ABSTRACT
The nonenveloped polyomavirus (PyV) simian virus 40 (SV40) traffics from the cell surface to the endoplasmic reticulum (ER), where it penetrates the ER membrane to reach the cytosol before mobilizing into the nucleus to cause infection. Prior to ER membrane penetration, ER lumenal factors impart structural rearrangements to the virus, generating a translocation-competent virion capable of crossing the ER membrane. Here we identify ERdj5 as an ER enzyme that reduces SV40's disulfide bonds, a reaction important for its ER membrane transport and infection. ERdj5 also mediates human BK PyV infection. This enzyme cooperates with protein disulfide isomerase (PDI), a redox chaperone previously implicated in the unfolding of SV40, to fully stimulate membrane penetration. Negative-stain electron microscopy of ER-localized SV40 suggests that ERdj5 and PDI impart structural rearrangements to the virus. These conformational changes enable SV40 to engage BAP31, an ER membrane protein essential for supporting membrane penetration of the virus. Uncoupling of SV40 from BAP31 traps the virus in ER subdomains called foci, which likely serve as depots from where SV40 gains access to the cytosol. Our study thus pinpoints two ER lumenal factors that coordinately prime SV40 for ER membrane translocation and establishes a functional connection between lumenal and membrane events driving this process.
IMPORTANCE PyVs are established etiologic agents of many debilitating human diseases, especially in immunocompromised individuals. To infect cells at the cellular level, this virus family must penetrate the host ER membrane to reach the cytosol, a critical entry step. In this report, we identify two ER lumenal factors that prepare the virus for ER membrane translocation and connect these lumenal events with events on the ER membrane. Pinpointing cellular components necessary for supporting PyV infection should lead to rational therapeutic strategies for preventing and treating PyV-related diseases.
INTRODUCTION
Viruses must penetrate host cell membranes to reach their proper intracellular destination where they replicate their genome, producing viral progenies used for the next round of infection. While enveloped viruses breach host cells by fusing their membrane with a target cell membrane, the mechanism by which nonenveloped viruses penetrate the host cell membrane must be distinct from that of enveloped viruses, as they lack a surrounding membrane. Despite being poorly characterized, a series of biochemical experiments provided a general model describing nonenveloped virus membrane penetration (1–5). In this model, the nonenveloped virus first traffics to the proper site for membrane penetration. Here the viral particle undergoes defined conformational changes induced by host environments and factors (e.g., low pH, proteases, reductases, and chaperones) that either expose hydrophobic moieties buried in the native virus or release small lytic peptides hidden in the intact virion (1–11). In the final step, the hydrophobic viral intermediate (or lytic factor) engages the limiting lipid bilayer, disrupting its integrity and enabling a subviral core particle to cross the membrane, in some cases aided by cellular membrane machineries (12).
The simian polyomavirus (PyV) simian virus 40 (SV40) is a nonenveloped virus that serves as the archetype for studying PyV entry. Well-established human PyVs known to cause debilitating human diseases include JC PyV (JCV) and BK PyV (BKV) (13). Structurally, the major capsid protein VP1 of SV40 forms 72 pentamers arranged as an icosahedral particle that encapsulates its double-stranded DNA genome (14, 15). Each VP1 pentamer also interacts with one copy of the internal protein VP2 or VP3 through hydrophobic interactions (16). Three major forces stabilize the overall capsid structure (14, 15). First, the C terminus of each VP1 protein “invades” a neighboring VP1 pentamer and makes extensive contacts with it. Second, a complex disulfide bond network is formed among the VP1 pentamers that further stabilize the capsid structure. Third, calcium ions bind to negatively charged residues in VP1, further supporting the overall viral architecture. When fully assembled, SV40 displays a near-spherical geometry, with a diameter of ∼45 nm (14, 15).
To infect cells, SV40 binds to the ganglioside GM1 receptor localized on the plasma membrane (17). This interaction induces membrane invagination (18), allowing the virus to enter cells via a lipid raft-dependent endocytic pathway (17–20). The virus is then transported through the classic endolysosome system, where it is sorted to the endoplasmic reticulum (ER) (21–23). In the ER, SV40 experiences conformational changes that enable it to penetrate the ER membrane and reach the cytosol. However, despite these conformational changes, it remains large and intact when crossing the ER membrane (24). Interestingly, thin-section electron microscopy (EM) analysis of ER-localized SV40 in infected cells suggests that the virus may become smaller in the ER, decreasing from its native diameter of 45 nm to 34 nm (25). Thus, the precise nature of SV40 in the ER is unclear. Regardless, upon reaching the cytosol, the virus undergoes significant disassembly to release the VP1 pentamers (24). The resulting smaller core viral particle harboring the DNA genome enters the nucleus to cause lytic infection or cellular transformation (26–28). In the context of this entry pathway, the molecular mechanism by which SV40 in the ER is primed for membrane transport is yet to be fully clarified.
One model posits that disrupting SV40's interpentamer disulfide bonds and unfolding VP1's C-terminal invading arm would induce a conformational change that destabilizes the viral particle, leading to the exposure of SV40's hydrophobic VP2/VP3 internal proteins (29). The resulting hydrophobic virus in turn integrates into the ER membrane, engaging selective ER membrane components that promote translocation into the cytosol. In accordance with this model, an ER lumenal protein disulfide isomerase (PDI) family member called ERp57 was shown to isomerize a subset of SV40's interpentamer disulfide bonds, while the canonical PDI itself is thought to unfold SV40 (30). Furthermore, the ER membrane protein BAP31 was found to interact with hydrophobic SV40 via the exposed VP2 protein (25). Despite these findings, what remains unknown is whether other PDI family members disrupt SV40's disulfide bonds and whether members within this family might cooperate with each other to fully prepare the virus for ER membrane penetration. Additionally, a functional link between lumenal and membrane events essential for ejecting SV40 from the ER to the cytosol has not been demonstrated.
Here we demonstrate that the PDI family member ERdj5 reduces SV40's disulfide bonds to prime the virus for ER membrane transport and infection. This reductase acts in concert with PDI to prepare SV40 for membrane penetration. Negative-stain EM analysis suggests that ERdj5 and PDI induce structural alterations to the virus. Importantly, this conformational change is required for SV40 to interact with BAP31. Our study thus identifies a new PDI family member that cooperates with PDI to facilitate SV40 ER membrane penetration and establishes a link between lumenal and membrane events essential for the translocation process.
MATERIALS AND METHODS
Materials.
Polyclonal Hsp90 and SV40 large T antigen antibodies were purchased from Santa Cruz Biotechnology (Santa Cruz, CA), polyclonal S tag and monoclonal PDI antibodies were obtained from Abcam (Cambridge, MA), monoclonal BiP antibody was obtained from BD (Franklin, NJ), monoclonal FLAG antibody was obtained from Sigma-Aldrich (St. Louis, MO), polyclonal ERdj5 antibody was obtained from Protein Tech Group (Chicago, IL), and monoclonal BAP31 antibody was obtained from Thermo Fisher Scientific (Waltham, MA). Monoclonal c-Myc antibody was isolated from 9E10 hybridomas. Monoclonal antibodies against SV40 VP1 were generous gifts from W. Scott (University of Miami). FLAG M2 antibody-conjugated agarose beads, FLAG peptides, Triton X-100, octyl-α-d-glucopyranoside, N-ethylmaleimide (NEM), and dithiothreitol (DTT) were purchased from Sigma-Aldrich; digitonin and S protein-conjugated agarose beads were obtained from EMD Millipore Chemicals (San Diego, CA); protein G magnetic beads were obtained from Life Technologies (Carlsbad, CA); and dithiobis-succinimidyl propionate (DSP) was obtained from Thermo Fisher Scientific. Purified BKV and pAb416 antibodies against BKV large T antigen were kindly provided by Michael Imperiale (University of Michigan).
siRNA transfection.
The small interfering RNAs (siRNAs) used in this study are as follows: PDI siRNA (5′-GACCUCCCCUUCAAAGUUGTT-3′) (Invitrogen), ERdj5 siRNA 1 (5′-GGACAAGGAACCAAAGAATT-3′) (Invitrogen), and ERdj5 siRNA 2 (5′-GCUCUUGGCUAGGAUGAUUTT-3′) (Invitrogen).
Duplex siRNA (30 or 50 nM for PDI and 10 nM for ERdj5) was reverse transfected into cells by using Lipofectamine RNAiMAX (Life Technologies) according to the manufacturer's protocol. Allstar negative-control siRNA (Qiagen, Hilden, Germany) was used as a control scrambled siRNA.
