Abstract
The expression of the yeast L2 r-protein gene is controlled at the level of mRNA accumulation. The product of the gene appears to participate in this regulation by an autogenous feedback mechanism. This control does not operate at the level of transcription but instead affects L2 mRNA accumulation. This autogenous regulation of mRNA accumulation provides an interesting analogy to the autogenous translational regulation of r-proteins in Escherichia coli.
Full text
PDF

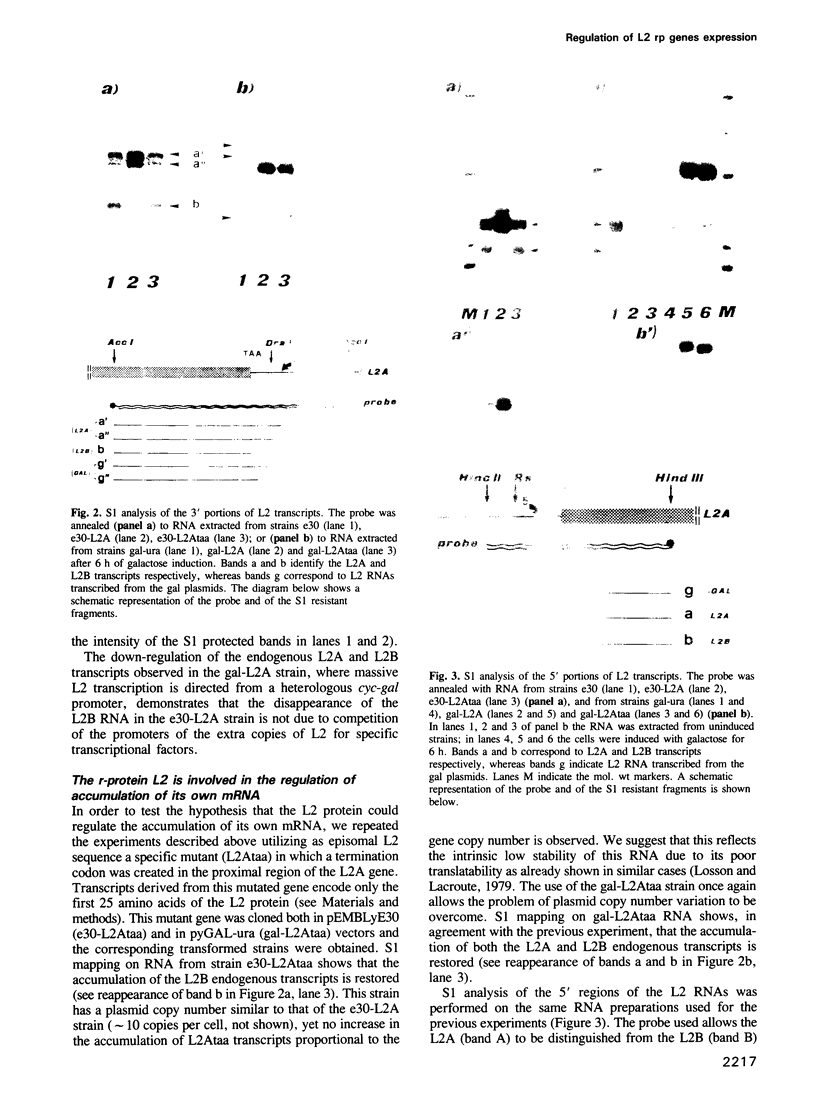




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abovich N., Gritz L., Tung L., Rosbash M. Effect of RP51 gene dosage alterations on ribosome synthesis in Saccharomyces cerevisiae. Mol Cell Biol. 1985 Dec;5(12):3429–3435. doi: 10.1128/mcb.5.12.3429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Al-Atia G. R., Fruscoloni P., Jacobs-Lorena M. Translational regulation of mRNAs for ribosomal proteins during early Drosophila development. Biochemistry. 1985 Oct 8;24(21):5798–5803. doi: 10.1021/bi00342a017. [DOI] [PubMed] [Google Scholar]
- Amaldi F., Bozzoni I., Beccari E., Pierandrei-Amaldi P. Expression of ribosomal protein genes and regulation of ribosome biosynthesis in Xenopus development. Trends Biochem Sci. 1989 May;14(5):175–178. doi: 10.1016/0968-0004(89)90269-7. [DOI] [PubMed] [Google Scholar]
- Baldari C., Cesareni G. Plasmids pEMBLY: new single-stranded shuttle vectors for the recovery and analysis of yeast DNA sequences. Gene. 1985;35(1-2):27–32. doi: 10.1016/0378-1119(85)90154-4. [DOI] [PubMed] [Google Scholar]
- Bowman L. H. The synthesis of ribosomal proteins S16 and L32 is not autogenously regulated during mouse myoblast differentiation. Mol Cell Biol. 1987 Dec;7(12):4464–4471. doi: 10.1128/mcb.7.12.4464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bozzoni I., Fragapane P., Annesi F., Pierandrei-Amaldi P., Amaldi F., Beccari E. Expression of two Xenopus laevis ribosomal protein genes in injected frog oocytes. A specific splicing block interferes with the L1 RNA maturation. J Mol Biol. 1984 Dec 25;180(4):987–1005. doi: 10.1016/0022-2836(84)90267-5. [DOI] [PubMed] [Google Scholar]
- Caffarelli E., Fragapane P., Gehring C., Bozzoni I. The accumulation of mature RNA for the Xenopus laevis ribosomal protein L1 is controlled at the level of splicing and turnover of the precursor RNA. EMBO J. 1987 Nov;6(11):3493–3498. doi: 10.1002/j.1460-2075.1987.tb02674.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Capieaux E., Vignais M. L., Sentenac A., Goffeau A. The yeast H+-ATPase gene is controlled by the promoter binding factor TUF. J Biol Chem. 1989 May 5;264(13):7437–7446. [PubMed] [Google Scholar]
- Cleveland D. W. Autoregulated instability of tubulin mRNAs: a novel eukaryotic regulatory mechanism. Trends Biochem Sci. 1988 Sep;13(9):339–343. doi: 10.1016/0968-0004(88)90103-x. [DOI] [PubMed] [Google Scholar]
- Dabeva M. D., Post-Beittenmiller M. A., Warner J. R. Autogenous regulation of splicing of the transcript of a yeast ribosomal protein gene. Proc Natl Acad Sci U S A. 1986 Aug;83(16):5854–5857. doi: 10.1073/pnas.83.16.5854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Della Seta F., Ciafré S. A., Marck C., Santoro B., Presutti C., Sentenac A., Bozzoni I. The ABF1 factor is the transcriptional activator of the L2 ribosomal protein genes in Saccharomyces cerevisiae. Mol Cell Biol. 1990 May;10(5):2437–2441. doi: 