 1.0, the cell cultures were subsequently diluted 10−1, 10−2, 10−3, and 10−4, and the diluted cells were spotted on synthetic complete glucose plates lacking leucine. The cell growth phenotypes were monitored for 3 days at 25°C, 30°C, and 37°C and for 5 days at 16°C.
Preinitiation complex isolation and BPA photo-cross-linking.
The Bdp1-amber strains were grown in YPD medium containing BPA until the OD600 was 1.5, and the harvested cells were subjected to whole-cell extract (WCE) preparation by grinding in liquid nitrogen (LN2) and subsequent ultracentrifugation. WCEs were applied to the immobilized template assay for PIC formation on the PCR-generated and 5′-biotinylated 603-bp DNA fragment containing the SUP4 tRNA gene. A detailed procedure for the immobilized template assay was described previously (24). For a typical photo-cross-linking experiment, 800 μg WCE was incubated with 4 μg of the DNA template immobilized on streptavidin magnetic beads (Dynal) for 30 min at 30°C. Each reaction mixture was washed three times with transcription buffer (20 mM HEPES [pH 7.9], 80 mM KCl, 5 mM MgCl2, 1 mM EDTA, 2% glycerol, 0.01% Tween 20). After the reaction mixture was washed, each reaction mixture was divided into two fractions, one for UV irradiation and the other as a control (no UV irradiation). UV irradiation was performed with a total energy of 9,000 μJ in a Spectrolinker XL-1500 UV oven (Spectronics). The PICs from photo-cross-linking were then resuspended in NuPAGE loading buffer (Invitrogen) for sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis (PAGE) and Western blot analysis. The Western blot results were visualized with a LI-COR BioScience Odyssey infrared imaging system with fluorescent dye-labeled secondary antibodies.
In vitro transcription.
The PICs for in vitro transcription were isolated by the immobilized template assay as described above. The isolated PICs were resuspended in 17 μl of transcription buffer containing 200 ng α-amanitin, 4 U of RNase inhibitor (Promega), and 1 mM dithiothreitol (DTT). Transcription was initiated by adding 3 μl of a nucleoside triphosphate mixture containing ATP (500 μM), UTP (500 μM), CTP (500 μM), GTP (50 μM), and [α-32P]GTP (0.16 μM, 3,000 Ci/mmol). The transcription reaction was performed for 30 min at 30°C. The reactions were stopped by adding 180 μl of 0.1 M sodium acetate, 10 mM EDTA, 0.5% SDS, and 200 μg/ml glycogen. The RNA product was extracted with phenol-chloroform, precipitated with ethanol, and then resolved by 8% denaturing urea-polyacrylamide gel electrophoresis. The results from gel electrophoresis were visualized by autoradiography.
Protein purification.
The DNA sequence encoding Bdp1 region II (RII; from amino acids 253 to 325) with a C-terminal single Flag epitope and six histidines was cloned into the pET21a vector (Novagen) to generate the plasmid pET21a-Bdp1 RII-Flag-His6. The plasmid was transformed into the Escherichia coli BL21(DE3) strain (Novagen) for recombinant protein expression. The transformed cells were grown to an OD600 of ∼0.8 at 37°C and were induced with 0.4 mM isopropyl-β-d-1-thiogalactopyranoside (IPTG). Cells were harvested at 2 h after induction, resuspended in lysis buffer (20 mM Tris-HCl [pH 7.5], 150 mM NaCl, 10% glycerol, 30 mM imidazole), and lysed with a Microfluidizer. The clarified lysate from centrifugation was subjected to initial affinity purification by Ni-Sepharose (GE Healthcare). Bdp1 RII was eluted with elution buffer (20 mM Tris-HCl [pH 7.5], 150 mM NaCl, 10% glycerol, 250 mM imidazole). DNA sequences encoding the Brf1 N-terminal half from aa 1 to 284 and the C128 N-terminal fragment from aa 1 to 535 were separately cloned into the pET21a vector to generate the plasmids pET21a-Brf1 Nt-V5-His6 and pET21a-C128-Nt-His6, respectively. The polypeptide consisting of the Brf1 N-terminal half was expressed in an E. coli BL21(DE3) strain that also allows coexpression of GroEL/ES (TaKaRa). The C128 N-terminal protein fragment was similarly expressed in the strain coexpressing GroEL/ES. In both protein induction experiments, the transformed cells, induced by 0.4 mM IPTG and 1 ng/ml tetracycline, were initially grown at 37°C to an OD600 of ∼0.8 and grown for another 4 h at 25°C. The cells were harvested and lysed, and the proteins were purified by Ni-Sepharose using the same protocol described above. All protein samples were then dialyzed in buffer containing 20 mM Tris-HCl (pH 7.9), 150 mM KCl, and 10% glycerol.
IP assay.
WCEs from yeast strains containing C-terminal Flag epitope-tagged C128 and C-terminal V5 epitope-tagged wild-type (WT) or mutant Bdp1 were used for immunoprecipitation (IP). WCE (1 mg) was incubated with 50 μl of anti-Flag–agarose beads (Sigma) in WCE dialysis buffer containing 20 mM HEPES (pH 7.9), 100 mM KCl, 5 mM MgCl2, 1 mM EDTA, and 20% glycerol for 1 h at 4°C. Following three washes in 800 μl of WCE dialysis buffer, the bound proteins were eluted by boiling the beads at 99°C for 5 min in 25 μl of NuPAGE loading buffer (Invitrogen). The eluted proteins were resolved by SDS-PAGE and analyzed by Western blotting with anti-Flag (for C128) and anti-V5 (for Bdp1) antibodies. For co-IP with protein fragments, the Bdp1 RII fragment sample (40 μg) was mixed with the Brf1 N-terminal half polypeptide or the C128 N-terminal fragment (20 μg for each sample), and the mixture was incubated with 50 μl anti-Flag–agarose beads (Sigma) in IP buffer (20 mM Tris-HCl [pH 7.9], 150 mM KCl, 10% glycerol, 0.5% Tween 20) at 4°C for 1 h. Following three washes with 800 μl IP buffer, the bound proteins were eluted by boiling the beads at 99°C for 5 min in 25 μl of NuPAGE loading buffer (Invitrogen) and analyzed by Western blotting.
