Abstract
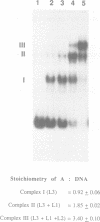
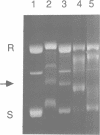
Initial events in Mu DNA transposition involve specific recognition of Mu DNA ends (att sites) and an internal enhancer site by the Mu transposase (A protein). This interaction between A protein and Mu DNA sequences present on a supercoiled DNA substrate leads to the formation of a stable synaptic complex in which the att ends are nicked, prior to DNA strand transfer. This study examines the properties of a synaptic complex proficient for DNA transposition. We show that the A protein binds as a monomer to its binding sites, and causes the DNA to bend through approximately 90 degrees at each site. All six att binding sites (three at each Mu end) are occupied by A within the synaptic complex. Three of these sites are loosely held and can be emptied of A upon challenge with heparin. A synaptic complex with only three sites occupied is stable and is fully competent in the subsequent strand-transfer step of transposition.
Full text
PDF
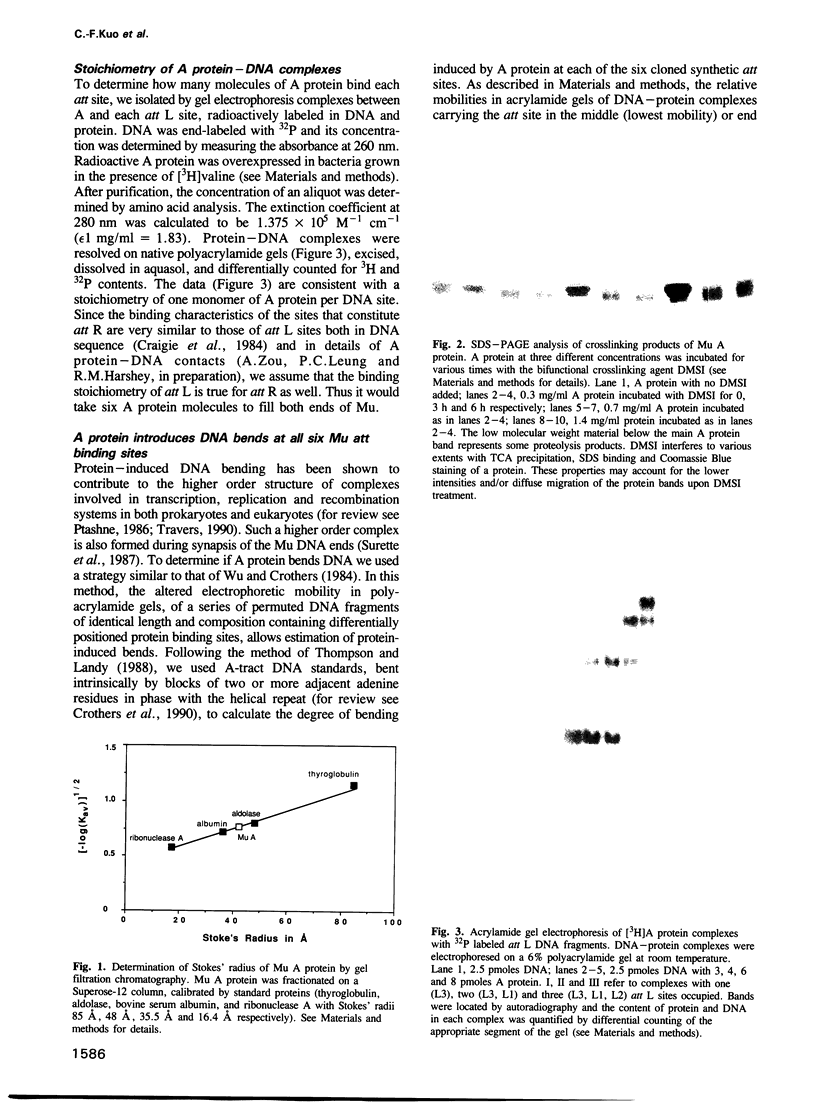
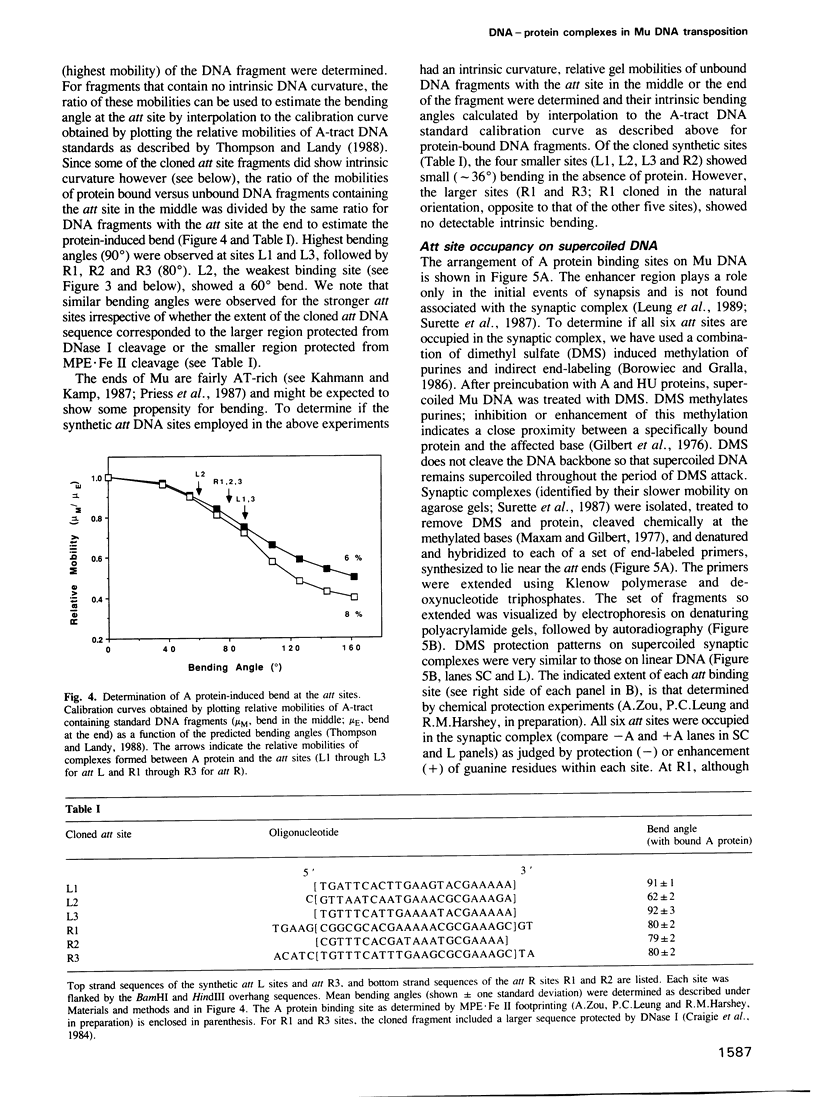




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Borowiec J. A., Gralla J. D. High-resolution analysis of lac transcription complexes inside cells. Biochemistry. 1986 Sep 9;25(18):5051–5057. doi: 10.1021/bi00366a012. [DOI] [PubMed] [Google Scholar]
- Chaconas G., de Bruijn F. J., Casadaban M. J., Lupski J. R., Kwoh T. J., Harshey R. M., DuBow M. S., Bukhari A. I. In vitro and in vivo manipulations of bacteriophage Mu DNA: cloning of Mu ends and construction of mini-Mu's carrying selectable markers. Gene. 1981 Jan-Feb;13(1):37–46. doi: 10.1016/0378-1119(81)90041-x. [DOI] [PubMed] [Google Scholar]
- Craigie R., Arndt-Jovin D. J., Mizuuchi K. A defined system for the DNA strand-transfer reaction at the initiation of bacteriophage Mu transposition: protein and DNA substrate requirements. Proc Natl Acad Sci U S A. 1985 Nov;82(22):7570–7574. doi: 10.1073/pnas.82.22.7570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Craigie R., Mizuuchi K. Cloning of the A gene of bacteriophage Mu and purification of its product, the Mu transposase. J Biol Chem. 1985 Feb 10;260(3):1832–1835. [PubMed] [Google Scholar]
- Craigie R., Mizuuchi K. Mechanism of transposition of bacteriophage Mu: structure of a transposition intermediate. Cell. 1985 Jul;41(3):867–876. doi: 10.1016/s0092-8674(85)80067-2. [DOI] [PubMed] [Google Scholar]
- Craigie R., Mizuuchi K. Role of DNA topology in Mu transposition: mechanism of sensing the relative orientation of two DNA segments. Cell. 1986 Jun 20;45(6):793–800. doi: 10.1016/0092-8674(86)90554-4. [DOI] [PubMed] [Google Scholar]
