Abstract
We quantitate the 'activating potentials' of deletion and point mutation variants of a 42 amino acid yeast transcriptional activating region excised from the yeast activator GAL4 and, using surface plasmon resonance, we measure the relative affinities of these molecules for a variety of proteins, including plausible target proteins as well as GAL80, a specific inhibitor of GAL4. We find a remarkable correlation between the relative activating potentials of the derivatives and their relative affinities for yeast TBP and for yeast TFIIB; other tested proteins interacted significantly more weakly, if at all. These results provide an especially strong argument that TBP and TFIIB are activating region targets. We also show, using one set of yeast activating region mutants, that activator-target interactions are strongly correlated with the length of the activating region, that the effect of point mutants is highly dependent on the length of the activating region mutated and that, unlike interactions with TBP and TFIIB, interaction with the specific inhibitor GAL80 is destroyed by deletion of certain critical residues in the C-terminal half of the 42 amino acid activating region.
Full text
PDF




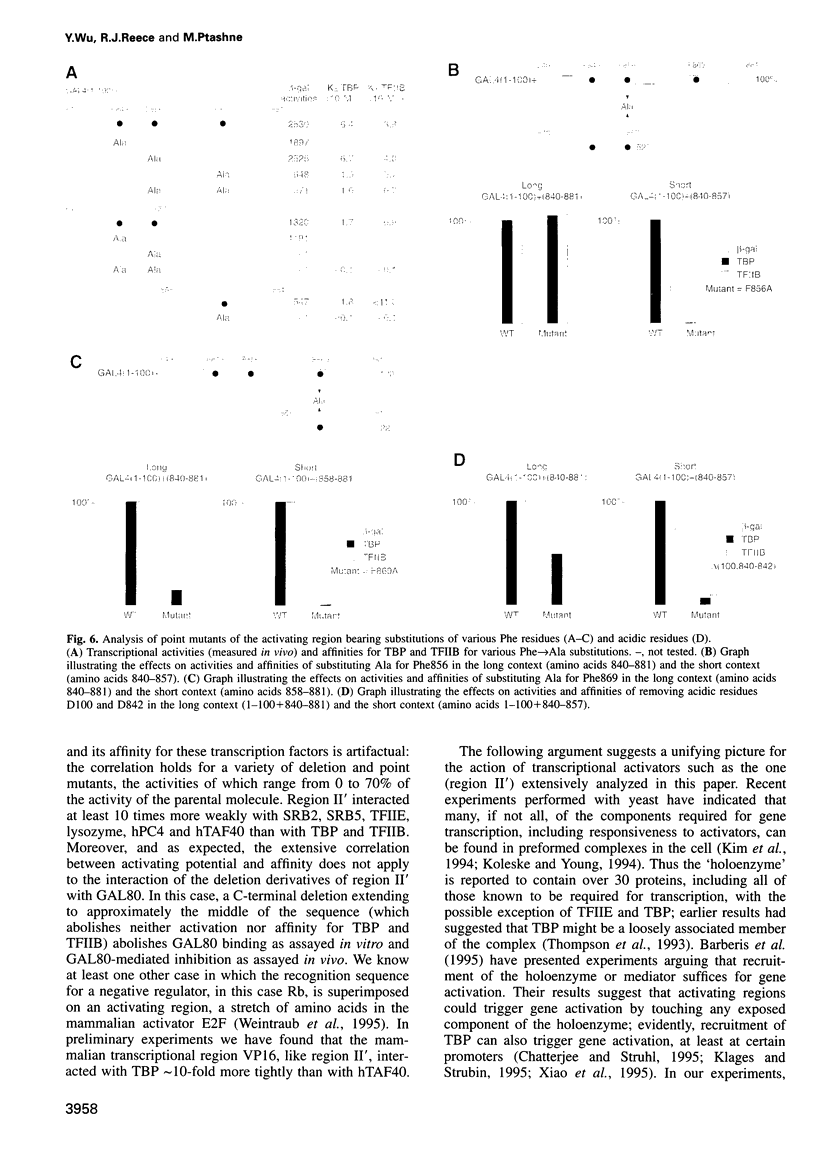
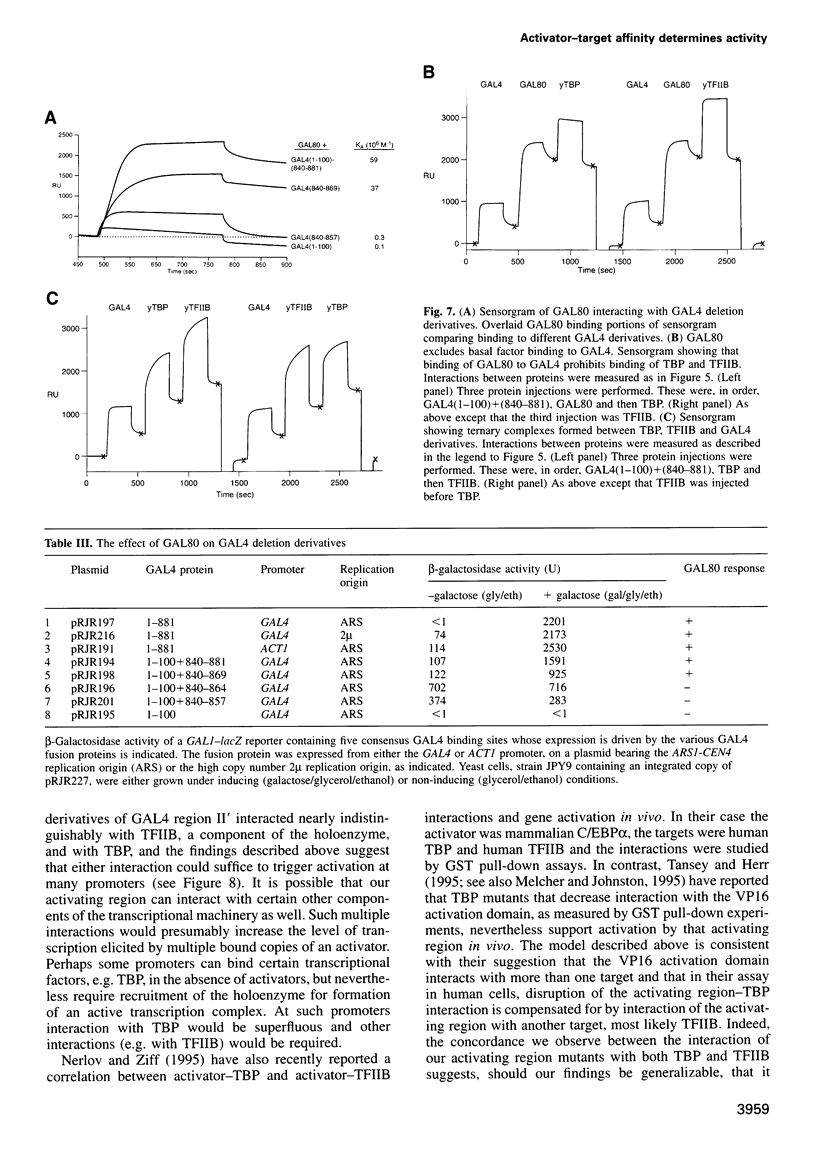




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alani E., Cao L., Kleckner N. A method for gene disruption that allows repeated use of URA3 selection in the construction of multiply disrupted yeast strains. Genetics. 1987 Aug;116(4):541–545. doi: 10.1534/genetics.112.541.test. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amann E., Brosius J., Ptashne M. Vectors bearing a hybrid trp-lac promoter useful for regulated expression of cloned genes in Escherichia coli. Gene. 1983 Nov;25(2-3):167–178. doi: 10.1016/0378-1119(83)90222-6. [DOI] [PubMed] [Google Scholar]
- Barberis A., Pearlberg J., Simkovich N., Farrell S., Reinagel P., Bamdad C., Sigal G., Ptashne M. Contact with a component of the polymerase II holoenzyme suffices for gene activation. Cell. 1995 May 5;81(3):359–368. doi: 10.1016/0092-8674(95)90389-5. [DOI] [PubMed] [Google Scholar]
- Blair W. S., Bogerd H. P., Madore S. J., Cullen B. R. Mutational analysis of the transcription activation domain of RelA: identification of a highly synergistic minimal acidic activation module. Mol Cell Biol. 1994 Nov;14(11):7226–7234. doi: 10.1128/mcb.14.11.7226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1006/abio.1976.9999. [DOI] [PubMed] [Google Scholar]
- Carey M., Leatherwood J., Ptashne M. A potent GAL4 derivative activates transcription at a distance in vitro. Science. 1990 Feb 9;247(4943):710–712. doi: 10.1126/science.2405489. [DOI] [PubMed] [Google Scholar]