FeBABE cleavage assay and determination of hydroxyl radical cleavage site.
Conjugation of the hydroxyl radical-generating probe FeBABE to the recombinant C37/C53 heterodimer and subsequent utilization of the FeBABE-tethered C37/C53 dimer for directed protein cleavage analysis of the Pol III PIC were performed as described previously (24). Briefly, in a typical FeBABE cleavage experiment, 400 μg yeast WCE containing C53 lacking aa 2 to 280 [C53 Δ(2–280)] and C-terminally Flag epitope-tagged Bdp1 was incubated with 4 μg the C37/C53-FeBABE derivative in a 200-μl immobilization template assay for PIC isolation. Pol III PICs were isolated with the immobilized SUP4 tDNA template, and sodium ascorbate and hydrogen peroxide were sequentially added to activate the iron-EDTA moiety for generating hydroxyl radicals (29). The resulting Bdp1 cleavage fragment was visualized following Western blotting with an anti-Flag antibody. The cleavage site was determined by comparing the mobility shift of the Bdp1 cleavage gel band with a ladder of in vitro-translated Bdp1 peptide fragments. Detailed information on the procedure for hydroxyl radical cleavage site determination was described previously (29).
RESULTS
The Bdp1 SANT domain interacts with Brf1 within the Pol III preinitiation complex.
To identify protein interaction targets for Bdp1, we site-specifically incorporated the photo-reactive cross-linker BPA as a nonnatural amino acid into Bdp1 by utilizing the nonsense suppression strategy developed by the Schultz laboratory (27). We obtained 141 viable mutant strains each with single-site BPA incorporation in Bdp1 (see Table S1 in the supplemental material). We applied whole-cell extracts (WCEs) from individual Bdp1 mutant strains to the immobilized template (IMT) assay coupled with UV irradiation to analyze the protein-binding targets in the PIC. We observed the most significant cross-linking results for BPA substitutions in the highly conserved SANT domain of Bdp1. As demonstrated in Fig. 1B, BPA replacements at amino acid positions Val422, Arg465, and Leu471 yielded a high-intensity cross-linked band with a molecular mass of ∼210 kDa. By subtracting the molecular mass of ∼120 kDa for the C-terminally Myc epitope-tagged Bdp1, the cross-linked polypeptide was estimated to have a size of ∼90 kDa. As the molecular mass is close to that of another TFIIIB subunit, Brf1, we applied a genetic method to insert the Flag epitope into Brf1 in the same Bdp1 mutant strains. By repeating the cross-linking experiment with WCEs from the newly derived strains, we confirmed that the cross-linked gel bands were Bdp1-Brf1 fusions by immunostaining with both anti-Myc (Bdp1) and anti-Flag (Brf1) antibodies (Fig. 1B). This result is consistent with protein pulldown and NMR analyses suggesting that the SANT domain interacts with the Brf1 C-terminal region (21, 30). In addition, Val422 and Arg465 are positioned, respectively, in α helices 1 and 3 of the SANT domain that form the Brf1-binding surface, as suggested by NMR (Fig. 1A) (21).
Internal deletions in the SANT domain, such as Bdp1Δ409–421 and Bdp1Δ424–438, lacking amino acids 409 to 421 and 424 to 438, respectively, have previously proven lethal (23). We therefore designed smaller deletions of residues involved in Bdp1-Brf1 cross-linking. However, deletions of short segments, including Bdp1Δ419–423, Bdp1Δ461–465, Bdp1Δ464–466, Bdp1Δ466–470, Bdp1Δ467–469, and Bdp1Δ470–472, were lethal as well. Although cell lethality was likely caused by the removal of Brf1-binding residues, the structural fold of the SANT domain might be disrupted. We thus additionally introduced point mutations into the conserved residues structurally located on the protein surface of the SANT domain. Although Trp420Glu and Pro466Glu single point mutations resulted in lethality, strains with two other mutations, Pro419Glu and Ile467Glu, showed slow growth at all temperatures, including 16, 25, 30, and 37°C (Fig. 1C). WCEs from these two mutants were applied to the IMT assay, and the isolated PICs were analyzed by Western blot analysis to determine individual protein levels in the PIC. As demonstrated in Fig. 1D, the two mutations resulted in a reduced Bdp1 protein level, indicating that the mutants might be defective in either the formation or stability of PICs. The results of further analysis by an in vitro transcription assay also correlated with the reduced Bdp1 protein level in the PIC (Fig. 1D, bottom). Together with the cross-linking data and the previous findings of biochemical and structural analyses, our study validates the Brf1-binding surface of the SANT domain and its functional importance in PIC formation.
Region II of Bdp1 interacts with Brf1 and the C128 subunit of Pol III.
Our BPA-cross-linking analysis identified another Brf1-binding site in the sequence N terminal to the SANT domain. As shown in Fig. 2A, a BPA substitution at residue Lys281 generated a cross-linked band confirmed to be Brf1. This Brf1-Bdp1 cross-link is positioned in the previously defined essential region II of Bdp1 (amino acids 269 to 312) (Fig. 1A) (23). In addition to the Brf1 cross-link, BPA substitutions at Glu291, Asn292, and Lys295 in region II yielded cross-linked polypeptides confirmed to be the Bdp1-C128 fusion product (Fig. 2B). Our cross-linking data thus suggest that Bdp1 region II functions as a structural domain for Brf1 and C128 interactions.
FIG 2.
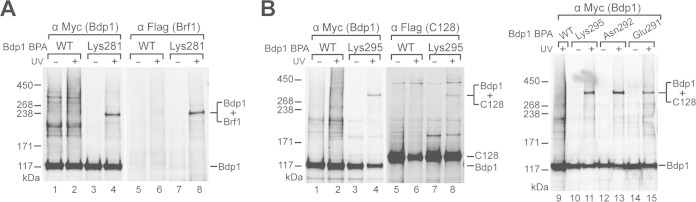
Bdp1 region II cross-links to Brf1 and C128. (A) Bdp1-Brf1 photo-cross-linking. Bdp1 and the photo-cross-linked bands were detected by anti-Myc antibody (lanes 1 to 4). Anti-Flag antibody against tagged Brf1 confirmed its presence in the cross-linked polypeptide (lanes 5 to 8). (B) BPA-substituted residues Glu291, Asn292, and Lys295 in Bdp1 cross-linked with C128 (right), as confirmed by probing with anti-Flag antibody against Flag-tagged C128 in the fusion protein band (left).