10.1128/mcb.10.5.2437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dorsman J. C., Doorenbosch M. M., Maurer C. T., de Winde J. H., Mager W. H., Planta R. J., Grivell L. A. An ARS/silencer binding factor also activates two ribosomal protein genes in yeast. Nucleic Acids Res. 1989 Jul 11;17(13):4917–4923. doi: 10.1093/nar/17.13.4917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dorsman J. C., van Heeswijk W. C., Grivell L. A. Identification of two factors which bind to the upstream sequences of a number of nuclear genes coding for mitochondrial proteins and to genetic elements important for cell division in yeast. Nucleic Acids Res. 1988 Aug 11;16(15):7287–7301. doi: 10.1093/nar/16.15.7287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- El-Baradi T. T., Raué H. A., De Regt C. H., Planta R. J. Stepwise dissociation of yeast 60S ribosomal subunits by LiCl and identification of L25 as a primary 26S rRNA binding protein. Eur J Biochem. 1984 Oct 15;144(2):393–400. doi: 10.1111/j.1432-1033.1984.tb08477.x. [DOI] [PubMed] [Google Scholar]
- Huet J., Cottrelle P., Cool M., Vignais M. L., Thiele D., Marck C., Buhler J. M., Sentenac A., Fromageot P. A general upstream binding factor for genes of the yeast translational apparatus. EMBO J. 1985 Dec 16;4(13A):3539–3547. doi: 10.1002/j.1460-2075.1985.tb04114.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacobs F. A., Bird R. C., Sells B. H. Differentiation of rat myoblasts. Regulation of turnover of ribosomal proteins and their mRNAs. Eur J Biochem. 1985 Jul 15;150(2):255–263. doi: 10.1111/j.1432-1033.1985.tb09015.x. [DOI] [PubMed] [Google Scholar]
- Leer R. J., Van Raamsdonk-Duin M. M., Mager W. H., Planta R. J. Conserved sequences upstream of yeast ribosomal protein genes. Curr Genet. 1985;9(4):273–277. doi: 10.1007/BF00419955. [DOI] [PubMed] [Google Scholar]
- Losson R., Lacroute F. Interference of nonsense mutations with eukaryotic messenger RNA stability. Proc Natl Acad Sci U S A. 1979 Oct;76(10):5134–5137. doi: 10.1073/pnas.76.10.5134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lucioli A., Presutti C., Ciafrè S., Caffarelli E., Fragapane P., Bozzoni I. Gene dosage alteration of L2 ribosomal protein genes in Saccharomyces cerevisiae: effects on ribosome synthesis. Mol Cell Biol. 1988 Nov;8(11):4792–4798. doi: 10.1128/mcb.8.11.4792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mager W. H. Control of ribosomal protein gene expression. Biochim Biophys Acta. 1988 Jan 25;949(1):1–15. doi: 10.1016/0167-4781(88)90048-6. [DOI] [PubMed] [Google Scholar]
- Maicas E., Pluthero F. G., Friesen J. D. The accumulation of three yeast ribosomal proteins under conditions of excess mRNA is determined primarily by fast protein decay. Mol Cell Biol. 1988 Jan;8(1):169–175. doi: 10.1128/mcb.8.1.169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mariottini P., Amaldi F. The 5' untranslated region of mRNA for ribosomal protein S19 is involved in its translational regulation during Xenopus development. Mol Cell Biol. 1990 Feb;10(2):816–822. doi: 10.1128/mcb.10.2.816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mattheakis L., Vu L., Sor F., Nomura M. Retroregulation of the synthesis of ribosomal proteins L14 and L24 by feedback repressor S8 in Escherichia coli. Proc Natl Acad Sci U S A. 1989 Jan;86(2):448–452. doi: 10.1073/pnas.86.2.448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Meyuhas O., Thompson E. A., Jr, Perry R. P. Glucocorticoids selectively inhibit translation of ribosomal protein mRNAs in P1798 lymphosarcoma cells. Mol Cell Biol. 1987 Aug;7(8):2691–2699. doi: 10.1128/mcb.7.8.2691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Myers A. M., Tzagoloff A., Kinney D. M., Lusty C. J. Yeast shuttle and integrative vectors with multiple cloning sites suitable for construction of lacZ fusions. Gene. 1986;45(3):299–310. doi: 10.1016/0378-1119(86)90028-4. [DOI] [PubMed] [Google Scholar]
