Abstract
The N-methyl-D-aspartate (NMDA) receptor NR2B subunit on neurons in the anterior cingulate cortex (ACC) is implicated in the affective response to noxious stimuli. Selectively silencing this NR2B subunit in ACC neurons could therefore alleviate pain-related aversion. However, to date, there is no optimal approach to selectively silence the NR2B gene in ACC neurons. In the present study, we constructed lentiviral vectors and delivered shRNA (NR2B-RNAi-LV) to effectively silence the NR2B gene in ACC neurons. The use of lentivirus resulted in 95% transfection efficiency and 83% silencing of the NR2B gene in ACC neurons. Electrophysiological experiments showed that the total INMDA was similarly reduced by 48% in lentivirus-transfected ACC neurons. The biochemical and functional data demonstrated that lentiviral shRNA delivery produced a high transfection and silencing efficiency in the ACC neurons. SNI rats weighting 220-250 g were randomly divided into three groups: normal saline group (NS), lenti-siRNA/NC (LV-NC) group, and lenti-siRNA/NR2B (LV-NR2B) group, and conditioned place avoidance was conducted. The results indicated that NR2B-RNAi-LV decreased greatly the conditioning scores of F-CPA while NC-GFP-LV has no effects. NR2B mRNA expression in the NR2B-RNAi-LV group was significantly lower than that in the control group and NC-GFP-LV group. This novel approach of silencing the NR2B gene in ACC neuron could potentially be used to alleviate pain-related aversion.
Keywords: RNA interference, NR2B, ACC, gene silencing
Introduction
Anterior cingulate cortex (ACC) neurons are involved in emotional processing of pain, making it an attractive target for therapeutic intervention on pain-related aversion [1-3]. Within ACC regions, excitatory amino acids such as glutamate, were involved in synaptic transmission and signal transduction [4,5]. Previous studies about formalin-conditioned place avoidance (F-CPA), a protocol where nociceptive stimulation produces aversive learning, suggested that glutamatergic activation of rostral-ACC (rACC) neurons was necessary for the acquisition of CPA. The research of Lei et al further confirmed that the blocking of the N-methyl-D-aspartate (NMDA) receptors in rACC brain regions, but not AMPA receptor, significantly inhibited F-CPA expression and attenuated F-CPA retrieval-induced Fos expression in the ACC [6]. It is therefore suggested that the generation and maintenance of pain-related emotion is mediated by the activation of NMDA receptors in the ACC.
Functional NMDA receptors consist of NR1 subunit and one or more NR2A-D subunits [7,8], of which the NR2B subunits are highly expressed in forebrain [9]. It has been shown that both NR2A and NR2B subunits contribute to the formation of long-term potentiation in the ACC [10]. Ren et al. have also demonstrated that the glycine site in the NR1 subunit of NMDA receptors in the rostral ACC is involved in the production of F-CPA [11]. Interestingly, the blockade of the NR2B subunit of NMDA receptors in the rACC with Ifenprodil or Ro 25-698 completely blocked the F-CPA acquisition. Based on these findings, we proposed that NMDA NR2B subunits might be involved in the formation of pain-related negative affect, making them ideal therapeutic targets.
Although most NMDA receptor antagonists have therapeutic effects, the clinically effective dosage is easy to cause side effects due to their lower therapeutic efficiency. Thus, the initial enthusiasm for NMDA receptor antagonists has been reduced. Alternatively, selectively silencing the NR2B gene in ACC neurons may be a potential treatment of pain-related negative affect, which has not been successfully carried out yet. The present study was the first to demonstrate the selective silencing of the NR2B gene (NR2B RNAi LV) in ACC neurons via lentiviral construction and delivery of shRNA.
Materials and methods
Animal protocol
All procedures involving the use of animals were conducted in compliance with the guidelines of the National Institutes of Health and were approved by the Animal Care and Use Committee of the School of Medicine of Shandong University. The study adhered to the Ethical Guidelines of the International Association for the Study of Pain.
Animals and spinal nerve ligation
Male Sprague-Dawley rats (250-300 g) were purchased from the Experimental Animal Center of the Shandong University. All animals were raised at a controlled temperature of 20±0.5°C in cages and maintained on a 12 h light/dark cycle, with free access to food and water. Rats were allowed to acclimate for 1 week after arrival.
The surgical procedure for L5/L6 spinal nerve ligation was performed [12]. Sham-operated control rats were prepared in an identical manner except that the L5/L6 spinal nerves were not ligated. The behavior of the rats was monitored carefully for any visual indication of motor disorders or change in weight or general health.
Culture and identification of ACC neurons
Primary ACC cultures were prepared from 1 to 2 day old Sprague-Dawley rats according to a previously described procedure with some modification. Briefly, the ACCs were digested for 30 min by the trypsin (0.125%, Sigma, St. Louis, MO) and the digestion was stopped by additional calf serum. The digested tissue was passed through a 100 mesh sieve. Cells in suspension were collected by centrifugation at 200 g for 8 min, washed three times with a standard culture medium, inoculated (at density 1.5-1.8×106/ml) in a neurobasal medium (consisting of medium with a B27 supplement, 10% fetal bovine serum, 1% glutamine [Gibco, Grand Island, NY], 100 U/ml penicillin, and 100 mg/ml streptomycin, pH 7.2-7.4), incubated 24 h, and then treated with 0.1 mg/ml of cytosine arabinoside (Sigma) to inhibit the proliferation of non-neuronal cells. The growth medium was changed twice per week.
