Abstract
Treating Shiga toxin-producing Escherichia coli (STEC) gastrointestinal infections is difficult. The utility of antibiotics for STEC treatment is controversial, since antibiotic resistance among STEC isolates is widespread and certain antibiotics dramatically increase the expression of Shiga toxins (Stxs), which are some of the most important virulence factors in STEC. Stxs contribute to life-threatening hemolytic uremic syndrome (HUS), which develops in considerable proportions of patients with STEC infections. Understanding the antibiotic resistance profiles of STEC isolates and the Stx induction potential of promising antibiotics is essential for evaluating any antibiotic treatment of STEC. In this study, 42 O157:H7 or non-O157 STEC isolates (including the “big six” serotypes) were evaluated for their resistance against 22 antibiotics by using an antibiotic array. Tigecycline inhibited the growth of all of the tested STEC isolates and also inhibited the production of Stxs (Stx2 in particular). In combination with neutralizing antibodies to Stx1 and Stx2, the tigecycline-antibody treatment fully protected Vero cells from Stx toxicity, even when the STEC bacteria and the Vero cells were cultured together. The combination of an antibiotic such as tigecycline with neutralizing antibodies presents a promising strategy for future STEC treatments.
INTRODUCTION
An estimated one in six residents of the United States contracts some form of food poisoning each year (1). Shiga toxin-producing Escherichia coli (STEC) is responsible for approximately 100,000 cases of illness, 3,000 hospitalizations, and 90 deaths each year in the United States alone (2). Eight percent of patients hospitalized with STEC infections develop the symptoms of hemolytic uremic syndrome (HUS), a life-threatening condition. STEC possesses a number of virulence factors, but Shiga toxins (Stxs) are considered the most critical in disease pathogenesis and are closely associated with HUS (3). Shiga toxin (Stx) was originally discovered in the Shigella genus (4), and similar toxins were later found in STEC strains (5). There are two types of Stxs in E. coli, namely, Stx1, which is nearly identical to the Stx from the Shigella genus, and Stx2, which is considerably different from Stx1 in terms of amino acid sequence, receptor preference, and toxicity (6, 7). Four subtypes of Stx1 (Stx1a, Stx1c, Stx1d, and Stx1e) and seven subtypes of Stx2 (Stx2a through Stx2g) have been identified (8, 9). All Stxs have an AB5 structure; they consist of a single catalytic A subunit and a receptor-binding B subunit pentamer.
The stx genes are carried by lambdoid phages, and these phages have disseminated stx genes among a wide variety of bacterial species, including E. coli (i.e., enterohemorrhagic E. coli [EHEC] and enteroaggregative hemorrhagic E. coli [EAHEC]) (10), Enterobacter cloacae (9, 11), Escherichia albertii (12), and even Gram-positive Enterococcus (13). EHEC, so named because it frequently causes hemorrhagic colitis (bloody diarrhea), often harbors one or several stx genes. The most clinically relevant serotypes of EHEC are O157:H7, the “big six” (i.e., O26, O103, O45, O111, O121, and O145), and the EAHEC serotype O104:H4, which was responsible for the German outbreak of STEC in 2011. EAHEC can also carry stx genes and attach to the intestinal lining, causing hemorrhagic colitis (10). The EAHEC strain O104:H4, which has a stx2 gene (14), is responsible for the worst STEC outbreak in recent history, an outbreak in Germany in 2011 in which 3,816 people were affected and nearly 25% of the patients developed HUS (15).
Stx expression is driven by a lambdoid phage late-phase promoter; when the lytic cycle of the phage is activated, the expression of Stx is also activated (16). The lytic phase of lambdoid phages can be induced by cellular stresses, especially genotoxic stress and DNA-damaging agents such as UV light or certain antibiotics. Mitomycin C (MITO) is often used to induce Stx production in the laboratory (17). Unlike most other bacterial infections, there is no widely accepted antibiotic treatment for STEC, due to a combination of antibiotic-initiated stx gene induction and emerging antibiotic resistance. Some antibiotics that showed promise against STEC infections (e.g., gentamicin [GEN]) are now less active against certain strains and serotypes (18, 19), and treatment of STEC with certain antibiotics resulted in significantly higher risk of developing HUS (20, 21).
Antibodies have played important roles in the field of therapeutics. However, only low levels of endogenous serum antibodies against Stxs are induced in STEC infections (22, 23). Passively administered Stx-specific monoclonal antibodies (MAbs) have been shown to be highly successful in clearing Stx2 completely from intoxicated mouse blood and in protecting mice for up to 4 weeks (24). Stx-specific MAbs also could neutralize Stx cytotoxicity in Vero cell assays (25). Therefore, antibody therapy could be evaluated as an option for treating STEC infections.
In this study, we identified an antibiotic that inhibited the growth of all 42 STEC isolates we tested (from the environment and clinical samples) but did not induce Stx production. When this antibiotic was combined with a mixture of MAbs against Stx1 and Stx2 and used to treat Vero cells cocultured with STEC strains, we were able to inhibit fully the growth of the bacteria and to neutralize the cytotoxicity of the Stxs.
MATERIALS AND METHODS
E. coli strains and growth conditions.
All E. coli strains were grown in Luria-Bertani (LB) medium (Luria-Bertani broth; Miller, Fisher Scientific). E. coli isolates were arrayed in a 96-well cell culture plate (Fisher) and stored at −80°C in LB medium with 10% glycerol. Fresh cultures for subsequent experiments were established using this frozen array. For each experiment using the array (subinhibitory antibiotic treatments and PCR characterization of stx genes), bacteria were inoculated into 100 μl/well LB medium on a 96-well plate and were grown overnight at 37°C, with shaking at 150 rpm. This overnight culture of E. coli was used as the inoculum for the array experiments. The STEC strains used are listed in Table 1 and are from a previous study (26).
TABLE 1.
Isolates and Stx genes
| Strain | Serotype | Origin | stx1a | stx2a | Reference |
|---|---|---|---|---|---|
| 8385 | O103 | + | − | 26 | |
| 9882 | O103 | Cow feces | + | − | 26 |
| 10061 | O103 | Feral pig | + | − | 26 |
| 10408 | O103 | Crow | + | − | 26 |
| 13508 | O103 | Human | + | − | 26 |
| 7370 | O111 | Water | + | + | 26 |
| 9322 | O111 | Water | + | − | 26 |
| 9907 | O111 | Feral pig | + | − | 26 |
| 9975 | O111 | Crow | + | − | 26 |
| 12788 | O111 | Human | + | + | 26 |
| 7783 | O113 | Crow | − | + | 26 |
| 7788 | O113 | Water | − | + | 26 |
| 7958 | O113 | Cow feces | + | stx2d | 26 |
| 10466 | O113 | Cow feces | − | + | 26 |
| 10940 | O113 | Cow feces | − | + | 26 |
| 6848 | O121 | Lettuce | − | + | 26 |
| 8082 | O121 | Cow feces | stx1d | − | 26 |
| 8352 | O121 | Sediment | − | + | 26 |
| 13504 | O121 | − | + | 26 | |
| 5856 | O121:H19 | Human | − | + | 26 |
| 8876 | O145 | Water | + | − | 26 |
| 9306 | O145 | Cow feces | + | − | 26 |
| 9872 | O145 | Cow feces | + | + | 26 |
| 9917 | O145 | Feral pig | + | − | 26 |
| 12238 | O145 | Human | − | + | 26 |
| 1239 | O157H7 | Human | − | + | 26 |
| 1913 | O157H7 | Human | − | + | 26 |
| 2367 | O157H7 | Human | + | + | 26 |
| 6649 | O157H7 | Human | + | + | 26 |
| 7543 | O157H7 | Human | + | + | 26 |
| 7375 | O26 | Human | + | − | 26 |
| 7927 | O26 | Water | + | − | 26 |
| 8426 | O26 | Water | + | − | 26 |
| 10817 | O26 | Cow feces | + | − | 26 |
| 13151 | O26 | Human | + | − | 26 |
| 7103 | O45 | − | − | 26 | |
| 9413 | O45 | Cow feces | + | − | 26 |
| 13506 | O45 | Human | + | − | 26 |
| 13752 | O45 | Cow feces | + | − | 26 |
| 15012 | O104:H4 | Human | − | + | 45 |
| 15014 | O104:H4 | Human | − | + | 45 |
| ATCC 25922 | O6 | Human | − | − | 28 |
Preparation of Shiga toxins and toxoids.
