Abstract
The binuclear title complex, [Cu2I2(C26H24P2)2(C5H12N2S)2], lies about an inversion centre. The CuI atom displays a distorted tetrahedral coordination geometry defined by one S atom of an N,N′-diethylthiourea ligand, two P atoms derived from two bridging 1,2-bis(diphenylphosphanyl)ethane (dppe) ligands and one iodide ion. The dppe ligand bridges two symmetry-related CuI ions, forming a 10-membered Cu2P4C4 ring. An intramolecular N—H⋯I hydrogen bond is noted. In the crystal, N—H⋯I hydrogen bonds link complex molecules into layers parallel to (-101).
Keywords: crystal structure; copper(I) complex; N,N′-diethylthiourea; N—H⋯I hydrogen bonding
Related literature
For background to the coordination chemistry of copper(I) halides and pseudohalides, see: Dennehy et al. (2011 ▸); Oshio et al. (1996 ▸); Seward et al. (2003 ▸). For their potential applications, see: Corey et al. (1987 ▸); Dias et al. (2006 ▸). For relevant examples of discrete complexes, see: Dennehy et al. (2009 ▸).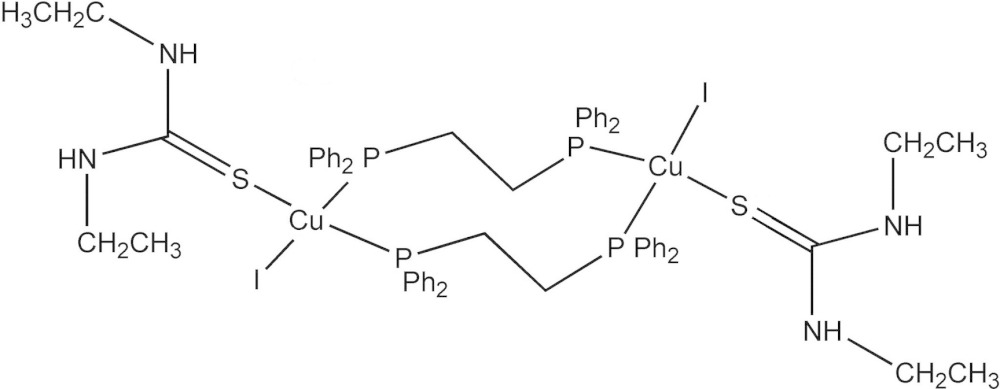
Experimental
Crystal data
[Cu2I2(C26H24P2)2(C5H12N2S)2]
M r = 1442.11
Monoclinic,

a = 12.2150 (8) Å
b = 15.1836 (9) Å
c = 17.1801 (10) Å
β = 96.414 (2)°
V = 3166.4 (3) Å3
Z = 2
Cu Kα radiation
μ = 10.37 mm−1
T = 100 K
0.16 × 0.15 × 0.08 mm
Data collection
Bruker Prospector CCD diffractometer
Absorption correction: multi-scan (SADABS; Bruker, 2013 ▸) T min = 0.433, T max = 0.753
23236 measured reflections
5564 independent reflections
5556 reflections with I > 2σ(I)
R int = 0.028
Refinement
R[F 2 > 2σ(F 2)] = 0.025
wR(F 2) = 0.063
S = 1.13
5564 reflections
345 parameters
H-atom parameters constrained
Δρmax = 0.98 e Å−3
Δρmin = −0.92 e Å−3
Data collection: APEX2 (Bruker, 2013 ▸); cell refinement: SAINT (Bruker, 2013 ▸); data reduction: SAINT; program(s) used to solve structure: SHELXS97 (Sheldrick, 2008 ▸); program(s) used to refine structure: SHELXL2015 (Sheldrick, 2015 ▸) and SHELXLE (Hübschle et al., 2011 ▸); molecular graphics: Mercury (Macrae et al., 2008 ▸); software used to prepare material for publication: publCIF (Westrip, 2010 ▸).
Supplementary Material
Crystal structure: contains datablock(s) I. DOI: 10.1107/S2056989015014176/tk5371sup1.cif
Structure factors: contains datablock(s) I. DOI: 10.1107/S2056989015014176/tk5371Isup2.hkl
. DOI: 10.1107/S2056989015014176/tk5371fig1.tif
The structure of title complex with displacement ellipsoids drawn at the 50% propbability level. All H atoms are omitted for clarity.
. DOI: 10.1107/S2056989015014176/tk5371fig2.tif
Part of the crystal structure showing intra/inter-molecular N—H⋯I hydrogen bonds forming a layers as dashed lines.
CCDC reference: 1415379
Additional supporting information: crystallographic information; 3D view; checkCIF report
Table 1. Hydrogen-bond geometry (, ).
| DHA | DH | HA | D A | DHA |
|---|---|---|---|---|
| N1H1I1i | 0.88 | 2.80 | 3.622(2) | 156 |
| N2H2I1 | 0.88 | 2.70 | 3.5517(19) | 162 |
Symmetry code: (i)  .
.
