Abstract
Proteolytic fragments of amyloid and post-translational modification of tau species in Cerebrospinal fluid (CSF) as well as cerebral amyloid deposition are important biomarkers for Alzheimer’s Disease. We conducted genome-wide association study to identify genetic factors influencing CSF biomarker level, cerebral amyloid deposition, and disease progression. The genome-wide association study was performed via a meta-analysis of two non-overlapping discovery sample sets to identify genetic variants other than APOE ε4 predictive of the CSF biomarker level (Aβ1–42, t-Tau, p-Tau181P, t-Tau:Aβ1–42 ratio, and p-Tau181P:Aβ1–42 ratio) in patients enrolled in the Alzheimer’s Disease Neuroimaging Initiative (ADNI) study. Loci passing a genome-wide significance threshold of P < 5 x 10−8 were followed-up for replication in an independent sample set. We also performed joint meta-analysis of both discovery sample sets together with the replication sample set. In the discovery phase, we identified variants in FRA10AC1 associated with CSF Aβ1–42 level passing the genome-wide significance threshold (directly genotyped SNV rs10509663 P FE = 1.1 x 10−9, imputed SNV rs116953792 P FE = 3.5 x 10−10), rs116953792 (P one-sided = 0.04) achieved replication. This association became stronger in the joint meta-analysis (directly genotyped SNV rs10509663 P FE = 1.7 x 10−9, imputed SNV rs116953792 P FE = 7.6 x 10−11). Additionally, we identified locus 15q21 (imputed SNV rs1503351 P FE = 4.0 x 10−8) associated with CSF Aβ1–42 level. No other variants passed the genome-wide significance threshold for other CSF biomarkers in either the discovery sample sets or joint analysis. Gene set enrichment analyses suggested that targeted genes mediated by miR-33, miR-146, and miR-193 were enriched in various GWAS analyses. This finding is particularly important because CSF biomarkers confer disease susceptibility and may be predictive of the likelihood of disease progression in Alzheimer’s Disease.
Introduction
Alzheimer’s Disease (AD) is the most common form of dementia and to date there is still no cure. Understanding the factors influencing cognitive decline of AD will enable us to better search for therapeutics to intervene or preempt this process. It is well established now that the CSF biosignature of increased total tau (t-tau) and phosphor-tau (p-tau) species especially tau phosphorylated at the threonine 181 (p-tau181p) and decreased amyloid-β 1–42 peptide (Aβ1–42), is predictive of Alzheimer’s related amyloid pathology in the brain.[1–4] Using the Alzheimer’s Disease Neuroimaging Initiative (ADNI) samples from ADNI-1 sample set, the APOE ε4 allele best discriminated AD from controls (CN) in a logistic regression model including Aβ1–42, t-tau, while the AD-like t-tau/Aβ1–42 profile was detected among mild cognitive impairment (MCI) patients who converted to AD in one year follow up.[5] Samtani et al., used the ADNI-1 data to modeled disease progression using the Alzheimer’s Disease Assessment Scale-Cognitive Subscale (ADAS-Cog) in MCI patients using a mixture model and proposed log CSF p-tau181p:Aβ1–42 ratio > -1.86 to be the predictor discriminant progressers from non-progressers.[6] Other CSF biomarkers such as Aβ1–42, p-tau181p, and t-tau have also been linked to disease progression as measured by conversion from MCI to AD and/or cognitive decline. For example, the combination of CSF Aβ1–42 (or Aβ1-42/p-tau181p) and t-tau predicted the conversion from MCI to AD.[7] In a small study, non-demented patients with severely impaired episodic memory (SIM) but with no moderate impairment (MIM) or no impairment (NIM) at baseline declined cognitively over time and progressed to dementia at a high rate, and this was accompanied by significant increase in CSF p-tau181P but not t-tau or Aβ1–42 during approximately 3 year follow-up.[8]
Most genetic studies in AD thus far have been utilizing clinical diagnosis to define cases and controls. These studies are inherently limited by accuracy of clinical diagnosis. For example, we know that up to 36.1% of APOE ε4 allele non-carriers clinically diagnosed with Alzheimer’s Dementia do not have Alzheimer’s pathology as measured by PIB-PET.[9] GWAS studies that utilize PET amyloid signal or a CSF biosignture as quantitative endophenotype offer an opportunity, in principle, to define more objective phenotype, and establish direct associations between genetic variations, disease state biomarkers and disease progression. Kim et al. performed a GWAS to identify genetic risk factors for the three singular CSF biomarkers (Aβ1–42, p-tau181p, and t-tau) and two ratios (p-tau181p:Aβ1–42 and t-tau:Aβ1–42), utilized the ADNI1 samples genotyped with the Illumina Human 610-Quad BeadChip [10], they implicated one novel gene, EPC2 that reached genome-wide significance for association with t-tau while confirming the expected association of CSF biomarkers with the APOE/TOMM40 region. Cruchaga et al. identified a SNP located in intron 5 of the regulatory subunit of the PPP3R1 gene associated with CSF p-tau181 levels in two independent CSF sample sets.[11] The largest CSF biomarker GWAS (N = 1,269) was based on a one stage analysis of four datasets leading to the identification of three novel genome-wide significant loci, an intronic imputed SNP in GLIS3 (associated with p-tau181p and t-tau), an intergenic imputed SNP between GMNC and OSTN (associate with t-tau), and an intergenic genotyped SNP (associated with p-tau181p) near NCR2 and within the TREM gene cluster.[12] Thus far, the ADNI-GO/-2 CSF samples have not been included in any CSF GWAS analysis. In this study, we expanded the reported studies by including additional samples genotyped by the ADNI study and thus aimed to identify genetic variants predictive of CSF biomarkers independent of the APOE ε4 effect.
Similar to CSF Aβ1–42, cerebral amyloid deposition as measured by PET imaging has been used as a quantitative trait (QT) in GWAS, in addition to the APOE loci, rs509208 near butyrylcholinesterase (BCHE) was identified as a hit passing the genome-wide significant threshold for association with the florbetapir PET QT.[13] Hu et al., assessed the rate of disease progression in MCI subjects, as measured by changes in the Clinical Dementia Rating-sum of boxes (CDR-SB), by performing a GWAS in two independent sample sets and identified several novel loci that achieved genome-wide significance (intronic SNPs in UBR5 and PARP6, and an intergenic SNP near ACOT11).[14] In this report, we also describe GWAS or GWAS meta-analysis using florbetapir PET as a quantitative trait and using a dichotomized measure of amyloid positivity, and disease progression/rate of cognitive decline in late mild cognitive impairment (LMCI) population.