Stx2 toxin was purified from the Stx2a-expressing RM10638 strain as described previously (27). Partially purified Stx1 (approximately 14% pure; 0.5 mg/ml Stx1, 3.5 mg/ml total protein) was purchased from Toxin Technologies. Stx2a (E167Q) and Stx1 (E167Q) toxoids were purified as described previously (27, 28). Protein concentrations were determined using a bicinchoninic acid (BCA) protein assay (Pierce).
Probiotic strains and growth conditions.
Lactobacillus acidophilus was purchased from the American Type Culture Collection (ATCC) (catalog no. 4356). L. acidophilus was grown overnight in lactobacilli MRS broth (Benton Dickenson), diluted to an absorbance at 600 nm (A600) of 0.0002 (∼1 × 105 cells/ml) in MRS broth, inoculated onto GN3F Sensititre plates at 50 μl bacterial suspension/well, and grown for 48 h at 37°C in 5% CO2. Plates were read at 660 nm on a Victor 3 plate reader. All wells with A660 values of >0.075 were considered positive; A660 values ranged from 0.15 (full growth) to 0.035 (no growth). Saccharomyces boulardii (catalog no. MYA796; ATCC) was generously donated by Luisa Cheng. S. boulardii was grown overnight in YPD (yeast extract-peptone-2% dextrose) medium, diluted to an A600 of 0.0002 (∼104 cells/ml) in YPD medium, inoculated onto GN3F plates at 50 μl/well, and grown for 48 h at 37°C, with shaking at 150 rpm. These plates were also read at 660 nm on a Victor 3 plate reader. All wells with A660 values of >0.075 were considered positive; A660 values ranged from 0.2 (full growth) to 0.031 (no growth).
Antibiotic susceptibility screen.
To identify antibiotics that were inhibitory against a variety of STEC strains, GN3F Sensititre Gram-negative antibiotic array plates were used, as suggested by the manufacturer (Thermo Scientific). Briefly, an overnight culture of E. coli cells was diluted to an A600 of 0.0002 (∼1.6 × 105 cells/ml; for E. coli, an A600 value of 1 indicates approximately 8 × 108 cells/ml) in LB broth. This suspension was added to GN3F plates at 50 μl/well. Plates were sealed with tape and incubated for 16 h at 37°C, with shaking at 150 rpm. Phosphate-buffered saline (PBS) was added at 50 μl per well, the cells were resuspended by being pipetted up and down, and the absorbance was read at 660 nm. All wells with A660 values of >0.1 were considered positive for bacterial growth; A660 values ranged from 0.4 (full growth) to 0.075 (no growth) (plates were read with the lids on, because they contained pathogens). Wells with A660 values of 0.1 were slightly turbid by visual inspection. All experiments using GN3F plates were performed once (one isolate per plate), and then the GN3F antibiotics were resuspended in LB medium and the results were confirmed twice using those antibiotic preparations. A description of the GN3F plates can be found at http://www.trekds.com/products/sensititre/files/GN3F.pdf.
Stx PCR confirmation.
Stx diagnostic PCRs were performed as described previously (29). Previously described primers and reaction conditions were used (8). Briefly, stx1-det-F1 and stx1-det R1 primers were used to determine whether the isolates had a stx1 gene, and isolates that possessed a stx1 gene were analyzed with a Stx1 variant multiplex PCR assay to determine the subtype. Isolates were then analyzed for the presence of a stx2 gene. Isolates positive for stx2 were subjected to individual subtype PCRs for Stx2a, Stx2b, Stx2c, Stx2d, Stx2e, Stx2f, and Stx2g. All PCR products were run for 20 min on a 0.8% agarose gel with ethidium bromide added and were visualized under UV illumination with an U:Genius imager (Syngene, Cambridge, United Kingdom). PCR results were confirmed a total of three times.
Enzyme-linked immunosorbent assays.
For sandwich enzyme-linked immunosorbent assays (ELISAs), coating antibody (MAb Stx1-3 for Stx1 or MAb Stx2-1 for Stx2) (25, 28) was diluted in phosphate-buffered saline (PBS) to 1 μg/ml, and then 100 μl/well was allowed to bind to black Nunc Maxisorb ELISA plates overnight at 4°C. This was followed by two washes with PBS with 0.05% Tween 20 (PBST), blocking solution (3% bovine serum albumin [BSA] in PBST) was added at 200 μl/well, and the plates were incubated for 1 h at room temperature. During this blocking step, 100 μl/well PBS was added to 96-well culture plates containing an overnight culture of bacteria (grown overnight at 37°C; 100 μl LB medium/well). The plates were centrifuged at 2,000 rpm for 20 min, and then 100 μl/well of bacterial cell-free supernatant was transferred to a separate culture plate. After the ELISA plate was washed twice more with PBST, 95 μl/well PBS was added, followed by 4 μl/well bacterial cell-free supernatant; this mixture was incubated for 1 h at room temperature. The ELISA plate was then washed six more times with PBST, 0.5 μg/ml secondary antibody (biotinylated MAb Stx1-1 for Stx1 or biotinylated MAb Stx2-5 for Stx2) (25, 28) diluted in BSA/PBST was added at 100 μl/well, and the plate was incubated for 1 h at room temperature. After six washes, 100 μl/well of 0.2 μg/ml streptavidin-horseradish peroxidase (HRP) diluted in PBST with BSA was added to the plates. After incubation at room temperature for 1 h and six washes, the plates were developed with Pico chemiluminescent substrate (Thermo Scientific). Luminescence was measured using a Victor 3 plate reader (PerkinElmer). Biotinylations were conducted using a Lightning-Link biotin conjugation kit type A (Innova Biosciences, Cambridge, United Kingdom), according to the manufacturer's instructions. All ELISAs were conducted three times for confirmation, which yielded similar results. Concentrations of Stx1 and Stx2 based on optical density (OD), as shown in Fig. 1 and 2, were calculated by dividing the ELISA result by the A595 value (which is proportional to the cell density) for each culture well and then extrapolating the values using a Stx1 (E167Q) or Stx2 (E167Q) toxoid standard curve. P values were derived using Student's t test.
FIG 1.
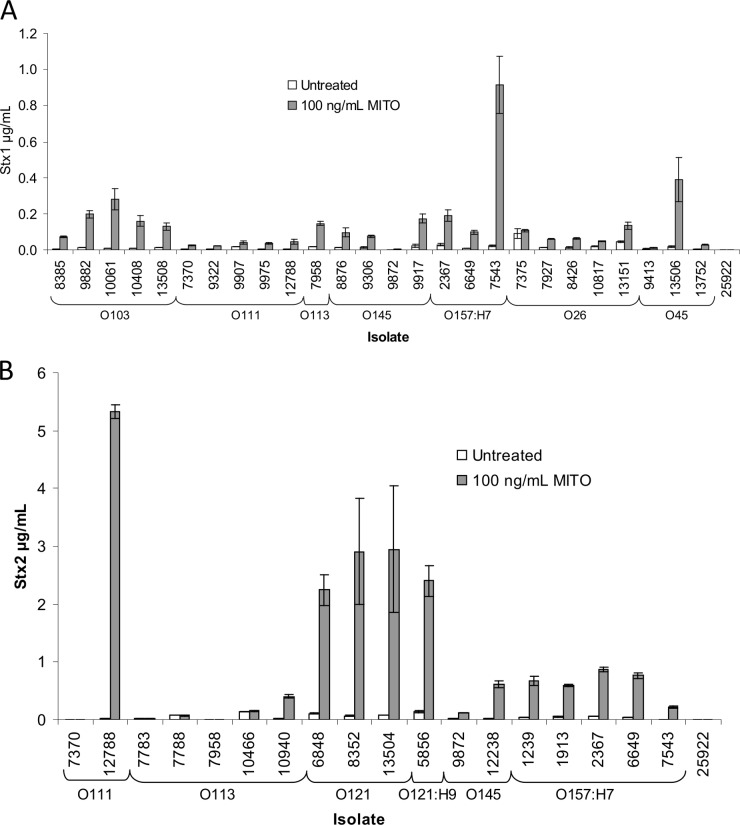
Stx induction by MITO. (A) Stx1 sandwich ELISAs were conducted with cell-free supernatants from all Stx1-containing STEC cultures. The Stx1 sandwich ELISAs used MAb Stx1-3 (1 μg/ml) as a capture antibody and Stx1-1-biotin (0.5 μg/ml) as a detection antibody. All MITO-treated samples underwent the same treatment conditions and preparations, except that they were grown in 100 ng/ml MITO-containing medium. (B) Stx2 sandwich ELISAs were conducted with cell-free supernatants from all Stx2-containing STEC cultures. The Stx2 sandwich ELISAs used Stx2-1 (1 μg/ml) as a capture antibody and Stx2-5-biotin (0.5 μg/ml) as a detection antibody. All MITO-treated samples underwent the same treatment conditions and preparations, except that they were grown in 100 ng/ml MITO-containing medium.