Acknowledgments
We gratefully acknowledge financial support from the Strategic Scholarships Fellowships Frontier Research Networks (Specific for Southern Region), the Commission on Higher Education, Ministry of Education, and the Department of Chemistry and Graduate School, Prince of Songkla University. LK would like to thank Dr Matthias Zeller of Youngstown State University, Ohio, USA, for suggestions and assistance with the X-ray structure refinement.
supplementary crystallographic information
S0.1. Synthesis and crystallization
N,N'-Diethylthiourea (0.07 g, 0.5 mmol) was dissolved in 30 cm3 of acetonitrile in a round flask equipped with reflux condenser and magnetic stirrer at 333 K and then CuI (0.1 g, 0.5 mmol) was added. The mixture was stirred for 2 h. 1,2-bis(diphenylphosphanyl)ethane (0.2 g, 0.5 mmol) was added and the reaction mixture was heated under reflux for 5 h where upon the precipitate gradually disappeared. The resulting clear solution was filtered and left to evaporate at room temperature. The colorless crystals, which deposited after standing for several days were filtered off and washed with acetone and dried in vacuo (M. pt = 557 K). Elemental analysis, calculated for [Cu2I2(C26H24P2)2(C5H12N2S)2]: C, 51.59; H, 4.97; N, 3.88; S, 4.44%, found: C, 55.69; H, 5.22; N, 3.62; S, 4.61%.
S0.2. Refinement
The (-1 8 3) reflection was affected by the beam-stop and was omitted from the final cycles of refinement. H atoms bonded to C and N atoms were included in their calculated positions and were refined using a riding model using bond lengths of 0.95–0.99 Å and Uiso(H) = 1.2–1.5Ueq(C), and N—H = 0.88 Å (NH) and Uiso(H) = 1.2Ueq(N). The (-1 8 3) reflection was omitted owing to poor agreement.
S1. Comment
Coordination complexes of copper(I) halides or pseudo-halides with mixed P and S donor ligands have been of interest in coordination chemistry (Dennehy et al., 2011; Oshio et al., 1996; Sewead et al., 2003) due to their applications such as magnetism (Oshio et al., 1996) and biological or medicinal activities (Corey et al., 1987; Dias et al., 2006). In this work, a mixed ligand complex of copper(I) iodide with 1,2-bis(diphenylphosphanyl)ethane (dppe) and N,N'-diethylthiourea (detu) is reported. The binuclear copper(I) complex lies across an inversion center. The µ2-dppe bridges between CuI centers leads to a 10-membered Cu2P4C4 rhomboid, see Fig. 1. The Cu1—P1 and Cu1—P2 bond lengths are 2.2681 (6) and 2.2813 (6)Å, respectively. These values are slightly shorter than the equivalent distances found in [Cu(tsac)(PPh3)2], [Cu4(tsac)4(PPh3)3], [Cu2(tsac)2(dppm)2] and [Cu4(tsac)4(dppm)2], which are in the range 2.2799 (5) and 2.3119 (5) Å (Dennehy et al., 2009). There is an intramolecular N2—H2···I1 hydrogen bond. In the crystal, intermolecular N1—H1···I1 hydrogen bonds link complex molecules into a two-dimensional supramolecular network parallel to (-101) (Fig. 2. and Table 1)
Figures
Fig. 1.

The structure of title complex with displacement ellipsoids drawn at the 50% propbability level. All H atoms are omitted for clarity.
Fig. 2.

