Abstract
The c-Myc protein is involved in cell proliferation, differentiation and apoptosis though heterodimerization with Max to form a transcriptionally active sequence-specific DNA binding complex. By means of sequential immunoprecipitation of chromatin using anti-Max and anti-Myc antibodies, we have identified a Myc-regulated gene and genomic sites occupied by Myc-Max in vivo. Four of 27 sites recovered by this procedure corresponded to the highest affinity 'canonical' CACGTG sequence. However, the most common in vivo binding sites belonged to the group of 'non-canonical' E box-related binding sites previously identified by in vitro selection. Several of the genomic fragments isolated contained transcribed sequences, including one, MrDb, encoding an evolutionarily conserved RNA helicase of the DEAD box family. The corresponding mRNA was induced following activation of a Myc-estrogen receptor fusion protein (Myc-ER) in the presence of a protein synthesis inhibitor, consistent with this helicase gene being a direct target of Myc-Max. In addition, as for c-Myc, the expression of MrDb is induced upon proliferative stimulation of primary human fibroblasts as well as B cells and down-regulated during terminal differentiation of HL60 leukemia cells. Our results indicate that Myc-Max heterodimers interact in vivo with a specific set of E box-related DNA sequences and that Myc is likely to activate multiple target genes including a highly conserved DEAD box protein. Therefore, Myc may exert its effects on cell behavior through proteins that affect RNA structure and metabolism.
Full text
PDF




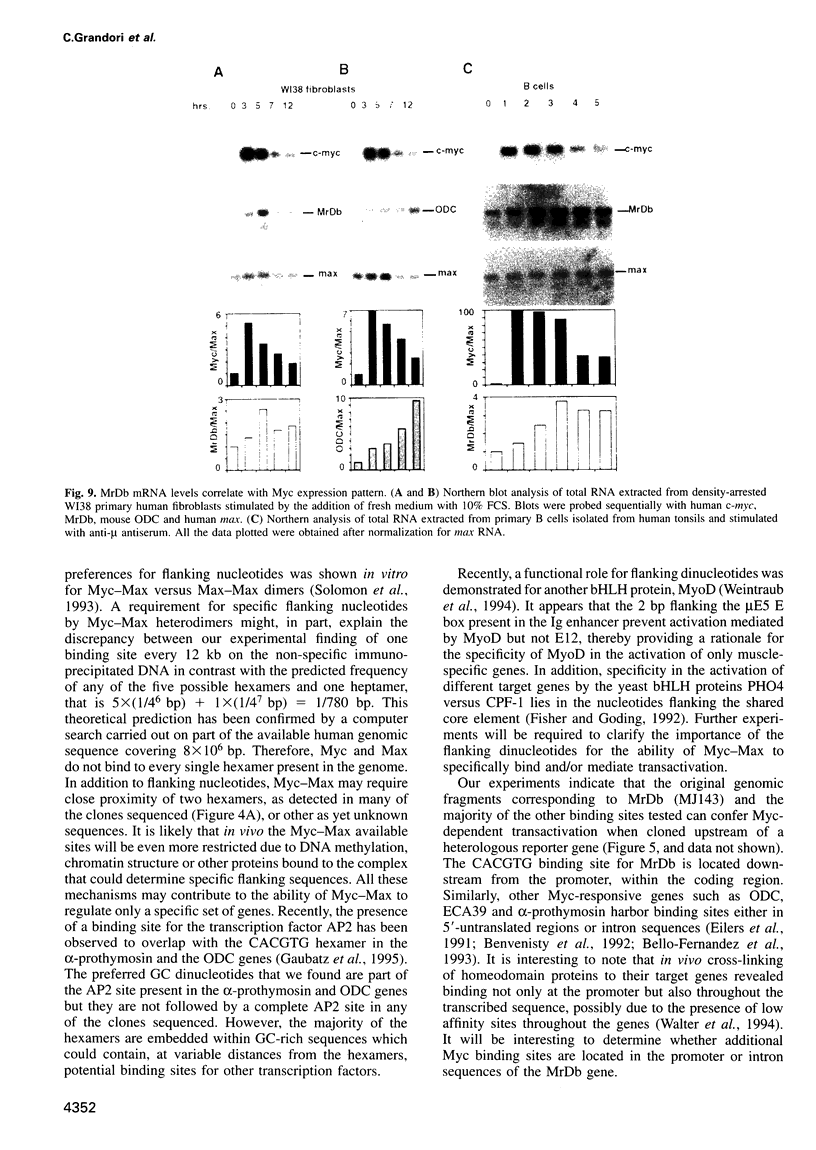
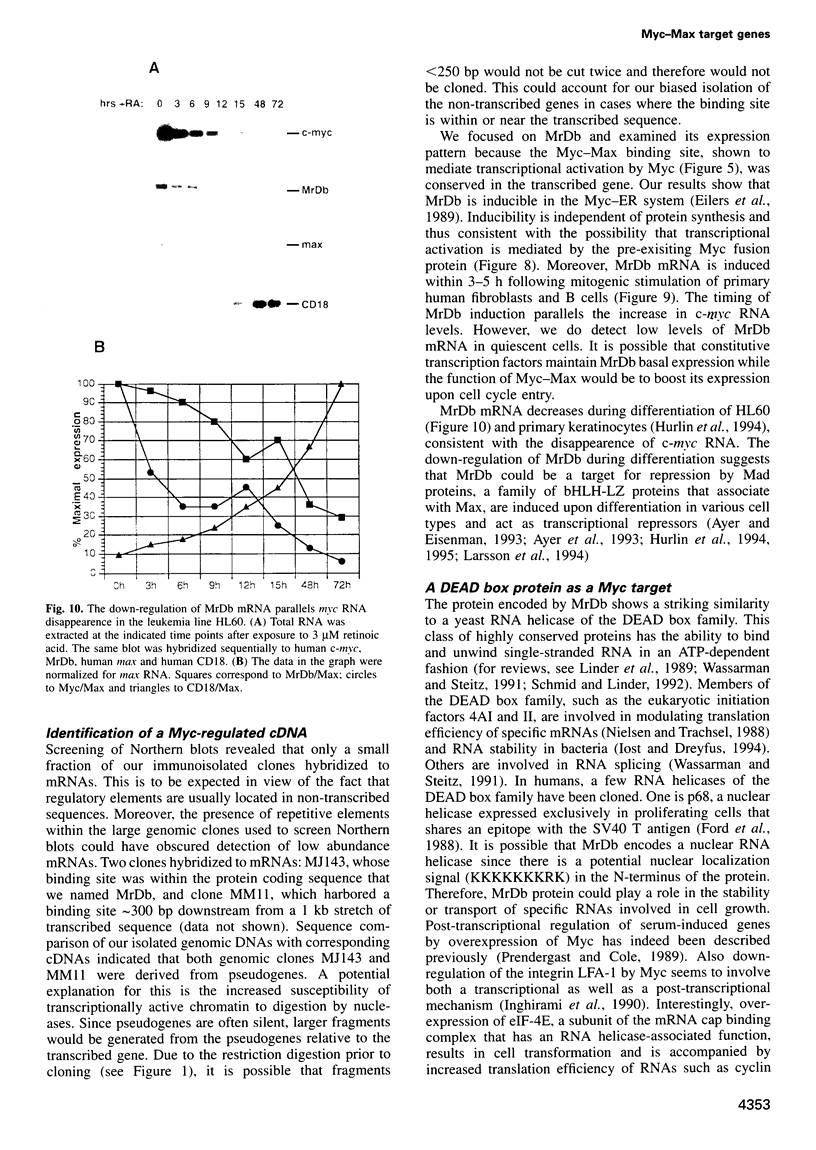




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Akao Y., Seto M., Yamamoto K., Iida S., Nakazawa S., Inazawa J., Abe T., Takahashi T., Ueda R. The RCK gene associated with t(11;14) translocation is distinct from the MLL/ALL-1 gene with t(4;11) and t(11;19) translocations. Cancer Res. 1992 Nov 1;52(21):6083–6087. [PubMed] [Google Scholar]
- Amati B., Brooks M. W., Levy N., Littlewood T. D., Evan G. I., Land H. Oncogenic activity of the c-Myc protein requires dimerization with Max. Cell. 1993 Jan 29;72(2):233–245. doi: 10.1016/0092-8674(93)90663-b. [DOI] [PubMed] [Google Scholar]
- Amati B., Land H. Myc-Max-Mad: a transcription factor network controlling cell cycle progression, differentiation and death. Curr Opin Genet Dev. 1994 Feb;4(1):102–108. doi: 10.1016/0959-437x(94)90098-1. [DOI] [PubMed] [Google Scholar]
- Amati B., Littlewood T. D., Evan G. I., Land H. The c-Myc protein induces cell cycle progression and apoptosis through dimerization with Max. EMBO J. 1993 Dec 15;12(13):5083–5087. doi: 10.1002/j.1460-2075.1993.tb06202.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Auvinen M., Paasinen A., Andersson L. C., Hölttä E. Ornithine decarboxylase activity is critical for cell transformation. Nature. 1992 Nov 26;360(6402):355–358. doi: 10.1038/360355a0. [DOI] [PubMed] [Google Scholar]
