Abstract
A previously unrecognized enzyme acting on damaged termini in DNA is present in Escherichia coli. The enzyme catalyses the hydrolytic release of 2-deoxyribose-5-phosphate from single-strand interruptions in DNA with a base-free residue on the 5' side. The partly purified protein appears to be free from endonuclease activity for apurinic/apyrimidinic sites, exonuclease activity and DNA 5'-phosphatase activity. The enzyme has a mol. wt of approximately 50,000-55,000 and has been termed DNA deoxyribophosphodiesterase (dRpase). The protein presumably is active in DNA excision repair to remove a sugar-phosphate residue from an endonucleolytically incised apurinic/apyrimidinic site, prior to gap filling and ligation.
Full text
PDF

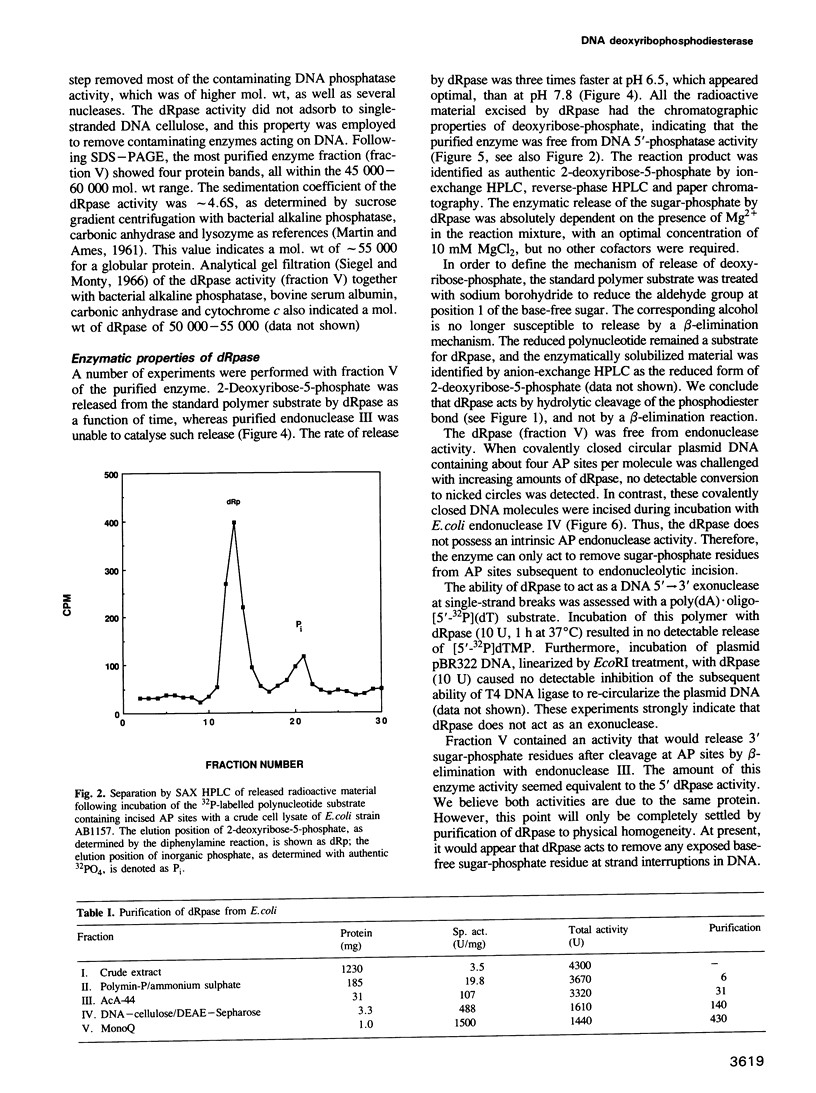
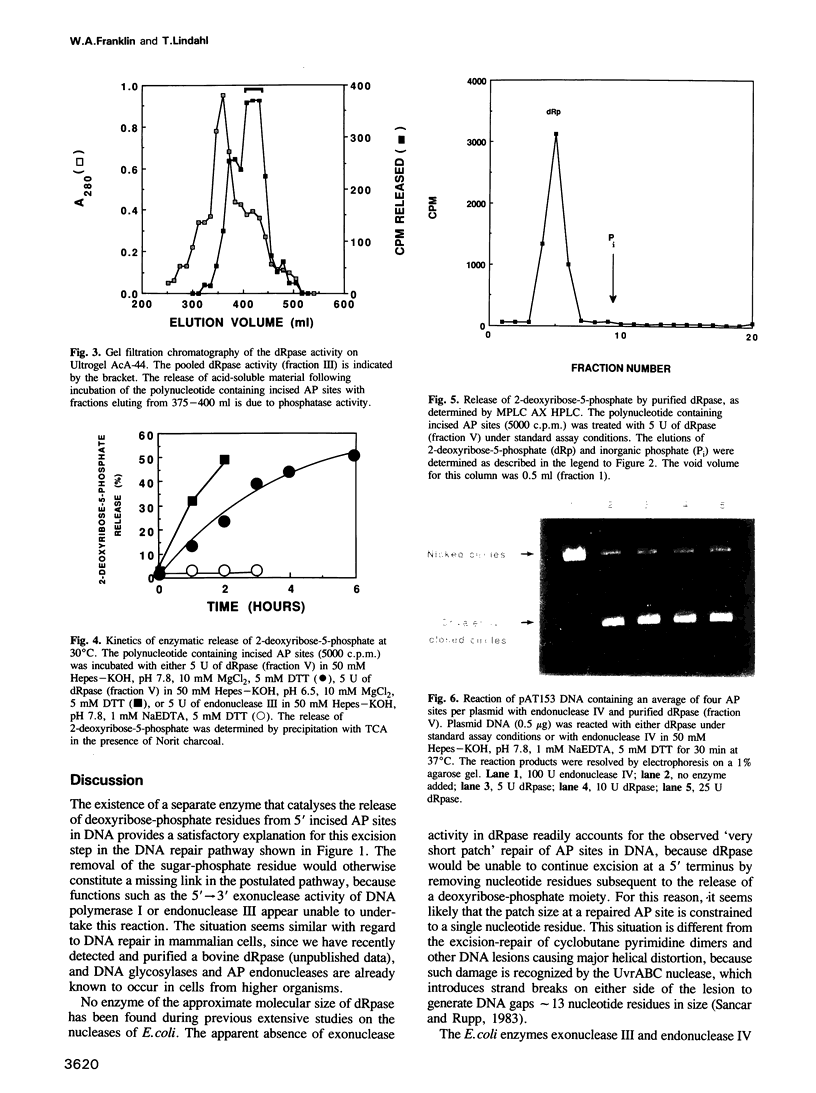


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arrand J. E., Willis A. E., Goldsmith I., Lindahl T. Different substrate specificities of the two DNA ligases of mammalian cells. J Biol Chem. 1986 Jul 15;261(20):9079–9082. [PubMed] [Google Scholar]
- BURTON K. A study of the conditions and mechanism of the diphenylamine reaction for the colorimetric estimation of deoxyribonucleic acid. Biochem J. 1956 Feb;62(2):315–323. doi: 10.1042/bj0620315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bailly V., Verly W. G. Escherichia coli endonuclease III is not an endonuclease but a beta-elimination catalyst. Biochem J. 1987 Mar 1;242(2):565–572. doi: 10.1042/bj2420565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Breimer L. H., Lindahl T. DNA glycosylase activities for thymine residues damaged by ring saturation, fragmentation, or ring contraction are functions of endonuclease III in Escherichia coli. J Biol Chem. 1984 May 10;259(9):5543–5548. [PubMed] [Google Scholar]
- Chan E., Weiss B. Endonuclease IV of Escherichia coli is induced by paraquat. Proc Natl Acad Sci U S A. 1987 May;84(10):3189–3193. doi: 10.1073/pnas.84.10.3189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chase J. W., Richardson C. C. Exonuclease VII of Escherichia coli. Purification and properties. J Biol Chem. 1974 Jul 25;249(14):4545–4552. [PubMed] [Google Scholar]
- Clements J. E., Rogers S. G., Weiss B. A DNase for apurinic/apyrimidinic sites associated with exonuclease III of Hemophilus influenzae. J Biol Chem. 1978 May 10;253(9):2990–2999. [PubMed] [Google Scholar]