Results
We performed a genome-wide association study via a meta-analysis of two discovery sample sets (discovery sample set 1: genotyped using the Illumina Human610-Quad; discovery sample set 2: genotyped using Illumina Omni2.5, Table 1 and Table A in S1 File) to identify genetic variants other than APOE ε4 predictive of the CSF biomarker level (Aβ1–42, t-Tau, p-Tau181P, t-Tau:Aβ1–42 ratio, and p-Tau181P:Aβ1–42 ratio) in patients enrolled in the ADNI study, and followed-up by replication of any locus passing the genome-wide significance threshold of P < 5 x 10−8. In the discovery phase, we identified variants from one single locus associated with CSF Aβ1–42 level passing the genome-wide significance threshold. The most significantly associated markers predictive of CSF Aβ1–42 level in our meta-analysis of discovery sample sets are, directly genotyped intronic SNV rs10509663 P FE = 1.1 x 10−9 and imputed putative promoter region SNV rs116953792 P FE = 3.5 x 10−10, they are located in a ~60kb interval in chromosome 10 that contains the rare FRA10A folate-sensitive fragile site FRA10AC1 (Figure B1 in S1 File). This genetic association was replicated (rs116953792 P one-sided = 0.04) in the replication sample set (Table 1 and Table A in S1 File, n = 172 genotyped using Illumina OmniExpress). We also performed joint meta-analysis of both discovery sample sets together with the replication sample set. The FRA10AC1 association became stronger in the joint analysis (directly genotyped SNV rs10509663 P FE = 1.7 x 10−9, imputed SNV rs116953792 P FE = 7.6 x 10−11, Figs 1 and 2A). We identified an additional genome wide-significant locus within the15q21 locus (directly genotyped SNV rs4301994 P FE = 6.5 x 10−8; imputed SNV rs1503351 P FE = 4.0 x 10−8, Figs 1 and 2B) also associated with CSF Aβ1–42 level. This locus is in the intergenic region of chromosome 15 (15q21) between spermatogenesis associated 8 (SPATA8) and hypothetical LOC91948. There are spliced yet uncharacterized EST in this “intergenic” region. No other variants passed the genome-wide significance threshold level for any of the other CSF biomarkers (t-Tau, p-Tau181P, t-Tau:Aβ1–42 ratio, and p-Tau181P:Aβ1–42 ratio, Figures B2-5 in S1 File). As expected, APOE ε4, age, and clinical diagnosis were predictive of CSF Aβ1–42 level in all three sample sets. The association between FRA10AC1 locus and CSF Aβ1–42 is primarily driven by the first two sample sets with uncorrected significance levels of p = 0.0006 for the Upennbiomk_Human610-Quad sample set and p = 1.86 x 10−6 for the Upennbiomk5_Omni2.5 sample set, and P two-sided = 0.2 (P one-sided = 0.1) for the Upennbiomk6_OmniExpress sample set for the directly genotyped marker rs10509663. The imputed SNV rs116953792 however achieved P one-sided = 0.04 for the Upennbiomk6_OmniExpress sample set. The association between rs4301994 and CSF Aβ1–42 is supported by all three sample sets with uncorrected significance levels of P = 0.003 for the Upennbiomk_Human610-Quad sample set, P = 6.6 x 10−5 for the Upennbiomk5_Omni2.5 sample set, and P = 0.03 in the Upennbiomk6_OmniExpress sample set. The regression beta coefficients and least square means of the minor allele dosage for rs10509663 (FRA10AC1) and rs4301994 (15q21 locus) are shown in the forest plot (Fig 3A and 3B) and Table 2, displaying a consistent trend across the three sample sets. Both directly genotyped markers rs10509663 and rs4301994 did not deviate from Hardy-Weinberg Equilibrium (P = 0.38 and 1, respectively based on Omni2.5 data among cognitively normal controls). Table 3 contains those mostly independent variants (both the most significant directly genotyped and imputed markers are retained) that are significantly associated with Aβ1–42 level from our meta-analysis, along with other variants with uncorrected P ≤ 1x10-6 in any CSF biomarker meta-analysis. The full list of variants meeting this more liberal threshold appears in the S1 Table.
Table 1. Basic demographic information of the three ADNI CSF Sample Sets.
| Upennbiomk_Human610-Quad (N = 391) | Upennbiomk5_Omni2.5 (N = 385) | Upennbiomk6_OmniExpress (N = 204) | |
|---|---|---|---|
| Sex, n (%) | |||
| F | 155 (39.6) | 176 (45.7) | 87 (42.6) |
| M | 236 (60.4) | 209 (54.3) | 117 (57.4) |
| Age at baseline, years | |||
| Mean (SD) | 74..8 (7.1) | 72.5 (7.5) | 72.1 (7.6) |
| Median (Range) | 75 (54.4, 89.6) | 72.6 (55.0, 91.4) | 72.3 (48.1, 89.3) |
| Baseline Clinical Diagnosis, n (%) | |||
| CN | 109 (27.9) | 106 (27.5) | 21 (10.3) |
| SMC | N/A | N/A | 3 (1.5) |
| EMCI | N/A | 189 (49.1) | 57 (27.9) |
| LMCI | 186 (47.6) | 65 (16.9) | 61 (29.9) |
| AD | 96 (24.6) | 25 (6.5) | 62 (30.4) |
| APOE ε4, copy, n (%) | |||
| 0 | 198 (50.6) | 234 (60.9) | 76 (39.8) |
| 1 | 149 (38.1) | 122 (31.8) | 82 (42.9) |
| 2 | 44 (11.3) | 28 (7.3) | 33 (17.3) |
| Missing Data | 0 | 1 | 13 |
| CDR-SB 0 n (%) | 103 (26.3) | 100 (26.0) | 24 (11.8) |
| Aβ1–42 (pg/ml) | |||
| Mean (SD) | 170.2 (56.6) | 176.6 (50.3) | 160.4 (51.7) |
| Median (Range) | 152.5 (53, 300) | 175.5 (82.5, 313.6) | 146.3 (40.5, 301.9) |
| p-Tau181P (pg/ml) | |||
| Mean (SD) | 33.8 (18.5) | 39.6 (23.2) | 44.2 (25.9) |
| Median (Range) | 30 (8, 115) | 33.7 (9.4. 173.3) | 38.9 (6.9, 151.2) |
| t-tau (pg/ml) | |||
| Mean (SD) | 98.4 (55.8) | 80.3 (47.4) | 106.8 (64.1) |
| Median (Range) | 84 (28, 495) | 66 (19.9, 300.5) | 89.1 (23.2, 360.0) |
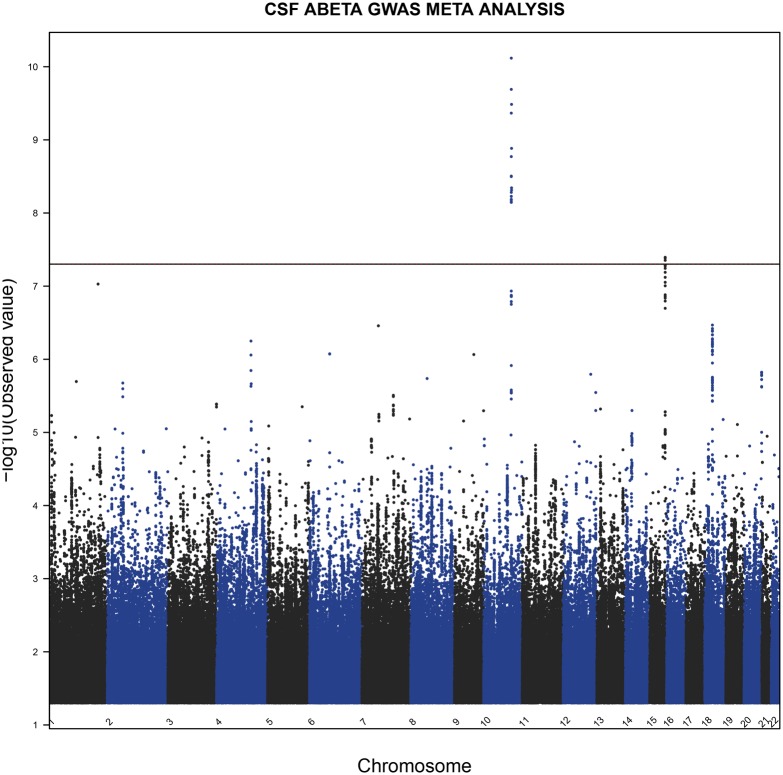
Fig 1. Manhattan plot of the CSF Biomarker Aβ1–42 GWAS Meta-Analysis.