- Ayer D. E., Eisenman R. N. A switch from Myc:Max to Mad:Max heterocomplexes accompanies monocyte/macrophage differentiation. Genes Dev. 1993 Nov;7(11):2110–2119. doi: 10.1101/gad.7.11.2110. [DOI] [PubMed] [Google Scholar]
- Ayer D. E., Kretzner L., Eisenman R. N. Mad: a heterodimeric partner for Max that antagonizes Myc transcriptional activity. Cell. 1993 Jan 29;72(2):211–222. doi: 10.1016/0092-8674(93)90661-9. [DOI] [PubMed] [Google Scholar]
- Beckmann H., Su L. K., Kadesch T. TFE3: a helix-loop-helix protein that activates transcription through the immunoglobulin enhancer muE3 motif. Genes Dev. 1990 Feb;4(2):167–179. doi: 10.1101/gad.4.2.167. [DOI] [PubMed] [Google Scholar]
- Bellard M., Dretzen G., Giangrande A., Ramain P. Nuclease digestion of transcriptionally active chromatin. Methods Enzymol. 1989;170:317–346. doi: 10.1016/0076-6879(89)70054-9. [DOI] [PubMed] [Google Scholar]
- Bello-Fernandez C., Packham G., Cleveland J. L. The ornithine decarboxylase gene is a transcriptional target of c-Myc. Proc Natl Acad Sci U S A. 1993 Aug 15;90(16):7804–7808. doi: 10.1073/pnas.90.16.7804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benvenisty N., Leder A., Kuo A., Leder P. An embryonically expressed gene is a target for c-Myc regulation via the c-Myc-binding sequence. Genes Dev. 1992 Dec;6(12B):2513–2523. doi: 10.1101/gad.6.12b.2513. [DOI] [PubMed] [Google Scholar]
- Berberich S., Hyde-DeRuyscher N., Espenshade P., Cole M. max encodes a sequence-specific DNA-binding protein and is not regulated by serum growth factors. Oncogene. 1992 Apr;7(4):775–779. [PubMed] [Google Scholar]
- Bigler J., Eisenman R. N. Isolation of a thyroid hormone-responsive gene by immunoprecipitation of thyroid hormone receptor-DNA complexes. Mol Cell Biol. 1994 Nov;14(11):7621–7632. doi: 10.1128/mcb.14.11.7621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bigler J., Eisenman R. N. Novel location and function of a thyroid hormone response element. EMBO J. 1995 Nov 15;14(22):5710–5723. doi: 10.1002/j.1460-2075.1995.tb00258.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blackwell T. K., Huang J., Ma A., Kretzner L., Alt F. W., Eisenman R. N., Weintraub H. Binding of myc proteins to canonical and noncanonical DNA sequences. Mol Cell Biol. 1993 Sep;13(9):5216–5224. doi: 10.1128/mcb.13.9.5216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blackwell T. K., Kretzner L., Blackwood E. M., Eisenman R. N., Weintraub H. Sequence-specific DNA binding by the c-Myc protein. Science. 1990 Nov 23;250(4984):1149–1151. doi: 10.1126/science.2251503. [DOI] [PubMed] [Google Scholar]
- Blackwood E. M., Eisenman R. N. Max: a helix-loop-helix zipper protein that forms a sequence-specific DNA-binding complex with Myc. Science. 1991 Mar 8;251(4998):1211–1217. doi: 10.1126/science.2006410. [DOI] [PubMed] [Google Scholar]
- Blackwood E. M., Lüscher B., Eisenman R. N. Myc and Max associate in vivo. Genes Dev. 1992 Jan;6(1):71–80. doi: 10.1101/gad.6.1.71. [DOI] [PubMed] [Google Scholar]