FIG 2.
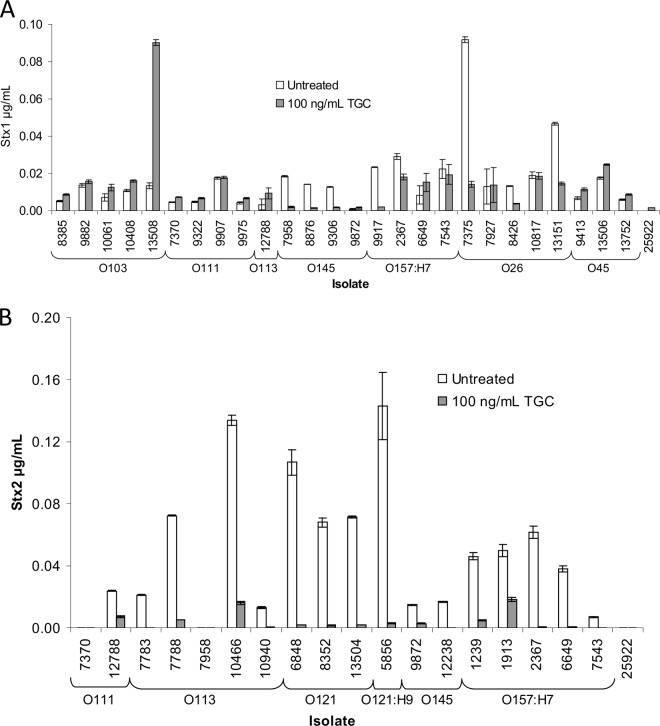
Stx repression by TGC. (A) Stx1 sandwich ELISAs were conducted with TGC-treated and untreated cell-free supernatants from all Stx1-containing STEC cultures, using MAb Stx1-3 (1 μg/ml) as a capture antibody and Stx1-1-biotin (0.5 μg/ml) as a detection antibody. All TGC-treated samples were grown in 100 ng/ml TGC-containing medium. (B) Stx2 sandwich ELISAs were conducted with TGC-treated (100 ng/ml) and untreated cell-free supernatants from all Stx2-containing STEC cultures, using Stx2-1 (1 μg/ml) as a capture antibody and Stx2-5-biotin (0.5 μg/ml) as a detection antibody.
Antibiotic preparations.
Antibiotics were harvested from GN3F plates by adding 100 μl/well LB broth, incubating the plates for 10 min, and then mixing and removing the solutions under sterile conditions. All concentrations (two to seven concentrations) of an antibiotic were pooled in a single sterile tube, and the resulting antibiotic concentration was calculated to be one-half the average of the concentrations (wells of GN3F plates are supposed to contain only 50 μl of medium). Antibiotic solutions were stored in LB broth at 4°C for up to 1 week in the dark and then were discarded.
Vero cell cytotoxicity assays.
Vero (African green monkey adult kidney) cells (ATCC CCL-81) were obtained from the American Type Culture Collection (Manassas, VA) (30). Vero cells were prepared and grown as described previously (29). Briefly, the medium used for Vero cell propagation and growth (in a humidified cell culture incubator at 37°C with 5% CO2), was Dulbecco's modified Eagle's medium (DMEM) (Invitrogen) with 10% fetal bovine serum (FBS) (Invitrogen). In preparation for cytotoxicity assays, the cells were trypsinized, diluted to 105 cells/ml, and dispensed into 96-well cell-culture-treated plates, and these plates were incubated for 24 h. For cytotoxicity assays using subinhibitory concentrations of antibiotic, antibiotic-induced filtered STEC medium was diluted in fresh Vero medium (5 nl or 25 nl STEC medium per 100 μl Vero medium). A sterile mixture of MAb Stx1-3 and MAb Stx2-5 (25) was then added for a final concentration of 1 μg/ml each (PBS was added instead for the samples without antibodies). The medium in the Vero cell plate was then removed and replaced with the STEC medium-antibody mixture (100 μl/well). Twenty-four hours after treatment, the Vero cells were lysed using 100 μl/well of a 1:5 dilution of CellTitre-Glo reagent (Promega), with 3 min of shaking. Luminescence was measured using a Victor II plate reader. All Vero cell toxicity assays were conducted three times, with similar results.
For cytotoxicity experiments using inhibitory concentrations of antibiotic, ceftazidime (CAZ) plus gentamicin (GEN), tigecycline (TGC), and ciprofloxacin (CIP) were added to Vero medium for final concentrations of 1 μg/ml each, 2 μg/ml, and 0.2 μg/ml, respectively. To this medium, a sterile mixture of MAbs Stx1-3 and Stx2-5 was added at a final concentration of 10 μg/ml (or PBS was added instead of antibody). An overnight culture of STEC (strains 1913, 2367, 13151, and ATCC 25922) was then added to the antibiotic- and antibody-containing medium (5 nl per 100 μl medium). The medium in the Vero cell plate was then removed and replaced with the STEC cell suspension-antibody mixture (100 μl/well). Twenty-four hours after treatment, the wells were photographed (×200 magnification, using a Leica microscope), and the cells were lysed using 100 μl/well of a 1:5 dilution of CellTitre-Glo reagent (Promega). Luminescence was measured using a Victor II plate reader as described above. These assays were conducted three times, and a representative experiment is shown.
RESULTS
Antibiotic susceptibility of STEC strains and probiotics.
To identify antibiotics that can inhibit the growth of STEC, 42 STEC strains (Table 1), including the O157, non-O157 “big six,” and O113 serotypes, that were collected from clinical or environmental settings were screened for their antibiotic susceptibility by using a commercially available Gram-negative antibiotic array (GN3F) (26). Properties of the antibiotics tested are summarized in Table S1 in the supplemental material. At least one strain grew at the lowest concentrations of the majority of antibiotics in the array (see Table S2 in the supplemental material). For ertapenem (ETP), piperacillin-tazobactam (TZP) constant 4, trimethoprim-sulfamethoxazole (SXT), cefpodoxime (CPD), ceftazidime (CAZ), tigecycline (TGC), and ceftriaxone (CRO), however, no strain survived even at the lowest concentration of each antibiotic. Further dilutions revealed the MICs for these antibiotics (see Table S2 in the supplemental material). Only ≤10% of the strains tested grew in the presence of the lowest concentrations of ampicillin-sulbactam (SAM), aztreonam (ATM), cefazolin (CFZ), cefepime (FEP), meropenem (MEM), ciprofloxacin (CIP), and ticarcillin-clavulanic acid (TIM) constant 2. Despite the occasional use of gentamicin (GEN) as a treatment for STEC, almost all strains tested in this study grew at the lowest concentrations of GEN. All strains tested grew at the lowest concentration of tobramycin (TOB). All recorded MICs are listed in Table S3 in the supplemental material.
A successful treatment regimen for STEC should not disrupt the intestinal commensal microflora or at least should allow its rapid recovery. Accordingly, we grew two probiotics on GN3F antibiotic plates to determine their sensitivities to promising antibiotics for STEC treatments. Lactobacillus acidophilus, a popular Gram-positive probiotic, was sensitive to many of the antibiotics that were inhibitory for the STEC strains (see Table S2 in the supplemental material). However, L. acidophilus notably grew in the presence of 1 μg/ml CAZ (see Table S2 in the supplemental material), a concentration that inhibited all of the STEC strains tested. Saccharomyces boulardii, a eukaryotic (yeast) probiotic, was resistant to every antibiotic on the GN3F plates (see Table S2 in the supplemental material), which suggests that it could recolonize the intestinal environment regardless of which antibiotic is used to treat STEC.
Induction of Stx by antibiotics.
