Abstract
Background and Aims The genus Rosa (150–200 species) is widely distributed throughout temperate and sub-tropical habitats from the northern hemisphere to tropical Asia, with only one tropical African species. In order to better understand the evolution of roses, this study examines infrageneric relationships with respect to conventional taxonomy, considers the extent of allopolyploidization and infers macroevolutionary processes that have led to the current distribution of the genus.
Methods Phylogenetic relationships among 101 species of the genus Rosa were reconstructed using sequences from the plastid psbA-trnH spacer, trnL intron, trnL-F spacer, trnS-G spacer and trnG intron, as well as from nuclear glyceraldehyde 3-phosphate dehydrogenase (GAPDH), which was used to identify putative allopolyploids and infer their possible origins. Chloroplast phylogeny was used to estimate divergence times and reconstruct ancestral areas.
Key Results Most subgenera and sections defined by traditional taxonomy are not monophyletic. However, several clades are partly consistent with currently recognized sections. Allopolyploidy seems to have played an important role in stabilizing intersectional hybrids. Biogeographic analyses suggest that Asia played a central role as a genetic reservoir in the evolution of the genus Rosa.
Conclusions The ancestral area reconstruction suggests that despite an early presence on the American continent, most extant American species are the results of a later re-colonization from Asia, probably through the Bering Land Bridge. The results suggest more recent exchanges between Asia and western North America than with eastern North America. The current distribution of roses from the Synstylae lineage in Europe is probably the result of a migration from Asia approx. 30 million years ago, after the closure of the Turgai strait. Directions for a new sectional classification of the genus Rosa are proposed, and the analyses provide an evolutionary framework for future studies on this notoriously difficult genus.
Keywords: Rosa, phylogeny, taxonomy, biogeography, ancestral area reconstruction, divergence time, allopolyploidy, hybridization
INTRODUCTION
The genus Rosa L. (roses; Rosoideae: Rosaceae) comprises about 150–200 species widely distributed throughout the temperate and sub-tropical habitats of the northern hemisphere (Rehder, 1940; Gu and Robertson, 2003), with the exception of one tropical African species. Approximately half of the rose species occur in Asia, while Europe and North America host approximately a quarter of the species each. The species of this genus are difficult to identify because of the homogeneity in morphology (see Fig. 1) associated with hybridization. Cultivated for >2000 years (Guoliang, 2003), roses are economically important as ornamental shrubs and cut flowers, as well as for perfumes, cosmetics and pharmaceutical research (see Cutler, 2003; and for recent pharmaceutical research, see Jager et al., 2007; Yi et al., 2007; Guimaraes et al., 2010). Moreover, numerous traits (small nuclear genome, extensive cross-species fertility and advanced industrial horticultural and micropropagation techniques) as well as their close affinity with several important woody Rosaceae crop species (e.g. raspberries, apples, almonds, cherries and peaches) suggest that roses could provide an ideal model for exploring woody plant genomes (see Bruneau et al., 2007; Debener and Linde, 2009).
Fig. 1.
Morphological diversity of flowers and fruits of a few representatives of Rosa. (A) Flower of Rosa hugonis. (B) Flower of R. moyesii. (C) Flower of R. multibracteata. (D) Flower of R. odorata. (E) Flower of R. praelucens. (F) Flower of R. prattii. (G) Flower of R. tsinglingensis. (H) Flower of R. roxburghii. (I) Flower of R. rubus. (J) Flower of R. roxburghii. (K) Flower of R. villosa. (L) Flower of R. primula. (M) Fruit of R. sikangensis. (N) Fruits of R. omeiensis. (O) Fruit of R. mairei. (P) Fruit of R. macrophylla. (Q) Fruits of R. sweginzowii.
Roses have captured the interest of scientists in various modern genetic fields (e.g. quantitative genetics and functional genomics; see Debener, 2009). However, these studies have been conducted within the framework of a classification that is probably obsolete because the most recent taxonomic treatment (Wissemann, 2003) still relies largely on the sub-divisions made >70 years ago (Rehder, 1940), and that are themselves adapted from 19th century arrangements (Crépin, 1889, 1891). Wissemann’s (2003) system divides the genus Rosa into four subgenera [R. subgen. Rosa, R. subgen. Hulthemia (Dumort.) Focke, R. subgen. Platyrhodon (Hurst) Rheder and R. subgen. Hesperhodos Cockerell] and the main subgenus Rosa into ten sections [R. sect. Pimpinellifoliae (DC.) Ser., R. sect. Rosa, R. sect. Caninae (DC.) Ser., R. sect. Carolinae Crép., R. sect. Cinnamomeae (DC.) Ser., R. sect. Synstylae DC., R. sect. Indicae Thory, R. sect. Banksianae Lindl., R. sect. Laevigatae Thory and R. sect. Bracteatae Thory], all subgenera and sections being identical to those of Rehder (1940). In addition, Wissemann (2003) defined six new sub-sections in R. sect. Caninae (R. sub-sect. Trachyphyllae H. Christ, R. sub-sect. Rubrifoliae Crép., R. sub-sect. Vestitae H. Christ, R. sub-sect. Rubiginae H. Christ, R. sub-sect. Tomentellae H. Christ and R. sub-sect. Caninae).
There have been numerous attempts at building a phylogeny that would give a new perspective on the genus [rapidly amplified polymorphic DNA (RAPD) data, Debbener et al., 1996; Millan et al., 1996; Jan et al., 1999; DNA sequences, Matsumoto et al., 1998, 2000, 2001; Iwata et al., 2000; Wu et al., 2000, 2001; Wissemann and Ritz, 2005; Bruneau et al., 2007; Qiu, 2012; microsatellite analyses, Scariot et al., 2006; amplified fragment length polymorphism (AFLP) data, Koopman et al., 2008]. However, these studies are contradictory and only a few of them support the monophyly of Wissemann’s sections. These previous studies faced numerous problems. First, in most of these studies, phylogenetic resolution is poor and, where clades are resolved, support is often weak. This is explained partly by the extremely low levels of sequence divergence observed across the genus (e.g. Matsumoto et al., 1998; Wissemann and Ritz, 2005). Secondly, hybridization complicates phylogeny reconstruction in roses. Several studies have confirmed that interspecific hybridization is frequent in the genus (Ritz et al., 2005; Joly and Bruneau, 2006; Joly et al., 2006; Schanzer and Vagina, 2007; Mercure and Bruneau, 2008; Schanzer and Kutlunina, 2010; Ritz and Wissemann, 2011; Kellner et al., 2012b). Indeed, Wissemann and Ritz (2005) noted several contradictions between their plastid and nuclear gene phylogenies. Thirdly, identification can also be problematic because Rosa taxonomy is further complicated by the publication of numerous names given to morphological variants and hybrids (Wissemann, 2003). Problems with the identification of plant material or sequencing of hybrids could explain why conspecific samples sometimes fall into distinct clades in some studies (see Bruneau et al., 2007). Fourthly, sampling has often been incomplete and biased toward cultivated varieties or specific geographic areas. When several geographic areas were represented, usually only one of them was effectively represented by wild-collected samples, and the rest were represented by garden-grown specimens. Given the strong ability of roses to hybridize, the use of garden-grown specimens is questionable unless a wild origin is clearly established.
In this study, we present a robust molecular phylogeny of the genus Rosa to provide the genus-wide perspective necessary to determine the origins of wild roses and to orient future studies on Rosa properly (floral evolution, traits genomics, conservation studies, rose breeding, etc.). Wild samples from Asia, Europe and North America were collected and were supplemented by a few herbarium or garden-grown samples (usually from wild origin). We use sequences from the chloroplast trnL intron and trnL-F and psbA-trnH intergenic spacers that have been found to be relatively variable in Rosa (Bruneau et al., 2007), but also include sequences from the trnS-G spacer and trnG intron, which Shaw et al. (2007) suggested were highly informative regions. The use of chloroplast data seems to be appropriate in Rosa to draw a first phylogenetic hypothesis without taking into account reticulate evolution, but we also present phylogenetic relationships obtained with the nuclear cytosolic glyceraldehyde 3-phosphate dehydrogenase (GAPDH) gene to highlight evidence for reticulate evolution in the genus. Finally, we infer a biogeographic history of the genus and draw general conclusions on our current knowledge of Rosa phylogeny in regards to the currently accepted taxonomy.
MATERIALS AND METHODS
Taxon sampling
A total of 101 rose species are represented in our sampling, corresponding to more than half of the species reported in the genus (Supplementary Data S1). At least one species from each of the subgenera and each of the sections belonging to the genus Rosa were sampled. Our sampling includes North American, European and Asian species, as well as the African species R. abyssinica R. Br. Most samples were collected in the field, and material was preserved in silica gel for molecular analyses while vouchers were deposited in herbaria. This fairly complete sampling of samples collected in natural habitats was supplemented by samples obtained from plants cultivated in gardens (from wild origin) or from herbarium specimens. When possible, we avoided garden-grown specimens obtained from seeds because they are more likely to have undergone hybridization.