- Carr C. S., Sharp P. A. A helix-loop-helix protein related to the immunoglobulin E box-binding proteins. Mol Cell Biol. 1990 Aug;10(8):4384–4388. doi: 10.1128/mcb.10.8.4384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark E. A., Shu G. L., Lüscher B., Draves K. E., Banchereau J., Ledbetter J. A., Valentine M. A. Activation of human B cells. Comparison of the signal transduced by IL-4 to four different competence signals. J Immunol. 1989 Dec 15;143(12):3873–3880. [PubMed] [Google Scholar]
- Collins S. J., Ruscetti F. W., Gallagher R. E., Gallo R. C. Terminal differentiation of human promyelocytic leukemia cells induced by dimethyl sulfoxide and other polar compounds. Proc Natl Acad Sci U S A. 1978 May;75(5):2458–2462. doi: 10.1073/pnas.75.5.2458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cosenza S. C., Owen T. A., Soprano D. R., Soprano K. J. Evidence that the time of entry into S is determined by events occurring in early G1. J Biol Chem. 1988 Sep 5;263(25):12751–12758. [PubMed] [Google Scholar]
- Cowan N. J., Dobner P. R., Fuchs E. V., Cleveland D. W. Expression of human alpha-tubulin genes: interspecies conservation of 3' untranslated regions. Mol Cell Biol. 1983 Oct;3(10):1738–1745. doi: 10.1128/mcb.3.10.1738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dingwall C., Laskey R. A. Nuclear targeting sequences--a consensus? Trends Biochem Sci. 1991 Dec;16(12):478–481. doi: 10.1016/0968-0004(91)90184-w. [DOI] [PubMed] [Google Scholar]
- Eilers M., Picard D., Yamamoto K. R., Bishop J. M. Chimaeras of myc oncoprotein and steroid receptors cause hormone-dependent transformation of cells. Nature. 1989 Jul 6;340(6228):66–68. doi: 10.1038/340066a0. [DOI] [PubMed] [Google Scholar]
- Eilers M., Schirm S., Bishop J. M. The MYC protein activates transcription of the alpha-prothymosin gene. EMBO J. 1991 Jan;10(1):133–141. doi: 10.1002/j.1460-2075.1991.tb07929.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisenman R. N., Tachibana C. Y., Abrams H. D., Hann S. R. V-myc- and c-myc-encoded proteins are associated with the nuclear matrix. Mol Cell Biol. 1985 Jan;5(1):114–126. doi: 10.1128/mcb.5.1.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evan G. I., Wyllie A. H., Gilbert C. S., Littlewood T. D., Land H., Brooks M., Waters C. M., Penn L. Z., Hancock D. C. Induction of apoptosis in fibroblasts by c-myc protein. Cell. 1992 Apr 3;69(1):119–128. doi: 10.1016/0092-8674(92)90123-t. [DOI] [PubMed] [Google Scholar]
- Fisher F., Goding C. R. Single amino acid substitutions alter helix-loop-helix protein specificity for bases flanking the core CANNTG motif. EMBO J. 1992 Nov;11(11):4103–4109. doi: 10.1002/j.1460-2075.1992.tb05503.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ford M. J., Anton I. A., Lane D. P. Nuclear protein with sequence homology to translation initiation factor eIF-4A. Nature. 1988 Apr 21;332(6166):736–738. doi: 10.1038/332736a0. [DOI] [PubMed] [Google Scholar]