The dotted line indicates genome wide significance threshold of 5x10-8. Only variants with p < 0.05 are shown.
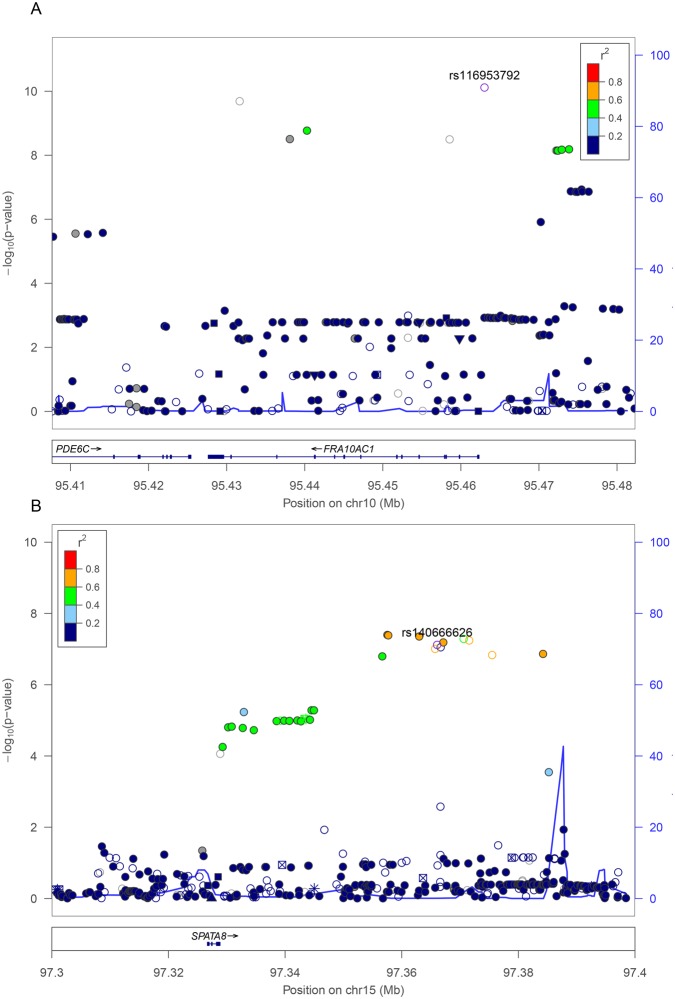
Fig 2. Regional Plot for the CSF Aβ1–42 Loci.
(A) FRA10AC1 (B) 15q21 Association results (-log10 p) are plotted for all single nucleotide polymorphisms (SNPs) passing quality control. Chromosome position is plotted with reference to the NCBI build 37. Recombination rate as estimated from the HapMap Project is plotted in light blue. SNPs are color coded according to the LD measure (r2) with reference SNP based on the reference panel of CEU population from the 1000 Genome Project (March 2012 release). SNP annotation for all 1000GP SNPs are represented by the annotation categories: framestop (triangle), splice (triangle), non-synonymous (inverted triangle), synonymous (square), UTR (square), TFBScons (star), MCS44 Placental (square with diagonal lines) and none-of-the-above (filled circle).
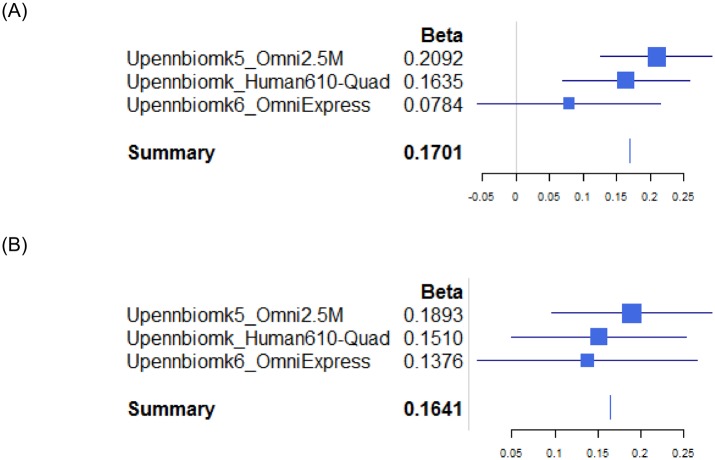
Fig 3. Forest Plot for the CSF Aβ1–42 Loci.
(A) rs10509663 in FRA10AC1 (B) rs4301994 in 15q21 showing the effect of genotype on Aβ1–42 by Individual sample set.
Table 2. Summary Characteristics of top CSF Aβ1–42 variants by Sample Set averaging over the levels of APOE ε4 and baseline clinical diagnosis assuming the baseline age was 60.
| Sample Set | Upennbiomk_Human610-Quad | Upennbiomk5_Omni2.5 | Upennbiomk6_OmniExpress | |||
|---|---|---|---|---|---|---|
| Copy | N | Least square means ± standard error (SE) | N | Least square means ± standard error (SE) | N | Least square means ± standard error (SE) |
| rs10509663-G | ||||||
| 0 | 313 | 161.8 ± 5.8 | 313 | 171.2 ± 4.9 | 154 | 159.6 ± 6.3 |
| 1 | 24 | 146.7 ± 10.6 | 30 | 141.6 ± 8.2 | 16 | 149.8 ± 11.7 |
| 2 | 3 | 96.3 ± 27.1 | 1 | 62.9 ± 39.5 | - | - |
| rs4301994-C | ||||||
| 0 | 311 | 162.5 ± 5.9 | 316 | 169.9 ± 4.9 | 154 | 159.9 ± 6.0 |
| 1 | 28 | 141.1 ± 10.1 | 28 | 139.6 ± 8.6 | 18 | 139.2 ± 11.2 |
| 2 | 1 | 151.3 ± 46.5 | - | - | - | - |
Table 3. Summary of CSF biomarker GWAS meta-analyses—SNPs with uncorrected p-value less than 1x10-6.
