Abstract
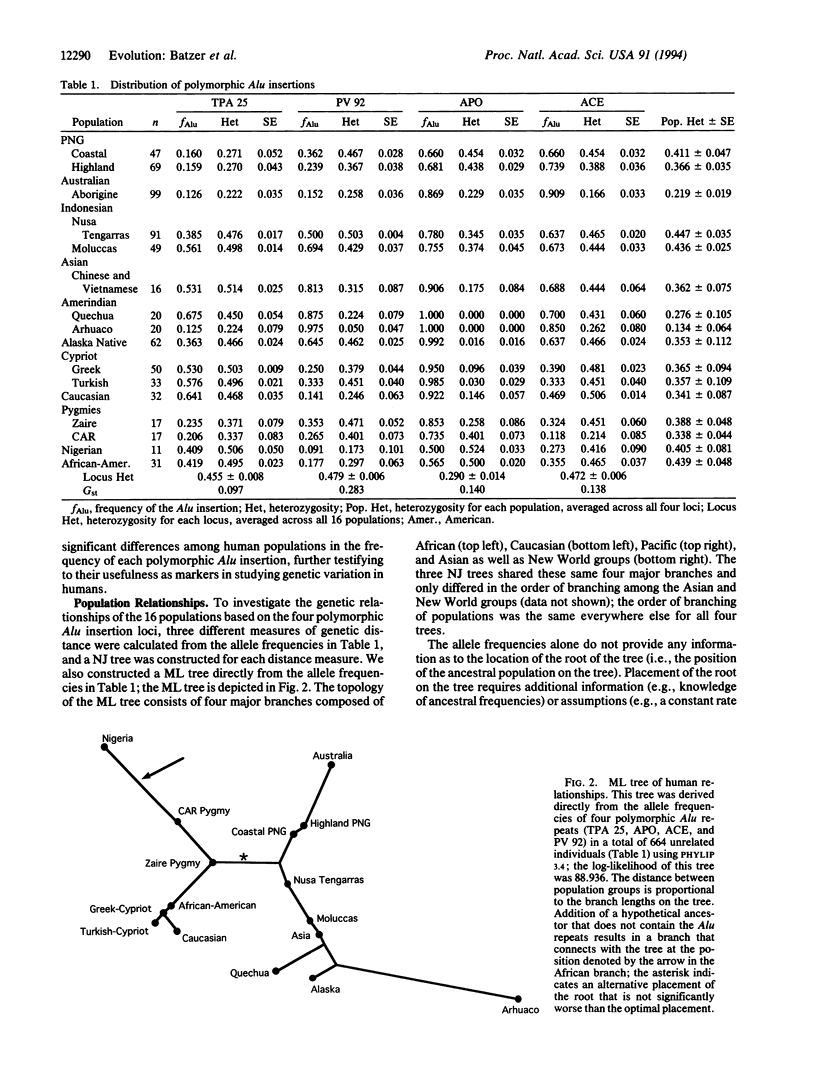
Alu elements are a family of interspersed repeats that have mobilized throughout primate genomes by retroposition from a few "master" genes. Among the 500,000 Alu elements in the human genome are members of the human-specific subfamily that are not fixed in the human species; that is, not all chromosomes carry an Alu element at a particular locus. Four such polymorphic human-specific Alu insertions were analyzed by a rapid, PCR-based assay that uses primers that flank the insertion point to determine genotypes based on the presence or absence of the Alu element. These four polymorphic Alu insertions were shown to be absent from the genomes of a number of nonhuman primates, consistent with their arising as human genetic polymorphisms sometime after the human/African ape divergence. Analysis of 664 unrelated individuals from 16 population groups from around the world revealed substantial levels of variation within population groups and significant genetic differentiation among groups. No significant associations were found among the four loci, consistent with their location on different chromosomes. A maximum-likelihood tree of population relationships showed four major groupings consisting of Africa, Europe, Asia/Americas, and Australia/New Guinea, which is concordant with similar trees based on other loci. A particularly useful feature of the polymorphic Alu insertions is that the ancestral state is known to be the absence of the Alu element, and the presence of the Alu element at a particular chromosomal site reflects a single, unique event in human evolution. A hypothetical ancestral group can then be included in the tree analysis, with the frequency of each insertion set to zero. The ancestral group connected to the maximum-likelihood tree within the African branch, which suggests an African origin of these polymorphic Alu insertions. These data are concordant with other diverse data sets, which lends further support to the recent African origin hypothesis for modern humans. Polymorphic Alu insertions represent a source of genetic variation for studying human population structure and evolution.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bailey A. D., Shen C. K. Sequential insertion of Alu family repeats into specific genomic sites of higher primates. Proc Natl Acad Sci U S A. 1993 Aug 1;90(15):7205–7209. doi: 10.1073/pnas.90.15.7205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Batzer M. A., Deininger P. L. A human-specific subfamily of Alu sequences. Genomics. 1991 Mar;9(3):481–487. doi: 10.1016/0888-7543(91)90414-a. [DOI] [PubMed] [Google Scholar]
- Batzer M. A., Gudi V. A., Mena J. C., Foltz D. W., Herrera R. J., Deininger P. L. Amplification dynamics of human-specific (HS) Alu family members. Nucleic Acids Res. 1991 Jul 11;19(13):3619–3623. doi: 10.1093/nar/19.13.3619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Batzer M. A., Kilroy G. E., Richard P. E., Shaikh T. H., Desselle T. D., Hoppens C. L., Deininger P. L. Structure and variability of recently inserted Alu family members. Nucleic Acids Res. 1990 Dec 11;18(23):6793–6798. doi: 10.1093/nar/18.23.6793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Botstein D., White R. L., Skolnick M., Davis R. W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet. 1980 May;32(3):314–331. [PMC free article] [PubMed] [Google Scholar]