Sequences of the trnL region (including the trnL-F spacer and trnL intron) and the psbA-trnH intergenic spacer were generated for 107 samples of roses, and these were added to the 50 sequences published by Bruneau et al. (2007) for 25 more samples. For the chloroplast trnG region (including the trnG intron and the trnG-trnS spacer), a total of 129 sequences were generated and included in the analyses. Sequences of the three regions generated for Rubus biflorus were used as the outgroup. Most samples included were sequenced for the three different regions. However, three samples (one for R. foetida Herrm. and two for R. minutifolia Engelm.) are represented only by two sequences because the trnG region sequences could not be obtained. In addition, a sub-set of 55 of the 101 species were also sequenced for the nuclear cytosolic GAPDH gene. The GAPDH gene was amplified from the end of exon 7 [according to the Arabidopsis thaliana (L.) Heynh. sequence; GenBank locus tag: At3g04120] to the beginning of exon 11 (which is exon 9 in A. thaliana; see Joly et al., 2006). Each of the 55 samples sequenced was represented by 1–5 sequences. We also included sequences previously published by Joly et al. (2006) and Meng et al. (2011) for 14 taxa. New sequences were submitted to GenBank, and accession numbers are given in Supplementary Data S1.
Molecular methods
DNA was extracted either using a modification of the Doyle and Doyle (1987) cetyltrimethylammonium bromide (CTAB) protocol or, notably for herbarium specimens, using the Tiangen Plant Genomic DNA Kit (Beijing) following instructions from the manufacturer. The PCR amplification mix contained 4–5 U of Taq DNA polymerase, Taq DNA polymerase buffer (Tiangen, Beijing) with 2 mm MgCl2, 200 μm of each dNTP and 0·4 μm of each primer. Amplifications were conducted using a PTC-0200 thermocycler (Bio-Rad, Beijing). Conditions for amplification of the trnL region were 3 min of initial denaturation at 94 °C, followed by 40 cycles of 30 s at 94 °C, 30 s at 47·5 °C and 1 min 30 s at 72 °C, with a final step of 7 min at 72 °C. Similar conditions were used for the other three regions, except that the annealing temperature was 52·5 °C for the psbA-trnH intergenic spacer, 54 °C for the trnG region and 49 °C for the GAPDH region, and the elongation time was 30 s for the psbA-trnH intergenic spacer and the trnG region and 2 min for the GAPDH region. For the GAPDH region, when multiple copies were suspected (presence of double peaks in direct sequencing reads), cloning was performed. PCR products were cloned in pMD19-T vector (TaKaRa Biotechnology, Dalian, China) following the instructions of the manufacturer. Colonies were screened by PCR and selected colonies were incubated overnight in LB broth with appropriate antibiotics. Two (for diploids) to 18 clones (for pentaploids) were selected depending on the ploidy level reported in the species. PCR products or clones were sent to Invitrogen (Shanghai) for purification and sequencing.
The trnL intron and trnL-F spacer were amplified using the ‘c’ and ‘f’ primer pair (Taberlet et al., 1991) then sequenced using the amplification pair as well as internal primers ‘d’ and ‘e’ as described by Taberlet et al. (1991). The psbA-trnH spacer was amplified and sequenced using ‘psbAF’ and ‘trnHR’ as described by Sang et al. (1997). The trnS-G spacer and trnG intron were amplified using the ‘trnS’ and ‘trnG’ primer pair as described by Shaw et al. (2007) then sequenced using those two primers and internal primers ‘trnG2S’ and ‘trnG2G’, as described by Shaw et al. (2005). The GAPDH region was amplified using GPDX7F (Strand et al., 1997) and GPDX11R (Joly et al., 2006). The strands were combined and edited using Sequencher (version 4.14, Gene Codes Corporation, Ann Arbor, MI, USA).
Phylogenetic analyses
Sequence alignments were performed using Clustal W (Larkin et al., 2007) with the default parameters as implemented in MEGA 4 (Tamura et al., 2007). Alignments were then verified and modified manually where inconsistencies were found. The alignments for the trnL and GAPDH were straightforward, but for the psbA-trnH spacer and trnG regions homology was difficult to assess in three highly repetitive regions that were subsequently removed from the analyses (182 bp of aligned sequences in the psbA-trnH spacer and 65 plus 50 bp of aligned sequences in the trnG regions).
Phylogenetic analyses of the concatenated chloroplast sequence data were performed under Maximum Likelihood optimization using RAxML 7.2.7 on CIPRES Science Gateway (Stamatakis, 2006; Stamatakis et al., 2008) with separate GTR + Γ substitution models for each region and the fast bootstrap option using 1000 replicates. Bootstrap values were considered low when strictly inferior to 65 %, moderate between 65 and 80 % and strong when superior to 80 %. The plastid matrices are available in TreeBASE (http://purl.org/phylo/treebase/phylows/study/TB2:S15526).
In vitro recombination of DNA sequences is a problem when cloning products of PCRs in which multiple alleles or paralogous copies have been amplified (Cronn et al., 2002; Russell et al., 2010). For this reason, we screened the sequences and eliminated chimeric sequences as described in Russell et al. (2010). For polyploids, single nucleotide substitutions [single nucleotide polymorphisms (SNPs)] were considered as PCR errors when not shared by several clones. For diploids, SNPs were considered as PCR errors when no double peaks were present in direct sequencing. Nuclear analyses were performed using the NeighborNet algorithm implemented in SplitsTree 4.13.1 (Huson and Bryant, 2006) to reconstruct a network that would summarize better than a tree the complex relationships between the different copies of GAPDH present in diploids and polypploids.
Divergence time analyses
Because of the complex nature of nuclear results and because we did not want to exclude the numerous polyploid species in the genus, we used the plastid sequence data with greater taxon sampling to perform biogeographical analyses. Preliminary analyses using a relaxed lognormal clock as implemented in BEAST 1.7.3 (Drummond et al., 2006; Drummond and Rambaut, 2007) failed to converge. A likelihood ratio test performed on the three plastid regions independently failed to reject the molecular clock hypothesis. As Brown and Yang (2011) suggested that a strict molecular clock could perform well when rate variation is low and that the likelihood ratio test was suitable to test the strict molecular clock hypothesis, divergence time analyses were conducted using a strict molecular clock as implemented in BEAST 1.7.3. The best partition scheme and the substitution models for each partition were selected using PartitionFinder (Lanfear et al., 2012). Consequently, the TVM + G, TVM + G and TVM + I + G substitution models were used for the psbA-trnH spacer, the trnL region and the trnG region, respectively, with a Birth Death process tree prior. Markov chain Monte Carlo (MCMC) runs were extended for 20 million generations (burn-in 10 %), with parameters and trees sampled every 1000 generations. Convergence was assessed using Tracer v1.5 (Rambaut et al., 2013) and then the maximum clade credibility tree was selected using TreeAnnotator from the BEAST package.
We applied two calibration points. The first calibration point was the stem node of the genus Rosa. The oldest known fossils in the genus are R. germerensis (Edelman, 1975) from the Germer Tuffaceous member, Challis volcanic formation [Idaho, 55·8–48·6 million years ago (Ma)] and R. hilliae Lesq. (Anonymous, 1978) from the Jijuntun formation (Fushun, 51–45 Ma; Meng et al., 2012). To accommodate those ages, we used a normal prior distribution centred on 50·5 Ma and a 97·5 % confidence interval between 55 and 45 Ma. The second calibration point was the stem node of the Synstylae and allies (see the Results). A recent study by Kellner et al. (2012a) reported that some Rosa lignitum Heer fossils (30·44 ± 1·52 Ma of age) exhibit a particular semi-craspedodromous leaf venation pattern that seem to be restricted to extant species of our Synstylae and allies clade. This character could be an adaptation to warmer climate but, first, our own observations suggest that this adaptation to warmer climate is also restricted to the Synstylae and allies clade and, secondly, Kellner et al. (2012a) showed that R. stellata Wooton (R. subgen. Hesperhodos) lacks this particular leaf venation while being adapted to warm climate. Other fossils from Colorado (Florissant) around the same epoch also lack this particular leaf venation (see Becker, 1963). The separation of this clade from the rest of the genus can be considered older than the first semi-craspedodromous fossil. The stem node of the Synstylae and allies clade was therefore constrained using a normal prior distribution centred on 30·5 Ma with a 97·5 % confidence interval between 28·99 and 32 Ma.
Ancestral area reconstructions
We performed ancestral area reconstruction analyses on the plastid chronogram obtained with BEAST. We used the dispersal–extinction–cladogenesis (DEC; Ree and Smith, 2008) model as implemented in RASP 2.1b (Yu et al., 2013). It uses the information contained in genetic branch lengths and allows the incorporation of changing dispersal probabilities across area and time. Areas were defined as described below. North America included Canada, the USA and northern Mexico. The southern boundary for Rosa distribution in North America is Baja California, so Central and South America are not considered. Africa included sub-Saharan Africa only. A few species occur in northern Africa (Atlas Mountains) and are also distributed in Europe. Similarly, the Northern part of the Arabian Peninsula is also included in the European distribution because only species distributed otherwise in the rest of Europe were found. The limit between Europe and Asia was drawn on the Caspian Sea, and Kazakhstan and Iran are considered as parts of Asia. Russia is divided into two parts: one Asian (Siberia and Far East) and one European (for the rest of the country). The maximum number of areas at each node was set to four because the most widespread species in the genus occur in four of our areas. We tested different time-slice models (one to four, see Supplementary Data S2), which yielded identical results. Fossil distribution information was alternatively excluded or included. Early Rosa fossils from China and the USA were considered to reconstruct the ancestral area for the genus. The presence of R. lignitum with a semi-craspedodromous venation in Europe during the Oligocene provides evidence for the presence of the Synstylae and allies in this area, but the fossils from the other areas have not been examined for semi-craspedodromous venation. It is thus difficult to know whether the Synstylae and allies were genuinely absent outside of Europe. Because the ancestral area reconstruction (AAR) excluding fossil information suggested Asia to be the ancestral area for the Synstylae and allies clade, we considered this clade to be present in both Europe and Asia.