Top variants were clumped using parameters—clump-p1 0.000001—clump-p2 0.05—clump-r2 0.2—clump-range entrez.gene.map—clump-range-border 20.
| SNP a | CHR | BP b | A1 | A2 | Func | Gene | MAF c | Imputed2.5M/610K/OmniExpress | PDiscovery d | Preplication, 1-sided d | Pjoint d | βjoint e |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Aβ1–42 | ||||||||||||
| rs116953792 | 10 | 95463026 | G | T | upstream | FRA10AC1 | 0.01 | Y/Y/Y | 3.47E-10 | 0.039115 | 7.64E-11 | 0.2395 |
| rs10509663 | 10 | 95440286 | A | G | intronic | FRA10AC1 | 0.14 | N/N/N | 1.08E-09 | 0.12455 | 1.69E-09 | 0.1701 |
| rs1503351 | 15 | 97357520 | A | G | intergenic | SPATA8(dist = 28675),LOC91948(dist = 928326) | 0.05 | Y/Y/Y | 4.03E-08 | 0.1699 | ||
| rs4301994 | 15 | 97367115 | T | C | intergenic | SPATA8(dist = 38270),LOC91948(dist = 918731) | 0.06 | N/N/N | 6.47E-08 | 0.1641 | ||
| rs188308056 | 15 | 97366666 | T | A | intergenic | SPATA8(dist = 37821),LOC91948(dist = 919180) | 0.03 | Y/Y/Y | 8.88E-08 | 0.3122 | ||
| rs75849835 | 1 | 221475899 | A | G | intergenic | HLX(dist = 417499),C1orf140(dist = 27371) | 0.02 | Y/Y/Y | 9.38E-08 | 0.2042 | ||
| rs7098209 | 10 | 95475470 | T | C | intergenic | FRA10AC1(dist = 13141),LGI1(dist = 42096) | 0.25 | N/N/N | 1.17E-07 | 0.0885 | ||
| rs509477 | 18 | 32559295 | C | G | intronic | MAPRE2 | 0.6 | Y/Y/Y | 3.41E-07 | -0.0653 | ||
| rs55704525 | 7 | 43568566 | G | A | intronic | HECW1 | 0.01 | Y/Y/Y | 3.48E-07 | 0.3463 | ||
| rs140913323 | 4 | 125740709 | T | G | intergenic | ANKRD50(dist = 106822),FAT4(dist = 496858) | 0.02 | Y/Y/Y | 5.64E-07 | 0.4822 | ||
| rs2493168 | 6 | 53118235 | A | G | intergenic | GCM1(dist = 104611),ELOVL5(dist = 13961) | 0.71 | Y/Y/Y | 8.39E-07 | -0.0653 | ||
| rs8190569 | 9 | 98998061 | T | G | intronic | HSD17B3 | 0.01 | Y/Y/Y | 8.61E-07 | 0.24 | ||
| p-Tau 181P | ||||||||||||
| rs6005807 | 22 | 28934313 | T | C | intronic | TTC28 | 0.9 | Y/Y/Y | 2.74E-07 | -0.213 | ||
| rs75213930 | 1 | 83125608 | G | A | intergenic | LPHN2(dist = 667501),MIR548AP(dist = 1133990) | 0.03 | Y/Y/Y | 3.59E-07 | 0.299 | ||
| rs76478271 | 19 | 41325199 | T | A | intergenic | RAB4B-EGLN2(dist = 10853),CYP2A6(dist = 24244) | 0.12 | Y/Y/Y | 4.08E-07 | -0.3309 | ||
| t-Tau | ||||||||||||
| rs76137255 | 19 | 40783832 | T | G | intronic | AKT2 | 0.02 | Y/Y/Y | 2.45E-07 | -0.3236 | ||
| rs79811809 | 7 | 140633481 | A | G | intergenic | BRAF(dist = 8917),MRPS33(dist = 72480) | 0.04 | Y/Y/Y | 3.31E-07 | 0.2394 | ||
| rs36056951 | 8 | 139965798 | C | T | intergenic | COL22A1(dist = 39562),KCNK9(dist = 659006) | 0.03 | Y/Y/Y | 3.88E-07 | -0.3291 | ||
| rs138451097 | 19 | 2873629 | A | G | intronic | ZNF556 | 0.01 | Y/Y/Y | 5.39E-07 | -1.0045 | ||
| rs8045334 | 16 | 63573376 | G | A | intergenic | CDH8(dist = 1502637),CDH11(dist = 1407307) | 0.19 | N/N/N | 8.15E-07 | -0.126 | ||
| p-Tau 181P:Aβ 1–42 ratio | ||||||||||||
| rs2301659 | 19 | 19035354 | G | T | intronic | DDX49 | 0.32 | Y/Y/Y | 2.66E-07 | -0.1679 | ||
| rs117025875 | 8 | 6999135 | G | A | intergenic | DEFA5(dist = 84876),LOC349196(dist = 119006) | 0.02 | Y/Y/Y | 9.33E-07 | -0.8594 | ||
| t-Tau:Aβ 1–42 ratio | ||||||||||||
| rs113027826 | 2 | 207549512 | T | C | intronic | DYTN | 0.14 | Y/Y/Y | 3.71E-07 | -0.4164 | ||
| rs114162361 | 1 | 174575642 | C | T | intronic | RABGAP1L | 0.01 | Y/Y/Y | 9.39E-07 | -0.6189 | ||
Chr, chromosome; A1, first allele code; A2, second allele code
a Indexed SNPs with uncorrected p < 1x10-6 in any of the CSF Biomarker GWAS Meta-Analyses
b Build 37, assembly hg19
c based on 2012 Apr release of 1000genome and all population
d Fixed-effects meta-analysis p-value
e Beta coefficient of for the SNP assuming additive genetic model
Although the CSF Aβ1–42 measurement using Upennbiomk5 (baseline Aβ1–42 for 117 ADNI-GO subjects and 272 ADNI-2 subjects) were not taken at the exact same visit as the florbetapir PET imaging, the overall correlation for the overlapping subjects between the two measurements was negatively correlated (r = -0.72). We expected the results from the cerebral amyloid deposition GWAS to provide complementary evidence with CSF Aβ1–42 meta-analysis even if the sample sets are not identical and the endpoints are not perfectly negatively correlated. In our florbetapir PET GWAS meta-analysis without correcting for APOE ε4 dosage, the APOE locus predicted florbetapir PET SUVR value (directly genotyped SNV rs429358 P = 7.99 x 10−32) as expected from the published results (Figure C1 in S1 File). There are uncommon variants (such as rs76117213, an intronic variant in WD repeat and FYVE domain containing 3 (WDFY3), P = 1.39 x 10−7 without correction for APOE ε4 dosage, P = 4.08 x 10−6 with correction for APOE ε4 dosage). The uncommon variant results, however, shall be interpreted with caution as these variants occurred at low minor allele frequency and the association statistics are based on small sample size for rare genotype groups. Table 4 contains the variants that were significant in florbetapir PET GWAS along with other variants with uncorrected P < 1x10-6 in any florbetapir PET GWAS analysis, most variants are independent except for the chromosome 19 region. The full list of variants meeting this more liberal threshold appears in the S2 Table. Top 15q21 locus variants associated with CSF Aβ1–42 exhibited a nominal association with florbetapir PET SUVR value (P = 0.002 for directly genotyped SNV rs4301994; P = 0.001 for imputed SNV rs1503351). Similarly, the imputed variant rs116953792 from FRA10AC1 also exhibited a nominal association with florbetapir PET SUVR value (P = 0.01). Results for the top CSF Aβ1–42 variant rs10509663 from FRA10AC1 in this and other analyses (S3 Table) are discussed in the S1 File.
Table 4. Summary of cerebral amyloid deposition florbetapir PET quantitative traits—SNPs with uncorrected p-value less than 1x10-6.
