Abstract
To identify the relationship between selenium intake and breast cancer progression, SeMet supplement was applied to 4T1 murine mammary carcinoma for 28 days and the effect of SeMet on tumor growth was assessed. Combined with SXRF mapping, trace metal distribution in tumor tissues in relation to selenium-associated protein expressions and cellular transcription factors, HIF-1α accumulation and its targeted molecule expressions involved in tumor progression, were further investigated using immunohistochemical staining. Our results showed that tumor growth was inhibited significantly and tumor cells apoptosis was promoted after SeMet supplement. High Se intakes in tumor tissues overlaid with high iron distribution were observed by SXRF mapping. SeMet supplement increased SBP1 expression and decreased GPx1 expression, and greatly inhibited VEGF expression and tumor immune suppression cells accumulation in tumor tissues. In conclusion, SeMet-mediated anti-cancer effect was likely to inhibit tumor growth through high expression of SBP1 inhibiting GPx1 activity and HIF-1α, which could be used as chemopreventive strategies in breast cancer.
Keywords: SeMet, synchrotron X-ray radiation fluorescence mapping (SXRF), glutathione peroxidase (GPx-1), selenium-binding protein 1 (SBP-1), hypoxia inducible factors 1 (HIF-1α), breast cancer
Introduction
Selenium is a very important micronutrient. Dietary Se supplementation could increase intracellular Se levels to produce useful ‘bioactive’ intermediates and selenoproteins. It is clear that organic forms of selenium are more efficient utilized than inorganic forms, and effective chemoprevention for a number of cancers [1]. Organic selenium compounds such as selenomethionine (SeMet) has been shown less cellular toxicity than sodium selenite (inorganic forms), and is more useful than sodium selenite in chemoprevention [2]. So SeMet is currently being used as the Se intervention agent in the ongoing SELECT (The Selenium and Vitamin E Cancer Prevention Trial) in the USA.
Selenium could exist in the forms of specific selenoproteins, and nonspecific selenium-binding proteins by binding to SBP1 in the body [3]. One of these is a family consisting of 25 human Se-containing proteins in which selenocysteine is incorporated co-translationally in the selenoprotein mRNA [4]. Among the best studied of these is the ubiquitously expressed cytosolic glutathione peroxidase (GPx-1). Up-regulated GPx-1 in carcinoma tissues at both the protein and mRNA levels has been reported to result in protection against DNA damage [5], which are also associated with increased risks of cancer [6-8]. The other class of selenoprotein is typified by selenium-binding protein 1 (SBP-1) which does not contain selenocysteine [9]. Low levels of SBP1 in the cancer were associated with poor clinical outcome [10-12]. In addition, dietary selenium had been shown to increase SBP1 expression. The close association of SBP1 with ovarian cancer suggested a possible involvement of SBP1 in the pathway of selenium-mediated anticancer activity [13]. Thus, SBP1 may mediate the effect of selenium on tumor inhibition. Therefore, SBP1 and GPx1 are identified to opposite potential mediators of the chemopreventive effects of selenium. Increasing the levels of SBP1 in either colon-derived HCT116 cells or MCF-7 breast carcinoma cells resulted in a consequential decline in GPx1 enzyme activity [14].
There is increasing evidence that selenium exhibits anticarcinogenic properties [15]. Currently, there are convincing epidemiological data showing a significant inverse relationship between selenium intake and some cancer risk, i.e. an inverse association between selenium exposure and prostate cancer risk, and possible reduction in lung cancer risk as well [16]. However, most of studies about potential anti-tumor efficacy of selenium were investigated in cell lines in vitro, only few studies in vivo. In particular, its association with breast cancer remains a controversial issue, although low selenium level was found to increase breast cancer risk [17]. Thus, the role of selenium in breast cancer is worthy for further investigation, especially in constructing chemopreventive strategies.
Until now, very little work has been conducted to determine the concentration and retention of selenium in tumor and correlation between selenium concentration and selenium related proteins after selenium supplement. In order to improve insight into the mechanism of selenium in tumor progression, it is essential to study the selenium distributions in tumor tissues and concentrations accompanying expression of related proteins after selenium supplement, which will allow us to better understand the role of selenium on tumor progression events. Of many methods suitable for detection of trace metals distribution in biological tissues, the most preferred is multi-elemental and highly sensitive imaging technique such as synchrotron radiation X-ray fluorescence mapping (SXRF). SXRF is the most direct and sensitive tool for metals imaging in cells and tissue, it could be used to image and quantify the distribution of metals with high spatial resolution in sub-microns [18].