- Craigie R., Mizuuchi K. Transposition of Mu DNA: joining of Mu to target DNA can be uncoupled from cleavage at the ends of Mu. Cell. 1987 Nov 6;51(3):493–501. doi: 10.1016/0092-8674(87)90645-3. [DOI] [PubMed] [Google Scholar]
- Craigie R., Mizuuchi M., Mizuuchi K. Site-specific recognition of the bacteriophage Mu ends by the Mu A protein. Cell. 1984 Dec;39(2 Pt 1):387–394. doi: 10.1016/0092-8674(84)90017-5. [DOI] [PubMed] [Google Scholar]
- Crothers D. M., Haran T. E., Nadeau J. G. Intrinsically bent DNA. J Biol Chem. 1990 May 5;265(13):7093–7096. [PubMed] [Google Scholar]
- Davies G. E., Stark G. R. Use of dimethyl suberimidate, a cross-linking reagent, in studying the subunit structure of oligomeric proteins. Proc Natl Acad Sci U S A. 1970 Jul;66(3):651–656. doi: 10.1073/pnas.66.3.651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Groenen M. A., Timmers E., van de Putte P. DNA sequences at the ends of the genome of bacteriophage Mu essential for transposition. Proc Natl Acad Sci U S A. 1985 Apr;82(7):2087–2091. doi: 10.1073/pnas.82.7.2087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harshey R. M., Getzoff E. D., Baldwin D. L., Miller J. L., Chaconas G. Primary structure of phage mu transposase: homology to mu repressor. Proc Natl Acad Sci U S A. 1985 Nov;82(22):7676–7680. doi: 10.1073/pnas.82.22.7676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ITZHAKI R. F., GILL D. M. A MICRO-BIURET METHOD FOR ESTIMATING PROTEINS. Anal Biochem. 1964 Dec;9:401–410. doi: 10.1016/0003-2697(64)90200-3. [DOI] [PubMed] [Google Scholar]
- Kostrewa D., Granzin J., Koch C., Choe H. W., Raghunathan S., Wolf W., Labahn J., Kahmann R., Saenger W. Three-dimensional structure of the E. coli DNA-binding protein FIS. Nature. 1991 Jan 10;349(6305):178–180. doi: 10.1038/349178a0. [DOI] [PubMed] [Google Scholar]
- Leung P. C., Teplow D. B., Harshey R. M. Interaction of distinct domains in Mu transposase with Mu DNA ends and an internal transpositional enhancer. Nature. 1989 Apr 20;338(6217):656–658. doi: 10.1038/338656a0. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. A new method for sequencing DNA. Proc Natl Acad Sci U S A. 1977 Feb;74(2):560–564. doi: 10.1073/pnas.74.2.560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxwell A., Craigie R., Mizuuchi K. B protein of bacteriophage mu is an ATPase that preferentially stimulates intermolecular DNA strand transfer. Proc Natl Acad Sci U S A. 1987 Feb;84(3):699–703. doi: 10.1073/pnas.84.3.699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakayama C., Teplow D. B., Harshey R. M. Structural domains in phage Mu transposase: identification of the site-specific DNA-binding domain. Proc Natl Acad Sci U S A. 1987 Apr;84(7):1809–1813. doi: 10.1073/pnas.84.7.1809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ptashne M. Gene regulation by proteins acting nearby and at a distance. Nature. 1986 Aug 21;322(6081):697–701. doi: 10.1038/322697a0. [DOI] [PubMed] [Google Scholar]