Top variants were clumped using parameters—clump-p1 0.000001—clump-p2 0.05—clump-r2 0.2—clump-range entrez.gene.map—clump-range-border 20.
| SNP a | CHR | BP b | Func | Gene | MAF c | Imputed2.5M/0mni Express | A1 | A2 | P d | β e | ExonicFunc |
|---|---|---|---|---|---|---|---|---|---|---|---|
| AV45 (not correcting APOE ε4) | |||||||||||
| rs76117213 | 4 | 85596236 | intronic | WDFY3 | 0.01 | Y/Y | G | A | 1.39E-07 | -0.26 | |
| rs362902 | 6 | 146681157 | intronic | GRM1 | 0.01 | Y/Y | T | C | 6.30E-07 | -0.27 | |
| rs36116061 | 8 | 89373327 | intergenic | MMP16(d ist = 33610), RIPK2(dist = 1396648) | 0.18 | N/Y | T | G | 6.93E-07 | 0.06 | |
| rs708886 | 12 | 119958702 | intronic | CCDC60 | 0.44 | Y/Y | T | G | 8.47E-07 | -0.05 | |
| rs28810 | 16 | 3500515 | intronic | NAA60 | 0.12 | Y/Y | A | G | 4.78E-07 | 0.06 | |
| rs589398 | 18 | 5311892 | intergenic | ZBTB14(dist = 14840),EPB41 L3(dist = 80496) | 0.99 | Y/Y | C | T | 3.48E-07 | 0.19 | |
| rs7407664 | 18 | 9031719 | intergenic | SOGA2(d ist = 198944), NDUFV2(dist = 70909) | 0.03 | Y/Y | A | G | 3.01 E-07 | -0.14 | |
| rs6859 | 19 | 45382034 | UTR3 | PVRL2 | 0.61 | N/N | A | G | 1.68E-07 | 0.05 | |
| rs157580 | 19 | 45395266 | intronic | T0MM40 | 0.63 | N/N | G | A | 1.63E-07 | -0.05 | |
| rs429358 | 19 | 45411941 | exonic | APOE | 0.15 | Y/Y | T | C | 7.99E-32 | -0.13 | nonsynonymous SNV |
| rs1081105 | 19 | 45412955 | dow nstream | APOE | 0.02 | Y/Y | A | C | 1.48E-08 | -0.13 | |
| rs12721052 | 19 | 45421972 | intronic | AP0C1 | 0.2568 | Y/Y | AG | A | 1.65E-07 | 0.06 | |
| rs60049679 | 19 | 45429708 | upstream | APOC1P1 | 0.11 | Y/Y | G | C | 1.30E-08 | -0.14 | |
| AV45 (correcting APOE ε4) | |||||||||||
| rs200527573 | 1 | 212742242 | intronic | ATF3 | 0.0042 | Y/Y | C | CTATT | 7.90E-07 | -0.28 | |
| rs57450513 | 5 | 141225446 | intergenic | ARAP3(dist = 163646), PCDH1 (dist = 7227) | 0.01 | Y/Y | C | A | 2.42E-07 | -0.22 | |
| rs708886 | 12 | 119958702 | intronic | CCDC60 | 0.44 | Y/Y | T | G | 9.33E-07 | -0.05 | |
| rs28810 | 16 | 3500515 | intronic | NAA60 | 0.12 | Y/Y | A | G | 5.90E-07 | 0.06 |
Chr, chromosome; A1, first allele code; A2, second allele code
a Representative SNPs with uncorrected p < 1x10-6 in any of the Cerebral amyloid deposition GWAS Analyses
b Build 37, assembly hg19
c based on 2012 Apr release of 1000genome and all population
d Fixed-effects meta-analysis p-value
e Beta coefficient of for the SNP assuming additive genetic model
For the rate of cognitive decline GWAS in the late-MCI subgroup, there were some variants that achieved the conventional genome-wide significance threshold (Figure D in S1 File), those variants occurred at low frequency (MAF < 0.05). An intronic variant rs2694777 in the GDNF family receptor alpha 1 gene (GFRA1) is among the common variants with suggestive association with rate of cognitive decline as measured by rate of change in CDR-SB (P = 1.2 x 10−5 European ancestry sample set and P = 6.77 x 10−6 in all races). The full list of variants with P ≤ 10−6 appears in the S4 Table.
The results from this study, for variants reported in the literature relevant/associated with CSF biomarkers, florbetapir PET, and disease progression were also reported in the S1 File.
Gene set enrichment analysis may yield signals of enriched gene sets in GWAS analysis despite the individual variants not reaching genome wide significance. Applying INRICH [15] enrichment analysis to the CSF biomarker GWAS results yields gene sets such as miR-33 target genes (P empirical = 0.0002, P corrected = 0.03) being enriched among p-tau181p:Aβ1–42 suggestive association hits (P < 0.0005) (Table 5 and S5 Table). miR-33 was identified to be a potent post-transcriptional regulator of lipid metabolism genes [16, 17] and cause disruption of cellular cholesterol homeostasis leading to pathologic processes including AD. Among, the potential targets of miR-33, PRKAA1 (Protein kinase, AMP-activated, alpha 1 catalytic subunit) mediates an autophagic process to clear extracellular Aβ fibrils by microglia, the immune cells in the brain.[18] Other potential targets of miR-33 included ARID5B (AT rich interactive domain 5B (MRF1-like)), KCNMA1 (Potassium large conductance calcium-activated channel, subfamily M, alpha member 1), and LGI1 (leucine-rich, glioma inactivated). ARID5B was previously implicated in AD. ARID5B variants (rs2588969 and rs494288) showed marginally significant association with LOAD in meta-analysis of 2,634 LOAD and 4,201 controls (P = 0.046 and 0.008, respectively), although the associations did not survive adjustment for covariates (P = 0.30 and 0.11, respectively).[19] Other inconclusive association of ARID5B variants included Naj et al. (OR = 0.93, P = 0.001) and Hollingworth et al. (OR = 1.06, P = 0.03) for rs2588969 with LOAD.[20, 21] rs16934131 in KCNMA1 was significantly associated with age at onset (AAO) of LOAD (P = 0.0066) and disease duration (P = 0.0002). LGI1 is an extracellular matrix (ECM) molecule forming a complex with postsynaptic scaffolding proteins (postsynaptic density proteins 95 and 93, and the synapse-associated protein 97), presynaptic scaffolding proteins (Ca2+/calmodulinactivated serine-threonine kinase and Lin7), and presynaptic K+ channels (Kv1.1, Kv1.4 and Kvβ1 subunits) and demonstrated to be important in epilepsy, but ECM and chondroitin sulfate proteoglycans (CSPGs), one of the most abundant glycanated protein types found in the nervous system and a major ECM component, which form dense lattice-like structures, termed perineuronal nets (PNNs), are thought to be neuroprotective in AD.[22] Application of Aβ1–42 to rodent primary neuronal cultures caused neuronal death of neurons not associated with PNNs, while the neurons expressing PNNs were not affected.[23] The same miR-33 target genes were observed to be enriched in other CSF biomarker suggestive hits, although the corrected enrichment p-value is greater than 0.05. In addition, miR-193 and miR-146 target genes are also enriched, miR-193 was one of the nine down-regulated miRNA identified in adult-onset AD Drosophila brains.[24] miR-193b were all upregulated in oxidative stressed (i.e. H2O2-induced) primary hippocampal neurons and different strains of senescence accelerated mice.[25]. miR-146 was reported to be related to up-regulated immune and inflammatory in Alzheimer's disease. [26]
Table 5. Inrich Analysis Results (Pcorrected < 0.05).