Part of the crystal structure showing intra/inter-molecular N—H···I hydrogen bonds forming a layers as dashed lines.
Crystal data
| [Cu2I2(C26H24P2)2(C5H12N2S)2] | F(000) = 1456 |
| Mr = 1442.11 | Dx = 1.513 Mg m−3 |
| Monoclinic, P21/n | Cu Kα radiation, λ = 1.54178 Å |
| a = 12.2150 (8) Å | Cell parameters from 9846 reflections |
| b = 15.1836 (9) Å | θ = 3.9–66.7° |
| c = 17.1801 (10) Å | µ = 10.37 mm−1 |
| β = 96.414 (2)° | T = 100 K |
| V = 3166.4 (3) Å3 | Block, colourless |
| Z = 2 | 0.16 × 0.15 × 0.08 mm |
Data collection
| Bruker Prospector CCD diffractometer | 5564 independent reflections |
| Radiation source: I-mu-S microsource X-ray tube | 5556 reflections with I > 2σ(I) |
| Laterally graded multilayer (Goebel) mirror monochromator | Rint = 0.028 |
| ω and phi scans | θmax = 67.0°, θmin = 3.9° |
| Absorption correction: multi-scan (SADABS; Bruker, 2013) | h = −14→12 |
| Tmin = 0.433, Tmax = 0.753 | k = −18→17 |
| 23236 measured reflections | l = −20→20 |
Refinement
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.025 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.063 | H-atom parameters constrained |
| S = 1.13 | w = 1/[σ2(Fo2) + (0.0315P)2 + 3.4263P] where P = (Fo2 + 2Fc2)/3 |
| 5564 reflections | (Δ/σ)max = 0.003 |
| 345 parameters | Δρmax = 0.98 e Å−3 |
| 0 restraints | Δρmin = −0.92 e Å−3 |
Special details
| Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | ||
| I1 | 0.71658 (2) | 0.56133 (2) | 0.16309 (2) | 0.01039 (6) | |
| Cu1 | 0.51704 (2) | 0.64264 (2) | 0.12681 (2) | 0.00798 (8) | |
| S1 | 0.48147 (4) | 0.74104 (3) | 0.22620 (3) | 0.01238 (12) | |
| P1 | 0.39142 (4) | 0.53264 (3) | 0.12684 (3) | 0.00711 (11) | |
| P2 | 0.52134 (4) | 0.70642 (3) | 0.00696 (3) | 0.00768 (11) | |
| N1 | 0.58767 (16) | 0.88958 (13) | 0.27366 (12) | 0.0163 (4) | |
| H1 | 0.6493 | 0.9171 | 0.2902 | 0.020* | |
| N2 | 0.69626 (15) | 0.77334 (13) | 0.24764 (11) | 0.0122 (4) | |
| H2 | 0.7019 | 0.7167 | 0.2380 | 0.015* | |
| C1 | 0.59546 (18) | 0.80593 (15) | 0.25058 (12) | 0.0109 (4) | |
| C2 | 0.4851 (2) | 0.93892 (15) | 0.27369 (17) | 0.0198 (5) | |
| H2A | 0.4929 | 0.9817 | 0.3174 | 0.024* | |
| H2B | 0.4250 | 0.8977 | 0.2827 | 0.024* | |
| C3 | 0.4547 (3) | 0.9875 (2) | 0.19789 (19) | 0.0348 (7) | |
| H3A | 0.5135 | 1.0289 | 0.1891 | 0.052* | |
| H3B | 0.3859 | 1.0199 | 0.2007 | 0.052* | |
| H3C | 0.4448 | 0.9452 | 0.1546 | 0.052* | |
| C4 | 0.79787 (18) | 0.82441 (16) | 0.25912 (14) | 0.0149 (5) | |
| H4A | 0.8076 | 0.8487 | 0.3129 | 0.018* | |
| H4B | 0.7936 | 0.8743 | 0.2218 | 0.018* | |
| C5 | 0.89514 (18) | 0.76646 (16) | 0.24646 (14) | 0.0162 (5) | |
| H5A | 0.8813 | 0.7370 | 0.1956 | 0.024* | |
| H5B | 0.9055 | 0.7222 | 0.2881 | 0.024* | |
| H5C | 0.9617 | 0.8027 | 0.2475 | 0.024* | |
| C11 | 0.38548 (18) | 0.47611 (14) | 0.22045 (13) | 0.0103 (4) | |
| C12 | 0.45710 (18) | 0.50116 (14) | 0.28549 (13) | 0.0124 (4) | |
| H12 | 0.5106 | 0.5456 | 0.2807 | 0.015* | |
| C13 | 0.4500 (2) | 0.46061 (16) | 0.35789 (13) | 0.0161 (5) | |
| H13 | 0.4989 | 0.4777 | 0.4022 | 0.019* | |
| C14 | 0.3723 (2) | 0.39590 (16) | 0.36541 (14) | 0.0196 (5) | |
