Abstract
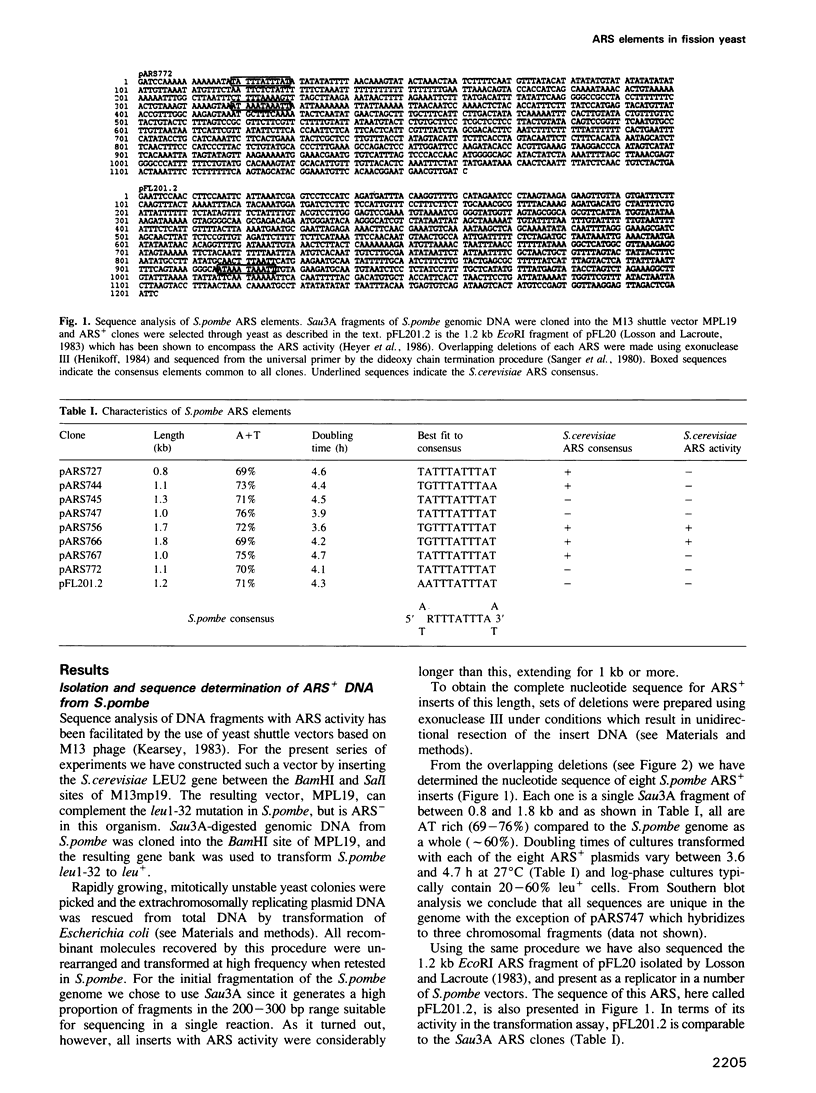
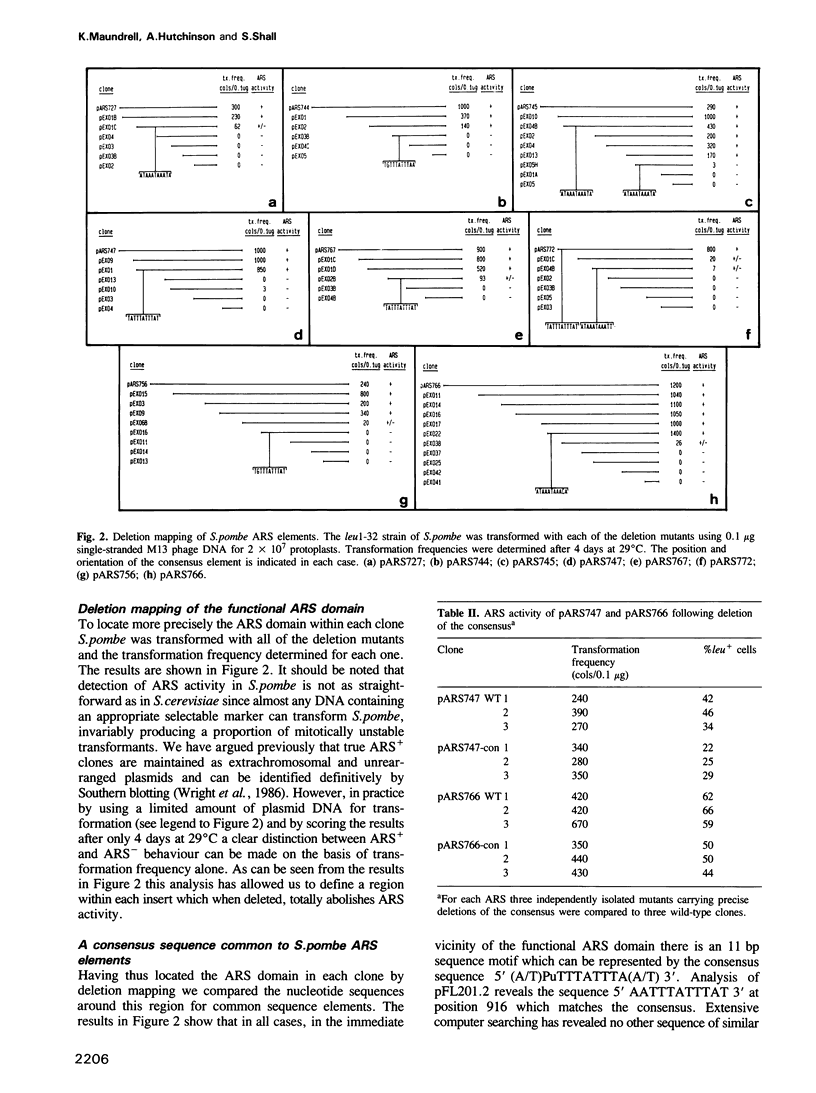
Chromosomal DNA of Schizosaccharomyces pombe contains sequences with properties analogous to ARS elements of Saccharomyces cerevisiae. Following Sau3A fragmentation of the S. pombe genome we have recovered a number of such fragments in an M13-based shuttle vector, suitable for subsequent sequence analysis. The complete nucleotide sequence has been obtained for eight ARS+ inserts derived from the Sau3A cloning and for the ARS present in pFL20 isolated previously by Losson and Lacroute (Cell, 32, 371-377, 1983). The Sau3A clones are single fragments between 0.8 and 1.8 kb. No ARS+ clones smaller than this were recovered even though the average size Sau3A fragment in S. pombe is approximately 200-300 bp. The sequence analysis revealed that all clones are AT-rich (69-75% A + T residues), and all contain a particularly AT-rich 11 bp core element represented by the consensus sequence 5' (A/T)PuTT-TATTTA(A/T) 3'. Deletion mapping indicates that the consensus in all cases is in the vicinity of a functional ARS domain. However precise excision of the consensus by in vitro mutagenesis has little effect on ARS activity as judged by the transformation assay. We argue that the association of the consensus with the ARS domain occurs too reproducibly to be explained by chance alone. We suggest that although it may not be essential for the extrachromosomal maintenance of plasmids in S. pombe, the consensus does have a function in situ in the chromosome and thus is always present as a cryptic sequence in the isolated ARS element.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Beach D., Piper M., Shall S. Isolation of chromosomal origins of replication in yeast. Nature. 1980 Mar 13;284(5752):185–187. doi: 10.1038/284185a0. [DOI] [PubMed] [Google Scholar]
- Bouton A. H., Stirling V. B., Smith M. M. Analysis of DNA sequences homologous with the ARS core consensus in Saccharomyces cerevisiae. Yeast. 1987 Jun;3(2):107–115. doi: 10.1002/yea.320030207. [DOI] [PubMed] [Google Scholar]
- Brewer B. J., Fangman W. L. The localization of replication origins on ARS plasmids in S. cerevisiae. Cell. 1987 Nov 6;51(3):463–471. doi: 10.1016/0092-8674(87)90642-8. [DOI] [PubMed] [Google Scholar]
- Broach J. R., Li Y. Y., Feldman J., Jayaram M., Abraham J., Nasmyth K. A., Hicks J. B. Localization and sequence analysis of yeast origins of DNA replication. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 2):1165–1173. doi: 10.1101/sqb.1983.047.01.132. [DOI] [PubMed] [Google Scholar]