In this work, synchrotron radiation X-ray fluorescence mapping was used to detect selected trace elements distribution after SeMet supplement for 4 weeks, such as iron, copper, zinc and selenium in 4T1 murine mammary carcinoma tissues, a common animal model of human metastatic breast cancer with highly invasive and poorly immunogenic characters [19]. Correspondingly, expressions of selenium related proteins, such as SBP1, GPx1, and cellular transcription factors, HIF-1α and its downstream targeted molecules involved in tumor progression were also analyzed using immunohistochemical staining. Further, correlations were investigated among selenium, selenium related proteins and highly related factors involved in tumor progression. The results could provide new insights into the possible mechanisms of the anticancer properties of SeMet.
Materials and methods
Animal model
All experimental animals were handled following the guidelines provided by the Animal Welfare Committee at the Shanghai Sixth People’s Hospital, affiliated to Shanghai Jiao Tong University. Female Balb/c mice (20+2 g, 6 weeks) were obtained from Animal Laboratory of Shanghai Medical College, China and were housed in the isolated cages with sterile food and water, with a 12-h light/dark cycle. The murine mammary carcinoma 4T1 cells were kindly provided by Shanghai First People’s Hospital, China and maintained in DMEM medium (Hyclone SH30243.01B) supplemented penicillin streptomycin and fetal bovine serum (GIBCO). To prepare tumor-bearing mice, approximately 1×106 cells were injected subcutaneously.
SeMet supplement
Seleno-L-Methionine (SeMet) (Sigma, S3132) was dissolved in sterile saline to prepare for the solution in the concentration of 1 mg/ml. After tumor inoculation, SeMet was administered orally at a dose of 0.2 mg/mouse/day using irrigation stomach needle for 4 weeks. PBS supplement was performed in the control group.
Tumor measurement
Tumor sizes were measured every 3 days starting in one week after tumor inoculation. The length, width and height of tumors were measured with the aid of vernier caliper. Tumor volume (cm3) was estimated using the formula: tumor volume = π/6 (length*width*height).
Preparation of cryosections
On the 28th day after tumor inoculation, tumors were removed and fixed with O.C.T compound at -80°C immediately. The frozen tumor tissues were sliced into 10 μm in thickness at -20°C (LEICA CM1900, Germany), and mounted on the 3525 Ultralene XRF film (SPEX CertiPrep, Metuchen, NJ) and air-dried. In addition, several adjacent sections of the same thickness were placed on microscopic slides, and used for immunohistochemical and immunofluorescence staining. Slides were stored frozen until usage.
Synchrotron radiation X-ray fluorescence imaging
Synchrotron radiation X-ray fluorescence mapping was carried out at the BL15U beamline of Shanghai Synchrotron Radiation Facilities (SSRF), China. X-rays of 14 keV energy were focused to a 100 micron-diameter spot on the specimen. X-ray fluorescence emission was collected by an energy-dispersive Li-ion drifted detector [seven-element Si (Li) detector, E2V]. X-ray fluorescence emission spectra were collected for 1.5 s.
Detection of apoptosis
Apoptosis was detected using the ApopTag Plus Peroxidase In Situ Apoptosis Detection Kit (Millipore). Terminal deoxynucleotidyl transferase mediated dUTP nick end labeling (TUNEL) is a method of choice for rapid identification and quantification of the apoptotic cell fraction in tissues. Cryosections were fixed in 4% paraformaldehyde. TdT enzyme was then added for 1 hour at 37°C to label fragmented DNA ends with digoxigenin nucleotides. Anti-digoxigenin antibody conjugated to a peroxidase reporter molecule was then added. Methyl green was used for counterstain. The slides were observed using optical microscopy.
Immunohistochemical staining
For IHC staining, specimens were dried at room temperature for approximately 1h and then fixed in cold acetone for 10 min at 4°C. 0.1% Triton X-100 were used for permeabilization and 0.3% H2O2 in methanol for peroxidase quenched. Then the specimens were blocked by 5% Bovine Serum Albumin (BSA) for 30 min. In order to investigate the changes of selenium related proteins (SBP1 and GPx1), HIF-1α, VEGF, CD31 in the control and SeMet groups, the primary antibodies which include anti-SBP1 (Abcam, 1:120) and anti-GPx1 (Abcam, 1:500), anti-HIF-1α (Abcam, 1:1000) anti-VEGF (Abcam, 1:100), and anti-CD31 (Abcam 1:50) were applied respectively overnight at 4°C. Subsequently, iotinylated secondary antibody, streptavidin-HRP and diaminobenzidine (DAB) peroxidase substrate were added. Finally nuclei were counterstained with hematoxylin.