The cells cultured on day 7 (7 DIV) were identified by the anti-MAP2 monoclonal antibody (Neomarkes). MAP2 is specifically expressed in the neuron but minimally expressed in the astrocyte and oligodendrocyte. Therefore, a neuron is identified by positive MAP2 expression while an astrocyte is identified by positive GFAP expression.
Design and construction of plasmid pFU-GW-iRNA-NR2B
The siRNA specifically targeting the NR2B gene was purchased from Shanghai Jikai Gene Chemical Co., Ltd. (Silencer Pre-designed siRNA; Shanghai, China). The oligonucleotides were selected by Blast homology search from cDNA sequences of target genes of various species including mouse, rat, and human. Oligonucleotides were designed to 50 phosphates, synthesized by Integrated DNA Technologies, Inc. (Coralville, IA), and purified by polyacrylamide gel electrophoresis. The oligo format was as follows: Sense Oligo: 5’ T-(19nt)-TTCAAGAGA)-(91nt)-TTTTTTC-3’ [Sense siRNA-Loop-Antisense siRNA-Stop]. Anti-sense oligo: complement of the sense oligo but with additional nucleotides at the 5’ end to form an XhoI site.
The sense and anti-sense oligo were resuspended in water at a concentration of 60 pmol/μl and annealed in annealing buffer (100 nM K-acetate, 30 nM HEPES-KOH, pH 7.4, 2 mM Mg-acetate) by incubating at 95°C for 4 min, 70°C for 10 min, and then slowly decreasing temperature till 4°C (0.1°C/min). The annealed oligo was subcloned into the Hpa I/Xho I site of the pFU-GW-iRNA vector which encodes the cytomegalovirus promoter driving eGFP (enhanced green fluorescent protein) marker expression (the internal control). The resulting lentiviral siRNA vector was confirmed by restriction enzyme digestion with Hpa I/Xho I, showing a 343 bp band shift on 2% agarose gel when compared with the parental vector pFU-GW-iRNA. The constructed vector was also confirmed by DNA sequencing with primers (5’-GCCCCGGTTAATTTGCATAT-3’ [forward] and 5’-GAGGCCAGATCTTGGGTG-3’ [reverse]).
The inhibitory effect of plasmid pFU-GW-iRNA-NR2B on NR2B protein expression was measured by co-transfecting 293T cells with pFU-GW-iRNA-NR2B and the plasmid expressing NR2B protein, collecting the cells 48 h later, isolating total protein, and visualizing NR2B protein expression on Western blots.
Lentivirus (NR2B-RNAi-LV) packaging
Packaging, purification, and titer determination of the lentivirus (NR2B-RNAi-LV) were performed [14,15]. The recombinant viruses were produced by Ca phosphate transfection of HEK 293T cells using standard protocols [16]. Briefly, 293T cells were cultured in Dulbecco’s modified Eagles’s medium supplemented with 10% fetal bovine serum and penicillin/streptomycin (100 U/ml). At around 70% culture confluency in a 10 cm culture dish, the cells were co-transfected with the plasmid pFU-GW-iRNA-NR2B (10 μg) and the NR2B-expression vector pcDNA3.1-NR2B (2 μg). The viruses were collected from the culture supernatant 48 h post-transfection, which was concentrated by ultracentrifugation for 2 h at 1200× g, and resuspended in phosphate-buffered saline (PBS). Titers were determined by infecting 293T cells with serial dilutions of concentrated lentivirus, incubating them for 48 h, and counting eGFP positive cells under a fluorescence microscope.
Lentiviral (NR2B-RNAi-LV) transduction in primary cultured ACC neurons
To determine the NR2B-RNAi-LV transduction efficiency in ACC neurons, GFP expression was examined by microscopy at different multiplicities of infection (MOIs) on day 3 and day 4 after infection.
Isolation of RNA and real-time RT-PCR
Total cellular RNA was extracted from neurons on day 3 post-transfection using Trizol reagent (Invitrogen, Carlsbad, CA), and reversely transcribed into cDNA synthesis using a reverse transcriptase (RT) premix. The amount of transcripts in the cDNA samples was measured by RT-PCR with primers (Table 1).
Table 1.