RESULTS
Plastid DNA phylogeny
The plastid DNA analyses (Fig. 2) suggest that R. subgen. Rosa is not monophyletic but instead resolved two main clades: the Cinnamomeae clade and the Synstylae clade. The Cinnamomeae clade is resolved with low support (55 %), and includes all members of R. sects Cinnamomeae and Carolinae (R. subgen. Rosa), as well as a few species from other sections (three from R. sect. Pimpinellifoliae species and one from R. sect. Synstylae) and one species from R. subgen. Platyrhodon. The Cinnamomeae clade includes a strongly supported clade (97 %) containing almost all American species of R. sects Cinnamomeae and Carolinae (except R. nutkana C. Presl var. hispida Fernald), the two European species of R. sect. Cinnamomeae and some Asian species. The Cinnamomeae clade together with the Pimpinnelifoliae clade and R. subgen. Hesperhodos and Hulthemia form an unsupported clade, and will be designated by the name Cinnamomeae and allies. The Synstylae clade is resolved with moderate support (76 %) and comprises nearly all members of R. sect. Synstylae and all members of R. sects Indicae, Caninae and Rosa. Within the Synstylae clade, a clade containing all the Asian species of R. sects Synstylae and Indicae is resolved with strong support (88 %) and another clade containing the other species of R. sect. Synstylae (from America and Europe) as well as R. gallica L. (monotypic R. sect. Rosa) and all members of R. sect. Caninae is resolved with strong support (89 %). Rosa sect. Caninae itself is not resolved as monophyletic and is divided into two clades. The first clade, the Caninae clade, comprises the members of R. sub-sect. Caninae with low support (53 %) but is strongly supported (93 %) as closely related to other European roses from the Synstylae clade. The second clade, the Rubigineae clade, gathers the members of the four other sub-sections with strong support (93 %). A poorly supported clade named Synstylae and allies includes the Synstylae clade and members of R. sects Laevigatae, Bracteatae and Banksianae as well as R. subgen. Platyrhodon, with no resolution within this larger clade.
Fig. 2.
Phylogenetic relationships among Rosa species as reconstructed by Maximum Likelihood analyses of three chloroplast regions (psbA-trnH spacer, trnL region and trnG region). Bootstrap values are placed as close as possible to the node supported. The ploidy level of each species is given after its name (see Erlanson, 1929, 1934, 1938; Roberts, 1977; Yokoya et al., 2000; Roberts et al., 2009; Jian et al., 2010). The names of known polyploids are in bold (in R. sect. Caninae all species are presumed to be polyploids even when the ploidy number is not exactly known). Wissemann’s (2003) classification is compared with our clades. A P designates our Pimpinellifoliae clade, a C our Cinnamomeae clade, an S our Synstylae clade, Ca our Caninae clade and Ru our Rubiginae clade.
Nuclear analyses
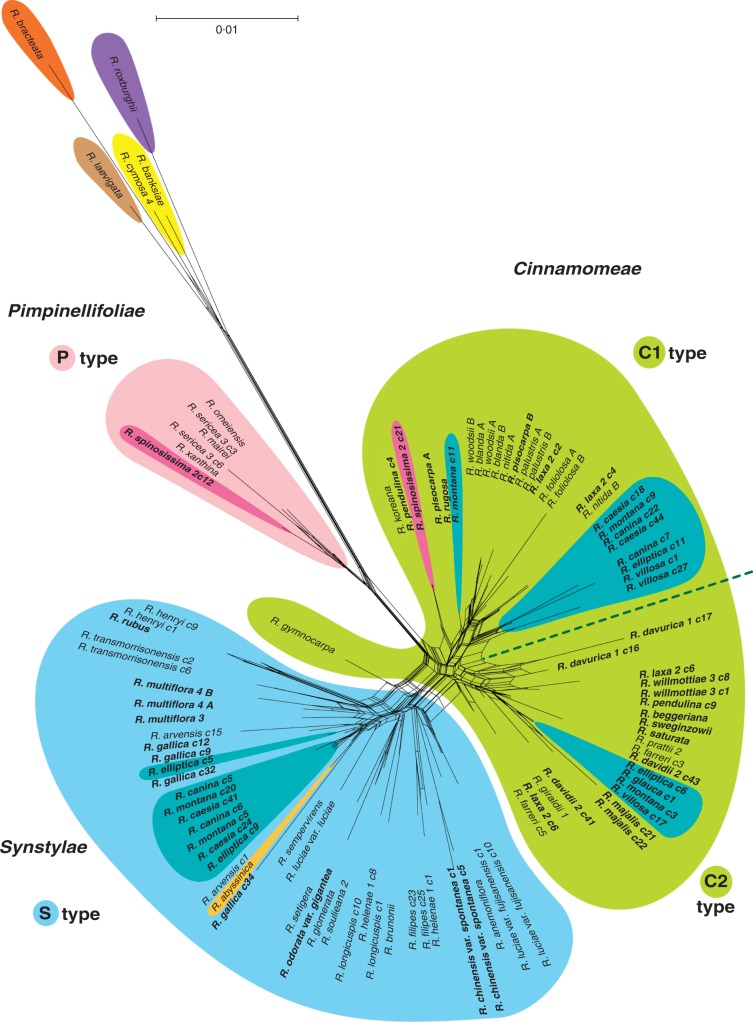
The NeighborNet analyses of the GAPDH nuclear sequences (Fig. 3) suggest that sequences from R. roxburghii Tratt. (R. subgen. Platyrhodon) and those from R. sect. Bracteatae, R. sect. Banksianae and R. sect. Laevigatae, as well as those from R. sect. Pimpinellifoliae are each well differentiated from each other and also from the rest of the genus. Moreover, sequences from Cinnamomeae and Synstylae clade members seem to have diversified independently except that the R. gymnocarpa Nutt. (R. sect. Cinnamomeae) GAPDH sequence is oddly placed close to those of the Synstylae clade. Other sequences available in GenBank for this species are not included in the analysis but gave the same result (not shown for the sake of clarity). Inside the Cinnamomeae group, two groups can be distinguished and we designate them as C1 and C2 types (see Fig. 3). The relationships between the Cinnamomeae clade sequences and the Synstylae clade sequences are very complex. The C1 and C2 groups are not supported by the chloroplast data, and some polyploids (Rosa laxa Retz. and R. pendulina L.) have copies from both groups. Members of R. sect. Caninae seem to possess a complex genome including alleles related to the Synstylae group and alleles related to the Cinnamomeae group. The alleles from R. sect. Caninae related to the Cinnamomeae group are divided into the two C1 and C2 groups, indicating that the genome from members of R. sect. Caninae contains three kinds of alleles (see Table 1). Despite the high number of clones sequenced, it was not always possible to recover all the alleles expected. Sometimes one of the alleles was more frequently sequenced than the other (nine of 13 clones sequenced were the same in R. villosa L.).
Fig. 3.
Network representing the relationships among copies of GAPDH obtained from Rosa species. A ‘c’ followed by a number indicates the number attributed to one particular clone sequenced. The groups are compared with our main clades from the chloroplast analyses. Purple is attributed to Rosa subgen. Platyrhodon, yellow to R. sect. Banksianae, bright orange to R. sect. Bracteatae, brown to R. sect. Laevigatae, light pink to Pimpinellifoliae clade, green to Cinnamomeae clade, light blue to Synstylae clade and a deeper blue to R. sect. Caninae. Two species have a particular colour, R. spinosissima is highlighted with a deeper pink colour and R. abyssinica with a light orange colour. Two types of copies, C1 and C2, are distinguished in our Cinnamomeae group. Some polyploids have several copies with different affinities. The names of known polyploids are in bold (in R. sect. Caninae all species are presumed to be polyploids even when the ploidy number is not exactly known).
Table 1.
Composition of the Caninae genome for some of the species studied
| Sub-section | Species | Ploidy level | GAPDH copies | Chloroplast |
|---|---|---|---|---|
| Caninae | R. montana | ? | SSC1C1C2 | Caninae, Synstylae |
| Caninae | R. canina | 4, 5 or 6 | SSC1C1? | Caninae, Synstylae |
| Caninae | R. caesia | 5 or 6 | SSC1C1? | Caninae, Synstylae |
| Rubiginae | R. elliptica | ? | SSC1C2? | Rubiginae, Synstylae |
| Tomentosae | R. villosa | 4 or 5 | ??C1C1C2 | Rubiginae, Synstylae |
Letters in the ‘GAPDH copies’ column correspond to the different types of GAPDH copies as defined in Fig. 3 (S for Synstylae type; C1 and C2 for two different groups of Cinnamomeae type). Question marks in this column indicate that some copies may not have been recovered.
In the ‘Chloroplast’ columns, the phylogenetic origin is described according to clades defined in the chloroplast phylogeny (Fig. 2). Ploidy levels are detailed in Roberts et al. (2009).