- Gaubatz S., Imhof A., Dosch R., Werner O., Mitchell P., Buettner R., Eilers M. Transcriptional activation by Myc is under negative control by the transcription factor AP-2. EMBO J. 1995 Apr 3;14(7):1508–1519. doi: 10.1002/j.1460-2075.1995.tb07137.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaubatz S., Meichle A., Eilers M. An E-box element localized in the first intron mediates regulation of the prothymosin alpha gene by c-myc. Mol Cell Biol. 1994 Jun;14(6):3853–3862. doi: 10.1128/mcb.14.6.3853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Godbout R., Squire J. Amplification of a DEAD box protein gene in retinoblastoma cell lines. Proc Natl Acad Sci U S A. 1993 Aug 15;90(16):7578–7582. doi: 10.1073/pnas.90.16.7578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gould A. P., Brookman J. J., Strutt D. I., White R. A. Targets of homeotic gene control in Drosophila. Nature. 1990 Nov 22;348(6299):308–312. doi: 10.1038/348308a0. [DOI] [PubMed] [Google Scholar]
- Gregor P. D., Sawadogo M., Roeder R. G. The adenovirus major late transcription factor USF is a member of the helix-loop-helix group of regulatory proteins and binds to DNA as a dimer. Genes Dev. 1990 Oct;4(10):1730–1740. doi: 10.1101/gad.4.10.1730. [DOI] [PubMed] [Google Scholar]
- Gu W., Cechova K., Tassi V., Dalla-Favera R. Opposite regulation of gene transcription and cell proliferation by c-Myc and Max. Proc Natl Acad Sci U S A. 1993 Apr 1;90(7):2935–2939. doi: 10.1073/pnas.90.7.2935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halazonetis T. D., Kandil A. N. Determination of the c-MYC DNA-binding site. Proc Natl Acad Sci U S A. 1991 Jul 15;88(14):6162–6166. doi: 10.1073/pnas.88.14.6162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hann S. R., King M. W., Bentley D. L., Anderson C. W., Eisenman R. N. A non-AUG translational initiation in c-myc exon 1 generates an N-terminally distinct protein whose synthesis is disrupted in Burkitt's lymphomas. Cell. 1988 Jan 29;52(2):185–195. doi: 10.1016/0092-8674(88)90507-7. [DOI] [PubMed] [Google Scholar]
- Henikoff S., Eghtedarzadeh M. K. Conserved arrangement of nested genes at the Drosophila Gart locus. Genetics. 1987 Dec;117(4):711–725. doi: 10.1093/genetics/117.4.711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene. 1984 Jun;28(3):351–359. doi: 10.1016/0378-1119(84)90153-7. [DOI] [PubMed] [Google Scholar]
- Hewish D. R., Burgoyne L. A. Chromatin sub-structure. The digestion of chromatin DNA at regularly spaced sites by a nuclear deoxyribonuclease. Biochem Biophys Res Commun. 1973 May 15;52(2):504–510. doi: 10.1016/0006-291x(73)90740-7. [DOI] [PubMed] [Google Scholar]
- Hickstein D. D., Back A. L., Collins S. J. Regulation of expression of the CD11b and CD18 subunits of the neutrophil adherence receptor during human myeloid differentiation. J Biol Chem. 1989 Dec 25;264(36):21812–21817. [PubMed] [Google Scholar]