SV40 preparation, ER-to-cytosol transport, and infection assays.
SV40 preparation, ER-to-cytosol transport, and infection assays were performed as described previously by Inoue and Tsai (24, 31). Approximately 1,000 cells were scored in each experiment, and each experiment was repeated at least three times.
Rescue experiment.
CV-1 cells transfected with siRNA against PDI were further transfected with a different DNA construct by using Fugene HD (Promega, Madison, WI), incubated for 24 h, and infected with SV40. At 20 h postinfection (p.i.), cells were fixed, and infection was assessed by immunofluorescence microscopy, as previously described (31). In FLAG-positive cells, the extent of T antigen expression in the nuclei was scored. For the siRNA-resistant PDI construct, the following silent mutations were introduced into the C-terminally FLAG-tagged mouse PDI construct by overlapping PCR: 650-AATTTCGAGGGTGAAATA-666 (underlining indicates the introduced silent mutations). At least 100 cells were scored in each experiment.
Purification of recombinant proteins.
FLAG-tagged ERdj5 and green fluorescent protein (GFP) were purified as described previously by Williams et al. (32), with minor modifications. ERdj5-bound beads were treated with 1 mM reduced glutathione, washed, and incubated with FLAG peptide to elute the bound proteins.
In vitro SV40 reduction assay.
Purified SV40 (50 ng) was incubated with GFP-FLAG (1 μM) or ERdj5-FLAG (1 μM) at 37°C for 30 min, followed by the addition of SDS sample buffer containing 10 mM NEM. Samples were analyzed by nonreducing SDS-PAGE and immunoblotted.
Isolation of endosome- and ER-localized SV40 from infected cells.
For isolation of endosome-localized SV40, cells were infected with SV40 (multiplicity of infection [MOI] of ∼30) for 2 h in a synchronized manner, harvested, treated with 0.1% digitonin, and centrifuged at 16,100 × g for 10 min. The resulting supernatant fraction containing SV40 in endosomes was further centrifuged at 50,000 × g in a Beckman TLA100 rotor for 30 min, and the resulting pellet fraction was resuspended in a buffer containing 50 mM HEPES (pH 7.5), 150 mM NaCl, 1% Triton X-100, and 1% octyl-d-glucopyranoside to solubilize membranes. Following solubilization, SV40 was immunoprecipitated with SV40 monoclonal antibodies (CC10 and BC11), and the immune complexes captured by magnetic beads. For isolation of ER-localized SV40, cells infected with SV40 (MOI of ∼30) for 6 or 12 h were treated with 0.1% digitonin and centrifuged at 16,100 × g for 10 min. The resulting pellet fraction was further treated with a buffer containing 50 mM HEPES (pH 7.5), 150 mM NaCl, and 1% Triton X-100 and centrifuged. The Triton X-100-extracted material was further fractionated by Optiprep gradient centrifugation, as described previously by Goodwin et al. (33), and subjected to immunoprecipitation with SV40 monoclonal antibodies. For samples transfected with siRNA, the Optiprep gradient centrifugation step was omitted, and ER-localized SV40 was isolated directly from the Triton X-100-extracted material. As a control, wild-type (WT) SV40 was incubated in a buffer containing 50 mM HEPES (pH 7.5), 150 mM NaCl, and 1% Triton X-100, followed by immunoprecipitation as described above but without the Optiprep gradient centrifugation step.
Coprecipitation of ER-localized SV40 with S-tagged BiP.
CV-1 cells transfected with S-tagged BiP using Fugene HD (Promega, Madison, WI) were reverse transfected with scrambled or ERdj5 siRNA 1 for 24 h and infected with SV40 (MOI of ∼10) for 6 h. Cells were subjected to the fractionation method outlined in Fig. 1A, and the resulting Triton X-100-extracted material containing ER-localized SV40 was incubated with S protein-conjugated agarose beads at 4°C for 2 h. The bound proteins were washed, eluted with SDS sample buffer, and subjected to SDS-PAGE followed by immunoblotting with the appropriate antibodies.
FIG 1.
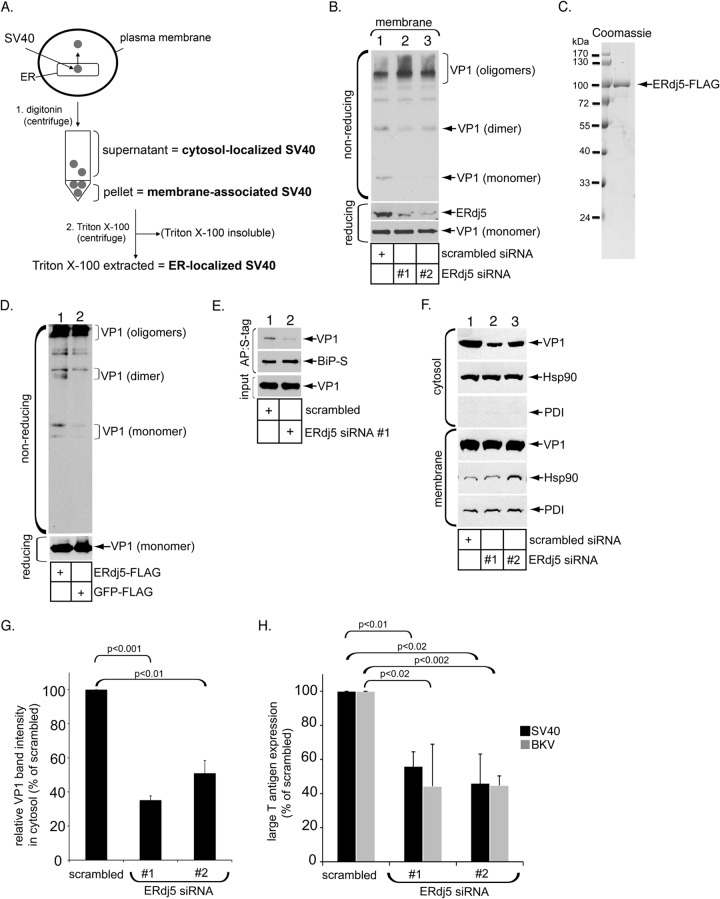
ERdj5 promotes SV40 disulfide bond reduction, ER membrane transport, and infection. (A) Flow diagram of the ER-to-cytosol transport assay depicting the cell fractionation protocol used in this study. In step 1, SV40-infected cells are treated with digitonin and centrifuged to generate a supernatant fraction (cytosol) containing cytosol-localized SV40 and a pellet fraction (membrane) containing membrane-associated SV40. In step 2, the pellet fraction can be further treated with Triton X-100 to extract ER-localized SV40 (Fig. 3). (B) CV-1 cells transfected with scrambled siRNA, ERdj5 siRNA 1, or ERdj5 siRNA 2 were infected with SV40 and fractionated as outlined above for panel A to generate Triton X-100-extracted material containing ER-localized SV40. Samples were subjected to reducing or nonreducing SDS-PAGE followed by immunoblotting with VP1-specific antibodies. (C) Coomassie staining of C-terminally FLAG-tagged ERdj5 (ERdj5-FLAG) purified from HEK 293T cells. (D) WT SV40 was incubated with ERdj5-FLAG or GFP-FLAG and subjected to nonreducing SDS-PAGE followed by immunoblotting with VP1-specific antibodies. (E) CV-1 cells expressing BiP-S and transfected with either scrambled siRNA or ERdj5 siRNA 1 were infected with SV40 for 6 h, harvested, and fractionated as described above for panel A. The resulting Triton X-100-extracted material containing ER-localized SV40 was incubated with S protein-conjugated agarose beads, and the isolated proteins were subjected to SDS-PAGE followed by immunoblotting with anti-VP1 and anti-S tag antibodies. AP, affinity purification. (F) CV-1 cells transfected with scrambled siRNA, ERdj5 siRNA 1, or ERdj5 siRNA 2 were infected with SV40 for 12 h and subjected to fractionation as outlined above for panel A. The cytosol and membrane fractions were analyzed by reducing SDS-PAGE and immunoblotted with the indicated antibodies. (G) The VP1 band intensity in the cytosol fraction shown in panel F was quantified with ImageJ (NIH). Data represent the means ± standard deviations of data from at least 3 independent experiments. A two-tailed t test was used. (H) Cells transfected with scrambled siRNA, ERdj5 siRNA 1, or ERdj5 siRNA 2 were infected with SV40 for 24 h or BKV for 48 h, fixed, and subjected to immunofluorescence staining using a SV40 or BKV large T antigen antibody. Large T antigen-positive nuclei were scored. Data represent the means ± standard deviations of data from at least 3 independent experiments. A two-tailed t test was used.