- Chatterjee S., Struhl K. Connecting a promoter-bound protein to TBP bypasses the need for a transcriptional activation domain. Nature. 1995 Apr 27;374(6525):820–822. doi: 10.1038/374820a0. [DOI] [PubMed] [Google Scholar]
- Ge H., Roeder R. G. Purification, cloning, and characterization of a human coactivator, PC4, that mediates transcriptional activation of class II genes. Cell. 1994 Aug 12;78(3):513–523. doi: 10.1016/0092-8674(94)90428-6. [DOI] [PubMed] [Google Scholar]
- Gietz R. D., Sugino A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene. 1988 Dec 30;74(2):527–534. doi: 10.1016/0378-1119(88)90185-0. [DOI] [PubMed] [Google Scholar]
- Gill G., Sadowski I., Ptashne M. Mutations that increase the activity of a transcriptional activator in yeast and mammalian cells. Proc Natl Acad Sci U S A. 1990 Mar;87(6):2127–2131. doi: 10.1073/pnas.87.6.2127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodrich J. A., Hoey T., Thut C. J., Admon A., Tjian R. Drosophila TAFII40 interacts with both a VP16 activation domain and the basal transcription factor TFIIB. Cell. 1993 Nov 5;75(3):519–530. doi: 10.1016/0092-8674(93)90386-5. [DOI] [PubMed] [Google Scholar]
- Hahn S., Guarente L. Yeast HAP2 and HAP3: transcriptional activators in a heteromeric complex. Science. 1988 Apr 15;240(4850):317–321. doi: 10.1126/science.2832951. [DOI] [PubMed] [Google Scholar]
- Himmelfarb H. J., Pearlberg J., Last D. H., Ptashne M. GAL11P: a yeast mutation that potentiates the effect of weak GAL4-derived activators. Cell. 1990 Dec 21;63(6):1299–1309. doi: 10.1016/0092-8674(90)90425-e. [DOI] [PubMed] [Google Scholar]
- Hope I. A., Mahadevan S., Struhl K. Structural and functional characterization of the short acidic transcriptional activation region of yeast GCN4 protein. Nature. 1988 Jun 16;333(6174):635–640. doi: 10.1038/333635a0. [DOI] [PubMed] [Google Scholar]
- Johnston S. A., Salmeron J. M., Jr, Dincher S. S. Interaction of positive and negative regulatory proteins in the galactose regulon of yeast. Cell. 1987 Jul 3;50(1):143–146. doi: 10.1016/0092-8674(87)90671-4. [DOI] [PubMed] [Google Scholar]
- Kakidani H., Ptashne M. GAL4 activates gene expression in mammalian cells. Cell. 1988 Jan 29;52(2):161–167. doi: 10.1016/0092-8674(88)90504-1. [DOI] [PubMed] [Google Scholar]
- Kim Y. J., Björklund S., Li Y., Sayre M. H., Kornberg R. D. A multiprotein mediator of transcriptional activation and its interaction with the C-terminal repeat domain of RNA polymerase II. Cell. 1994 May 20;77(4):599–608. doi: 10.1016/0092-8674(94)90221-6. [DOI] [PubMed] [Google Scholar]