Subtyping PCRs were performed for all strains, to confirm the presence of stx1 and stx2 genes. All strains tested contained stx1a, stx2a, or both stx1a and stx2a genes, with the exceptions of strains 8082 and 7958, which contained a stx1d gene and a stx2d gene, respectively. None of these strains appeared to contain more than one subtype of Stx1 or Stx2. Strain 7103 did not contain any stx gene (Table 1). These PCRs were conducted using primers reported previously for subtyping (8), in a single-amplicon or multiplex format.
The relative concentrations of Stx1 and Stx2 in bacterial media were determined by ELISA and then corrected for bacterial growth (by dividing the values by the optical density at 660 nm). For Stx1 ELISAs, MAbs that recognize the B subunit of Stx1 were used (Stx1-3 for capture and Stx1-1 for detection) (28). For Stx2 ELISAs, MAbs that recognize Stx2 were used (Stx2-1 for capture and Stx2-5 for detection) (25). It was found that the Stx levels varied significantly among these 40 strains in the absence or presence of antibiotics. The serotype with the most abundant soluble Stx1 in antibiotic-free medium was O26 (average for five strains; P < 0.01), followed by O157:H7 (Fig. 1A and 2A). The serotype with the highest levels of released Stx2 was O121 (average for four strains; P = 0.01), followed by O157:H7 (Fig. 1B and 2B). As reported previously (31, 32), treatment of STEC strains with a subinhibitory concentration of mitomycin C (MITO) strongly induced Stx production (both Stx1 and Stx2) for most strains, with average induction levels of 8.3-fold for Stx1 (Fig. 1A) and 38.2-fold for Stx2 (Fig. 1B).
We then examined the induction of Stx production by the aforementioned seven antibiotics (ETP, TZP, SXT, CPD, CAZ, TGC, and CRO), which inhibited the growth of all tested strains at concentrations below the MIC breakpoints for susceptible strains of Enterobacteriaceae (see Table S2 in the supplemental material) (33). All bacterial strains were grown overnight in LB medium containing one of the antibiotics at its highest subinhibitory concentration, and the Stx levels in the medium were determined. The six antibiotics other than TGC induced Stx1 and/or Stx2 production in multiple strains (>2-fold increases in Stx levels with antibiotic treatment were considered induced), but none of them reached the extent of MITO (Table 2). CPD induced Stx1 production in all of the tested strains and Stx2 production in 28% of the tested strains. ETP, TZP, CAZ, and CRO increased Stx1 production in 46 to 92% of the Stx1-expressing strains and Stx2 production in less than 25% of the Stx2-producing strains. SXT induced Stx1 and Stx2 production in 19% and 39% of the strains, respectively. Overall, TGC did not induce Stx1 production except in strains 12788 and 13508. In many strains, TGC inhibited Stx1 production. The production of Stx2 was inhibited strongly by TGC in all of the strains tested (Fig. 2A and B).
TABLE 2.
Stx induction by first-tier antibioticsa
| Antibiotic | Antibiotic concentration (μg/ml) | No. of isolates induced/total no. of isolatesb |
% of isolates induced | Average fold induction | ||
|---|---|---|---|---|---|---|
| Stx1 | Stx2 | Total Stxc | ||||
| ETP | 0.004 | 24/26 | 1/18 | 25/44 | 57 | 5.7 |
| TZP | 0.5 | 23/26 | 1/18 | 24/44 | 55 | 6.2 |
| SXT | 0.03 | 5/26 | 7/18 | 12/44 | 27 | 2.6 |
| CPD | 0.1 | 26/26 | 5/18 | 31/44 | 70 | 6.2 |
| CAZ | 0.03 | 14/26 | 4/18 | 18/44 | 41 | 2.9 |
| TGC | 0.1 | 2/26 | 0/18 | 2/44 | 5 | 0.8 |
| CRO | 0.01 | 12/26 | 2/18 | 14/44 | 32 | 2.7 |
| MITO | 0.1 | 24/26 | 16/18 | 37/44 | 89 | 22.9 |
All strains were tested except strains 7103, 8082, 15012, 15014, and 25922.
More than 2-fold increases in Stx levels with antibiotic treatment indicated induction.
A total of 37 STEC strains were tested; seven of them expressed both Stx1 and Stx2.
In order to simplify analysis, a collection of eight strains from different serotypes was used to study the effects of the rest of the antibiotics (at subinhibitory concentrations) on Stx production (strain RM2367 expresses both Stx1 and Stx2) (see Table S4 in the supplemental material). Overall, treatment with most of the antibiotics induced production of Stx1 only, while CIP strongly induced production of both Stx1 and Stx2 and tetracycline (TET), GEN, and TOB inhibited Stx production.
In vitro STEC treatment regimen.
An ideal antibiotic for the treatment of STEC infections should kill or inhibit the bacteria but should not induce Stx expression at any concentration of the antibiotic. Our results indicated that TGC effectively prevented bacterial growth at its inhibitory concentration (see Table S2 in the supplemental material) but did not induce Stx1 or Stx2 production in general at a universally subinhibitory concentration (Table 2). Therefore, we chose TGC for further study of its utility in STEC treatments, using an in vitro cell-based toxicity assay.
In order to assess the effect of TGC on the production of Stxs in a mammalian cell-based toxicity model, we grew STEC strains overnight with a subinhibitory concentration of TGC (100 ng/ml) and then applied the filtered bacterial culture medium to Vero cells. CIP (at a subinhibitory concentration of 10 ng/ml) was included for comparison because it is also active in killing bacteria (see Table S2 in the supplemental material) but induces Stx production (see Table S4 in the supplemental material). Four representative strains of STEC with relatively high levels of Stx expression (strain 1913, Stx2 only; strain 2367, Stx1 and Stx2; strain 13151, Stx1 only; strain 25922, no toxin) were used. TGC-treated STEC culture medium increased Vero cell viability, compared to STEC culture medium without antibiotics (Fig. 3A), for both Stx2-expressing strains (P values of 0.02 [strain 1913] and 0.004 [strain 2367]), while CIP-treated STEC culture medium significantly decreased Vero cell viability (P values of 0.03 [strain 1913], 0.003 [strain 2367], and 0.02 [strain 13151]). This confirmed that TGC did not induce production of additional Stxs or other toxic components and TGC was not harmful to Vero cells at a subinhibitory concentration.
FIG 3.
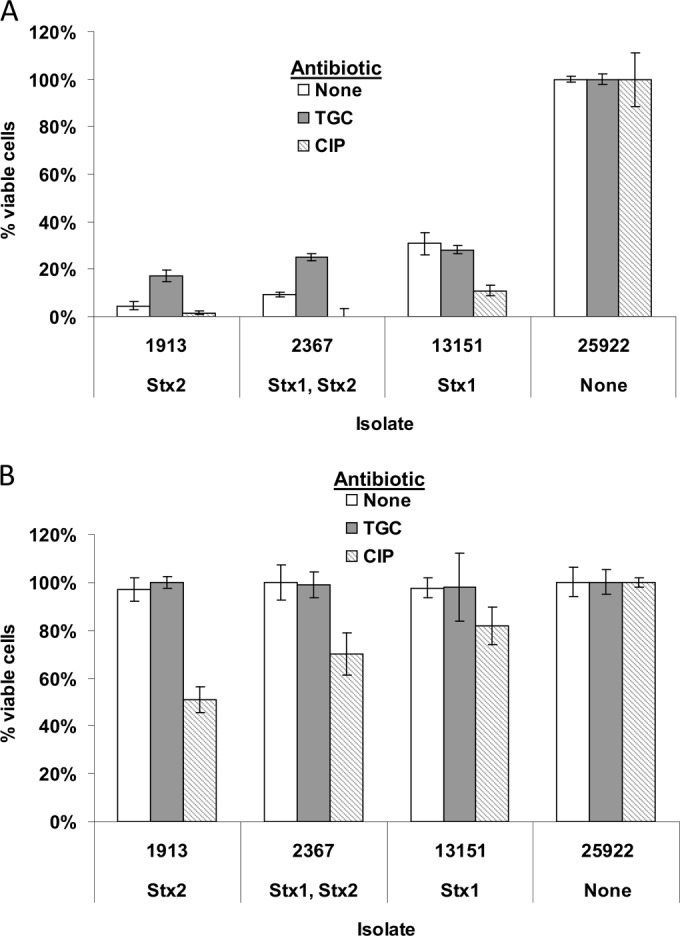
Cytotoxicity of Stxs from culture medium from bacteria incubated with antibiotics at subinhibitory concentrations. (A) Cell-free supernatants from isolates expressing Stx2 only (RM1913), Stx1 and Stx2 (RM2367), Stx1 only (RM13151), or no toxin (ATCC 25922), grown with 100 ng/ml TGC, 10 ng/ml CIP, or no antibiotic, were added to cell culture wells containing 105 Vero cells, and plates were incubated for 24 h. All samples were added at 25 nl cell-free supernatant/well. (B) Cell-free supernatants from the experiment shown in panel A were added to Vero cell medium containing a mixture of anti-Stx antibodies (MAbs Stx1-1 and Stx2-5). This cell supernatant-antibody mixture was then added to cell culture wells containing 105 Vero cells, and plates were incubated for 24 h. All samples were added at 25 nl cell-free supernatant/well. Antibodies were provided at 1 μg/ml each.