Bdp1 region II functions in a step after PIC formation.
With the cross-linking data indicating Brf1 and C128 interactions in Bdp1 region II, we proceeded with a mutagenesis study to further analyze the functional importance of this region. We first targeted residues flanking Lys281, where BPA substitution yielded Brf1 cross-linking (Fig. 2A). As summarized in Fig. 3A, the single point mutations Leu279Ala, Cys280Ala, and Phe284Ala were lethal. Several other point mutations, including Ala277Glu, Glu278Ala, and Pro282Ala, resulted in a temperature-sensitive (ts) phenotype (Fig. 3B; summarized in Fig. 3A). We subsequently utilized whole-cell extracts from the ts mutants for in vitro transcription analysis. As shown in Fig. 3C, these mutations severely affected tRNA and U6 RNA synthesis. However, these Bdp1 mutations did not affect protein levels for Bdp1, Brf1, Pol III (C128), and TFIIIC (Tfc4) in the analysis of PIC formation using the IMT assay (Fig. 3D). We conclude that the Bdp1-Brf1-binding interface in region II appears to function only in a step after PIC formation, in contrast to the functional contribution of the SANT domain in PIC formation or stability (Fig. 1D).
FIG 3.
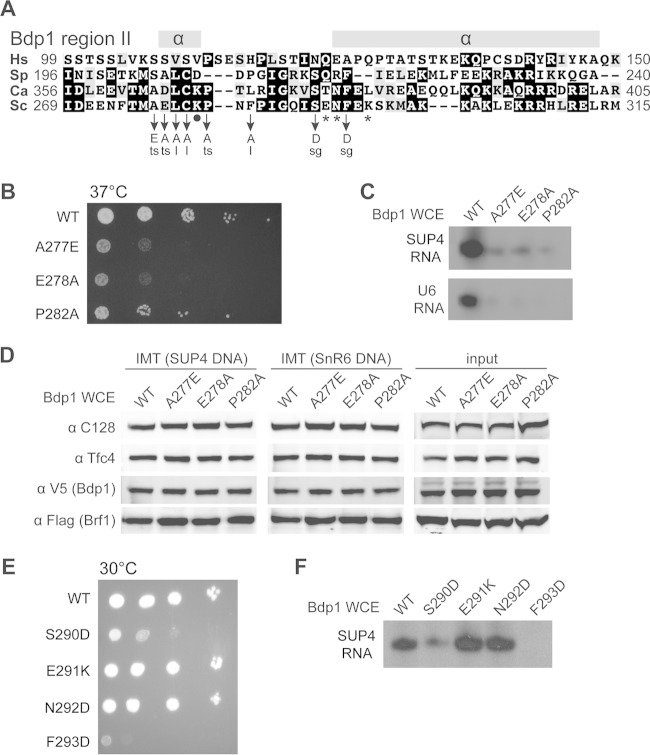
Functional analysis of Bdp1 region II. (A) Bdp1 region II multiple-sequence alignment. As depicted in Fig. 1A, region II of Saccharomyces cerevisiae (Sc) Bdp1 ranges from amino acids 269 to 315. Hs, Homo sapiens; Sp, Schizosaccharomyces pombe; Ca, Candida albicans. The secondary structure of region II was predicted by use of the PSIPRED protein analysis workbench (http://bioinf.cs.ucl.ac.uk/psipred/). Symbols below the sequence alignment: asterisks, residues with BPA-cross-linking to C128; dot, the residue with BPA-cross-linking to Brf1. Point mutations and associated cell growth phenotypes are listed below. ts, temperature-sensitive (37°C) phenotype; sg, slow growth at all temperatures (16, 25, 30, and 37°C); l, amino acid substitution causing lethality. (B) Cell growth analysis of mutants with mutations in Bdp1 region II residues flanking the Brf1 BPA-cross-linking site. The three mutations caused a temperature-sensitive phenotype, as shown by serial dilution cell growth at 37°C. (C) In vitro transcription analysis for Bdp1 region II mutations flanking the residue cross-linking to Brf1. The autoradiograms show RNAs generated from the PICs of the IMT assay. As indicated, whole-cell extracts containing either WT or Bdp1 mutants were utilized in the in vitro transcription analysis. (D) PIC formation analysis of Bdp1 region II mutants. As indicated, Bdp1 mutant or WT whole-cell extracts were used in the IMT assay to isolate PICs on DNAs containing the SUP4 or SnR6 (U6) gene. Individual proteins of the isolated PICs were probed with antibodies, as indicated, in the Western blot analysis. (E) Cell growth analysis of mutations in Bdp1 region II residues involved in the C128 interaction. Representative slow cell growth at 30°C is shown. (F) In vitro transcription analysis for mutations in Bdp1 region II residues involved in the C128 interaction. The autoradiogram shows SUP4 tRNA from an in vitro transcription assay using WT or Bdp1 mutant whole-cell extracts.
We next turned our attention to the region that interacted with C128 in our BPA-cross-linking experiments (Fig. 2B; summarized in Fig. 3A). While double point mutations (Ser290Asp, Phe293Asp) or short internal deletion of amino acids 290 to 293 (Δ290–293) were lethal, mutants with single point mutations, including Ser290Asp and Phe293Asp, were viable but exhibited a slow-growth phenotype (Fig. 3E; summarized in Fig. 3A). In vitro transcription activity was analyzed with WCEs derived from these single point mutants. In agreement with the observed cell growth defects, mutants exhibited compromised SUP4 tRNA synthesis (Fig. 3F). Further analysis by the immobilized template assay indicated that these mutations did not appear to affect the protein levels of Bdp1 and Pol III (C128) in the isolated PICs (data not shown). In summary, although region II serves as a functionally important protein-binding region for Brf1 and C128, the interactions do not appear to be responsible for the recruitment of polymerase or the stable association of proteins in the PIC. We suggest that Bdp1 region II is involved in a network of protein interactions for a critical step after PIC formation.