- Bowcock A. M., Bucci C., Hebert J. M., Kidd J. R., Kidd K. K., Friedlaender J. S., Cavalli-Sforza L. L. Study of 47 DNA markers in five populations from four continents. Gene Geogr. 1987 Apr;1(1):47–64. [PubMed] [Google Scholar]
- Bowcock A. M., Kidd J. R., Mountain J. L., Hebert J. M., Carotenuto L., Kidd K. K., Cavalli-Sforza L. L. Drift, admixture, and selection in human evolution: a study with DNA polymorphisms. Proc Natl Acad Sci U S A. 1991 Feb 1;88(3):839–843. doi: 10.1073/pnas.88.3.839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bowcock A. M., Ruiz-Linares A., Tomfohrde J., Minch E., Kidd J. R., Cavalli-Sforza L. L. High resolution of human evolutionary trees with polymorphic microsatellites. Nature. 1994 Mar 31;368(6470):455–457. doi: 10.1038/368455a0. [DOI] [PubMed] [Google Scholar]
- Cann R. L., Stoneking M., Wilson A. C. Mitochondrial DNA and human evolution. Nature. 1987 Jan 1;325(6099):31–36. doi: 10.1038/325031a0. [DOI] [PubMed] [Google Scholar]
- Cavalli-Sforza L. L., Piazza A., Menozzi P., Mountain J. Reconstruction of human evolution: bringing together genetic, archaeological, and linguistic data. Proc Natl Acad Sci U S A. 1988 Aug;85(16):6002–6006. doi: 10.1073/pnas.85.16.6002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chakraborty R., Kamboh M. I., Nwankwo M., Ferrell R. E. Caucasian genes in American blacks: new data. Am J Hum Genet. 1992 Jan;50(1):145–155. [PMC free article] [PubMed] [Google Scholar]
- Edwards M. C., Gibbs R. A. A human dimorphism resulting from loss of an Alu. Genomics. 1992 Nov;14(3):590–597. doi: 10.1016/s0888-7543(05)80156-9. [DOI] [PubMed] [Google Scholar]
- Karathanasis S. K. Apolipoprotein multigene family: tandem organization of human apolipoprotein AI, CIII, and AIV genes. Proc Natl Acad Sci U S A. 1985 Oct;82(19):6374–6378. doi: 10.1073/pnas.82.19.6374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kishino H., Hasegawa M. Evaluation of the maximum likelihood estimate of the evolutionary tree topologies from DNA sequence data, and the branching order in hominoidea. J Mol Evol. 1989 Aug;29(2):170–179. doi: 10.1007/BF02100115. [DOI] [PubMed] [Google Scholar]
- Leeflang E. P., Liu W. M., Hashimoto C., Choudary P. V., Schmid C. W. Phylogenetic evidence for multiple Alu source genes. J Mol Evol. 1992 Jul;35(1):7–16. doi: 10.1007/BF00160256. [DOI] [PubMed] [Google Scholar]
- Matera A. G., Hellmann U., Hintz M. F., Schmid C. W. Recently transposed Alu repeats result from multiple source genes. Nucleic Acids Res. 1990 Oct 25;18(20):6019–6023. doi: 10.1093/nar/18.20.6019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matera A. G., Hellmann U., Schmid C. W. A transpositionally and transcriptionally competent Alu subfamily. Mol Cell Biol. 1990 Oct;10(10):5424–5432. doi: 10.1128/mcb.10.10.5424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merriwether D. A., Clark A. G., Ballinger S. W., Schurr T. G., Soodyall H., Jenkins T., Sherry S. T., Wallace D. C. The structure of human mitochondrial DNA variation. J Mol Evol. 1991 Dec;33(6):543–555. doi: 10.1007/BF02102807. [DOI] [PubMed] [Google Scholar]
- Muratani K., Hada T., Yamamoto Y., Kaneko T., Shigeto Y., Ohue T., Furuyama J., Higashino K. Inactivation of the cholinesterase gene by Alu insertion: possible mechanism for human gene transposition. Proc Natl Acad Sci U S A. 1991 Dec 15;88(24):11315–11319. doi: 10.1073/pnas.88.24.11315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakamura Y., Leppert M., O'Connell P., Wolff R., Holm T., Culver M., Martin C., Fujimoto E., Hoff M., Kumlin E. Variable number of tandem repeat (VNTR) markers for human gene mapping. Science. 1987 Mar 27;235(4796):1616–1622. doi: 10.1126/science.3029872. [DOI] [PubMed] [Google Scholar]