Primer sequences
| Gene | Primer sequences |
|---|---|
| NR2B | Forward: 5-AATGGCGGATAAGGATGAGT-3 |
| Reverse: 5-GGGAAGTAGGTGGTGACGAT-3 | |
| β-actin | Forward: 5-CCCATCTATGAGGGTTACGC-3 |
| Reverse: 5-TTTAATGTCAXGCACGATTTC-3 |
Protein extraction and western blotting
Cells were collected on day 3 post-transfection. Total protein was prepared as previously described [16]. Neonatal ACC neurons were lysed in RIPA buffer (50 mM Tris-Cl, pH 7.4, 150 mM NaCl, 1% NP40, 0.25% Na-deoxycholate, 1 mM phenylmethylsulfonyl fluoride [PMSF]) with protease inhibitors (in mmol/L: leupeptin 0.1 and PMSF 1.0) and kept for 30 min on ice with vortexing every 5 min. The supernatant was collected after centrifugation at 400 g for 15 min at 4°C. The protein concentration was measured for three times using a Bio-Rad DC protein assay kit. The same amount of membrane protein (50 μg) was loaded onto each lane of standard 4-12% SDS-polyacrylamide gels. After electrophoresis, proteins were transferred to a nitrocellulose membrane, which was blocked in PBS containing 0.1% Tween and 5% non-fat milk, incubated overnight at 4°C in 1:200 primary anti-NR2B channel antibody (Abcam, Cambridge, England), incubated with horseradish peroxidase-labeled goat anti-rabbit IgG (Chemicon, Temecula, CA) in PBS-Tween as a secondary antibody (1:2000) for 1 h, followed by enhanced chemiluminescence (ECL; Amersham, Little Chalfont, UK) for immunoblot development. The density of protein bands was quantified using NIH image software.
Patch-clamp recording of NMDA-activated currents in ACC neurons following lentiviral (NR2B-RNAi-LV) transduction
Current responses to NMDA were evoked in cultured ACC neurons (on day 4 or day 5 after successful transfection). Patch pipettes had resistances of 5-8 MΩ when filled with the following solution (in mmol/L): cesium methane sulfonate (CH3O3SCs) 135, NaCl 8, HEPES 10, EGTA 0.5, Mg-ATP 4, and Na-GTP 0.3 (adjusted to pH 7.2 with 1 mol/L CsOH, and adjusted to 300-310 mOsm with sucrose). NMDA receptor-mediated currents (INMDA) were induced in the following bath solution (in mmol/L): NaCl 150, KCl 3, HEPES 10, CaCl2 × 2H2O 3, and glucose 8 (adjusted to pH 7.4 with 1 mol/L NaOH, and adjusted to 300-310 mOsm with sucrose) and recorded by using a Multi-Clamp 700 A amplifier, Digidata 1320 A AD converter and pCLAMP 8.0 software (Axon Instruments, Foster City, CA). The junction potential between micropipette and bath solution was -9.9 mV, as calculated by using Clampex 8.0 (in pCLAMP 8.0) and nulled just before the formation of a gigaseal. In most experiments, the series resistance (Rs) before compensation was 10-20 MΩ. Routinely, 70-80% of the Rs was compensated. Only neurons with an Rs <20 MΩ were selected for further tests.
All NMDA currents were recorded while the membrane potential was held at -60 mV, except as otherwise noted. Bath application of NMDA in cultured neurons was performed via a perfusion system (MPS-2; World Precision Instruments, Sarasota, FL) at an ambient temperature of 23-25°C. The tip (diameter 250 μm) of the drug application pipette was usually placed 100 μm away from the recorded cell. To avoid drug accumulation in the bath, each 2 s pulse of drug was followed by a 2 min rinsing.
Experimental grouping
SNI rats weighting 220-250 g were randomly divided into three groups: normal saline group (NS), lenti-siRNA/NC (LV-NC) group, and lenti-siRNA/NR2B (LV-NR2B) group. Rats in each group were intra-rACC administered with NS, lenti-siRNA/NC or lenti-siRNA/NR2B in a total volume of 0.6 μl per hemisphere.
CPA
CPA was conducted [13]. Place conditioning apparatus consists of three opaque acrylic compartments (a neutral compartment and two conditioning compartments with distinctive visual, olfactory and tactile cues) with removable doors to allow room isolation when necessary. The experimental process consists of three distinct sessions: a pre-conditioning session (day 1 and day 2), conditioning session (day 3), and post-conditioning session (day 4). On day 1, rats were individually placed in the neutral compartment and were allowed to explore the two conditioning compartments freely for 15 min so that they could get habituated to the apparatus. The time spent in each of the compartments was automatically recorded by a timer with a blind method. On day 2, the same trial was performed and the time spent in each compartment was measured. Rats that spent more than 80% (720 s) on one side on day 2, or that spent more than 600 s one side on day 1 and more than 600 s on the other side on day 2 were eliminated from the experiment. On day 3, place conditioning was performed. In the morning, rats received nothing, and were randomly confined in one of the conditioning compartments for 45 min. After at least 4 h, rats received a noxious stimulus (i.pl. 5% formalin 50 μl) or a control treatment (i.pl. NS), and were then confined in the other conditioning compartment for 45 min in formalin experiments. In the electric foot-shock-induced CPA (S-CPA) experiments, rats received an electric shock (0.5 mA for 2 s) every 8-10 min in the other conditioning compartment during the 45 min training session. On day 4, the same trial as on day 1 or 2 was performed. The time spent in each compartment was measured.
Statistical analysis
Data were presented as means ± SD. The results were evaluated by independent-sample T test, one-way analysis of variance, followed by the Student-Newman-Keuls test for multiple comparisons using SPSS13.0 software (SPSS Inc., Chicago, IL). A P value <0.05 was considered statistically significant.