- Celniker S. E., Sweder K., Srienc F., Bailey J. E., Campbell J. L. Deletion mutations affecting autonomously replicating sequence ARS1 of Saccharomyces cerevisiae. Mol Cell Biol. 1984 Nov;4(11):2455–2466. doi: 10.1128/mcb.4.11.2455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan C. S., Tye B. K. Autonomously replicating sequences in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1980 Nov;77(11):6329–6333. doi: 10.1073/pnas.77.11.6329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. "A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity". Addendum. Anal Biochem. 1984 Feb;137(1):266–267. doi: 10.1016/0003-2697(84)90381-6. [DOI] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene. 1984 Jun;28(3):351–359. doi: 10.1016/0378-1119(84)90153-7. [DOI] [PubMed] [Google Scholar]
- Heyer W. D., Sipiczki M., Kohli J. Replicating plasmids in Schizosaccharomyces pombe: improvement of symmetric segregation by a new genetic element. Mol Cell Biol. 1986 Jan;6(1):80–89. doi: 10.1128/mcb.6.1.80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hinnen A., Hicks J. B., Fink G. R. Transformation of yeast. Proc Natl Acad Sci U S A. 1978 Apr;75(4):1929–1933. doi: 10.1073/pnas.75.4.1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huberman J. A., Spotila L. D., Nawotka K. A., el-Assouli S. M., Davis L. R. The in vivo replication origin of the yeast 2 microns plasmid. Cell. 1987 Nov 6;51(3):473–481. doi: 10.1016/0092-8674(87)90643-x. [DOI] [PubMed] [Google Scholar]
- Kearsey S. Analysis of sequences conferring autonomous replication in baker's yeast. EMBO J. 1983;2(9):1571–1575. doi: 10.1002/j.1460-2075.1983.tb01626.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kearsey S. Structural requirements for the function of a yeast chromosomal replicator. Cell. 1984 May;37(1):299–307. doi: 10.1016/0092-8674(84)90326-x. [DOI] [PubMed] [Google Scholar]
- Losson R., Lacroute F. Plasmids carrying the yeast OMP decarboxylase structural and regulatory genes: transcription regulation in a foreign environment. Cell. 1983 Feb;32(2):371–377. doi: 10.1016/0092-8674(83)90456-7. [DOI] [PubMed] [Google Scholar]
- Maundrell K., Wright A. P., Piper M., Shall S. Evaluation of heterologous ARS activity in S. cerevisiae using cloned DNA from S. pombe. Nucleic Acids Res. 1985 May 24;13(10):3711–3722. doi: 10.1093/nar/13.10.3711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Messing J. New M13 vectors for cloning. Methods Enzymol. 1983;101:20–78. doi: 10.1016/0076-6879(83)01005-8. [DOI] [PubMed] [Google Scholar]
- Murray A. W., Szostak J. W. Pedigree analysis of plasmid segregation in yeast. Cell. 1983 Oct;34(3):961–970. doi: 10.1016/0092-8674(83)90553-6. [DOI] [PubMed] [Google Scholar]