- Klages N., Strubin M. Stimulation of RNA polymerase II transcription initiation by recruitment of TBP in vivo. Nature. 1995 Apr 27;374(6525):822–823. doi: 10.1038/374822a0. [DOI] [PubMed] [Google Scholar]
- Klemm R. D., Goodrich J. A., Zhou S., Tjian R. Molecular cloning and expression of the 32-kDa subunit of human TFIID reveals interactions with VP16 and TFIIB that mediate transcriptional activation. Proc Natl Acad Sci U S A. 1995 Jun 20;92(13):5788–5792. doi: 10.1073/pnas.92.13.5788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koleske A. J., Young R. A. An RNA polymerase II holoenzyme responsive to activators. Nature. 1994 Mar 31;368(6470):466–469. doi: 10.1038/368466a0. [DOI] [PubMed] [Google Scholar]
- Leuther K. K., Johnston S. A. Nondissociation of GAL4 and GAL80 in vivo after galactose induction. Science. 1992 May 29;256(5061):1333–1335. doi: 10.1126/science.1598579. [DOI] [PubMed] [Google Scholar]
- Leuther K. K., Salmeron J. M., Johnston S. A. Genetic evidence that an activation domain of GAL4 does not require acidity and may form a beta sheet. Cell. 1993 Feb 26;72(4):575–585. doi: 10.1016/0092-8674(93)90076-3. [DOI] [PubMed] [Google Scholar]
- Lillie J. W., Green M., Green M. R. An adenovirus E1a protein region required for transformation and transcriptional repression. Cell. 1986 Sep 26;46(7):1043–1051. doi: 10.1016/0092-8674(86)90704-x. [DOI] [PubMed] [Google Scholar]
- Lin Y. S., Carey M. F., Ptashne M., Green M. R. GAL4 derivatives function alone and synergistically with mammalian activators in vitro. Cell. 1988 Aug 26;54(5):659–664. doi: 10.1016/s0092-8674(88)80010-2. [DOI] [PubMed] [Google Scholar]
- Ma J., Ptashne M. Deletion analysis of GAL4 defines two transcriptional activating segments. Cell. 1987 Mar 13;48(5):847–853. doi: 10.1016/0092-8674(87)90081-x. [DOI] [PubMed] [Google Scholar]
- Ma J., Ptashne M. The carboxy-terminal 30 amino acids of GAL4 are recognized by GAL80. Cell. 1987 Jul 3;50(1):137–142. doi: 10.1016/0092-8674(87)90670-2. [DOI] [PubMed] [Google Scholar]
- Melcher K., Johnston S. A. GAL4 interacts with TATA-binding protein and coactivators. Mol Cell Biol. 1995 May;15(5):2839–2848. doi: 10.1128/mcb.15.5.2839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nerlov C., Ziff E. B. CCAAT/enhancer binding protein-alpha amino acid motifs with dual TBP and TFIIB binding ability co-operate to activate transcription in both yeast and mammalian cells. EMBO J. 1995 Sep 1;14(17):4318–4328. doi: 10.1002/j.1460-2075.1995.tb00106.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikolov D. B., Chen H., Halay E. D., Usheva A. A., Hisatake K., Lee D. K., Roeder R. G., Burley S. K. Crystal structure of a TFIIB-TBP-TATA-element ternary complex. Nature. 1995 Sep 14;377(6545):119–128. doi: 10.1038/377119a0. [DOI] [PubMed] [Google Scholar]