EM analyses.
SV40-bound magnetic beads were directly applied onto a glow-discharged copper grid. The grid was washed with water three times and stained with 0.75% uranyl formate. The samples were observed with a Morgagni transmission electron microscope at 120 kV.
XBP1 splicing.
XBP1 splicing was performed as described previously by Uemura et al. (34), except that CV-1 cells were used.
Coprecipitation of ER-localized SV40 with endogenous BAP31.
Cells transfected with scrambled siRNA, PDI siRNA/ERdj5 siRNA 1 were infected with SV40 (MOI of ∼10) for 16 h, washed with Hanks balanced salt solution (HBSS), and incubated in HBSS containing 0.2 mM DSP at 37°C for 30 min. Following extensive washes with phosphate-buffered saline (PBS), cells were harvested and lysed in a buffer containing 50 mM HEPES (pH 7.5), 150 mM NaCl, 1% Triton X-100, 10 mM NEM, and 1 mM phenylmethylsulfonyl fluoride (PMSF). The resulting whole-cell lysate was incubated with a c-Myc or BAP31 antibody, and the immunoprecipitates were captured with protein G magnetic beads. After the beads were extensively washed with a buffer containing 50 mM Tris (pH 7.5), 150 mM NaCl, 1% Triton X-100, and 0.5% sodium deoxycholate, the bound proteins were eluted with SDS sample buffer and subjected to SDS-PAGE followed by immunoblotting.
Visualization of SV40-induced focus formation.
Cells transfected with the indicated siRNAs were infected with SV40 (MOI of ∼10) for 16 h, fixed, and processed by using the immunofluorescence methods described previously by Walczak et al. (12).
RESULTS
ERdj5 promotes SV40 disulfide bond reduction, ER membrane transport, and infection.
Despite the ER being oxidized, the PDI family protein ERdj5 was reported to act as an ER-resident reductase in this environment (35), reducing disulfide bonds within substrates. Accordingly, we asked if ERdj5 reduces SV40's disulfide bonds during infection. Simian CV-1 cells transfected with a control (scrambled) siRNA or either of the two ERdj5 siRNAs (siRNA 1 or 2) were infected with SV40 for 12 h. SV40 in membranes (including the ER) was then isolated from these cells by using an established protocol designed to monitor SV40 ER-to-cytosol transport (24), as outlined in Fig. 1A. Briefly, infected cells were treated with the detergent digitonin to semipermeabilize the plasma membrane without damaging internal membranes. Centrifugation of the cells generates two fractions, a supernatant fraction (cytosol), which contains cytosolic proteins and SV40 that reached the cytosol from the ER, and a pellet fraction (membrane), which harbors membranes, including the ER, and membrane-associated SV40.
The membrane fractions derived from control and ERdj5-knocked-down cells infected with SV40 were subjected to nonreducing or reducing SDS-PAGE followed by immunoblotting. Strikingly, under nonreducing conditions, less of the VP1 monomer was observed in ERdj5 knockdown cells than in control cells (Fig. 1B, top, compare lanes 2 and 3 to lane 1). This decrease in the amount of the VP1 monomer was accompanied by a corresponding increase in the amount of VP1 oligomers in the ERdj5-depleted cells. The total amount of VP1 in the membrane fraction, as assessed by the VP1 level in reducing SDS-PAGE gels, was the same between control and ERdj5 knockdown cells (Fig. 1B, bottom, compare lanes 2 and 3 to lane 1). These findings suggest that ERdj5 reduces SV40 to generate disulfide bond-free VP1 monomers during infection. In vitro, the formation of VP1 monomers (and dimers) was readily detected when SV40 was incubated with ERdj5 (FLAG-tagged ERdj5) purified from mammalian cells (Fig. 1C) but not with the control GFP-FLAG (Fig. 1D, compare lane 1 to lane 2). Together, these findings demonstrate that ERdj5 acts as a reductase to reduce the disulfide bonds present in SV40.
We next asked if the ERdj5-dependent reduction of SV40's disulfide bonds affects the ability of SV40 to bind to the ER-resident Hsp70 BiP (31, 33). Affinity purification (AP) of S-tagged BiP (BiP-S) derived from infected cells transfected with either scrambled siRNA or ERdj5 siRNA 1 revealed that knockdown of ERdj5 decreased the SV40-BiP interaction (Fig. 1E, top, compare lane 2 to lane 1). Because BiP binds to hydrophobic moieties present in unfolded/misfolded proteins (36–38), this result suggests that the reduction of SV40's disulfide bonds by ERdj5 triggers a conformational change in SV40 to generate a hydrophobic virus that in turn recruits BiP. We note that as ERdj5 also contains a so-called J domain that stimulates BiP's ATPase activity to promote BiP-substrate interactions (35, 39), the lack of this J domain when ERdj5 is silenced might also contribute to the decreased SV40-BiP interactions.
We hypothesized that the action of ERdj5 on SV40 primes the viral particle for translocation to the cytosol from the ER. To test this, SV40-infected cells transfected with scrambled siRNA or ERdj5 siRNA 1 or 2 were subjected to the semipermeabilized ER-to-cytosol transport assay (Fig. 1A). We found that knockdown of ERdj5 decreased the SV40 level in the cytosol fraction (Fig. 1F, top, compare lanes 2 and 3 to lane 1; the VP1 band intensity is quantified in Fig. 1G). These data indicate that ERdj5 mediates ER-to-cytosol entry of SV40. Consistent with these results, SV40 infection, as assessed by the expression of the virally encoded large T antigen in the host nucleus, was also decreased when ERdj5 was downregulated (Fig. 1H, black bars). This observation extended to human BKV (Fig. 1H, gray bars), demonstrating a more general role of ERdj5 in facilitating PyV infection. Collectively, our findings suggest a scenario in which ERdj5, by reducing disulfide bonds in SV40, prepares the viral particle for ER membrane translocation into the cytosol, leading to successful infection.
ERdj5 cooperates with PDI to stimulate SV40 ER membrane transport and infection.
Because ERdj5 knockdown blocked SV40 ER membrane transport and infection only partially, we speculated that ERdj5 might cooperate with another PDI family member to more robustly support this entry step. Canonical PDI was previously reported to mediate SV40 infection (30). However, in PDI knockdown cells, we found that the level of the VP1 monomer was not significantly affected (Fig. 2A, top, compare lane 2 to lane 1), consistent with data from a previous report (30). These findings suggest that PDI might utilize its nonenzymatic unfoldase activity against SV40 to prime the virus for membrane transport. As silencing of PDI partially impaired SV40 infection (30), we postulated that PDI might act in concert with ERdj5 to fully support the cytosol entry of SV40 from the ER.
FIG 2.