- Rao R. N. Construction and properties of plasmid pKC30, a pBR322 derivative containing the pL-N region of phage lambda. Gene. 1984 Nov;31(1-3):247–250. doi: 10.1016/0378-1119(84)90216-6. [DOI] [PubMed] [Google Scholar]
- Reyes O., Gottesman M., Adhya S. Formation of lambda lysogens by IS2 recombination: gal operon--lambda pR promoter fusions. Virology. 1979 Apr 30;94(2):400–408. doi: 10.1016/0042-6822(79)90470-7. [DOI] [PubMed] [Google Scholar]
- Shuey D. J., Parker C. S. Bending of promoter DNA on binding of heat shock transcription factor. Nature. 1986 Oct 2;323(6087):459–461. doi: 10.1038/323459a0. [DOI] [PubMed] [Google Scholar]
- Surette M. G., Buch S. J., Chaconas G. Transpososomes: stable protein-DNA complexes involved in the in vitro transposition of bacteriophage Mu DNA. Cell. 1987 Apr 24;49(2):253–262. doi: 10.1016/0092-8674(87)90566-6. [DOI] [PubMed] [Google Scholar]
- Surette M. G., Chaconas G. A protein factor which reduces the negative supercoiling requirement in the Mu DNA strand transfer reaction is Escherichia coli integration host factor. J Biol Chem. 1989 Feb 15;264(5):3028–3034. [PubMed] [Google Scholar]
- Surette M. G., Lavoie B. D., Chaconas G. Action at a distance in Mu DNA transposition: an enhancer-like element is the site of action of supercoiling relief activity by integration host factor (IHF). EMBO J. 1989 Nov;8(11):3483–3489. doi: 10.1002/j.1460-2075.1989.tb08513.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teplow D. B., Nakayama C., Leung P. C., Harshey R. M. Structure-function relationships in the transposition protein B of bacteriophage Mu. J Biol Chem. 1988 Aug 5;263(22):10851–10857. [PubMed] [Google Scholar]
- Thompson J. F., Landy A. Empirical estimation of protein-induced DNA bending angles: applications to lambda site-specific recombination complexes. Nucleic Acids Res. 1988 Oct 25;16(20):9687–9705. doi: 10.1093/nar/16.20.9687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Travers A. A. Why bend DNA? Cell. 1990 Jan 26;60(2):177–180. doi: 10.1016/0092-8674(90)90729-x. [DOI] [PubMed] [Google Scholar]
- Wu H. M., Crothers D. M. The locus of sequence-directed and protein-induced DNA bending. Nature. 1984 Apr 5;308(5959):509–513. doi: 10.1038/308509a0. [DOI] [PubMed] [Google Scholar]