To analyze SBP1 and GPx1 expression in the control group and SeMet groups, the whole tissue was observed using upright microscope (10×) to overview stained brown positive area of SBP1 and GPx1. Quantitative analysis of positive areas of proteins was used by the Image-Pro Plus software. 10 randomly selected vision fields were analyzed per mouse using microscopy.
Immunofluorescence staining
For immunofluorescence staining, slides were first fixed 10 minutes in cold acetone, washed 3 times for 2 minutes in PBS, and permeabilized with 0.1% Triton X-100. Then, slides were blocked for 30 minutes in 10% Bovine Serum Albumin (BSA). Antibodies were applied overnight at 4°C and include: APC-conjugated anti-mouse CD4 (BioLegend, 1:200), Alexa Fluor®488-conjugated anti-mouse CD25 (BioLegend, 1:200), FITC-conjugated anti-mouse CD11b (BioLegend, 1:200) and PE-conjugated anti-mouse Gr-1 (BioLegend, 1:400). VECTASHIELD Mounting Medium with DAPI (Vector Laboratories) was used as a counterstain. Slides were viewed using a confocal microscope (Leica TCS SP5). Quantitative analysis of positive cells was used by the Image-Pro Plus software. 10 randomly selected vision fields per mouse were analyzed.
Statistical analysis
Data were analyzed using Student’s t-test and p values of ≤0.05 were considered statistically significant by SPSS software. Results were displayed as mean ± SEM.
Results
Effect of SeMet supplement on tumor growth
To investigate anti-tumor effect induced by the SeMet, 4T1 tumor volume in the control and SeMet groups were measured along with time. Three mice were included in the control and SeMet group, respectively. As shown in Figure 1, tumor volume increased quickly with time in the control group, while its growth was obviously slower in the SeMet group. On the 19th day after tumor inoculation, tumor volume in the SeMet group was much smaller than that in the control group. There was a significant difference between tumor volumes in the SeMet and control groups (P<0.05), indicating the SeMet effect on tumor growth inhibition.
Figure 1.

Representative tumor volume curves in the control group (blue) and SeMet group mice (red). The tumor volume has a significant reduction in the SeMet group as compared to control group. There was a significant difference (P<0.05) between SeMet and untreated control group.
SXRF imaging of iron, copper and zinc and selenium distribution in 4T1 tumor tissues after SeMet supplement
The distributions of iron, copper, zinc and selenium in the cryosections of 4T1 tumor tissues of the control and SeMet group on the 28th day after tumor inoculation were obtained using SXRF imaging (Figure 2). In the control group, there were some iron distribution in tumor tissue, but copper, zinc and selenium were not observed obviously. In contrast, iron and selenium were found significantly distributed in the tumor tissues of the SeMet group. Moreover, the distribution of selenium was more dispersive in most of tumor tissue, and high selenium distribution area was almost overlapped with the high iron distribution area.
Figure 2.
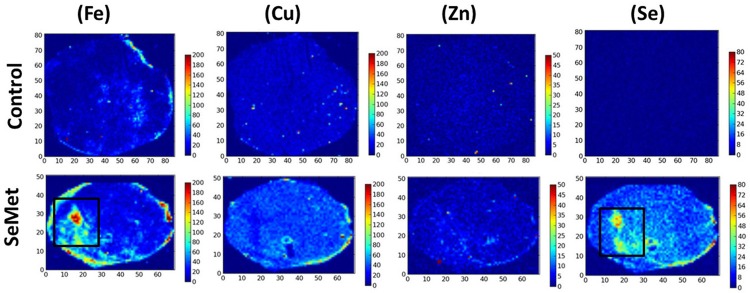
Fe, Cu, Zn and Se distributions in tumor tissues by SRXF imaging in two groups: control group and SeMet group on the 28th day after tumor inoculation. Iron and selenium significantly distributed in the tumor tissues of the SeMet group compared to the control. Moreover, selenium distributed in most of tumor tissue, and high selenium distribution area was almost overlapped in the high iron distribution area (black rectangle).