Discussion
In this study we have analyzed the ADNI Cohort to predict CSF and PET biomarker status and cognitive decline rates from genetic data and to assess if use of molecular biomarkers as quantitative traits provides extra power to uncover novel genotype-phenotype relationships in AD. Unequivocally, APOE ε4 allele is the strongest genetic predictor of CSF biomarker level, cerebral amyloid deposition, and disease progression. The effect size for other genetic markers is much smaller.
CSF biomarker measurements have the advantage that their measurements are widely sensitive across different patient subpopulations ranging from cognitive normal to AD. Cognitive measurements on the other hand are sensitive for different patient population segments and this will limit the sample size available for GWAS if we study the rate of cognitive decline directly in the selected patient population. For example, the Clinical Dementia Rating-Sum of Boxes (CDR-SB) is most sensitive for mild cognitive impairment (MCI) patients, while the Alzheimer’s Disease Assessment Scale-Cognitive Subscale (ADAS-Cog) will be more appropriate for AD patients. In this sample set, variants in FRA10AC1 and 15q21 were associated with CSF Aβ1–42 reaching genome-wide significance, replication was achieved with the FRA10AC1 variant. FRA10AC1 (fragile site, folic acid type, rare, fra(10)(q23.3) or fra(10)(q24.2) candidate 1) encodes a nuclear phosphorprotein of unknown function. The 5' UTR of this gene is part of a CpG island and contains a tandem CGG repeat region that normally consists of 8–14 repeats but can expand to over 200 repeats. The CGG repeat is ~723 base pairs away from the rs116953792, the most significant SNP (P = 2.0 x 10−10, in LD with the directly genotyped variant rs10509663) associated with CSF Aβ1–42 level. The expanded allele becomes hypermethylated and is not transcribed and an expanded repeat region has not been associated with any disease phenotype. The extent of LD between the CpG repeat and rs116953792, is unclear, given their close proximity.
WDFY3 is one of the biologically most interesting genes identified to have suggestive association with the florbetapir PET quantitative trait with or without correction for APOE ε4 dosage. WDFY3 encodes a phosphatidylinositol 3-phosphate-binding protein that functions as a master conductor for aggregate clearance by autophagy. This protein shuttles from the nuclear membrane to co-localize with aggregated proteins, and complexes with other autophagic components to achieve macroautophagy-mediated clearance of aggregated proteins. This protein is of particular interest given the proposed synergy between amyloid and tau aggregates in driving AD progression. Another variant (rs17009220, P = 0.00484, http://www.gwascentral.org/marker/HGVM16286779/results?t=2) in WDFY3 exhibited nominally significant interaction (SNP*APOE ε4) in an AD case-control GWAS study.[27]
GFRA1 gene, which encodes GDNF family receptor alpha 1, a member of the GDNF receptor family, is among the few genes with suggestive association to rate of cognitive decline in the LMCI subgroup. The GDNF family receptor alpha 1 is a glycosylphosphatidylinositol(GPI)-linked cell surface receptor for both glial cell line-derived neurotrophic factor (GDNF) and neurturin (NTN), and mediates activation of the RET tyrosine kinase receptor. GDNF and NTN are two structurally related, potent neurotrophic factors that play key roles in the control of neuron survival and differentiation. Neuronal loss is a hallmark of AD, a neurodegenerative disease.
Human and mouse experiments have implicated the role of miRNA in the regulation of Aβ, tau, inflammation, and cell death as the main disease mechanisms of AD [28]. In our GWAS and INRICH analyses, it was intriguing that molecules involved in Aβ autophagy, inflammation, cell death and proliferation are enriched in INRICH analysis or among the top hits of GWAS analyses. APOE ε4 has been the most convincing genome wide signal in AD, and variants in APOE-APOC1-APOC4-APOC2 and TOMM40-APOE have previously been associated with total cholesterol, LDL cholesterol, and triglyceride concentrations.[29, 30] Cholesterol metabolism was implicated to be enriched in the etiology of AD in previous study.[31] In the INRICH analysis, miR-33 targets are enriched in CSF GWAS analysis and miR-33 is critical in regulating cholesterol metabolism, affirming the interrelationship between cholesterol metabolism and AD process.
Studies comparing (a) non-demented individuals free of substantial Alzheimer’s pathology (controls), (b) non-demented individuals with equivalent loads of amyloid-β plaques (“mismatches”) and tangles, and (c) demented Alzheimer’s cases, observed four main phenotypic differences between the groups, which include demented cases had significantly higher burdens of fibrillar thioflavin-S positive plaques and of oligomeric amyloid-β deposits reactive to conformer-specific antibody NAB61 than “mismatches”.[32] Thus, florbetapir PET most likely could not distinguish between these different forms of amyloid deposition. Therefore, future amyloid phenotype differentiating the pathological forms of amyloid deposition will help genetic association study.
Future studies with larger sample sizes or replication samples are needed to further dissect the genetic basis of CSF and cerebral biomarkers. AD progression is a challenging problem as the patients are likely to be at different stages of the disease continuum, additionally, deterioration is not linear over the course of the disease progression. Furthermore, different neuropsycogntive instruments are sensitive for patients at different stages of the disease continuum, thus, genetic association studies further suffers from the sample size after stratification by disease stage.
Methods
Alzheimer’s Disease Neuroimaging Initiative
Data used in this study were obtained from the ADNI database (adni.loni.usc.edu). The ADNI study was launched in 2003 by the National Institute on Aging (NIA), the National Institute of Biomedical Imaging and Bioengineering (NIBIB), the Food and Drug Administration (FDA), private pharmaceutical companies and non-profit organizations, as a $60 million dollar, 5-year public private partnership. The primary goal of ADNI study has been to test whether serial magnetic resonance imaging (MRI), positron emission tomography (PET), other biological markers, and clinical and neuropsychological assessments can be combined to measure the progression of MCI and early AD. For up-to-date information, see www.adni-info.org.
The data from ADNIMERGE R package dated 2014-06-11 together with genetic data (Figure A in S1 File) (http://www.loni.ucla.edu/ADNI) were utilized in this manuscript. The adnimerge table merges together several of the key variables from various case report forms and biomarker lab summaries across the ADNI protocols (ADNI1, ADNIGO, and ADNI2). The details of genotype data quality control (QC), imputation, and genetic association analysis are described in the S1 File.
Inference of APOE ε4 genotype
A fraction of subjects with OmniExpress genotype data had missing APOE ε4 dosage in the ADNIMERGE database. Genotype dosage for rs429358 which is the defining variant for APOE ε4 dosage were imputed using IMPUTE2 [33–37] (v2.3.0) and the rs429358 genotype was inferred using the best guessed genotype if the probability having that best guessed genotype exceeds 90%.