- Hurlin P. J., Ayer D. E., Grandori C., Eisenman R. N. The Max transcription factor network: involvement of Mad in differentiation and an approach to identification of target genes. Cold Spring Harb Symp Quant Biol. 1994;59:109–116. doi: 10.1101/sqb.1994.059.01.014. [DOI] [PubMed] [Google Scholar]
- Hurlin P. J., Quéva C., Koskinen P. J., Steingrímsson E., Ayer D. E., Copeland N. G., Jenkins N. A., Eisenman R. N. Mad3 and Mad4: novel Max-interacting transcriptional repressors that suppress c-myc dependent transformation and are expressed during neural and epidermal differentiation. EMBO J. 1995 Nov 15;14(22):5646–5659. doi: 10.1002/j.1460-2075.1995.tb00252.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hümbelin M., Safer B., Chiorini J. A., Hershey J. W., Cohen R. B. Isolation and characterization of the promoter and flanking regions of the gene encoding the human protein-synthesis-initiation factor 2 alpha. Gene. 1989 Sep 30;81(2):315–324. doi: 10.1016/0378-1119(89)90192-3. [DOI] [PubMed] [Google Scholar]
- Inghirami G., Grignani F., Sternas L., Lombardi L., Knowles D. M., Dalla-Favera R. Down-regulation of LFA-1 adhesion receptors by C-myc oncogene in human B lymphoblastoid cells. Science. 1990 Nov 2;250(4981):682–686. doi: 10.1126/science.2237417. [DOI] [PubMed] [Google Scholar]
- Iost I., Dreyfus M. mRNAs can be stabilized by DEAD-box proteins. Nature. 1994 Nov 10;372(6502):193–196. doi: 10.1038/372193a0. [DOI] [PubMed] [Google Scholar]
- Kahana C., Nathans D. Isolation of cloned cDNA encoding mammalian ornithine decarboxylase. Proc Natl Acad Sci U S A. 1984 Jun;81(12):3645–3649. doi: 10.1073/pnas.81.12.3645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kato G. J., Barrett J., Villa-Garcia M., Dang C. V. An amino-terminal c-myc domain required for neoplastic transformation activates transcription. Mol Cell Biol. 1990 Nov;10(11):5914–5920. doi: 10.1128/mcb.10.11.5914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozak M. Point mutations define a sequence flanking the AUG initiator codon that modulates translation by eukaryotic ribosomes. Cell. 1986 Jan 31;44(2):283–292. doi: 10.1016/0092-8674(86)90762-2. [DOI] [PubMed] [Google Scholar]
- Kretzner L., Blackwood E. M., Eisenman R. N. Myc and Max proteins possess distinct transcriptional activities. Nature. 1992 Oct 1;359(6394):426–429. doi: 10.1038/359426a0. [DOI] [PubMed] [Google Scholar]
- Larsson L. G., Pettersson M., Oberg F., Nilsson K., Lüscher B. Expression of mad, mxi1, max and c-myc during induced differentiation of hematopoietic cells: opposite regulation of mad and c-myc. Oncogene. 1994 Apr;9(4):1247–1252. [PubMed] [Google Scholar]
- Lasko P. F., Ashburner M. The product of the Drosophila gene vasa is very similar to eukaryotic initiation factor-4A. Nature. 1988 Oct 13;335(6191):611–617. doi: 10.1038/335611a0. [DOI] [PubMed] [Google Scholar]
- Lazaris-Karatzas A., Montine K. S., Sonenberg N. Malignant transformation by a eukaryotic initiation factor subunit that binds to mRNA 5' cap. Nature. 1990 Jun 7;345(6275):544–547. doi: 10.1038/345544a0. [DOI] [PubMed] [Google Scholar]