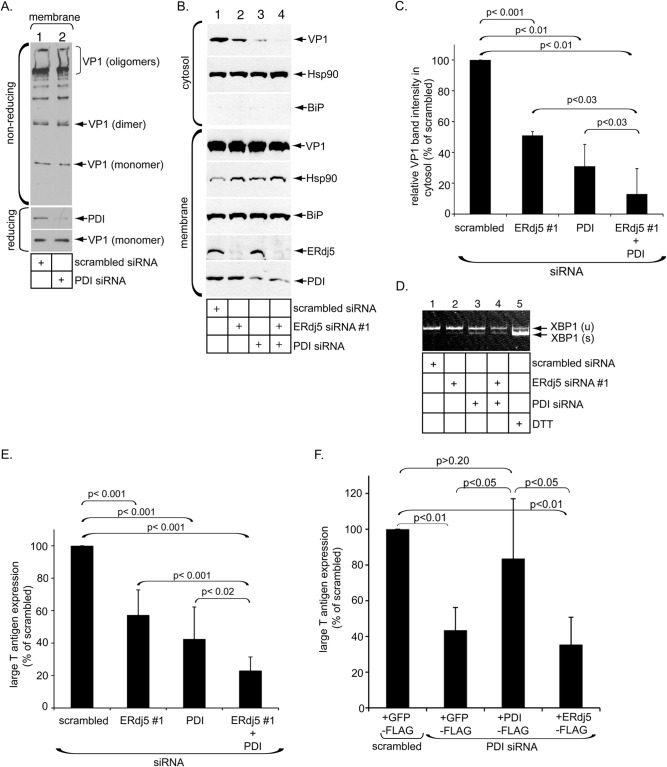
ERdj5 cooperates with PDI to stimulate SV40 ER membrane transport and infection. (A) Same as Fig. 1B except that cells were transfected with PDI siRNA. (B) Same as Fig. 1F except that cells were transfected with scrambled siRNA, ERdj5 siRNA 1, PDI siRNA, or ERdj5 siRNA 1 plus PDI siRNA. The resulting cytosol and membrane fractions were analyzed by immunoblotting with the indicated antibodies. (C) The VP1 band intensity in the cytosol fraction in panel A was quantified as described in the legend of Fig. 1G. Data represent the means ± standard deviations of data from at least 3 independent experiments. A two-tailed t test was used. (D) Reverse transcription-PCR analysis of the unspliced (u) and spliced (s) forms of XBP1 mRNA from cells transfected with the indicated siRNAs or treated with DTT. (E) Same as Fig. 1H except that cells were transfected with scrambled siRNA, ERdj5 siRNA 1, PDI siRNA, or ERdj5 siRNA 1 plus PDI siRNA. A two-tailed t test was used. (F) CV-1 cells initially transfected with scrambled or PDI siRNA were subsequently transfected with GFP-FLAG, PDI-FLAG, or ERdj5-FLAG. Cells were then infected with SV40 and subjected to immunofluorescence analyses using anti-FLAG and anti-T antigen antibodies. FLAG and T antigen double-positive cells were counted and analyzed as described above for panel E. Data represent the means ± standard deviations of data from at least 3 independent experiments. A two-tailed t test was used.
To test this idea, we asked if the simultaneous depletion of ERdj5 and PDI more potently disrupts SV40 ER membrane transport and infection than the single knockdown. Cells transfected with scrambled siRNA, ERdj5 siRNA 1, PDI siRNA, or a combination of ERdj5 siRNA 1 and PDI siRNA were infected with SV40 and subjected to the semipermeabilized ER-to-cytosol transport assay. Indeed, we found that the concurrent depletion of ERdj5 and PDI led to a more severe block in the arrival of SV40 in the cytosol than did the single knockdown (Fig. 2B, top, compare lane 4 to lanes 2 and 3; the VP1 band intensity is quantified in Fig. 2C). No significant XBP1 splicing, a sensitive method to monitor ER stress, was observed with the double knockdown (Fig. 2D, compare lane 4 to lanes 1 to 3), in contrast to cells treated with the chemical ER stress inducer dithiothreitol (DTT) (Fig. 2D, compare lane 5 to lane 1). These results suggest that the simultaneous depletion of ERdj5 and PDI did not trigger robust ER stress that might indirectly influence viral membrane transport. As expected, the double knockdown of ERdj5 and PDI markedly impaired SV40 infection compared to their single knockdowns (Fig. 2E). Thus, ERdj5 acts coordinately with PDI to prepare SV40 for successful ER membrane transport and infection.
A knockdown rescue experiment was performed to clarify whether ERdj5 and PDI act in a parallel or linear fashion during SV40 entry. We reasoned that if ERdj5 and PDI are indeed functioning in a parallel pathway, reexpression of either (siRNA-resistant) PDI or ERdj5 in PDI-depleted cells should restore infection because SV40 could use the ERdj5-dependent route when PDI is absent. However, if only PDI but not ERdj5 rescues infection in PDI knockdown cells, the data would suggest that SV40 cannot bypass the PDI-dependent route by relying on the ERdj5-directed pathway. This finding would then indicate that PDI and ERdj5 operate in a linear pathway, where both factors have unique and nonoverlapping roles. Indeed, we found that only PDI but not ERdj5 restored SV40 infection in PDI-depleted cells (Fig. 2F), implicating that ERdj5 and PDI function in a linear pathway and play distinct roles to promote SV40 entry. This interpretation is consistent with our hypothesis that ERdj5 reduces the disulfide bonds in the virus, while PDI unfolds the viral particle.
ERdj5 and PDI impart structural instability to ER-localized SV40, as revealed by EM analysis.
We next used EM analysis to evaluate the impact of ERdj5 and PDI on the conformation of SV40. We first examined the structure of ER-localized SV40 in control cells. ER-localized SV40 can be extracted from the pellet (membrane) fraction by treatment of this fraction with Triton X-100 (Fig. 1A). This is because during entry, SV40 binds to its ganglioside GM1 receptor on the cell surface, remaining attached to it until the virus-receptor complex traffics to the ER, where the virus is released from its receptor. As GM1 is highly concentrated in lipid rafts, SV40 persists in lipid rafts until it reaches the ER, and because lipid rafts are resistant to Triton X-100 extraction, while ER lumenal proteins are extracted by this detergent, SV40 extracted from the pellet fraction by Triton X-100 represents virus that reached the ER, detached from its receptor, and was released into the ER lumen (24).
Because SV40 begins to arrive at the ER from the cell surface at ∼6 h p.i. (24, 30), CV-1 cells were infected with SV40 for either 6 or 12 h, and the ER-localized virus was extracted by using the above-mentioned methods. Next, monoclonal VP1 antibodies were added to this sample to immunoprecipitate the virus, and the immune complex was captured with protein G-coated magnetic beads. When SV40 bound to magnetic beads is applied onto EM grids, a pool of the viral particles is released spontaneously to the grid surface. Although the efficiency of spontaneous release is low, this process largely preserves the structural integrity of the viral particles, which would otherwise be compromised if harsher methods were used to elute the viral particles from the immune complex. The released particles were embedded in negative stain and subsequently visualized by EM. As a control, input wild-type (WT) SV40 was incubated with the same extraction buffer, isolated, and subjected to EM analysis. To verify the quality of SV40 after immunoprecipitation, the samples were eluted with SDS and subjected to silver staining. VP1 monomers, VP1 oligomers, and, to a lesser extent, VP2 and VP3 were present in the samples (Fig. 3A, lanes 2 and 3). A band at ∼80 kDa coprecipitated with ER-localized SV40 (6 and 12 h p.i.) but not WT SV40 (Fig. 3A, compare lanes 2 and 3 to lane 1); this band was previously identified as the ER-resident BiP (33). These findings support the premise that precipitated SV40 is localized predominantly at the ER. As SV40 transits through the endosome before reaching the ER (21), we also isolated endosome-localized SV40 for structural comparison (see Materials and Methods for the isolation procedure).
FIG 3.
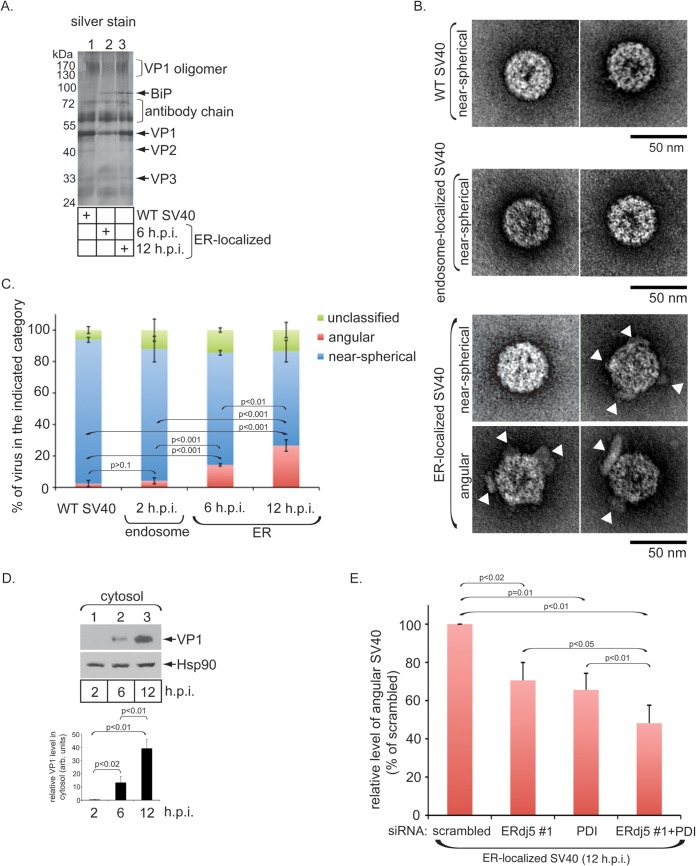
ERdj5 and PDI impart structural instability to ER-localized SV40, as revealed by EM analysis. (A) ER-localized SV40 (at 6 and 12 h p.i.) was immunoprecipitated with VP1-specific antibodies from the Triton X-100-extracted material, and the immunoprecipitate was captured by magnetic beads and subjected to SDS-PAGE followed by silver staining. WT SV40 (incubated with a buffer containing Triton X-100) was also subjected to the same immunoprecipitation method. (B) WT SV40, endosome-localized SV40 (at 2 h p.i.) (see Materials and Methods for the method of isolation of endosome-localized virus), and ER-localized SV40 were negatively stained and visualized by EM. Shown are representative images of viruses from each category. White arrowheads point to virus-associated materials. (C) Quantification of WT, endosome-localized (2 h p.i.), and ER-localized (6 and 12 h p.i.) SV40 as near-spherical, angular, or unclassified particle projections. Data represent the means ± standard deviations of data from at least 3 independent experiments. Each experiment was performed with ∼100 viral particles. A two-tailed t test was used. (D) VP1 levels in the cytosol fraction derived from cells infected with SV40 for the indicated times (immunoblots) were quantified (graph). Data represent the means ± standard deviations of data from at least 3 independent experiments. A two-tailed t test was used. (E) ER-localized SV40 (12 h p.i.) isolated from cells transfected with the indicated siRNAs was subjected to EM analysis as described above for panel B. Angular particles were analyzed and scored as described above for panel C. Data are normalized to the level of angular particles found in the scrambled control. A two-tailed t test was used.