Biogeographic analyses
The plastid analyses suggest that the Synstylae and allies and the Cinnamomeae and allies are characterized by contrasting geographic patterns (Fig. 4). Members of Cinnamomeae and allies are mostly Asian and American (only one African species, two European species and two widely distributed) and neither of these two geographic origins is monophyletic, implying multiple dispersal events. Members of Synstylae and allies occur mostly in Asia and Europe (only one American species) and the European origin is monophyletic while Asian species are paraphyletic. The Pimpinellifolieae clade and most of the species from early diverging lineages (R. sects Pimpinellifolieae, Hulthemia, Platyrhodon, Bracteatae, Laevigatae and Banksianae) are Asian, except R. subgen. Hesperhodos. Our ancestral area reconstruction including fossil information suggests that early distribution of the genus included Asia and America. Combined with the divergence time analyses, our results also suggest that the Synstylae lineage and its allies extended their distribution from Asia to Europe around 30·1 Ma and then that part of the lineage reached eastern North America around 17·4 Ma. The exchanges between Asia and the rest of the range were interrupted at 13·1 Ma and the exchanges between Europe and eastern North America persisted until 8·4 Ma. Despite the early presence of the genus Rosa on the American continent, the ancestor of the Pimpinellifoliae and Cinnamomeae clade was strictly Asian. This means that among the extant species of American roses, only the species of R. subgen. Hesperhodos results from this ancestral widespread distribution while the other American species result from a later (at 13·4 Ma) re-colonization from Asia. Exchanges between western North America and eastern North America seem to persist even today. Exchanges between eastern North America and Asia were interrupted at 5·3 Ma but exchanges between western North America and Asia lasted longer and were finally interrupted at 4·1 Ma. Disjunctions between eastern Asia and eastern North America are not represented in our results. Instead, eastern North American species seem to be closely related to western North American or European species. The African R. abyssinica seems to have captured its chloroplast genome from Cinnamomeae between 9·3 and 6·9 Ma. European Cinnamomeae species seem to be the results of colonization from Asia between 2·5 and 0·6 Ma.
Fig. 4.
Chronogram obtained from BEAST analyses of chloroplast regions. Branches are coloured according to the ancestral area reconstruction analyses (DEC model) including fossils information. A P designates our Pimpinellifoliae clade, a C our Cinnamomeae clade and an S our Synstylae clade. The names of known polyploids are in bold (in R. sect. Caninae all species are presumed to be polyploids even when the ploidy number is not exactly known).
DISCUSSION
Our study has generated the most comprehensively sampled and well-resolved phylogeny of the genus Rosa to date, although support for the deeper nodes of the phylogeny remains low. Some clades roughly corresponding to sections described by Wissemann (2003) are supported. Although some relationships obtained here are consistent with previous studies, others are new or inconsistent with previous results (see Table 2 for example) and these inconsistencies are likely to be the result of hybridization.
Table 2.
Comparison of the recurrent Cinnamomeae and Synstylae clades from different studies
| Study | Cinnamomeae clade | Synstylae clade |
|---|---|---|
| Zhang et al. (2013) | Cinnamomeae and Carolinae | Synstylae and Indicae |
| SSR and flanking regions | Some Caninae haplotypes | Some Caninae haplotypes |
| Bootstrap: no support | R. pimpinellifolia | R. pimpinellifolia var. spinosissima |
| Qiu et al. (2012) | Cinnamomeae* | Synstylae and Indicae |
| ITS and matK | R. praelucens (subgen. Platyrhodon) | R. alba (sect. Rosa) |
| Bootstrap: 79 and 83 resp. | ||
| Meng et al. (2011) | Absent | Synstylae and Indicae |
| GAPDH | (paraphyletic) | R. gallica (sect. Rosa) |
| Bootstrap: no and 56 | ||
| Koopman et al. (2008) | Cinnamomeae and Carolinae | Synstylae† |
| AFLP; BA | Caninae and sect. Rosa | |
| PP: 59 and no, respectively | ||
| Koopman et al. (2008) | Cinnamomeae and Carolinae | Synstylae† |
| AFLP; MP | Most Caninae | Some Caninae and sect. Rosa |
| Bootstrap: no support | R. spinosissima | |
| Bruneau et al. (2007) | Cinnamomeae and Carolinae | Synstylae and Indicae |
| trnL region, psbA-trnH | R. koreana and R. spinosissima | Caninae and sect. Rosa |
| Bootstrap: no support | var. altaica (Pimpinellifoliae) | R. xanthina, R. bracteata, R. laevigata and R. cymosa |
| PP: 92 and 96, respectively | ||
| Scariot et al. (2006) | Cinnamomeae and Carolinae | Synstylae and Indicae |
| SSR | Pimpinellifoliae | Caninae and sect. Rosa |
| Bootstrap: no and 92 | ||
| Wissemann and Ritz (2005) | Cinnamomeae and Carolinae (both in part) | Synstylae and Indicae |
| atpB-rbcL | R. altaica | Caninae and sect. Rosa |
| PP: 83 and 70, respectively | ||
| Wissemann and Ritz (2005) | Cinnamomeae (in part) and Carolinae and Bracteatae | Synstylae (in part) and Indicae |
| ITS | R. hugonis | Caninae |
| PP: 50 and no, respectively | ||
| Wu et al. (2001) | Absent | Synstylae only |
| ITS | (polyphyletic) | |
| Bootstrap: no and 68 | ||
| Jan et al. (1999) | Cinnamomae sister to Carolinae | Synstylae sister to Indicae |
| RAPD | ||
| Matsumoto et al. (1998) | Cinnamomae and Carolinae | Synstylae and Indicae |
| matK | R. spinosissima var. pimpinellifolia | Caninae and sect. Rosa |
| Bootstrap: 80 and 61, respectively | R. californica ‘Plena’ (Cinnamomae) |
For each study the type of data used and the supports are given when available (bootstrap or posterior probability, PP).
AFLP, amplified fragment length polymorphism; BA, Bayesian analyses, GAPDH, glyceraldehyde 3-phosphate dehydrogenase; ITS, internal transcribed spacer; MP, Maximum Parsimony; RAPD, random amplified polymorphic DNA; SSR, simple sequence repeat.
*Rosa sect. Carolinae not sampled.
†Rosa sect. Indicae not sampled.
Subgenus status
The two arid-adapted subgenera (R. subgen. Hulthemia and Hesperhodos) are suggested as closely related despite low clade support, a relationship previously suggested by Wissemann and Ritz (2005).
Rosa subgen. Platyrhodon (R. roxburghii and R. praelucens Bijh.) is not resolved as monophyletic in our plastid analyses. While R. roxburghii is resolved as an independent lineage in both our plastid and nuclear analyses, R. praelucens is supported as a member of the Cinnamomeae clade in our plastid analyses. We consider the diploid R. roxburghii as a typical member of R. subgen. Platyrhodon. However, because allopolyploids have often been reported in the genus (Joly and Bruneau, 2006; Joly et al., 2006; Mercure and Bruneau, 2008; Schanzer and Kutlunina, 2010), we suspect that the decaploid R. praelucens (Jian et al., 2010) is an allopolyploid resulting from a hybridization event (or multiple hybridization events) involving the diploid R. roxburghii and at least one member of the Cinnamomeae clade. Based on karyomorphology, Jian et al. (2010) also suggested that R. praelucens is of allopolyploid origin, and a study in progress using GAPDH sequences supports the same conclusion (X. F. Gao, Chengdu Institute of Biology, CAS, Chengdu, China, pers. commun.).
Wissemann’s (2003) classification, based on morphology, divides the genus Rosa into four subgenera (R. subgen. Hesperhodos, Platyrhodon, Hulthemia and Rosa), but to date most phylogenetic studies (Matsumoto et al., 1998; Jan et al., 1999; Wu et al., 2000, 2001; Wissemann and Ritz, 2005; Bruneau et al., 2007; Koopman et al., 2008) have failed to recover a monophyletic R. subgen. Rosa. For this reason, Wissemann and Ritz (2005) suggested that the four subgenera would be best treated at the sectional level. Similarly, our results suggest that R. subgen. Rosa is not monophyletic with R. subgen. Platyrhodon as a member of the Synstylae and allies clade. Despite low support, the occurrence of a semi-craspedodromous venation in R. subgen. Platyrhodon and other members of the Synstylae and allies clade further support this result. So the status of the four subgenera remains questionable.
Wissemann’s (2003) sections
Our plastid sequence analyses resolved R. sect. Banksianae as monophyletic (both species sampled from wild origin) and closely related to but not embedded in the Synstylae clade. The nuclear GAPDH analyses confirm the close relationship of the two species and suggest that they are distinct from the Synstylae clade, as noted by other studies [Wu et al., 2001, internal transcribed spacer (ITS); Qiu et al., 2012, ITS and matK].
Rosa sect. Bracteatae (one of two species sampled) and R. sect. Laevigatae (monospecific) are resolved as close relatives of the Synstylae clade, and our nuclear GAPDH sequences suggest that they form a distinct lineage. Several studies obtained results similar to ours for R. sect. Bracteatae (Matsumoto et al., 1998; Wu et al., 2001), for R. sect. Laevigatae (Jan et al., 1999; chloroplast result only, in Wissemann and Ritz, 2005) or for the two sections (Qiu et al., 2012).
Regarding the other sections, our results are consistent with other studies in which the genus Rosa generally is resolved into two main clades (see Table 2), named Cinnamomeae and Synstylae clades in this study but, for example, named Clade I and Clade II in Bruneau et al. (2007). Similar results are obtained with the analyses of the nuclear GAPDH sequences, suggesting that the two groups are distinct but the pattern of relationships within each group is complex, even when putative allopolyploids are removed from the analysis (Appendix).