To examine whether the toxins produced by these bacteria could be neutralized by MAbs against Stxs, a mixture of anti-Stx1 and anti-Stx2 antibodies (MAbs Stx1-1 and Stx2-5) was premixed with filter-sterilized culture medium from antibiotic-treated or untreated STEC strains (the same medium as used in Fig. 3A) and then added to Vero cells. This mixture of MAbs (at 1 μg/ml each) was able to neutralize the toxicities of the untreated and TGC-treated STEC media effectively, protecting >75% of Vero cells (Fig. 3B). However, the amounts of toxins in the bacterial medium from CIP-treated strains (strains 1913 and 2367) overwhelmed the antibodies, and significant cell death was once again observed (Fig. 3B) (P values of 0.006 [strain 1913], 0.03 [strain 2367], and 0.01 [strain 13151], comparing untreated versus CIP treated).
To study more directly whether the combination of antibiotic and antibody could protect mammalian cells from STEC in culture, we cocultured bacteria with Vero cells. E. coli grows robustly in the medium used to culture Vero cells (see Fig. S1 in the supplemental material), and microscopic evaluations and assays for viability of the Vero cells are impossible to conduct in the absence of sufficient antibiotic, due to bacterial overgrowth. Therefore, bacteria were inoculated into a Vero cell culture well containing a concentration of antibiotic just above the MIC for the tested STEC strains. We tested three antibiotic regimens, i.e., a combination of CAZ and GEN (at 1 μg/ml each), TGC (at 2 μg/ml), and CIP (at 0.2 μg/ml). The CAZ-GEN combination (GEN was added to inhibit Stx production) was included because it is inhibitory against all STEC strains tested and serves for comparison with TGC and CIP. The same four STEC strains used in the subinhibitory experiments were used again in this study (strains 1913, 2367, 13151, and 25922). Although the bacteria appeared to be completely inhibited in the presence of antibiotics (as observed microscopically), many fewer Vero cells survived in the antibiotic-treated wells containing Stx-expressing strains 1913, 2367, and 13151; strain 25922 (containing no stx genes) had no effect on Vero cell viability (Fig. 4A; see also Fig. S2 in the supplemental material). Even fewer Vero cells survived in the CIP-treated wells than in the CAZ- and GEN-treated wells, and the most Vero cells survived in the wells treated with TGC (Fig. 4A). We suggest that the reduction of Vero cell viability in wells cocultured with bacteria may be caused by a combination of the Stxs present in the initial bacterial inocula and the Stxs induced during bacteria death.
FIG 4.
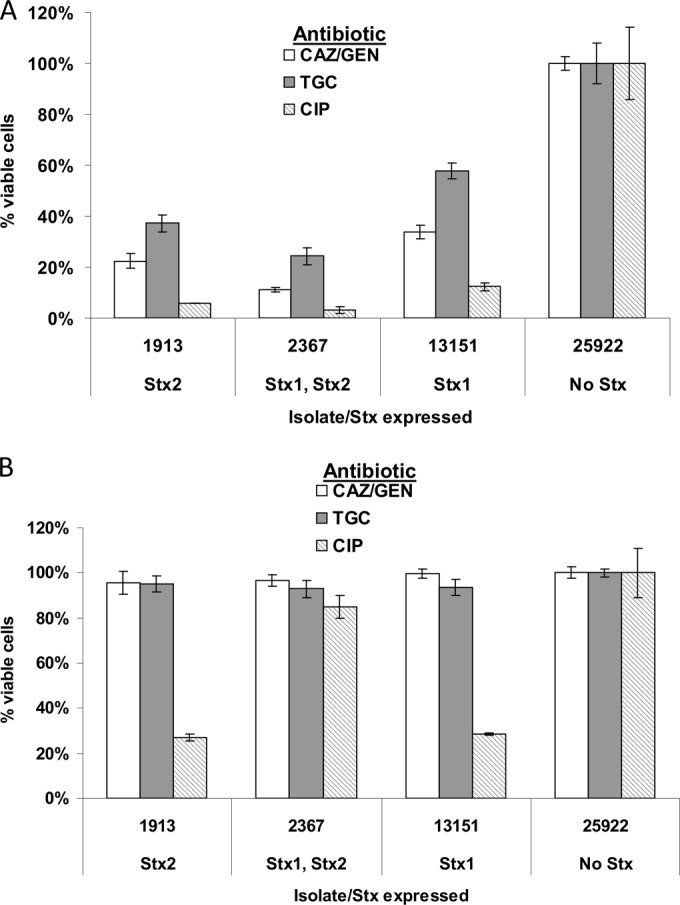
Effects of antibiotics at inhibitory concentrations on the viability of Vero cells cocultured with STEC. (A) STEC isolates expressing Stx2 only (RM1913), Stx1 and Stx2 (RM2367), Stx1 only (RM13151), or no toxin (ATCC 25922) were inoculated into Vero cell medium containing inhibitory concentrations of the indicated antibiotics (1 μg/ml each for CAZ and GEN, 2 μg/ml for TGC, and 0.2 μg/ml for CIP). (B) STEC isolates were combined with antibiotic, a mixture of neutralizing anti-Stx MAbs (Stx1-1 and Stx2-5 at 10 μg/ml each), and Vero cell medium and then were added immediately to Vero cells already present in cell culture wells. All Vero cell culture wells received 5 nl of stationary-phase bacteria (A600 of 3) per well.
In order to develop a treatment regimen that is useful not only in combating STEC but also in neutralizing the toxicity derived from the bacteria, we cocultured Vero cells with one of the four bacterial strains in the presence of antibiotic (TGC at 2 μg/ml, CAZ and GEN at 1 μg/ml each, or CIP at 0.2 μg/ml) and the same MAb mixture (at 10 μg/ml each) as used in the subinhibitory experiments. Our results indicated that treatment with TGC and MAbs or CAZ, GEN, and MAbs successfully controlled bacterial growth and protected the Vero cells from Stx intoxication (P values of 0.002 [strain 1913], 0.008 [strain 2367], and 0.001 [strain 13151], comparing CIP and TGC). Treatment with CIP and MAbs rescued fewer Vero cells from death (P values of 0.001 [strain 1913], 0.06 [strain 2367], and 0.001 [strain 13151], comparing CIP versus TGC with MAbs) (Fig. 4B; see also Fig. S2 in the supplemental material).
O104:H4, an attaching and effacing EAHEC serotype, can promote greater secretion of mucus in the intestine, possibly extending the amount of time it can persist in the digestive tract and increasing the amount of toxin it can release (34). Therefore, antibiotic treatment may be even more useful for treating O104:H4 infections than for treating other STEC infections. Using two isolates of O104:H4 from the German outbreak in 2011, we evaluated the efficacy of treatment with TGC and antibodies in Vero cell culture. A subinhibitory concentration of TGC significantly reduced the toxicity of O104:H4 to Vero cells, suggesting that TGC inhibited Stx2 production for both isolates (average P value of 0.02) (Fig. 5A); with neutralizing MAbs (a mixture of Stx1-1 and Stx2-5 at 1 μg/ml each), however, Vero cells were fully rescued from O104:H4 cell-free medium. The TGC-antibody mixture similarly protected Vero cells from O104:H4 bacteria when they were treated with an inhibitory concentration of antibiotic (Fig. 5B). Treatment of O104:H4 isolates with CIP at an inhibitory concentration resulted in greater cytotoxicity to Vero cells (average P value of 0.02), and cell viability did not fully recover with the addition of neutralizing MAbs (average P value of 0.01). CAZ and GEN treatment of O104:H4 isolates was not attempted, since this O104:H4 strain expresses an extended-spectrum beta-lactamase (ESBL).