Bdp1 region II interacts with the N-terminal half of Brf1 and the Pol III active site cleft.
Based on our photo-cross-linking and mutational data, we surmised that Bdp1 region II is an important binding interface for Brf1 and C128. The new results prompted us to investigate what structural regions of Brf1 and C128 interact directly with region II. We first purified the peptide fragments of Brf1 and tested their binding with the purified Bdp1 region II peptide fragment (aa 253 to 325). Following coimmunoprecipitation, Brf1 N-terminal fragments containing amino acids 1 to 90 and 1 to 284 showed substantial interaction with Bdp1 region II (Fig. 4A, lanes 5 and 6). We subsequently introduced mutations into the Bdp1 region II peptide fragment and tested the binding affinity with Brf1. As shown in Fig. 4A (lanes 7 and 8), Bdp1 mutations in Leu279, Cys280, and Phe284, previously shown to be important for cell viability (Fig. 3A), severely compromised the Brf1-Bdp1 interaction. This was also consistent with previous mutational analysis suggesting interaction between the Bdp1 N-terminal region and the Brf1 N-terminal half (30).
FIG 4.
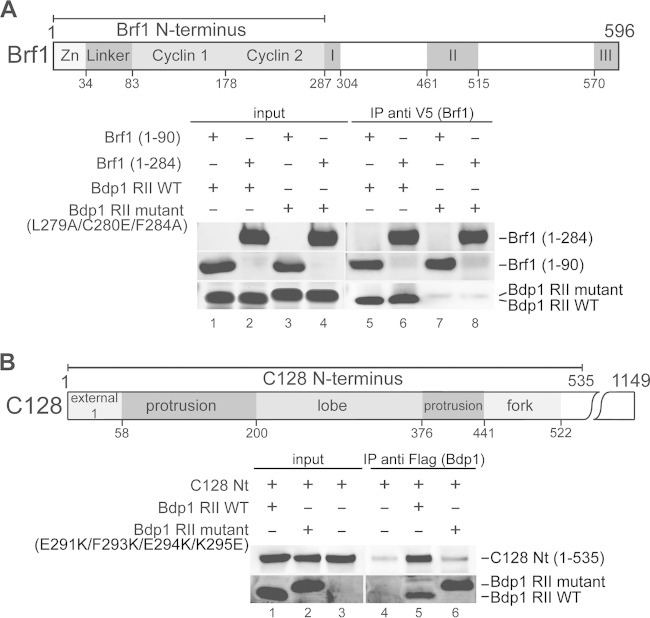
Bdp1 region II interacts with the Brf1 N terminus and the C128 N terminus. (A) (Top) A schematic of Brf1 indicates the N-terminal zinc ribbon, linker, and cyclin fold repeat domains as well as the C-terminal conserved sequence blocks I, II, and III. (Bottom) Western blot analysis of coimmunoprecipitation with V5 epitope-tagged Brf1 N-terminal fragments from aa 1 to 90 and aa 1 to 284 and a Bdp1 region II peptide fragment containing aa 253 to 325. Anti-V5 antibody-agarose beads were used to precipitate the V5-tagged Brf1 fragments. As indicated, either the WT Bdp1 region II peptide or the mutant peptide with triple point mutations was applied in the coimmunoprecipitation assay. (B) (Top) A schematic of the C128 N-terminal region includes the external 1, protrusion, lobe, and fork domains. (Bottom) Western blot analysis of coimmunoprecipitation with Bdp1 region II and C128 N-terminal (Nt) peptide fragments. Anti-Flag antibody-agarose beads were applied to precipitate the Flag-tagged Bdp1 peptide fragment. As indicated, either the WT Bdp1 region II fragment or the Bdp1 region II fragment with a quadruple point mutation was used. Coimmunoprecipitated polypeptides were probed with anti-Flag and anti-His antibodies, revealing the Bdp1 region and the C128 N terminus, respectively.
Our binding analysis was next extended to the C128 subunit of Pol III. We purified a peptide fragment of the C128 N-terminal region encompassing the lobe, protrusion, and fork domains that contribute to the formation of the Pol III active site cleft. As demonstrated in Fig. 4B (lane 5), the purified C128 peptide fragment showed substantial coimmunoprecipitation with the immobilized Bdp1 region II peptide fragment. Mutations were further introduced into Bdp1 at residues found to be involved in the C128 interaction on the basis of the BPA-cross-linking analysis. In agreement with the cross-linking data, the Bdp1 region II mutant peptide fragment failed to pull down comparable amounts of the C128 N-terminal fragment (Fig. 4B, lane 6). Based on the binding analysis with the C128 N-terminal fragment, we conclude that Bdp1 region II is possibly localized in the Pol III active site cleft. Supporting this conclusion, Bdp1 region II also interacts with the Brf1 N-terminal half proposed to be positioned in the Pol III active site cleft as well (17).
Bdp1 region II further interacts with the C-terminal domain of C37 in the Pol III active site cleft.
On the basis of the findings of our previous cross-linking and hydroxyl radical probing analyses, the C37 subunit of Pol III dimerizes with another subunit, C53, to position itself on the lobe domain of C128. Extending from the dimerization domain, the C37 C-terminal domain is positioned in the Pol III active site cleft to interact with the lobe, protrusion, and fork domains of C128 (24). The C37 C-terminal domain also interacts with C34 and Bdp1, as evidenced by the cross-linking data showing that BPA substitutions in residues Met215 and Ser217 of C37 simultaneously cross-link with C128, C34, and Bdp1 (24). To further characterize the interaction between C37 and Bdp1, we conducted a site-directed hydroxyl radical analysis to map the C37 binding position on Bdp1. We conjugated the hydroxyl radical-generating probe FeBABE to C37 single cysteine mutants and applied the FeBABE conjugates to the immobilized template assay. On the basis of the results of multiple site-directed hydroxyl radical experiments, a specific Bdp1 cleavage fragment was identified when the FeBABE-conjugated Ser215Cys mutant of C37 was applied in the PIC formation analysis (Fig. 5A). We subsequently determined the cleavage site in Bdp1 by comparing the size of the cleaved peptide fragment in gel electrophoresis with the sizes of the fragments on the Bdp1 peptide ladder generated by in vitro translation. The cleavage site in Bdp1 was calculated to be at approximately aa 264, which is located immediately N terminal to region II. This directed hydroxyl radical analysis thus localizes a possible binding site for the C37 C-terminal domain in Bdp1.