- Cunningham R. P., Saporito S. M., Spitzer S. G., Weiss B. Endonuclease IV (nfo) mutant of Escherichia coli. J Bacteriol. 1986 Dec;168(3):1120–1127. doi: 10.1128/jb.168.3.1120-1127.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cunningham R. P., Weiss B. Endonuclease III (nth) mutants of Escherichia coli. Proc Natl Acad Sci U S A. 1985 Jan;82(2):474–478. doi: 10.1073/pnas.82.2.474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gossard F., Verly W. G. Properties of the main endonuclease specific for apurinic sites of Escherichia coli (endonuclease VI). Mechanism of apurinic site excision from DNA. Eur J Biochem. 1978 Jan 16;82(2):321–332. doi: 10.1111/j.1432-1033.1978.tb12026.x. [DOI] [PubMed] [Google Scholar]
- Grafstrom R. H., Shaper N. L., Grossman L. Human placental apurinic/apyrimidinic endonuclease. Mechanism of action. J Biol Chem. 1982 Nov 25;257(22):13459–13464. [PubMed] [Google Scholar]
- Gross J., Gross M. Genetic analysis of an E. coli strain with a mutation affecting DNA polymerase. Nature. 1969 Dec 20;224(5225):1166–1168. doi: 10.1038/2241166a0. [DOI] [PubMed] [Google Scholar]
- Kataoka H., Sekiguchi M. Are purine bases enzymatically inserted into depurinated DNA in Escherichia coli? J Biochem. 1982 Sep;92(3):971–973. doi: 10.1093/oxfordjournals.jbchem.a134014. [DOI] [PubMed] [Google Scholar]
- Kim J., Linn S. The mechanisms of action of E. coli endonuclease III and T4 UV endonuclease (endonuclease V) at AP sites. Nucleic Acids Res. 1988 Feb 11;16(3):1135–1141. doi: 10.1093/nar/16.3.1135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LEHMAN I. R., ROUSSOS G. G., PRATT E. A. The deoxyribonucleases of Escherichia coli. II. Purification and properties of a ribonucleic acid-inhibitable endonuclease. J Biol Chem. 1962 Mar;237:819–828. [PubMed] [Google Scholar]
- Levin J. D., Johnson A. W., Demple B. Homogeneous Escherichia coli endonuclease IV. Characterization of an enzyme that recognizes oxidative damage in DNA. J Biol Chem. 1988 Jun 15;263(17):8066–8071. [PubMed] [Google Scholar]
- Lindahl T., Andersson A. Rate of chain breakage at apurinic sites in double-stranded deoxyribonucleic acid. Biochemistry. 1972 Sep 12;11(19):3618–3623. doi: 10.1021/bi00769a019. [DOI] [PubMed] [Google Scholar]
- Lindahl T. DNA repair enzymes. Annu Rev Biochem. 1982;51:61–87. doi: 10.1146/annurev.bi.51.070182.000425. [DOI] [PubMed] [Google Scholar]