FIG 5.
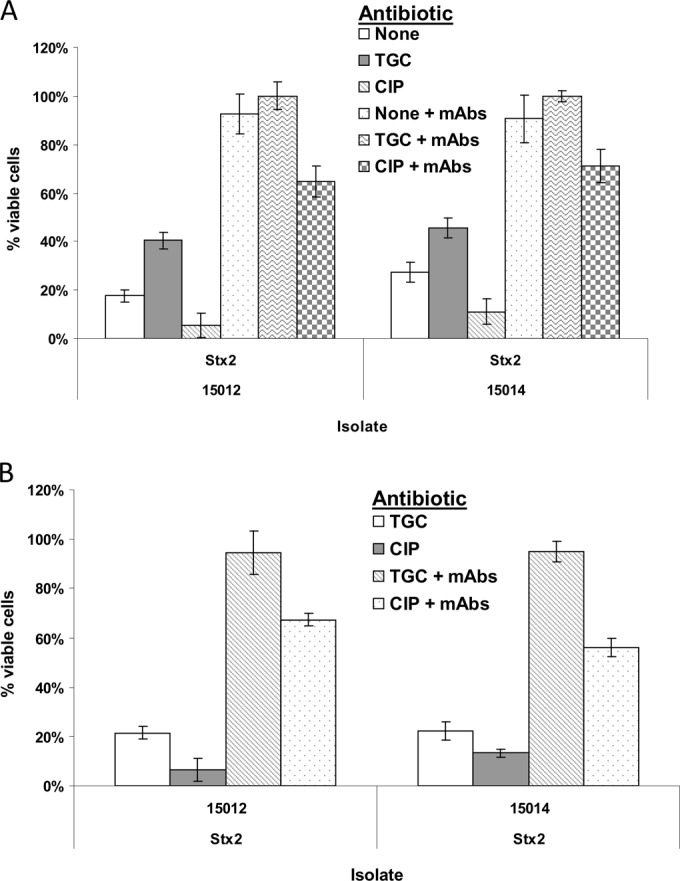
Effects of antibiotics and antibodies on the viability of Vero cells exposed to O104:H4 strains. (A) Cell-free medium from O104:H4 isolates (from the German outbreak in 2011) grown with 100 ng/ml TGC, 10 ng/ml CIP, or no antibiotic was added to cell culture wells containing 105 Vero cells, and plates were incubated for 24 h. A subset of cell-free medium samples were also incubated with a MAb mixture (1 μg/ml each of Stx1-1 and Stx2-5). All samples were added at 25 nl cell-free supernatant/well. (B) O104:H4 isolates were inoculated into Vero cell medium containing inhibitory concentrations of the indicated antibiotics (1 μg/ml each for CAZ and GEN, 2 μg/ml for TGC, and 0.2 μg/ml for CIP). A subset of samples also received a MAb mixture (10 μg/ml each of Stx1-1 and Stx2-5). All Vero cell culture wells received 5 nl of stationary-phase bacteria per well.
DISCUSSION
STEC infections can have a variety of clinical manifestations, ranging from diarrhea to life-threatening HUS. Although these infections can be deadly, there is still no widely accepted universal treatment. Therefore, we sought to find antibiotics that would be inhibitory to the majority of STEC strains. To accomplish this goal, we applied a diverse collection of STEC isolates (including O157:H7 and the “big six” serotypes) to a Gram-negative antibiotic array, looking for both inhibition of bacterial growth and reduction of toxin production. All tested isolates were sensitive to ETP, TZP, SXT, CPD, CAZ, TGC, and CRO. Of these, however, only TGC did not induce Stx production, with the exception of two isolates; instead, TGC inhibited Stx production in almost one-half of the isolates. TGC at either a subinhibitory or inhibitory concentration synergized remarkably well with Stx-neutralizing antibodies to protect fully Vero cells in culture, suggesting that the combination of TGC with neutralizing antibodies ameliorates the effects and aftereffects of STEC infections better than antibiotic treatment alone. This TGC-antibody combination also fully protected Vero cells from O104:H4 toxicity. Additionally, we determined that TGC did not affect the growth of Saccharomyces boulardii, a yeast probiotic that could be administered with the TGC-antibody combination to initiate the recovery of intestinal flora.
Whether antibiotic treatment of STEC increases or decreases the risk of developing HUS is still a matter of debate, but our data suggest that the risk may depend on more than just the antibiotic used. MITO and CIP strongly induced Stx1 and Stx2 production in a majority of the strains we tested, but they did not affect Stx levels in some strains (such as strains 7375, 7783, and 7788). Therefore, the risk of exacerbating HUS with antibiotic use might be dependent on which antibiotic is used and the characteristics of the particular strain (including whether Stx production is inducible). This added complexity could account for the contrasting reports on the efficacy of STEC treatment with antibiotics. While treatment of O157:H7 with ciprofloxacin is occasionally reported to exacerbate HUS symptoms (21), a recent report indicated that antibiotic treatment (with ciprofloxacin) of German O104:H4 EAHEC was effective in preventing the development of HUS (35). This strain, as an enteroaggregative strain of STEC, embeds itself in intestinal mucus and may persist longer than EHEC, resulting in larger proportions of patients with HUS (∼25%) due to longer exposure to Stxs. For enteroaggregative STEC, it might be more important to eliminate the pathogenic E. coli, but selecting an antibiotic that does not induce Stx production is still a concern. Inclusion of the proper dose of neutralizing antibodies in an antibiotic regimen, however, might turn even antibiotics that strongly induce Stx production (such as CIP, CPD, and SXT) into potent treatments (Fig. 4B). A neutralizing antibody mixture may even be useful as a treatment on its own.
Bactericidal antibiotic treatments of STEC could liberate additional Stx from the bacteria (beyond that from phage-mediated lysis). However, bacteriostatic antibiotics, which inhibit the growth of bacteria while not necessarily killing them, may not liberate Stxs from STEC, allowing the toxins to pass harmlessly, encapsulated within the bacteria, from the intestine. Therefore, the bacteriostatic property of TGC may be essential for its mild inhibition of Stx levels. Bactericidal antibiotics such as CRO, MEM, and CIP would be expected to release more Stxs into the medium through antibiotic-associated lysis, complementing or enhancing phage-associated lysis. Serendipitously, the three strongest inducers of Stx production (ATM, CIP, and CPD) are bactericidal antibiotics but induce Stx production even at subinhibitory levels, suggesting that they also promote phage-associated lysis. The strongest repressors of Stx (GEN, TOB, TGC, and TET) are all bacteriostatic antibiotics, although they all belong to the aminoglycoside or tetracycline classes and inhibit protein synthesis; therefore, their repression of Stx may be due to their function rather than their ultimate effects on the bacteria.
From the data presented here, it appears that combined treatment with TGC and anti-Stx antibodies would be effective in neutralizing Stx toxicity and eliminating the bacterial infection for the majority of STEC strains and serotypes. This treatment strategy has several advantages, i.e., TGC does not induce Stx production at inhibitory or subinhibitory concentrations and is broadly inhibitory to bacteria, and anti-Stx IgG can neutralize the remainder of the toxins produced. Additionally, TGC is active against Clostridium difficile (36), a common secondary infection, and does not inhibit the probiotic S. boulardii, which, in a clinical setting, might be used concurrently to repopulate the intestinal flora. TGC is even active against strains of E. coli that express extended-spectrum beta-lactamases (ESBLs) (37) or carbapenemases (38, 39). TGC has several drawbacks, however. It is a very broad-spectrum antibiotic, suppressing the growth of both Gram-positive and Gram-negative bacteria and likely leaving many vacancies for colonization with resistant opportunistic microorganisms. It inhibits Lactobacillus acidophilus very effectively (see Table S2 in the supplemental material) and probably would do the same to other nonpathogenic intestinal microflora. In addition, there is currently no oral formulation for TGC, and it is not known whether TGC can enter the intestine effectively after intravenous administration, although some amount might cross the intestinal barrier during hemorrhagic colitis. A recent report stated that orally administered TGC was effective in Clostridium difficile infections in pigs (40), however, and certain formulations of TGC have increased stability (41); therefore, TGC could conceivably be effective when delivered orally to humans. TGC is not currently approved for use in children, in whom the effects of HUS are most severe. This may simply be a matter of finding the right dosage (42).