Both the plastid and nuclear analyses resolve R. sect. Indicae as embedded in R. sect. Synstylae. Similar relationships between these two sections have been reported in several studies (see Table 2: Matsumoto, 1998; Wissemann and Ritz, 2005; Scariot et al., 2006; Bruneau et al., 2007; Meng et al., 2011; Qiu et al., 2012; Zhang et al., 2013), but other studies suggested that these two sections are independently monophyletic (RAPD – Millan et al., 1996; Jan et al., 1999; ITS –Matsumoto et al., 2000; Wu et al., 2001). Only one sample of each species of R. sect. Indicae, R. odorata (Andrews) Sweet and R. chinensis Jacq. were included here, but other samples of these two species yielded identical sequences to the samples included. They were not included because data were available for only one or two plastid regions.
The chloroplast sequences of the two samples of the Asian R. multiflora Thunb. collected in China formed a clade while the third one collected in North America, where it is naturalized, occurred in a clade with other Asian species of R. sect. Synstylae. The nuclear GAPDH sequences from another American sample were closely related to the sequences from our Asian sample. In North America, this species was introduced as rootstock for cultivated roses and propagated for soil conservation. It is possible that the individuals introduced were already horticultural hybrids and not pure R. multiflora.
The only sub-Saharan species of the genus Rosa, R. abyssinica, is resolved by chloroplast data as embedded in the Cinnamomeae clade but clearly shows the synstyly consistent with R. sect. Synstylae. The nuclear GAPDH sequences resolve this species as closely related to R. gallica (R. sect. Rosa) and European species of R. sect. Synstylae, suggesting a hybrid origin.
Within the Synstylae clade, a few members of R. sect. Synstylae from Europe and North America, the only member of R. sect. Rosa and all the species sampled in R. sect. Caninae form a sub-clade. The North American R. setigera Michx. (R. sect. Synstylae) is supported as sister to all the other species of this sub-clade, all of which are European, but GAPDH sequences suggest it is more closely related to Asian species of R. sect. Synstylae. Species of R. sect. Synstylae of this clade (three species) are all diploids, while R. gallica (R. sect. Rosa) is a tetraploid and most members of R. sect. Caninae for which the ploidy level is known are usually pentaploids. This ploidy level is rather unusual especially for species with a low level of apomixis compared with sexual reproduction (Wissemann and Hellwig, 1997; Werlemark et al., 1999; Werlemark, 2000; Werlemark and Nybom, 2001; Nybom et al., 2004, 2006). Various authors (see Lim et al., 2005) have described a peculiar heterogamous meiosis in species of R. sect. Caninae, which more or less maintains pentaploidy. This heterogamous meiosis produces tetraploid ovules and haploid pollen grains because two sets of chromosomes form bivalents and segregate while three sets remain as univalents. Similar asymmetrically compensating allopolyploids are described in Onosma L. (Kolarčik et al., 2014).
Our results suggest that the genome of species in R. sect. Caninae is indeed complex, with two different kinds of chloroplast genomes and possibly a nuclear genome with three distinct origins (Figs 2 and 3; Table 1). Two chloroplast lineages within R. sect. Caninae were also reported by Wissemann and Ritz (2005), but they consider section Caninae as monophyletic based on the presence of a unique type of ITS sequences. The different sub-sections are not very well resolved by our phylogeny but De Riek et al. (2013) managed to delineate three sub-sections using a different approach. Our analyses indicate that, in R. sect. Caninae, several copies of GAPDH originate from R. sect. Cinnamomeae while the other copies originate from R. sect. Synstylae (Fig. 3; Table 1), which is consistent with results from microsatellite analyses (Zhang et al., 2013) and ITS analyses (Ritz et al., 2005).
The Pimpinellifoliae clade includes most species of R. sect. Pimpinellifoliae, but other species of this section occur in the Cinnamomeae clade or have an unresolved position (R. foetida). These results are similar to those of Matsumoto (2001) and Wissemann and Ritz (2005; chloroplast analysis).
The species resolved in the Pimpinellifoliae clade all are Asian and diploids (or of unknown ploidy), with a consistent morphology (Table 4) that usually included them in R. sect. Pimpinellifoliae (see Table 3). Some diploid species (R. farreri Cox and R. koreana Kom.) that were previously included in R. sect. Pimpinellifoliae but are resolved in the Cinnamomeae clade are morphologically distinct from other members of the section by having bracts and sometimes pink or red flowers. These distinct species seem to be genetically close to R. sect. Cinnamomeae as suggested by the GAPDH analyses and should probably be transferred to R. sect. Cinnamomeae, as noted in the taxonomic revision by Roberts (1977).
Table 4.
Comparative morphology of section Pimpinellifoliae (R. forrestiana from Cinnamomae is included for comparison)
| Position | Species | Ploidy level | Bracts | Petals | Prickles | Stipules margin | Sepals | Hip |
|---|---|---|---|---|---|---|---|---|
| Pimpinellifoliae Clade | R. omeiensis | 2 | Absent | White | Flat/terete | Sinuous, enrolled | Not leafy | Globose |
| R. sericea | 2 | Absent | White | Flat/terete | Sinuous, enrolled | Not leafy | Globose | |
| R. hugonis | 2 | Absent | Yellow | Flat/terete | Sinuous, enrolled | Not leafy | Globose | |
| R. xanthina | 2 | Absent | Yellow | Flat/terete | Sinuous, enrolled | Not leafy | Globose | |
| R. mairei | ? | Absent | White | Flat/terete | Sinuous, enrolled | Not leafy | Globose/ovoid | |
| R. taronensis | ? | Absent | White/yellow | Flat/terete | Sinuous, enrolled | Not leafy | Globose | |
| R. sikangensis | ? | Absent | White | Flat/terete | Sinuous, enrolled | Not leafy | Globose | |
| Unresolved | R. foetida | 4 | Sometimes present | Yellow | Terete | Sinuous, slightly enrolled | Leafy | Globose |
| Cinnamomae Clade | R. spinosissima | 4 | Absent | Pink/white/yellow | Terete | Sinuous, enrolled | Not leafy | Globose |
| R. tsinglingensis | ? | Sometimes present | White | Terete | Entire, not enrolled | Leafy | Ovoid | |
| R. farreri | 2 | Present | Pink/white | Terete | Entire, not enrolled | Leafy | Ovoid | |
| R. koreana | 2 | Present | Pinkish | Terete | Entire, not enrolled | Leafy | Ovoid | |
| R. forrestiana | 2 | Present | Red | Terete | Entire, not enrolled | Leafy | Ovoid |
Table 3.
Evolution of Pimpinellifoliae taxonomy and comparison with our results
| Rehder (1940) | Roberts (1977) | Wissemann (2003) | Flora of China (Gu and Robertson, 2003) | This study | Species |
|---|---|---|---|---|---|
| Pimpinellifoliae | Pimpinellifoliae | Pimpinellifoliae | Pimpinellifoliae | Pimpinellifoliae Clade | R. omeiensis |
| – | – | – | – | – | R. sericea |
| – | – | – | – | – | R. hugonis |
| – | – | – | – | – | R. xanthina |
| Not cited | Not cited | Not cited | – | – | R. mairei |
| – | – | – | – | – | R. taronensis |
| – | – | – | – | – | R. sikangensis |
| Pimpinellifoliae | Pimpinellifoliae | Pimpinellifoliae | – | Unresolved | R. foetida |
| – | – | – | – | Cinnamomae clade | R. spinosissima |
| Not cited | Not cited | Not cited | – | – | R. tsinglingensis |
| Pimpinellifoliae | Cinnamomae | – | – | – | R. farreri |
| – | – | Pimpinellifoliae | – | – | R. koreana |
Interestingly, two other species that were included in R. sect. Pimpinellifoliae but that are not in the Pimpinellifoliae clade are polyploids or of unknown ploidy. These species have some morphological characters consistent with R. sect. Pimpinellifoliae and some characters that are inconsistent with it (Table 4). We suspect that those species are allopolyploids. Our results from the nuclear GAPDH analyses (Fig. 3) confirm the presence of two different copies in R. spinosissima L. One of these copies is consistent with other members of the Pimpinellifoliae clade and the other is consistent with Cinnamomeae clade members particularly close to R. pendulina. Zhang et al. (2013) also suggested a close relationship between R. spinosissima and R. pendulina but could not find any evidence of a relationship between R. spinosissima and other members of R. sect. Pimpinellifoliae. Instead, the same authors and Wissemann and Ritz (2005) seem to find that some individuals from these species are genetically close to R. sect. Caninae. This could be explained by some degree of hybridization between those two kinds of polyploids.
Species of R. sect. Carolinae are embedded in the Cinnamomeae clade. Bruneau et al. (2007) discussed in detail the possible merging of R. sect. Carolinae in R. sect Cinnamomeae based on evidence from plastid DNA sequences but also morphological data (Lewis, 1957; Robertson, 1974), biochemical compounds (Grossi et al., 1998) and nuclear gene sequences (Joly et al., 2006).
Conspecific individuals are not always monophyletic in the Cinnamomeae clade. This may be related to incomplete lineage sorting due to rapid speciation events or to high incidence of polyploids with frequent hybridization in this clade. Indeed, recurrently formed allopolyploids from different maternal species, as reported for North American allopolyploid species (Joly et al., 2006), could explain this pattern. Members of the Cinnamomeae clade show a highly variable ploidy level. They mostly have an even ploidy level (2–10) but a few species have been reported occasionally to have an uneven ploidy level (e.g. Rosa acicularis Lindl., R. pendulina and R. willmottiae Hemsl.; Roberts et al., 2009). Alternatively, Joly (2012) has shown that it is not always possible to reject the null hypothesis of incomplete lineage sorting when testing for hybridization between non-monophyletic North American species, suggesting that a variety of processes might explain these patterns.