Selenium-associated protein expressions in 4T1 tumor tissues after SeMet supplement
To investigate if high selenium distribution in the tumor tissues after SeMet supplement interfere with selenium-associated proteins expression in the tumor tissues, the two selenium-associated proteins, Selenium-binding protein (SBP) 1 and another selenium-containing protein, glutathione peroxidase 1 (GPx1) expressions in the control group and SeMet group on the 28th day after tumor inoculation were studied using immunohistochemical staining. GPx1 expression in the control group was obviously much higher than that in the SeMet group (Figure 3A), while SBP1 expression in the SeMet group was obviously much higher than that in the control (Figure 3B). The results showed that there was an inversed association between GPx1 and selenium-binding protein 1 levels in the tumor tissues. High GPx1 expressed in tumor tissues with tumor growth, while after SeMet supplement, SBP1 expressed highly in tumor tissue and GPx1 expression decreased. High SBP1 expression after SeMet supplement might inhibit high GPx1 expression in tumor tissues.
Figure 3.
(A) Glutathione peroxidase 1 (GPx1) expression in the two groups: control group and SeMet group on the 28th day after tumor inoculation. (B) Selenium-binding protein (SBP1) expression in the two groups: control group and SeMet group on the 28th day after tumor inoculation. GPx1 expression in the control group was obviously much higher than that in SeMet group (A), while SBP1 expression in SeMet group was obviously much higher than that in the control group (B). Data were displayed as mean ± SEM and represented 10 images per tumor and three tumors per group in the right panel. *P<0.05.
Apoptosis of tumor cells after SeMet supplement
To investigate effects of SeMet supplement on tumor cells growth, cryosections of the control and SeMet groups on the 28th day after tumor inoculation were used for apoptosis analysis by TUNEL method. In Figure 4, brown staining revealed cells apoptosis. In the control group, apoptosis signals were only scattered in tumor tissues, while brown staining of apoptosis signals were observed in most of tumor tissues in SeMet group, implying most of tumor cells gone through apoptosis after SeMet intakes. It was clear that SeMet could promote tumor cells apoptosis.
Figure 4.

Apoptosis in tumor tissues of the control (A) and SeMet group (B). Brown staining revealed cells apoptosis signals. In the control group, apoptosis signals were only scattered in tumor tissues, while apoptosis almost covered whole tumor tissues in SeMet group. ×200; Scale bar: 100 μm. Data were displayed as mean ± SEM and represented 10 images per tumor and three tumors per group in the right panel. *P<0.05.
Effect of SeMet supplement on VEGF expression and tumor blood vessel intensity in 4T1 tumor tissues
VEGF is one of the most potent and specific angiogenic factors that promote formation of angiogenesis for continued tumor growth and metastasis. To investigate whether SeMet supplement could alter the expression of VEGF in tumor tissues, immunohistochemical staining of VEGF in 4T1 tumor tissues of the control and SeMet groups on the 28th day after tumor inoculation were performed (Figure 5). Brown staining indicated positive VEGF staining. In the control group, there was high VEGF expression in tumor tissues. But in the SeMet group, brown staining could not be observed, and the VEGF expression was found low in tumor tissues after SeMet supplement. The results showed that high SeMet intakes in tumor tissues after SeMet supplement could greatly inhibit VEGF expression.
Figure 5.

Immunohistochemical staining of VEGF expression in tumor tissues of the control (A) and SeMet group (B). ×200; Scale bar: 100 μm. In the control group, there was high VEGF expression in the tumor tissues. But in SeMet group, brown staining could not be observed, and there was low VEGF expression in the tumor tissues after SeMet supplement. Data was displayed as mean ± SEM and represented 5 images per tumor and three tumors per group in the right panel. *P<0.05.
With decreased VEGF expression, its correlation with CD31-positive microvessel density (MVD) also decreased significantly in the SeMet group, as compared to that in the control group (Figure 6). The results indicated that SeMet supplement inhibited tumor angiogenesis growth by inhibiting VEGF expression in tumor tissues.
Figure 6.
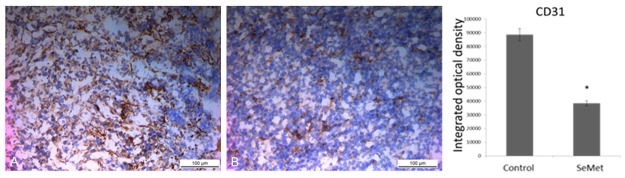
Immunohistochemical staining of CD31 in tumor tissues of the control (A) and SeMet group (B). ×200; Scale bar: 100 μm. Data was displayed as mean ± SEM and represented 10 images per tumor and three tumors per group in the right panel. *P<0.05.