Although CAZ did induce Stx production in some strains, it effectively inhibited bacterial growth. By incorporating CAZ with GEN, Stx1 and Stx2 production might be reduced, at least to levels low enough to be neutralized by antibody treatment (Fig. 4). Therefore, treatment with CAZ, GEN, and anti-Stx antibodies may be an alternative to the TGC-antibody combination. One of the unique characteristics of CAZ is that it is broadly active against Gram-negative bacteria but is not especially toxic to Gram-positive bacteria. Thus, CAZ treatment would enable Gram-positive commensal organisms such as Lactobacillus acidophilus to initially recolonize the intestine. GEN, at concentrations lower than 4 μg/ml, is not toxic to L. acidophilus. One of the disadvantages of this treatment regimen is that it may not kill STEC strains expressing extended-spectrum beta-lactamases, although pairing it with a beta-lactamase inhibitor could overcome this limitation (43). Another disadvantage is that low-dose CAZ treatment induces filament formation in E. coli, which can result in increased endotoxin release (44).
An ideal universal STEC treatment would include an antibiotic that both eliminates the STEC bacteria and inhibits Stx production, antibodies that neutralize residual toxins in the bloodstream and intestine, and a method for rapidly recovering intestinal microflora. Although the combined TGC-antibody treatment we describe requires further modification, testing, and validation before being considered for clinical use, our data suggest that TGC and CAZ are broadly inhibitory against most STEC strains and TGC also inhibits the O104:H4 serotype. A cocktail of antibodies against Stx1 and Stx2 would serve to inactivate residual toxins, and a eukaryotic commensal organism such as S. boulardii could be used to repopulate the intestine initially, allowing swifter recovery after STEC infections.
Supplementary Material
ACKNOWLEDGMENTS
We thank Christopher Silva and Bradley Hernlem for their critical reviews and suggestions and Luisa Cheng for providing reagents and advice. We also thank Anna Bates and Robert Mandrell for the use of their bacterial isolates.
This project was supported by USDA-ARS National Program NP108 (CRIS project 5325-42000-048-00D).
Footnotes
Supplemental material for this article may be found at http://dx.doi.org/10.1128/AAC.00763-15.
REFERENCES
- 1.Scharff RL. 2012. Economic burden from health losses due to foodborne illness in the United States. J Food Protect 75:123–131. doi: 10.4315/0362-028X.JFP-11-058. [DOI] [PubMed] [Google Scholar]
- 2.Gould LH, Bopp C, Strockbine N, Atkinson R, Baselski V, Body B, Carey R, Crandall C, Hurd S, Kaplan R, Neill M, Shea S, Somsel P, Tobin-D'Angelo M, Griffin PM, Gerner-Smidt P. 2009. Recommendations for diagnosis of Shiga toxin-producing Escherichia coli infections by clinical laboratories. MMWR Recomm Rep 58(RR-12):1–14. [PubMed] [Google Scholar]
- 3.Friedrich AW, Bielaszewska M, Zhang WL, Pulz M, Kuczius T, Ammon A, Karch H. 2002. Escherichia coli harboring Shiga toxin 2 gene variants: frequency and association with clinical symptoms. J Infect Dis 185:74–84. doi: 10.1086/338115. [DOI] [PubMed] [Google Scholar]
- 4.Trofa AF, Ueno-Olsen H, Oiwa R, Yoshikawa M. 1999. Dr. Kiyoshi Shiga: discoverer of the dysentery bacillus. Clin Infect Dis 29:1303–1306. [DOI] [PubMed] [Google Scholar]
- 5.Tesh VL, O'Brien AD. 1991. The pathogenic mechanisms of Shiga toxin and the Shiga-like toxins. Mol Microbiol 5:1817–1822. doi: 10.1111/j.1365-2958.1991.tb00805.x. [DOI] [PubMed] [Google Scholar]
- 6.Fraser ME, Fujinaga M, Cherney MM, Melton-Celsa AR, Twiddy EM, O'Brien AD, James MN. 2004. Structure of Shiga toxin type 2 (Stx2) from Escherichia coli O157:H7. J Biol Chem 279:27511–27517. doi: 10.1074/jbc.M401939200. [DOI] [PubMed] [Google Scholar]
- 7.Tesh VL, Burris JA, Owens JW, Gordon VM, Wadolkowski EA, O'Brien AD, Samuel JE. 1993. Comparison of the relative toxicities of Shiga-like toxins type I and type II for mice. Infect Immun 61:3392–3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Scheutz F, Teel LD, Beutin L, Pierard D, Buvens G, Karch H, Mellmann A, Caprioli A, Tozzoli R, Morabito S, Strockbine NA, Melton-Celsa AR, Sanchez M, Persson S, O'Brien AD. 2012. Multicenter evaluation of a sequence-based protocol for subtyping Shiga toxins and standardizing Stx nomenclature. J Clin Microbiol 50:2951–2963. doi: 10.1128/JCM.00860-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Probert WS, McQuaid C, Schrader K. 2014. Isolation and identification of an Enterobacter cloacae strain producing a novel subtype of Shiga toxin type 1. J Clin Microbiol 52:2346–2351. doi: 10.1128/JCM.00338-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Brzuszkiewicz E, Thurmer A, Schuldes J, Leimbach A, Liesegang H, Meyer FD, Boelter J, Petersen H, Gottschalk G, Daniel R. 2011. Genome sequence analyses of two isolates from the recent Escherichia coli outbreak in Germany reveal the emergence of a new pathotype: entero-aggregative-haemorrhagic Escherichia coli (EAHEC). Arch Microbiol 193:883–891. doi: 10.1007/s00203-011-0725-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Paton AW, Paton JC. 1996. Enterobacter cloacae producing a Shiga-like toxin II-related cytotoxin associated with a case of hemolytic-uremic syndrome. J Clin Microbiol 34:463–465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ooka T, Seto K, Kawano K, Kobayashi H, Etoh Y, Ichihara S, Kaneko A, Isobe J, Yamaguchi K, Horikawa K, Gomes TA, Linden A, Bardiau M, Mainil JG, Beutin L, Ogura Y, Hayashi T. 2012. Clinical significance of Escherichia albertii. Emerg Infect Dis 18:488–492. doi: 10.3201/eid1803.111401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Casas V, Sobrepena G, Rodriguez-Mueller B, Ahtye J, Maloy SR. 2011. Bacteriophage-encoded Shiga toxin gene in atypical bacterial host. Gut Pathog 3:10. doi: 10.1186/1757-4749-3-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Delannoy S, Beutin L, Burgos Y, Fach P. 2012. Specific detection of enteroaggregative hemorrhagic Escherichia coli O104:H4 strains by use of the CRISPR locus as a target for a diagnostic real-time PCR. J Clin Microbiol 50:3485–3492. doi: 10.1128/JCM.01656-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Frank C, Werber D, Cramer JP, Askar M, Faber M, an der Heiden M, Bernard H, Fruth A, Prager R, Spode A, Wadl M, Zoufaly A, Jordan S, Kemper MJ, Follin P, Muller L, King LA, Rosner B, Buchholz U, Stark K, Krause G. 2011. Epidemic profile of Shiga-toxin-producing Escherichia coli O104:H4 outbreak in Germany. N Engl J Med 365:1771–1780. doi: 10.1056/NEJMoa1106483. [DOI] [PubMed] [Google Scholar]
- 16.Plunkett G III, Rose DJ, Durfee TJ, Blattner FR. 1999. Sequence of Shiga toxin 2 phage 933W from Escherichia coli O157:H7: Shiga toxin as a phage late-gene product. J Bacteriol 181:1767–1778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wagner PL, Livny J, Neely MN, Acheson DW, Friedman DI, Waldor MK. 2002. Bacteriophage control of Shiga toxin 1 production and release by Escherichia coli. Mol Microbiol 44:957–970. doi: 10.1046/j.1365-2958.2002.02950.x. [DOI] [PubMed] [Google Scholar]
- 18.Singh R, Schroeder CM, Meng J, White DG, McDermott PF, Wagner DD, Yang H, Simjee S, Debroy C, Walker RD, Zhao S. 2005. Identification of antimicrobial resistance and class 1 integrons in Shiga toxin-producing Escherichia coli recovered from humans and food animals. J Antimicrob Chemother 56:216–219. doi: 10.1093/jac/dki161. [DOI] [PubMed] [Google Scholar]