| H14 | 0.3673 | 0.3692 | 0.4149 | 0.024* | |
| C15 | 0.3015 (2) | 0.36998 (17) | 0.30053 (15) | 0.0201 (5) | |
| H15 | 0.2486 | 0.3250 | 0.3055 | 0.024* | |
| C16 | 0.30793 (19) | 0.40978 (16) | 0.22830 (14) | 0.0149 (5) | |
| H16 | 0.2594 | 0.3918 | 0.1840 | 0.018* | |
| C21 | 0.24971 (18) | 0.57205 (14) | 0.10553 (14) | 0.0109 (4) | |
| C22 | 0.1845 (2) | 0.55542 (16) | 0.03495 (15) | 0.0188 (5) | |
| H22 | 0.2117 | 0.5190 | −0.0035 | 0.023* | |
| C23 | 0.0792 (2) | 0.59235 (18) | 0.02086 (16) | 0.0240 (5) | |
| H23 | 0.0359 | 0.5819 | −0.0277 | 0.029* | |
| C24 | 0.0375 (2) | 0.64380 (17) | 0.07685 (17) | 0.0246 (6) | |
| H24 | −0.0346 | 0.6678 | 0.0674 | 0.029* | |
| C25 | 0.1021 (2) | 0.66008 (18) | 0.14715 (17) | 0.0245 (6) | |
| H25 | 0.0739 | 0.6953 | 0.1860 | 0.029* | |
| C26 | 0.2072 (2) | 0.62539 (17) | 0.16094 (14) | 0.0177 (5) | |
| H26 | 0.2510 | 0.6381 | 0.2088 | 0.021* | |
| C27 | 0.39162 (19) | 0.44094 (13) | 0.05650 (13) | 0.0098 (4) | |
| H27A | 0.3228 | 0.4068 | 0.0578 | 0.012* | |
| H27B | 0.3910 | 0.4658 | 0.0032 | 0.012* | |
| C28 | 0.51050 (17) | 0.62207 (14) | −0.07137 (12) | 0.0090 (4) | |
| H28A | 0.4406 | 0.5891 | −0.0713 | 0.011* | |
| H28B | 0.5110 | 0.6507 | −0.1231 | 0.011* | |
| C31 | 0.40910 (19) | 0.78228 (15) | −0.02656 (12) | 0.0126 (4) | |
| C32 | 0.4258 (2) | 0.87216 (18) | −0.03567 (19) | 0.0302 (6) | |
| H32 | 0.4977 | 0.8963 | −0.0252 | 0.036* | |
| C33 | 0.3369 (3) | 0.9266 (2) | −0.0601 (2) | 0.0432 (8) | |
| H33 | 0.3488 | 0.9880 | −0.0658 | 0.052* | |
| C34 | 0.2325 (3) | 0.8932 (2) | −0.07599 (18) | 0.0343 (7) | |
| H34 | 0.1728 | 0.9310 | −0.0933 | 0.041* | |
| C35 | 0.2148 (2) | 0.8042 (2) | −0.06672 (16) | 0.0279 (6) | |
| H35 | 0.1427 | 0.7805 | −0.0779 | 0.033* | |
| C36 | 0.3023 (2) | 0.74917 (18) | −0.04106 (15) | 0.0205 (5) | |
| H36 | 0.2893 | 0.6883 | −0.0333 | 0.025* | |
| C41 | 0.64463 (18) | 0.76671 (14) | −0.01297 (13) | 0.0114 (4) | |
| C42 | 0.7191 (2) | 0.79247 (18) | 0.04993 (15) | 0.0239 (6) | |
| H42 | 0.7029 | 0.7809 | 0.1018 | 0.029* | |
| C43 | 0.8164 (3) | 0.8347 (2) | 0.03845 (17) | 0.0333 (7) | |
| H43 | 0.8658 | 0.8524 | 0.0823 | 0.040* | |
| C44 | 0.8419 (2) | 0.8513 (2) | −0.03605 (18) | 0.0315 (7) | |
| H44 | 0.9092 | 0.8794 | −0.0440 | 0.038* | |
| C45 | 0.7686 (3) | 0.8267 (3) | −0.09912 (18) | 0.0423 (9) | |
| H45 | 0.7856 | 0.8383 | −0.1508 | 0.051* | |
| C46 | 0.6708 (3) | 0.7853 (2) | −0.08813 (15) | 0.0316 (7) | |
| H46 | 0.6209 | 0.7694 | −0.1323 | 0.038* |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| I1 | 0.01029 (9) | 0.00674 (9) | 0.01363 (9) | 0.00312 (4) | −0.00089 (6) | −0.00191 (4) |
| Cu1 | 0.00902 (16) | 0.00693 (16) | 0.00820 (15) | −0.00076 (11) | 0.00197 (12) | −0.00022 (11) |
| S1 | 0.0096 (2) | 0.0109 (3) | 0.0175 (3) | −0.00149 (19) | 0.0049 (2) | −0.0069 (2) |
| P1 | 0.0074 (2) | 0.0068 (3) | 0.0076 (2) | −0.00001 (19) | 0.00262 (19) | −0.00070 (19) |
| P2 | 0.0097 (3) | 0.0057 (2) | 0.0078 (2) | 0.00084 (19) | 0.00168 (19) | 0.00046 (19) |
| N1 | 0.0105 (9) | 0.0122 (10) | 0.0262 (11) | −0.0006 (8) | 0.0025 (8) | −0.0094 (8) |