- Li L. H., Nerlov C., Prendergast G., MacGregor D., Ziff E. B. c-Myc represses transcription in vivo by a novel mechanism dependent on the initiator element and Myc box II. EMBO J. 1994 Sep 1;13(17):4070–4079. doi: 10.1002/j.1460-2075.1994.tb06724.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linder P., Lasko P. F., Ashburner M., Leroy P., Nielsen P. J., Nishi K., Schnier J., Slonimski P. P. Birth of the D-E-A-D box. Nature. 1989 Jan 12;337(6203):121–122. doi: 10.1038/337121a0. [DOI] [PubMed] [Google Scholar]
- Littlewood T. D., Hancock D. C., Danielian P. S., Parker M. G., Evan G. I. A modified oestrogen receptor ligand-binding domain as an improved switch for the regulation of heterologous proteins. Nucleic Acids Res. 1995 May 25;23(10):1686–1690. doi: 10.1093/nar/23.10.1686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lombardi L., Newcomb E. W., Dalla-Favera R. Pathogenesis of Burkitt lymphoma: expression of an activated c-myc oncogene causes the tumorigenic conversion of EBV-infected human B lymphoblasts. Cell. 1987 Apr 24;49(2):161–170. doi: 10.1016/0092-8674(87)90556-3. [DOI] [PubMed] [Google Scholar]
- Lu D., Yunis J. J. Cloning, expression and localization of an RNA helicase gene from a human lymphoid cell line with chromosomal breakpoint 11q23.3. Nucleic Acids Res. 1992 Apr 25;20(8):1967–1972. doi: 10.1093/nar/20.8.1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manohar C. F., Salwen H. R., Brodeur G. M., Cohn S. L. Co-amplification and concomitant high levels of expression of a DEAD box gene with MYCN in human neuroblastoma. Genes Chromosomes Cancer. 1995 Nov;14(3):196–203. doi: 10.1002/gcc.2870140307. [DOI] [PubMed] [Google Scholar]
- Meichle A., Philipp A., Eilers M. The functions of Myc proteins. Biochim Biophys Acta. 1992 Dec 16;1114(2-3):129–146. doi: 10.1016/0304-419x(92)90011-m. [DOI] [PubMed] [Google Scholar]
- Nielsen P. J., Trachsel H. The mouse protein synthesis initiation factor 4A gene family includes two related functional genes which are differentially expressed. EMBO J. 1988 Jul;7(7):2097–2105. doi: 10.1002/j.1460-2075.1988.tb03049.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niwa O., Kumazaki T., Tsukiyama T., Soma G., Miyajima N., Yokoro K. A cDNA clone overexpressed and amplified in a mouse teratocarcinoma line. Nucleic Acids Res. 1990 Nov 25;18(22):6709–6709. doi: 10.1093/nar/18.22.6709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nottenburg C., Gallatin W. M., St John T. Lymphocyte HEV adhesion variants differ in the expression of multiple gene sequences. Gene. 1990 Nov 15;95(2):279–284. doi: 10.1016/0378-1119(90)90372-x. [DOI] [PubMed] [Google Scholar]
- Packham G., Cleveland J. L. Ornithine decarboxylase is a mediator of c-Myc-induced apoptosis. Mol Cell Biol. 1994 Sep;14(9):5741–5747. doi: 10.1128/mcb.14.9.5741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papoulas O., Williams N. G., Kingston R. E. DNA binding activities of c-Myc purified from eukaryotic cells. J Biol Chem. 1992 May 25;267(15):10470–10480. [PubMed] [Google Scholar]