Results
Morphological observation and identification of cultured ACC neurons
At 24 h after inoculation, neonatal rat ACC neurons were seen that most of the cells had attached to the wall of the culture flask, and even a few of cells began to grow one or two processes under the inverted phase contrast microscope. After 2 days of culture (2 DIV), cells grew rapidly and the cell bodies of neurons and length of the processes increased further. At 5 DIV, more processes sprouted and began to form clusters, and the neuronal bodies kept growing bigger. At 7 DIV the processes extended and branched further to form a dense web and various synapses between cells began to form. The neurons appeared no significant morphological changes even after two weeks in culture (Figure 1).
Figure 1.
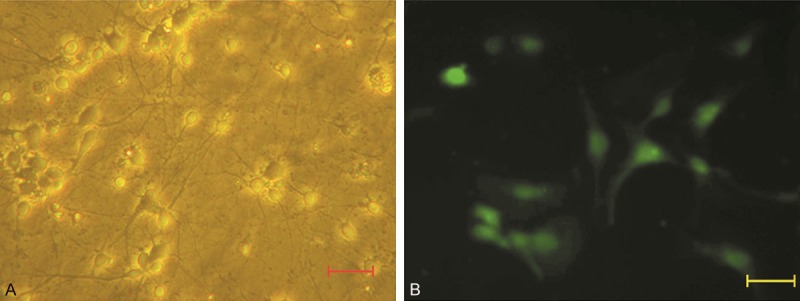
Culture and identification of ACC neurons. A. ACC neurons on day 7 of culture in vitro (7 DIV). The clear pale halo was around each neurons. Neuronal processes mutually interconnected into a network. B. Identification of neurons by immunofluorescence staining. More than 95% of cultured neurons were MAP2 positive at 7 DIV. (scale bar: 50 μm).
At 7 DIV, immunofluorescence staining with anti-MAP2 monoclonal antibody demonstrated that more than 95% of cells were MAP2 positive (Figure 1).
Design and construction of NR2B-RNAi-LV vectors
The siRNA specifically targeting the NR2B gene was synthesized and then subcloned into the multiple cloning sites of lentiviral transfer vector pFU-GW-iRNA. The insertion of NR2B-shRNA was confirmed by PCR, double-enzyme digestion (Figure 2), and gene sequencing (data not shown). Western blotting showed that more than 95% of the NR2B protein expressed by the plasmid delivered NR2B gene was silenced by using the plasmid pFU-GW-iRNA-NR2B, which indicated that plasmid pFU-GW-iRNA-NR2B could efficiently silence NR2B protein expressed by the plasmid delivered NR2B gene (Figure 2).
Figure 2.
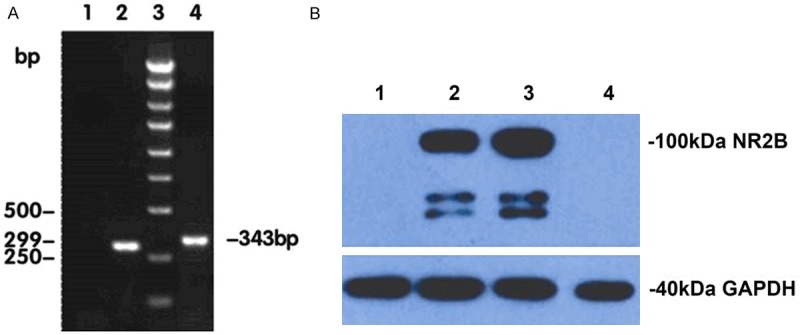
Design and construction of pFU-GW-iRNA-NR2B. A. The identification of plasmid pFU-GW-iRNA-NR2B by PCR analysis. Lane 1: negative control (ddH2O); Lane 2: negative control (pFU-GW-iRNA); Lane 3: 250-bp DNA ladder; Lane 4: PCR analysis of the recombinant plasmid pFU-GW-iRNA-NR2B. B. Inhibitory effects of the plasmid pFU-GW-iRNA-NR2B on NR2B protein expression were observed by Western blotting. GAPDH protein was used as a control. Lane 1: 293T cells; Lane 2: plasmid expressing NR2B protein; Lane 3: plasmid expressing NR2B protein and the plasmid pFU-GW-iRNA; Lane 4: plasmid expressing NR2B protein and the plasmid pFU-GW-iRNA-NR2B.
Transduction efficiency of NR2B-RNAi-LV vector in ACC neurons
To determine the efficiency of NR2B-RNAi-LV transduction (assessed on days 3-4 after infection [MOI 3]), GFP expression was examined under a microscope. Up to 95% of ACC neurons were successfully transduced with NR2B-RNAi-LV (Figure 3).
Figure 3.
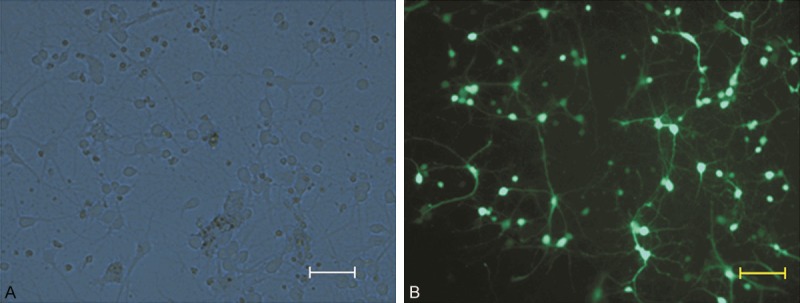
Determination of NR2B-RNAi-LV vector transduction efficiency in ACC neurons. GFP expression was observed under a light microscope (A) or fluorescence microscope (B). The increase in GFP expression in ACC neurons peaked when the MOI was 3 (scale bar: 50 μm).