Input WT SV40 was first analyzed. The negative-stain EM projection images of the structure of SV40 revealed that the majority of the virus (>93%) appeared as near-spherical particles ∼45 nm in diameter, as previously described (40, 41) (Fig. 3B and C and 4). Quantification of WT SV40 EM images represents data from at least three independent experiments, with analysis of at least 100 viral particles in each experiment. A very small fraction (<2%) was not near spherical but instead appeared more angular, displaying an enhanced hexagonal projection profile (see below). While the actual three-dimensional (3D) structure of the viral particle represented by virus with this enhanced hexagonal projection is unknown, we refer to this category as “angular” SV40 for simplicity. A small WT SV40 pool (<5%) was found to be distorted and was categorized as “unclassified” (Fig. 3C). Importantly, if WT SV40 was placed directly onto the EM grid, the same proportions of virus in each category were observed (not shown), indicating that our immunoprecipitation protocol did not artificially introduce any structural alterations to the viral particles. We note that the surfaces of the majority of the WT SV40 particles were smooth but infrequently displayed visible associated material of unknown identity, likely representing attached proteins and lipids (Fig. 4, far right, arrowheads).
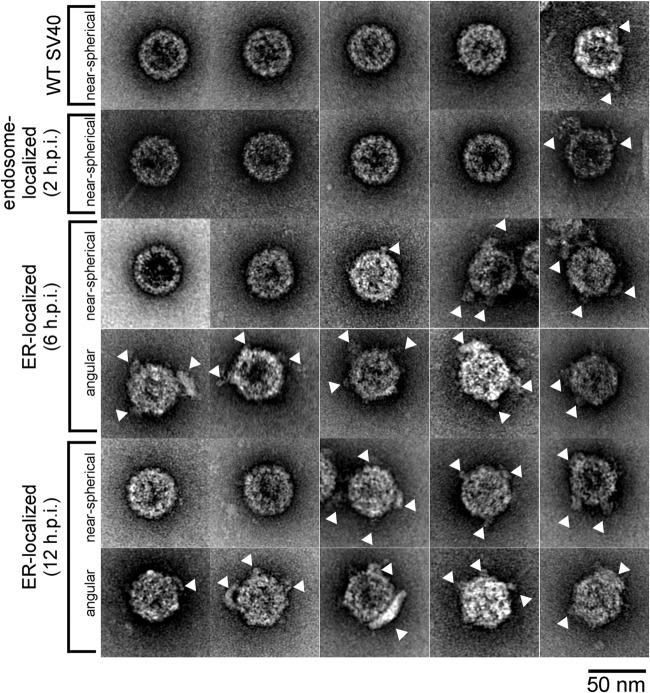
FIG 4.
Additional representative EM images of WT, endosome-localized (2 h p.i.), and ER-localized (6 and 12 h p.i.) SV40 particles. White arrowheads point to virus-associated materials.
What conformational changes are imparted to native SV40 when it reaches the endosome? Similarly to WT SV40, EM analysis indicated that the majority of endosome-localized SV40 particles are near spherical (Fig. 3B and C and 4), with only a very small pool that is categorized as angular or unclassified (Fig. 3C). The average diameter of these particles is similar to that of WT SV40. Again, comparable to WT SV40, the surfaces of most endosome-localized virus particles were smooth but occasionally exhibited associated material (Fig. 4, far right, arrowheads). Thus, when SV40 reaches the endosomes, any conformational changes that it might experience are likely subtle and not readily observed by low-resolution EM visualization.
From endosomes, SV40 sorts to the ER. We therefore asked whether ER-localized SV40 experiences any significant structural changes. While a majority of the ER-localized virus particles retained their near-spherical shape (Fig. 3B, top, and C and 4), there was a significant increase in the fraction of angular SV40 particles in ER-localized SV40 particles isolated after 6 h p.i. (15%) and 12 h p.i. (27%) compared to that of WT SV40 particles (<2%) (Fig. 3B, middle, and C and 4). When ER-localized SV40 isolated after 16 h p.i. was analyzed, the fraction of angular SV40 particles did not increase beyond 27% (not shown). Importantly, the average diameter of ER-localized SV40 particles was similar to those of the WT and endosome-localized virus particles, supporting our previous report that SV40 does not undergo significant disassembly in the ER (24). Virus surface-associated material was more frequently observed in ER-localized SV40 (Fig. 3B, top and middle, arrowheads, and Fig. 4, arrowheads). Because negative-stain EM visualization records projection images of dehydrated and partially flattened specimens, whether the observed angular-shaped SV40 particle is indeed a specific morphology generated in the ER requires additional investigation. Nonetheless, EM analysis of ER-localized SV40 and comparison with WT/endosome-localized SV40 demonstrate that the angular-shaped ER-localized SV40 particles are at least indicative of structural rearrangements imposed on the virus in the ER.
Using the cell-based ER-to-cytosol membrane transport assay, we found that the time-dependent increased formation of angular SV40 observed by EM analysis correlates strongly with a similar time-dependent increase in virus arrival to the cytosol (Fig. 3D). These findings raise the possibility that angular SV40 particles may act as a functional intermediate during ER membrane penetration.
To test this idea, we asked whether depletion of ERdj5, PDI, or both might prevent the formation of angular viral particles, given that these ER components can impart the conformational instability to SV40 that is necessary for its transport to the ER membrane and infection. Whereas silencing of ERdj5 or PDI modestly decreased the formation of the angular particles (Fig. 3E), the simultaneous knockdown of both factors more effectively impaired the formation of angular viral particles (Fig. 3E). These findings suggest that ERdj5 and PDI act in a cooperative manner to impart structural changes to ER-localized SV40, causing the formation of the SV40 hexagonal particle projections observed by EM. As ERdj5 and PDI promote SV40 ER-to-cytosol transport and infection, we propose that the destabilized virions observed as angular particles represent membrane penetration intermediates essential for the ER membrane translocation process. While the amount of angular particles formed in the ER was not large (only 27%), it must be noted that that successful arrival of one SV40 particle to the host nucleus from the cytosol is sufficient to promote infection (42).
Physical and functional relationship between ERdj5/PDI and the ER membrane protein BAP31.