Polyploidy has been recognized as a prominent phenomenon in evolution and as an important cytogenetic mechanism in speciation (Wood et al., 2009). In the genus Rosa, hybridity is often accompanied by polyploidy and it may have helped stabilize hybrids between distantly related species of the genus. Polyploidy may also have favoured the rate of diversification (R. sect. Caninae), an increase in the geographic range (R. acicularis, R. spinosissima) and the colonization of high-altitude habitats (R. praelucens) or of high latitudes (R. acicularis). Ritz et al. (2011) suggested that the success of Caninae could be caused by its peculiar reproduction.
Biogeography
Our results (Fig. 4) suggest that the genus Rosa probably evolved during the Eocene in Asia and western North America, with the Bering Land Bridge enabling genetic exchanges between the two areas. This is supported by the previously cited oldest fossils of the genus Rosa, from Idaho and China (see the Materials and Methods), and by the presence of Rosa fossils in Alaska (see Becker, 1963; Hollick, 1936) from the Paleo-Eocene. At that time, the climate at those latitudes was temperate to warm temperate, which is consistent with the climate where most species of roses grow today. From Asia, approx. 30 Ma, the Synstylae lineage and its allies extended their distribution westward into Europe. This period corresponds to the closure of the Turgai strait that would have facilitated this migration. Following this western migration, part of the lineage extended its range into Eastern North America. The exchanges between Europe and Eastern North America persisted until the upper Miocene and were interrupted around 8 Ma. This is consistent with the hypothesis of Denk et al. (2011) based on fossils from Iceland that the North Atlantic Land Bridge had been available until 9–8 Ma by providing a sub-aerial route with mild conditions. Rosaceae fossils are known in this area but difficult to attribute to the genus Rosa.
Our ancestral area reconstruction suggests that despite an early presence on the American continent, most extant American species are the results of a later re-colonization from Asia. Therefore, R. subgen. Hesperhodos could be considered as a relic of this early American presence of the genus Rosa. The ancestors of the Pimpinellifoliae and Cinnamomeae clades seem to have been exclusively Asian.
The Cinnamomeae clade extended its distribution eastward, possibly through the Bering Land Bridge, into western North America then eastern North America. Exchanges between western North America and eastern North America seem to persist even today. The chloroplast data suggest fairly recent (until early Pliocene) exchanges between Asia and western North America. This suggests that the Bering Land Bridge may have been available for a long time for Rosa despite the climatic cooling that occurred during the Miocene. The absence of eastern North America–eastern Asia disjunction in our result contrasts with results from other plants (Donoghue and Smith, 2004). Those authors invoke greater extinction in western North America and Europe to explain eastern North America–eastern Asia disjunctions. However, our results suggest that roses may have not suffered such extinction.
European species of the Cinnamomeae clade seem to be the results of recent colonizations from Asia during the Pleistocene. The Caninae lineages diversification pre-dates those colonizations, which means it is unlikely that the Cinnamomeae type GAPDH sequences of species in sub-sect. Caninae and sub-sect. Rubigineae clades come from the European species of the Cinnamomeae clade. More probably, these Cinnamomeae type sequences come from previous genetic exchanges with Asian species of the Cinnamomeae clade during the latest Miocene or the Pliocene.
Exchanges between Asia and Africa occurred during the late Miocene, at some time between 9·3 Ma and 6·3 Ma. This period is consistent with the closure of the Parathetys, which enabled exchanges (Rögl, 1999). It is possible that these exchanges resulted in the capture of a chloroplast genome from a member of Cinnamomeae clade by R. abyssinica, whose nuclear GAPDH sequence is close to that of European species of the Synstylae clade. Some of those (R. sempervirens L., R. phoenicea Boiss.) occur today in Lebanon (both species), Syria and Israel (R. phoenicia).
Conclusions
With the most comprehensively sampled and well resolved phylogeny of the genus Rosa, we provide an evolutionary framework that will prove useful for the study of this difficult genus. Our phylogenetic results of the genus Rosa allow us to redefine sections in the genus Rosa (Table 5); however, a proper revision of the genus is needed to name and typify the newly defined sections. For example, since Wissemann’s (2003) publication, The type for the genus Rosa is now in R. sect. Cinnamomeae, which means that this section should be named R. sect. Rosa. We suggest that the subgenera could be treated at the sectional level and that R. sects Bracteatae, Laevigatae and Banksianae could be left unchanged. Rosa sect. Pimpinellifoliae should include members of our Pimpinellifoliae clade only, but this means that the type of the section would not be included. Rosa sects Cinnamomeae and Carolinae should be merged. There is a complex situation in the Synstylae clade due to the close relationship with the allopolyploid R. sect. Caninae. This allopolyploid section could be treated as one or two nothosections. Rosa sect. Synstylae should include R. sect. Indicae and include or exclude non-Asian members.
Table 5.
Taxonomic modifications proposed based on our phylogenetic results
| Wissemann (2003) | Tentative 1 | Tentative 2 |
|---|---|---|
| Subgenus: Hulthemia | Could be best treated as a section | |
| Subgenus: Hesperhodos | Could be best treated as a section | |
| Subgenus: Platyrhodon | Would be best treated as a section (see Flora of China; Gu and Robertson, 2003) | |
| Subgenus: Rosa | – | |
| Section: Bracteatae | Unchanged | |
| Section: Laevigatae | Unchanged | |
| Section: Banksianae | Unchanged | |
| Section: Pimpinellifoliae | Include members of our Pimpinellifoliae clade only | |
| Section: Cinnamomeae | Merge | |
| Section: Carolinae | ||
| Section: Synstylae: | Treat as four sections: | |
| - American (R. setigera) | R. setigera | |
| - European (R. sempervirens, R. arvensis) | R. sempervirens | |
| R. arvensis | Merge | |
| - Asian | Asian Synstylae (including Indicae) | |
| Section: Indicae | ||
| Section: Rosa | Unchanged | |
| Section: Caninae | – | Consider as a nothosection (Synstylae × Cinnamomae) |
| Sub-section: Rubrifoliae | Create a nothosection | Create a sub-nothosection |
| Sub-section: Vestitae | ||
| Sub-section: Trachyphyllae | ||
| Sub-section: Rubigineae | ||
| Sub-section: Tomentellae | ||
| Sub-section: Caninae | Create a nothosection | Create a sub-nothosection |
| Not included (putative intersectional hybrids): | ||
| R. spinosissima (Cinnamomae member × Pimpinellifoliae member?), | ||
| R. abyssinica (Cinnamomae member × European Synstylae member?), | ||
| R. praelucens (Cinnamomae member × Microphyllae member?), | ||
| R. foetida (more data needed) | ||
Tentative 1 uses monophyletic groups from the chloroplast phylogeny. Tentative 2 considers the results from chloroplast and nuclear data for a synthetic approach.
Our results also provide useful information for the studies of polyploids that are frequent in the genus. We formulate hypotheses about the origin of the genomes of several allopolyploids (R. sect. Caninae, R. praelucens and R. spinosissima) and diploid hybrids (R. abyssinica).
Our ancestral area reconstruction suggests that despite an early presence on the American continent, most extant American species are the results of a later re-colonization from Asia probably through the Bering Land Bridge. Our results suggest more recent exchanges between Asia and western North America than with eastern North America. The current distribution of roses from the Synstylae lineage in Europe is probably the result of a migration from Asia approx. 30 Ma ago, after the closure of the Turgai strait.
SUPPLEMENTARY DATA
Supplementary data are available online at www.aob.oxfordjournals.org and consist of the following. S1: list of samples used in this study with voucher information and GenBank accession numbers for the trnL region, the psbA-trnH intergenic spacer, the trnG region and GAPDH sequences. S2: dispersal matrices used for the DEC ancestral area reconstructions used in the four time-slice model (A, C, D, E) or the two time-slice model (A, B).
ACKNOWLEDGEMENTS
We thank our colleagues Yaling Cao, Yonghua He and Jian Gu from CIB who created the first wild rose garden in Maoxian Ecological Station 20 years ago; without their hard work this study would not have been possible. We also thank the Royal Botanic Gardens of Kew and Edinburgh, the Montreal Botanical Garden, Mark P. Simmons, Rene Smulder and Jan Wieringa for providing some samples. M.F.-D. thanks Pierre Honoré for his help with the figures, Luc Garraud, the Conservatoire Botanique Gap Charance, Marie-France Tarbouriech and René Roux for their help during field collection in France, and Michael McLeish for useful discussion about molecular dating. We also thank Anne Brysting and two anonymous reviewers for their useful comments. This work was supported by the National Natural Science Foundation of China [grant no. 31070173]; the Knowledge Innovation Program of the Chinese Academy of Sciences [grant no. KSCX2-EW-J-22]; the Project of Platform Construction for Plant Resources of Sichuan Province to X.F.-G.; a Natural Sciences and Engineering Research Council of Canada grant to A.B. and a Chinese Academy of Sciences Research Fellowship for International Young Researchers to M.F.-D.
APPENDIX
The network representing the relationships among copies of GAPDH obtained from Rosa species excluding obvious allopolyploids. The names of other polyploids are in bold. A ‘c’ followed by a number indicates the number attributed to one particular clone sequenced. The groups are compared with our main clades from the chloroplast analyses. Purple is attributed to Rosa subgen. Platyrhodon, yellow to R. sect. Banksianae, bright orange to R. sect. Bracteatae, brown to R. sect. Laevigatae, light pink to the Pimpinellifoliae clade, green to the Cinnamomeae clade and light blue to the Synstylae clade. Rosa abyssinica is highlighted with a light orange colour. Two types of copies, C1 and C2, are distinguished in our Cinnamomeae group.