| N2 | 0.0100 (9) | 0.0092 (9) | 0.0174 (9) | 0.0010 (7) | 0.0012 (7) | −0.0073 (7) |
| C1 | 0.0121 (10) | 0.0117 (11) | 0.0096 (10) | 0.0003 (8) | 0.0036 (8) | −0.0022 (8) |
| C2 | 0.0172 (12) | 0.0122 (12) | 0.0315 (14) | 0.0030 (9) | 0.0091 (11) | −0.0075 (9) |
| C3 | 0.0314 (15) | 0.0299 (16) | 0.0441 (17) | 0.0096 (13) | 0.0083 (13) | 0.0068 (13) |
| C4 | 0.0112 (11) | 0.0149 (11) | 0.0185 (11) | −0.0007 (9) | 0.0012 (9) | −0.0063 (9) |
| C5 | 0.0105 (10) | 0.0172 (12) | 0.0208 (12) | −0.0002 (9) | 0.0011 (9) | −0.0089 (9) |
| C11 | 0.0117 (10) | 0.0088 (10) | 0.0112 (10) | 0.0041 (8) | 0.0052 (8) | 0.0014 (8) |
| C12 | 0.0144 (10) | 0.0090 (10) | 0.0146 (11) | 0.0020 (8) | 0.0051 (9) | −0.0004 (8) |
| C13 | 0.0214 (12) | 0.0149 (11) | 0.0124 (11) | 0.0049 (10) | 0.0030 (9) | −0.0003 (9) |
| C14 | 0.0248 (12) | 0.0197 (13) | 0.0164 (11) | 0.0054 (10) | 0.0115 (10) | 0.0085 (9) |
| C15 | 0.0170 (11) | 0.0186 (12) | 0.0260 (13) | −0.0024 (10) | 0.0088 (10) | 0.0076 (10) |
| C16 | 0.0106 (10) | 0.0166 (12) | 0.0181 (11) | −0.0015 (9) | 0.0036 (9) | 0.0029 (9) |
| C21 | 0.0088 (10) | 0.0093 (10) | 0.0151 (11) | −0.0017 (8) | 0.0033 (9) | 0.0016 (8) |
| C22 | 0.0156 (12) | 0.0219 (13) | 0.0185 (12) | 0.0035 (9) | 0.0002 (10) | −0.0044 (9) |
| C23 | 0.0166 (12) | 0.0239 (14) | 0.0297 (14) | 0.0046 (11) | −0.0059 (10) | 0.0002 (11) |
| C24 | 0.0120 (11) | 0.0191 (13) | 0.0425 (16) | 0.0048 (10) | 0.0023 (11) | 0.0007 (11) |
| C25 | 0.0191 (12) | 0.0202 (13) | 0.0358 (15) | 0.0057 (10) | 0.0105 (11) | −0.0064 (11) |
| C26 | 0.0175 (12) | 0.0161 (12) | 0.0200 (12) | 0.0017 (9) | 0.0042 (10) | −0.0048 (9) |
| C27 | 0.0112 (11) | 0.0082 (11) | 0.0101 (10) | 0.0002 (8) | 0.0017 (8) | −0.0020 (7) |
| C28 | 0.0111 (10) | 0.0075 (10) | 0.0085 (10) | 0.0002 (8) | 0.0016 (8) | −0.0017 (8) |
| C31 | 0.0175 (11) | 0.0123 (11) | 0.0088 (10) | 0.0062 (9) | 0.0039 (8) | 0.0011 (8) |
| C32 | 0.0274 (14) | 0.0171 (13) | 0.0456 (17) | 0.0059 (11) | 0.0025 (12) | 0.0054 (12) |
| C33 | 0.045 (2) | 0.0210 (15) | 0.064 (2) | 0.0149 (14) | 0.0092 (17) | 0.0142 (15) |
| C34 | 0.0319 (15) | 0.0399 (18) | 0.0315 (15) | 0.0273 (14) | 0.0049 (12) | 0.0107 (13) |
| C35 | 0.0188 (13) | 0.0409 (17) | 0.0238 (13) | 0.0122 (12) | 0.0018 (10) | −0.0045 (12) |
| C36 | 0.0179 (12) | 0.0221 (13) | 0.0218 (12) | 0.0057 (10) | 0.0042 (10) | −0.0024 (10) |
| C41 | 0.0149 (11) | 0.0054 (10) | 0.0140 (11) | −0.0001 (8) | 0.0027 (9) | 0.0020 (8) |
| C42 | 0.0278 (14) | 0.0278 (14) | 0.0154 (12) | −0.0125 (11) | 0.0003 (10) | 0.0057 (10) |
| C43 | 0.0305 (15) | 0.0388 (17) | 0.0285 (15) | −0.0211 (13) | −0.0067 (12) | 0.0108 (13) |
| C44 | 0.0245 (14) | 0.0330 (16) | 0.0371 (16) | −0.0154 (12) | 0.0042 (12) | 0.0131 (13) |
| C45 | 0.0434 (18) | 0.063 (2) | 0.0220 (14) | −0.0303 (17) | 0.0116 (13) | 0.0075 (14) |
| C46 | 0.0358 (16) | 0.0455 (18) | 0.0128 (12) | −0.0229 (14) | 0.0002 (11) | 0.0039 (12) |
Geometric parameters (Å, º)
| I1—Cu1 | 2.7412 (4) | C21—C26 | 1.394 (3) |
| Cu1—P1 | 2.2681 (6) | C21—C22 | 1.398 (4) |
| Cu1—P2 | 2.2813 (6) | C22—C23 | 1.399 (4) |
| Cu1—S1 | 2.3457 (6) | C22—H22 | 0.9500 |