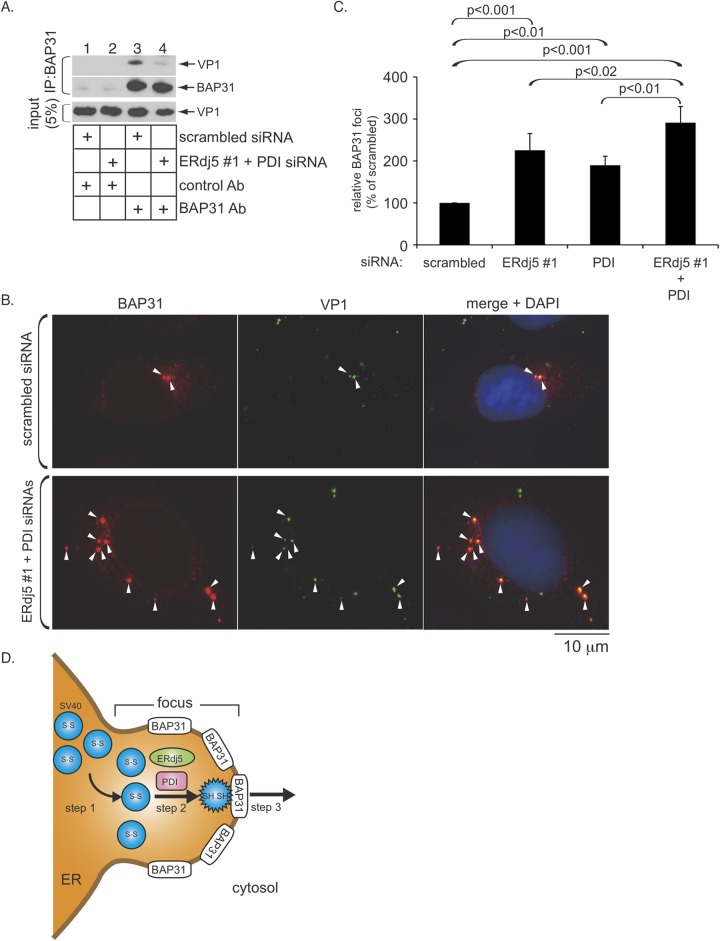
Our data thus far depict a scenario in which ERdj5 and PDI coordinately act on SV40 in the ER lumen to prepare the virus for ER membrane translocation. A previous report suggested that during infection, SV40 induces the ER membrane protein BAP31 to accumulate in discrete ER subdomains called foci, where the virus also accumulates (25). Here hydrophobic SV40 binds to BAP31 and becomes poised for ejection to the cytosol. We therefore asked if the actions of ERdj5 and PDI on SV40 prepare the virus for subsequent interactions with BAP31. Coimmunoprecipitation experiments revealed that knockdown of ERdj5 and PDI significantly blocked the BAP31-SV40 interaction (Fig. 5A, top, compare lane 4 to lane 3), suggesting that the activities of these PDI family proteins on SV40 in the ER lumen are necessary for the virus to engage BAP31 on the ER membrane.
FIG 5.
Physical and functional relationship between ERdj5/PDI and the ER membrane protein BAP31. (A) Cells transfected with the indicated siRNAs were infected with SV40 for 16 h, treated with DSP for 30 min, and harvested. The resulting whole-cell lysate was immunoprecipitated (IP) with a control anti-c-Myc tag (9E10) or anti-BAP31 antibody (Ab). The immunoprecipitates were subjected to SDS-PAGE and analyzed by immunoblotting with the indicated antibodies. (B) Cells transfected with the indicated siRNAs were infected with SV40 for 16 h, fixed, and subjected to immunofluorescence staining with anti-BAP31 and -VP1 antibodies. DAPI, 4′,6-diamidino-2-phenylindole. (C) Numbers of cells with at least one BAP31-positive focus in SV40-infected cells transfected with the indicated siRNAs were determined. Data are normalized to values for scrambled siRNA and represent the means ± standard deviations of data from at least 3 independent experiments. A two-tailed t test was used. (D) Model depicting the ERdj5- and PDI-dependent conformational change of SV40 and its relationship to the BAP31 ER membrane protein (see the text).
Finally, we reasoned that by uncoupling the SV40-BAP31 interaction (via depletion of ERdj5 and PDI), the virus should remain trapped in the foci, thereby enhancing the overall focus structure. In cells transfected with scrambled siRNA, SV40 infection caused BAP31 to accumulate in foci, as previously described (25) (Fig. 5B, top); the foci contained SV40, as revealed by colocalization with the VP1 signal (Fig. 5B, top). Strikingly, in ERdj5/PDI knockdown cells, the focus structure was markedly enhanced. In fact, under knockdown conditions, more foci were observed per cell, and the size of the foci appeared to be larger than those observed under control conditions (Fig. 5B, compare bottom to top). Quantification revealed that the number of cells containing at least one BAP31-positive focus increased by ∼3-fold when ERdj5 and PDI were concurrently depleted (Fig. 5C); depletion of single component resulted in a more modest effect (Fig. 5C). These results establish a physical and functional link between ER lumenal and membrane events that are coupled to drive SV40 ER membrane translocation.
DISCUSSION
A summary of our model describing how ERdj5 and PDI promote SV40 penetration of the ER membrane is depicted in Fig. 5D. After reaching the ER, SV40 is first delivered to discrete ER subdomains containing accumulated BAP31, called foci, which serve as depots for the virus prior to ER membrane translocation (step 1). Here ERdj5 reduces SV40's disulfide bonds, while PDI likely unfolds the viral particle. These two events coordinately induce conformational changes in the virus, generating a hydrophobic viral particle (step 2). The structurally altered hydrophobic virus binds to BAP31, where it is positioned to cross the ER membrane to reach the cytosol (step 3).
A major finding of this study is that ERdj5 regulates SV40 ER-to-cytosol transport and infection by reducing virus disulfide bonds, reminiscent of its role as a reductase in the ER-associated degradation (ERAD) pathway (35). ERAD is a cellular quality control pathway in which misfolded ER proteins are recognized and ejected to the cytosol for proteasomal degradation (43–45). In many instances, the misfolded substrates contain nonnative disulfide bonds that must be reduced in the ER prior to their retrotranslocation to the cytosol. ERdj5 contains four canonical thioredoxin domains that harbor the redox-active cysteines used during a typical reduction reaction (35, 39). Pinpointing which of these domains is used to reduce SV40's disulfide bonds requires further investigation.
In addition to ERdj5, other PDI family members are also hijacked by PyVs to disrupt their disulfide bonds. In general, the use of a selective PDI family protein appears to depend on the specific PyV or cell system under study. For instance, while ERdj5 reduces SV40's disulfide bonds in permissive simian CV-1 cells to stimulate infection (this study), it does not appear to play a major role in mediating SV40 infection in human HeLa cells (25). Rather, ERp57 is used in HeLa cells to break and isomerize SV40's disulfide bonds (30). Our experiment also revealed a role of ERdj5 in BKV infection, suggesting that this PDI family member reduces BKV's disulfide bonds during infection. Intriguingly, we reported previously that ERp57 isomerizes the disulfide bonds in murine PyV, while PDI reduces them (11); whether ERdj5 acts on this virus is not known. The different disulfide bond arrangements in murine PyV and SV40 (11, 14, 15, 46) might explain the differential use of PDI family members prior to ER membrane translocation.
Another key finding of this study is that ERdj5 acts coordinately with PDI to regulate SV40 ER-to-cytosol transport and infection. We found that silencing of PDI did not affect SV40's disulfide bonds in infected cells, consistent with data from a previous report (30). This result indicates that PDI does not function as a reductase against SV40, in contrast to its action on murine PyV (11). Instead, in addition to its enzymatic function, PDI is also known to possess a chaperone activity, unfolding ERAD substrates and the bacterial cholera toxin prior to retrotranslocation to the cytosol (47–50). If PDI's unfoldase activity operates on SV40, how then might this activity cooperate with ERdj5 to regulate SV40 ER-to-cytosol transport? One possibility is that because ERdj5 reduces the interpentamer disulfide bonds of the SV40 capsid to partially liberate some VP1 pentamers from each other, this reaction could destabilize the virus, which in turn allows PDI to more efficiently unfold the virus. Untangling the VP1 C-terminal arm using PDI's unfoldase activity would indeed be one attractive scenario to further destabilize the viral particle. In this context, another PDI family protein, called ERp29, was shown previously to unfold the VP1 C-terminal arm of murine PyV (8). However, this protein does not appear to play any significant role during SV40 infection (30).
Regardless of the precise region of SV40 affected by ERdj5 and PDI, their concerted actions impose a structural rearrangement on the viral particle. This idea is supported by the negative-stain EM images of ER-localized virus, suggesting that ERdj5 and PDI impart structural instability to the virus, as indicated by the decreased formation of the angular-shaped virus in the ER when these host components are depleted. We hypothesize that this conformational change leads to the formation of a hydrophobic virus, in line with the finding that depletion of ERdj5 alone is sufficient to impair the interaction between SV40 and BiP, an ER-resident chaperone with an intrinsic affinity for hydrophobic proteins. Exposure of hydrophobic regions of VP1 hidden in the native virion and extrusion of the hydrophobic VP2/VP3 proteins buried within the VP1 pentamer (see below) are two potential mechanisms to engender a hydrophobic virus.