- Nomura M., Gourse R., Baughman G. Regulation of the synthesis of ribosomes and ribosomal components. Annu Rev Biochem. 1984;53:75–117. doi: 10.1146/annurev.bi.53.070184.000451. [DOI] [PubMed] [Google Scholar]
- Pachter J. S., Yen T. J., Cleveland D. W. Autoregulation of tubulin expression is achieved through specific degradation of polysomal tubulin mRNAs. Cell. 1987 Oct 23;51(2):283–292. doi: 10.1016/0092-8674(87)90155-3. [DOI] [PubMed] [Google Scholar]
- Pierandrei-Amaldi P., Beccari E., Bozzoni I., Amaldi F. Ribosomal protein production in normal and anucleolate Xenopus embryos: regulation at the posttranscriptional and translational levels. Cell. 1985 Aug;42(1):317–323. doi: 10.1016/s0092-8674(85)80127-6. [DOI] [PubMed] [Google Scholar]
- Planta R. J., Raué H. A. Control of ribosome biogenesis in yeast. Trends Genet. 1988 Mar;4(3):64–68. doi: 10.1016/0168-9525(88)90042-x. [DOI] [PubMed] [Google Scholar]
- Presutti C., Lucioli A., Bozzoni I. Ribosomal protein L2 in Saccharomyces cerevisiae is homologous to ribosomal protein L1 in Xenopus laevis. Isolation and characterization of the genes. J Biol Chem. 1988 May 5;263(13):6188–6192. [PubMed] [Google Scholar]
- Rotenberg M. O., Woolford J. L., Jr Tripartite upstream promoter element essential for expression of Saccharomyces cerevisiae ribosomal protein genes. Mol Cell Biol. 1986 Feb;6(2):674–687. doi: 10.1128/mcb.6.2.674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teem J. L., Abovich N., Kaufer N. F., Schwindinger W. F., Warner J. R., Levy A., Woolford J., Leer R. J., van Raamsdonk-Duin M. M., Mager W. H. A comparison of yeast ribosomal protein gene DNA sequences. Nucleic Acids Res. 1984 Nov 26;12(22):8295–8312. doi: 10.1093/nar/12.22.8295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teem J. L., Rosbash M. Expression of a beta-galactosidase gene containing the ribosomal protein 51 intron is sensitive to the rna2 mutation of yeast. Proc Natl Acad Sci U S A. 1983 Jul;80(14):4403–4407. doi: 10.1073/pnas.80.14.4403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsay Y. F., Thompson J. R., Rotenberg M. O., Larkin J. C., Woolford J. L., Jr Ribosomal protein synthesis is not regulated at the translational level in Saccharomyces cerevisiae: balanced accumulation of ribosomal proteins L16 and rp59 is mediated by turnover of excess protein. Genes Dev. 1988 Jun;2(6):664–676. doi: 10.1101/gad.2.6.664. [DOI] [PubMed] [Google Scholar]
- el-Baradi T. T., Raué H. A., Linnekamp M., Planta R. J. Identification of yeast 60 S ribosomal proteins crosslinked to rRNA by 2-iminothiolane. FEBS Lett. 1985 Jul 1;186(1):26–30. doi: 10.1016/0014-5793(85)81332-6. [DOI] [PubMed] [Google Scholar]
- elBaradi T. T., van der Sande C. A., Mager W. H., Raué H. A., Planta R. J. The cellular level of yeast ribosomal protein L25 is controlled principally by rapid degradation of excess protein. Curr Genet. 1986;10(10):733–739. doi: 10.1007/BF00405095. [DOI] [PubMed] [Google Scholar]