| S1—C1 | 1.719 (2) | C23—C24 | 1.380 (4) |
| P1—C21 | 1.829 (2) | C23—H23 | 0.9500 |
| P1—C11 | 1.832 (2) | C24—C25 | 1.389 (4) |
| P1—C27 | 1.844 (2) | C24—H24 | 0.9500 |
| P2—C41 | 1.827 (2) | C25—C26 | 1.384 (4) |
| P2—C31 | 1.834 (2) | C25—H25 | 0.9500 |
| P2—C28 | 1.852 (2) | C26—H26 | 0.9500 |
| N1—C1 | 1.337 (3) | C27—C28i | 1.530 (3) |
| N1—C2 | 1.459 (3) | C27—H27A | 0.9900 |
| N1—H1 | 0.8800 | C27—H27B | 0.9900 |
| N2—C1 | 1.333 (3) | C28—C27i | 1.530 (3) |
| N2—C4 | 1.458 (3) | C28—H28A | 0.9900 |
| N2—H2 | 0.8800 | C28—H28B | 0.9900 |
| C2—C3 | 1.507 (4) | C31—C32 | 1.391 (4) |
| C2—H2A | 0.9900 | C31—C36 | 1.394 (4) |
| C2—H2B | 0.9900 | C32—C33 | 1.391 (4) |
| C3—H3A | 0.9800 | C32—H32 | 0.9500 |
| C3—H3B | 0.9800 | C33—C34 | 1.371 (5) |
| C3—H3C | 0.9800 | C33—H33 | 0.9500 |
| C4—C5 | 1.514 (3) | C34—C35 | 1.381 (5) |
| C4—H4A | 0.9900 | C34—H34 | 0.9500 |
| C4—H4B | 0.9900 | C35—C36 | 1.389 (4) |
| C5—H5A | 0.9800 | C35—H35 | 0.9500 |
| C5—H5B | 0.9800 | C36—H36 | 0.9500 |
| C5—H5C | 0.9800 | C41—C42 | 1.389 (3) |
| C11—C12 | 1.393 (3) | C41—C46 | 1.393 (3) |
| C11—C16 | 1.399 (3) | C42—C43 | 1.384 (4) |
| C12—C13 | 1.399 (3) | C42—H42 | 0.9500 |
| C12—H12 | 0.9500 | C43—C44 | 1.374 (4) |
| C13—C14 | 1.382 (4) | C43—H43 | 0.9500 |
| C13—H13 | 0.9500 | C44—C45 | 1.378 (5) |
| C14—C15 | 1.389 (4) | C44—H44 | 0.9500 |
| C14—H14 | 0.9500 | C45—C46 | 1.381 (4) |
| C15—C16 | 1.390 (3) | C45—H45 | 0.9500 |
| C15—H15 | 0.9500 | C46—H46 | 0.9500 |
| C16—H16 | 0.9500 | ||
| P1—Cu1—P2 | 113.37 (2) | C15—C16—H16 | 119.8 |
| P1—Cu1—S1 | 106.77 (2) | C11—C16—H16 | 119.8 |
| P2—Cu1—S1 | 114.30 (2) | C26—C21—C22 | 118.6 (2) |
| P1—Cu1—I1 | 104.637 (18) | C26—C21—P1 | 118.01 (18) |
| P2—Cu1—I1 | 106.669 (17) | C22—C21—P1 | 123.33 (18) |
| S1—Cu1—I1 | 110.709 (17) | C21—C22—C23 | 120.1 (2) |
| C1—S1—Cu1 | 109.37 (7) | C21—C22—H22 | 119.9 |
| C21—P1—C11 | 101.52 (10) | C23—C22—H22 | 119.9 |
| C21—P1—C27 | 100.74 (10) | C24—C23—C22 | 120.6 (2) |
| C11—P1—C27 | 102.98 (10) | C24—C23—H23 | 119.7 |
| C21—P1—Cu1 | 112.59 (7) | C22—C23—H23 | 119.7 |
| C11—P1—Cu1 | 116.01 (8) | C23—C24—C25 | 119.3 (2) |
| C27—P1—Cu1 | 120.36 (7) | C23—C24—H24 | 120.4 |
| C41—P2—C31 | 103.26 (10) | C25—C24—H24 | 120.4 |
| C41—P2—C28 | 101.76 (10) | C26—C25—C24 | 120.5 (2) |
| C31—P2—C28 | 102.47 (10) | C26—C25—H25 | 119.8 |
| C41—P2—Cu1 | 118.97 (7) | C24—C25—H25 | 119.8 |
| C31—P2—Cu1 | 117.28 (7) | C25—C26—C21 | 120.9 (2) |
| C28—P2—Cu1 | 110.82 (7) | C25—C26—H26 | 119.5 |
| C1—N1—C2 | 125.3 (2) | C21—C26—H26 | 119.5 |
| C1—N1—H1 | 117.4 | C28i—C27—P1 | 114.89 (15) |
| C2—N1—H1 | 117.4 | C28i—C27—H27A | 108.5 |
| C1—N2—C4 | 124.98 (19) | P1—C27—H27A | 108.5 |
| C1—N2—H2 | 117.5 | C28i—C27—H27B | 108.5 |
| C4—N2—H2 | 117.5 | P1—C27—H27B | 108.5 |
| N2—C1—N1 | 117.4 (2) | H27A—C27—H27B | 107.5 |
| N2—C1—S1 | 120.29 (17) | C27i—C28—P2 | 108.66 (14) |
| N1—C1—S1 | 122.29 (17) | C27i—C28—H28A | 110.0 |
| N1—C2—C3 | 112.4 (2) | P2—C28—H28A | 110.0 |
| N1—C2—H2A | 109.1 | C27i—C28—H28B | 110.0 |
| C3—C2—H2A | 109.1 | P2—C28—H28B | 110.0 |
| N1—C2—H2B | 109.1 | H28A—C28—H28B | 108.3 |