What might be the functional consequence of this structural rearrangement? Our experiments indicated that the ERdj5- and PDI-dependent conformational change in SV40 enables the virus to bind to BAP31, an ER membrane protein critical for SV40 membrane penetration (25). Because BAP31 interacts with the N terminus of SV40 VP2 (25), the actions of ERdj5 and PDI may lead to the exposure of this minor protein. How VP2 exposure is achieved is unknown, but a recent study of human JCV using mutations that clog the pore of the VP1 pentamer suggests that a portion of VP2 may slip out from this pore (51). Other ERdj5- and PDI-induced structural changes in SV40 that are potentially recognized by BAP31 deserve additional investigation.
Finally, our analyses revealed a previously uncharacterized connection between the activities of ERdj5 and PDI on SV40 and virus-induced foci. Our laboratory and others previously demonstrated that SV40 infection triggered selective ER membrane (e.g., BAP31) and soluble proteins to reorganize into these foci, where the virus also accumulates (12, 25, 31, 52); these foci are thought to represent entry sites from where the viral particle penetrates the cytosol. If this is the case, we envision that impairment of SV40 entry into the cytosol should cause the virus to accumulate in the foci. Indeed, preventing SV40 from entering the cytosol (via depletion of ERdj5 and PDI) entraps the virus in the foci, leading to a significant enhancement of the focus structure. Interestingly, our observation that the focus structure can be enlarged suggests that it has the capacity to “expand” and accommodate numerous viral particles. Precisely how many viral particles are harbored within the foci requires higher-resolution microscopy analysis. Nevertheless, these findings further support the notion that the foci serve as depots where SV40 prepares to enter the cytosol. We note that SV40 can induce focus formation without the ERdj5- and PDI-dependent conformational change that enables the altered virus to bind to BAP31. This indicates that the ability of SV40 to interact with BAP31 does not serve as the trigger for focus formation. We therefore speculate that there is likely an upstream event that targets the virus to the foci; the precise trigger for this reaction remains unknown. Finally, an important question raised by this study that extends beyond the field of viral entry is whether large cellular aggregates in the ER also induce focus formation. If so, the underlying principle guiding SV40 ER-to-cytosol transport may be applicable to the disposal of these aggregates in the cytosol.
ACKNOWLEDGMENTS
B.T. is funded by the NIH (RO1 AI064296-08). This works is also partially supported by the Protein Folding Disease Initiative at the University of Michigan Medical School. G.S. acknowledges support from the Pew Scholars Program in Biomedical Sciences.
REFERENCES
- 1.Chandran K, Farsetta DL, Nibert ML. 2002. Strategy for nonenveloped virus entry: a hydrophobic conformer of the reovirus membrane penetration protein micro 1 mediates membrane disruption. J Virol 76:9920–9933. doi: 10.1128/JVI.76.19.9920-9933.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ivanovic T, Agosto MA, Zhang L, Chandran K, Harrison SC, Nibert ML. 2008. Peptides released from reovirus outer capsid form membrane pores that recruit virus particles. EMBO J 27:1289–1298. doi: 10.1038/emboj.2008.60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Moyer CL, Wiethoff CM, Maier O, Smith JG, Nemerow GR. 2011. Functional genetic and biophysical analyses of membrane disruption by human adenovirus. J Virol 85:2631–2641. doi: 10.1128/JVI.02321-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Trask SD, Kim IS, Harrison SC, Dormitzer PR. 2010. A rotavirus spike protein conformational intermediate binds lipid bilayers. J Virol 84:1764–1770. doi: 10.1128/JVI.01682-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wiethoff CM, Wodrich H, Gerace L, Nemerow GR. 2005. Adenovirus protein VI mediates membrane disruption following capsid disassembly. J Virol 79:1992–2000. doi: 10.1128/JVI.79.4.1992-2000.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bubeck D, Filman DJ, Cheng N, Steven AC, Hogle JM, Belnap DM. 2005. The structure of the poliovirus 135S cell entry intermediate at 10-angstrom resolution reveals the location of an externalized polypeptide that binds to membranes. J Virol 79:7745–7755. doi: 10.1128/JVI.79.12.7745-7755.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bubeck D, Filman DJ, Hogle JM. 2005. Cryo-electron microscopy reconstruction of a poliovirus-receptor-membrane complex. Nat Struct Mol Biol 12:615–618. doi: 10.1038/nsmb955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Magnuson B, Rainey EK, Benjamin T, Baryshev M, Mkrtchian S, Tsai B. 2005. ERp29 triggers a conformational change in polyomavirus to stimulate membrane binding. Mol Cell 20:289–300. doi: 10.1016/j.molcel.2005.08.034. [DOI] [PubMed] [Google Scholar]
- 9.Rainey-Barger EK, Magnuson B, Tsai B. 2007. A chaperone-activated nonenveloped virus perforates the physiologically relevant endoplasmic reticulum membrane. J Virol 81:12996–13004. doi: 10.1128/JVI.01037-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Tsai B. 2007. Penetration of nonenveloped viruses into the cytoplasm. Annu Rev Cell Dev Biol 23:23–43. doi: 10.1146/annurev.cellbio.23.090506.123454. [DOI] [PubMed] [Google Scholar]
- 11.Walczak CP, Tsai B. 2011. A PDI family network acts distinctly and coordinately with ERp29 to facilitate polyomavirus infection. J Virol 85:2386–2396. doi: 10.1128/JVI.01855-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Walczak CP, Ravindran MS, Inoue T, Tsai B. 2014. A cytosolic chaperone complexes with dynamic membrane J-proteins and mobilizes a nonenveloped virus out of the endoplasmic reticulum. PLoS Pathog 10:e1004007. doi: 10.1371/journal.ppat.1004007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.DeCaprio JA, Garcea RL. 2013. A cornucopia of human polyomaviruses. Nat Rev Microbiol 11:264–276. doi: 10.1038/nrmicro2992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Liddington RC, Yan Y, Moulai J, Sahli R, Benjamin TL, Harrison SC. 1991. Structure of simian virus 40 at 3.8-A resolution. Nature 354:278–284. doi: 10.1038/354278a0. [DOI] [PubMed] [Google Scholar]
- 15.Stehle T, Gamblin SJ, Yan Y, Harrison SC. 1996. The structure of simian virus 40 refined at 3.1 A resolution. Structure 4:165–182. doi: 10.1016/S0969-2126(96)00020-2. [DOI] [PubMed] [Google Scholar]
- 16.Chen XS, Stehle T, Harrison SC. 1998. Interaction of polyomavirus internal protein VP2 with the major capsid protein VP1 and implications for participation of VP2 in viral entry. EMBO J 17:3233–3240. doi: 10.1093/emboj/17.12.3233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Tsai B, Gilbert JM, Stehle T, Lencer W, Benjamin TL, Rapoport TA. 2003. Gangliosides are receptors for murine polyoma virus and SV40. EMBO J 22:4346–4355. doi: 10.1093/emboj/cdg439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ewers H, Romer W, Smith AE, Bacia K, Dmitrieff S, Chai W, Mancini R, Kartenbeck J, Chambon V, Berland L, Oppenheim A, Schwarzmann G, Feizi T, Schwille P, Sens P, Helenius A, Johannes L. 2010. GM1 structure determines SV40-induced membrane invagination and infection. Nat Cell Biol 12:11–18. doi: 10.1038/ncb1999. [DOI] [PubMed] [Google Scholar]
- 19.Anderson HA, Chen Y, Norkin LC. 1996. Bound simian virus 40 translocates to caveolin-enriched membrane domains, and its entry is inhibited by drugs that selectively disrupt caveolae. Mol Biol Cell 7:1825–1834. doi: 10.1091/mbc.7.11.1825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Damm EM, Pelkmans L, Kartenbeck J, Mezzacasa A, Kurzchalia T, Helenius A. 2005. Clathrin- and caveolin-1-independent endocytosis: entry of simian virus 40 into cells devoid of caveolae. J Cell Biol 168:477–488. doi: 10.1083/jcb.200407113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Engel S, Heger T, Mancini R, Herzog F, Kartenbeck J, Hayer A, Helenius A. 2011. Role of endosomes in simian virus 40 entry and infection. J Virol 85:4198–4211. doi: 10.1128/JVI.02179-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kartenbeck J, Stukenbrok H, Helenius A. 1989. Endocytosis of simian virus 40 into the endoplasmic reticulum. J Cell Biol 109:2721–2729. doi: 10.1083/jcb.109.6.2721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Pelkmans L, Kartenbeck J, Helenius A. 2001. Caveolar endocytosis of simian virus 40 reveals a new two-step vesicular-transport pathway to the ER. Nat Cell Biol 3:473–483. doi: 10.1038/35074539. [DOI] [PubMed] [Google Scholar]