CSF biomarker GWAS meta-analysis
To minimize differences due to different CSF assay batches, three sets of samples (Table 1) were used in the GWAS and referred to as Upennbiomk_Human610-Quad, Upennbiomk5_Omni2.5, and Upennbiomk6_OmniExpress. Sample sizes for CSF biomarkers after sample level QC and with phenotype data are listed in Table A in S1 File. CSF biomarker data at baseline visit of ADNI study were log transformed to approximate a normal distribution. APOE ε4 allele dosage, age and clinical diagnosis group (CN, EMCI, LMCI, or AD) at baseline visit were included as covariates. Fixed effects meta-analyses were carried out using PLINK.[38] Regional visualization of genome-wide association scan results was plotted using LocusZoom.[39]
Cerebral amyloid deposition (florbetapir PET) quantitative trait GWAS
The cerebral amyloid deposition quantitative trait as measured by florbetapir PET was taken from adnimerge table. The AV45 value in the adnimerge table is the average AV45 SUVR of frontal, anterior cingulate, precuneus, and parietal cortex relative to the cerebellum. Independent GWA analyses using florbetapir PET SUVR as a quantitative trait were performed for subjects genotyped with the Omni2.5 (N = 661) or the OmniExpress (N = 291) platform (see Table B in S1 File for basic demographic characteristics), followed by meta-analysis across the two sample sets. For patients with more than one florbetapir PET imaging data, the peak measurement was used in this analysis. Gender, age and clinical diagnosis (NL, MCI, or AD) at the time of peak florbetapir PET measurement were included as covariates. Other auxiliary analyses are described in the S1 File.
Rate of cognitive decline GWAS
LMCI subjects (N = 540) with genetic data were included in this analysis. The subset of overlapping markers shared across all samples was used to impute unobserved genotype data since chip platform is correlated with rate of cognitive decline to avoid the situation of using different sets of markers to infer unobserved genotyping being confounded with the phenotypic endpoint. The rate of cognitive decline is defined by the yearly rate of CDR-SB score change over one of the following: (1) the first two year period since baseline (if month 24 data are available); or (2) the first one year (if month 24 data are not available but month 12 data are available); or (3) the first 6 month (if only month 6 data are available). The rate of cognitive decline was also log transformed after addition of 3.5 to avoid log transformation of negative numbers using this formula ln (ΔCDRSB/duration + 3.5) to approximate normal distribution. Gender, age, baseline CDR-SB score, baseline MMSE score, and APOE ε4 allele dosage were included as covariates.
Gene Set Enrichment Analysis
INRICH is a pathway analysis tool for genome wide association studies, designed for detecting enriched association signals of LD-independent genomic regions within biologically relevant gene sets.[15] Reference gene sets used in the INRICH analysis include KEGG, Gene Ontology, and Molecular Signature Database. Top variants from CSF and florbetapir PET SUVR analyses with nominal association p-values less than 0.0005, 0.0001, 0.00005, 0.00001 were separately fed into PLINK to clump the variants into LD-independent genomic intervals (r2 threshold using 0.2, 0.3, and 0.5 respectively), then LD-independent genomic regions were used for INRICH (version 1.0) analyses. No multiple testing correction was applied for running INRICH against multiple reference gene sets or for using multiple parameters (p-value cut-off and LD threshold).
Supporting Information
(DOCX)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
Acknowledgments
Data collection and sharing for this project was funded by the Alzheimer's Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: Alzheimer’s Association; Alzheimer’s Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen Idec Inc.; Bristol-Myers Squibb Company; Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Medpace, Inc.; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Synarc Inc.; and Takeda Pharmaceutical Company. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer's Disease Cooperative Study at the University of California, San Diego. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California.
Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at: http://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf
Data Availability
Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu).
Funding Statement
The study was funded by Janssen Research & Development, LLC, Titusville, N.J., USA. Drs. Li, Parrado, Samtani, and Narayan are employees of Janssen Research & Development, LLC. The funder provided support in the form of salaries for authors QSL, ARP, MNS and VAN, but did not have any additional role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript. The funder however approved the intention to publish the final manuscript. The specific roles of these authors are articulated in the ‘author contributions’ section.
References
- 1. Consensus report of the Working Group on: "Molecular and Biochemical Markers of Alzheimer's Disease". The Ronald and Nancy Reagan Research Institute of the Alzheimer's Association and the National Institute on Aging Working Group. Neurobiology of aging. 1998;19(2):109–16. . [PubMed] [Google Scholar]
- 2. Blennow K, Hampel H. CSF markers for incipient Alzheimer's disease. The Lancet Neurology. 2003;2(10):605–13. . [DOI] [PubMed] [Google Scholar]
- 3. Frank RA, Galasko D, Hampel H, Hardy J, de Leon MJ, Mehta PD, et al. Biological markers for therapeutic trials in Alzheimer's disease. Proceedings of the biological markers working group; NIA initiative on neuroimaging in Alzheimer's disease. Neurobiology of aging. 2003;24(4):521–36. . [DOI] [PubMed] [Google Scholar]
- 4. Sunderland T, Linker G, Mirza N, Putnam KT, Friedman DL, Kimmel LH, et al. Decreased beta-amyloid1-42 and increased tau levels in cerebrospinal fluid of patients with Alzheimer disease. Jama. 2003;289(16):2094–103. 10.1001/jama.289.16.2094 . [DOI] [PubMed] [Google Scholar]
- 5. Shaw LM, Vanderstichele H, Knapik-Czajka M, Clark CM, Aisen PS, Petersen RC, et al. Cerebrospinal fluid biomarker signature in Alzheimer's disease neuroimaging initiative subjects. Annals of neurology. 2009;65(4):403–13. 10.1002/ana.21610 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Samtani MN, Raghavan N, Shi Y, Novak G, Farnum M, Lobanov V, et al. Disease progression model in subjects with mild cognitive impairment from the Alzheimer's disease neuroimaging initiative: CSF biomarkers predict population subtypes. British journal of clinical pharmacology. 2013;75(1):146–61. 10.1111/j.1365-2125.2012.04308.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Hansson O, Zetterberg H, Buchhave P, Londos E, Blennow K, Minthon L. Association between CSF biomarkers and incipient Alzheimer's disease in patients with mild cognitive impairment: a follow-up study. The Lancet Neurology. 2006;5(3):228–34. 10.1016/S1474-4422(06)70355-6 . [DOI] [PubMed] [Google Scholar]
- 8. Andersson C, Blennow K, Almkvist O, Andreasen N, Engfeldt P, Johansson SE, et al. Increasing CSF phospho-tau levels during cognitive decline and progression to dementia. Neurobiology of aging. 2008;29(10):1466–73. 10.1016/j.neurobiolaging.2007.03.027 . [DOI] [PubMed] [Google Scholar]