| C3—C2—H2B | 109.1 | C32—C31—C36 | 118.7 (2) |
| H2A—C2—H2B | 107.9 | C32—C31—P2 | 122.5 (2) |
| C2—C3—H3A | 109.5 | C36—C31—P2 | 118.77 (18) |
| C2—C3—H3B | 109.5 | C33—C32—C31 | 119.8 (3) |
| H3A—C3—H3B | 109.5 | C33—C32—H32 | 120.1 |
| C2—C3—H3C | 109.5 | C31—C32—H32 | 120.1 |
| H3A—C3—H3C | 109.5 | C34—C33—C32 | 121.2 (3) |
| H3B—C3—H3C | 109.5 | C34—C33—H33 | 119.4 |
| N2—C4—C5 | 109.97 (19) | C32—C33—H33 | 119.4 |
| N2—C4—H4A | 109.7 | C33—C34—C35 | 119.6 (3) |
| C5—C4—H4A | 109.7 | C33—C34—H34 | 120.2 |
| N2—C4—H4B | 109.7 | C35—C34—H34 | 120.2 |
| C5—C4—H4B | 109.7 | C34—C35—C36 | 120.0 (3) |
| H4A—C4—H4B | 108.2 | C34—C35—H35 | 120.0 |
| C4—C5—H5A | 109.5 | C36—C35—H35 | 120.0 |
| C4—C5—H5B | 109.5 | C35—C36—C31 | 120.8 (3) |
| H5A—C5—H5B | 109.5 | C35—C36—H36 | 119.6 |
| C4—C5—H5C | 109.5 | C31—C36—H36 | 119.6 |
| H5A—C5—H5C | 109.5 | C42—C41—C46 | 117.8 (2) |
| H5B—C5—H5C | 109.5 | C42—C41—P2 | 118.50 (18) |
| C12—C11—C16 | 119.4 (2) | C46—C41—P2 | 123.68 (18) |
| C12—C11—P1 | 119.57 (17) | C43—C42—C41 | 121.2 (2) |
| C16—C11—P1 | 121.03 (17) | C43—C42—H42 | 119.4 |
| C11—C12—C13 | 119.8 (2) | C41—C42—H42 | 119.4 |
| C11—C12—H12 | 120.1 | C44—C43—C42 | 120.3 (3) |
| C13—C12—H12 | 120.1 | C44—C43—H43 | 119.8 |
| C14—C13—C12 | 120.5 (2) | C42—C43—H43 | 119.8 |
| C14—C13—H13 | 119.7 | C43—C44—C45 | 119.2 (3) |
| C12—C13—H13 | 119.7 | C43—C44—H44 | 120.4 |
| C13—C14—C15 | 119.9 (2) | C45—C44—H44 | 120.4 |
| C13—C14—H14 | 120.0 | C44—C45—C46 | 120.8 (3) |
| C15—C14—H14 | 120.0 | C44—C45—H45 | 119.6 |
| C14—C15—C16 | 120.0 (2) | C46—C45—H45 | 119.6 |
| C14—C15—H15 | 120.0 | C45—C46—C41 | 120.7 (3) |
| C16—C15—H15 | 120.0 | C45—C46—H46 | 119.6 |
| C15—C16—C11 | 120.3 (2) | C41—C46—H46 | 119.6 |
| C4—N2—C1—N1 | −7.2 (3) | P1—C21—C26—C25 | 177.6 (2) |
| C4—N2—C1—S1 | 173.37 (18) | C21—P1—C27—C28i | −165.70 (16) |
| C2—N1—C1—N2 | 174.1 (2) | C11—P1—C27—C28i | −61.09 (18) |
| C2—N1—C1—S1 | −6.5 (3) | Cu1—P1—C27—C28i | 69.95 (17) |
| Cu1—S1—C1—N2 | −33.6 (2) | C41—P2—C28—C27i | 66.78 (16) |
| Cu1—S1—C1—N1 | 146.99 (17) | C31—P2—C28—C27i | 173.40 (15) |
| C1—N1—C2—C3 | −90.6 (3) | Cu1—P2—C28—C27i | −60.70 (15) |
| C1—N2—C4—C5 | −176.4 (2) | C41—P2—C31—C32 | −20.2 (2) |
| C21—P1—C11—C12 | −123.32 (18) | C28—P2—C31—C32 | −125.7 (2) |
| C27—P1—C11—C12 | 132.67 (18) | Cu1—P2—C31—C32 | 112.8 (2) |
| Cu1—P1—C11—C12 | −0.9 (2) | C41—P2—C31—C36 | 161.93 (18) |
| C21—P1—C11—C16 | 54.8 (2) | C28—P2—C31—C36 | 56.5 (2) |
| C27—P1—C11—C16 | −49.2 (2) | Cu1—P2—C31—C36 | −65.11 (19) |
| Cu1—P1—C11—C16 | 177.20 (16) | C36—C31—C32—C33 | −1.1 (4) |
| C16—C11—C12—C13 | −0.7 (3) | P2—C31—C32—C33 | −178.9 (3) |
| P1—C11—C12—C13 | 177.45 (17) | C31—C32—C33—C34 | −0.4 (5) |
| C11—C12—C13—C14 | −0.1 (3) | C32—C33—C34—C35 | 0.8 (5) |
| C12—C13—C14—C15 | 0.8 (4) | C33—C34—C35—C36 | 0.3 (4) |
| C13—C14—C15—C16 | −0.7 (4) | C34—C35—C36—C31 | −1.9 (4) |
| C14—C15—C16—C11 | −0.1 (4) | C32—C31—C36—C35 | 2.2 (4) |
| C12—C11—C16—C15 | 0.8 (3) | P2—C31—C36—C35 | −179.84 (19) |
| P1—C11—C16—C15 | −177.35 (18) | C31—P2—C41—C42 | 114.9 (2) |