- Nurse P. Genetic control of cell size at cell division in yeast. Nature. 1975 Aug 14;256(5518):547–551. doi: 10.1038/256547a0. [DOI] [PubMed] [Google Scholar]
- Palzkill T. G., Oliver S. G., Newlon C. S. DNA sequence analysis of ARS elements from chromosome III of Saccharomyces cerevisiae: identification of a new conserved sequence. Nucleic Acids Res. 1986 Aug 11;14(15):6247–6264. doi: 10.1093/nar/14.15.6247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakaguchi J., Yamamoto M. Cloned ural locus of Schizosaccharomyces pombe propagates autonomously in this yeast assuming a polymeric form. Proc Natl Acad Sci U S A. 1982 Dec;79(24):7819–7823. doi: 10.1073/pnas.79.24.7819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Coulson A. R., Barrell B. G., Smith A. J., Roe B. A. Cloning in single-stranded bacteriophage as an aid to rapid DNA sequencing. J Mol Biol. 1980 Oct 25;143(2):161–178. doi: 10.1016/0022-2836(80)90196-5. [DOI] [PubMed] [Google Scholar]
- Shore D., Stillman D. J., Brand A. H., Nasmyth K. A. Identification of silencer binding proteins from yeast: possible roles in SIR control and DNA replication. EMBO J. 1987 Feb;6(2):461–467. doi: 10.1002/j.1460-2075.1987.tb04776.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stinchcomb D. T., Struhl K., Davis R. W. Isolation and characterisation of a yeast chromosomal replicator. Nature. 1979 Nov 1;282(5734):39–43. doi: 10.1038/282039a0. [DOI] [PubMed] [Google Scholar]
- Strich R., Woontner M., Scott J. F. Mutations in ARS1 increase the rate of simple loss of plasmids in Saccharomyces cerevisiae. Yeast. 1986 Sep;2(3):169–178. doi: 10.1002/yea.320020305. [DOI] [PubMed] [Google Scholar]
- Umek R. M., Kowalski D. The ease of DNA unwinding as a determinant of initiation at yeast replication origins. Cell. 1988 Feb 26;52(4):559–567. doi: 10.1016/0092-8674(88)90469-2. [DOI] [PubMed] [Google Scholar]
- Williamson D. H. The yeast ARS element, six years on: a progress report. Yeast. 1985 Sep;1(1):1–14. doi: 10.1002/yea.320010102. [DOI] [PubMed] [Google Scholar]
- Wright A. P., Maundrell K., Shall S. Transformation of Schizosaccharomyces pombe by non-homologous, unstable integration of plasmids in the genome. Curr Genet. 1986;10(7):503–508. doi: 10.1007/BF00447383. [DOI] [PubMed] [Google Scholar]
- Zoller M. J., Smith M. Oligonucleotide-directed mutagenesis using M13-derived vectors: an efficient and general procedure for the production of point mutations in any fragment of DNA. Nucleic Acids Res. 1982 Oct 25;10(20):6487–6500. doi: 10.1093/nar/10.20.6487. [DOI] [PMC free article] [PubMed] [Google Scholar]