FIG 5.
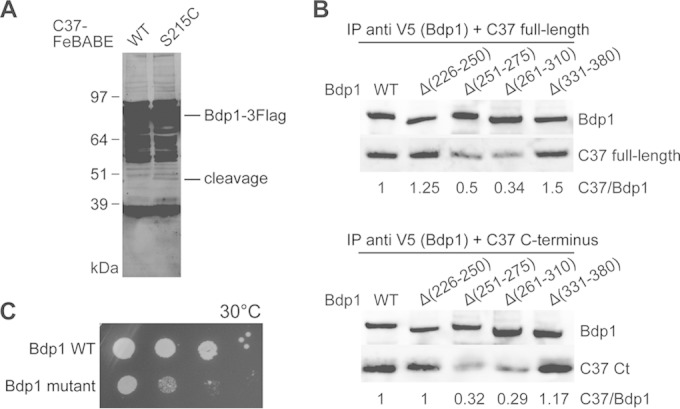
The C37 C terminus interacts with Bdp1 region II. (A) Western blot analysis of hydroxyl radical cleavage of Bdp1 by FeBABE-conjugated C37. As indicated, either the wild type or the C37 mutant with a single cysteine mutation at Ser215 was used in the FeBABE conjugation and subsequent hydroxyl radical protein cleavage assay. C-terminally Flag-tagged full-length Bdp1 and the cleaved peptide fragments were visualized by probing with an anti-Flag antibody. The corresponding cleavage site for the Bdp1 peptide fragment is aa 264. (B) Western blot analysis of coimmunoprecipitation of Bdp1 and C37. Anti-V5 antibody beads were used to precipitate the C-terminally V5 epitope-tagged Bdp1 and internal deletion mutants. Either the C37 full-length protein (top) or the C37 C-terminal (Ct) peptide fragment from aa 181 to 281 (bottom) was used in the coimmunoprecipitation assay. Bdp1 and C37 were probed with anti-V5 and anti-His antibodies, respectively. The relative intensity of C37 immunostaining is normalized to the intensity of Bdp1 immunostaining and is listed below each lane. (C) The cell growth phenotype was analyzed by a serial dilution spot assay. As indicated, Bdp1 quadruple point mutant R260E/D261K/E263K/K266E and WT strains were grown at 30°C.
To further analyze the direct C37-Bdp1 protein interaction, we purified a series of Bdp1 mutant protein samples and examined their abilities to bind C37 by coimmunoprecipitation. As shown in Fig. 5B, two Bdp1 internal deletion mutants, Bdp1 Δ251–275 and Bdp1 Δ261–310, exhibited reduced binding with both purified C37 and the C37 C-terminal fragment (amino acids 181 to 281). In contrast, the Bdp1 Δ226–250 and Bdp1 Δ331–380 mutants did not exhibit a compromised Bdp1-C37 interaction. The binding data thus validate that C37 binds Bdp1 at the sequence centered at about aa 264.
Based on the hydroxyl radical probing and coimmunoprecipitation data, we introduced Bdp1 mutations for cell growth analysis. Mutant strains with single point mutations in residues flanking the hydroxyl radical cleavage site (aa 264) were generated and showed no pronounced growth defects (data not shown). In a further internal deletion mutant, removal of amino acids 255 to 270 of Bdp1 resulted in cell lethality, and the Bdp1 mutant with multiple point radical substitutions for charged residues Arg260, Asp261, Glu263, and Lys266 (Bdp1 R260E/D261K/E263K/K266E) had a slow-growth phenotype (Fig. 5C). The mutational data thus suggest the functional importance of the C37-binding sequence region at about aa 264 of Bdp1.
DISCUSSION
Here we identified the protein targets of Bdp1 in PIC by site-specific photo-cross-linking, and we characterized the functional importance of the Bdp1 protein network by genetic and in vitro transcription analyses. Our results confirmed the interaction between the Bdp1 SANT domain and Brf1 (21, 30). We further identified Bdp1 region II to be an important protein-binding module for Brf1, C128, and C37. On the basis of our protein-binding analysis, the region is likely localized within the active site cleft of Pol III. The SANT domain and region II are added into the model of the Pol III PIC derived from previous structural and biochemical studies (31–33). As illustrated in Fig. 6, the SANT domain is positioned next to the Brf1 C-terminal block II motif that forms a complex with TBP and upstream promoter DNA. Bdp1 region II, meanwhile, is positioned within the Pol III active site cleft and is in direct contact with C128, the N-terminal region of Brf1, and the C-terminal domain of C37.
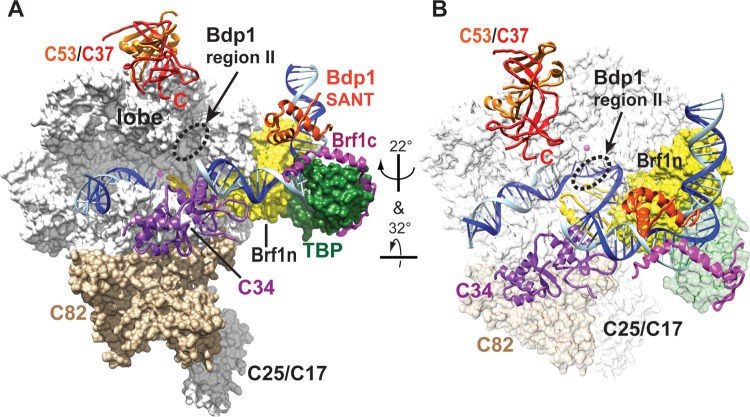
FIG 6.