- Ohashi Y., Brickman J. M., Furman E., Middleton B., Carey M. Modulating the potency of an activator in a yeast in vitro transcription system. Mol Cell Biol. 1994 Apr;14(4):2731–2739. doi: 10.1128/mcb.14.4.2731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ponticelli A. S., Struhl K. Analysis of Saccharomyces cerevisiae his3 transcription in vitro: biochemical support for multiple mechanisms of transcription. Mol Cell Biol. 1990 Jun;10(6):2832–2839. doi: 10.1128/mcb.10.6.2832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ptashne M. How eukaryotic transcriptional activators work. Nature. 1988 Oct 20;335(6192):683–689. doi: 10.1038/335683a0. [DOI] [PubMed] [Google Scholar]
- Reece R. J., Maxwell A. The C-terminal domain of the Escherichia coli DNA gyrase A subunit is a DNA-binding protein. Nucleic Acids Res. 1991 Apr 11;19(7):1399–1405. doi: 10.1093/nar/19.7.1399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reece R. J., Ptashne M. Determinants of binding-site specificity among yeast C6 zinc cluster proteins. Science. 1993 Aug 13;261(5123):909–911. doi: 10.1126/science.8346441. [DOI] [PubMed] [Google Scholar]
- Roberts S. G., Ha I., Maldonado E., Reinberg D., Green M. R. Interaction between an acidic activator and transcription factor TFIIB is required for transcriptional activation. Nature. 1993 Jun 24;363(6431):741–744. doi: 10.1038/363741a0. [DOI] [PubMed] [Google Scholar]
- Stringer K. F., Ingles C. J., Greenblatt J. Direct and selective binding of an acidic transcriptional activation domain to the TATA-box factor TFIID. Nature. 1990 Jun 28;345(6278):783–786. doi: 10.1038/345783a0. [DOI] [PubMed] [Google Scholar]
- Strubin M., Struhl K. Yeast and human TFIID with altered DNA-binding specificity for TATA elements. Cell. 1992 Feb 21;68(4):721–730. doi: 10.1016/0092-8674(92)90147-5. [DOI] [PubMed] [Google Scholar]
- Studier F. W., Rosenberg A. H., Dunn J. J., Dubendorff J. W. Use of T7 RNA polymerase to direct expression of cloned genes. Methods Enzymol. 1990;185:60–89. doi: 10.1016/0076-6879(90)85008-c. [DOI] [PubMed] [Google Scholar]
- Swaffield J. C., Melcher K., Johnston S. A. A highly conserved ATPase protein as a mediator between acidic activation domains and the TATA-binding protein. Nature. 1995 Mar 2;374(6517):88–91. doi: 10.1038/374088a0. [DOI] [PubMed] [Google Scholar]