- Pause A., Méthot N., Sonenberg N. The HRIGRXXR region of the DEAD box RNA helicase eukaryotic translation initiation factor 4A is required for RNA binding and ATP hydrolysis. Mol Cell Biol. 1993 Nov;13(11):6789–6798. doi: 10.1128/mcb.13.11.6789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Penn L. J., Brooks M. W., Laufer E. M., Littlewood T. D., Morgenstern J. P., Evan G. I., Lee W. M., Land H. Domains of human c-myc protein required for autosuppression and cooperation with ras oncogenes are overlapping. Mol Cell Biol. 1990 Sep;10(9):4961–4966. doi: 10.1128/mcb.10.9.4961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Philipp A., Schneider A., Väsrik I., Finke K., Xiong Y., Beach D., Alitalo K., Eilers M. Repression of cyclin D1: a novel function of MYC. Mol Cell Biol. 1994 Jun;14(6):4032–4043. doi: 10.1128/mcb.14.6.4032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prendergast G. C., Cole M. D. Posttranscriptional regulation of cellular gene expression by the c-myc oncogene. Mol Cell Biol. 1989 Jan;9(1):124–134. doi: 10.1128/mcb.9.1.124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prendergast G. C., Lawe D., Ziff E. B. Association of Myn, the murine homolog of max, with c-Myc stimulates methylation-sensitive DNA binding and ras cotransformation. Cell. 1991 May 3;65(3):395–407. doi: 10.1016/0092-8674(91)90457-a. [DOI] [PubMed] [Google Scholar]
- Prendergast G. C., Ziff E. B. Methylation-sensitive sequence-specific DNA binding by the c-Myc basic region. Science. 1991 Jan 11;251(4990):186–189. doi: 10.1126/science.1987636. [DOI] [PubMed] [Google Scholar]
- Reisman D., Elkind N. B., Roy B., Beamon J., Rotter V. c-Myc trans-activates the p53 promoter through a required downstream CACGTG motif. Cell Growth Differ. 1993 Feb;4(2):57–65. [PubMed] [Google Scholar]
- Rosenwald I. B., Lazaris-Karatzas A., Sonenberg N., Schmidt E. V. Elevated levels of cyclin D1 protein in response to increased expression of eukaryotic initiation factor 4E. Mol Cell Biol. 1993 Dec;13(12):7358–7363. doi: 10.1128/mcb.13.12.7358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosenwald I. B., Rhoads D. B., Callanan L. D., Isselbacher K. J., Schmidt E. V. Increased expression of eukaryotic translation initiation factors eIF-4E and eIF-2 alpha in response to growth induction by c-myc. Proc Natl Acad Sci U S A. 1993 Jul 1;90(13):6175–6178. doi: 10.1073/pnas.90.13.6175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmid S. R., Linder P. D-E-A-D protein family of putative RNA helicases. Mol Microbiol. 1992 Feb;6(3):283–291. doi: 10.1111/j.1365-2958.1992.tb01470.x. [DOI] [PubMed] [Google Scholar]
- Solomon D. L., Amati B., Land H. Distinct DNA binding preferences for the c-Myc/Max and Max/Max dimers. Nucleic Acids Res. 1993 Nov 25;21(23):5372–5376. doi: 10.1093/nar/21.23.5372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Solomon D. L., Philipp A., Land H., Eilers M. Expression of cyclin D1 mRNA is not upregulated by Myc in rat fibroblasts. Oncogene. 1995 Nov 2;11(9):1893–1897. [PubMed] [Google Scholar]