Positions of Bdp1 region II and the SANT domain in the Pol III open promoter complex model. (A) The model of the Pol III-Brf1-TBP-DNA open promoter complex was initially assembled with the structures of the Pol II-TFIIB-TBP complex, the Brf1-TBP-DNA complex, and Pol I (31, 43–46). The C82 and C34 structures were generated on the basis of homology modeling using structures of human homologues as the templates. The C53/C37 dimerization module was generated using the human TFIIF (Rap74/Rap30) dimer structure as the homology modeling template. C82, C34, and the C53/C37 dimerization module were added to Pol III on the basis of biochemical cross-linking and binding data, as described previously (31). Template and nontemplate DNA strands are in blue and cyan, respectively. The Pol III 12-subunit core structure (white), C82 (tan), the Brf1 cyclin fold repeat domain (yellow), and TBP (dark green) are shown as a molecular surface representation. The magenta sphere in the Pol III core structure represents the active site magnesium ion. C34 (purple), C53 (orange), C37 (red), the Bdp1 SANT domain (orange-red), and Brf1 C-terminal block II (magenta) are shown as backbone trace representations. TFIIB N-terminal zinc ribbon, reader, and linker domains (yellow) are placed in the Pol III active site cleft to indicate likely positions for Brf1 N-terminal zinc ribbon and linker domains. (B) Different orientation of the open promoter complex. Pol III core, C82, and TBP are in semitransparent surface representations.
In the structure proposed here, Bdp1 region II may participate in a network comprising proteins and promoter DNA within the active site cleft that orchestrates transcription initiation. As shown in Fig. 6, the network also includes C34 winged helix (WH) 1 and 2 domains, the C37 C-terminal domain, and the Brf1 linker and cyclin fold domains. Both Brf1 and C34 are involved in promoter DNA melting (18, 34, 35). In contrast, the C37 C-terminal domain functions in transcription termination (24, 36, 37). On the basis of previous genetic and biochemical analyses, aa 355 to 421 of the essential region I of Bdp1 (Fig. 1A) were proposed to function in promoter DNA melting (12, 34). BPA substitutions in this sequence commonly cross-linked to a protein with an estimated molecular mass larger than the molecular masses of Brf1 and C128 (data not shown; Table S1 in the supplemental material). However, due to weak cross-linking intensity, we were unable to determine the identity of the cross-linked protein.
Genetic analysis also linked Bdp1 with Rpr1, the RNA component of RNase P, as well as Tfc4, a subunit of the transcription factor TFIIIC (23). However, our cross-linking analysis was not applied to the protein-RNA interaction, and no Tfc4 cross-linking was identified. In addition, human Bdp1 is targeted by the CK2 kinase (38), which targets the region between aa 355 and 470 for phosphorylation. On the basis of sequence alignment, this is homologous to aa 472 to 578, the region immediately C terminal to the SANT domain in S. cerevisiae Bdp1. Human Bdp1 phosphorylation was further proposed to dissociate Bdp1 from the U6 gene promoter. However, whether Bdp1 is phosphorylated by CK2 or other kinases in yeast to regulate transcription remains to be determined. In addition to Pol III transcription initiation, Bdp1 has also been implicated in the integration of the Ty1 retrotransposon near tDNA genes in Saccharomyces cerevisiae (39). Although genetic analysis linked the chromatin remodeling factor ISW2 and the N terminus of Bdp1 with Ty1 integration, the possibility of a direct protein interaction was not characterized. Recently, genome-wide binding analysis revealed that the protein arginine methyltransferase family protein Hmt1 interacts with Bdp1 (40). Hmt1 was proposed to repress tRNA transcription by methylating protein components of the Pol III machinery. The Hmt1 interaction with Bdp1 in transcription initiation was not identified in this analysis and remains to be characterized in the future.
In Pol II transcription initiation, the structural modules of transcription factors TFIIB, TFIIE, and TFIIF are localized within the polymerase active site cleft to promote DNA melting, template strand loading, transcription start site selection, and promoter escape (41–43). However, the DNA melting process is not spontaneous upon PIC formation, and an additional transcription factor, TFIIH, is required to unwind the promoter DNA with its DNA translocase function. In the Pol III system, the TFIIB-related factor Brf1 and the TFIIE- and TFIIF-related Pol III subcomplexes C82/C34/C31 and C53/C37 provide structural modules for open promoter complex formation. Although Bdp1 contains no structural regions homologous to transcription factors in Pol II, our study revealed that Bdp1 region II is a new protein module in the polymerase active center. Bdp1 region II might contribute to specific functions in Pol III transcription initiation, such as spontaneous DNA melting and template strand loading.
Supplementary Material
ACKNOWLEDGMENTS
We thank Bruce Knutson (Upstate Medical University, SUNY) for valuable comments. We thank AndreAna Peña for English editing.
This work was supported by grants MOST 100-2311-B-001-013-MY3 and 103-2311-B-001-021-MY3 from the Ministry of Science and Technology, Republic of China, and a career development award to H.-T.C. from the Academia Sinica, Taiwan, Republic of China.
Footnotes
Supplemental material for this article may be found at http://dx.doi.org/10.1128/MCB.00263-15.