- Tanaka M. Modulation of promoter occupancy by cooperative DNA binding and activation-domain function is a major determinant of transcriptional regulation by activators in vivo. Proc Natl Acad Sci U S A. 1996 Apr 30;93(9):4311–4315. doi: 10.1073/pnas.93.9.4311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tansey W. P., Herr W. The ability to associate with activation domains in vitro is not required for the TATA box-binding protein to support activated transcription in vivo. Proc Natl Acad Sci U S A. 1995 Nov 7;92(23):10550–10554. doi: 10.1073/pnas.92.23.10550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson C. M., Koleske A. J., Chao D. M., Young R. A. A multisubunit complex associated with the RNA polymerase II CTD and TATA-binding protein in yeast. Cell. 1993 Jul 2;73(7):1361–1375. doi: 10.1016/0092-8674(93)90362-t. [DOI] [PubMed] [Google Scholar]
- Thut C. J., Chen J. L., Klemm R., Tjian R. p53 transcriptional activation mediated by coactivators TAFII40 and TAFII60. Science. 1995 Jan 6;267(5194):100–104. doi: 10.1126/science.7809597. [DOI] [PubMed] [Google Scholar]
- Triezenberg S. J. Structure and function of transcriptional activation domains. Curr Opin Genet Dev. 1995 Apr;5(2):190–196. doi: 10.1016/0959-437x(95)80007-7. [DOI] [PubMed] [Google Scholar]
- Van Hoy M., Leuther K. K., Kodadek T., Johnston S. A. The acidic activation domains of the GCN4 and GAL4 proteins are not alpha helical but form beta sheets. Cell. 1993 Feb 26;72(4):587–594. doi: 10.1016/0092-8674(93)90077-4. [DOI] [PubMed] [Google Scholar]
- Vashee S., Kodadek T. The activation domain of GAL4 protein mediates cooperative promoter binding with general transcription factors in vivo. Proc Natl Acad Sci U S A. 1995 Nov 7;92(23):10683–10687. doi: 10.1073/pnas.92.23.10683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weintraub S. J., Chow K. N., Luo R. X., Zhang S. H., He S., Dean D. C. Mechanism of active transcriptional repression by the retinoblastoma protein. Nature. 1995 Jun 29;375(6534):812–815. doi: 10.1038/375812a0. [DOI] [PubMed] [Google Scholar]
- Xiao H., Friesen J. D., Lis J. T. Recruiting TATA-binding protein to a promoter: transcriptional activation without an upstream activator. Mol Cell Biol. 1995 Oct;15(10):5757–5761. doi: 10.1128/mcb.15.10.5757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao H., Pearson A., Coulombe B., Truant R., Zhang S., Regier J. L., Triezenberg S. J., Reinberg D., Flores O., Ingles C. J. Binding of basal transcription factor TFIIH to the acidic activation domains of VP16 and p53. Mol Cell Biol. 1994 Oct;14(10):7013–7024. doi: 10.1128/mcb.14.10.7013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yocum R. R., Hanley S., West R., Jr, Ptashne M. Use of lacZ fusions to delimit regulatory elements of the inducible divergent GAL1-GAL10 promoter in Saccharomyces cerevisiae. Mol Cell Biol. 1984 Oct;4(10):1985–1998. doi: 10.1128/mcb.4.10.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]