- Lindahl T., Ljungquist S., Siegert W., Nyberg B., Sperens B. DNA N-glycosidases: properties of uracil-DNA glycosidase from Escherichia coli. J Biol Chem. 1977 May 25;252(10):3286–3294. [PubMed] [Google Scholar]
- Ljungquist S. A new endonuclease from Escherichia coli acting at apurinic sites in DNA. J Biol Chem. 1977 May 10;252(9):2808–2814. [PubMed] [Google Scholar]
- Ljungquist S., Lindahl T., Howard-Flanders P. Methyl methane sulfonate-sensitive mutant of Escherichia coli deficient in an endonuclease specific for apurinic sites in deoxyribonucleic acid. J Bacteriol. 1976 May;126(2):646–653. doi: 10.1128/jb.126.2.646-653.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loeb L. A., Preston B. D. Mutagenesis by apurinic/apyrimidinic sites. Annu Rev Genet. 1986;20:201–230. doi: 10.1146/annurev.ge.20.120186.001221. [DOI] [PubMed] [Google Scholar]
- MARTIN R. G., AMES B. N. A method for determining the sedimentation behavior of enzymes: application to protein mixtures. J Biol Chem. 1961 May;236:1372–1379. [PubMed] [Google Scholar]
- Mackay V., Linn S. Molecular structure of exonuclease I from Escherichia coli B. Biochim Biophys Acta. 1974 Apr 27;349(1):131–134. doi: 10.1016/0005-2787(74)90016-1. [DOI] [PubMed] [Google Scholar]
- Mosbaugh D. W., Linn S. Characterization of the action of Escherichia coli DNA polymerase I at incisions produced by repair endodeoxyribonucleases. J Biol Chem. 1982 Jan 10;257(1):575–583. [PubMed] [Google Scholar]
- SCHACHMAN H. K., ADLER J., RADDING C. M., LEHMAN I. R., KORNBERG A. Enzymatic synthesis of deoxyribonucleic acid. VII. Synthesis of a polymer of deoxyadenylate and deoxythymidylate. J Biol Chem. 1960 Nov;235:3242–3249. [PubMed] [Google Scholar]
- Sancar A., Rupp W. D. A novel repair enzyme: UVRABC excision nuclease of Escherichia coli cuts a DNA strand on both sides of the damaged region. Cell. 1983 May;33(1):249–260. doi: 10.1016/0092-8674(83)90354-9. [DOI] [PubMed] [Google Scholar]
- Siegel L. M., Monty K. J. Determination of molecular weights and frictional ratios of proteins in impure systems by use of gel filtration and density gradient centrifugation. Application to crude preparations of sulfite and hydroxylamine reductases. Biochim Biophys Acta. 1966 Feb 7;112(2):346–362. doi: 10.1016/0926-6585(66)90333-5. [DOI] [PubMed] [Google Scholar]
- Weiss B., Grossman L. Phosphodiesterases involved in DNA repair. Adv Enzymol Relat Areas Mol Biol. 1987;60:1–34. doi: 10.1002/9780470123065.ch1. [DOI] [PubMed] [Google Scholar]