LITERATURE CITED
- Anonymous. 1978. Fossils of China III. Cenozoic plants of China. Beijing: Science Press. [Google Scholar]
- Becker HF. 1963. The fossil record of the genus Rosa. Bulletin of the Torrey Botanical Club 90: 99–110. [Google Scholar]
- Brown RP, Yang Z. 2011. Rate variation and estimation of divergence times using strict and relaxed clocks. BMC Evolutionary Biology 11: 271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruneau A, Starr JR, Joly S. 2007. Phylogenetic relationships in the genus Rosa: new evidence from chloroplast DNA sequences and an appraisal of current knowledge. Systematic Botany 32: 366–378. [Google Scholar]
- Crépin F. 1889. Sketch of a new classification of roses. Journal of the Royal Horticultural Society 11: 217–228. [Google Scholar]
- Crépin F. 1891. Nouvelle classification des roses. Journal des Roses 15: 41–43, 53–55, 76–77. [Google Scholar]
- Cronn R, Cedroni M, Haselkorn T, Grover C, Wendel JF. 2002. PCR-mediated recombination in amplification products derived from polyploid cotton. Theoretical and Applied Genetics 104: 482–489. [DOI] [PubMed] [Google Scholar]
- Cutler RR. 2003. Medicinal and pharmaceutical uses. In: Roberts A, Debener T, Gudin S, eds. Encyclopedia of rose science. Amsterdam: Elsevier, 716–726. [Google Scholar]
- De Riek J, De Cock K, Smulders MJM, Nybom H. 2013. AFLP-based population structure analysis as a means to validate the complex taxonomy of dogroses (Rosa section Caninae). Molecular Phylogenetics and Evolution 67: 547–559. [DOI] [PubMed] [Google Scholar]
- Debener T, Linde M. 2009. Exploring complex ornamental ge nomes: the rose as a model plant. Critical Reviews in Plant Science 28: 267–280. [Google Scholar]
- Debener T, Bartels C, Mattiesch L. 1996. RAPD analysis of genetic variation between a group of rose cultivars and selected wild rose species. Molecular Breeding 2: 321–327. [Google Scholar]
- Denk T, Grimsson F, Zetter R, Simonarson LA. 2011. Late Cainozoic floras of Iceland: 15 million years of vegetation and climate history in the northern North Atlantic. Dordrecht, The Netherlands: Springer. [Google Scholar]
- Donoghue MJ, Smith SA. 2004. Patterns in the assembly of temperate forests around the Northern Hemisphere. Philosophical Transactions of the Royal Society B: Biological Sciences 359: 1633–1644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doyle JJ, Doyle JL. 1987. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin 19: 11–15. [Google Scholar]
- Drummond AJ, Ho SYW, Phillips MJ, Rambaut A. 2006. Relaxed phylogenetics and dating with confidence. PLoS Biology 4: e88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drummond AJ, Rambaut A. 2007. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evolutionary Biology 7: 214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edelman D. 1975. The Eocene Germer Basin Flora of South-Central Idaho. MS thesis, University of Idaho, Moscow, Idaho. [Google Scholar]
- Erlanson EW. 1929. Cytological conditions and evidences for hybridity in North American wild roses. Botanical Gazette 87: 443–506. [Google Scholar]
- Erlanson EW. 1934. Experimental data for a revision of the North American wild roses. Botanical Gazette 96: 197–259. [Google Scholar]
- Erlanson EW. 1938. Phylogeny and polyploidy in Rosa . New Phytologist 37: 72–81. [Google Scholar]
- Grossi C, Raymond O, Jay M. 1998. Flavonoid and enzyme polymorphisms and taxonomic organisation of Rosa sections: Carolinae, Cinnamomeae, Pimpinellifoliae and Synstylae. Biochemical Systematics and Ecology 26: 857–871. [Google Scholar]
- Gu C, Robertson KR. 2003. Rosa L. In: Team FoCe, ed. Flora of China. St. Louis, MO: Missouri Botanical Garden Press. [Google Scholar]
- Guimarães R, Barros L, Carvalho AM, Ferreira ICFR. 2010. Studies on chemical constituents and bioactivity of Rosa micrantha: an alternative antioxidants source for food, pharmaceutical, or cosmetic application. Journal of Agricultural and Food Chemistry 58: 6277–6284. [DOI] [PubMed] [Google Scholar]
- Guoliang W. 2003. Ancient Chinese roses. In: Roberts A, Debener T, Gudin S, eds. Encyclopedia of rose science. Amsterdam: Elsevier, 387–395. [Google Scholar]
- Hollick AA. 1936. The Tertiary flora of Alaska. United States Geological Survey Professional Paper 182: 1–185. [Google Scholar]
- Huson DH, Bryant D. 2006. Application of phylogenetic networks in evolutionary studies. Molecular Biology and Evolution 23: 254–267. [DOI] [PubMed] [Google Scholar]
- Iwata H, Kato T, Ohno S. 2000. Triparental origin of Damask roses. Gene 259: 53–59. [DOI] [PubMed] [Google Scholar]
- Jäger AK, Eldeen IMS, van Staden J. 2007. COX-1 and -2 activity of rose hip. Phytotherapy Research 21: 1251–1252. [DOI] [PubMed] [Google Scholar]
- Jan CH, Byrne DH, Manhart J, Wilson H. 1999. Rose germplasm analysis with RAPD markers. Hortscience 34: 341–345. [Google Scholar]
- Jian HY, Zhang H, Tang KX, et al. 2010. Decaploidy in Rosa praelucens Byhouwer (Rosaceae) endemic to Zhongdian Plateau, Yunnan, China. Caryologia 63: 162–167. [Google Scholar]
- Joly S. 2012. JML: testing hybridization from species trees. Molecular Ecology Resources 12: 179–184. [DOI] [PubMed] [Google Scholar]
- Joly S, Bruneau A. 2006. Incorporating allelic variation for reconstructing the evolutionary history of organisms from multiple genes: an example from Rosa in North America. Systematic Biology 55: 623–636. [DOI] [PubMed] [Google Scholar]
- Joly S, Starr JR, Lewis WH, Bruneau A. 2006. Polyploid and hybrid evolution in roses east of the Rocky Mountains. American Journal of Botany 93: 412–425. [DOI] [PubMed] [Google Scholar]
- Kellner A, Benner M, Walther H, Kunzmann L, Wissemann V, Ritz CM. 2012a. Leaf architecture of extant species of Rosa L. and the Paleogene species Rosa lignitum Heer (Rosaceae). International Journal of Plant Sciences 173: 239–250. [Google Scholar]
- Kellner A, Ritz CM, Wissemann V. 2012b. Hybridization with invasive Rosa rugosa threatens the genetic integrity of native Rosa mollis. Botanical Journal of the Linnean Society 170: 472–484. [Google Scholar]
- Kolarčik V, Zozomová-Lihová J, Ducár E, Mártonfi P. 2014. Evolutionary significance of hybridization in Onosma (Boraginaceae): analyses of stabilized hemisexual odd polyploids and recent sterile hybrids. Biological Journal of the Linnean Society 112: 89–107. [Google Scholar]
- Koopman WJL, Wissemann V, De Cock K, et al. 2008. AFLP markers as a tool to reconstruct complex relationships: a case study in Rosa (Rosaceae). American Journal of Botany 95: 353–366. [DOI] [PubMed] [Google Scholar]
- Lanfear R, Calcott B, Ho SYW, Guindon S. 2012. PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Molecular Biology and Evolution 29: 1695–1701. [DOI] [PubMed] [Google Scholar]
- Larkin MA, Blackshields G, Brown NP, et al. 2007. Clustal W and Clustal X version 2.0. Bioinformatics 23: 2947–2948. [DOI] [PubMed] [Google Scholar]
- Lewis WH. 1957. Revision of the genus Rosa in eastern North America: a review. American Rose Annual 42: 116–126. [Google Scholar]
- Lim KY, Werlemark G, Matyasek R, et al. 2005. Evolutionary implications of permanent odd polyploidy in the stable sexual, pentaploid of Rosa canina L. Heredity 94: 501–506. [DOI] [PubMed] [Google Scholar]
- Matsumoto S, Kouchi M, Yabuki J, Kusunoki M, Ueda Y, Fukui H. 1998. Phylogenetic analyses of the genus Rosa using the matK sequence: molecular evidence for the narrow genetic background of modern roses. Scientia Horticulturae 77: 73–82. [Google Scholar]
- Matsumoto S, Kouchi M, Fukui H. 2000. Phylogenetic analyses of the subgenus Eurosa using the ITS nrDNA sequence. Acta Horticulturae 521: 193–202. [Google Scholar]
- Matsumoto S, Nishio H, Fukui H. 2001. Phylogenetic analyses of genus Rosa: polyphyly of section Pimpinellifoliae and origin of Rosa × fortuniana. Acta Horticulturae 547: 357–363. [Google Scholar]
- Meng J, Fougère-Danezan M, Zhang L-B, Li D-Z, Yi T-S. 2011. Untangling the hybrid origin of the Chinese tea roses: evidence from DNA sequences of single-copy nuclear and chloroplast genes. Plant Systematics and Evolution 297: 157–170. [Google Scholar]