The effect on immunosuppressive cells infiltration in tumor tissues after SeMet supplement
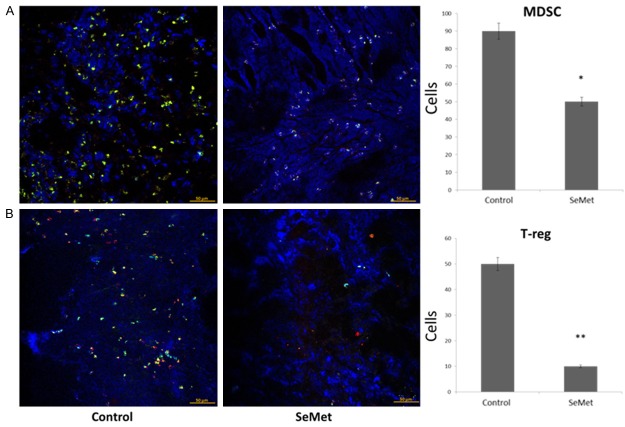
Immunosuppressive cells which include myeloid-derived suppressor cells (MDSC), and regulatory T cells (T-regs) etc. normally infiltrate in tumor tissues, which generate a highly immune suppression, and promote tumor progression and metastasis. To examine whether the SeMet supplement effect on immune suppression cells, immunofluorescence staining was performed in tumor tissues on the 28th day after SeMet supplement. In the control group, immune suppression cells, such as MDSCs and T-regs abundantly infiltrated into in tumor tissues, however, MDSCs and T-regs decreased significantly after the SeMet supplement (Figure 7). The results indicated SeMet supplement could significantly reduce immunosuppression cells infiltration in tumor tissues, and thus relief tumor immune suppression.
Figure 7.
Confocal microscopy immunofluorescence images of immune suppressive cells in tumor tissues of the control and SeMet groups. Scale bar: 50 μm. A. Immunofluorescence staining for MDSCs defined as co-localization of CD11b+ (green) and Gr-1+ (red) cells. B. Immunofluorescence staining for T-regs defined as co-localization of CD4+ (yellow), CD25+ (green) and Foxp3+ (red) cells. The number of eligible cells was counted and shown in the right panel. Data was displayed as mean ± SEM and represented 10 images (total magnification, 400×) per tumor and three tumors per group. MDSCs, Myeloid derived suppressor cells; T-regs, regulatory T cells. *P<0.05, **P<0.01.
Effect of SeMet supplement on HIF-1α accumulation in 4T1 tumor tissues
Hypoxia-inducible factor (HIF) 1α was found to transcriptionally regulate VEGF, and be responsible for immune suppression cells differentiation and function in the tumor microenvironment. To examine if SeMet supplement could affect HIF-1α accumulation, immunohistochemical analysis were performed. Specific positive staining (brown) of HIF-1α was found in tumor tissues (Figure 8). The data presented HIF-1α staining intensity was obviously strong on the 28th day of the control group as compared to that in SeMet group, indicating significant HIF-1α accumulation decrease after SeMet supplement.
Figure 8.

Immunohistochemical staining of HIF-1α in tumor tissues of the control (A) and SeMet (B) groups. ×200; Scale bar: 100 μm. Brown staining meant positive HIF-1α in the tumor tissues. Data were displayed as mean ± SEM from 10 images per tumor and three tumors per group in the right panel. *P<0.05.
Discussion
Currently, the effect of selenium supplement on breast cancer remains a controversial issue. The present study was to identify the relationship between selenium intake and breast cancer progression, to gain a better understanding of the relationships between Se and selenium related proteins in tumor tissues, and thus to find correlations between Se levels and some parameters and their relations to breast cancer progression. The results indicated that tumor growth was inhibited significantly after SeMet supplement. High Se intakes in tumor tissues was observed using SXRF imaging, and found its distribution overlapped with iron after SeMet supplement. Further, SeMet supplement increased SBP1 expression and decreased GPx1 expression in tumor tissues, and SeMet could promote most of tumor cells apoptosis, inhibited HIF-1α significantly. Accordingly, VEGF expression was greatly inhibited, and tumor immune suppression cells accumulation was relieved in tumor tissues. We postulated that high SBP1 expression after high Se intakes in tumor tissues by oral SeMet supplement played a crucial role in inhibiting tumor growth. SBP1 could promote most of tumor cells apoptosis by inhibiting expression of GPx1, at the same time, SBP1 decreased HIF-1α accumulation in tumor tissues by enhancing HIF-1α degradation resulting in inhibiting VEGF expression and relieving tumor immune suppression. So elucidation of the interrelation of selenium level, SBP1, GPx1 and HIF-1α may aid in understanding the mechanisms which selenium exerts its chemopreventive effects.