Inhibition of NR2B mRNA expression by NR2B-RNAi-LV
The mRNA expression of NR2B gene, specifically inhibited by NR2B shRNA in ACC neurons, was analyzed by quantitative RT-PCR. ACC neurons were infected by lentivirus containing NR2B-RNAi (NR2B-RNAi-LV) or the scrambled control (NC-GFP-LV), respectively. We found that NR2B mRNA expression was down-regulated by almost 83% in NR2B-RNAi-LV-infected ACC neurons compared with ACC neurons exposed to normal saline or NC-GFP-LV (Figure 4), but which was similar between ACC neurons exposed to normal saline or NC-GFP-LV. These results suggest that the NR2B-RNAi-LV was a highly efficient gene-silencer.
Figure 4.

NR2B-RNAi-LV transfection silenced the expression of NR2B mRNA in ACC neurons. These data were analyzed by the relative gene expression (2-ΔΔCt) method. The figures show that NR2B-RNAi-LV decreased significantly NR2B mRNA expression in neurons (# vs. control group, P<0.05) or negative transfection (* vs. NC-GFP-LV group, P<0.05).
Knockdown of NR2B protein expression by NR2B-RNAi-LV vectors
The effect of NR2B-RNAi-LV on NR2B protein expression on day 3 post-transfection was evaluated by western blot (Figure 5). Levels of NR2B protein were similar in the untransduced group and NC-GFP-LV group (P>0.05), but significantly reduced in the NR2B-RNAi-LV group (P<0.05), showing that transfection with NR2B shRNA significantly decreased the expression of the NR2B protein in ACC neurons.
Figure 5.
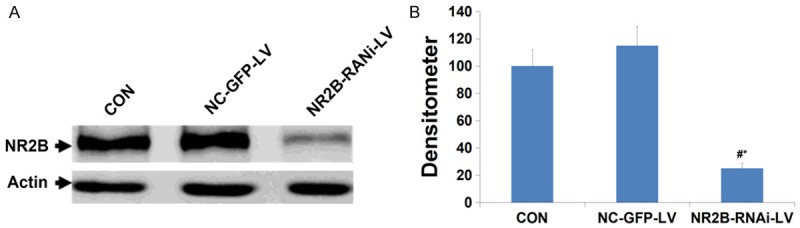
The levels of NR2B protein in NR2B-RNAi-LV-transduced cells was significantly lower than that in NC-GFP-LV-transduced or Control cells on 72 h after treatment. CON = ACC neurons uninfected with lentivirus; NC-GFP-LV = ACC neurons infected with control lentivirus; NR2B-RNAi-LV = ACC neurons infected with lentivirus carrying NR2B shRNA. A and B show the protein level measured by Western blotting and band densitometry, respectively. # vs. control group, P<0.05, * vs. NC-GFP-LV group, P<0.05.
Attenuation of NMDA-activated currents in ACC neurons by the NR2B-RNAi-LV vector
Using whole cell patch clamp recording technique, it was examined for the effects of NR2B gene knockout on NMDA receptor function in ACC neurons. Figure 6A shows representative INMDA traces measured at -60 mV. Silencing of NR2B gene expression in ACC neurons reduced NMDA-induced current to 52% of its control level. Thus, we demonstrated an optimal approach for silencing of NR2B gene expression by NR2B-RNAi-LV vectors in ACC neurons at both mRNA and protein levels (Figure 6).
Figure 6.

Functional evidence of silencing of NR2B gene expression in ACC neurons provided by whole cell patch-clamp recordings. A. Representative traces of the peak NMDA current elicited in control cells by 10 μmol/L NMDA and after silencing of the NR2B gene. The peak NMDA current was significantly decreased in silenced ACC neurons. B. The averaged data from patch-clamp experiments. Knockdown of NR2B protein expression by NR2B-RNAi-LV vectors reduced NMDA responses to 52% of that in control neurons. # vs. control group, P<0.05, * vs. NC-GFP-LV group, P<0.05.
Reduction of NR2B within rACC neurons by NR2B-RNAi-LV alleviates pain-related aversion
In CPA experiments, the CPA scores represented the time spent in the treatment-paired compartment on the pre-conditioning day (day 1) minus the time spent in the same conditioning compartment on the post-conditioning day (day 4), which indicated the index of emotional pain.