- 19.Canizalez-Roman A, Gonzalez-Nunez E, Vidal JE, Flores-Villasenor H, Leon-Sicairos N. 2013. Prevalence and antibiotic resistance profiles of diarrheagenic Escherichia coli strains isolated from food items in northwestern Mexico. Int J Food Microbiol 164:36–45. doi: 10.1016/j.ijfoodmicro.2013.03.020. [DOI] [PubMed] [Google Scholar]
- 20.Slutsker L, Ries AA, Maloney K, Wells JG, Greene KD, Griffin PM. 1998. A nationwide case-control study of Escherichia coli O157:H7 infection in the United States. J Infect Dis 177:962–966. doi: 10.1086/515258. [DOI] [PubMed] [Google Scholar]
- 21.Wong CS, Jelacic S, Habeeb RL, Watkins SL, Tarr PI. 2000. The risk of the hemolytic-uremic syndrome after antibiotic treatment of Escherichia coli O157:H7 infections. N Engl J Med 342:1930–1936. doi: 10.1056/NEJM200006293422601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Bitzan M, Klemt M, Steffens R, Muller-Wiefel DE. 1993. Differences in verotoxin neutralizing activity of therapeutic immunoglobulins and sera from healthy controls. Infection 21:140–145. doi: 10.1007/BF01710530. [DOI] [PubMed] [Google Scholar]
- 23.Scotland SM, Willshaw GA, Smith HR, Rowe B. 1987. Properties of strains of Escherichia coli belonging to serogroup O157 with special reference to production of Vero cytotoxins VT1 and VT2. Epidemiol Infect 99:613–624. doi: 10.1017/S0950268800066462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Cheng LW, Henderson TD, Patfield S, Stanker LH, He X. 2013. Mouse in vivo neutralization of Escherichia coli Shiga toxin 2 with monoclonal antibodies. Toxins 5:1845–1858. doi: 10.3390/toxins5101845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.He X, McMahon S, Skinner C, Merrill P, Scotcher MC, Stanker LH. 2013. Development and characterization of monoclonal antibodies against Shiga toxin 2 and their application for toxin detection in milk. J Immunol Methods 389:18–28. doi: 10.1016/j.jim.2012.12.005. [DOI] [PubMed] [Google Scholar]
- 26.He X, Patfield S, Hnasko R, Rasooly R, Mandrell RE. 2013. A polyclonal antibody based immunoassay detects seven subtypes of Shiga toxin 2 produced by Escherichia coli in human and environmental samples. PLoS One 8:e76368. doi: 10.1371/journal.pone.0076368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Skinner C, McMahon S, Rasooly R, Carter JM, He X. 2013. Purification and characterization of Shiga toxin 2f, an immunologically unrelated subtype of Shiga toxin 2. PLoS One 8:e59760. doi: 10.1371/journal.pone.0059760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Skinner C, Patfield S, Stanker LH, Fratamico P, He X. 2014. New high-affinity monoclonal antibodies against Shiga toxin 1 facilitate the detection of hybrid Stx1/Stx2 in vivo. PLoS One 9:e99854. doi: 10.1371/journal.pone.0099854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Skinner C, Patfield S, Stanker L, He X. 2013. Development of monoclonal antibodies and immunoassays for sensitive and specific detection of Shiga toxin Stx2f. PLoS One 8:e76563. doi: 10.1371/journal.pone.0076563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Rasooly R, Do PM. 2010. Shiga toxin Stx2 is heat-stable and not inactivated by pasteurization. Int J Food Microbiol 136:290–294. doi: 10.1016/j.ijfoodmicro.2009.10.005. [DOI] [PubMed] [Google Scholar]
- 31.Shimizu T, Ohta Y, Noda M. 2009. Shiga toxin 2 is specifically released from bacterial cells by two different mechanisms. Infect Immun 77:2813–2823. doi: 10.1128/IAI.00060-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.He X, Quinones B, McMahon S, Mandrell RE. 2012. A single-step purification and molecular characterization of functional Shiga toxin 2 variants from pathogenic Escherichia coli. Toxins 4:487–504. doi: 10.3390/toxins4070487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Clinical and Laboratory Standards Institute. 2012. Performance standards for antimicrobial susceptibility testing; 22nd informational supplement, p 44–49. CLSI M100-S22. Clinical and Laboratory Standards Institute, Wayne, PA. [Google Scholar]
- 34.Muniesa M, Hammerl JA, Hertwig S, Appel B, Brussow H. 2012. Shiga toxin-producing Escherichia coli O104:H4: a new challenge for microbiology. Appl Environ Microbiol 78:4065–4073. doi: 10.1128/AEM.00217-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Geerdes-Fenge HF, Lobermann M, Nurnberg M, Fritzsche C, Koball S, Henschel J, Hohn R, Schober HC, Mitzner S, Podbielski A, Reisinger EC. 2013. Ciprofloxacin reduces the risk of hemolytic uremic syndrome in patients with Escherichia coli O104:H4-associated diarrhea. Infection 41:669–673. doi: 10.1007/s15010-012-0387-6. [DOI] [PubMed] [Google Scholar]
- 36.Larson KC, Belliveau PP, Spooner LM. 2011. Tigecycline for the treatment of severe Clostridium difficile infection. Ann Pharmacother 45:1005–1010. doi: 10.1345/aph.1Q080. [DOI] [PubMed] [Google Scholar]
- 37.Silva-Sanchez J, Reyna-Flores F, Velazquez-Meza ME, Rojas-Moreno T, Benitez-Diaz A, Sanchez-Perez A. 2011. In vitro activity of tigecycline against extended-spectrum β-lactamase-producing Enterobacteriaceae and MRSA clinical isolates from Mexico: a multicentric study. Diagn Microbiol Infect Dis 70:270–273. doi: 10.1016/j.diagmicrobio.2010.12.018. [DOI] [PubMed] [Google Scholar]
- 38.Rasheed JK, Kitchel B, Zhu W, Anderson KF, Clark NC, Ferraro MJ, Savard P, Humphries RM, Kallen AJ, Limbago BM. 2013. New Delhi metallo-β-lactamase-producing Enterobacteriaceae, United States. Emerg Infect Dis 19:870–878. doi: 10.3201/eid1906.121515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Du XX, Fu Y, Yu YS. 2013. Tigecycline treatment of infection caused by KPC-producing Escherichia coli in a pediatric patient. Ann Clin Microbiol Antimicrob 12:19. doi: 10.1186/1476-0711-12-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kim HB, Zhang Q, Sun X, Beamer G, Wang Y, Tzipori S. 2014. Beneficial effect of oral tigecycline treatment on Clostridium difficile infection in gnotobiotic piglets. Antimicrob Agents Chemother 58:7560–7564. doi: 10.1128/AAC.03447-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Jitkova Y, Gronda M, Hurren R, Wang X, Goard CA, Jhas B, Schimmer AD. 2014. A novel formulation of tigecycline has enhanced stability and sustained antibacterial and antileukemic activity. PLoS One 9:e95281. doi: 10.1371/journal.pone.0095281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Purdy J, Jouve S, Yan JL, Balter I, Dartois N, Cooper CA, Korth-Bradley J. 2012. Pharmacokinetics and safety profile of tigecycline in children aged 8 to 11 years with selected serious infections: a multicenter, open-label, ascending-dose study. Clin Ther 34:496–507.e1. doi: 10.1016/j.clinthera.2011.12.010. [DOI] [PubMed] [Google Scholar]
- 43.Zhanel GG, Lawson CD, Adam H, Schweizer F, Zelenitsky S, Lagace-Wiens PR, Denisuik A, Rubinstein E, Gin AS, Hoban DJ, Lynch JP III, Karlowsky JA. 2013. Ceftazidime-avibactam: a novel cephalosporin/β-lactamase inhibitor combination. Drugs 73:159–177. doi: 10.1007/s40265-013-0013-7. [DOI] [PubMed] [Google Scholar]
- 44.Eng RH, Smith SM, Fan-Havard P, Ogbara T. 1993. Effect of antibiotics on endotoxin release from Gram-negative bacteria. Diagn Microbiol Infect Dis 16:185–189. doi: 10.1016/0732-8893(93)90109-K. [DOI] [PubMed] [Google Scholar]
- 45.Scholl D, Gebhart D, Williams SR, Bates A, Mandrell R. 2012. Genome sequence of E. coli O104:H4 leads to rapid development of a targeted antimicrobial agent against this emerging pathogen. PLoS One 7:e33637. doi: 10.1371/journal.pone.0033637. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.