Glutathione peroxidase-1 (GPx-1) is a crucial antioxidant enzyme, which could reduce H2O2 to water and protect cells under oxidative Stress. In this study, high iron accumulation in 4T1 tumor tissues was observed. Iron might participate in facilitating cancer cell proliferation and promoting cell survival as an essential nutrient [20]. A recent study reported that iron levels in breast cancer biopsy tissue were 5 times higher than levels in breast tissue from women without breast cancer [21]. Avoiding excess iron accumulation to produce high ROS level, over-expression of GPx-1 protects the cell from damage due to ROS [22]. GPx-1 over-expression was observed in tumor tissues of the control group in our study. Moreover, due to the inhibition of SBP1 transcription by the GPX1 mediated reduction in both SBP1 mRNA and expression from an SBP1 promoter-driven reporter gene [14], over-expression of GPX1 resulted in the reduction of SBP1 in the control group.
SBP1, which has been suggested to mediate the intracellular transport of selenium, bond to selenium and formed high SBP1 expression [23]. The levels of SBP1 expression are directly increasing with elevated media selenium concentration [24]. After SeMet supplement, high Se distribution in tumor tissues was in accordance with high SBP1 expression found in our study, indicating that high Se existed as the form of SBP1. High SBP1 might inhibit GPx1 expression and activity in tumor tissues involving a direct physical interaction between the SBP1 and GPx1 forming a complex [14]. So, high Se almost overlapped with high iron distribution, which led to high SBP1 expression and inhibiting GPx1 expression in tumor tissues, thus, H2O2 could not be cleaned by GPx1 and H2O2-mediated apoptosis was induced. With SeMet supplement, most of tumor cells apoptosis could be observed in our study. Pohl NM indicated over-expression of SBP1 sensitized colon cancer cells to H2O2-induced apoptosis [25].
Hypoxia inducible factors 1 (HIF-1α) are transcription factors that regulate several genes involved in angiogenesis, cell proliferation, metabolism, drug resistance and other cellular processes [26]. Stable expression of HIF-1α is regulated by synthesis and degradation pathways. Increased cellular accumulation of ROS contributes to the stability of HIF-1α [27]. Thus, high HIF-1α accumulation was observed in the control group. Recently, SBP1 was indentified to contain hypoxia response elements (HREs) in its promoter region and was found to be a target gene for HIF-1α. SBP1 mRNA and protein expression was increased with expressing HIF-1α [28]. Chintala et al. have demonstrated HIF-1α inhibition by selenium as a post-translational degradation mechanism [29]. With SeMet supplement, high SBP1 expression would be mediated by expressing HIF-1α, while high SBP1 level conversely promoted HIF-1α degradation resulting in high SBP1 and low HIF-1α level in SeMet group as compared to that in the control group. Due to HIF-1α being inhibited through SBP1 promoting HIF-1α degradation, some events in cancer progression mediated by HIF-1α, such as angiogenesis growth and tumor immunosuppression cells recruitment were also inhibited in our study. Our studies indicated that selenium intake through the diet could act as a chemopreventive agent in reducing the growth of breast cancer.
Conclusions
In summary, combined with highly sensitive SXRF mapping, results from this study provided evidences that SeMet-mediated anti-cancer protection was through inhibiting tumor growth via promoting tumor apoptosis, and greatly decreasing VEGF expression and relief of tumor immune suppression via high SBP1 expression inhibiting GPx1 activity and HIF-1α. SeMet may be used as dietary supplement combined with chemotherapy in adjuvant therapy or prevention.
Acknowledgements
This work was supported by The National Basic Research Program of China (973 Program 2010CB834303), National Natural Science Foundation of China (11275126). The authors would especially like to thank Beamline scientists at BL15U Beamline of Shanghai Synchrotron Radiation Facilities (SSRF), China.
Disclosure of conflict of interest
None.