REFERENCES
- 1.Dieci G, Fiorino G, Castelnuovo M, Teichmann M, Pagano A. 2007. The expanding RNA polymerase III transcriptome. Trends Genet 23:614–622. doi: 10.1016/j.tig.2007.09.001. [DOI] [PubMed] [Google Scholar]
- 2.Vannini A, Cramer P. 2012. Conservation between the RNA polymerase I, II, and III transcription initiation machineries. Mol Cell 45:439–446. doi: 10.1016/j.molcel.2012.01.023. [DOI] [PubMed] [Google Scholar]
- 3.Geiduschek EP, Kassavetis GA. 2001. The RNA polymerase III transcription apparatus. J Mol Biol 310:1–26. doi: 10.1006/jmbi.2001.4732. [DOI] [PubMed] [Google Scholar]
- 4.Schramm L, Hernandez N. 2002. Recruitment of RNA polymerase III to its target promoters. Genes Dev 16:2593–2620. doi: 10.1101/gad.1018902. [DOI] [PubMed] [Google Scholar]
- 5.Kassavetis GA, Bartholomew B, Blanco JA, Johnson TE, Geiduschek EP. 1991. Two essential components of the Saccharomyces cerevisiae transcription factor TFIIIB: transcription and DNA-binding properties. Proc Natl Acad Sci U S A 88:7308–7312. doi: 10.1073/pnas.88.16.7308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Naidu S, Friedrich JK, Russell J, Zomerdijk JC. 2011. TAF1B is a TFIIB-like component of the basal transcription machinery for RNA polymerase I. Science 333:1640–1642. doi: 10.1126/science.1207656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Knutson BA, Hahn S. 2011. Yeast Rrn7 and human TAF1B are TFIIB-related RNA polymerase I general transcription factors. Science 333:1637–1640. doi: 10.1126/science.1207699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bartholomew B, Kassavetis GA, Geiduschek EP. 1991. Two components of Saccharomyces cerevisiae transcription factor IIIB (TFIIIB) are stereospecifically located upstream of a tRNA gene and interact with the second-largest subunit of TFIIIC. Mol Cell Biol 11:5181–5189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Persinger J, Bartholomew B. 1996. Mapping the contacts of yeast TFIIIB and RNA polymerase III at various distances from the major groove of DNA by DNA photoaffinity labeling. J Biol Chem 271:33039–33046. doi: 10.1074/jbc.271.51.33039. [DOI] [PubMed] [Google Scholar]
- 10.Colbert T, Lee S, Schimmack G, Hahn S. 1998. Architecture of protein and DNA contacts within the TFIIIB-DNA complex. Mol Cell Biol 18:1682–1691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Persinger J, Sengupta SM, Bartholomew B. 1999. Spatial organization of the core region of yeast TFIIIB-DNA complexes. Mol Cell Biol 19:5218–5234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kassavetis GA, Kumar A, Letts GA, Geiduschek EP. 1998. A post-recruitment function for the RNA polymerase III transcription-initiation factor IIIB. Proc Natl Acad Sci U S A 95:9196–9201. doi: 10.1073/pnas.95.16.9196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kassavetis GA, Kumar A, Ramirez E, Geiduschek EP. 1998. Functional and structural organization of Brf, the TFIIB-related component of the RNA polymerase III transcription initiation complex. Mol Cell Biol 18:5587–5599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kassavetis GA, Nguyen ST, Kobayashi R, Kumar A, Geiduschek EP, Pisano M. 1995. Cloning, expression, and function of TFC5, the gene encoding the B″ component of the Saccharomyces cerevisiae RNA polymerase III transcription factor TFIIIB. Proc Natl Acad Sci U S A 92:9786–9790. doi: 10.1073/pnas.92.21.9786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kumar A, Kassavetis GA, Geiduschek EP, Hambalko M, Brent CJ. 1997. Functional dissection of the B″ component of RNA polymerase III transcription factor IIIB: a scaffolding protein with multiple roles in assembly and initiation of transcription. Mol Cell Biol 17:1868–1880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kassavetis GA, Bardeleben C, Kumar A, Ramirez E, Geiduschek EP. 1997. Domains of the Brf component of RNA polymerase III transcription factor IIIB (TFIIIB): functions in assembly of TFIIIB-DNA complexes and recruitment of RNA polymerase to the promoter. Mol Cell Biol 17:5299–5306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Khoo SK, Wu CC, Lin YC, Lee JC, Chen HT. 2014. Mapping the protein interaction network for TFIIB-related factor Brf1 in the RNA polymerase III preinitiation complex. Mol Cell Biol 34:551–559. doi: 10.1128/MCB.00910-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hahn S, Roberts S. 2000. The zinc ribbon domains of the general transcription factors TFIIB and Brf: conserved functional surfaces but different roles in transcription initiation. Genes Dev 14:719–730. [PMC free article] [PubMed] [Google Scholar]
- 19.Roberts S, Miller SJ, Lane WS, Lee S, Hahn S. 1996. Cloning and functional characterization of the gene encoding the TFIIIB90 subunit of RNA polymerase III transcription factor TFIIIB. J Biol Chem 271:14903–14909. doi: 10.1074/jbc.271.25.14903. [DOI] [PubMed] [Google Scholar]
- 20.Aasland R, Stewart AF, Gibson T. 1996. The SANT domain: a putative DNA-binding domain in the SWI-SNF and ADA complexes, the transcriptional co-repressor N-CoR and TFIIIB. Trends Biochem Sci 21:87–88. doi: 10.1016/S0968-0004(96)30009-1. [DOI] [PubMed] [Google Scholar]
- 21.Saida F. 2008. Structural characterization of the interaction between TFIIIB components Bdp1 and Brf1. Biochemistry 47:13197–13206. doi: 10.1021/bi801406z. [DOI] [PubMed] [Google Scholar]