| C11—P1—C21—C26 | 56.7 (2) | C28—P2—C41—C42 | −139.1 (2) |
| C27—P1—C21—C26 | 162.44 (19) | Cu1—P2—C41—C42 | −17.1 (2) |
| Cu1—P1—C21—C26 | −68.06 (19) | C31—P2—C41—C46 | −67.9 (2) |
| C11—P1—C21—C22 | −127.0 (2) | C28—P2—C41—C46 | 38.1 (3) |
| C27—P1—C21—C22 | −21.2 (2) | Cu1—P2—C41—C46 | 160.1 (2) |
| Cu1—P1—C21—C22 | 108.27 (19) | C46—C41—C42—C43 | −0.4 (4) |
| C26—C21—C22—C23 | 0.3 (4) | P2—C41—C42—C43 | 177.0 (2) |
| P1—C21—C22—C23 | −176.0 (2) | C41—C42—C43—C44 | −0.7 (5) |
| C21—C22—C23—C24 | −1.5 (4) | C42—C43—C44—C45 | 1.1 (5) |
| C22—C23—C24—C25 | 1.2 (4) | C43—C44—C45—C46 | −0.4 (6) |
| C23—C24—C25—C26 | 0.2 (4) | C44—C45—C46—C41 | −0.7 (6) |
| C24—C25—C26—C21 | −1.3 (4) | C42—C41—C46—C45 | 1.1 (5) |
| C22—C21—C26—C25 | 1.1 (4) | P2—C41—C46—C45 | −176.2 (3) |
Symmetry code: (i) −x+1, −y+1, −z.
Hydrogen-bond geometry (Å, º)
| D—H···A | D—H | H···A | D···A | D—H···A |
| N1—H1···I1ii | 0.88 | 2.80 | 3.622 (2) | 156 |
| N2—H2···I1 | 0.88 | 2.70 | 3.5517 (19) | 162 |
Symmetry code: (ii) −x+3/2, y+1/2, −z+1/2.
Footnotes
Supporting information for this paper is available from the IUCr electronic archives (Reference: TK5371).
References
- Bruker (2013). APEX2, SAINT and SADABS. Bruker AXS Inc., Madison, Wisconsin, USA.
- Corey, E. J., Wess, G., Xiang, Y. B. & Singh, A. K. (1987). J. Am. Chem. Soc. 109, 4717–4718.
- Dennehy, M., Quinzani, O. V., Mandolesi, S. D. & Burrow, R. A. (2011). J. Mol. Struct. 998, 119–125.
- Dennehy, M., Tellería, G. P., Quinzani, O. V., Echeverría, G. A., Piro, O. E. & Castellano, E. E. (2009). Inorg. Chim. Acta, 362, 2900–2908.
- Dias, H. V. R., Batdorf, K. H., Fianchini, M., Diyabalanage, H. V. K., Carnahan, S., Mulcahy, R., Rabiee, A., Nelson, K. & van Waasbergen, L. G. (2006). J. Inorg. Biochem. 100, 158–160. [DOI] [PubMed]
- Hübschle, C. B., Sheldrick, G. M. & Dittrich, B. (2011). J. Appl. Cryst. 44, 1281–1284. [DOI] [PMC free article] [PubMed]
- Macrae, C. F., Bruno, I. J., Chisholm, J. A., Edgington, P. R., McCabe, P., Pidcock, E., Rodriguez-Monge, L., Taylor, R., van de Streek, J. & Wood, P. A. (2008). J. Appl. Cryst. 41, 466–470.
- Oshio, H., Watanabe, T., Ohto, A., Ito, T. & Masuda, H. (1996). Inorg. Chem. 35, 472–479. [DOI] [PubMed]
- Seward, C., Chan, J., Song, D. & Wang, S. (2003). Inorg. Chem. 42, 1112–1120. [DOI] [PubMed]
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
- Sheldrick, G. M. (2015). Acta Cryst. C71, 3–8.
- Westrip, S. P. (2010). J. Appl. Cryst. 43, 920–925.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablock(s) I. DOI: 10.1107/S2056989015014176/tk5371sup1.cif
Structure factors: contains datablock(s) I. DOI: 10.1107/S2056989015014176/tk5371Isup2.hkl
. DOI: 10.1107/S2056989015014176/tk5371fig1.tif
The structure of title complex with displacement ellipsoids drawn at the 50% propbability level. All H atoms are omitted for clarity.
. DOI: 10.1107/S2056989015014176/tk5371fig2.tif
Part of the crystal structure showing intra/inter-molecular N—H⋯I hydrogen bonds forming a layers as dashed lines.
CCDC reference: 1415379
Additional supporting information: crystallographic information; 3D view; checkCIF report