- Squire J. A., Thorner P. S., Weitzman S., Maggi J. D., Dirks P., Doyle J., Hale M., Godbout R. Co-amplification of MYCN and a DEAD box gene (DDX1) in primary neuroblastoma. Oncogene. 1995 Apr 6;10(7):1417–1422. [PubMed] [Google Scholar]
- Stone J., de Lange T., Ramsay G., Jakobovits E., Bishop J. M., Varmus H., Lee W. Definition of regions in human c-myc that are involved in transformation and nuclear localization. Mol Cell Biol. 1987 May;7(5):1697–1709. doi: 10.1128/mcb.7.5.1697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tikhonenko A. T., Hartman A. R., Linial M. L. Overproduction of v-Myc in the nucleus and its excess over Max are not required for avian fibroblast transformation. Mol Cell Biol. 1993 Jun;13(6):3623–3631. doi: 10.1128/mcb.13.6.3623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tobias K. E., Shor J., Kahana C. c-Myc and Max transregulate the mouse ornithine decarboxylase promoter through interaction with two downstream CACGTG motifs. Oncogene. 1995 Nov 2;11(9):1721–1727. [PubMed] [Google Scholar]
- Tokunaga K., Nakamura Y., Sakata K., Fujimori K., Ohkubo M., Sawada K., Sakiyama S. Enhanced expression of a glyceraldehyde-3-phosphate dehydrogenase gene in human lung cancers. Cancer Res. 1987 Nov 1;47(21):5616–5619. [PubMed] [Google Scholar]
- Wagner A. J., Meyers C., Laimins L. A., Hay N. c-Myc induces the expression and activity of ornithine decarboxylase. Cell Growth Differ. 1993 Nov;4(11):879–883. [PubMed] [Google Scholar]
- Walter J., Dever C. A., Biggin M. D. Two homeo domain proteins bind with similar specificity to a wide range of DNA sites in Drosophila embryos. Genes Dev. 1994 Jul 15;8(14):1678–1692. doi: 10.1101/gad.8.14.1678. [DOI] [PubMed] [Google Scholar]
- Wassarman D. A., Steitz J. A. RNA splicing. Alive with DEAD proteins. Nature. 1991 Feb 7;349(6309):463–464. doi: 10.1038/349463a0. [DOI] [PubMed] [Google Scholar]
- Weintraub H., Genetta T., Kadesch T. Tissue-specific gene activation by MyoD: determination of specificity by cis-acting repression elements. Genes Dev. 1994 Sep 15;8(18):2203–2211. doi: 10.1101/gad.8.18.2203. [DOI] [PubMed] [Google Scholar]
- Wessel D., Flügge U. I. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal Biochem. 1984 Apr;138(1):141–143. doi: 10.1016/0003-2697(84)90782-6. [DOI] [PubMed] [Google Scholar]
- Westin E. H., Gallo R. C., Arya S. K., Eva A., Souza L. M., Baluda M. A., Aaronson S. A., Wong-Staal F. Differential expression of the amv gene in human hematopoietic cells. Proc Natl Acad Sci U S A. 1982 Apr;79(7):2194–2198. doi: 10.1073/pnas.79.7.2194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu C. Analysis of hypersensitive sites in chromatin. Methods Enzymol. 1989;170:269–289. doi: 10.1016/0076-6879(89)70052-5. [DOI] [PubMed] [Google Scholar]