Abstract
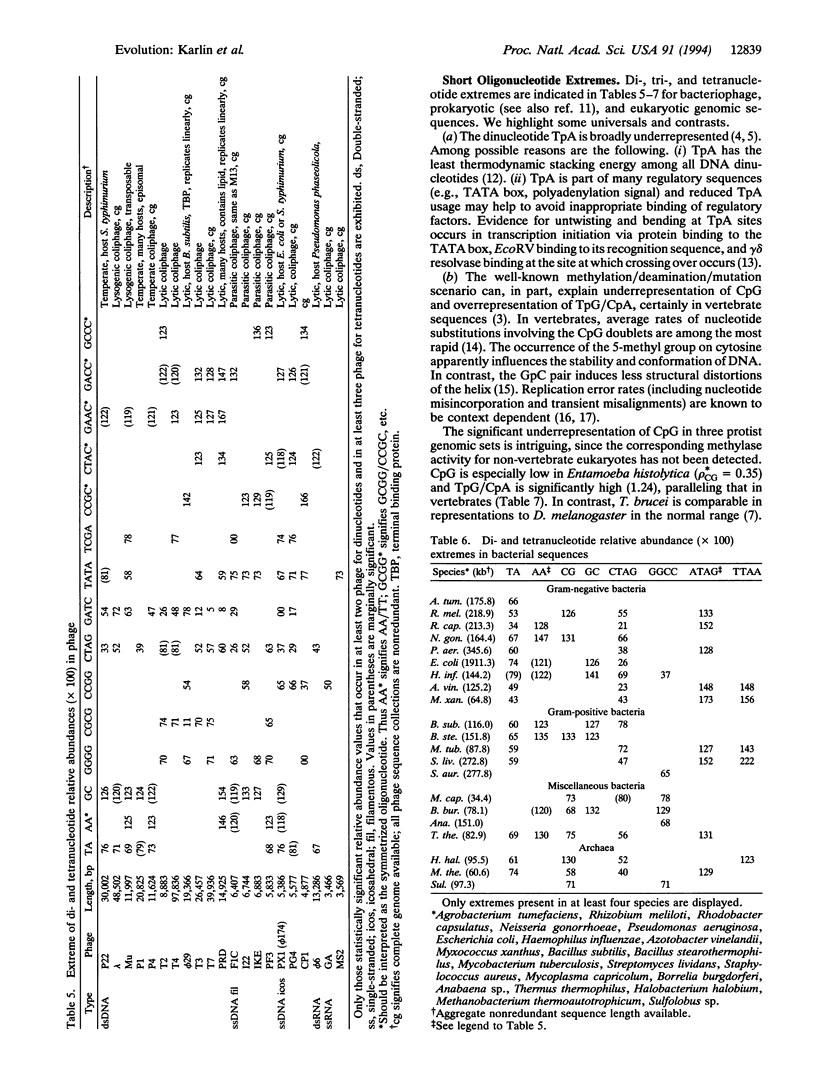
Genomic homogeneity is investigated for a broad base of DNA sequences in terms of dinucleotide relative abundance distances (abbreviated delta-distances) and of oligonucleotide compositional extremes. It is shown that delta-distances between different genomic sequences in the same species are low, only about 2 or 3 times the distance found in random DNA, and are generally smaller than the between-species delta-distances. Extremes in short oligonucleotides include underrepresentation of TpA and overrepresentation of GpC in most temperate bacteriophage sequences; underrepresentation of CTAG in most eubacterial genomes; underrepresentation of GATC in most bacteriophage; CpG suppression in vertebrates, in all animal mitochondrial genomes, and in many thermophilic bacterial sequences; and overrepresentation of GpG/CpC in all animal mitochondrial sets and chloroplast genomes. Interpretations center on DNA structures (dinucleotide stacking energies, DNA curvature and superhelicity, nucleosome organization), context-dependent mutational events, methylation effects, and processes of replication and repair.