- Nei M., Roychoudhury A. K. Evolutionary relationships of human populations on a global scale. Mol Biol Evol. 1993 Sep;10(5):927–943. doi: 10.1093/oxfordjournals.molbev.a040059. [DOI] [PubMed] [Google Scholar]
- Newman W. P., Middaugh J. P., Propst M. T., Rogers D. R. Atherosclerosis in Alaska Natives and non-natives. Lancet. 1993 Apr 24;341(8852):1056–1057. doi: 10.1016/0140-6736(93)92413-n. [DOI] [PubMed] [Google Scholar]
- Perna N. T., Batzer M. A., Deininger P. L., Stoneking M. Alu insertion polymorphism: a new type of marker for human population studies. Hum Biol. 1992 Oct;64(5):641–648. [PubMed] [Google Scholar]
- Reynolds J., Weir B. S., Cockerham C. C. Estimation of the coancestry coefficient: basis for a short-term genetic distance. Genetics. 1983 Nov;105(3):767–779. doi: 10.1093/genetics/105.3.767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saitou N., Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987 Jul;4(4):406–425. doi: 10.1093/oxfordjournals.molbev.a040454. [DOI] [PubMed] [Google Scholar]
- Sawada I., Schmid C. W. Primate evolution of the alpha-globin gene cluster and its Alu-like repeats. J Mol Biol. 1986 Dec 20;192(4):693–709. doi: 10.1016/0022-2836(86)90022-7. [DOI] [PubMed] [Google Scholar]
- Schmid C., Maraia R. Transcriptional regulation and transpositional selection of active SINE sequences. Curr Opin Genet Dev. 1992 Dec;2(6):874–882. doi: 10.1016/s0959-437x(05)80110-8. [DOI] [PubMed] [Google Scholar]
- Slagel V., Flemington E., Traina-Dorge V., Bradshaw H., Deininger P. Clustering and subfamily relationships of the Alu family in the human genome. Mol Biol Evol. 1987 Jan;4(1):19–29. doi: 10.1093/oxfordjournals.molbev.a040422. [DOI] [PubMed] [Google Scholar]
- Tiret L., Rigat B., Visvikis S., Breda C., Corvol P., Cambien F., Soubrier F. Evidence, from combined segregation and linkage analysis, that a variant of the angiotensin I-converting enzyme (ACE) gene controls plasma ACE levels. Am J Hum Genet. 1992 Jul;51(1):197–205. [PMC free article] [PubMed] [Google Scholar]
- Vigilant L., Stoneking M., Harpending H., Hawkes K., Wilson A. C. African populations and the evolution of human mitochondrial DNA. Science. 1991 Sep 27;253(5027):1503–1507. doi: 10.1126/science.1840702. [DOI] [PubMed] [Google Scholar]
- Wainscoat J. S., Hill A. V., Boyce A. L., Flint J., Hernandez M., Thein S. L., Old J. M., Lynch J. R., Falusi A. G., Weatherall D. J. Evolutionary relationships of human populations from an analysis of nuclear DNA polymorphisms. Nature. 1986 Feb 6;319(6053):491–493. doi: 10.1038/319491a0. [DOI] [PubMed] [Google Scholar]
- Wallace M. R., Andersen L. B., Saulino A. M., Gregory P. E., Glover T. W., Collins F. S. A de novo Alu insertion results in neurofibromatosis type 1. Nature. 1991 Oct 31;353(6347):864–866. doi: 10.1038/353864a0. [DOI] [PubMed] [Google Scholar]
- Willard C., Nguyen H. T., Schmid C. W. Existence of at least three distinct Alu subfamilies. J Mol Evol. 1987;26(3):180–186. doi: 10.1007/BF02099850. [DOI] [PubMed] [Google Scholar]
- Yang-Feng T. L., Opdenakker G., Volckaert G., Francke U. Human tissue-type plasminogen activator gene located near chromosomal breakpoint in myeloproliferative disorder. Am J Hum Genet. 1986 Jul;39(1):79–87. [PMC free article] [PubMed] [Google Scholar]