- 9. Salloway S, Sperling R, Fox NC, Blennow K, Klunk W, Raskind M, et al. Two phase 3 trials of bapineuzumab in mild-to-moderate Alzheimer's disease. The New England journal of medicine. 2014;370:322–33. 10.1056/NEJMoa1304839 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Kim S, Swaminathan S, Shen L, Risacher SL, Nho K, Foroud T, et al. Genome-wide association study of CSF biomarkers Abeta1-42, t-tau, and p-tau181p in the ADNI cohort. Neurology. 2011;76(1):69–79. 10.1212/WNL.0b013e318204a397 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Cruchaga C, Kauwe JS, Mayo K, Spiegel N, Bertelsen S, Nowotny P, et al. SNPs associated with cerebrospinal fluid phospho-tau levels influence rate of decline in Alzheimer's disease. PLoS genetics. 2010;6(9):e1001101 10.1371/journal.pgen.1001101 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Cruchaga C, Kauwe JS, Harari O, Jin SC, Cai Y, Karch CM, et al. GWAS of cerebrospinal fluid tau levels identifies risk variants for Alzheimer's disease. Neuron. 2013;78(2):256–68. 10.1016/j.neuron.2013.02.026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Ramanan VK, Risacher SL, Nho K, Kim S, Swaminathan S, Shen L, et al. APOE and BCHE as modulators of cerebral amyloid deposition: a florbetapir PET genome-wide association study. Molecular psychiatry. 2014;19(3):351–7. 10.1038/mp.2013.19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Hu X, Pickering EH, Hall SK, Naik S, Liu YC, Soares H, et al. Genome-wide association study identifies multiple novel loci associated with disease progression in subjects with mild cognitive impairment. Translational psychiatry. 2011;1:e54 10.1038/tp.2011.50 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Lee PH, O'Dushlaine C, Thomas B, Purcell SM. INRICH: interval-based enrichment analysis for genome-wide association studies. Bioinformatics. 2012;28(13):1797–9. 10.1093/bioinformatics/bts191 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Rayner KJ, Suarez Y, Davalos A, Parathath S, Fitzgerald ML, Tamehiro N, et al. MiR-33 contributes to the regulation of cholesterol homeostasis. Science. 2010;328:1570–3. 10.1126/science.1189862 Epub 2010 May 13. . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Rotllan N, Fernandez-Hernando C. MicroRNA Regulation of Cholesterol Metabolism. Cholesterol. 2012;2012:847849 10.1155/2012/847849 Epub 2012 Aug 5. . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Cho MH, Cho K, Kang HJ, Jeon EY, Kim HS, Kwon HJ, et al. Autophagy in microglia degrades extracellular beta-amyloid fibrils and regulates the NLRP3 inflammasome. Autophagy. 2014;10(10):1761–75. 10.4161/auto.29647 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Carrasquillo MM, Belbin O, Hunter TA, Ma L, Bisceglio GD, Zou F, et al. Replication of EPHA1 and CD33 associations with late-onset Alzheimer's disease: a multi-centre case-control study. Molecular neurodegeneration. 2011;6:54 10.1186/1750-1326-6-54 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Naj AC, Jun G, Beecham GW, Wang LS, Vardarajan BN, Buros J, et al. Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer's disease. Nature genetics. 2011;43:436–41. 10.1038/ng.801 Epub 2011 Apr 3. . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Hollingworth P, Harold D, Sims R, Gerrish A, Lambert JC, Carrasquillo MM, et al. Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer's disease. Nature genetics. 2011;43:429–35. 10.1038/ng.803 Epub 2011 Apr 3. . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Soleman S, Filippov MA, Dityatev A, Fawcett JW. Targeting the neural extracellular matrix in neurological disorders. Neuroscience. 2013;253:194–213. 10.1016/j.neuroscience.2013.08.050 . [DOI] [PubMed] [Google Scholar]
- 23. Miyata S, Nishimura Y, Nakashima T. Perineuronal nets protect against amyloid beta-protein neurotoxicity in cultured cortical neurons. Brain research. 2007;1150:200–6. 10.1016/j.brainres.2007.02.066 . [DOI] [PubMed] [Google Scholar]
- 24. Kong Y, Wu J, Yuan L. MicroRNA expression analysis of adult-onset Drosophila Alzheimer's disease model. Current Alzheimer research. 2014;11(9):882–91. . [PubMed] [Google Scholar]
- 25. Zhang R, Zhang Q, Niu J, Lu K, Xie B, Cui D, et al. Screening of microRNAs associated with Alzheimer's disease using oxidative stress cell model and different strains of senescence accelerated mice. Journal of the neurological sciences. 2014;338(1–2):57–64. 10.1016/j.jns.2013.12.017 . [DOI] [PubMed] [Google Scholar]
- 26. Wang LL, Huang Y, Wang G, Chen SD. The potential role of microRNA-146 in Alzheimer's disease: biomarker or therapeutic target? Medical hypotheses. 2012;78(3):398–401. 10.1016/j.mehy.2011.11.019 . [DOI] [PubMed] [Google Scholar]
- 27. Li H, Wetten S, Li L, St Jean PL, Upmanyu R, Surh L, et al. Candidate single-nucleotide polymorphisms from a genomewide association study of Alzheimer disease. Archives of neurology. 2008;65(1):45–53. 10.1001/archneurol.2007.3 . [DOI] [PubMed] [Google Scholar]
- 28. Tan L, Yu JT, Hu N, Tan L. Non-coding RNAs in Alzheimer's disease. Molecular neurobiology. 2013;47(1):382–93. 10.1007/s12035-012-8359-5 . [DOI] [PubMed] [Google Scholar]
- 29. Aulchenko YS, Ripatti S, Lindqvist I, Boomsma D, Heid IM, Pramstaller PP, et al. Loci influencing lipid levels and coronary heart disease risk in 16 European population cohorts. Nature genetics. 2009;41(1):47–55. 10.1038/ng.269 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Kathiresan S, Melander O, Guiducci C, Surti A, Burtt NP, Rieder MJ, et al. Six new loci associated with blood low-density lipoprotein cholesterol, high-density lipoprotein cholesterol or triglycerides in humans. Nature genetics. 2008;40(2):189–97. 10.1038/ng.75 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Jones L, Holmans PA, Hamshere ML, Harold D, Moskvina V, Ivanov D, et al. Genetic evidence implicates the immune system and cholesterol metabolism in the aetiology of Alzheimer's disease. PloS one. 2010;5:e13950 10.1371/journal.pone.0013950 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Perez-Nievas BG, Stein TD, Tai HC, Dols-Icardo O, Scotton TC, Barroeta-Espar I, et al. Dissecting phenotypic traits linked to human resilience to Alzheimer's pathology. Brain: a journal of neurology. 2013;136(Pt 8):2510–26. 10.1093/brain/awt171 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Marchini J, Howie B. Genotype imputation for genome-wide association studies. Nature reviews Genetics. 2010;11(7):499–511. 10.1038/nrg2796 . [DOI] [PubMed] [Google Scholar]
- 34. Marchini J, Howie B, Myers S, McVean G, Donnelly P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nature genetics. 2007;39(7):906–13. 10.1038/ng2088 . [DOI] [PubMed] [Google Scholar]
- 35. Howie B, Fuchsberger C, Stephens M, Marchini J, Abecasis GR. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nature genetics. 2012;44(8):955–9. 10.1038/ng.2354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Howie B, Marchini J, Stephens M. Genotype imputation with thousands of genomes. G3. 2011;1(6):457–70. 10.1534/g3.111.001198 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Howie BN, Donnelly P, Marchini J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS genetics. 2009;5(6):e1000529 10.1371/journal.pgen.1000529 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. American journal of human genetics. 2007;81(3):559–75. . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Pruim RJ, Welch RP, Sanna S, Teslovich TM, Chines PS, Gliedt TP, et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics. 2010;26(18):2336–7. 10.1093/bioinformatics/btq419 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
(DOCX)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
(XLSX)
Data Availability Statement
Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu).