Abstract
Sequence heterogeneity at the ends of mature microRNAs (miRNAs) is well documented, but its effects on miRNA function are largely unexplored. Here we studied the impact of miRNA 5′-heterogeneity, which affects the seed region critical for target recognition. Using the example of miR-142-3p, an emerging regulator of the hematopoietic lineage in vertebrates, we show that naturally coexpressed 5′-variants (5′-isomiRs) can recognize largely distinct sets of binding sites. Despite this, both miR-142-3p isomiRs regulate exclusive and shared targets involved in actin dynamics. Thus, 5′-heterogeneity can substantially broaden and enhance regulation of one pathway. Other 5′-isomiRs, in contrast, recognize largely overlapping sets of binding sites. This is exemplified by two herpesviral 5′-isomiRs that selectively mimic one of the miR-142-3p 5′-isomiRs. We hypothesize that other cellular and viral 5′-isomiRs can similarly be grouped into those with divergent or convergent target repertoires, based on 5′-sequence features. Taken together, our results provide a detailed characterization of target recognition by miR-142-3p and its 5′-isomiR-specific viral mimic. We furthermore demonstrate that miRNA 5′-end variation leads to differential targeting and can thus broaden the target range of miRNAs.
Keywords: isomiR, miR-142-3p, miR-K10a, Kaposi's sarcoma associated herpesvirus, miRNA
INTRODUCTION
miRNAs are a class of ∼22-nucleotide (nt) long noncoding RNAs that negatively regulate mRNA expression and translation (Bartel 2009). Mature miRNAs are generally derived from one arm of an imperfect stem–loop precursor contained within longer primary miRNAs by sequential endonucleolytic processing (Ha and Kim 2014). First, the microprocessor complex, comprised of Drosha and its cofactor DGCR8, introduces staggered cuts near the base of the stem to liberate the stem–loop pre-miRNA. Upon pre-miRNA export from the nucleus, Dicer makes a second set of staggered cuts to excise an ∼22-nt imperfect duplex with 3′-overhangs. Mature miRNAs can originate from either the 5p or 3p arm of this duplex and are loaded into one of four Argonaute proteins (Ago1-4) to form active RNA-induced silencing complexes (RISCs). miRNA-mediated mRNA repression is primarily facilitated by base-pairing of the miRNA seed region (Lewis et al. 2003), that is, nucleotides (nts) 2–7, with sites in the 3′-untranslated regions (3′-UTRs) of target mRNAs. To achieve more than marginal regulation, seed matches are accompanied by an adenosine (A) immediately 3′ to the binding site (“7merA1” match, referred to here as “2–7A” to indicate base-paired nts and the A across from nt 1 of the miRNA) and/or extended base-pairing including at least nt 8 of the miRNA (“7mer-m8 match,” referred to as “2–8” to indicate base-paired nts) (Lewis et al. 2005; Bartel 2009). The importance of these empirical seed rules is supported by the evolutionary conservation of these seed motifs in the targets of conserved miRNAs. The crystal structure of human Ago2 bound to an RNA guide suggests that the miRNA seed region is specifically displayed for target recognition (Schirle et al. 2014). The structure also supports a preference of Ago for an A opposite nt 1 of the miRNA, which furthermore enhanced the in vitro target affinity approximately threefold over sites with non-A bases in this position. While studies agree that seed matches are the most common miRNA-binding sites, noncanonical interactions with sub-optimal seed base-pairing and compensatory features have been described (Ha et al. 1996; Vella et al. 2004; Grimson et al. 2007; Lal et al. 2009; Shin et al. 2010; Chi et al. 2012; Loeb et al. 2012; Helwak et al. 2013; Khorshid et al. 2013; Majoros et al. 2013; Grosswendt et al. 2014).
Small RNA sequencing studies in multiple species have yielded an increasingly detailed understanding of the miRNA repertoire (Ruby et al. 2006, 2007; Landgraf et al. 2007; Morin et al. 2008; Chiang et al. 2010; Berezikov et al. 2011; Cloonan et al. 2011; Loher et al. 2014; Xia and Zhang 2014). These studies have shown that miRNA 3′-ends are generated with low stringency resulting in the frequent coexpression of 3′-variants. In contrast, miRNA 5′-ends are more uniform, reflecting the need for defined seed sequences through high fidelity of miRNA 5′-processing. miRNA variants derived from the same arm of one pre-miRNA are referred to as isomiRs (Morin et al. 2008). Despite the typical uniformity of miRNA 5′-ends, several miRNAs are expressed with more than one defined 5′-end, giving rise to 5′-isomiRs or seed-isomiRs. Because 5′-end variation redefines the miRNA seed region, a microRNA with 5′-isomiRs of significant abundance could have a substantially different target repertoire and functional impact compared to a miRNA with a single seed. Abundant 5′-isomiRs have been documented for Caenorhabditis elegans (Ruby et al. 2006), Drosophila melanogaster (Berezikov et al. 2011), mice (Chiang et al. 2010), humans (Morin et al. 2008; Cloonan et al. 2011), and herpesviruses (Umbach and Cullen 2010; Gottwein et al. 2011), among other organisms. In many cases, 5′-isomiR expression appears to be evolutionarily conserved and 5′-isomiRs associate with Ago proteins (Azuma-Mukai et al. 2008; Lee et al. 2010; Berezikov et al. 2011; Cloonan et al. 2011; Tan et al. 2014; Xia and Zhang 2014). Evidence that endogenously expressed 5′-isomiRs could indeed have functional impact comes from gene expression studies in miR-223-deleted murine neutrophils (Baek et al. 2008). Up-regulated mRNAs were significantly enriched not only for those with seed matches to miR-223, but also for seed matches exclusive to a minor miR-223 variant that accounted for only 12% of all miR-223 sequences and lacks the 5′-terminal uridine (U) of miR-223 (Chiang et al. 2010). Thus, miR-223 5′-isomiR expression appears to broaden the overall range of miR-223 targets. The prediction that 5′-isomiR expression can impact miRNA target ranges is further supported by the confirmation of small sets of differentially regulated targets for an aberrant miR-307 5′-isomiR in flies (Fukunaga et al. 2012) and for transfected 5′-isomiRs of miR-133a, miR-101, and miR-9 (Humphreys et al. 2012; Llorens et al. 2013; Tan et al. 2014). On the other hand, it has been suggested that 5′-isomiRs have highly overlapping targets (Cloonan et al. 2011; Llorens et al. 2013) and could therefore act redundantly, to increase the effective miRNA dosage or reduce off-target effects (Cloonan et al. 2011). Thus, a clear understanding of the impact of miRNA 5′-variants is still lacking.
Our interest in understanding the impact of 5′-isomiR expression was prompted by our functional work on two herpesviral 5′-isomiRs that share identical and offset seed sequences with two miR-142-3p 5′-isomiRs. miR-142-3p expression is specific to the vertebrate hematopoietic lineage, where it is among the most highly expressed miRNAs (Chen et al. 2004; Landgraf et al. 2007). Overexpression of the miR-142 precursor in mouse hematopoietic progenitor cells substantially increases the T-cell population in vitro (Chen et al. 2004). The inactivation of miR-142-3p causes defects in hematopoiesis in zebrafish (Nishiyama et al. 2012) and prevents the specification of definitive hemangioblasts in Xenopus (Nimmo et al. 2013). In mice, ablation of the miR-142 locus results in reduced numbers of CD4+ dendritic cells (Mildner et al. 2013) and a severe impairment of platelet formation (Chapnik et al. 2014). This latter phenotype is a consequence of the dysregulation of the actin cytoskeleton in megakaryocytes, the cell type responsible for platelet production (Chapnik et al. 2014). miR-142-3p is coexpressed with an abundant 5′-isomiR that lacks the 5′-terminal U (Fig. 1A, referred to as miR-142-3p−1 here) and both 5′-isomiRs are found in the RISC (Wu et al. 2007, 2009; Azuma-Mukai et al. 2008; Chiang et al. 2010; Gottwein et al. 2011). miR-142-3p 5′-isomiR expression has been suggested to result from differential processing of the primary miR-142 transcript by Drosha, leading to the production of two major pre-miRNAs (Wu et al. 2009; Ma et al. 2013). These are then each precisely processed by Dicer to define the 5′-ends of each 5′-isomiR. Functional studies of miR-142-3p to date have not considered the impact of miR-142-3p 5′-isomiR expression.
FIGURE 1.
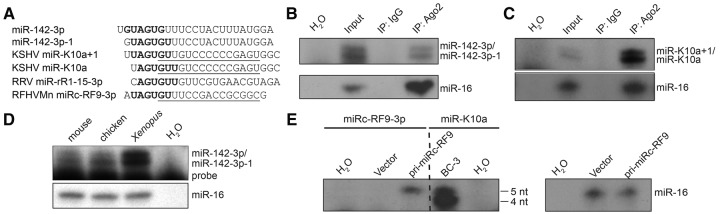
The miR-142-3p and miR-K10a 5′-isomiRs. (A) Sequences of the miR-142-3p and miR-K10a 5′-isomiRs, RRV miR-rR1-15-3p, and RFHVMn miRc-RF9-3p. Seed sequences are in bold. Underlined sequences indicate hybridization of the primers used for primer extension analysis of RFHVMn miRc-RF9-3p and KSHV miR-K10a/+1 shown in panel E. (B,C) Primer extension analyses of (B) miR-142-3p/−1 and (C) miR-K10a/+1 in Ago2-immunoprecipitates from PEL cell lines. (D) Primer extension analysis of miR-142-3p/−1 in total RNA from mouse, chicken, and Xenopus spleens. (E) Primer extension analysis of RFHVMn miRc-RF9-3p, KSHV miR-K10a/+1, and miR-16 in total RNA from rhesus LLC-MK2 cells transfected with a construct expressing pri-miRc-RF9. The miR-K10a/+1 extension products from the KSHV-infected primary effusion lymphoma (PEL) cell line BC-3 served as a molecular weight ladder for 5- or 4-nt extension products.
Interestingly, the miR-142-3p miRNAs are subject to herpesviral mimicry (Gottwein et al. 2011). The oncogenic Kaposi's sarcoma-associated herpesvirus (KSHV) infects B cells and endothelial cells to cause lymphoproliferative disorders and the AIDS-defining cancer Kaposi's sarcoma, respectively. Like many other herpesviruses, KSHV expresses its own set of ∼20 mature miRNAs (Cai et al. 2005; Pfeffer et al. 2005; Samols et al. 2005; Grundhoff et al. 2006; Umbach and Cullen 2010). These microRNAs include known viral mimics of cellular miR-155 and miR-23 (Gottwein et al. 2007; Skalsky et al. 2007; Manzano et al. 2013). One of the KSHV miRNA precursors is processed to two approximately equally abundant 5′-isomiRs called miR-K10a and miR-K10a+1 (Fig. 1A; Umbach and Cullen 2010; Gottwein et al. 2011). The seed region of miR-K10a+1 is identical to that of miR-142-3p−1 (Gottwein et al. 2011), while the seed sequence of miR-K10a is offset by 1 nt. This has led us to hypothesize that the miR-K10a 5′-isomiRs are at least partial viral mimics of miR-142-3p (Gottwein et al. 2011). Luciferase reporter assays following the ectopic coexpression of miR-142-3p/−1 or miR-K10a/+1 from pri-miRNA-derived expression cassettes has suggested that the miR-K10a isomiRs together can indeed regulate several targets also regulated by the miR-142-3p isomiRs (Gottwein et al. 2011). However, the extent of this mimicry and the contributions of the individual isomiRs remain unaddressed.
Here we used the miR-K10a and miR-142-3p 5′-isomiRs to characterize how miRNA 5′-variation affects the range of targets recognized. We show that 5′-isomiRs can target largely overlapping or largely discrepant sets of binding sites, depending on 5′-sequence features. We specifically demonstrate that the 5′-isomiRs of cellular miR-142-3p act as distinct regulatory units. In contrast, the two viral miR-K10a 5′-isomiRs are largely similar. Together, they not only regulate targets of mainly the miR-142-3p−1 isomiR but also extend their range of targets beyond this mimicry. Thus, KSHV appears to cherry-pick and customize a preexisting regulatory network of miR-142-3p targets. Finally, we hypothesize that other 5′-isomiRs also have convergent or divergent target ranges, depending on seed sequence features.
RESULTS
miR-142-3p 5′-isomiR expression is conserved in vertebrates
It has previously been demonstrated that both miR-142-3p 5′-isomiRs are expressed in humans and mice (Supplemental Table S1). The reported relative abundance of the miR-142-3p 5′-isomiRs varies slightly between settings and miR-142-3p−1 is often detected at slightly higher frequency than miR-142-3p in total RNA or endogenous RISCs (Supplemental Table S1; Azuma-Mukai et al. 2008). We independently confirmed RISC-association of both 5′-isomiRs using primer extension analyses of Ago2-immunoprecipitates from latently KSHV-infected primary effusion lymphoma (PEL) B-cell lines (Fig. 1B). These data confirm that both miR-142-3p and miR-142-3p−1 are loaded into Ago2. Similarly, both miR-K10a and miR-K10a+1 were present in the RISC (Fig. 1C). Consistent with the ratio of the miR-K10a 5′-isomiRs detected by deep sequencing (Supplemental Table S2; Gottwein et al. 2011), the miR-K10a 5′-isomiR is slightly more abundant in PEL cell lines than miR-K10a+1. Because of the emerging pivotal role of miR-142-3p in the vertebrate hematopoietic lineage (Nishiyama et al. 2012; Mildner et al. 2013; Nimmo et al. 2013; Chapnik et al. 2014), we assessed whether the expression of mature miR-142-3p−1 is conserved beyond mice and humans. The predicted mature miR-142-3p/−1 sequences are invariant across vertebrates (http://genome.ucsc.edu/), but less conserved portions of the pri-miRNA could lead to differences in isomiR expression. Primer extension analysis confirmed that both miR-142-3p 5′-isomiRs are coexpressed in Xenopus and chicken (Fig. 1D). Thus, we conclude that miR-142-3p/−1 5′-isomiR expression is very likely to be conserved across vertebrates.
Old World primate rhadinovirus miRNAs with miR-142-3p-like seed sequences
Herpesviruses are evolutionarily ancient viruses that have co-evolved with their host species (Pellett and Roizman 2013). While most herpesviruses encode miRNAs, only the most closely related species sometimes share homologous miRNAs (Cai et al. 2006). The two human γ-herpesviruses, KSHV and Epstein–Barr virus (EBV), representing the rhadino- and lymphocryptovirus genera, do not share either miRNA homologs or miRNAs with identical seed sequences. However, the primate Rhesus rhadinovirus (RRV) encodes miR-rR1-15, which has a seed sequence identical to that of miR-K10a (Fig. 1A; Umbach et al. 2010). Another primate rhadinovirus, retroperitoneal fibromatosis-associated herpesvirus Macaca nemestrina (RFHVMn), carries a predicted miRNA precursor, miRc-RF9, with the potential to generate miRc-RF9-3p miRNAs with seed sequences identical to miR-K10a and/or miR-K10a+1 (Fig. 1A; Bruce et al. 2013). To determine the 5′-end(s) of miRc-RF9-3p, we cloned and expressed a 250-nt fragment of pri-miRc-RF9. Primer extension analysis yielded only one 5-nt extension product, indicating that miRc-RF9-3p shares its seed with miR-K10a+1 (Fig. 1E). Thus, Old World primate rhadinoviruses analyzed to date encode either miR-K10a or miR-K10a+1-like miRNAs. Other KSHV miRNAs have not been found to have counterparts in primate rhadinoviruses (Umbach et al. 2010; Bruce et al. 2013), suggesting that miR-K10a/+1-like miRNAs may be particularly important components of the genetic makeup of Old World primate rhadinoviruses and KSHV.
The seed region governs differential target recognition by the miR-142-3p and miR-K10a 5′-isomiRs
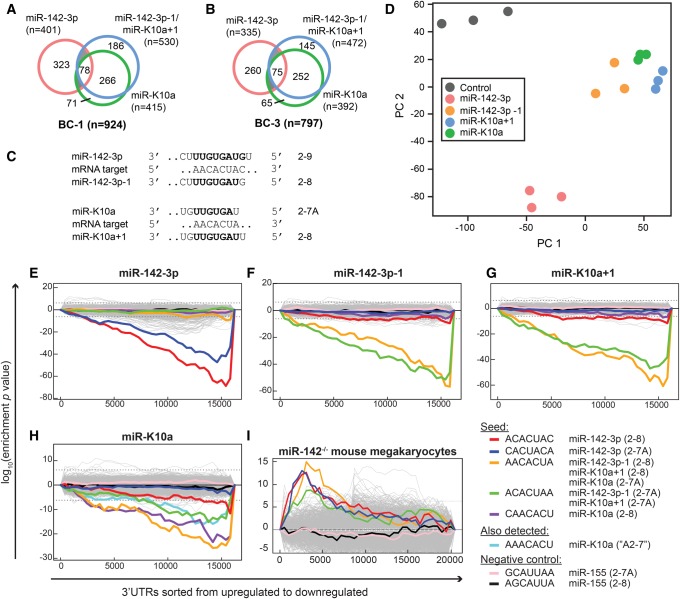
To begin to investigate how the miR-142-3p and miR-K10a 5′-isomiRs are functionally related, we compared the binding sites assigned to each of these miRNAs in Ago2-PAR-CLIP (photoactivatable-ribonucleoside-enhanced crosslinking and immunoprecipitation) data sets from the KSHV-infected PEL cell lines BC-1 and BC-3 (Supplemental Table S3; Gottwein et al. 2011). BC-1 and BC-3 cells express all four miRNAs. PAR-CLIP in combination with next generation sequencing and computational analysis identifies miRNA-binding sites at nucleotide resolution (Hafner et al. 2010; Corcoran et al. 2011). The targeting miRNAs are computationally assigned using seed matching. Of note, we did not detect significant numbers of miRNA:binding site chimeric reads from a reanalysis of our Ago2-PAR-CLIP data set that would allow the unambiguous experimental assignment of the targeting miR-K10a or miR-142-3p 5-isomiR to their binding sites (Grosswendt et al. 2014). Because miR-142-3p−1 and miR-K10a+1 have identical seeds, they are predicted to share their canonical target sites. Our analysis suggested that binding sites of the miR-K10a and miR-142-3p 5′-isomiRs fall into five groups that were roughly similarly represented in BC-1 and BC-3 cells. These are target sites of (1) miR-142-3p only; (2) miR-142-3p−1/miR-K10a+1 only; (3) miR-K10a only; (4) all except miR-142-3p; and (5) all four miRNAs (Fig. 2A,B).
FIGURE 2.
Target recognition by the miR-142-3p/−1 and miR-K10a/+1 5′-isomiRs. (A,B) Overlap of binding sites for the miR-142-3p and miR-K10a 5′-isomiRs identified in Ago2-PAR-CLIP data from (A) BC-1 and (B) BC-3 PEL cell lines (Gottwein et al. 2011). (C) Minimum base-pairing required for the miR-142-3p (upper panel) and miR-K10a (lower panel) 5′-isomiRs to share canonical binding sites. (D) Principal component analysis of microarray data of HEK293T cells transfected with individual 5′-isomiRs. (E–I) Sylamer (van Dongen et al. 2008) enrichment landscape plots for 7mer 3′-UTR matches to miR-142-3p and miR-K10a 5′-isomiRs using microarray data from 293T cells transfected with individual miRNA mimics (E–H, this study) or from published microarray data from miR-142−/− mouse megakaryocytes (GEO data set GSE52141, Chapnik et al. 2014). The x-axis represents the ranked gene lists. miR-155 sites (pink and black lines) served as negative controls in addition to all random 7mers (gray). Enrichment plots for hexamer motifs are shown in Supplemental Figure S2.
Strikingly, the majority of PAR-CLIP-assigned binding sites for miR-142-3p and miR-142-3p−1 (≥75% for miR-142-3p and ≥84% for miR-142-3p−1) are distinct from one another (Fig. 2A,B). This suggests that these 5-isomiRs represent distinct regulatory entities and is in stark contrast to other reports, which concluded that 5′-isomiRs mostly share their targets (Cloonan et al. 2011; Llorens et al. 2013).
We next considered the relationship between the two miR-K10a 5′-isomiRs. While the miR-K10a 5′-isomiRs are also offset from each other by 1 nt, the overlap of their binding sites is remarkably greater than between the miR-142-3p 5′-isomiRs (∼65%–70% shared sites for miR-K10a+1 and >80% for miR-K10a, Fig. 2A,B). This is because nt 2 of miR-K10a+1 is a U, which implies that all of its ≥2–8 seed matches are also candidate ≥2–7A binding sites of miR-K10a (Fig. 2C). Finally, we considered how the miR-K10a 5′-isomiRs relate to the miR-142-3p 5′-isomiRs. Based on the assignment of PAR-CLIP sites, miR-K10a and miR-K10a+1 together may selectively mimic miR-142-3p−1 and share only a minor subset of their targets with miR-142-3p (Fig. 2A,B). These data also predict that miR-K10a extends the range of targets beyond that of the miR-142-3p miRNAs.
Because PAR-CLIP binding sites were assigned computationally using seed rules, this analysis cannot definitively distinguish the targets of each isomiR. Furthermore, this analysis did not account for potential non-seed interactions. For an unbiased comparison of the regulatory potential of these isomiRs, we therefore performed Illumina microarray gene expression analyses of HEK293T cells transfected with mimics of the individual isomiRs. 293T cells do not express endogenous miR-142-3p/−1 or miR-K10a/+1 and therefore represent a clean background for this experiment. Hierarchal clustering (Supplemental Fig. S1) and principal component analysis (PCA, Fig. 2D) showed that expression profiles following transfection of miR-142-3p and miR-142-3p−1 were quite distinct, while those for viral miR-K10a and miR-K10a+1 were similar. In addition, these data clearly showed that miR-K10a or miR-K10a+1 affect gene expression similarly to miR-142-3p−1, but not miR-142-3p. The microarray data are therefore consistent with the PAR-CLIP binding site assignment presented above (Fig. 2A,B) and further suggest that miR-K10a/+1 specifically mimic miR-142-3p−1 expression.
To test whether these changes in gene expression are indeed driven by the miRNA seed sequences, we performed Sylamer analysis (van Dongen et al. 2008). The Sylamer algorithm computationally identifies overrepresented sequences in the 3′-UTRs of differentially expressed mRNAs in an unbiased manner. For miR-142-3p, miR-142-3p−1, and miR-K10a+1, the only detected enrichments were for the exact 2–7, 2–8, and 2–7A seed matches among down-regulated mRNAs (Fig. 2E–G; Supplemental Fig. S2a–c). The analysis easily distinguished the signatures of miR-142-3p and miR-142-3p−1, which underscores that these miRNAs are different regulatory entities (Fig. 2E,F; Supplemental Fig. S2a,b). Moreover, the enriched seed match motifs for miR-142-3p−1 and miR-K10a+1 are shared, confirming that miR-K10a+1 is a direct mimic of miR-142-3p−1 (Fig. 2F,G; Supplemental Fig. S2b,c). For miR-K10a, we also detected an enrichment of the canonical hexa- and heptameric seed matches among down-regulated genes (Fig. 2H; Supplemental Fig. S2d). In addition to the 2–7A seed match to miR-K10a (AACACUA), which is shared with the 2–8 seed match of miR-142-3p−1/miR-K10a+1, the 2–7A seed match of miR-K10+1/miR-142-3p−1 (ACACUAA) was also enriched among down-regulated genes in miR-K10a transfected cells. Similarly, mRNAs with an AAACACU motif, corresponding to a 2–7 seed match with an A across from G8 of miR-K10a, were depleted upon miR-K10a expression. The enrichment of this motif is interesting, given that this A does not pair to miR-K10a, but could potentially base pair to the U at position 9 of miR-142-3p−1. The enrichment of these two additional motifs therefore suggests that miR-K10a indeed taps into the target pool of miR-142-3p−1. Taken together, these data strongly support the notion that the miR-142-3p isomiRs have distinct targets, while the miR-K10a isomiRs share their targets and together selectively mimic miR-142-3p−1. These relationships are driven by the seed sequences of these miRNAs.
miR-142-3p−1 is a functional miRNA in vivo
Having established that Sylamer readily distinguishes signatures of miR-142-3p and miR-142-3p−1, we queried published microarray data from miR-142−/− mice for hexa- or heptamer motifs that were overrepresented in the 3′-UTRs of up-regulated mRNAs (Chapnik et al. 2014). The 6mer seed matches to both miR-142-3p 5′-isomiRs were among the three most highly enriched hexamers in mRNAs that were up-regulated in cells lacking miR-142 expression (Supplemental Fig. S2e). Similarly, 2–7A and 2–8 matches to either miR-142-3p or miR-142-3p−1 were significantly enriched in the 3′-UTRs of up-regulated mRNAs (Fig. 2I). Thus, these data further support the notion that both 5′-isomiRs are functional and contribute to the regulatory potential of miR-142 in vivo.
miR-142-3p/−1 and miR-K10a/+1 result in only marginal regulation of non-seed sites
The PCA and Sylamer analyses clearly indicated that the seed is the major determinant of target recognition for the miR-142-3p and miR-K10a miRNAs. These data, however, do not exclude the possibility of functional noncanonical interactions. To determine how 5′-isomiR expression by miR-142-3p and miR-K10a would affect the regulation of noncanonical sites, we constructed optimal in tandem reporters for several types of previously described noncanonical sites. These included “centered sites” (Shin et al. 2010), “3′-supplementary sites” (Grimson et al. 2007; Bartel 2009), and “pivot sites” (Chi et al. 2012; Fig. 3; Supplemental Fig. S3). We also constructed a positive control vector with a seed site that is predicted to be shared by all miR-142-3p and miR-K10a 5′isomiRs. Resulting reporters were tested under conditions that result in robust repression of canonical seed sites (Figs. 3B, 4, 5). Under these conditions, the large majority of noncanonical sites did not result in significant reporter inhibition. Marginal, but significant, regulation was observed for two sites that have a 1-nt bulge in the seed match for miR-K10a+1 (“PIV-G” and “PIV-U,” Fig. 3E; Supplemental Fig. S3d). One of these sites, i.e., “PIV-U,” can be considered a pivot site. The non-pivot control “PIV-G” unexpectedly resulted in similar reporter repression as “PIV-U,” suggesting that some bulged nts are tolerated whether or not they can nucleate a pivot interaction as previously described (Chi et al. 2012). It is also possible that these interactions are facilitated by extended seed base-pairing or 3′-supplementary interactions of miR-K10a+1 in this sequence context. Taken together, the data presented so far strongly support the idea that these four miRNAs recognize their binding sites through canonical seed interactions, which drive their impact on gene expression. We cannot exclude that individual non-seed interactions exist that could lead to subtle functional differences between miR-K10a+1 and miR-142-3p−1 (see below).
FIGURE 3.
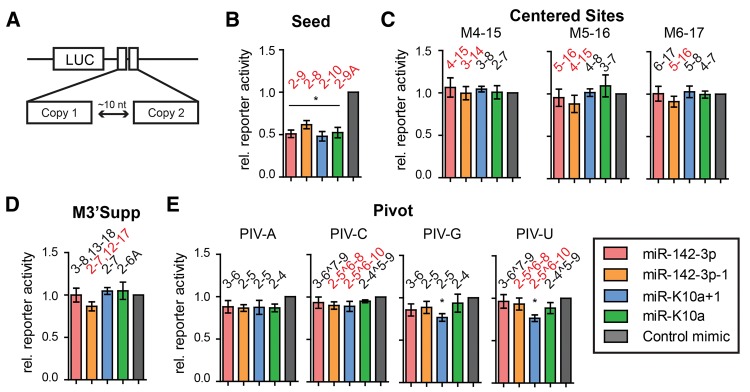
Regulation of noncanonical binding sites by miR-142-3p/−1 and miR-K10a/+1 is weak or not detected. (A) Schematic of the 3′-UTR reporter constructs used in this figure. To ensure optimal sensitivity, two identical miRNA-binding sites were cloned in tandem, separated by an ∼10-nt spacer. Firefly luciferase data were normalized to values from an internal Renilla luciferase control, a construct containing only the ∼10-nt spacer sequence, and control mimic transfections. (B) A canonical seed site was repressed by all four miRNAs, as expected. (C–E) Prototypical noncanonical sites were designed and tested for previously reported noncanonical modes of interaction, including “centered sites” (C, Shin et al. 2010); “3′ Supplementary Sites” (D, Grimson et al. 2007); and “Pivot Sites” (E, Chi et al. 2012). PIV-C and PIV-U are pivot sites, while PIV-A and PIV-G are not predicted to function as pivot sites and represent controls (see Supplemental Fig. S3). Predicted base-paired nts are indicated above columns; configurations predicted to be functional based on previous reports are in red. (^) Bulged target nts. (*) P < 0.05 by Student's t-test, error bars indicate SEM; n ≥ 3 biological replicates. Sequences of the miRNA-binding sites are found in Supplemental Figure S3, cloning details in Supplemental Table S4.
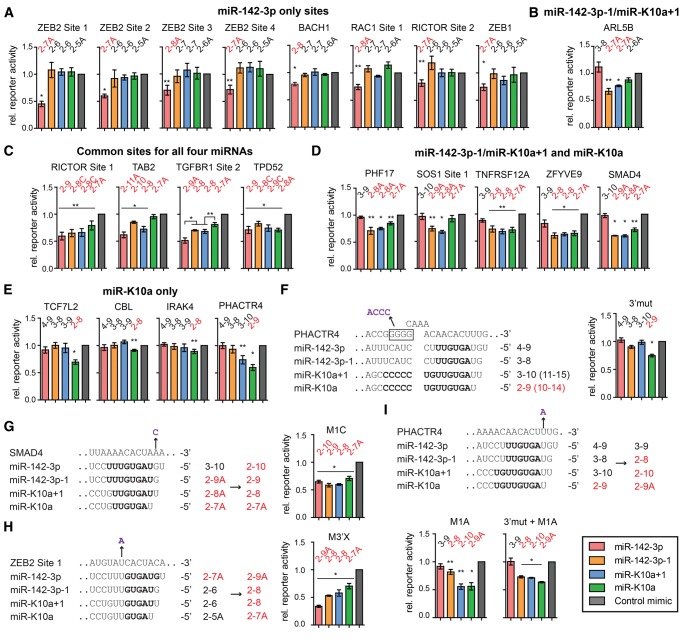
FIGURE 4.
Differential target repression by the miR-142-3p and miR-K10a 5′-isomiRs. (A–E) Dual luciferase 3′-UTR reporter assays performed in 293T cells. Data from single copy, full length, or substantial length 3′-UTR reporters were normalized to values from an internal Renilla luciferase control, control mimic transfection, and matched seed mutant constructs. This assay therefore isolates the binding site under investigation. We tested representative candidate binding sites of (A) only miR-142-3p; (B) only miR-142-3p−1/miR-K10a+1; (C) all four miRNAs; (D) miR-142-3p−1/miR-K10a+1 and miR-K10a; and (E) only miR-K10a. (F) Predicted supplementary pairing of miR-K10a/+1 to PHACTR4 is shown in the left panel. Also shown is a 4-nt mutation that disrupts this supplementary interaction, as shown on the right. (G–I) One-nucleotide gain of function mutations of the wt reporters alter isomiR-specific regulation of the indicated binding sites. Gain of function mutants were normalized to seed mutants. Nucleotides in bold denote residues expected to pair to the binding site. Mutations are in purple and indicated by arrows. Predicted base-paired nts are indicated above columns, canonical seed match types predicted to be functional based on previous reports are in red. (*) P < 0.01; (**) P < 0.05, by Student's t-test. Error bars, SEM (n ≥ 3 biological replicates). Additional results from less functional sites are shown in Supplemental Figure S4. Characteristics of each site and primers used for cloning are listed in Supplemental Table S5.
FIGURE 5.
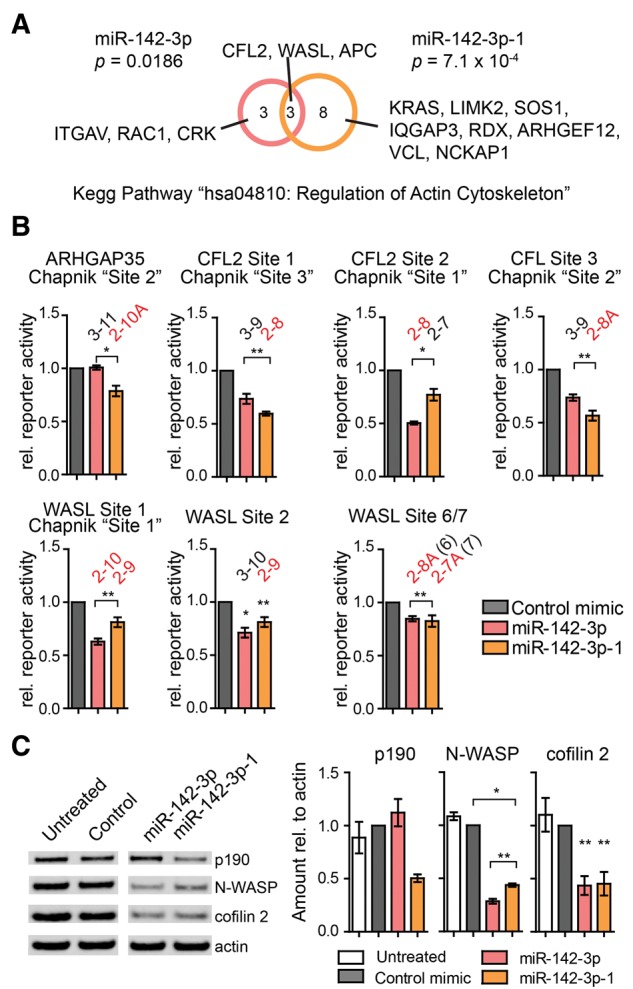
Both miR-142-3p 5′-isomiRs repress regulators of the actin cytoskeleton. (A) Pathway analysis using DAVID reveals that high confidence target mRNAs of miR-142-3p and/or miR-142-3p-1 identified by both Ago2-PAR-CLIP and predicted by TargetScan v5.2 are significantly enriched for genes in the KEGG pathway hsa04810 (regulation of actin cytoskeleton). This analysis is at the mRNA level and several mRNAs have more than one binding site for one or both miR-142-3p 5′-isomiRs (see Supplemental Table S6). (B,C) Differential regulation of p190 (ARHGAP35), N-WASP (WASL), and cofilin 2 (CFL2) by the miR-142-3p 5′-isomiRs in (B) 3′-UTR reporter assays performed as described in Figure 4 and (C) quantitative Western blot analysis for endogenous proteins in iHMVECs transfected with miRNA mimics. One representative blot is shown on the left. All lanes were run together on the same gel and membrane and irrelevant lanes were cropped from the picture. Quantitative analysis of three independent Western blots is shown on the right. (*) P < 0.05; (**) P < 0.01, by Student's t-test. Error bars, SEM (n ≥ 3 biological replicates).
Divergent reporter regulation by the miR-142-3p 5′-isomiRs and selective viral mimicry
We next tested differential target recognition by the miR-142-3p and miR-K10a 5′-isomiRs directly using 3′-UTR reporter assays. Reporters contained either full length 3′-UTRs or sites in their authentic ≥400-nt sequence context. Data from wild-type (wt) reporters were normalized to those from constructs with mutated miRNA-binding sites and therefore represent the activity of the specific binding sites. We selected several sites from each of the five subsets (Fig. 2A,B) to represent the entire range of seed match types observed. Sites were not only predominantly chosen from PAR-CLIP-identified sites, but also included several candidate sites of miR-142-3p/−1 predicted only by TargetScan (Lewis et al. 2005) and previously reported miR-142-3p binding sites (Supplemental Table S5; Wu et al. 2011; Kwanhian et al. 2012). We furthermore focused this analysis on mRNAs with known functions and likely significance to biological roles of miR-142-3p/−1 and/or miR-K10a/+1. Properties of the tested sites are reported in Supplemental Table S5 and resulting data are reported in Figure 4 and Supplemental Figure S4, regardless of regulatory outcome. Resulting data generally confirmed the hypotheses and binding site subsets outlined above. We successfully validated sites that are exclusively regulated by each 5′-isomiR (Fig. 4A,B,E). Regulation observed in the “miR-142-3p−1/miR-K10a+1-only” group of reporters was weak or did not reach statistical significance (Fig. 4B; Supplemental Fig. S4b; Supplemental Table S5). This is most likely because all sites in this group are 2–7A sites, which are often relatively weak sites (Grimson et al. 2007; Nielsen et al. 2007). In addition, we validated sites that were shared between all four miRNAs or between the two miR-K10a 5′-isomiRs and miR-142-3p−1, but not miR-142-3p (Fig. 4C,D).
At least for some sites, including TGFBR1 Site 2, we were able to resolve the expected differential regulatory potencies of different types of seed matches (Fig. 4C; Bartel 2009). In this example, a 2–7A site (miR-K10a) resulted in weak (∼20%) reporter inhibition. Inhibition increased to ∼30% and ∼50% for 2–8 (miR-K10a+1 and miR-142-3p−1) and 2–9A (miR-142-3p), respectively. This is consistent with previous studies that established the hierarchy of miRNA seed match efficacy: 2–8A > 2–8 > 2–7A (Grimson et al. 2007; Nielsen et al. 2007). Interestingly, the absolute efficacy of each type of site showed strong variation between different sites. For example, some 2–7A matches in the 3′-UTR of ZEB2 resulted in surprisingly robust regulation by miR-142-3p, while this type of site had little regulatory potency in several other reporters (Fig. 4; Supplemental Fig. S4; Supplemental Table S5). This suggests, in agreement with published reports (Grimson et al. 2007), that the sequence context of the seed match affects the efficacy of repression.
We noticed that the PHACTR4 reporter was regulated by miR-K10a+1 despite having an offset 3–10 seed match (Fig. 4E). Further examination of the nts surrounding the binding site suggested that miR-K10a+1 might engage in 3′ supplementary pairing (Fig. 4F). Indeed, mutation of the complementary sequence in the PHACTR4 3′-UTR eliminated regulation by miR-K10+1, suggesting that this site is a functional 3′ supplementary site for miR-K10a+1 (Fig. 4F). The identification of this site shows that a small subset of sites might be differentially regulated between miR-K10a+1 and miR-142-3p−1.
Taken together, these results provide direct validation of our finding that many binding sites of the two miR-142-3p 5′-isomiRs are distinct. miR-K10a+1 functions as a straightforward mimic of miR-142-3p−1 and miR-K10a shares most of its binding sites with miR-142-3p−1/miR-K10a+1. Our data furthermore confirm that miR-K10a/+1 has a small set of unique binding sites and therefore extends the regulatory capacity of miR-K10a/+1 beyond that of miR-142-3p−1.
Gain of function mutations alter 5′-isomiR target specificity
To further confirm that target regulation by the miR-142-3p and miR-K10a isomiRs is governed by their seed sequences, we introduced 1-nt mutations into ZEB2 Site 1, SMAD4, and PHACTR4 to change the regulatory potential of these sites (Fig. 4F–I). Resulting data confirm that the target specificity can be redirected as predicted by the seed rules. For PHACTR4, these data also clearly resolve the regulatory contribution of 3′ supplementary pairing by miR-K10a+1 compared to miR-142-3p−1 (Fig. 4I). Together, data from the gain of function mutants further confirm that the seed sequence is the main determinant of 5′-isomiRs target recognition.
Both miR-142-3p 5′-isomiRs repress actin regulators using distinct binding sites
Our finding that the two miR-142-3p miRNAs regulate mostly disparate sites immediately raises the question of the functional impact of miR-142-3p 5′-isomiR expression. To address this question, we performed pathway analyses of miR-142-3p and miR-142-3p−1 targets using DAVID (Huang da et al. 2009). We only considered high confidence targets identified by both PAR-CLIP and TargetScan. The top enriched pathway for each isomiR was “Regulation of actin cytoskeleton” (Fig. 5A). We considered this pathway in detail, because the thrombocytopenia of miR-142 knockout mice has recently been linked to defects in actin dynamics in megakaryocytes (Chapnik et al. 2014). While Chapnik et al. (2014) had detected a significant up-regulation of TargetScan-predicted miR-142-3p targets that are regulators of the actin cytoskeleton, these authors had not considered targets of miR-142-3p−1. The enrichments we observed for high confidence targets of each of the miR-142-3p 5′-isomiRs were due to both shared and unique targets (Fig. 5A). Thus, it appears that the two miR-142-3p 5′-isomiRs may function nonredundantly in the regulation of the actin cytoskeleton.
This is nicely illustrated by considering the three targets that were functionally linked to actin defects in megakaryocytes in miR-142 knockout mice, i.e., p190RhoGap (ARHGAP35, formerly Grlf1 in mice), cofilin 2 (CFL2), and neuronal Wiskott-Aldrich Syndrome protein (N-WASP or WASL). We further investigated the human counterparts of these three target mRNAs by reporter assays and Western blotting analyses (Fig. 5B,C; Supplemental Table S6). For Western analysis, we chose immortalized human microvascular endothelial cells (iHMVECs, Shao and Guo 2004), because these cells do not express endogenous miR-142-3p and are considered close to the physiological target cell type of KSHV. p190/ARHGAP35 was missed by the analysis in Figure 5A, because the gene symbol changed since the release of TargetScan v5.2. Of the three previously identified miR-142-3p binding sites in the murine Arhgap35 3′-UTR, only one is conserved in humans. While this site matches the seeds of either miR-142-3p or miR-142-3p−1 in mice, the human site has retained only its capacity for regulation by miR-142-3p−1 (Fig. 5B,C). Accordingly, p190 protein expression is repressed only by miR-142-3p−1 (Fig. 5C). Thus, p190/Arhgap35 is potentially regulated by both 5′-isomiRs in mice, but is regulated exclusively by miR-142-3p−1 in humans.
All three previously identified candidate miR-142-3p binding sites in the murine 3′-UTR of CFL2 are conserved in humans. Site 1 is preferentially regulated by miR-142-3p−1 and site 2 mostly by miR-142-3p (Fig. 5B). Both sites are expected to have the same specificities in mice and humans. Site 3 matches the seeds of either miR-142-3p or miR-142-3p−1 in mice, while human site 3 is a canonical target of only miR-142-3p−1. We do indeed detect preferential regulation of this site by miR-142-3p−1, although we also detect significant regulation of the 3–9 offset seed match for miR-142-3p. Both miRNAs were equally efficient repressors of cofilin 2 protein expression (Fig. 5C).
Finally, we considered N-WASP/WASL. The human WASL 3′-UTR has seven seed matches to one or both of the miR-142-3p isomiRs (Supplemental Table S6), with sites 6 and 7 too closely spaced to be occupied at the same time. Site 1 is a canonical site for both isomiRs. Sites 3–6 are canonical sites for miR-142-3p, but not miR-142-3p−1. Sites 2 and 7 are canonical sites for miR-142-3p−1, but not miR-142-3p. To examine the differential regulation by the miR-142-3p isomiRs, we conducted 3′-UTR reporter assays for sites that are predicted to be bound by miR-142-3p−1 (Sites 1, 2, and 6/7). We confirmed Site 1 as a target of both isomiRs (Fig. 5B). Site 2 was surprisingly well regulated by the offset miR-142-3p seed match, while the overlapping sites 6/7 were regulated by either isomiR as predicted (Fig. 5B). N-WASP expression was significantly more repressed by miR-142-3p than by miR-142-3p−1, presumably due to the presence of more binding sites (five for miR-142-3p versus three for miR-142-3p−1). Together, these data further suggest that the miR-142-3p isomiRs mostly target discrete sites but ultimately both modulate multiple regulators of the actin cytoskeleton. We speculate that coexpression of both isomiRs may be a requirement to access and efficiently regulate a large set of mRNAs encoding regulators of the actin cytoskeleton. It will be interesting for future studies to separate effects of the two miR-142-3p 5′-isomiRs in phenotypic experiments and in miR-142 knockout models.
5′-isomiRs have either “convergent” or “divergent” target ranges
We noted above that the miR-142-3p 5′-isomiRs regulate largely discrete sets of sites, while the miR-K10a 5′-isomiRs access mostly the same sites. We furthermore noted that the convergence of the miR-K10a 5′-isomiR target ranges is due to the presence of a U at the second position of the longer isomiR. Given these opposite trends, we hypothesize that other 5′-isomiRs that are offset from each other by 1 nt could similarly fall into these two classes. “Convergent” 5′-isomiRs have mostly identical binding sites like miR-K10a/+1. On the other hand, “divergent” 5-isomiRs have mostly discrete target ranges like miR-142-3p/−1. Like for the miR-K10a and miR-142-3p isomiRs, we propose that this generalized 5′-isomiR relationship is driven by nt 2 of the longer variant (Fig. 6A). In the divergent class, nt 2 is A/C/G (V). Extensive seed pairing (≥2–9) of the longer variant is thus required for the binding site to be shared with the shorter 5′-isomiR. This imposes a more stringent requirement for a site to be bound by both 5′-isomiRs. Hence, isomiRs in this class will generally bind to different sites. In contrast, the longer variant of convergent 5′-isomiRs has a U at nt 2. As in miR-K10a/+1, this U allows 2–8 matches of the longer 5′-isomiR to be 2–7A sites for the shorter variant, resulting in a greater overlap between their binding sites.
FIGURE 6.
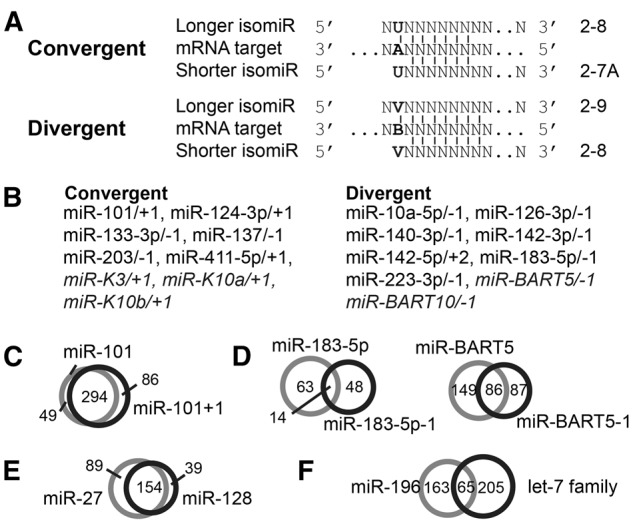
We hypothesize that 5′-isomiRs and unrelated miRNAs with offset seed sequences have convergent or divergent target ranges. (A) Schematic of a shared binding site for proposed convergent and divergent 5′-isomiRs. Bold letters highlight the importance of nt 2 of the longer 5′-variant. V: A, C, or G; B: C, G, or U. (B) Well-documented (Chiang et al. 2010; Cloonan et al. 2011; Gottwein et al. 2011; Xia and Zhang 2014) 5′-isomiRs that are conserved in at least humans and mice or of herpesviral origin (italics) were grouped into convergent and divergent 5′-isomiRs, based on the identity of nt 2 of the longer 5′ sequence. Offsets were designated relative to the isoform annotated in miRBase v21. (C,D) Comparison of Ago2-PAR-CLIP-identified miRNA-binding sites of (C) convergent or (D) divergent 5′-isomiRs. (E,F) Comparison of Ago2-PAR-CLIP-identified miRNA-binding sites of unrelated miRNAs that have offset seed sequence and behave like (E) convergent or (F) divergent 5′-isomiRs.
We compiled and classified a list of 5′-isomiRs that are conserved in at least humans and mouse from comprehensive miRNA sequencing studies (Fig. 6B; Chiang et al. 2010; Cloonan et al. 2011; Gottwein et al. 2011; Xia and Zhang 2014). We also included KSHV- and EBV-encoded isomiRs (Gottwein et al. 2011). While the relationship of these abundant 5′-isomiRs will have to be validated in future experiments, at the level of PAR-CLIP-identified binding sites, the convergent miR-101/-1 overwhelmingly share their assigned binding sites (69%) (Fig. 6C). This is consistent with the previous observation that the miR-101 5′-isomiRs have mostly similar targets (Cloonan et al. 2011). Similarly, our earlier analysis of KSHV miR-K3/+1 has already demonstrated a remarkably strong target overlap between these two isomiRs (∼70%, Manzano et al. 2013). In contrast, the divergent isomiRs miR-183/-1 and miR-BART10/-1 have only 11% and 27% shared PAR-CLIP binding sites, respectively (Fig. 6D). Taken together, these data support our hypothesis that 5′-offsets by 1 nt result in isomiRs with a majority of discrete or identical binding sites. These relationships are governed by the identity of the second nt of the variant with the longer 5′-end.
Unrelated miRNAs with offset seeds might act like 5′-isomiRs
Our results predict that unrelated miRNAs with offset seed sequences may act like 5′-isomiRs. Our analysis revealed only a handful of examples of unrelated cellular miRNAs that could potentially have similar functions due to overlapping target ranges (Supplemental Table S7). In addition, miRNAs expressed by different vertebrate herpesviruses also have seed regions offset by 1 nt from cellular or other viral miRNAs (Supplemental Table S7). For an initial confirmation of our hypothesis, we specifically examined the PAR-CLIP-identified binding sites of the miRNA pairs miR-27/miR-128 and miR-196/let-7 family miRNAs and found that these miRNAs are likely to have convergent and divergent target ranges, as predicted (Fig. 6E,F). The convergent-like miR-27/miR-128 share 55% of their assigned PAR-CLIP target sites while the divergent-like miR-196/let-7 have 15% of their sites in common. In sum, it appears likely that unrelated cellular and viral miRNAs have the potential to act as partial functional mimics of each other and have 5′-isomiR-like properties with convergent or divergent target ranges. Future studies will have to validate these predictions experimentally.
DISCUSSION
Here we studied the consequences of 5′-isomiR expression on miRNA target repertoires, using the specific case of the vertebrate miR-142-3p and KSHV miR-K10a 5′-isomiRs as examples. Our data show that 5′-isomiRs that are offset by 1 nt from each other can have highly divergent or convergent target ranges. The main sequence determinant of 5′-isomiR behavior is the second nt of the longer variant. Specifically, a U at position 2 of the longer 5′-isomiR results in shared sites with base-pairing of ≥2–8 of the longer variant and with an A across from nt 1 of the shorter variant. Such “-1A” sites can facilitate target regulation, even when nt 8 of the miRNA is not base paired (Lewis et al. 2005).
Our findings strongly support the idea that 5′-isomiR expression broadens the binding site repertoire of a miRNA, for both classes of 5′-isomiRs. The functional consequences of 5′-isomiR expression will presumably differ for each miRNA. In the case of the highly divergent miR-142-3p 5′-isomiRs, both miRNAs target mRNAs encoding factors involved in actin dynamics through exclusive and shared sites. These 5′-isomiRs are therefore likely to function additively or cooperatively in this pathway. While this remains to be tested, it is conceivable that there are 5′-isomiR-specific functions. Even for shared targets, functional difference between 5′-isomiRs could result from differential efficacies of different types of seed matches. In addition, competition of two well-expressed 5′-isomiRs for their shared sites could result in an intermediate regulatory output compared to the expression of either 5′-isomiR alone. The relationships of 5′-isomiRs uncovered here are likely to also apply to unrelated miRNAs with offset seed sequences as well as miRNA paralogs that have undergone seed shifting during evolution (Wheeler et al. 2009).
While abundantly expressed 5′-isomiRs are generated by only a minority of animal miRNAs (Fig. 6B), our findings have important implications for functional studies of these miRNAs. For instance, previous functional studies of miR-142-3p have considered targets of only the miR-142-3p 5′-isomiR, which is specifically annotated in miRBase. Thus, key target mRNAs of miR-142-3p−1 may have been missed. For example, high confidence targets of miR-142-3p−1 included several additional regulators of the actin cytoskeleton, suggesting that both 5′-isomiRs function nonredundantly in the regulation of actin dynamics. Similarly, a contribution of miR-142-3p−1 to additional already identified functions of miR-142-3p appears likely. In recent work done in Xenopus, knockdown of miR-142-3p by morpholinos (which is expected to also affect miR-142-3p−1) blocks the specification of definitive hemangioblasts, the pluripotent progenitors of endothelial cells and the hematopoietic cell lineage (Nimmo et al. 2013). This phenotype was attributed to increased TGF-β signaling in miR-142-3p morphants and a small molecular inhibitor of TGF-β signaling fully restored the hemangioblast population. Accordingly, the authors showed that miR-142-3p has several potential direct targets in this signaling cascade, including the TGF-β receptor 1 (encoded by TGFBR1). A similar result was obtained following inhibition of DGCR8, but microinjection of a mimic of the miR-142-3p 5′-isomiR in these morphants was able to only partially rescue hemangioblast specification. It would be interesting to see if coexpression of miR-142-3p−1 would allow a full restoration of the DGCR8 morphants. As in the case of regulation of the actin cytoskeleton, both miR-142-3p and miR-142-3p−1 target mRNAs with roles in TGF-β signaling. This is suggested by our validation of key components of the TGF-β cascade as targets of miR-142-3p−1, but not miR-142-3p. For example, ZFYVE9 is a target of miR-142-3p−1 that is conserved in Xenopus and encodes a protein important for the recruitment of Smad2 to the TGF-β receptor complex (Tsukazaki et al. 1998). Similarly, SMAD4 is a target of miR-142-3p−1, but not miR-142-3p, through a binding site conserved in mouse but not in Xenopus. A potential function of miR-142-3p−1 in the inhibition of TGF-β is furthermore suggested by the finding that the miR-K10a miRNAs, which mimic mostly miR-142-3p−1, inhibit TGF-β in the context of KSHV infection (Lei et al. 2012). Similar to the Xenopus study, miR-142 knockout mice lack both miR-142-3p isomiRs and it will be interesting to see how the two 5′-isomiRs control miR-142-3p functions in vivo.
Other cellular miRNAs with conserved coexpression of abundant divergent 5′-isomiRs, including miR-10a, miR-126, and miR-140, have likewise been shown to carry out important biological functions. Expression of miR-10a in porcine arterial endothelial cells promotes an anti-inflammatory state that prevents atherosclerotic plaque formation through modulation of the NF-κB pathway (Fang et al. 2010). Knockdown and knockout studies in zebrafish and mouse, respectively, have demonstrated that the endothelial cell-specific miR-126 is critical for vascular integrity and angiogenesis (Fish et al. 2008; Wang et al. 2008). These studies have identified sprouty-related, EVH1 domain-containing 1 (SPRED1), a negative regulator of vascular endothelial growth factor (VEGF) signaling, as a direct target of miR-126. Finally, miR-140 has been linked to cartilage formation and homeostasis during mouse development (Miyaki et al. 2010). Given that these miRNAs are coexpressed with divergent 5′-isomiRs, future phenotypic work should consider the contributions of these 5′-variants and their specific targets.
Expression of 5′-isomiR will also have to be considered in studies of the targets and function of KSHV miR-K10a+1. A pivotal importance for the miR-K10a 5′-isomiRs is suggested by the fact that a number of Old World primate rhadinoviruses carry miRNAs with a seed sequence identical to either miR-K10a or miR-K10a+1. This finding suggests that these viruses as well, selectively mimic the miR-142-3p−1 5′-isomiR. While miRc-RF9-3p shares its extended seed sequence with miR-142-3p−1 (nts 2–11), this extended sequence similarity does not increase the expected functional overlap with the miR-142-3p 5′-isomiR. This prediction will have to be confirmed, however, given that we did detect some regulatory potential for extended (≥3–9) offset seed matches to miR-142-3p. These miR-142-3p−1-like viral miRNAs could, in principle, function in the lymphoid target cells of rhadinoviruses, which express abundant miR-142-3p/−1, to repress non-miR-142-3p−1-targets, skew the regulation of miR-142-3p−1 targets or overexpress miR-142-3p−1-like functions. Alternatively, these viral mimics of miR-142-3p−1 could introduce miR-142-3p−1-like functions into cell types where miR-142-3p/−1 are not expressed, including the differentiated endothelium, the cell type underlying the pathogenesis of Kaposi's sarcoma. It will be interesting to study the functional relevance of miR-K10a/+1 in the context of KSHV-infected endothelial cells. Identifying roles of miR-K10a/+1 in KSHV infection should also shed light on why rhadinoviruses selectively mimic miR-142-3p−1. For example, a previous report has shown that miR-K10a inhibits the tumor necrosis factor-like weak inducer of apoptosis receptor (TWEAKR/TNFRSF12A), which impacts cytokine signaling in KSHV-infected endothelial cells (Abend et al. 2010). It was suggested that the down-regulation of TNFRSF12A protects infected cells from apoptosis-induced cell death and suppress the expression of pro-inflammatory cytokines. This function is likely due to both 5′-isomiRs, which we have shown are equally potent repressors of TNFRSF12A reporters (Fig. 4D). Interestingly, TNFRSF12A reporters were also repressed by miR-142-3p−1, but not miR-142-3p, suggesting that this and other 5′-isomiR-specific targets of miR-142-3p−1 might underlie the selective pressure to selectively mimic miR-142-3p−1. It has previously been suggested that direct repression of TGF-β receptor 2 expression by both 5′-isomiRs underlies the inhibition of this TGF-β signaling by miR-K10a/+1 (Lei et al. 2012). The TGFBR2 3′-UTR, however, lacks canonical seed matches to either the miR-K10a or miR-142-3p 5′-isomiRs. Finally, it will be interesting to see if the miR-K10a isomiRs, like the miR-142-3p miRNAs, are regulators of the actin cytoskeleton, which is known to be deregulated during infection by KSHV and many other viruses (Taylor et al. 2011).
Several important questions remain for future studies. These include whether 5′-isomiR expression is regulated in specific tissues, at different developmental stages, or in disease. Instances of spatial and temporal variation in the relative expression of isomiRs (isomiR switching) have been described (Fukunaga et al. 2012; Tan et al. 2014). For instance, the relative expression of the convergent miR-101 and miR-101+1 isomiRs changes from 1:2 in the brain to 3:1 in fetal lung fibroblasts (Tan et al. 2014). Similarly, the expression of the divergent 5′-isomiRs miR-140-3p and miR-140-3p-1 change from ∼6:1 in the nasal epithelium and mucosa to 1:1.6 in the heart. The dsRNA binding domain-containing partner proteins of Drosha and Dicer are important determinants of cleavage site selection by these enzymes (Han et al. 2006; Fukunaga et al. 2012; Lee and Doudna 2012), and are candidates for players in regulated isomiR expression. In Drosophila, the level of the Dicer binding partner loqs-PA relative to its isoform loqs-PB increases during development (Fukunaga et al. 2012). These two isoforms bind Dicer separately and alter the processing of a specific subset of miRNAs, including miR-307a.
Finally, our work highlights the importance of the careful annotation of miRNA ends. To date no unified nomenclature for miRNA sequence variation exists, although several have been proposed (Ruby et al. 2006; Cloonan et al. 2011). The annotation, detection, and validation of 5′-offsets are critically important for functional work as 5′-offsets are clearly consequential in terms of target recognition. In addition, the annotation of 3′-offsets is important, because these affect the accurate measurement of miRNA abundance using hybridization-based assays.
MATERIALS AND METHODS
Cloning of 3′-UTR reporters and miRNA expression vectors
Candidate miRNA-binding sites were identified from previously published Ago2-PAR-CLIP data sets from primary effusion lymphoma (PEL) B cell lines (Gottwein et al. 2011), or predicted by TargetScan 6.2 (for miR-142-3p) and/or custom TargetScan 5.2 (for miR-K10a and miR-142-3p−1/miR-K10a+1). Full length or substantial length 3′-UTR sequences were PCR-amplified from BC-1 genomic DNA and cloned into the reporter vector pLSG (Gottwein et al. 2007) expressing firefly luciferase using XhoI and NotI sites. Predicted seed sites in each 3′-UTR were mutated from 5′-…AACACU…-3 to 5′-…AAGACA…-3′ using mutated primers to abolish binding of the four miRNAs. For reporters with noncanonical sites, primers were designed to contain two predicted binding sites separated by a 10 bp linker (Fig. 3). These binding sites were modeled from previously reported noncanonical interactions (Grimson et al. 2007; Shin et al. 2010; Chi et al. 2012). Primers that were used for cloning or mutagenesis are detailed in Supplemental Tables S4, S5; primer sequences are listed in Supplemental Table S8. Reporter assays were conducted as described using mirVana miRNA mimics (Life Technologies) (Manzano et al. 2013). To express miRc-RF9-3p, a 250-bp fragment centered on the predicted pre-miRc-RF9 was synthesized as a gBlock (IDT), PCR-amplified using primers 2079 and 2080, and inserted between XhoI and NotI sites of the published lentiviral vector pLCE (Gottwein et al. 2011).
RNA preparation for primer extension
Frog spleens were harvested from 1- to 2-yr-old male X. laevis. Chicken spleens were dissected from E13 to E16 embryos. Mouse spleen was from a 7–8 wk female A/J mouse. For Ago2-IP experiments, ∼5 × 107 cells of PEL-derived cell lines VG-1 or BC-3 were lysed and incubated with anti-Ago2- or control IgG1-bound protein G beads as previously described (Gottwein et al. 2011). For the RFHV primer extension, LLC-MK2 cells were transfected with the cloned pri-miRc-RF9-3p. RNAs were prepared using TRIzol (Life Technologies), with the ethanol wash omitted.
Primer extension
Primer extension analyses were conducted as described using denaturing polyacrylamide gel-purified probes (Manzano et al. 2013). Primers for miR-K10a, miR-142-3p, and 5S RNA have been previously published (Gottwein et al. 2011). Primers used for hsa-miR-16 (1580), xtr-miR-16 (2078), and miRc-RF9-3p (2083) are found in Supplemental Table S8.
Western blotting
iHMVEC cells (7 × 105 cells/10 cm2 dish) were transfected with 100 pmol of control 1, miR-142-3p and miR-142-3p−1 mirVana miRNA mimics using Lipofectamine RNAiMAX (Life Technologies). Two days after transfection, cells were harvested in 20 mM Tris pH 7.5, 100 mM NaCl, 1% Triton X-100, 10% glycerol, 1 mM EDTA, and Complete EDTA-free protease inhibitors (Roche). Quantitative Western blotting using the Odyssey Fc Imaging System (LI-COR Biosciences) was performed as described (Manzano et al. 2013) using anti-p190 (BD Biosciences, Cat. 610149), anti-cofilin 2 (EMD Millipore, Cat. 07-300), anti-N-WASP (30D10, Cell Signaling Technology), and anti-actin (C-2, Santa Cruz Biotechnology) antibodies.
Gene expression profiling
HEK293T cells (5 × 105) in six-well plates were transfected with 50 pmol of mirVana miRNA mimics (negative control 2, miR-142-3p, miR-142-3p−1, miR-K10a+1, or miR-K10a) using the RNAiMAX transfection reagent in triplicate wells. The medium was replaced after 8 h. Two days post-transfection, total RNA was harvested with 0.5 mL TRIzol reagent and further purified using RNeasy MinElute Cleanup Kit (Qiagen). Total RNA was eluted in 15 µL water from the spin columns. Samples were processed by the Northwestern University Genomics Core Facility for Illumina Gene Expression Array using the HumanHT-12 v4 Expression BeadChip (Illumina).
Analysis of microarray data
Quality checks and probe level processing of the Illumina microarray data were made with the R Bioconductor package lumi (Du et al. 2008). Data processing included a normalization method to reduce the obscuring variation between microarrays (Bolstad et al. 2003), which might be introduced during the processes of sample preparation, manufacturing, fluorescence labeling, hybridization, and/or scanning. Probes that were expressed in none of the samples, as judged by an Illumina detection P > 0.01, were filtered out. Probes that lacked annotated gene symbol, description and Entrez Gene ID were also excluded from further analyses. A total of 17,242 probes remained after the filtering processes and were used for further analyses. Hierarchical clustering was performed on the filtered data with the plotSampleRelation function of the lumi package to assess sample relationships. Principal component analysis was performed on the scaled version of the filtered data to evaluate sample variation. Differential gene expression between the different conditions was assessed by statistical linear model analysis using the Bioconductor package limma (Smyth 2004; Wettenhall and Smyth 2004), in which an empirical Bayes method is used to moderate the standard errors of the estimated log-fold changes of gene expression. The moderated T statistic P-value derived from the limma analysis above was further adjusted for multiple testing by Benjamini and Hochberg's method to control false discovery rate (FDR). The list of differentially expressed genes was obtained by the FDR criterion of <5%.
Sylamer analysis
Gene lists from published GEO data set GSE52141 and newly generated data set GSE66489 were ranked by fold change. 3′-UTR sequences were obtained from the UCSC Genome database. For genes with multiple transcripts and 3′-UTR sequences, the longest sequence was retained. Lists containing gene names and log-fold expression values were prepared along with their longest corresponding 3′-UTR sequences. The relative enrichment of sequences in the 3′-UTR was identified using Sylamer (van Dongen et al. 2008) version 08-123 and plotted in R.
DATA DEPOSITION
Microarray data were submitted to the Gene Expression Omnibus under accession number GSE66489.
SUPPLEMENTAL MATERIAL
Supplemental material is available for this article.
Supplementary Material
ACKNOWLEDGMENTS
We thank Dr. Nikolaus Rajewsky and Andrei Filipchyck (Max Delbruck Center for Molecular Medicine, Berlin) for helpful discussions and sharing unpublished data, Dr. Phillip Zamore (University of Massachusetts, Worcester) for helpful suggestions, Matt Bonakdarpour (University of Chicago) for analysis of miRNA homologies, Dr. Brian Mitchell and Dr. Philip Spear (NU) for frog spleens, Dr. Gregory Smith and Gina Daniels (NU) for chick embryos, Dr. Nicholas Cianciotto and Richard C. White (NU) for mouse spleen, Dr. Radhakrishnan Padmanabhan (Georgetown University) for LLC-MK2 cells, Nakisha Rutledge (NU) for contributions during the initial stages of the project, and Kylee Morrison for critical reading of the manuscript. We also thank Drs. Nadereh Jafari and Chunfa Jie of the Northwestern University Genomics Core Facility for microarray data generation and processing. This work was supported by the National Institutes of Health (R01 CA180813 to E.G.), the Zell Family Foundation, and the Robert H. Lurie Comprehensive Cancer Center. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health, the Zell Family Foundation, or the Robert H. Lurie Comprehensive Cancer Center.
Footnotes
Article published online ahead of print. Article and publication date are at http://www.rnajournal.org/cgi/doi/10.1261/rna.048876.114.
REFERENCES
- Abend JR, Uldrick T, Ziegelbauer JM. 2010. Regulation of tumor necrosis factor-like weak inducer of apoptosis receptor protein (TWEAKR) expression by Kaposi's sarcoma-associated herpesvirus microRNA prevents TWEAK-induced apoptosis and inflammatory cytokine expression. J Virol 84: 12139–12151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Azuma-Mukai A, Oguri H, Mituyama T, Qian ZR, Asai K, Siomi H, Siomi MC. 2008. Characterization of endogenous human Argonautes and their miRNA partners in RNA silencing. Proc Natl Acad Sci 105: 7964–7969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baek D, Villén J, Shin C, Camargo FD, Gygi SP, Bartel DP. 2008. The impact of microRNAs on protein output. Nature 455: 64–71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartel DP. 2009. MicroRNAs: target recognition and regulatory functions. Cell 136: 215–233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berezikov E, Robine N, Samsonova A, Westholm JO, Naqvi A, Hung JH, Okamura K, Dai Q, Bortolamiol-Becet D, Martin R, et al. 2011. Deep annotation of Drosophila melanogaster microRNAs yields insights into their processing, modification, and emergence. Genome Res 21: 203–215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolstad BM, Irizarry RA, Astrand M, Speed TP. 2003. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19: 185–193. [DOI] [PubMed] [Google Scholar]
- Bruce AG, Ryan JT, Thomas MJ, Peng X, Grundhoff A, Tsai CC, Rose TM. 2013. Next-generation sequence analysis of the genome of RFHVMn, the macaque homolog of Kaposi's sarcoma (KS)-associated herpesvirus, from a KS-like tumor of a pig-tailed macaque. J Virol 87: 13676–13693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai X, Lu S, Zhang Z, Gonzalez CM, Damania B, Cullen BR. 2005. Kaposi's sarcoma-associated herpesvirus expresses an array of viral microRNAs in latently infected cells. Proc Natl Acad Sci 102: 5570–5575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai X, Schafer A, Lu S, Bilello JP, Desrosiers RC, Edwards R, Raab-Traub N, Cullen BR. 2006. Epstein-Barr virus microRNAs are evolutionarily conserved and differentially expressed. PLoS Pathog 2: e23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chapnik E, Rivkin N, Mildner A, Beck G, Pasvolsky R, Metzl-Raz E, Birger Y, Amir G, Tirosh I, Porat Z, et al. 2014. miR-142 orchestrates a network of actin cytoskeleton regulators during megakaryopoiesis. Elife 3: e01964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen CZ, Li L, Lodish HF, Bartel DP. 2004. MicroRNAs modulate hematopoietic lineage differentiation. Science 303: 83–86. [DOI] [PubMed] [Google Scholar]
- Chi SW, Hannon GJ, Darnell RB. 2012. An alternative mode of microRNA target recognition. Nat Struct Mol Biol 19: 321–327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiang HR, Schoenfeld LW, Ruby JG, Auyeung VC, Spies N, Baek D, Johnston WK, Russ C, Luo S, Babiarz JE, et al. 2010. Mammalian microRNAs: experimental evaluation of novel and previously annotated genes. Genes Dev 24: 992–1009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cloonan N, Wani S, Xu Q, Gu J, Lea K, Heater S, Barbacioru C, Steptoe AL, Martin HC, Nourbakhsh E, et al. 2011. MicroRNAs and their isomiRs function cooperatively to target common biological pathways. Genome Biol 12: R126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corcoran DL, Georgiev S, Mukherjee N, Gottwein E, Skalsky RL, Keene JD, Ohler U. 2011. PARalyzer: definition of RNA binding sites from PAR-CLIP short-read sequence data. Genome Biol 12: R79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du P, Kibbe WA, Lin SM. 2008. lumi: a pipeline for processing Illumina microarray. Bioinformatics 24: 1547–1548. [DOI] [PubMed] [Google Scholar]
- Fang Y, Shi C, Manduchi E, Civelek M, Davies PF. 2010. MicroRNA-10a regulation of proinflammatory phenotype in athero-susceptible endothelium in vivo and in vitro. Proc Natl Acad Sci 107: 13450–13455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fish JE, Santoro MM, Morton SU, Yu S, Yeh RF, Wythe JD, Ivey KN, Bruneau BG, Stainier DY, Srivastava D. 2008. miR-126 regulates angiogenic signaling and vascular integrity. Dev Cell 15: 272–284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukunaga R, Han BW, Hung JH, Xu J, Weng Z, Zamore PD. 2012. Dicer partner proteins tune the length of mature miRNAs in flies and mammals. Cell 151: 533–546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gottwein E, Mukherjee N, Sachse C, Frenzel C, Majoros WH, Chi JT, Braich R, Manoharan M, Soutschek J, Ohler U, et al. 2007. A viral microRNA functions as an orthologue of cellular miR-155. Nature 450: 1096–1099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gottwein E, Corcoran DL, Mukherjee N, Skalsky RL, Hafner M, Nusbaum JD, Shamulailatpam P, Love CL, Dave SS, Tuschl T, et al. 2011. Viral microRNA targetome of KSHV-infected primary effusion lymphoma cell lines. Cell Host Microbe 10: 515–526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grimson A, Farh KK, Johnston WK, Garrett-Engele P, Lim LP, Bartel DP. 2007. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell 27: 91–105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grosswendt S, Filipchyk A, Manzano M, Klironomos F, Schilling M, Herzog M, Gottwein E, Rajewsky N. 2014. Unambiguous identification of miRNA:target site interactions by different types of ligation reactions. Mol Cell 54: 1042–1054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grundhoff A, Sullivan CS, Ganem D. 2006. A combined computational and microarray-based approach identifies novel microRNAs encoded by human gamma-herpesviruses. RNA 12: 733–750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ha M, Kim VN. 2014. Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15: 509–524. [DOI] [PubMed] [Google Scholar]
- Ha I, Wightman B, Ruvkun G. 1996. A bulged lin-4/lin-14 RNA duplex is sufficient for Caenorhabditis elegans lin-14 temporal gradient formation. Genes Dev 10: 3041–3050. [DOI] [PubMed] [Google Scholar]
- Hafner M, Landthaler M, Burger L, Khorshid M, Hausser J, Berninger P, Rothballer A, Ascano M Jr, Jungkamp AC, Munschauer M, et al. 2010. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141: 129–141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han J, Lee Y, Yeom KH, Nam JW, Heo I, Rhee JK, Sohn SY, Cho Y, Zhang BT, Kim VN. 2006. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 125: 887–901. [DOI] [PubMed] [Google Scholar]
- Helwak A, Kudla G, Dudnakova T, Tollervey D. 2013. Mapping the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell 153: 654–665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang da W, Sherman BT, Lempicki RA. 2009. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4: 44–57. [DOI] [PubMed] [Google Scholar]
- Humphreys DT, Hynes CJ, Patel HR, Wei GH, Cannon L, Fatkin D, Suter CM, Clancy JL, Preiss T. 2012. Complexity of murine cardiomyocyte miRNA biogenesis, sequence variant expression and function. PLoS One 7: e30933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khorshid M, Hausser J, Zavolan M, van Nimwegen E. 2013. A biophysical miRNA-mRNA interaction model infers canonical and noncanonical targets. Nat Methods 10: 253–255. [DOI] [PubMed] [Google Scholar]
- Kwanhian W, Lenze D, Alles J, Motsch N, Barth S, Döll C, Imig J, Hummel M, Tinguely M, Trivedi P, et al. 2012. MicroRNA-142 is mutated in about 20% of diffuse large B-cell lymphoma. Cancer Med 1: 141–155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lal A, Navarro F, Maher CA, Maliszewski LE, Yan N, O'Day E, Chowdhury D, Dykxhoorn DM, Tsai P, Hofmann O, et al. 2009. miR-24 Inhibits cell proliferation by targeting E2F2, MYC, and other cell-cycle genes via binding to “seedless” 3′UTR microRNA recognition elements. Mol Cell 35: 610–625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Landgraf P, Rusu M, Sheridan R, Sewer A, Iovino N, Aravin A, Pfeffer S, Rice A, Kamphorst AO, Landthaler M, et al. 2007. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129: 1401–1414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee HY, Doudna JA. 2012. TRBP alters human precursor microRNA processing in vitro. RNA 18: 2012–2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee LW, Zhang S, Etheridge A, Ma L, Martin D, Galas D, Wang K. 2010. Complexity of the microRNA repertoire revealed by next-generation sequencing. RNA 16: 2170–2180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lei X, Zhu Y, Jones T, Bai Z, Huang Y, Gao SJ. 2012. A Kaposi's sarcoma-associated herpesvirus microRNA and its variants target TGF-beta pathway to promote cell survival. J Virol 86: 11698–11711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB. 2003. Prediction of mammalian microRNA targets. Cell 115: 787–798. [DOI] [PubMed] [Google Scholar]
- Lewis BP, Burge CB, Bartel DP. 2005. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120: 15–20. [DOI] [PubMed] [Google Scholar]
- Llorens F, Banez-Coronel M, Pantano L, del Rio JA, Ferrer I, Estivill X, Marti E. 2013. A highly expressed miR-101 isomiR is a functional silencing small RNA. BMC Genomics 14: 104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loeb GB, Khan AA, Canner D, Hiatt JB, Shendure J, Darnell RB, Leslie CS, Rudensky AY. 2012. Transcriptome-wide miR-155 binding map reveals widespread noncanonical microRNA targeting. Mol Cell 48: 760–770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loher P, Londin ER, Rigoutsos I. 2014. IsomiR expression profiles in human lymphoblastoid cell lines exhibit population and gender dependencies. Oncotarget 5: 8790–8802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma H, Wu Y, Choi JG, Wu H. 2013. Lower and upper stem-single-stranded RNA junctions together determine the Drosha cleavage site. Proc Natl Acad Sci 110: 20687–20692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Majoros WH, Lekprasert P, Mukherjee N, Skalsky RL, Corcoran DL, Cullen BR, Ohler U. 2013. MicroRNA target site identification by integrating sequence and binding information. Nat Methods 10: 630–633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manzano M, Shamulailatpam P, Raja AN, Gottwein E. 2013. Kaposi's sarcoma-associated herpesvirus encodes a mimic of cellular miR-23. J Virol 87: 11821–11830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mildner A, Chapnik E, Manor O, Yona S, Kim KW, Aychek T, Varol D, Beck G, Itzhaki ZB, Feldmesser E, et al. 2013. Mononuclear phagocyte miRNome analysis identifies miR-142 as critical regulator of murine dendritic cell homeostasis. Blood 121: 1016–1027. [DOI] [PubMed] [Google Scholar]
- Miyaki S, Sato T, Inoue A, Otsuki S, Ito Y, Yokoyama S, Kato Y, Takemoto F, Nakasa T, Yamashita S, et al. 2010. MicroRNA-140 plays dual roles in both cartilage development and homeostasis. Genes Dev 24: 1173–1185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morin RD, O'Connor MD, Griffith M, Kuchenbauer F, Delaney A, Prabhu AL, Zhao Y, McDonald H, Zeng T, Hirst M, et al. 2008. Application of massively parallel sequencing to microRNA profiling and discovery in human embryonic stem cells. Genome Res 18: 610–621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nielsen CB, Shomron N, Sandberg R, Hornstein E, Kitzman J, Burge CB. 2007. Determinants of targeting by endogenous and exogenous microRNAs and siRNAs. RNA 13: 1894–1910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nimmo R, Ciau-Uitz A, Ruiz-Herguido C, Soneji S, Bigas A, Patient R, Enver T. 2013. MiR-142-3p controls the specification of definitive hemangioblasts during ontogeny. Dev Cell 26: 237–249. [DOI] [PubMed] [Google Scholar]
- Nishiyama T, Kaneda R, Ono T, Tohyama S, Hashimoto H, Endo J, Tsuruta H, Yuasa S, Ieda M, Makino S, et al. 2012. miR-142-3p is essential for hematopoiesis and affects cardiac cell fate in zebrafish. Biochem Biophys Res Commun 425: 755–761. [DOI] [PubMed] [Google Scholar]
- Pellett PE, Roizman B. 2013. Herpesviridae. In Fields virology (ed. Knipe DM, Howley PM), pp. 1802–1822. Lippincott Williams & Wilkins, Philadelphia, PA. [Google Scholar]
- Pfeffer S, Sewer A, Lagos-Quintana M, Sheridan R, Sander C, Grässer FA, van Dyk LF, Ho CK, Shuman S, Chien M, et al. 2005. Identification of microRNAs of the herpesvirus family. Nat Methods 2: 269–276. [DOI] [PubMed] [Google Scholar]
- Ruby JG, Jan C, Player C, Axtell MJ, Lee W, Nusbaum C, Ge H, Bartel DP. 2006. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans. Cell 127: 1193–1207. [DOI] [PubMed] [Google Scholar]
- Ruby JG, Stark A, Johnston WK, Kellis M, Bartel DP, Lai EC. 2007. Evolution, biogenesis, expression, and target predictions of a substantially expanded set of Drosophila microRNAs. Genome Res 17: 1850–1864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samols MA, Hu J, Skalsky RL, Renne R. 2005. Cloning and identification of a microRNA cluster within the latency-associated region of Kaposi's sarcoma-associated herpesvirus. J Virol 79: 9301–9305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schirle NT, Sheu-Gruttadauria J, MacRae IJ. 2014. Structural basis for microRNA targeting. Science 346: 608–613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shao R, Guo X. 2004. Human microvascular endothelial cells immortalized with human telomerase catalytic protein: a model for the study of in vitro angiogenesis. Biochem Biophys Res Commun 321: 788–794. [DOI] [PubMed] [Google Scholar]
- Shin C, Nam JW, Farh KK, Chiang HR, Shkumatava A, Bartel DP. 2010. Expanding the microRNA targeting code: functional sites with centered pairing. Mol Cell 38: 789–802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skalsky RL, Samols MA, Plaisance KB, Boss IW, Riva A, Lopez MC, Baker HV, Renne R. 2007. Kaposi's sarcoma-associated herpesvirus encodes an ortholog of miR-155. J Virol 81: 12836–12845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smyth GK. 2004. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3: Article3. [DOI] [PubMed] [Google Scholar]
- Tan GC, Chan E, Molnar A, Sarkar R, Alexieva D, Isa IM, Robinson S, Zhang S, Ellis P, Langford CF, et al. 2014. 5′ isomiR variation is of functional and evolutionary importance. Nucleic Acids Res 42: 9424–9435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor MP, Koyuncu OO, Enquist LW. 2011. Subversion of the actin cytoskeleton during viral infection. Nat Rev Microbiol 9: 427–439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsukazaki T, Chiang TA, Davison AF, Attisano L, Wrana JL. 1998. SARA, a FYVE domain protein that recruits Smad2 to the TGFβ receptor. Cell 95: 779–791. [DOI] [PubMed] [Google Scholar]
- Umbach JL, Cullen BR. 2010. In-depth analysis of Kaposi's sarcoma-associated herpesvirus microRNA expression provides insights into the mammalian microRNA-processing machinery. J Virol 84: 695–703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Umbach JL, Strelow LI, Wong SW, Cullen BR. 2010. Analysis of rhesus rhadinovirus microRNAs expressed in virus-induced tumors from infected rhesus macaques. Virology 405: 592–599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Dongen S, Abreu-Goodger C, Enright AJ. 2008. Detecting microRNA binding and siRNA off-target effects from expression data. Nat Methods 5: 1023–1025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vella MC, Choi EY, Lin SY, Reinert K, Slack FJ. 2004. The C. elegans microRNA let-7 binds to imperfect let-7 complementary sites from the lin-41 3′UTR. Genes Dev 18: 132–137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang S, Aurora AB, Johnson BA, Qi X, McAnally J, Hill JA, Richardson JA, Bassel-Duby R, Olson EN. 2008. The endothelial-specific microRNA miR-126 governs vascular integrity and angiogenesis. Dev Cell 15: 261–271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wettenhall JM, Smyth GK. 2004. limmaGUI: a graphical user interface for linear modeling of microarray data. Bioinformatics 20: 3705–3706. [DOI] [PubMed] [Google Scholar]
- Wheeler BM, Heimberg AM, Moy VN, Sperling EA, Holstein TW, Heber S, Peterson KJ. 2009. The deep evolution of metazoan microRNAs. Evol Dev 11: 50–68. [DOI] [PubMed] [Google Scholar]
- Wu H, Neilson JR, Kumar P, Manocha M, Shankar P, Sharp PA, Manjunath N. 2007. miRNA profiling of naive, effector and memory CD8 T cells. PLoS One 2: e1020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu H, Ye C, Ramirez D, Manjunath N. 2009. Alternative processing of primary microRNA transcripts by Drosha generates 5′ end variation of mature microRNA. PLoS One 4: e7566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu L, Cai C, Wang X, Liu M, Li X, Tang H. 2011. MicroRNA-142-3p, a new regulator of RAC1, suppresses the migration and invasion of hepatocellular carcinoma cells. FEBS Lett 585: 1322–1330. [DOI] [PubMed] [Google Scholar]
- Xia J, Zhang W. 2014. A meta-analysis revealed insights into the sources, conservation and impact of microRNA 5′-isoforms in four model species. Nucleic Acids Res 42: 1427–1441. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.