For NC-GFP-LV group and Control group, the time spent in the formalin-paired compartment was 868.30±13.2 s pre-conditioning vs. 115.83±21.3 s post-conditioning (independent-sample t test, t=7.804, P<0.05, n=10), 967.05±12.2 s preconditioning vs. 135.50±21.3 s post-conditioning (independent-sample t test, t=17.844, P<0.05, n=10), indicating that rats spent less time in formalin-paired compartment. The conditioning score was 752.47±24.6 s and 831.55±12.3 s. The times spent in the formalin-paired compartment were 1060.78±19.8 s for pre-conditioning, 1035.74±29.7 s for post-conditioning in NR2B-RNAi-LV group. The conditioning scores were 124.45±23.3 s. The conditioning scores of rats in the Control groups and NC-GFP-LV groups were similar (Independent-sample T test, t=0.738, P>0.05, n=10), but reduced significantly in NR2B-RNAi-LV group (independent-sample t test, t=7.815, P<0.05, n=10; t=9.477, P<0.05, n=10). The results indicated that NR2B-RNAi-LV decreased greatly the conditioning scores of F-CPA while NC-GFP-LV has no effects (Figures 7 and 8).
Figure 7.
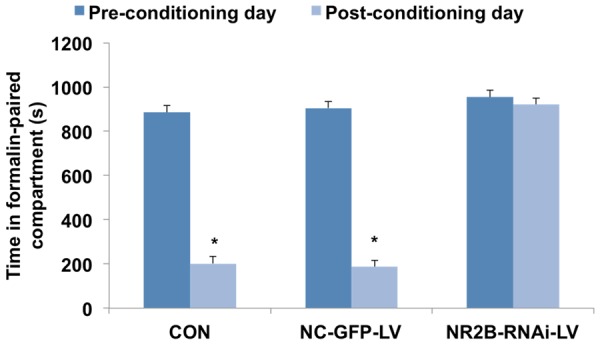
Comparison of the time spent in the formalin-paired compartment among three groups. For NC-GFP-LV group and Control group rats, the time spent in the formalin-paired compartment was 868.30±13.2 s preconditioning vs. 115.83±21.3 s post-conditioning (independent-sample t test, t=7.804, P<0.05, n=10), 967.05±12.2 s preconditioning vs. 135.50±21.3 s post-conditioning (independent-sample t test, t=17.844, P<0.05, n=10) was 868.30±13.2 s preconditioning vs. 115.83±21.3 s post-conditioning (independent-sample t test, t=7.804, P<0.05, n=10), 967.05±12.2 s preconditioning vs. 135.50±21.3 s post-conditioning (Independent-sample T test, t=17.844, P<0.05, n=10). *P<0.05 compared with pre-conditioning day.
Figure 8.

Comparison of the conditioning score among three groups. The conditioning score was 752.47±24.6 s and 831.55±12.3. The times spent in the formalin-paired compartment were 1060.78±19.8 s for preconditioning, 1035.74±29.7 s for post-conditioning in NR2B-RNAi-LV group. The conditioning scores were 124.45±23.3 s. The conditioning scores of rats in the Control groups and NC-GFP-LV groups were similar (independent-sample t test, t=0.738, P>0.05, n=10), but reduced significantly in NR2B-RNAi-LV groups (independent-sample t test, t=7.815, P<0.05, n=10; t=9.477, P<0.05, n=10). *P<0.05 compared with CON group, #P<0.05 compared with NC-GFP-LV group.
Analysis of NR2B mRNA expression in the ACC
ACC NR2B mRNA expression differed significantly among three groups. NR2B mRNA expression in the NR2B-RNAi-LV group was significantly lower than that in the control group and NC-GFP-LV group (independent-sample t test, t=24.149, P<0.05; t=17.976, P<0.05) (Figure 9).
Figure 9.
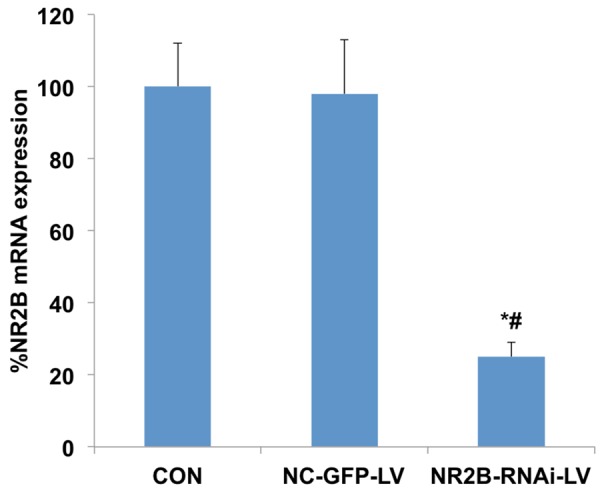
ACC NR2B mRNA expression among three groups. NR2B mRNA expression in the NR2B-RNAi-LV groups was significantly lower than levels in the control groups and NC-GFP-LV groups (independent-sample t test, t=24.149, P<0.05; independent-sample t test, t=17.976, P<0.05). *P<0.05 compared with CON group, #P<0.05 compared with NC-GFP-LV group.
Discussion
This study is the first to establish an optimal method of selectively silencing the NR2B gene in ACC neurons by transfecting them with NR2B-RNAi-LV vectors. The results demonstrated that shRNA could mediate selective silence of NR2B expression in primary-cultured rat neonatal ACC neurons. The use of lentivirus to mediate delivery of shRNA achieved 95% transfection efficiency and 83% silencing of NR2B at the mRNA and protein level. Whole cell patch-clamp electrophysiology further demonstrated that silence of NR2B gene significantly attenuated the post-synaptic NMDA-mediated responses of ACC neurons.
The present study selected the lentiviral vector-mediated gene delivery system to silence the NR2B gene on ACC neurons. It is well-known that an RNAi technique is more effective than antisense oligonucleotides or ribozymes in the specific knockdown of target mRNA, which is widely applied to various studies such as human gene function research, signal transduction research, and gene therapy [17,18]. And some studies showed that RNAi could be produced in a variety of ways, such as small interfering RNAs, plasmids expressing small hairpin RNAs transiently transfecting mammalian cells or selecting transfected or infected cells that have stably incorporated the shRNA plasmid or retrovirus into the genome [17,19-21].
The lentiviral vector-mediated gene delivery system used in the present study, compared with traditional transfection methods such as lipofection and electroporation, has the following advantages, such as effectively transfecting non-dividing cells (neurons, etc), the ability of a long-term stable expression of RNAi in host cells and low toxicity to the host cells [22,23]. Thus, we eventually selected the lentivirus-mediated RNAi system targeting NR2B gene expression to efficiently down regulate NR2B expression in ACC neurons.
A number of studies showed that NMDA receptor in ACC neurons, but not AMPA/KA receptor, mediated pain-related mood processing and the upregulation of NR2B enhanced pain-emotion production. It is therefore implicated that inhibiting NR2B expression in ACC neurons is a new therapeutic strategy. In the present study, we successfully used lentivirus-mediated RNAi to knockdown NR2B expression in ACC neurons. This work provided a foundation for future in vivo studies in which RNAi knock-down of NR2B expression in ACC will be directly evaluated as a pain-related aversion treatment.
Using whole-cell patch clamp electrophysiology, the effect of gene silencing on NMDA-evoked currents were examined. The results showed that following NR2B-RNAi-LV transfection, the NMDA-evoked currents in ACC neurons decreased significantly. The results suggested that silencing of NR2B gene by NR2B-RNAi-LV transfection decreased the excitability of ACC neurons. Besides, results of this in vivo study indicated that NR2B-RNAi-LV decreased greatly the conditioning scores of F-CPA while NC-GFP-LV has no effects and that NR2B mRNA expression in the NR2B-RNAi-LV group was significantly lower than that in the control group and NC-GFP-LV group, which suggested that the silencing of NR2B gene by NR2B-RNAi-LV transfection might be involved in the formation of pain-related negative affects.
In conclusion, the novel approach reported in this study will allow further investigation into the role of the NR2B NMDA receptor as target in pain processing, and represents significant progress in the potential treatment of pain-related aversion.
Acknowledgements
We are very grateful for the great help and technical support provided by the Department of Laboratory Medicine, Shandong Provincial Hospital, Shandong University. This work was supported by a grant from the National Natural Science Foundation of China (30872433, 81171171, 81100802) and a grant from the Science and Technology Foundation of Shandong Province (2014GSF118160).
Disclosure of conflict of interest
None.
References
- 1.Koyama T, Tanaka YZ, Mikami A. Nociceptive neurons in the macaque anterior cingulate activate during anticipation of pain. Neuroreport. 1998;9:2663–2667. doi: 10.1097/00001756-199808030-00044. [DOI] [PubMed] [Google Scholar]
- 2.Gao YJ, Ren WH, Zhang YQ, Zhao ZQ. Contributions of the anterior cingulate cortex and amygdala to pain-and fear-conditioned place avoidance in rats. Pain. 2004;110:343–353. doi: 10.1016/j.pain.2004.04.030. [DOI] [PubMed] [Google Scholar]
- 3.Johansen JP, Fields HL, Manning BH. The affective component of pain in rodents: direct evidence for a contribution of the anterior cingulate cortex. Proc Natl Acad Sci U S A. 2001;98:8077–8082. doi: 10.1073/pnas.141218998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Higashi H, Tanaka E, Nishi S. Synaptic responses of guinea pig cingulate cortical neurons in vitro. J Neurophysiol. 1991;65:822–833. doi: 10.1152/jn.1991.65.4.822. [DOI] [PubMed] [Google Scholar]
- 5.Wei F, Wang GD, Kerchner GA, Kim SJ, Xu HM, Chen ZF, Zhuo M. Genetic enhancement of inflammatory pain by forebrain NR2B overexpression. Nat Neurosci. 2001;4:164–169. doi: 10.1038/83993. [DOI] [PubMed] [Google Scholar]
- 6.Lei LG, Sun S, Gao YJ, Zhao ZQ, Zhang YQ. NMDA receptors in the anterior cingulate cortex mediate pain-related aversion. Exp Neurol. 2004;189:413–421. doi: 10.1016/j.expneurol.2004.06.012. [DOI] [PubMed] [Google Scholar]
- 7.Hollmann M, Boulter J, Maron C, Heinemann S. Molecular biology of glutamate receptors. Potentiation of N-methyl-D-aspartate receptor splice variants by zinc. Ren Physiol Biochem. 1994;17:182–183. [PubMed] [Google Scholar]
- 8.Nakanishi N, Axel R, Shneider NA. Alternative splicing generates functionally distinct N-methyl-D-aspartate receptors. Proc Natl Acad Sci U S A. 1992;89:8552–8556. doi: 10.1073/pnas.89.18.8552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Monyer H, Burnashev N, Laurie DJ, Sakmann B, Seeburg PH. Developmental and regional expression in the rat brain and functional properties of four NMDA receptors. Neuron. 1994;12:529–540. doi: 10.1016/0896-6273(94)90210-0. [DOI] [PubMed] [Google Scholar]
- 10.Zhao MG, Toyoda H, Lee YS, Wu LJ, Ko SW, Zhang XH, Jia YH, Shum F, Xu H, Li BM. Roles of NMDA NR2B subtype receptor in prefrontal long-term potentiation and contextual fear memory. Neuron. 2005;47:859–872. doi: 10.1016/j.neuron.2005.08.014. [DOI] [PubMed] [Google Scholar]
- 11.Ren WH, Guo JD, Cao H, Wang H, Wang PF, Sha H, Ji RR, Zhao ZQ, Zhang YQ. Is endogenous d-serine in the rostral anterior cingulate cortex necessary for pain-related negative affect? J Neurochem. 2006;96:1636–1647. doi: 10.1111/j.1471-4159.2006.03677.x. [DOI] [PubMed] [Google Scholar]
- 12.Kim SH, Chung JM. An experimental model for peripheral neuropathy produced by segmental spinal nerve ligation in the rat. Pain. 1992;50:355–363. doi: 10.1016/0304-3959(92)90041-9. [DOI] [PubMed] [Google Scholar]
- 13.Ren WH, Guo JD, Cao H, Wang H, Wang PF, Sha H, Ji RR, Zhao ZQ, Zhang YQ. Is endogenous D-serine in the rostral anterior cingulate cortex necessary for pain-related negative affect? J Neurochem. 2006;96:1636–1647. doi: 10.1111/j.1471-4159.2006.03677.x. [DOI] [PubMed] [Google Scholar]
- 14.Rubinson DA, Dillon CP, Kwiatkowski AV, Sievers C, Yang L, Kopinja J, Rooney DL, Zhang M, Ihrig MM, McManus MT. A lentivirus-based system to functionally silence genes in primary mammalian cells, stem cells and transgenic mice by RNA interference. Nat Genet. 2003;33:401–406. doi: 10.1038/ng1117. [DOI] [PubMed] [Google Scholar]
- 15.Follenzi A, Naldini L. Generation of HIV-1 derived lentiviral vectors. Methods Enzymol. 2002;346:454–465. doi: 10.1016/s0076-6879(02)46071-5. [DOI] [PubMed] [Google Scholar]
- 16.Qu Y, Karnabi E, Chahine M, Vassalle M, Boutjdir M. Expression of skeletal muscle Na V 1.4 Na channel isoform in canine cardiac Purkinje myocytes. Biochem Biophys Res Commun. 2007;355:28–33. doi: 10.1016/j.bbrc.2007.01.101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Dillon CP, Sandy P, Nencioni A, Kissler S, Rubinson DA, Van Parijs L. RNAi as an experimental and therapeutic tool to study and regulate physiological and disease processes. Annu Rev Physiol. 2005;67:147–173. doi: 10.1146/annurev.physiol.67.040403.130716. [DOI] [PubMed] [Google Scholar]
- 18.Cullen BR. Induction of stable RNA interference in mammalian cells. Gene Ther. 2006;13:503–508. doi: 10.1038/sj.gt.3302656. [DOI] [PubMed] [Google Scholar]
- 19.Sandy P, Ventura A, Jacks T. Mammalian RNAi: a practical guide. Biotechniques. 2005;39:215. doi: 10.2144/05392RV01. [DOI] [PubMed] [Google Scholar]
- 20.Brummelkamp TR, Bernards R, Agami R. A system for stable expression of short interfering RNAs in mammalian cells. Science. 2002;296:550–553. doi: 10.1126/science.1068999. [DOI] [PubMed] [Google Scholar]
- 21.Paddison PJ, Caudy AA, Hannon GJ. Stable suppression of gene expression by RNAi in mammalian cells. Proc Natl Acad Sci U S A. 2002;99:1443–1448. doi: 10.1073/pnas.032652399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Haga K, Lemp NA, Logg CR, Nagashima J, Faure-Kumar E, Gomez GG, Kruse CA, Mendez R, Stripecke R, Kasahara N. Permanent, lowered HLA class I expression using lentivirus vectors with shRNA constructs: averting cytotoxicity by alloreactive T lymphocytes. Transplant Proc. 2006;38:3184–3188. doi: 10.1016/j.transproceed.2006.10.158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Hacke K, Falahati R, Flebbe-Rehwaldt L, Kasahara N, Gaensler KML. Suppression of HLA expression by lentivirus-mediated gene transfer of siRNA cassettes and in vivo chemoselection to enhance hematopoietic stem cell transplantation. Immunol Res. 2009;44:112–126. doi: 10.1007/s12026-008-8088-z. [DOI] [PMC free article] [PubMed] [Google Scholar]