References
- 1.Ledesma MC, Jung-Hynes B, Schmit TL, Kumar R, Mukhtar H, Ahmad N. Selenium and vitamin E for prostate cancer: post-SELECT (Selenium and Vitamin E Cancer Prevention Trial) status. Mol Med. 2011;17:134–143. doi: 10.2119/molmed.2010.00136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Kim A, Oh JH, Park JM, Chung AS. Methylselenol generated from selenomethionine by methioninase downregulates integrin expression and induces caspase-mediated apoptosis of B16F10 melanoma cells. J Cell Physiol. 2007;212:386–400. doi: 10.1002/jcp.21038. [DOI] [PubMed] [Google Scholar]
- 3.Yang M, Sytkowski AJ. Differential expression and androgen regulation of the human selenium-binding protein gene hSP56 in prostate cancer cells. Cancer Res. 1998;58:3150–3153. [PubMed] [Google Scholar]
- 4.Hatfield DL, Gladyshev VN. How selenium has altered our understanding of the genetic code. Mol Cell Biol. 2002;22:3565–3576. doi: 10.1128/MCB.22.11.3565-3576.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Baliga MS, Diwadkar-Navsariwala V, Koh T, Fayad R, Fantuzzi G, Diamond AM. Selenoprotein deficiency enhances radiation-induced micronuclei formation. Mol Nutr Food Res. 2008;52:1300–1304. doi: 10.1002/mnfr.200800020. [DOI] [PubMed] [Google Scholar]
- 6.Zhuo P, Diamond AM. Molecular mechanisms by which selenoproteins affect cancer risk and progression. Biochimica et Biophysica Acta (BBA)-General Subjects. 2009;1790:1546–1554. doi: 10.1016/j.bbagen.2009.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lei XG, Cheng WH, McClung JP. Metabolic regulation and function of glutathione peroxidase-1. Annu Rev Nutr. 2007;27:41–61. doi: 10.1146/annurev.nutr.27.061406.093716. [DOI] [PubMed] [Google Scholar]
- 8.Yagublu V, Arthur J, Babayeva S, Nicol F, Post S, Keese M. Expression of Selenium-containing Proteins in Human Colon Carcinoma Tissue. Anticancer Res. 2011;31:2693–2698. [PubMed] [Google Scholar]
- 9.Jerome-Morais A, Wright ME, Liu R, Yang W, Jackson MI, Combs GF Jr, Diamond AM. Inverse association between glutathione peroxidase activity and both selenium-binding protein 1 levels and gleason score in human prostate tissue. Prostate. 2012;72:1006–1012. doi: 10.1002/pros.21506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chen G, Wang H, Miller CT, Thomas DG, Gharib TG, Misek DE, Giordano TJ, Orringer MB, Hanash SM, Beer DG. Reduced selenium-binding protein 1 expression is associated with poor outcome in lung adenocarcinomas. J Pathol. 2004;202:321–329. doi: 10.1002/path.1524. [DOI] [PubMed] [Google Scholar]
- 11.Kim H, Kang HJ, You KT, Kim SH, Lee KY, Kim TI, Kim C, Song SY, Kim HJ, Lee C, Kim H. Suppression of human selenium-binding protein 1 is a late event in colorectal carcinogenesis and is associated with poor survival. Proteomics. 2006;6:3466–3476. doi: 10.1002/pmic.200500629. [DOI] [PubMed] [Google Scholar]
- 12.Li T, Yang W, Li M, Byun DS, Tong C, Nasser S, Zhuang M, Arango D, Mariadason JM, Augenlicht LH. Expression of selenium-binding protein 1 characterizes intestinal cell maturation and predicts survival for patients with colorectal cancer. Mol Nutr Food Res. 2008;52:1289–1299. doi: 10.1002/mnfr.200700331. [DOI] [PubMed] [Google Scholar]
- 13.Huang KC, Park DC, Ng SK, Lee JY, Ni X, Ng WC, Bandera CA, Welch WR, Berkowitz RS, Mok SC, Ng SW. Selenium binding protein 1 in ovarian cancer. Int J Cancer. 2006;118:2433–40. doi: 10.1002/ijc.21671. [DOI] [PubMed] [Google Scholar]
- 14.Fang W, Goldberg ML, Pohl NM, Bi X, Tong C, Xiong B, Koh TJ, Diamond AM, Yang W. Functional and physical interaction between the selenium-binding protein 1 (SBP1) and the glutathione peroxidase 1 selenoprotein. Carcinogenesis. 2010;31:1360–1366. doi: 10.1093/carcin/bgq114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Li Z, Carrier L, Belame A, Thiyagarajah A, Salvo VA, Burow ME, Rowan BG. Combination of methylselenocysteine with tamoxifen inhibits MCF-7 breast cancer xenografts in nude mice through elevated apoptosis and reduced angiogenesis. Breast Cancer Res Treat. 2009;118:33–43. doi: 10.1007/s10549-008-0216-x. [DOI] [PubMed] [Google Scholar]