- Meng Q, Liu Z, Bruch AA, Liu R, Hu F. 2012. Palaeoclimatic evolution during Eocene and its influence on oil shale mineralisation, Fushun basin, China. Journal of Asian Earth Sciences 45: 95–105. [Google Scholar]
- Mercure M, Bruneau A. 2008. Hybridization between the escaped Rosa rugosa (Rosaceae) and native R . blanda in Eastern North America. American Journal of Botany 95: 597–607. [DOI] [PubMed] [Google Scholar]
- Millan T, Osuna F, Cobos S, Torres A, Cubero J. 1996. Using RAPDs to study phylogenetic relationships in Rosa. Theoretical and Applied Genetics 92: 273–277. [DOI] [PubMed] [Google Scholar]
- Nybom H, Esselink GD, Werlemark G, Vosman B. 2004. Microsatellite DNA marker inheritance indicates preferential pairing between two highly homologous genomes in polyploid and hemisexual dog-roses, Rosa L. Sect. Caninae DC. Heredity 92: 139–150. [DOI] [PubMed] [Google Scholar]
- Nybom H, Esselink GD, Werlemark G, Leus L, Vosman B. 2006. Unique genomic configuration revealed by microsatellite DNA in polyploid dogroses, Rosa sect. Caninae. Journal of Evolutionary Biology 19: 635–648. [DOI] [PubMed] [Google Scholar]
- Qiu X-Q, Zhang H, Wang Q-G, et al. 2012. Phylogenetic relationships of wild roses in China based on nrDNA and matK data. Scientia Horticulturae 140: 45–51. [Google Scholar]
- Rambaut A, Suchard M, Xie D, Drummond AJ. 2013. Tracer v1.5. Available from http://beast.bio.ed.ac.uk/Tracer (last accessed 5 January 2013). [Google Scholar]
- Ree RH, Smith SA. 2008. Maximum likelihood inference of geographic range evolution by dispersal, local extinction, and cladogenesis. Systematic Biology 57: 4–14. [DOI] [PubMed] [Google Scholar]
- Rehder A. 1940. Manual of cultivated trees and shrubs hardy in North America. New York: Macmillan. [Google Scholar]
- Ritz CM, Wissemann V. 2011. Microsatellites analyses of artificial and spontaneous dogrose hybrids reveal the hybridogenic origin of Rosa micrantha by the contribution of unreduced gametes. Journal of Heredity 102: 217–227. [DOI] [PubMed] [Google Scholar]
- Ritz CM, Schmuths H, Wissemann V. 2005. Evolution by reticulation: European dogroses originated by multiple hybridization across the genus Rosa. Journal of Heredity 96: 4–14. [DOI] [PubMed] [Google Scholar]
- Ritz CM, Koehnen I, Groth M, Theissen G, Wissemann V. 2011. To be or not to be the odd one out – allele-specific transcription in pentaploid dogroses (Rosa L. sect. Caninae (DC.) Ser). BMC Plant Biology 11: 37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts AV. 1977. Relationship between species in the genus Rosa, section Pimpinellifoliae. Botanical Journal of the Linnean Society 74: 309–328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts AV, Gladis T, Brumme H. 2009. DNA amounts of roses (Rosa L.) and their use in attributing ploidy levels. Plant Cell Reports 28: 61–71. [DOI] [PubMed] [Google Scholar]
- Robertson KR. 1974. Genera of Rosaceae in southeastern United-States. Journal of the Arnold Arboretum 55: 611–662. [Google Scholar]
- Rögl F. 1999. Mediterranean and Paratethys. Facts and hypotheses of an Oligocene to Miocene paleogeography (short overview). Geologica Carpathica 50: 339–349. [Google Scholar]
- Russell A, Samuel R, Klejna V, Barfuss MHJ, Rupp B, Chase MW. 2010. Reticulate evolution in diploid and tetraploid species of Polystachya (Orchidaceae) as shown by plastid DNA sequences and low-copy nuclear genes. Annals of Botany 106: 37–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sang T, Crawford DJ, Stuessy TF. 1997. Chloroplast DNA phylogeny, reticulate evolution, and biogeography of Paeonia (Paeoniaceae). American Journal of Botany 84: 1120–1136. [PubMed] [Google Scholar]
- Scariot V, Akkak A, Botta R. 2006. Characterization and genetic relationships of wild species and old garden roses based on microsatellite analysis. Journal of the American Society for Horticultural Science 131: 66–73. [Google Scholar]
- Schanzer IA, Kutlunina NA. 2010. Interspecific hybridization in wild roses (Rosa L. sect. Caninae DC.). Biology Bulletin 37: 480–488. [PubMed] [Google Scholar]
- Schanzer IA, Vagina AV. 2007. ISSR (inter simple sequence repeat) markers reveal natural intersectional hybridization in wild roses [Rosa L., sect. Caninae (DC.) Ser. and sect. Cinnamomeae (DC.) Ser.]. Wulfenia 14: 1–14. [Google Scholar]
- Shaw J, Lickey E, Beck J, et al. 2005. The tortoise and the hare II: relative utility of 21 noncoding chloroplast DNA sequences for phylogenetic analysis. American Journal of Botany 92: 142–166. [DOI] [PubMed] [Google Scholar]
- Shaw J, Lickey EB, Schilling EE, Small RL. 2007. Comparion of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. American Journal of Botany 94: 275–288. [DOI] [PubMed] [Google Scholar]
- Stamatakis A. 2006. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22: 2688–2690. [DOI] [PubMed] [Google Scholar]
- Stamatakis A, Hoover P, Rougemont J. 2008. A rapid bootstrap algorithm for the RAxML web servers. Systematic Biology 57: 758–771. [DOI] [PubMed] [Google Scholar]
- Strand A, LeebensMack J, Milligan B. 1997. Nuclear DNA-based markers for plant evolutionary biology. Molecular Ecology 6: 113–118. [DOI] [PubMed] [Google Scholar]
- Taberlet P, Gielly L, Pautou G, Bouvet J. 1991. Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Molecular Biology 17: 1105–1109. [DOI] [PubMed] [Google Scholar]
- Tamura K, Dudley J, Nei M, Kumar S. 2007. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Molecular Biology and Evolution 24: 1596–1599. [DOI] [PubMed] [Google Scholar]
- Werlemark G. 2000. Evidence of apomixis in hemisexual dogroses, Rosa section Caninae. Sexual Plant Reproduction 12: 353–359. [Google Scholar]
- Werlemark G, Nybom H. 2001. Skewed distribution of morphological character scores and molecular markers in three interspecific crosses in Rosa section Caninae. Hereditas 134: 1–13. [DOI] [PubMed] [Google Scholar]
- Werlemark G, Uggla M, Nybom H. 1999. Morphological and RAPD markers show a highly skewed distribution in a pair of reciprocal crosses between hemisexual dogrose species, Rosa sect. Caninae. Theoretical and Applied Genetics 98: 557–563. [Google Scholar]
- Wissemann V. 2003. Conventional taxonomy (wild roses). In: Roberts AV, Debener T, Gudin S, eds. Encyclopedia of rose science. Amsterdam: Elsevier, 111–117. [Google Scholar]
- Wissemann V, Hellwig FH. 1997. Reproduction and hybridisation in the genus Rosa, section Caninae (Ser) Rehd. Botanica Acta 110: 251–256. [Google Scholar]
- Wissemann V, Ritz CM. 2005. The genus Rosa (Rosoideae, Rosaceae) revisited: molecular analysis of nrITS-1 and atpB-rbcL intergenic spacer (IGS) versus conventional taxonomy. Botanical Journal of the Linnean Society 147: 275–290. [Google Scholar]
- Wood TE, Takebayashi N, Barker MS, Mayrose I, Greenspoon PB, Rieseberg LH. 2009. The frequency of polyploid speciation in vascular plants. Proceedings of the National Academy of Sciences, USA 106: 13875–13879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu S, Ueda Y, Nishihara S, Matsumoto S. 2000. Phylogenetic analysis of Japanese Rosa using matK sequences. Breeding Science 50: 275–281. [Google Scholar]
- Wu S, Ueda Y, Nishihara S, Matsumoto S. 2001. Phylogenetic analysis of Japanese Rosa species using DNA sequences of nuclear ribosomal internal transcribed spacer (ITS). Journal of Horticultural Science and Biotechnology 76: 127–132. [Google Scholar]
- Yi OS, Jovel EM, Towers GHN, Wahbe TR, Cho DW. 2007. Antioxidant and antimicrobial activities of native Rosa sp. from British Columbia, Canada. International Journal of Food Sciences and Nutrition 58: 178–189. [DOI] [PubMed] [Google Scholar]
- Yokoya K, Roberts AV, Mottley J, Lewis R, Brandham PE. 2000. Nuclear DNA amounts in roses. Annals of Botany 85: 557–561. [Google Scholar]
- Yu Y, Harris A, He X-J. 2013. RASP (Reconstruct Ancestral State in Phylogenies) 2.1 beta. Available at http://mnh.scu.edu.cn/soft/blog/RASP (last accessed 22 August 2013). [DOI] [PubMed] [Google Scholar]
- Zhang J, Esselink GD, Che D, Fougère-Danezan M, Arens P, Smulders MJM. 2013. The diploid origins of allopolyploid rose species studied using single nucleotide polymorphism haplotypes flanking a microsatellite repeat. Journal of Horticultural Science and Biotechnology 88: 85–92. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.