Full text
PDF



Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Beutler E., Gelbart T., Han J. H., Koziol J. A., Beutler B. Evolution of the genome and the genetic code: selection at the dinucleotide level by methylation and polyribonucleotide cleavage. Proc Natl Acad Sci U S A. 1989 Jan;86(1):192–196. doi: 10.1073/pnas.86.1.192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blumenthal A. B., Kriegstein H. J., Hogness D. S. The units of DNA replication in Drosophila melanogaster chromosomes. Cold Spring Harb Symp Quant Biol. 1974;38:205–223. doi: 10.1101/sqb.1974.038.01.024. [DOI] [PubMed] [Google Scholar]
- Breslauer K. J., Frank R., Blöcker H., Marky L. A. Predicting DNA duplex stability from the base sequence. Proc Natl Acad Sci U S A. 1986 Jun;83(11):3746–3750. doi: 10.1073/pnas.83.11.3746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burge C., Campbell A. M., Karlin S. Over- and under-representation of short oligonucleotides in DNA sequences. Proc Natl Acad Sci U S A. 1992 Feb 15;89(4):1358–1362. doi: 10.1073/pnas.89.4.1358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dickerson R. E. DNA structure from A to Z. Methods Enzymol. 1992;211:67–111. doi: 10.1016/0076-6879(92)11007-6. [DOI] [PubMed] [Google Scholar]
- Echols H., Goodman M. F. Fidelity mechanisms in DNA replication. Annu Rev Biochem. 1991;60:477–511. doi: 10.1146/annurev.bi.60.070191.002401. [DOI] [PubMed] [Google Scholar]
- Heinemann J. A., Sprague G. F., Jr Bacterial conjugative plasmids mobilize DNA transfer between bacteria and yeast. Nature. 1989 Jul 20;340(6230):205–209. doi: 10.1038/340205a0. [DOI] [PubMed] [Google Scholar]
- Hess S. T., Blake J. D., Blake R. D. Wide variations in neighbor-dependent substitution rates. J Mol Biol. 1994 Mar 4;236(4):1022–1033. doi: 10.1016/0022-2836(94)90009-4. [DOI] [PubMed] [Google Scholar]
- Hunter C. A. Sequence-dependent DNA structure. The role of base stacking interactions. J Mol Biol. 1993 Apr 5;230(3):1025–1054. doi: 10.1006/jmbi.1993.1217. [DOI] [PubMed] [Google Scholar]
- Karlin S., Burge C., Campbell A. M. Statistical analyses of counts and distributions of restriction sites in DNA sequences. Nucleic Acids Res. 1992 Mar 25;20(6):1363–1370. doi: 10.1093/nar/20.6.1363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karlin S., Campbell A. M. Which bacterium is the ancestor of the animal mitochondrial genome? Proc Natl Acad Sci U S A. 1994 Dec 20;91(26):12842–12846. doi: 10.1073/pnas.91.26.12842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karlin S., Cardon L. R. Computational DNA sequence analysis. Annu Rev Microbiol. 1994;48:619–654. doi: 10.1146/annurev.mi.48.100194.003155. [DOI] [PubMed] [Google Scholar]
- Karlin S., Ladunga I. Comparisons of eukaryotic genomic sequences. Proc Natl Acad Sci U S A. 1994 Dec 20;91(26):12832–12836. doi: 10.1073/pnas.91.26.12832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karlin S., Mocarski E. S., Schachtel G. A. Molecular evolution of herpesviruses: genomic and protein sequence comparisons. J Virol. 1994 Mar;68(3):1886–1902. doi: 10.1128/jvi.68.3.1886-1902.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krysan P. J., Smith J. G., Calos M. P. Autonomous replication in human cells of multimers of specific human and bacterial DNA sequences. Mol Cell Biol. 1993 May;13(5):2688–2696. doi: 10.1128/mcb.13.5.2688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kunkel T. A. Biological asymmetries and the fidelity of eukaryotic DNA replication. Bioessays. 1992 May;14(5):303–308. doi: 10.1002/bies.950140503. [DOI] [PubMed] [Google Scholar]
- Lupski J. R., Weinstock G. M. Short, interspersed repetitive DNA sequences in prokaryotic genomes. J Bacteriol. 1992 Jul;174(14):4525–4529. doi: 10.1128/jb.174.14.4525-4529.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merkl R., Kröger M., Rice P., Fritz H. J. Statistical evaluation and biological interpretation of non-random abundance in the E. coli K-12 genome of tetra- and pentanucleotide sequences related to VSP DNA mismatch repair. Nucleic Acids Res. 1992 Apr 11;20(7):1657–1662. doi: 10.1093/nar/20.7.1657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nussinov R. Theoretical molecular biology: prospectives and perspectives. J Theor Biol. 1987 Mar 21;125(2):219–235. doi: 10.1016/s0022-5193(87)80043-7. [DOI] [PubMed] [Google Scholar]
- Rafferty J. B., Somers W. S., Saint-Girons I., Phillips S. E. Three-dimensional crystal structures of Escherichia coli met repressor with and without corepressor. Nature. 1989 Oct 26;341(6244):705–710. doi: 10.1038/341705a0. [DOI] [PubMed] [Google Scholar]
- Selker E. U. Premeiotic instability of repeated sequences in Neurospora crassa. Annu Rev Genet. 1990;24:579–613. doi: 10.1146/annurev.ge.24.120190.003051. [DOI] [PubMed] [Google Scholar]
- Travers A. A., Schwabe J. W. Spurring on transcription? Curr Biol. 1993 Dec 1;3(12):898–900. doi: 10.1016/0960-9822(93)90231-c. [DOI] [PubMed] [Google Scholar]