- 16.Fleshner N, Zlotta AR. Prostate cancer prevention. Cancer. 2007;110:1889–1899. doi: 10.1002/cncr.23009. [DOI] [PubMed] [Google Scholar]
- 17.Suzana S, Cham B, Ahmad Rohi G, Mohd Rizal R, Fairulnizal M, Normah H, Fatimah A. Relationship between selenium and breast cancer: A case-control study in the Kiang Valley. Singapore Med J. 2009;50:265–9. [PubMed] [Google Scholar]
- 18.Sun Z. The promise of synchrotron radiation in medical science. Australasian Medical Journal. 2009;1:1–5. [Google Scholar]
- 19.Tao K, Fang M, Alroy J, Sahagian GG. Imagable 4T1 model for the study of late stage breast cancer. BMC Cancer. 2008;8:228. doi: 10.1186/1471-2407-8-228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Chua ACG, Klopcic B, Lawrance IC, Olynyk JK, Trinder D. Iron: An emerging factor in colorectal carcinogenesis. World J Gastroenterol. 2010;16:663–672. doi: 10.3748/wjg.v16.i6.663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Cavalieri E, Chakravarti D, Guttenplan J, Hart E, Ingle J, Jankowiak R, Muti P, Rogan E, Russo J, Santen R, Sutter T. Catechol estrogen quinones as initiators of breast and other human cancers: implications for biomarkers of susceptibility and cancer prevention. Biochim Biophys Acta. 2006;1766:63–78. doi: 10.1016/j.bbcan.2006.03.001. [DOI] [PubMed] [Google Scholar]
- 22.Chen J, Cao Q, Qin C, Shao P, Wu Y, Wang M, Zhang Z, Yin C. GPx-1 polymorphism (rs1050450) contributes to tumor susceptibility: evidence from meta-analysis. J Cancer Res Clin Oncol. 2011;137:1553–61. doi: 10.1007/s00432-011-1033-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Xia YJ, Ma YY, He XJ, Wang HJ, Ye ZY, Tao HQ. Suppression of selenium-binding protein 1 in gastric cancer is associated with poor survival. Hum Pathol. 2011;42:1620–1628. doi: 10.1016/j.humpath.2011.01.008. [DOI] [PubMed] [Google Scholar]
- 24.Morrison DG, Dishart MK, Medina D. Intracellular 58-kd selenoprotein levels correlate with inhibition of DNA synthesis in mammary epithelial cells. Carcinogenesis. 1988;9:1801–1810. doi: 10.1093/carcin/9.10.1801. [DOI] [PubMed] [Google Scholar]
- 25.Pohl NM, Tong C, Fang W, Bi X, Li T, Yang W. Transcriptional regulation and biological functions of selenium-binding protein 1 in colorectal cancer in vitro and in nude mouse xenografts. PLoS One. 2009;4:e7774. doi: 10.1371/journal.pone.0007774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Semenza GL. Targeting HIF-1 for cancer therapy. Nat Rev Cancer. 2003;3:721–732. doi: 10.1038/nrc1187. [DOI] [PubMed] [Google Scholar]
- 27.Finley LWS, Carracedo A, Lee J, Souza A, Egia A, Zhang J, Teruya-Feldstein J, Moreira PI, Cardoso SM, Clish CB, Pandolfi PP, Haigis MC. SIRT3 opposes reprogramming of cancer cell metabolism through HIF1α destabilization. Cancer Cell. 2011;19:416–28. doi: 10.1016/j.ccr.2011.02.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Scortegagna M, Martin RJ, Kladney RD, Neumann RG, Arbeit JM. Hypoxia-inducible factor-1α suppresses squamous carcinogenic progression and epithelial-mesenchymal transition. Cancer Res. 2009;69:2638–2646. doi: 10.1158/0008-5472.CAN-08-3643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Chintala S, Najrana T, Toth K, Cao S, Durrani FA, Pili R, Rustum YM. Prolyl hydroxylase 2 dependent and Von-HippelLindau independent degradation of Hypoxiainducible factor 1 and 2 alpha by selenium in clear cell renal cell carcinoma leads to tumor growth inhibition. BMC Cancer. 2012;12:293. doi: 10.1186/1471-2407-12-293. [DOI] [PMC free article] [PubMed] [Google Scholar]