- 22.Schramm L, Pendergrast PS, Sun Y, Hernandez N. 2000. Different human TFIIIB activities direct RNA polymerase III transcription from TATA-containing and TATA-less promoters. Genes Dev 14:2650–2663. doi: 10.1101/gad.836400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ishiguro A, Kassavetis GA, Geiduschek EP. 2002. Essential roles of Bdp1, a subunit of RNA polymerase III initiation factor TFIIIB, in transcription and tRNA processing. Mol Cell Biol 22:3264–3275. doi: 10.1128/MCB.22.10.3264-3275.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wu CC, Lin YC, Chen HT. 2011. The TFIIF-like Rpc37/53 dimer lies at the center of a protein network to connect TFIIIC, Bdp1, and the RNA polymerase III active center. Mol Cell Biol 31:2715–2728. doi: 10.1128/MCB.05151-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Christianson TW, Sikorski RS, Dante M, Shero JH, Hieter P. 1992. Multifunctional yeast high-copy-number shuttle vectors. Gene 110:119–122. doi: 10.1016/0378-1119(92)90454-W. [DOI] [PubMed] [Google Scholar]
- 26.Brachmann CB, Davies A, Cost GJ, Caputo E, Li J, Hieter P, Boeke JD. 1998. Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 14:115–132. doi: 10.1002/(SICI)1097-0061(19980130)14:2<115::AID-YEA204>3.0.CO;2-2. [DOI] [PubMed] [Google Scholar]
- 27.Chin JW, Cropp TA, Anderson JC, Mukherji M, Zhang Z, Schultz PG. 2003. An expanded eukaryotic genetic code. Science 301:964–967. doi: 10.1126/science.1084772. [DOI] [PubMed] [Google Scholar]
- 28.Chen HT, Warfield L, Hahn S. 2007. The positions of TFIIF and TFIIE in the RNA polymerase II transcription preinitiation complex. Nat Struct Mol Biol 14:696–703. doi: 10.1038/nsmb1272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Chen HT, Hahn S. 2003. Binding of TFIIB to RNA polymerase II: mapping the binding site for the TFIIB zinc ribbon domain within the preinitiation complex. Mol Cell 12:437–447. doi: 10.1016/S1097-2765(03)00306-X. [DOI] [PubMed] [Google Scholar]
- 30.Kassavetis GA, Driscoll R, Geiduschek EP. 2006. Mapping the principal interaction site of the Brf1 and Bdp1 subunits of Saccharomyces cerevisiae TFIIIB. J Biol Chem 281:14321–14329. doi: 10.1074/jbc.M601702200. [DOI] [PubMed] [Google Scholar]
- 31.Wu CC, Herzog F, Jennebach S, Lin YC, Pai CY, Aebersold R, Cramer P, Chen HT. 2012. RNA polymerase III subunit architecture and implications for open promoter complex formation. Proc Natl Acad Sci U S A 109:19232–19237. doi: 10.1073/pnas.1211665109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Vannini A, Ringel R, Kusser AG, Berninghausen O, Kassavetis GA, Cramer P. 2010. Molecular basis of RNA polymerase III transcription repression by Maf1. Cell 143:59–70. doi: 10.1016/j.cell.2010.09.002. [DOI] [PubMed] [Google Scholar]
- 33.Fernandez-Tornero C, Bottcher B, Rashid UJ, Steuerwald U, Florchinger B, Devos DP, Lindner D, Muller CW. 2010. Conformational flexibility of RNA polymerase III during transcriptional elongation. EMBO J 29:3762–3772. doi: 10.1038/emboj.2010.266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kassavetis GA, Letts GA, Geiduschek EP. 2001. The RNA polymerase III transcription initiation factor TFIIIB participates in two steps of promoter opening. EMBO J 20:2823–2834. doi: 10.1093/emboj/20.11.2823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Brun I, Sentenac A, Werner M. 1997. Dual role of the C34 subunit of RNA polymerase III in transcription initiation. EMBO J 16:5730–5741. doi: 10.1093/emboj/16.18.5730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Rijal K, Maraia RJ. 2013. RNA polymerase III mutants in TFIIFalpha-like C37 that cause terminator readthrough with no decrease in transcription output. Nucleic Acids Res 41:139–155. doi: 10.1093/nar/gks985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Landrieux E, Alic N, Ducrot C, Acker J, Riva M, Carles C. 2006. A subcomplex of RNA polymerase III subunits involved in transcription termination and reinitiation. EMBO J 25:118–128. doi: 10.1038/sj.emboj.7600915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hu P, Samudre K, Wu S, Sun Y, Hernandez N. 2004. CK2 phosphorylation of Bdp1 executes cell cycle-specific RNA polymerase III transcription repression. Mol Cell 16:81–92. doi: 10.1016/j.molcel.2004.09.008. [DOI] [PubMed] [Google Scholar]
- 39.Bachman N, Gelbart ME, Tsukiyama T, Boeke JD. 2005. TFIIIB subunit Bdp1p is required for periodic integration of the Ty1 retrotransposon and targeting of Isw2p to S. cerevisiae tDNAs. Genes Dev 19:955–964. doi: 10.1101/gad.1299105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Milliman EJ, Hu Z, Yu MC. 2012. Genomic insights of protein arginine methyltransferase Hmt1 binding reveals novel regulatory functions. BMC Genomics 13:728. doi: 10.1186/1471-2164-13-728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.He Y, Fang J, Taatjes DJ, Nogales E. 2013. Structural visualization of key steps in human transcription initiation. Nature 495:481–486. doi: 10.1038/nature11991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Murakami K, Elmlund H, Kalisman N, Bushnell DA, Adams CM, Azubel M, Elmlund D, Levi-Kalisman Y, Liu X, Gibbons BJ, Levitt M, Kornberg RD. 2013. Architecture of an RNA polymerase II transcription pre-initiation complex. Science 342:1238724. doi: 10.1126/science.1238724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Sainsbury S, Niesser J, Cramer P. 2013. Structure and function of the initially transcribing RNA polymerase II-TFIIB complex. Nature 493:437–440. doi: 10.1038/nature11715. [DOI] [PubMed] [Google Scholar]
- 44.Kostrewa D, Zeller ME, Armache KJ, Seizl M, Leike K, Thomm M, Cramer P. 2009. RNA polymerase II-TFIIB structure and mechanism of transcription initiation. Nature 462:323–330. doi: 10.1038/nature08548. [DOI] [PubMed] [Google Scholar]
- 45.Juo ZS, Kassavetis GA, Wang J, Geiduschek EP, Sigler PB. 2003. Crystal structure of a transcription factor IIIB core interface ternary complex. Nature 422:534–539. doi: 10.1038/nature01534. [DOI] [PubMed] [Google Scholar]
- 46.Fernandez-Tornero C, Moreno-Morcillo M, Rashid UJ, Taylor NM, Ruiz FM, Gruene T, Legrand P, Steuerwald U, Muller CW. 2013. Crystal structure of the 14-subunit RNA polymerase I. Nature 502:644–649. doi: 10.1038/nature12636. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.