- 24.Inoue T, Tsai B. 2011. A large and intact viral particle penetrates the endoplasmic reticulum membrane to reach the cytosol. PLoS Pathog 7:e1002037. doi: 10.1371/journal.ppat.1002037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Geiger R, Andritschke D, Friebe S, Herzog F, Luisoni S, Heger T, Helenius A. 2011. BAP31 and BiP are essential for dislocation of SV40 from the endoplasmic reticulum to the cytosol. Nat Cell Biol 13:1305–1314. doi: 10.1038/ncb2339. [DOI] [PubMed] [Google Scholar]
- 26.Kuksin D, Norkin LC. 2012. Disassembly of simian virus 40 during passage through the endoplasmic reticulum and in the cytoplasm. J Virol 86:1555–1562. doi: 10.1128/JVI.05753-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kuksin D, Norkin LC. 2012. Disassociation of the SV40 genome from capsid proteins prior to nuclear entry. Virol J 9:158. doi: 10.1186/1743-422X-9-158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Nakanishi A, Shum D, Morioka H, Otsuka E, Kasamatsu H. 2002. Interaction of the Vp3 nuclear localization signal with the importin alpha 2/beta heterodimer directs nuclear entry of infecting simian virus 40. J Virol 76:9368–9377. doi: 10.1128/JVI.76.18.9368-9377.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Inoue T, Tsai B. 2013. How viruses use the endoplasmic reticulum for entry, replication, and assembly. Cold Spring Harb Perspect Biol 5:a013250. doi: 10.1101/cshperspect.a013250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Schelhaas M, Malmstrom J, Pelkmans L, Haugstetter J, Ellgaard L, Grunewald K, Helenius A. 2007. Simian virus 40 depends on ER protein folding and quality control factors for entry into host cells. Cell 131:516–529. doi: 10.1016/j.cell.2007.09.038. [DOI] [PubMed] [Google Scholar]
- 31.Inoue T, Tsai B. 2015. A nucleotide exchange factor promotes endoplasmic reticulum-to-cytosol membrane penetration of the nonenveloped virus simian virus 40. J Virol 89:4069–4079. doi: 10.1128/JVI.03552-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Williams JM, Inoue T, Banks L, Tsai B. 2013. The ERdj5-Sel1L complex facilitates cholera toxin retrotranslocation. Mol Biol Cell 24:785–795. doi: 10.1091/mbc.E12-07-0522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Goodwin EC, Lipovsky A, Inoue T, Magaldi TG, Edwards AP, Van Goor KE, Paton AW, Paton JC, Atwood WJ, Tsai B, DiMaio D. 2011. BiP and multiple DNAJ molecular chaperones in the endoplasmic reticulum are required for efficient simian virus 40 infection. mBio 2(3):e00101-11. doi: 10.1128/mBio.00101-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Uemura A, Oku M, Mori K, Yoshida H. 2009. Unconventional splicing of XBP1 mRNA occurs in the cytoplasm during the mammalian unfolded protein response. J Cell Sci 122:2877–2886. doi: 10.1242/jcs.040584. [DOI] [PubMed] [Google Scholar]
- 35.Ushioda R, Hoseki J, Araki K, Jansen G, Thomas DY, Nagata K. 2008. ERdj5 is required as a disulfide reductase for degradation of misfolded proteins in the ER. Science 321:569–572. doi: 10.1126/science.1159293. [DOI] [PubMed] [Google Scholar]
- 36.Brodsky JL, Werner ED, Dubas ME, Goeckeler JL, Kruse KB, McCracken AA. 1999. The requirement for molecular chaperones during endoplasmic reticulum-associated protein degradation demonstrates that protein export and import are mechanistically distinct. J Biol Chem 274:3453–3460. doi: 10.1074/jbc.274.6.3453. [DOI] [PubMed] [Google Scholar]
- 37.Kabani M, Kelley SS, Morrow MW, Montgomery DL, Sivendran R, Rose MD, Gierasch LM, Brodsky JL. 2003. Dependence of endoplasmic reticulum-associated degradation on the peptide binding domain and concentration of BiP. Mol Biol Cell 14:3437–3448. doi: 10.1091/mbc.E02-12-0847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Nishikawa SI, Fewell SW, Kato Y, Brodsky JL, Endo T. 2001. Molecular chaperones in the yeast endoplasmic reticulum maintain the solubility of proteins for retrotranslocation and degradation. J Cell Biol 153:1061–1070. doi: 10.1083/jcb.153.5.1061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Hagiwara M, Maegawa K, Suzuki M, Ushioda R, Araki K, Matsumoto Y, Hoseki J, Nagata K, Inaba K. 2011. Structural basis of an ERAD pathway mediated by the ER-resident protein disulfide reductase ERdj5. Mol Cell 41:432–444. doi: 10.1016/j.molcel.2011.01.021. [DOI] [PubMed] [Google Scholar]
- 40.Li PP, Nguyen AP, Qu Q, Jafri QH, Aungsumart S, Cheng RH, Kasamatsu H. 2007. Importance of calcium-binding site 2 in simian virus 40 infection. J Virol 81:6099–6105. doi: 10.1128/JVI.02195-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Li PP, Naknanishi A, Tran MA, Ishizu K, Kawano M, Phillips M, Handa H, Liddington RC, Kasamatsu H. 2003. Importance of Vp1 calcium-binding residues in assembly, cell entry, and nuclear entry of simian virus 40. J Virol 77:7527–7538. doi: 10.1128/JVI.77.13.7527-7538.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Diacumakos EG, Gershey EL. 1977. Uncoating and gene expression of simian virus 40 in CV-1 cell nuclei inoculated by microinjection. J Virol 24:903–906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Brodsky JL. 2012. Cleaning up: ER-associated degradation to the rescue. Cell 151:1163–1167. doi: 10.1016/j.cell.2012.11.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Olzmann JA, Kopito RR, Christianson JC. 2013. The mammalian endoplasmic reticulum-associated degradation system. Cold Spring Harb Perspect Biol 5:a013185. doi: 10.1101/cshperspect.a013185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Smith MH, Ploegh HL, Weissman JS. 2011. Road to ruin: targeting proteins for degradation in the endoplasmic reticulum. Science 334:1086–1090. doi: 10.1126/science.1209235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Li PP, Nakanishi A, Fontanes V, Kasamatsu H. 2005. Pairs of Vp1 cysteine residues essential for simian virus 40 infection. J Virol 79:3859–3864. doi: 10.1128/JVI.79.6.3859-3864.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Grubb S, Guo L, Fisher EA, Brodsky JL. 2012. Protein disulfide isomerases contribute differentially to the endoplasmic reticulum-associated degradation of apolipoprotein B and other substrates. Mol Biol Cell 23:520–532. doi: 10.1091/mbc.E11-08-0704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Molinari M, Galli C, Piccaluga V, Pieren M, Paganetti P. 2002. Sequential assistance of molecular chaperones and transient formation of covalent complexes during protein degradation from the ER. J Cell Biol 158:247–257. doi: 10.1083/jcb.200204122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Tsai B, Rodighiero C, Lencer WI, Rapoport TA. 2001. Protein disulfide isomerase acts as a redox-dependent chaperone to unfold cholera toxin. Cell 104:937–948. doi: 10.1016/S0092-8674(01)00289-6. [DOI] [PubMed] [Google Scholar]
- 50.Wahlman J, DeMartino GN, Skach WR, Bulleid NJ, Brodsky JL, Johnson AE. 2007. Real-time fluorescence detection of ERAD substrate retrotranslocation in a mammalian in vitro system. Cell 129:943–955. doi: 10.1016/j.cell.2007.03.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Nelson CD, Stroh LJ, Gee GV, O'Hara BA, Stehle T, Atwood WJ. 2015. Modulation of a pore in the capsid of JC polyomavirus reduces infectivity and prevents exposure of the minor capsid proteins. J Virol 89:3910–3921. doi: 10.1128/JVI.00089-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Bagchi P, Walczak CP, Tsai B. 2015. The endoplasmic reticulum membrane J protein C18 executes a distinct role in promoting simian virus 40 membrane penetration. J Virol 89:4058–4068. doi: 10.1128/JVI.03574-14. [DOI] [PMC free article] [PubMed] [Google Scholar]